Abstract
Genomic prediction models that fit multiple environments globally are valuable tools for assessing cultivar performance across diverse and variable growing conditions. We analyzed 2,064 strawberry (Fragaria × ananassa) accessions genotyped with 12,591 SNP markers. Soluble solids content (SSC) was measured in multi-year trials conducted at seven locations spanning the U.S., Europe, and Australia. Population structure analysis grouped accessions into two major clusters corresponding to subtropical and temperate origins, which was confirmed by significant differences in allele frequency distributions. To improve prediction accuracy across environments, we developed factor analytic models focusing on genotype-by-environment interactions rather than covariance between sub-populations. We compared three genomic prediction approaches: (i) a standard GBLUP model (Gfa), (ii) a GBLUP model incorporating principal component analysis eigenvalues and re-parameterization (Pfa), and (iii) a multi-population GBLUP model that fits sub-population genomic relationship matrices (Wfa). The Pfa and Wfa models achieved the highest prediction accuracy (r = 0.8) for SSC, outperforming individual environment models and the standard GBLUP. These findings demonstrate that accounting for population structure and genotype-by-environment interactions enhances multi-environment genomic prediction and supports practical implementation of genomic selection in global strawberry improvement programs.
Similar content being viewed by others
Introduction
Global genomic prediction has been proposed as a means to integrate datasets from diverse environments and years in horticultural crops, thereby improving prediction accuracy and facilitating cultivar deployment across locations1. Horticultural crops, including high-value fruit and nut species such as strawberry (Fragaria × ananassa), depend on the adoption of improved germplasm that meets grower, consumer, and industry demands2,3. Genotype-by-environment (G×E) interactions are common in plants and must be understood to optimize breeding strategies and cultivar deployment. However, many breeding programs rely on relatively narrow genetic bases derived from a limited set of founding ancestors, which can constrain the ability to capture the full range of G×E interactions4,5,57. Leveraging historical datasets collected across global environments enables breeders to better characterize G×E patterns, understand the genetic basis of complex traits, and identify parents with broader adaptation.
Genomic best linear unbiased prediction (GBLUP) using a genomic relationship matrix (GRM) derived from entry-by-marker genotype data6,7 is widely applied in global genomic prediction because it offers a flexible mixed-model framework8. However, population structure caused by inbreeding, genetic drift, migration, or isolation can influence prediction accuracy by producing differences in allele frequencies and possibly in QTL effects among genetic groups9,10,11,12. Population structure can be quantified from geographic or breeding origin13, pedigree records14, or molecular markers12. If unaccounted for, these differences can inflate variance estimates and bias genomic estimated breeding values (GEBVs), heritability, and predictive ability7. Explicitly incorporating population structure into genomic prediction models may therefore improve accuracy and reduce bias.
PCA-based approach
One approach to account for population structure is to incorporate principal components (PCs) or principal coordinates (PCos) derived from genomic data into prediction models15. Fitting these components as fixed effects can correct for major sources of structure, but because PCs are derived from the same GRM used in the model, this method may result in “double counting” genetic information16,17. Janss et al.16 addressed this by reparameterizing the GBLUP model, partitioning genetic variance across and within subpopulations using eigenvalues from PCA. This PCA-derived relationship matrix in a Gaussian GBLUP framework has been shown to yield higher prediction accuracies than ridge regression or Bayesian methods (BayesA, BayesB)18, with dairy cattle studies reporting slightly higher accuracies compared to the standard GRM19,20,21.
Population-specific GRM approach
Another strategy is to construct a GRM using population-specific allele frequencies rather than overall means, thereby accounting for differences in allele distribution among subpopulations10,22. This method can capture situations where causal variants segregate in only one population. Simulated data suggest that this approach improves prediction accuracy by ~ 2% compared to the standard GRM10. These findings underscore the potential benefits of accounting for population structure, while also indicating that performance gains may vary by species and dataset.
Cultivated strawberry (F. × ananassa) originated in 18th-century France from a hybridization between F. virginiana(North America) and F. chiloensis (South America)23. Today, strawberry is a $15.9 billion global industry24, supported by numerous regionally focused breeding programs. Diversity analyses show that F. × ananassa has a broadly shared genetic base, with structure often aligned to geography or major breeding programs25,26. For example, germplasm from the University of Florida, University of California–Davis, and globally distributed “Cosmopolitan” material form distinct groups26. Additional fine-scale structuring within the USDA-ARS collection further highlights the need to account for population structure when modeling G×E interactions in strawberry25.
Strawberry flavor is a balance of sugars, acids, and aroma compounds27,28,29, with sweetness a key driver of consumer preference30,31,32,33,34. Soluble solids content (SSC), measured by refractometry, is widely used as a proxy for sweetness because sugars comprise 80–90% of SSC35. SSC is a quantitative trait controlled by many minor-effect loci, with few stable across environments27,28,29,36. It can also be negatively correlated with other desirable traits such as firmness and size27,29, making simultaneous improvement challenging. Genomic prediction offers a means to account for environmental and design-related variation, improving selection for SSC while managing trade-offs with other fruit quality traits37.
