Abstract
Due to the impact of various environmental factors, in conjunction with the ecological adaptability and biochemical diversity of Chinese medicinal materials, Codonopsis Radix from Shanxi Province has developed into two distinct commercial specifications, namely Tai Codonopsis Radix (TCP) and Lu Codonopsis Radix (LCP). The present study examined the relationships between TCP and LCP from Shanxi Province regarding their environment, secondary metabolites, and Traditional Chinese Medicinal plant authenticity using metagenomic, metabolomic, and comprehensive quality assessment methods. The findings revealed that the two types of CPs met the quality standards. With respect to the growing environment, TCP and LCP grow in the same longitudinal extent, with a maximum longitudinal span of only 53ʹ. Both regions presented similar natural environments conducive to the growth of CP. Additionally, the Lobetyolin content in TCP was greater than that in LCP. The soil microbial abundance in the TCP group was considerably significantly greater than that in the LCP group, and a new genus, Ascobolus, was found in TCP. Additionally, comparison of the metallic element data of the two groups soils revealed that the contents of Ti, Mg, Cr (P < 0.01) in the TCP group were significantly greater than those in the LCP group. Through correlation analysis, the ternary network of microbial-metal-metabolite interactions was constructed. The rhizosphere fungus Ascobolus was found to enhance the bioavailability of Zn via mineralization processes, thereby underscoring the significance of rhizosphere microecology in promoting plant growth. This study offers novel insights and approaches for the cultivation and processing of Codonopsis Radix.
Similar content being viewed by others
Introduction
Traditional herbal medicines (THMs) are derived from natural sources. Prior to the introduction of synthetic drugs in Western medicine, Chinese people frequently relied on natural remedies known as THMs to address illnesses, following Traditional Chinese medical (TCM) theories and principles. The assessment of THM quality and clinical effectiveness is influenced by various factors, including the cultivation environment, different processing methods and techniques employed in distinct planting areas or regions. These factors are crucial in determining the extrinsic characteristics of THMs as well as the content of intrinsic bioactive compounds or secondary metabolites. The aforementioned factors can also be encapsulated by an authentic TCM concept known as Daodi1.
Daodi medicines2,3,4 are synonymous with genuine THMs (GTHMs), which refer to those that have been selected and recognized through long-term application in TCM clinical practice. Compared with similar THMs grown elsewhere, it is cultivated in specific regions; has superior properties, quality, and efficacy; has consistent quality; and is more popular.
China possesses an extensive land area, and its distinctive natural and geographical conditions have fostered a wide range of GTHM resources. Shanxi Province is situated in an area that lies between the North China Plain and the Inner Mongolia Highlands and is located across the Yellow River system. This advantageous geographical position provides perfect conditions for the growth of GTHMs, including Astragalus membranaceus (Huangqi in Chinese), Codonopsis Radix (Dangshen in Chineses, CP), Forsythiae (Lianqiao in Chinese), Polygala (Yuanzhi in Chinese), Radix bupleuri (Chaihu in Chinese), Scutellaria baicalensis (Huangqin in Chinese), Jujube seed (Suanzaoren in Chinese), Kushen (Kushen in Chinese), Hawthorn (Shanzha in Chinese), and Peach kernel (Taoren in Chinese). CP can be divided into many categories5,6,7, such as Fang CP (Fang Dangshen in Chinese), Wen Yuan CP (Wenyuan Dangshen in Chinese), and Tiao CP (Tiao Dangshen in Chinese). According to its origin in Shanxi Province, geographical conditions and traits can be divided into two major Commodity Specifications: Lu CP (Lu dangshen in Chinese, LCP) and Tai CP (Tai dangshen in Chinese, TCP). Both of these categories are derived from Codonopsis pilosula (Franch.) Nannf., which belongs to the Campanulaceae family. LCP is cultivated mostly in Changzhi city in the Taihang Mountains area, with the most famous LCP found in Pingshun County. There are two main basins inside the county: the Zhuozhang River and the Wei River. The presence of mountains, hills, and river valleys in the area provides suitable habitat for the growth of the LCP. TCP is produced mainly in Xinzhou in the Wutai Mountain area of Wutai County. The TCP growing area is characterized by a network of intersecting gullies, and the TCP that grows in high-altitude cold mountain locations above 1,500 m is of exceptional quality. Currently, there is a lack of research on the evaluation of real regions via modern analytical techniques and methods.
The large-scale cultivation of LCP has been extensively practised in Shanxi, while TCP predominantly exists as a wild species. The artificial cultivation technology for TCP has not yet been fully developed, and the availability of wild resources for TCP is limited. Admittedly, there have been few comparative studies on the authentic evaluation of the TCP and LCP. The present study investigated the relationships of two Commodity Specifications CPs from Shanxi Province with the environment8, secondary metabolites, and THM authenticity via metagenomic, metabolomic9, and comprehensive quality assessment methods. These findings reveal the details of THMs and serve as valuable references for artificial cultivation, site selection, and processing in the Shanxi Province, which specializes in two commodity specifications of CPs. (The specific research approach is shown in the Fig. 14).
Results
TCP and LCP geographical environment analysis
The TCP and LCP acquisition sites are specified in section "Materials", and a comprehensive analysis of the growth environments of the two commodity specifications CP species in these regions was performed. The geographical environment map indicated that the transition from blue to orange depicted the increasing distributions of precipitation, temperature, and sunshine time. The comparison is as follows.
Wutai County is situated in the northeastern part of Shanxi Province between 112°57′41″ and 113°50′56″ east longitude and 38°28′ and 39°4′49″ north latitude. The precipitation in Wutai County shows a spatial distribution that is characterized by unevenness, with a pattern that extends from low in the south to high in the north. The maximum recorded precipitation is 69.5 mm. Precipitation is mostly concentrated in the Wutai Mountain region, particularly in the northern part of Wutai County, where the altitude is relatively high. The annual rainfall in this area can reach approximately 500 mm. The county experiences moderate temperatures during the summer and frigid temperatures during the winter. The primary factor contributing to the temperature variation in the county is the decrease in altitude. The temperature inversely correlates with altitude, exhibiting latitudinal and altitudinal distributions from high in the south to low in the north. The low-temperature area is mainly concentrated in the Wutai Mountain area, which is located at relatively high altitudes in the northern part of the county. The minimum temperature can decrease to as low as − 3.14 °C. The distribution of sunshine duration in Wutai County exhibited spatial disparities, varying from shorter in the south to longer in the north. The regions characterized by extended periods of sunlight were found primarily in the elevated terrain to the north of the county. The annual total duration of sunshine was 2400–2700 h. TCP growth may be facilitated by the natural conditions of Mount Wutai in Wutai County, such as its high altitude of 3061 m, which results in relatively low temperatures and lengthy periods of sunshine (Fig. 1A–C).
Precipitation distribution map of Wutai County, Shanxi Province (A); Ttemperature distribution map of Wutai County, Shanxi Province (B); Sunshine distribution map of Wutai County, Shanxi Province (C); Precipitation distribution map of Pingshun County, Shanxi Province (D); Temperature distribution map of Pingshun County, Shanxi Province (E); Sunshine duration distribution map of Pingshun County, Shanxi Province (F).
Pingshun County is located in southeastern Shanxi Province and spans from 113°11′45″ to 113°44′04″ east longitude and from 35°56′37″ to 36°27′44″ north latitude. Pingshun County experiences uneven precipitation distributions, with the rainfall density increasing from southwest to northeast. The northeastern mountainous area receives the highest amount of precipitation, reaching 65.38 mm, while the annual precipitation in the county is 584 mm. The temperature in the county is mild in summer and cold in winter, and the temperature gradient increases from south to north. The region with the lowest temperature is located primarily in the southern mountainous area of the county, where the minimum temperature is 7.66 °C. The distribution of sunshine duration varies from east to west, with longer durations predominantly found in the eastern portion of the county. The maximum duration recorded was 6.22 h, and the yearly average duration of sunshine was 2,517.8 h (Fig. 1D–F).
The average soil pH value was 7.34 ± 0.23 in Wutai County and 7.30 ± 0.17 in Pingshun County, with no significant difference observed between the two counties. In terms of soil organic matter content, Wutai County had a significantly higher level (35.25 ± 4.20) compared to Pingshun County (26.65 ± 2.14), and this difference was statistically highly significant (P < 0.0001). Regarding climatic factors, the average annual temperature (Temp) in Pingshun County (10.17 ± 0.35) was significantly higher than that in Wutai County (7.25 ± 0.44), with a statistical significance of P < 0.01. Additionally, the diurnal temperature range (DIF) was 11.09 ± 0.31 in Wutai County and 10.87 ± 0.3 in Pingshun County. For ultraviolet radiation intensity (UV-B), the respective values were 185.96 ± 2.23 in Wutai County and 184.69 ± 3.95 in Pingshun County.