Only a few studies have applied genomic selection in strawberry38,39,39, but results indicate it can shorten the breeding cycle from three to two years by enabling earlier selection of parents based on GEBVs. Osorio et al.40 reported that predictive ability averaged 0.35 for five polygenic traits when training and validation sets shared individuals, but dropped to 0.24 when they did not, underscoring the role of relatedness.
In this study, we investigate the effect of population structure on genomic prediction for SSC in a large, diverse strawberry panel combining germplasm from breeding programs in the USA, Europe, and Australia. To our knowledge, this is the first genomic prediction study for SSC in strawberry using such a broad and genetically diverse dataset. The results provide insights for the practical implementation of genomic selection for complex traits in strawberry and strategies to effectively control for population structure in global GS datasets.
Materials and methods
Phenotypic data
Soluble solids content was assessed via refractometry (McRoberts 1932) on 2,064 accessions planted in nine trials at seven locations across the U.S.A., Europe, and Australia (Tables 1 and 2). These locations were within regions considered both temperate and subtropical. Below details of experimental design:
Further details regarding the experimental trials are provided in Supplementary Note 1.
RosBREED trials (Corvallis, OR & Benton Harbor, MI)
As part of RosBREED41, 425 clonal strawberry entries were evaluated at USDA-ARS (Oregon) and Michigan State University (Michigan), with 399 and 369 genotypes assessed, respectively. Plantings included cultivars and bi-parental populations in randomized designs (2010–2011), with two adjacent clones per genotype forming one experimental unit. Ripe fruits were collected once per plant during peak season and stored at − 20 °C. Soluble solids content (SSC) was measured from thawed, homogenized fruit using a handheld refractometer.
UF trials (F4 & F5, Balm, Florida)
Conducted in 2014–2015 using randomized block designs with five blocks. Ripe berries were sampled from each plant in December–January, macerated, and SSC measured with a refractometer. Values were averaged over five sampling periods.
NIAB-EMR trial (East Malling, UK)
Clonal genotypes were planted in five blocks (two screenhouses) using a randomized design in 2018. SSC was measured on up to three ripe berries per plant and averaged per plant.
IFAPA trials (Málaga, Spain)
Two trials evaluated 66 genotypes in randomized plots. Two ripe fruits per plant were measured for SSC and averaged.
QLD-DAF trials (Australia)
Two trials were held in 2018 N8 (subtropical, Queensland) with 121 genotypes in two-replicate incomplete blocks, and W8 (temperate, Victoria) with 70 genotypes in randomized blocks. SSC was measured at three harvests; for N8, fruit was frozen, thawed, and homogenized, while for W8, juice was measured immediately.
Genotypic data, curation, and imputation
Genotyping for the Oregon USDA (ORUS) and Michigan State University (MSU) breeding programs (trials C1/2 and B1/2) was performed using the 90 K Strawberry Axiom array (Thermo Fisher, Santa Clara, CA, USA)42, while all other programs employed the IStraw35 384HT Axiom array, developed from a subset of probes on the 90 K array28. Allele calling was conducted using the Axiom Analysis Suite software (Thermo Fisher), and a total of 12,591 SNPs shared between the two arrays were retained for analysis.
Data curation involved removing markers not present on the IStraw35 array from the ORUS and MSU datasets, followed by filtration based on Axiom Analysis Suite quality classifications. Only markers classified as “poly high-resolution,” “no minor homozygous,” or “monomorphic high-resolution” across all datasets were retained18. Accessions appearing in multiple studies were compared for identity; those differing at > 5% of markers were considered distinct and assigned unique accession names (e.g., the ‘Mara des Bois’ genotype in Spain differed from the same cultivar in Michigan and Oregon). For accessions with < 5% differences, consensus genotypes were created by converting discordant calls to missing data. Markers with > 25% missing data and accessions with > 20% missing data were excluded, resulting in a final dataset of 2,064 samples and 12,591 SNPs for downstream analyses.
Missing genotypes were imputed using FImpute v343, applied both across the entire population and within sub-populations. Imputation accuracy was assessed by masking 2,000 genotypes, imputing them, and calculating Pearson correlations and concordance rates across 10 repetitions. SNP distribution was evaluated in 1 Mb windows (~ 830 Mb genome) using CMplot in R, providing a genome-wide view of marker density and ensuring adequate representation of genomic variation.
Population structure
Population structure was characterized using two complementary approaches: ADMIXTURE and principal coordinate analysis (PCoA) based on the genomic relationship matrix. ADMIXTURE analysis was performed with K = 2 ancestral populations, and individuals with ≥ 90% ancestry assigned to a single cluster were classified as “non-admixed,” while those with < 90% ancestry were considered “admixed.” PCoA was conducted using classical multidimensional scaling of the genomic relationship matrix, followed by k-means clustering (K = 2) on the first two principal coordinates. Cluster assignments from both methods were compared to assess concordance. For downstream genomic prediction, ADMIXTURE-based clusters were retained due to their clearer biological interpretability and direct estimation of ancestry proportions, with admixed individuals treated as a separate category.
The optimal number of clusters (K) was evaluated using two criteria: (i) the silhouette method (R package factoextra44‚54, where the k maximizing average silhouette width was selected, and (ii) ADMIXTURE v1.3.045 with 20-fold cross-validation, where the K with the lowest cross-validation error was chosen.
Statistical methods
General mixed model
A general linear mixed model was analyzed using ASReml-R55, incorporating data from all trials and environments.:
where \(\:\mathbf{y}\) is the vector of phenotypic observations, \(\:\mathbf{X}\) is the design matrix for fixed effects (trial × season, block within environment), and \(\:\mathbf{b}\) is the vector of fixed effects. The matrix \(\:{\mathbf{Z}}_{\text{g}}\) links observations to additive genetic effects \(\:\text{a}\), while \(\:{\mathbf{Z}}_{\text{u}}\) links to non-additive effects \(\:\mathbf{u}\). The residual term is \(\:\mathbf{e}\). Certain fixed or random effects were omitted depending on trial design; details are provided in Table 3.
Additive genetic effects and G×E covariance
Additive genetic effects were modelled as a genotype-by-environment (G×E) term:
where \(\:\varvec{G}\) is the genomic relationship matrix among individuals and \(\:{{\Sigma\:}}_{\varvec{A}}\) is the additive covariance matrix across environments. To parsimoniously capture cross-environment correlations, a factor-analytic (FA) decomposition was applied to \(\:{{\Sigma\:}}_{\varvec{A}}\):
where \(\:\varvec{\Lambda\:}\) is the environment-by-factor loading matrix and \(\:\varvec{\Psi\:}\) is diagonal, containing environment-specific variances. Competing FA models (FA1–FA3) were compared using AIC, and the most parsimonious was selected. Importantly, the FA was applied to \(\:{\varvec{\Sigma\:}}_{\varvec{A}}\) (the additive covariance).
Genetic correlations between trials
Additive genetic correlations between environments \(\:i\) and \(\:j\) were estimated from \(\:{{\Sigma\:}}_{\varvec{A}}\):
These additive correlations reflect the consistency of heritable effects across trials and are directly relevant for genomic prediction.
Genomic relationship matrices
To assess the impact of population structure, three approaches were used to construct \(\:\mathbf{G}\):
-
1.
Standard GBLUP: \(\:\mathbf{G}\) was computed from centered genotypes using allele frequencies across the entire population:
$$\:\mathbf{G}=\frac{\mathbf{M}{\mathbf{M}}^{\mathbf{\top\:}}}{2\sum\:_{i}pi(1-{p}_{i})}$$where \(\:\mathbf{M}\) is the centered marker matrix (columns = loci, rows = individuals) and \(\:{p}_{i}\) is the allele frequency at locus \(\:i\). When required, \(\:\mathbf{G}\) was bent to be positive-definite46.
-
2.
P-GBLUP: Principal components (PCs) derived from \(\:\mathbf{G}\) were included as fixed covariates to control for population structure. Enough PCs were retained to explain ~ 99% of the genetic variance. This model is equivalent to standard GBLUP with PCA covariates.
-
3.
Population-specific GRM: Separate GRMs were built for each subpopulation using population-specific allele frequencies10:
$${\mathbf{G}}_\text{pop}=\frac{{\mathbf{S}}_\text{pop}{\mathbf{S}}_\text{pop}^\text{T}}{\sum_{j}2{p}_{j,{pop}}(1-{p}_{j,{pop}})}$$where \(\:{\varvec{S}}_{\text{pop}}\) is the centered genotype matrix for the subpopulation and \(\:{p}_{j,\text{pop}}\) is the allele frequency at locus \(\:j\) in that group.
Within-trial genomic environments were defined as combinations of seasons within a location that exhibited homogeneous additive variance and near-unity pairwise additive correlations. Single-trial models were first fit to define environments and then combined across trials using the FA parameterization of \(\:{{\Sigma\:}}_{\mathbf{A}}\).
Generalized genomic heritability
Generalized genomic heritability was estimated to quantify the proportion of trait variability attributable to genetic differences. Heritability was calculated for each trial following the method described by Hardner et al4. The heritability for trial \(\:t\) was computed as:
where \({\overline{\sigma}}_{\varDelta{A,t}}^{2}\) is the mean variance of the difference of additive predictions at the \(\:{t}^{th}\) trial, estimated from the prediction error variance matrix of additive effects and \({\widehat{\sigma}}_{\varDelta{A,t}}^{2}\) is the estimated additive genetic variance at the \(\:{t}^{th}\) trial.