In conclusion, both Wutai and Pingshun Counties share the same longitude, with a maximum longitude difference of only 53’. TCP thrives in regions characterized by high elevation, low temperatures, approximately 500 mm of rainfall, and extended periods of sunlight. The natural environments of the two locations are similar and are conducive to CP cultivation. However, there is a considerable difference in height within the Wutai Mountain area of Wutai County. Furthermore, the soil organic matter content in Wutai County is markedly higher than that in Pingshun County (Table 1).
TCP and LCP quality complies with regulations
An analysis of the properties revealed that the circular patterns beneath the root head of the LCP decreased in thickness as they moved downwards, but the circular patterns beneath the root head of the TCP were denser. Microscopic analysis revealed that the LCP powder was a light yellow colour with many inulins, many fragmented laticiferous vessel containing light yellow particles, and reticular ducts that were easily observable. The similarity of the TCPs suggests that both varieties conform to the specifications outlined in the “Chinese Pharmacopoeia” (2020 edition) (Fig. 2A–B). Both CPs complied with the requirements stated in the Chinese Pharmacopoeia (2020 edition).
Characteristics and microscopic identification map of TCPs and characteristics (A); Microscopic identification map of LCPs (B); Lobetyolin content map of control substance, TCP and LCP (C). TCP: Tai Codonopsis Radix; LCP: Lu Codonopsis Radix.
The Lobetyolin content in TCP was determined to be 1.68% ± 0.03, whereas that in LCP was found to be 1.17% ± 0.05. These results indicate a higher concentration of codontoside in TCP than in LCP (Fig. 2C).
The secondary metabolites of LCP and TCP are similar
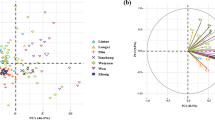
The Base peak chromatogram (BPC) of the QC (Quality Control) samples were compared via spectral overlap analysis.The results indicated that the response intensity and retention time of each chromatographic peak were essentially the same. PCA (principal component analysis) was performed on each sample, revealing a clear separation between LCP and TCP.
The PCA model parameters of TCP compared to LCP, which were obtained after sevenfold cross-validation, yielded R2X = 0.998. This finding indicates that the model is reliable and that there are significant differences between the groups (Fig. 3A,B).
TCP and LCP positive ion mode PCA scoring figure, PCA score plots of TCP and LCP in the negative ion mode of the implant platform (A); TCP and LCP positive ion mode OPLS-DA scoring figure, OPLS-DA score plots of TCP and LCP in the negative ion mode of the implant platform (B). Green: TCP, Tai Codonopsis Radix; Red: LCP, Lu Codonopsis Radix.
Simultaneously, OPLS-DA (Orthogonal Partial Least Squares Discriminant Analysis) was employed to further validate that the degree of dissociation between TCP and LCP was remarkable and that the experimental results were reliable. By comparing the secondary spectra of the database, the components with relatively high contents of TCP and LCP were identified. The peak labelling results for different species are as follows. The BPC diagram of TCP shows 17 chromatographic peaks (14 in the positive ion mode and 3 in the negative ion mode), and the BPC diagram of LCP shows 15 chromatographic peaks (11 in the positive ion mode and 4 in the negative ion mode) (Fig. 4A–B).
TCP active ingredient BPC standard peak figure (A,B); Material map of differences in secondary metabolites between TCP and LCP under positive and negative ions (C). TCP: Tai Codonopsis Radix; LCP: Lu Codonopsis Radix.(In this sampling and analysis process, the sample size was determined to be n = 6).
Through a comparative analysis of the differential substances found in TCP and LCP, using a screening condition of a response intensity greater than 3 times, one unknown component was found in TCP, and two unknown components were determined in LCP. Isovalerylcarnitine, 8-methoxyatractylenolide I, and atractylenolide I were identified as shared compounds found in TCP. Bufotenine, 3-indolyllactic acid, and sinapyl alcohols are the common compounds in wild varieties of TCP and LCP. Anileridine, 9-octadecenoic acid, 5,8,11-trihydroxy-, and hexyl 6-O-pentopyranosylhexopyranoside were found in both cultivated TCP and LCP. Lobetyolin and anileridine are common compounds found in all the two categories of CP. Both lobetyolin and hexyl 6-O-pentopyranosylhexopyranoside were detected in both positive and negative ion modes (Fig. 4C).
To elucidate the molecular basis underpinning the metabolic differences between TCP and LCP, we performed KEGG pathway enrichment analysis on the identified differentially metabolites. The results demonstrated that differential metabolites between the two groups were significantly enriched in several metabolic pathways, including phenylalanine metabolism, ubiquinone and other terpenoid-quinone biosynthesis, pyruvate metabolism, and glutathione metabolism (P < 0.05). Notably, phenylalanine metabolism was significantly enriched exclusively under positive ion mode (POS, − log10(p) = 3.2), whereas pyruvate metabolism and glutathione metabolism were significantly enriched under negative ion mode (NEG, − log10(p) = 2.9). These suggest that TCP may drive secondary metabolic differentiation through the synergistic regulation of “aromatic skeleton synthesis-carbon chain extension-redox homeostasis”. The coordinated alterations in these pathways collectively form the metabolic basis for the significantly higher accumulation of lobetyolin in TCP compared to LCP, which are highly consistent with the distinct separation of the two groups observed in the OPLS-DA analysis (Fig. 5).
KEGG enrichment pathway map of differential metabolites in the positive ion mode (A); KEGG enrichment pathway map of differential metabolites in the negative ion mode (B).
The abundance of microbial flora in TCP bulk soil was high
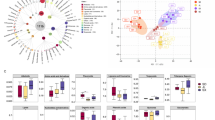
The locations for the collection of TCP and LCP soils are described in section "Materials" Bulk soil was investigated through metagenomics. The results revealed that the community in TCP (YF Group, TCP bulk soil group) presented more significant changes in diversity and species composition than did that in LCP (PF Group, LCP bulk soil group) (Fig. 6A).
Soil abundance Wayne figure (A); Bulk soil bacteria, archaea, and fungi PCA scoring figure (B); alpha diversity analysis of non-root soil, Blue: YF, Purple: PF (C); LEfSe difference analysis of bulk soil microorganisms, Purple: PF, Grey: YF (D). YF: Bulk system soil of Tai Codonopsis Radix; PF: Bulk system soil of Lu Codonopsis Radix. (In this sampling and analysis process, the sample size was determined to be n = 6).
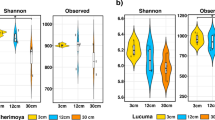
The analysis of alpha diversity in the soil samples from the two regions revealed similar distribution patterns but varying richness levels. Compared with the LCP samples, the TCP samples presented greater diversity (Fig. 6B).
Beta diversity analysis was employed to examine alterations in species composition across different time and spatial scales. PCA was conducted on bacteria, fungi, and archaea in each sample at the sublevel. The PCoA (Principal co-ordinates analysis) map clearly shows distinct separation of the YF and PF groups in the bacterial community, indicating significant variations in the soil microbial composition across locations. In the PCoA diagram, there is a noticeable tendency for the YF and PF groups to separate in terms of distribution. The proximity between each group is minimal, and the disparity between the groups is both slight and noteworthy (Fig. 6C).
In this study, LEfSe (LDA Effect Size analysis) (LDA > 3.5, P < 0.05) difference analysis of the microbial community was used to analyse the species with significant differences in the relative abundance of bulk soil microbial colonies in different regions. LEfSe revealed 101 significantly different populations, including 52 in the YF group and 22 in the PF group (Fig. 6D).
To investigate the effects of soil microbial diversity on CP growth in various production regions, the high-throughput sequencing results of bacteria, fungi, and archaea were annotated to 2663, 201, and 171 genera, respectively. The top 10 genera with clear annotation information were defined as the dominant genera.
The abundances of the bacterial genera Branchiibius, Candidatus Rokubacteria, Chloroflexi, Gemmatimonadetes, and Verrucomicrobia in the YF group significantly increased, and the abundances of Betaproteobacteria, Sphingosinicella, and Verrucomicrobia in the PF group significantly increased. The YF group presented a considerable increase in the abundance of the fungal genera Rhizophagus, Aspergillus, and Fusarium. Similarly, the PF group presented significant increases in the abundances of Rhizophagus, Fusarium, and Diversispora. The PF group presented an increase in the abundance of Thermoplasmata and Nitrosopumilales, whereas the abundance of the remaining genera decreased (Fig. 7A).
Difference map of the microbial abundances of bacteria, archaea and fungi in bulk soil (A); EggNOG annotation diagram (B). YF: Bulk system soil of Tai Codonopsis Radix; PF: Bulk system soil of Lu Codonopsis Radix. (In this sampling and analysis process, the sample size was determined to be n = 6).