Prediction accuracy
Expected accuracy
Expected prediction accuracy for an individual was computed as:
where \(\:\widehat{\mathbf{A}}\) is the predicted additive effect, \(\:\mathbf{A}\) is the true additive effect, and \(\:{\sigma\:}_{\varvec{A},t}^{2}\) is the additive genetic variance at trial t.
Realized accuracy
Realized accuracy was assessed by k-fold cross-validation within and across environments. For each fold, individuals in the validation set were excluded from model fitting, marker effects were estimated from the training set, and genomic breeding values were predicted for the validation set. Accuracy was calculated as the Pearson correlation between predicted breeding values and reference genotypic values57 . To account for the imperfect reliability of the phenotypes, this correlation was further divided by the square root of the generalized heritability at the corresponding trial.
Results
SNP distribution, allele frequency and imputation
SNP markers were evenly distributed across the genome in 1 Mbp windows (Figure S1), with an average density of 84 SNPs per 1 cM. The largest physical gaps between adjacent markers were observed on chromosomes 6 and 3 (up to 35 cM), while the smallest gaps occurred on chromosomes 1, 5, and 7. Allele frequencies differed markedly between the two sub-populations (SP1 and SP2), with 98% pairwise dissimilarity (Figure S3) and a fixation index (Fst) of 0.35. Across populations, 12.5% of loci contained missing genotypes (10% in SP1 and 15% in SP2). Genotype imputation achieved 90% concordance when performed population-wide but nearly 99% when performed within populations, and the latter results were used for downstream analyses.
Population structure
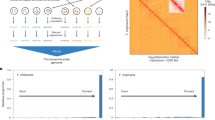
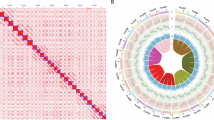
Clustering results from ADMIXTURE and PCoA were largely concordant, with the majority of individuals assigned to the same clusters across methods (Fig. 1and Figure S2). ADMIXTURE analysis (K = 2) with 20-fold cross-validation revealed two primary genetic clusters and a subset of individuals showing substantial mixed ancestry, defined here as having less than 90% ancestry assigned to either cluster (i.e., more than 10% from both clusters). Using this threshold, 1,111 individuals (54%) were classified as Cluster 1 (SP1), 387 individuals (19%) as Cluster 2 (SP2), and 566 individuals (27%) as admixed (Fig. 1C&D). In parallel, principal coordinate analysis (PCoA) of the genomic relationship matrix, followed by k-means clustering (K = 2) of the first two coordinates, produced similar groupings, with most discrepancies occurring near cluster boundaries and involving the admixed group identified by ADMIXTURE (Fig. 1B & S1). Across the 2,064 accessions planted in seven locations, these genetic clusters were also geographically structured (Figure S1C): SP1 consisted primarily of accessions tested in Florida (P), forming a distinct subclade with only a few accessions from Australian trials (Nambour, QLD [N], and Wandin, VIC [W]), whereas SP2 was composed almost entirely of accessions tested in Benton Harbor, MI (B); Corvallis, OR (C); East Malling, U.K. (E); and Málaga, Spain (M). The admixed group included accessions from multiple locations, consistent with their intermediate genetic composition. Silhouette analysis supported K = 2 as the optimal number of clusters (Figure S1A) and cross-validation error from ADMIXTURE (Figure S1C), with average silhouette widths of 0.09 for SP1 and 0.22 for SP2, indicating moderate within-cluster cohesion. Given the clearer biological interpretability and direct representation of ancestry proportions, we used ADMIXTURE-defined clusters including the admixed category for downstream genomic prediction to more accurately capture population structure and admixture in the dataset.
Comparison of population structure inferred by ADMIXTURE and Principal Coordinates Analysis (PCoA) in strawberry samples. Panel (A) shows the ADMIXTURE bar plot where individuals are represented by their ancestry proportions from two clusters. Individuals with less than 90% ancestry from any single cluster are classified as “Admixed,” shown as mixed proportions rather than a solid color. Panel (B) presents the PCoA scatter plot, where samples are grouped into three categories: Cluster 1 (blue), Cluster 2 (green), and Admixed (purple). Panel (C) compares the number of samples assigned to each category by ADMIXTURE and PCoA using side-by-side barplots. Panel (D) displays a confusion heatmap illustrating the correspondence between ADMIXTURE and PCoA group assignments, including the admixed group, highlighting both concordance and discrepancies between these complementary approaches. Information on the optimal K value is provided in Figure S2.
Standard GBLUP
Single location GBLUP
We have reduced the complexity of the models by removing factors, interactions and combining trial within locations (Table S3, Table 3). There was no interaction between genetic effects and year for the most parsimonious individual trial models (Table 3 and Table S1 & S3). Variance component estimates for the single-trial model were presented in Fig. 2 and Table S1. In some trials, the estimated additive genomic variance (vA) was relatively higher than the residual variance (vR), indicating that additive genetic effects contributed more to the observed variation in those specific cases (Fig. 2 and Table S1). In addition, genetic correlations between individual trials (Fig. 3) provide key insights into the stability of genetic values across environments. High positive correlations (such as those observed between the Nambour and Wandin trials with other individual trials) indicate strong consistency in genetic effects, suggesting shared genetic control and the potential for joint or across-environment model selection. In contrast, correlations close to zero (e.g., between the Corvallis and Kent trials) reflect minimal genetic overlap, implying that these environments differ substantially in their genetic architecture and may need to be analyzed separately in downstream applications. The proportion of total genomic variance explained by additive genomic effects was more variable for the single-trial models for Málaga and Balm, FL trials (Fig. 2 and Table S1). Generalized heritability was highest for the Florida trial (h2 = 0.45) followed by Málaga trial (h2 = 0.41) and the lowest was recorded at East Malling, U.K. and Benton Harbor, MI (h2 = 0.16 and 0.18, respectively) (Fig. 4). Realized prediction accuracy (square root of reliability) ranged from 0.44 at the Benton Harbor, MI trial to 0.72 for the Balm, FL trials (Fig. 4).
Variance component estimates for the single-location single-trial model (details in Table S1 and Figure S2).
Genetic correlations between individual trials. These estimates are central to the study, as they indicate the stability of genetic values across environments. Positive correlations suggest shared genetic information and potential for model selection, while correlations near zero indicate limited genetic overlap, implying that environments should be treated independently for downstream analysis.
Genomic heritability (h2) and reliability (A) for individual environment (IE) and multi-trial models. (B) * Trials from the same locations were combined (IE-M4&5 = M; IE-F4&5 = F) when genomic heritability and prediction accuracy are estimated. IE = individual environment (italic); Gfa = factor analytic model (FA) based on standard GBLUP model; Pfa = factor analytic model (FA) based on standard GBLUP + PCA model; Wfa = factor analytic model (FA) based on multi-population model (detail description of the model is provided in M&M section).
Multi-location GBLUP
Compared to the single location models, narrow sense heritability (h2) and prediction accuracy values were higher for the multiple locations standard GBLUP approach for all trials (Fig. 2 and Figure S4). Under the multi-location models, heritability was highest for the Florida trial (h2 = 0.61) and lowest for the East Malling, U.K. and Benton Harbor, MI U.S.A. trials (h2 = 0.27 and 0.28, respectively). This reflects what was observed when modeling environments individually. On average, the multi-location approach increased h2 estimates by 0.16. Prediction accuracies ranged from 0.53 at the Wandin, AUS trial to 0.75 at the Nambour, Australia. On average, prediction accuracies increased by 0.06 when incorporating multiple environments into the model.
P-GBLUP (Janss PCA method)
The relative size of realized prediction accuracy among trials for the GBLUP model that used a reparametrized GRM based on eigen decomposition (P-GBLUP) was similar to that observed for the standard multi-environment approach, where the Nambour and Wandin trials had the highest (r = 0.79) and lowest (r = 0.56) prediction accuracies, respectively (Fig. 4). The P-GBLUP model explained approximately 76% of the phenotypic variance. For all trials, realized prediction accuracies obtained from the GBLUP + PCA model were higher than the standard GBLUP (Fig. 4).
Population specific GRM approach
Realized genomic prediction accuracy for the multi-population model that accounted for population structure through the kinship matrix displayed the same relative prediction accuracy as the standard GBLUP + PCA approach. The population specific (Wfa) model explained approximately 39% of the phenotypic variance.
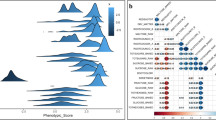
Distribution of best linear unbiased prediction (BLUP) of the Multi-location model (i.e., standard GBLUP model (Gfa1), standard GBLUP + PCA (Pfa1), and population specific model (Wfa1) model) and additional information is provided in Table S2. (A) Distribution between Gfa1 BLUP values vs Pfa1 BLUP values (B) Gfa1 BLUP values vs Wfa1 BLUP values (C) Gfa1 BLUP values vs Pfa1 BLUP values.