Moreover, the functional metabolism of microorganisms in non-root system soil was analysed via eggNOG (evolutionary genealogy of genes: Non-supervised Orthologous Groups) annotation. Analysis of eggNOG functional annotations revealed that functional categories Q (biosynthesis, transport, and catabolism of secondary metabolites), G (carbohydrate transport and metabolism), and E (amino acid transport and metabolism) were significantly enriched in the YF group. The eggNOG functional profile indicates that the YF group displays distinct metabolic features relative to the PF group, primarily through the upregulation of category Q and coordinated activation of categories C, E, G, and I, which are linked to carbon, energy, and reducing power supply. Furthermore, KEGG functional annotation analysis (with the horizontal axis denoting EC numbers and the vertical axis representing normalized average relative abundance, where a taller bar indicates a higher abundance of the corresponding enzyme in each group) showed that EC 2.3.1.9 (acetyl-CoA acetyltransferase, involved in fatty acid biosynthesis pathways) exhibited increased relative abundance in the YF group, which facilitates the supply of carbon chain extension substrates for polyacetylene synthesis. Additionally, EC 1.2.7.11 (pyruvate:ferredoxin/flavodoxin oxidoreductase, involved in pyruvate metabolism) was upregulated, further supporting this metabolic enhancement. Moreover, the upregulation of EC 1.1.1.100 (malate dehydrogenase, involved in TCA/redox pathways) and EC 7.1.1.1 (V-type ATPase, involved in proton pump pathways) ensured an adequate energy supply for the alkyne bond reduction step and ABC transporter activity. Finally, the increased abundance of EC 2.1.1.13 (COMT, involved in phenylpropanoid-polyacetylene modification pathways) resulted in elevated methylation levels of lobetyolin side chains, leading to reduced polarity and enhanced storage capacity. Collectively, these changes significantly promoted the synthesis and accumulation of lobetyolin in the YF group compared to the PF group (Fig. 7B).
Finally, the degree of species variation in bulk system soils may differ across different regions. Among all the groups, the YF group presented the greatest number of different colonies, suggesting that the soil microbial diversity of TCP was both abundant and distinctive. The expression of functional genes in soil bacteria from various places exhibited similar patterns, although the relative abundance of individual genes varied.
The TCP bulk soil dominant species comprised Branchiibius, Candidatus Rokubacteria, Chloroflexi, Gemmatimonadetes, and Verrucomicrobia. In contrast, the LCP bulk soil community was predominantly composed of Betaproteobacteria, Sphingosinicella, and Verrucomicrobia. The presence of Verrucomicrobia in both groups suggests that variations in its abundance may be influenced by environmental factors.
The abundance of microbial flora in TCP rooted soil was high
The analysis of soil microorganisms in the bulk system soil revealed that the community of TCP (YG group, TCP rooted soil group) presented more pronounced changes in community diversity and species composition than did the communities of LCP (PG group, LCP rooted soil group).
Alpha diversity analysis indicated that the bacterial populations in the soil samples from the root system soils in the two regions presented similar distribution patterns but varied in terms of abundance. However, there were distinct variations in the distribution and abundance of bacterial populations in the soil across different regions. The diversity of the soil microflora in TCP (YG group) was greater than that in the LCP soil (PG group). Beta diversity analysis revealed that the principal component analysis of bacteria, fungi and archaea in each sample was performed at the genus level, indicating that the YG and PG groups were obviously separated on the PCoA diagram in terms of the community distributions of bacteria, fungi and archaea, indicating that there were significant differences in the structure of rhizosphere soil microbial colonies in different regions. LEfSe (LDA > 3.5, P < 0.05) was employed to analyse the species that exhibited significant differences in the relative abundance of root soil microbial colonies across several locations. LEfSe analysis revealed 171 distinct bacterial populations at the genus level, including 81 in the YG group and 37 in the PG group (Fig. 8A–C).
Root soil alpha diversity analysis diagram (A); PCA score plot of bacteria, archaea and fungi in root soil, Blue: YG, Purple: PG (B); Root soil microbial LEfSe difference analysis diagram (C). YG: Root system soil of Tai Codonopsis Radix; PG: Root system soil of Lu Codonopsis Radix.(In this sampling and analysis process, the sample size was determined to be n = 6).
In the analysis of nonroot soil microorganisms described in section “The abundance of microbial flora in TCP bulk soil was high”, a new strain of Ascobolus was identified among the fungal species in the YG group (Fig. 9A).
Difference map of the microbial abundances of bacteria, archaea and fungi in root soil (A); EggNOG annotation diagram (B). YG: Root system soil of Tai Codonopsis Radix; PG: Root system soil of Lu Codonopsis Radix. (In this sampling and analysis process, the sample size was determined to be n = 6) .
Analysis of eggNOG functional annotations revealed that functional category Q was significantly upregulated in the YG group, whereas category G showed relatively higher expression in the PG group. Functional categories E, C, and I also exhibited intergroup variations. The eggNOG functional profile indicates that the YG group redirects carbon flux from primary metabolism toward polyacetylene-based secondary metabolism by upregulating functional classes Q, E, C, and I, thereby forming a distinct metabolic pattern compared to the PG group. Furthermore, KEGG functional annotation analysis revealed that the enzyme activities corresponding to EC 1.2.7.11, EC 2.3.1.9, EC 1.1.1.100, and EC 2.1.1.13 were markedly elevated in the YG group relative to the PG group. This enhancement suggests increased availability of the C2 unit derived from pyruvate-acetyl-CoA, which supports the elongation of polyacetylene side chains. Concurrently, the capacity to maintain a reductive environment conducive to alkyne bond stability via NADPH was also improved. Through modifications to the aromatic skeleton and significant upregulation of key enzymatic activities, the biosynthesis and accumulation of lobetyolin were effectively promoted, resulting in distinct metabolic characteristics in the YG group (Fig. 9B).
Ultimately, the degree of variation in root soil composition among different places varies. Among all the groups, the YG group presented the greatest variation in the number of colonies, suggesting that the soil microbial richness of TCP was high and possessed distinct characteristics. The expression of functional genes in soil bacteria from various places exhibited similar overall patterns, although the relative abundance of specific genes varied.
Meanwhile, a novel fungus genus, Ascobolus, was identified in the TCP-rooted soil. The TCMs quantitative analysis of components indicated that the elevated Lobetyolin in TCP may be associated with this fungal species.
Ecological function analysis
The top 10 genes were identified as the dominant species by analysing the nitrogen cycling of bulk and root system soils at the genus level, for example. The abundances of different species increased or decreased to different degrees during nitrogen cycling. Nitrospira and Bradyrhizobium are likely the primary genes involved. All of them exhibited an increasing pattern during nitrogen cycling, as depicted below (Fig. 10A–B).
Functional map of metabolic genes in bulk soil and nitrogen cycling ecosystems (A); Functional map of soil-nitrogen cycling ecosystem metabolic genes in roots (B) . YG: Root system soil of Tai Codonopsis Radix; PG: Root system soil of Lu Codonopsis Radix; YF: Bulk system soil of Tai Codonopsis Radix; PF: Bulk system soil of Lu Codonopsis Radix. (In this sampling and analysis process, the sample size was determined to be n = 6).
High contents of TCP roil metallic elements
The findings of the examination of metallic elements in the soil are presented in Table 2. A comparison of the metallic element data of the two groups of bulk system soils revealed that the contents of Ba, Sr (P < 0.01) in the YF group were significantly greater than those in the PF group. The contents of Cu, Na, Mn, V, Cr (P < 0.01) and Zn (P < 0.0001) in the PF group significantly increased. The contents of the other elements in each group were identical. The content of metallic elements in Group PF was the highest, and the types were the most abundant.
Analysis of the metallic element data from the two groups of root system soils clearly revealed that the YG group presented significantly higher concentrations of Ni, Ba, Sr (P < 0.0001) compared with the PG group. The PG group presented increased levels of V (P < 0.001) Fe, Cu, Mn,Cr (P < 0.0001) and K, Ti (P < 0.01) whereas the concentrations of the remaining elements in the other groups were similar. The YG group presented the highest concentrations of metallic elements and the most diverse range of metallic elements. Such differences may be the result of the combined influence of climatic factors and the dominant bacterial flora in the soil.
Correlation analysis
Through the study of the correlation of soil bacteria and metabolic material, in the top 20 bacterial groups, Bradyrhizobiaceae, Bradyrhizobium, Hyphomicrobiaceae, Rhodoplanes, Gaiellaceae, Gaiella, Streptomycetaceae, Streptomyces, Phyllobacteriaceae, Mesorhizobium and Bufotenine, and hexyl 6-O-pentopyranosylhexopyranoside components were positively correlated and negatively correlated with 1-isopropylbenzimidazole-2-sulfonic acid, 8-benzylcanadine, and 1-(2-chlorobenzyl)pyrrole-2-carboxamide in CP. Sphingomonadaceae, Sphingosinicella, Solirubrobacteraceae, Solirubrobacter, Geodermatophilaceae, Blastococcus, Gemmatimonadaceae, Gemmatirosa, Sinobacteraceae, Steroidobacter, Sphingomonadaceae, Sphingomonas, Xanthobacteraceae, Pseudolabrys and 1-isopropylbenzimidazole-2-sulfonic acid, 8-benzylcanadine, and 1-(2-chlorobenzyl) pyrrole-2-carboxamide components were positively correlated; however, with the CPs Bufotenine peak, hexyl 6-O-pentopyranosylhexopyranoside was negatively correlated (Fig. 11A-B).
Correlation analysis of soil microorganisms and pharmacodynamic components in TCP and LCP (A, B). TCP: Tai Codonopsis Radix; LCP: Lu Codonopsis Radix.
Through correlation analysis of metal elements and metabolic substances, the contents of the metal elements Zn, Mg, Mn, Ti, Mo, Co, Ni, and Cr in the soil and the hexyl CP composition 6-O-pentopyranosylhexopyranoside and Bufotenine were positively correlated and negatively correlated with the remaining CP composition. K, Al, Ca, Na, Cu, Ba, Cd, V, Sr and Lobetyolin, alpha-D-xylopyranose, 8-benzylcanadine, 1-isopropylbenzimidazole-2-sulfonic acid, 7-methoxy-2-propylquinolin-4-ol, and 1-(2-chlorobenzyl) pyrrole-2-carboxamide were positively correlated and negatively correlated with the other components of CP (Fig. 12A–B).
Correlation analysis diagram of soil metal elements and pharmacodynamic components in TCP and LCP. TCP: Tai Codonopsis Radix; LCP: Lu Codonopsis Radix.
Discussion
Key phenotypic differences and environmental drivers of CP quality
CP, a Medicinal Food Homologous Herb in TCM, has been utilized since ancient times in clinical practice, health products, and dietary regimens. Its quality is intrinsically linked to its geographical origin and growth environment. This study focused on two prevalent commodity specifications: TCP and LCP. Distinct morphological differences were observed in the root head region, particularly marked by a higher incidence and greater density of TCP rings. Further analysis indicates that these features are linked to the plant’s adaptation to the low-temperature plateau environment. Furthermore, significant differences were identified in Lobetyolin, soil microbial flora composition, and soil metal element profiles between TCP and LCP. These phenotypic and compositional differences are fundamentally driven by their distinct growth habitats: TCP thrives at high altitudes, characterized by low temperatures, intense ultraviolet radiation, and diurnal temperature fluctuations exceeding 15 °C, whereas LCP grows in warmer environments at moderate elevations. Understanding these fundamental differences provides essential context for elucidating the underlying mechanisms involved in quality formation. (The specific research approach is shown in Fig. 15).
Multi-omics profiling reveals systemic differences between TCP and LCP
To dissect the molecular and ecological basis underlying quality divergence, we adopted an integrative multi-omics approach to compare TCP and LCP across four key dimensions: climate, secondary metabolites, soil microorganisms, and soil metal elements. Non-targeted metabolomics analysis indicated that TCP was notably enriched in triterpene saponins (e.g., Lobetyolin) and glycosides (e.g., 6-O-hexyl pentopyranosylhexopyranoside), along with compounds such as 9-octadecenoic acid and 5,8,11-trihydroxy fatty acid. Conversely, LCP exhibited higher levels of compounds including Sinapyl alcohol and the alkaloid Bufotenine.
Analysis of the soil microbiome revealed distinct community structures. The bulk soil of TCP was enriched with bacterial phyla such as Gemmatimonadetes, Candidatus Rokubacteria, Chloroflexi, and Verrucomicrobia. Notably, a novel fungal genus, Ascobolus, was identified in the root soil of TCP. In contrast, the bulk soil of LCP showed higher abundances of Betaproteobacteria and the genus Sphingosinicella, with a significantly greater relative abundance of Verrucomicrobia compared to TCP. Soil metal element analysis further differentiated the habitats: the bulk soil of TCP contained elevated levels of Ti, Ni, and Cr, while the bulk soil of LCP was characterized by higher concentrations of Fe, Cu, K, Al, Ba, and Mn.
This multidimensional analysis has established a systematic approach to distinguish TCP from LCP, thus laying a robust foundation for investigating the underlying mechanisms.
Microbial-metal-metabolite interaction network underlying quality divergence
The observed systemic differences arise from a complex interplay between environmental stressors, soil metal elements, and the soil microbiome, which collectively shape the synthesis of key medicinal metabolites in TCP and LCP.
TCP quality formation: synergy of abiotic stress and metal-tolerant microbes driving triterpenoid synthesis
The high-altitude niche of TCP subjects plants to significant abiotic stresses: low temperatures, large diurnal temperature fluctuations, intense UV radiation, and elevated soil Ti, Ni, and Cr. This Selective pressure has favored the proliferation of metal-tolerant and stress-adapted microbial communities in bulk soil, such as Gemmatimonadetes positively correlated with As, Cd, and Pb, and likely adapting via metal resistance genes including metallothioneins10,11, and Rokubacteria which may dominate by competitively inhibiting metal-sensitive bacteria in environments with high levels of Ti, Ni, and Cr12. Chloroflexi, an oligotrophic group, likely thrives under low organic matter conditions induced by high metal stress. This high-metal, low-temperature environment may suppress pathogenic bacteria while potentially promoting beneficial interactions. A pivotal finding was the identification of the novel fungus Ascobolus in the rooted soil of TCP. Ascobolus exhibits remarkable tolerance to Zn, Ni, and Cr, achieved through mechanisms such as secreting oxalic and citric acids to form stable metal–organic acid chelates (thereby reducing the toxicity of free ions) and adsorbing metals (Ni, Ti, Cr) via abundant Carboxyl, Hydroxyl, and Thiol groups on its cell wall. Notably, heavy metal stress activates the SoxS/MerR family of transcription factors in Ascobolus, leading to the generation of reactive oxygen species (ROS) and nitric oxide (NO) signaling molecules13. These diffusible signals enter the plant rhizosphere, inducing the plant’s antioxidant defense systems and activating key secondary metabolic pathways. Specifically as follows:
Fungal-derived ROS/NO signals induce the expression of plant genes encoding 3-hydroxy-3-methylglutaryl coenzyme A reductase (HMGR) and squalene synthase (SQS), thereby significantly enhancing the metabolic flux of isopentenyl pyrophosphate (IPP) and dimethylallyl pyrophosphate (DMAPP). These compounds serve as critical precursors for the biosynthesis of the triterpene skeleton. Concurrently, Ascobolus species exhibit lignin-degrading enzymatic activities, including laccase and lignin peroxidase (LiP), which can oxidize sinapyl alcohol to generate free radical intermediates. These reactive intermediates act as secondary signaling molecules, further upregulating the expression of plant cytochrome P450 monooxygenases and thereby promoting hydroxylation and glycosylation modifications of the triterpene backbone.
Ultimately, this coordinated microbial signaling and enzymatic activity drives the synthesis of high-value triterpenoid saponins, particularly Lobetyolin and Hexyl glycosides (e.g., 6-O-hexyl pentopyranosylhexopyranoside), characteristic of TCP quality. Low temperatures and UV stress further contribute by inducing oxidative stress, activating CYP450 genes, and stimulating glycoside accumulation as photoprotectants. Thus, the TCP environment selects for a unique microbiome (Ascobolus, metal-tolerant bacteria) whose activities and stress-induced signals directly orchestrate the plant’s synthesis of stress-protective and medicinally important triterpenoids (Fig. 13).
A proposed schematic model illustrating the microbial-metal-metabolite interaction network underlying enhanced lobetyolin biosynthesis in Codonopsis Radix.