Comparing the multi-location approaches
In a multi-location model, the lowest prediction accuracies were achieved in the standard model (Gfa) whereas the two approaches that account for population structure in the prediction model (Pfa and Wfa) achieved higher and more stable accuracies across trials than the standard GBLUP approach (Figs. 4 and 5 and Figure S4). The total genomic correlation matrix across genomic environments estimated from the most parsimonious multivariate for SSC assessed across breeding trials. Genomic environments are defined as groupings of trial-by-seasons such that genomic variance is homogeneous, and genomic correlations are 1 within environments. The factor analytic (FA) model was selected after comparing FA1 to FA3, with the most parsimonious model chosen for subsequent genomic prediction scenarios (Table S2). In addition, the BLUP distribution and correlation between the multi-populations further confirmed the presence of variation in BLUP predictors (Figs. 4 and 5). A strong positive correlation of BLUPs was observed between the Gfa, Pfa, and Wfa approaches for the Florida (F; r = 0.83–0.96), Málaga (M; r = 0.75–0.9), and Corvallis, OR (r = 0.7–0.8) trials; whereas unstructured distribution and low correlation between BLUPs were observed across the multi-population approaches for the Nambour and Wandin trials (Fig. 5 & Figure S4). Genomic heritability followed the same trend as the prediction accuracy estimates with the exception of the Nambour, AUS trial. For this trial, heritability was noticeably lower for the Pfa and Wfa approaches compared to the standard multi-location GBLUP approach.
In most cases, the models that accounted for population structure (standard GBLUP + PCA [Pfa] and multi-population [Wfa]) approaches) generated the highest prediction accuracies (r = ~ 0.8) and showed the lowest variation across trials. Similarly, genomic heritability followed the same pattern as prediction accuracy, where the multi-population approach exhibited high heritability estimates.
Discussion
This study evaluated strategies to account for population structure in genomic prediction models, using a large and diverse panel of global strawberry clones for soluble solids content (SSC). We found that models explicitly accounting for population-specific genomic relationships (multi-population GBLUP) achieved higher prediction accuracies compared to standard GBLUP models that ignore structure. Prediction accuracy varied considerably across environments in the single-trial univariate models, with the highest accuracy observed in the Florida trials (F4 and F5) and the lowest in Benton Harbor, MI. Combining trials from the same location in a multi-trial model increased the size of the reference population and improved prediction accuracy. Further improvements were obtained when population structure was incorporated into the multi-trial analysis, highlighting the benefit of using population-specific genomic relationship matrices rather than a single matrix for the entire population.
Analysis of global genetic relatedness revealed two major sub-populations (SP1 and SP2), broadly associated with subtropical and temperate growing environments. This structure was consistent with previous findings25,26 and is likely the result of historical germplasm exchange, particularly between the Florida and Australian breeding programs. Genetic diversity between the two groups was further supported by differences in allele frequency distributions, which have implications for the unbiased estimation of genetic correlations47. Accounting for this structure proved critical for improving genomic prediction. In our data, correcting for population structure increased prediction accuracy by up to 20% in single-location models and by about 10% in multi-location models. Similar results have been reported in maize, wheat, and cattle, where ignoring structure reduced accuracy, particularly for across-population predictions7,9,10,13,48,49,50,51,52.
Our results confirm that population structure between temperate and subtropical germplasm directly influences prediction accuracy, and models such as Pfa and Wfa generally improved reliability. However, performance gains were not uniform across all locations, indicating that environmental and genetic factors may interact in complex ways. The observed variability in model performance underscores the importance of tailored model development. Environments with low BLUP correlations likely reflect situations where additional covariates or interaction terms are needed. Future research should aim to identify these location-specific factors to further refine model robustness and generalizability.
The implication in breeding
Strong population structure can lead to biased predictions if not addressed, potentially causing false positives and false negatives in marker–trait associations. Best practice involves evaluating population and family structure prior to genomic prediction, using population-specific allele frequencies to construct GRMs, and adopting reduced-dimensionality approaches to handle complex genotype-by-environment covariance structures. Equally important is the use of diverse and representative training populations to ensure shared genetic backgrounds between training and prediction sets4.
Data availability
Input and output data and codes used to analyze genomic prediction using strawberry global collection can be accessed: GitHub: [https://github.com/DrMulusewFikere/StrawberryGP].
References
Hardner, et al. Global genomic prediction in horticultural crops: promises, progress, challenges and outlook. Front. Agr Sci. Eng. 8(2), 353–355 (2021).
Goldschmidt, E. E. The evolution of fruit tree productivity: A review. Econ. Bot. 67 (1), 51–62 (2013).
Ashworth, V. E. T. M., Chen, N. H. & Clegg, M. T. Fruits and nuts. Berlin Springer (2017).
Hardner, C. Exploring opportunities for reducing complexity of genotype-by-environment interaction models. Euphytica 213 (11), 248 (2017).
Ru, S. et al. Current applications, challenges, and perspectives of marker-assisted seedling selection in rosaceae tree fruit breeding. Tree. Genet. Genomes. 11 (1), 8–19 (2015).