LCP quality formation: warm climate and metal-mediated regulation of alkaloid synthesis
LCP thrives under warmer conditions at moderate altitudes, with soils enriched in Fe, Cu, K, Al, Ba, and Mn. These environmental conditions support diverse microbial communities and distinct metabolic pathways. The increased abundance of Betaproteobacteria in LCP bulk soil may be associated with elevated Fe and Cu concentrations. Sphingosinicella, a genus recognized for its ability to degrade aromatic compounds and tolerate metal stress14, is also notably abundant. High levels of Al and Fe may enhance plant metal uptake through mechanisms such as siderophore secretion or metal dissolution via organic acids. Furthermore, the significantly higher abundance of Verrucomicrobia in LCP soils is likely attributable to elevated K levels, which play a critical role in regulating cellular osmotic pressure and promoting metabolic activity12. Elevated Al and Ba concentrations may activate microbial phosphatase genes, thereby promoting the mineralization of organic phosphate compounds15. This process releases bioavailable phosphorus, which in turn may support the biosynthesis of alkaloids such as Bufotenine in LCP. Warmer climatic conditions accelerate the metabolic transformation of compounds like sinapyl alcohol. Additionally, synergistic interactions between K and Mn may further enhance the production of specific bioactive constituents. Notably, high levels of K and Al in LCP soils may suppress the growth of certain fungal taxa, including Ascobolus16, thereby inhibiting the activation of TCP-type triterpenoid biosynthesis pathways. Consequently, the LCP environment—characterized by its unique metal composition and elevated temperatures—fosters microbial communities involved in nutrient cycling and metal metabolism. These communities synergistically enhance the biosynthetic pathways of alkaloids such as Bufotenine, ultimately shaping the quality profile of LCP. While interactions between soil microorganisms and metal elements are evident, the underlying signaling mechanisms may differ from those observed in TCP.
In summary, this study identifies a core “Microbial-Metal-Metabolite Interaction Network” that governs the quality divergence of CP. For TCP, the combined effects of elevated Ti/Ni/Cr levels, low temperatures, and UV stress drive the selection of a specific microbiome (including Ascobolus, Gemmatimonadetes, Rokubacteria, and Chloroflexi). Through mechanisms involving metal tolerance, stress signaling (ROS/NO), and enzymatic activity, this microbiome actively enhances the plant’s biosynthetic pathways for triterpenoid saponins (e.g., Lobetyolin, hexyl glycosides). In contrast, for LCP, warmer temperatures and higher concentrations of Fe/Cu/K/Al/Ba/Mn support microbial communities (Betaproteobacteria, Sphingosinicella, Verrucomicrobia) that facilitate metal metabolism, phosphorus mineralization, and nutrient uptake. These communities promote alkaloid (Bufotenine) synthesis while suppressing TCP associated fungi and their corresponding metabolic pathways.
Research implications: authentication paradigms and eco-cultivation strategies for Daodi herbs
The "Microbial-Metal-Metabolite" interaction network model unveiled herein, particularly the identification of keystone microbial taxa (e.g., the previously unrecognized genus Ascobolus) and signaling mechanisms (e.g., ROS/NO pathways) involved in microbe-plant communication, introduces a novel paradigm for assessing Daodi authenticity that extends beyond conventional morphological and chemical markers. Given the severe depletion of wild TCP resources, mechanistic insights derived from this research provide a solid foundation for developing targeted, microbe-mediated ecological cultivation strategies aimed at producing high-quality TCP with elevated Lobetyolin content. Based on the core interaction network identified (section “TCP quality formation: synergy of abiotic stress and metal-tolerant microbes driving triterpenoid synthesis”), the following cultivation protocol is proposed:
Cultivate at elevations ranging from 1800 to 2000 m, where the diurnal temperature fluctuation is at least 15 °C and the annual precipitation ranges between 450 and 550 mm. Implement drip irrigation to maintain approximately 70% soil moisture during dry periods, thereby simulating the natural habitat stressors of TCP. Scale up pure Ascobolus strains from authentic TCP root soil and apply them through seed coating or root dipping during transplantation to introduce signal-inducing and metabolite-modulating microbes into the rhizosphere.
Apply chelated Ni/Zn fertilizers, which are associated with TCP metal profiles and microbial interactions, in combination with microbial fertilizers containing Ascobolus throughout the growth period to support the plant and microbial requirements essential for Lobetyolin synthesis. Conduct trials in representative regions (e.g., Wutai, Pingshun) to assess the impacts on key components, particularly Lobetyolin, as well as the structure and function of the rhizosphere microecosystem, thereby providing a foundation for large-scale, standardized, and sustainable TCP cultivation.
This microbe-mediated cultivation strategy illustrates how mechanistic insights into Daodi formation can be translated from theoretical understanding to practical applications, offering a sustainable approach to resource conservation and quality assurance.
However, certain limitations were observed. During the sampling process, it was noted that the planting density of TCP is significantly lower compared to that of LCP. It is noteworthy that the observed differences in microbial communities and metabolite profiles between TCP and LCP remained consistent across varying planting densities. This suggests that environmental and edaphic factors—such as altitude, temperature, and soil metal content—exert a more pronounced influence on the quality of Codonopsis Radix compared to planting density under the conditions of this study. The inherent differences in planting density between wild-simulated TCP and cultivated LCP were not quantitatively characterized during sampling. While our discussion suggests that macro-environmental factors are the primary drivers of the observed differences, the specific effect of planting density warrants investigation in future studies designed with controlled density gradients to disentangle its influence from that of the environment.
Materials and methods
Materials
On the basis of the findings of the fourth Chinese materia medica resource survey, Wutai County in Shanxi Province was selected as the representative area for TCP sampling, and Pingshun in Changzhi city was designated for LCP collection. The specific sampling sites are detailed in Table 3.
The soil samples were collected in accordance with the Association Standard of the People’s Republic of China-2022 technical specifications for soil sampling, preparation and long-term storage (T/GSSF004). With respect to the S-shaped route, 12 soil samples and root samples were obtained from several sites in the sampling region, all of which had the same position and maturity. The soil samples from the rhizosphere and root system were obtained from four different vertical directions, approximately 1 m from the root, and were then deposited in sterile centrifuge tubes. The samples were promptly immersed in liquid nitrogen and preserved in dry ice, after which they were stored at − 80 °C.
The fresh CP samples from which the soil was collected were removed, and the various parts were then separated. These pieces were placed in sterile self-sealing bags, numbered, promptly immersed in liquid nitrogen, preserved on dry ice and finally stored − 80 °C. The collected samples were assigned numbers, and the precise information is displayed in Table 3.
All CP specimens used in this experiment were collected in August 2023, following a consistent sampling protocol. TCP samples were derived from three-year-old plants, while LCP samples were obtained from three-year-old plants that underwent a cultivation process encompassing one year of seed propagation, one year of seedling raising, and one year of field growth.
The CP was authenticated by Professor Zhang Shuosheng from Shanxi University of Chinese Medicine as the dried root of Codonopsis Radix (Franch.) Nannf. of the Campanulaceae family. The voucher samples were stored in the plant specimen room of the Shanxi College of Traditional Chinese Medicine (specimen number: SXTCM-Zhang-2023001–2023012).
Instruments and reagents
Instruments: The instruments used in this study were as follows: a Q Exactive HF-X mass spectrometer (Thermo Fisher Technology Co., Ltd., CA, USA); a Vanquish UHPLC (Thermo Scientific, Waltham, MA); a low-temperature high-speed Eppendorf 5430R centrifuge (Eppendorf Innovation Co., LTD., GER); a Mettler–Toledo one-thousandth LE204E balance (METTLER TOLEDO Co., USA); a NexlON 5000 ICP-MS (PerkinElmer Co., LTD, CA, USA); an Agilent 710 ICP-OE (Agilent Technologies, Palo Alto, Calif.); a Qubit 2.0 fluorometer (r Life Technologies, CA, USA); an Agilent 2100 bioanalyzer (Agilent Technologies, Palo Alto, Calif.); CFX96 Touch Real-Time PCR (Bio-Rad, Co., CA, USA); a KZ-III-F homogenizer (Servicebio Technology Co., Ltd., Wuhan, CN); an MX-S vortex mixer (Dalong Xingchuang Experimental Instrument Co., Ltd., Beijing, CN); an SB-5200 ultrasonic cleaner (Xinzhi Biotechnology Co., Ltd., Ningbo, CN); and a vacuum freeze dryer (Kangyibo Medical Devices Co., Ltd., Beijing, CN).
Chromatographic column: ACQUITY UPLC HSS T3 (2.1 mm × 100 mm, 1.8 µm, Waters Co., Ltd., MA, USA).
Reagents: Ultra-pure water (MS grade, W6-4, Thermo Fisher Technology Co., LTD, CA, USA); acetonitrile (MS grade, A955-4, Thermo Fisher Technology Co., Ltd., CA, USA); methanol (MS grade, A456-4, Thermo Fisher Technology Co., LTD, CA, USA); formic acid (GR, F0654-25 ml, Tixi Ai Chemical Industry Development Co., Ltd., Shanghai); hydrogen peroxide (GR, 20240103, National Pharmaceutical Group Chemical Reagent Co., Ltd., Shanghai, CN); concentrated nitric acid (GR, 20231220, National Pharmaceutical Group Chemical Reagent Co., Ltd., Shanghai, CN); and hydrofluoric acid (GR, C12897954, McLean Biochemical Technology Stock Co., Ltd., Shanghai, CN).