VanRaden, P. M. Efficient methods to compute genomic predictions. J. Dairy Sci. 91 (11), 4414–4423 (2008).
Hayes, B. J. et al. Accuracy of genomic breeding values in multi-breed dairy cattle populations. Genet. Selection Evol. 41 (1), 51 (2009).
Hardner, C. M. et al. Prediction of genetic value for sweet Cherry fruit maturity among environments using a 6K SNP array. Hortic. Res. 6 (1), 6–20 (2019).
Werner, C. R. et al. How population structure impacts genomic selection accuracy in cross-validation: Implications for practical breeding. 11 (2020), (2028).
Wientjes, Y. C. J. et al. Multi-population genomic relationships for estimating current genetic variances within and genetic correlations between populations. Genetics 207 (2), 503–515 (2017).
Guo, Z. et al. The impact of population structure on genomic prediction in stratified populations. Theor. Appl. Genet. 127 (3), 749–762 (2014).
Windhausen, V. S. et al. Effectiveness of genomic prediction of maize hybrid performance in different breeding populations and environments. G3 Genes|Genomes|Genetics. 2 (11), 1427–1436 (2012).
Albrecht, T. et al. Genome-based prediction of testcross values in maize. Theor. Appl. Genet. 123 (2), 339 (2011).
Saatchi, M. et al. Accuracies of genomic breeding values in American Angus beef cattle using K-means clustering for cross-validation. Genet. Sel. Evol. 40–43 (2011).
Price, A. L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38 (8), 904–909 (2006).
Janss, L. et al. Inferences from genomic models in stratified populations. Genetics 192 (2), 693 (2012).
Pocrnic, I. et al. Accuracy of genomic BLUP when considering a genomic relationship matrix based on the number of the largest eigenvalues: a simulation study. Genet. Selection Evol. 51 (1), 75 (2019).
Hosseini-Vardanjani, S. M. et al. Incorporating prior knowledge of principal components in genomic prediction. 9 (289), (2018).
Macciotta, N. P. P. et al. Using eigenvalues as variance priors in the prediction of genomic breeding values by principal component analysis. J. Dairy Sci. 93 (6), 2765–2774 (2010).
Dadousis, C. et al. A comparison of principal component regression and genomic REML for genomic prediction across populations. Genet. Selection Evolution: GSE. 46 (1), 60–60 (2014).
Du, C. et al. Genomic selection using principal component regression. Heredity 121 (1), 12–23 (2018).
Makgahlela, M. L. et al. The Estimation of genomic relationships using breedwise allele frequencies among animals in multibreed populations. J. Dairy Sci. 96 (8), 5364–5375 (2013).
Darrow, G. M. M. The strawberry: History, breeding, and physiology. Holt, Rinehart and Winston. (1966).
Food and agriculture organization of the United Nations, Crop report. (2018).
Zurn, J. H. M., Hummer, K., Knapp, S. & Bassil, N. Exploring the diversity and structure of the U.S. National cultivated strawberry collection. Unpublished (2021).
Hardigan, M. A. et al. Unraveling the complex hybrid ancestry and domestication history of cultivated strawberry. Mol. Biol. Evol. 38 (6), 2285–2305 (2021).
Lerceteau-Köhler, E. et al. Genetic dissection of fruit quality traits in the octoploid cultivated strawberry highlights the role of homoeo-QTL in their control. Theor. Appl. Genet. 124 (6), 1059–1077 (2012).
Verma, S. et al. Clarifying sub-genomic positions of QTLs for flowering habit and fruit quality in U.S. Strawberry (Fragaria×ananassa) breeding populations using pedigree-based QTL analysis. Hortic. Res. 4 (1), 17062 (2017).
Zorrilla-Fontanesi, Y. et al. Quantitative trait loci and underlying candidate genes controlling agronomical and fruit quality traits in octoploid strawberry (Fragaria × ananassa). Theor. Appl. Genet. 123 (5), 755–778 (2011).
Bhat, R. et al. Consumers perceptions and preference for strawberries A case study from Germany. Int. J. Fruit Sci. 15 (4), 405–424 (2015).
Colquhoun, T. A. et al. Framing the perfect strawberry: an exercise in consumer-assisted selection of fruit crops. J. Berry Res. 2, 45–61 (2012).
Jouquand, C. Chemical analysis of fresh strawberries over harvest dates and seasons reveals factors that affect eating quality. J. Am. Soc. Hortic. Sci. 133(6), 859–867 (2008)
Lewers, K. S. et al. Consumer preference and physiochemical analyses of fresh strawberries from ten cultivars. Int. J. Fruit Sci. 20 (sup2), 733–756 (2020).
Schwieterman, M. L. et al. Strawberry flavor: diverse chemical Compositions, a seasonal Influence, and effects on sensory perception. PLOS ONE. 9 (2), e88446 (2014).
Perkins-Veazie, P., Collins, J. K. & Cartwright, B. Ethylene production in watermelon fruit varies with cultivar and fruit tissue. HortScience HortSci. 30 (4), 825G–826 (1995).
Fan, Z. et al. Strawberry soluble solids QTL with inverse effects on yield. Hortic. Res. 11(2), (2023).
Gezan, S. A. et al. An experimental validation of genomic selection in octoploid strawberry. Hortic. Res. 4 (1), 16070 (2017).
Cockerton, H. M. et al. Genomic informed breeding strategies for strawberry yield and fruit quality traits. Front. Plant. Sci. 12, 724847 (2021).
Yamamoto, E. et al. Genomic selection for F1 hybrid breeding in strawberry (Fragaria × ananassa). Front. Plant Sci. 12, (2021).
Osorio, L. F. et al. Independent validation of genomic prediction in strawberry over multiple cycles. Front. Genet. 11(1862) (2021).
Iezzoni, A. F. et al. RosBREED: bridging the chasm between discovery and application to enable DNA-informed breeding in rosaceous crops. Hortic. Res. 7 (1), 177 (2020).
Bassil, N. V. et al. Development and preliminary evaluation of a 90 K Axiom® SNP array for the allo-octoploid cultivated strawberry Fragaria× Ananassa. BMC Genom. 16 (1), 155 (2015).
Sargolzaei, M., Chesnais, J. & Schenkel, F. FImpute -An efficient imputation algorithm for dairy cattle populations. J. Dairy. Sci. (94), 421 (2011).
Kassambara, A. & Mundt, F. Factoextra: Extract and visualize the results of multivariate data analyses (R package version 1.0.7) (2020).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based Estimation of ancestry in unrelated individuals. Genome Res. 19 (9), 1655–1664 (2009).
Nazarian, A. & Gezan, S. A. GenoMatrix: A software package for Pedigree-Based and genomic prediction analyses on complex traits. J. Hered. 107 (4), 372–379 (2016).
Wientjes, Y. C. J. et al. Required properties for markers used to calculate unbiased estimates of the genetic correlation between populations. Genet. Selection Evol. 50 (1), 65 (2018).
Hickey, J. M. et al. Evaluation of genomic selection training population designs and genotyping strategies in plant breeding programs using simulation. Crop Sci. 54 (4), 1476–1488 (2014).
Lehermeier, C. et al. Usefulness of multiparental populations of maize (Zea Mays L.) for genome-based prediction. Genetics 198 (1), 3–16 (2014).
Herter, C. P. et al. Accuracy of within- and among-family genomic prediction for fusarium head blight and septoria tritici blotch in winter wheat. Theor. Appl. Genet. 132 (4), 1121–1135 (2019).
de Roos, A. P. W., Hayes, B. J. & Goddard, M. E. Reliability of genomic predictions across multiple populations. Genetics 183 (4), 1545–1553 (2009).
Hayes, B. J., Visscher, P. M. & Goddard, M. E. Increased accuracy of artificial selection by using the realized relationship matrix. Genet. Res. 91, 47–60 (2009).
Massman, J. M. et al. Genomewide predictions from maize single-cross data. Theor. Appl. Genet. 126 (1), 13–22 (2013).
Lengyel, A. & Botta-Dukát, Z. Silhouette width using generalized mean A flexible method for assessing clustering efficiency. Ecol. Evol. 9 (23), 13231–13243 (2019).
Butler, D. G. et al. ASReml estimates variance components under a general linear mixed model by residual maximum likelihood (REML) ASReml-R Version 4.2. (2023).
Korsgaard, I. R., Andersen, A. H. & Jensen, J. Prediction error variance and expected response to selection, when selection is based on the best predictor – for Gaussian and threshold characters, traits following a Poisson mixed model and survival traits. Genet. Selection Evol. 34 (3), 307 (2002).
Branchereau, C. et al. Genotype-by-environment and QTL-by-environment interactions in sweet cherry (Prunus avium L.) for flowering date. Front. Plant Sci. 14 (1142974), (2023).
Author information
Authors and Affiliations
Contributions
CH conceptualized the study, led the project, and contributed to manuscript writing. MF proposed the study design, conducted data analysis, and wrote the manuscript. JDZ performed data quality control and contributed to manuscript writing. NVB also contributed to manuscript writing. SV, IA, PM, JFS, HMC, RJH, LLM, TMD, JFH, CEF, MMM, JN, HLK, and VMW and NVB led field experiments at their respective experimental sites and provided essential data for the study. All authors have read and approved the final manuscript for submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Fikere, M., Zurn, J.D., Verma, S. et al. Accounting for population structure in genomic prediction of strawberry sweetness at a global scale. Sci Rep 15, 40547 (2025). https://doi.org/10.1038/s41598-025-24188-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-24188-0