Test kits: Magnetic bead-based soil and faeces genomic DNA Extraction Kit (TIANGEN Biochemical Technology Co., Ltd., CN); RNase A (10 mg/ml, Cat: RT405-02, TIANGEN Biochemical Technology Co., Ltd., CN); Qubit dsDNA Assay Kit (Q32854, Life Technologies, CA, USA); and NEBNext Ultra DNA Library Prep Kit for Illumina (NEB, Ipswich, MA, USA).
Software and database
Geographical Database: National Tibetan Plateau Data Centre (http://data.tpdc.ac.cn), National Earth System Science Data Center-Soil Data Center (https://soil.geodata.cn/), Originating from the National Environmental Information Center (NCEI) under the National Oceanic and Atmospheric Administration (NOAA) of the United States (https://www.ncei.noaa.gov/data/global-summary-of-the-day/archive/), Climatology Lab (https://www.climatologylab.org/terraclimate.html).
Geographic information system: ArcGIS software (version 10.8, http://www.esri.com/zh-cn/home).
Bioinformatics software: fastp (https://github.com/OpenGene/fastp), Prodigal (https://github.com/hyattpd/Prodigal/releases/tag/v2.6.3), Bowtie2 (Bowtie download|SourceForge.net), DIAMOND (version 0.9.110, http://www.crystalimpact.com), MEGAN (https://software-ab.informatik.uni-tuebingen.de), Gephi (https://gephi.org), BWA (http://maq.sourceforge.net), cBot cluster generation system (https://support.illumina.com.cn), CD-HIT (https://github.com/weizhongli/cdhit), and ARG-OAP (http://smile.hku.hk/ARGs/indexing).
Functional annotation database: Non-redundant protein database (NR database, version 2021.11, wget-c fto://ftp.ncbi.nlm.nih.gov/blast/db/FASTA/nr.gz), Kyoto Encyclopedia of Genes and Genomes database (KEGG database, http://www.genome.jp/), evolutionary genealogy of genes: Non-supervised Orthologous Groups database (eggNOG database, http://eggnog5.embl.de), and Carbohydrate-Active enZYmes database (CAZy database, http://www.cazy.org).
Natural environmental data processing methods
The natural environmental data for Wutai and Pingshun Counties in Shanxi Province, including precipitation, sunshine, and temperature data, were obtained from the National Tibetan Plateau Data Centre. The acquired data were subsequently analysed via ArcGIS software. The processing methodology employed was as follows. A total of ten nc-formatted datasets spanning from 2012 to 2022 were selected, with each dataset containing twelve months of data. The mean temperature data for these years were extracted and stored in .tif format. Additionally, the R language was utilized to retrieve sunshine duration data from the Global Surface Summary of the Day (GSOD) meteorological stations in China covering the period between 2012 and 2022. The IDW inverse weight matrix was utilized for data interpolation, with the inclusion of only 5 adjacent stations to mitigate excessive smoothing. A downscaling operation using the delta algorithm was applied to increase the precision postinterpolation. The final raster data had a spatial resolution of 0.1°*0.1°.
The pH and organic matter content data for Wutai and Pingshun Counties in Shanxi Province from 2012 to 2022 were acquired from the National Earth System Science Data Center-Soil Data Center and processed using ArcGIS software. The datasets were generated by extracting information from the original raster data provided by the platform, and could be directly downloaded from the server and exported through the statistical analysis tools embedded in ArcGIS software.
The average annual temperature and diurnal temperature range data for Wutai County and Pingshun County in Shanxi Province from 2012 to 2022 were obtained from the National Environmental Information Center under the U.S. National Oceanic and Atmospheric Administration (NOAA). The data were provided in two formats—Shapefile (Shp) and Excel—and both were processed accordingly. In the Excel format, missing values were indicated as null entries. Monthly mean temperatures were calculated by averaging daily temperature observations within each month, while annual mean temperatures were derived by averaging the 12 monthly means for each year. Ultraviolet radiation intensity data were initially released in January 2018 by John T. Abatzoglou from the Department of Geography at the University of Idaho, United States, via the Scientific Data platform. The dataset originally covered the period from 1958 to 2015 and has since been updated incrementally to include data through 2024. The spatial coordinate system is based on WGS_1984, with a spatial resolution of 4 km and measurement units expressed in W/m2.
UPLC‒MS/MS analysis
CP root sample preparation: After the sample was pulverized in liquid nitrogen, 100 mg of the resulting powder was weighed and placed in a 1.5 mL centrifuge tube. Then, 1 mL of 70% methanol was added to the tube, and the mixture was vigorously mixed for 30 s by vortexing. The tube was subsequently subjected to sonication in a water bath for 90 min, followed by centrifugation for 10 min at a speed of 16,000 × g at a temperature of 4 °C. The supernatant was transferred to a 96-well protein filter plate and filtered with nitrogen under positive pressure, and the filtrate was transferred to a 2-mL EOP tube and dried under vacuum. Next, 300 μL of 40% methanol solution was added to the lyophilized sample to dissolve it. The mixture was then vigorously agitated for 30 s by vortexing, followed by centrifugation for 10 min (16,000 × g, 4 °C), and the supernatant was collected17.
Chromatographic conditions: The samples were separated via ultrahigh-performance liquid chromatography (UHPLC) in combination with an ACQUITY UPLC HSS T3 column (2.1 mm × 100 mm, 1.8 µm) with the column temperature maintained at 35 °C and a flow rate of 0.3 mL/min. The elution gradient comprised mobile phase A, which was a 0.1% aqueous solution of formic acid, and mobile phase B, which was a 0.1% acetonitrile solution of formic acid. The process of gradient elution was executed in accordance with the information provided in Table 4.
MS conditions: The primary and secondary spectra of the samples were collected via a Q Exactive HFX mass spectrometer, which was connected to a UHPLC instrument and equipped with an electrospray ionization (ESI) source operating in both positive and negative ionization modes. The spray voltage was 3800 V (ESI+)/3500 V (ESI-), the sheath gas flow rate was 40 L/min, the ion transfer tube temperature was 320 °C, and the atomization temperature was 350 °C. The detection mode was full-MS/dd-MS2 mode, MS/MS spectra were obtained from the top 10 MS1 ions, and the collision energy was normalized by stepwise energy levels 20, 40, and 60. The mass‒charge ratio of the first stage was scanned from 90 to 1300.
Properties and microscopic identification
The CP properties were determined in accordance with the criteria specified for the CP traits in the Chinese Pharmacopoeia (2020 edition). The microscopic identification was conducted following the guidelines outlined in the Microscopic Identification Method 2001 of the Chinese Pharmacopoeia (2020 edition).
HPLC method
Chromatographic conditions: The analysis was performed using a Hypersil GOLD aQ column at a temperature of 30 °C. The detection wavelength was set to 220 nm. The mobile phase consisted of acetonitrile (phase A) and an aqueous solution containing 0.3% phosphoric acid (phase B). Gradient elution, as shown in Table 1, was employed for separation. The flow rate was maintained at 1 mL/min, and the injection volume was set to 10 μL.The process of gradient elution was executed in accordance with the information provided in Table 518.
Three milligrams of the reference substance codontoside was precisely weighed and dissolved in 75% methanol. The solution was transferred to dry and clean volumetric flasks with a capacity of 10 mL, ensuring a constant volume. The mixture was shaken well to obtain the reference substance solution. Each reference solution was removed into the same volumetric flask and diluted with 75% methanol to prepare the reference mixture.
Metagenomic sequencing
DNA extraction and quality control
DNA was extracted from the soil samples via a magnetic soil and stool DNA kit (Cat: DP712, TIANGEN Biochemical Technology Co., Ltd., CN). The DNA concentration was determined via a Qubit dsDNA Assay Kit with a Qubit 2.0 fluorometer (Life Technologies, CA, USA). Only the DNA that successfully underwent the quality check was eligible for use in the construction of the metagenomic libraries. The NEBNext Ultra DNA Library Prep Kit for Illumina (NEB) was used to create metagenomic libraries. The attribute sequence of each sample was then supplemented with an index. The index sample generated clusters on the cBot cluster generation system. Following the generation of clusters, library preparation was carried out via a NovaSeq 6000.
Bioinformatics analysis
Data generated from the Illumina platform were used for bioinformatics analysis. The software and parameters are as follows.
Sequencing results preprocessing
The default parameters of fastp19 software were utilized to conduct preprocessing on the raw data acquired from the Illumina sequencing platform. The splicing software MEGAHIT20 was used to assemble the clean data. A de Brujin graph was constructed according to the overlap relationship between k-mers, resulting in the generation of contigs. Contigs exceeding a length of 800 bp were selected for data statistics and further analysis.
Gene prediction and abundance analyses
Prodigal21 was used to predict the ORF of the spliced contig sequence and then translate it into an amino acid sequence. The ORF prediction results were subsequently used to calculate the abundance information for genes in the corresponding samples. The abundance information, measured in transcripts per million (TPM), for each gene in each sample was computed via the following formula (where r represents the number of reads aligned to the gene and L represents the length of the gene)22,23,24,25,26:
Species annotation
DIAMOND27 (V0.9.9.110) was used to align the sequences of bacteria, fungi, Archaea and viruses. The alignment results of each sequence were filtered to include only those with an eigenvalue ≤ 1e−528. The species annotation information of the sequence was then determined via the LCA algorithm, which was applied to the taxonomic assignment of the MEGAN29. On the basis of the abundance tables at each taxonomic level, Krona analysis30 was used to determine the relative abundance profile. Gephi software was then employed to create a species association network map, with a threshold set at a species correlation coefficient > 0.7 and P < 0.01.
Functional annotation
DIAMOND (Version 0.9.9.110) was used to compare the nonredundant gene sets with the KEGG31,32, eggNOG33 and CAZy34 databases. For each sequence alignment result, the selected BestBlastHit results were utilized for further analysis35,36,37. From the results of the alignment, the relative abundance of different functional levels was determined. The number of annotated genes was counted, and the relative abundance profiles are displayed.
Statistical analysis
PCA (Principal Components Analysis)38 (Rade4 package, Version 2.15.3) and NMDS39 (Rvegan package, Version 2.15.3) dimension reduction analyses were applied to compare the species and functional compositions between different groups. ANOSIM and Adonis were employed to assess the dissimilarities between the groups. Then, Kruskal‒Wallis (multigroup) rank sum test (or STAMP analysis)40 and LEfSe analysis41 were used to identify distinct species or functions at each level between the groups.
In this study, each group had six samples. We used the Kruskal–Wallis rank sum test to check for differences between group averages. Our analysis included data from biological replicates. We examined two aspects: OTU levels of specific operational taxonomic units (OTUs) and taxonomic classifications of bacterial species. When we found p-values below 0.05, we considered these differences statistically meaningful.
Metallic element detection
After 0.1 g of the sample was weighed precisely, it was placed into the digesting inner tank. Then, 5 mL of concentrated nitric acid was added, and the mixture was left undisturbed overnight. The next day, 2 mL of hydrogen peroxide and 1 mL of hydrofluoric acid were introduced into the inner tank, and a stainless-steel jacket was firmly sealed. The tank was placed in a constant-temperature drying oven for digestion at 150–170 °C for 4 h. After cooling, the stainless-steel jacket was carefully unscrewed, and the digesting inner tank was subsequently removed. The inner tank was heated on an electric hot plate at 160 °C for 30 min, 1% nitric acid was added to 25 mL, and the content of each element was measured by the instrument. The results were calculated via the following formula:
where C is the concentration of elements in the solution, mg/L; V is the extraction volume, mL; D is the dilution factor; and m is the sample mass, g.
Data are expressed as mean ± standard deviation (SD). Statistical comparisons of metallic element concentrations across groups were conducted using GraphPad Prism 9 software. Normality was evaluated within each group using the Shapiro–Wilk test, and homogeneity of variances was assessed using Levene’s test. For data satisfying both assumptions (normality and homogeneity of variances, P > 0.05), one-way analysis of variance (ANOVA) was performed, followed by Tukey’s post-hoc test for multiple pairwise comparisons. When the assumption of homogeneity of variances was violated (P < 0.05), Welch’s ANOVA was applied, with the Games-Howell post-hoc test used for subsequent comparisons. For non-normally distributed data, the non-parametric Kruskal–Wallis H test was utilized, followed by Dunn’s post-hoc test. A significance level of P < 0.05 was adopted for all analyses.
Conclusion
Codonopsis Radix (Franch.) Nannf., indigenous to Shanxi province, has evolved under the influence of diverse environmental factors, notably climatic variations, soil microbial communities, and mineral composition, leading to the emergence of two distinct commercial cultivars: TCP and LCP. These ecotypic differentiations are manifested not only in their morphological attributes but also in significant variations in the concentration of bioactive compounds and the production of specialized metabolites during ontogeny (Figs. 14, 15). Such phenotypic plasticity reflects adaptive responses to local environmental pressures, underscoring the species’ ecological resilience and biochemical diversity for THMs.
Abstract diagram.
Mechanism diagram.
Data availability
The datasets analysed during the current study are available in the National Center for Biotechnology Information (NCBI), https://dataview.ncbi.nlm.nih.gov/object/PRJNA1267254?reviewer=sh7tg9h3durav6trum4r0mrg84
Abbreviations
- TCP:
-
Tai Codonopsis Radix
- LCP:
-
Lu Codonopsis Radix
- CP:
-
Codonopsis Radix
- THMs:
-
Traditional herbal medicines
- TCM:
-
Traditional Chinese medical
- GTHMs:
-
Genuine Traditional herbal medicines
- YF Group:
-
TCP bulk soil group
- PF Group:
-
LCP bulk soil group
- YG group:
-
TCP rooted soil group
- PG group:
-
LCP rooted soil group
- nc format:
-
Network Common Data Format
- IDW:
-
Inverse distance weighted
- GSOD:
-
Global surface summary of the day
- MS:
-
Mass spectroscopy
- EIS:
-
Electrochemical impedance spectroscopy
- HPLC:
-
High performance liquid chromatography
- DNA:
-
Deoxyribonucleic acid
- ORF:
-
Open reading frame
- LCA:
-
Lowest common ancestor
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- NMDS:
-
Nonmetric multidimensional scaling
- BPC:
-
Base peak chromatogram
- QC:
-
Quality control
- PCA:
-
Principal component analysis
- OPLS-DA:
-
Orthogonal partial least squares discriminant analysis
- PCoA:
-
Principal co-ordinates analysis
- LEfSe:
-
LDA effect size analysis
- eggNOG:
-
Evolutionary gene genealogy Nonsupervised Orthologous Groups
- UPLC-MS/MS:
-
Ultraperformance liquid chromatography tandem mass Spectrometry
- VIP:
-
Variable importance in projection
References
Yuan, Y., Han, Z. & Lu-qi, H. Review of contributing factors and research model of “excellent shape, high quality, and superior effect” of Dao-di herbs. China J. Chin. Mater. Med. 49, 3977–3985. https://doi.org/10.19540/j.cnki.cjcmm.20240516.103 (2024).
Fang, Q. M. et al. Research progress on production districts of Sichuan Dao-di herbs. Zhongguo Zhong Yao Za Zhi 45, 720–731. https://doi.org/10.19540/j.cnki.cjcmm.20200104.101 (2020).
Jin, A. Q. et al. Research on protection mode of Dao-di herbs-taking the protection mode of geographical indications as an example. Zhongguo Zhong Yao Za Zhi 44, 619–623. https://doi.org/10.19540/j.cnki.cjcmm.20181204.001 (2019).
Zhan, Z. L. et al. Study of genuineness based on changes of ancient herbal origin–taking Astragalus membranaceus and Salvia miltiorrhiza as examples. Zhongguo Zhong Yao Za Zhi 41, 3202–3208. https://doi.org/10.4268/cjcmm20161714 (2016).
Qingwen, S. Studies on Taxonomy and Content Analysis of Lobetyolin of the Codonopsis Plants in Guizhou (2007).
Kaixuan, Z., Qiuyang, Y., Yan, R., Sha, L. & Faming, W. Research progress on the origin, evolution, and current status of the authentic geographical varieties of Astragalus membranaceus. Chin. Tradit. Patent Med. 46, 211–216 (2024).
Zhang, K. et al. Integrated widely targeted metabolomics and network pharmacology revealed quality disparities between Guizhou and conventional producing areas of Codonopsis Radix. Front. Nutr. 10, 1271817. https://doi.org/10.3389/fnut.2023.1271817 (2023).
Wang, Z. et al. Study on environmental factors affecting the quality of Codonopsis Radix based on MaxEnt model and all-in-one functional factor. Sci. Rep. 13, 20726. https://doi.org/10.1038/s41598-023-46546-6 (2023).
Fan, L. et al. Culturable endophytic fungi community structure isolated from Codonopsis pilosula roots and effect of season and geographic location on their structures. BMC Microbiol. 23, 132. https://doi.org/10.1186/s12866-023-02848-3 (2023).
Sazykin, I., Khmelevtsova, L., Azhogina, T. & Sazykina, M. Heavy metals influence on the bacterial community of soils: A review. Agriculture 13, 653 (2023).
Hu, B. et al. Assessment of potentially toxic element pollution in soils and related health risks in 271 cities across China. Environ. Pollut. 270, 116196. https://doi.org/10.1016/j.envpol.2020.116196 (2021).
Zhao, P. Y. Responses of Foxtail millet and its rhizosphere soil microbial communities to chromium stress (2023).
Mao, S. B. et al. Isolation and identification of Enterobacter hormaechei BG200 with cadmium tolerance and promoting effect on rice growth under cadmium stress. Microbiol. China 52, 3039–3057. https://doi.org/10.13344/j.microbiol.china.240889 (2025).
Xiao, Y. et al. Two microbes assisting Miscanthus floridulus in remediating multi-metal(loid)s-contaminated soil. Environ. Sci. Pollut. Res. Int. 31, 28922–28938. https://doi.org/10.1007/s11356-024-33032-3 (2024).
Liang, Y. et al. Contents of lobetyolin, syringin, and atractylolide III in Codonopsis pilosula are related to dynamic changes of endophytes under drought stress. Chin. Med. 16, 122. https://doi.org/10.1186/s13020-021-00533-z (2021).
Jansson, J. K., McClure, R. & Egbert, R. G. Soil microbiome engineering for sustainability in a changing environment. Nat. Biotechnol. 41, 1716–1728. https://doi.org/10.1038/s41587-023-01932-3 (2023).
Kim, H. W. et al. NPClassifier: A deep neural network-based structural classification tool for natural products. J. Nat. Prod. 84, 2795–2807. https://doi.org/10.1021/acs.jnatprod.1c00399 (2021).
Hui, Z., Shaoming, S., Zhiqi, W., Luqi, Y. & Xinxin, Z. Content comparison of effective components for two kinds of Codonopsis pilosula cultivated in soil and hydroponics conditions. J. Northwest Normal Univ. (Nat. Sci.) 60, 86–92. https://doi.org/10.16783/j.cnki.nwnuz.2024.02.014 (2024).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: An ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890. https://doi.org/10.1093/bioinformatics/bty560 (2018).
Li, D., Liu, C. M., Luo, R., Sadakane, K. & Lam, T. W. MEGAHIT: An ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics 31, 1674–1676. https://doi.org/10.1093/bioinformatics/btv033 (2015).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinform. 11, 119. https://doi.org/10.1186/1471-2105-11-119 (2010).
Karlsson, F. H. et al. Symptomatic atherosclerosis is associated with an altered gut metagenome. Nat. Commun. 3, 1245. https://doi.org/10.1038/ncomms2266 (2012).
Qin, J. et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464, 59–65. https://doi.org/10.1038/nature08821 (2010).
Villar, E. et al. Ocean plankton. Environmental characteristics of Agulhas rings affect interocean plankton transport. Science 348, 1261447. https://doi.org/10.1126/science.1261447 (2015).
Cotillard, A. et al. Dietary intervention impact on gut microbial gene richness. Nature 500, 585–588. https://doi.org/10.1038/nature12480 (2013).
Le Chatelier, E. et al. Richness of human gut microbiome correlates with metabolic markers. Nature 500, 541–546. https://doi.org/10.1038/nature12506 (2013).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 12, 59–60. https://doi.org/10.1038/nmeth.3176 (2015).
Oh, J. et al. Biogeography and individuality shape function in the human skin metagenome. Nature 514, 59–64. https://doi.org/10.1038/nature13786 (2014).
Huson, D. H., Mitra, S., Ruscheweyh, H. J., Weber, N. & Schuster, S. C. Integrative analysis of environmental sequences using MEGAN4. Genome Res. 21, 1552–1560. https://doi.org/10.1101/gr.120618.111 (2011).
Ondov, B. D., Bergman, N. H. & Phillippy, A. M. Interactive metagenomic visualization in a Web browser. BMC Bioinform. 12, 385. https://doi.org/10.1186/1471-2105-12-385 (2011).
Kanehisa, M. et al. From genomics to chemical genomics: new developments in KEGG. Nucleic Acids Res. 34, D354-357. https://doi.org/10.1093/nar/gkj102 (2006).
Kanehisa, M. et al. Data, information, knowledge and principle: back to metabolism in KEGG. Nucleic Acids Res. 42, D199-205. https://doi.org/10.1093/nar/gkt1076 (2014).
Powell, S. et al. eggNOG v4.0: Nested orthology inference across 3686 organisms. Nucleic Acids Res. 42, D231-239. https://doi.org/10.1093/nar/gkt1253 (2014).
Cantarel, B. L. et al. The Carbohydrate-Active EnZymes database (CAZy): An expert resource for Glycogenomics. Nucleic Acids Res. 37, D233-238. https://doi.org/10.1093/nar/gkn663 (2009).
Li, J. et al. An integrated catalog of reference genes in the human gut microbiome. Nat. Biotechnol. 32, 834–841. https://doi.org/10.1038/nbt.2942 (2014).
Feng, Q. et al. Gut microbiome development along the colorectal adenoma-carcinoma sequence. Nat. Commun. 6, 6528. https://doi.org/10.1038/ncomms7528 (2015).
Bäckhed, F. et al. Dynamics and stabilization of the human gut microbiome during the first year of life. Cell Host Microbe 17, 852. https://doi.org/10.1016/j.chom.2015.05.012 (2015).
Avershina, E., Frisli, T. & Rudi, K. D. novo semi-alignment of 16S rRNA gene sequences for deep phylogenetic characterization of next generation sequencing data. Microbes Environ. 28, 211–216. https://doi.org/10.1264/jsme2.me12157 (2013).
Noval Rivas, M. et al. A microbiota signature associated with experimental food allergy promotes allergic sensitization and anaphylaxis. J. Allergy Clin. Immunol. 131, 201–212. https://doi.org/10.1016/j.jaci.2012.10.026 (2013).
Parks, D. H., Tyson, G. W., Hugenholtz, P. & Beiko, R. G. STAMP: Statistical analysis of taxonomic and functional profiles. Bioinformatics 30, 3123–3124. https://doi.org/10.1093/bioinformatics/btu494 (2014).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biol. 12, R60. https://doi.org/10.1186/gb-2011-12-6-r60 (2011).
Acknowledgements
Thanks to the village committee of Heiyatang Vlg., Menjiaoshi Twp., Wutai Cty., Wengzigou Vlg., Menjiaoshi Twp., Wutai Ct y., Xincheng Vlg., Pingshun city and Zhaodian Vlg., Pingshun Cty for the help in collecting soil and Codonopsis Radix samples needed in this experiment, and thanks for the permission for these samples collection. The authors also thank the National Sect. Plateau Data Center (http://data.tpdc.ac.cn), and the process (https://www.processon.com) provides part of the graphics data.
Funding
This research was funded by the characterize and correlate the changes in the properties of medicinal materials during the manufacturing process of "fire/temperature" in Shanxi Province, China (CZ2023041_019). Additionally, this research aimed to demonstrate the research and application of these materials, establish a TCM processing technology foundation, conduct a survey of TCM resources (CZ2023016), establish the Shanxi Key Laboratory of Chinese Medicinal Materials Processing (202104010910029), develop the Shanxi Modern Agricultural Industry Traditional Chinese Medicine Technology System (CZ2023018), collaborate with Jinyao Comprehensive Development and Utilization Collaborative Innovation Center (CZ2023056_003)(XN2024040_006), support pharmaceutical processing discipline construction at Shanxi University of Chinese Medicine (2023XKJS-22) through their innovation team (2022TD1014), and the Shanxi Key Laboratory of Traditional Chinese Medicine Processing (Shanxi Medicine Comprehensive Development and Utilization Collaborative Center) (XN2024040_006), and Construction of TCM processing discipline (XN2024093_024) with the Chinese Medicine—Key Research Laboratory (CZ2024010_002), and Taiyuan City unveiling-hanging project—Shanxi Authentic Chinese medicine slice quality improvement key technology research, industrialization (KY2024102), Innovation Team of the Administration of Traditional Chinese Medicine(ZYYTD2024034), Discipline of Chinese Materia Medica Processing(2025XK34), Shanxi Provincial Science and Technology Innovation Talent Team (Young) (202304051001044) and Graduate Student Practice and Innovation Project(X2025SJ034).
Author information
Authors and Affiliations
Contributions
Y.W. and X.M. contributed equally and will put their name first on the citation in their CVs. Conceptualization, Z.B. and Y.L.; Investigation, Methodology, Y.W. and X.M; Software, Validation, Visualization, Y.W. and X.H; formal analysis, X.H. writing—original draft preparation, Y.W.; writing—review and editing, X.M; project administration, H.Z. and S.Z.; funding acquisition, S.Z. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
All local, national or international guidelines and legislation were adhered to in the production of this study. Permission to collect plant specimens has been obtained from Changzhi City and Xinzhou City of Shanxi Province for this study.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wu, Y., Meng, X., Han, X. et al. Integrated metagenomic and metabolomic analysis reveals microbial metabolite interactions involved in the quality formation of Codonopsis Radix. Sci Rep 15, 40278 (2025). https://doi.org/10.1038/s41598-025-24002-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-24002-x