Abstract
Wheat leaf rust, caused by Puccinia triticina Eriks., is recognized as one of the most destructive diseases affecting wheat worldwide, including Iran, resulting in substantial losses in grain yield and quality. This research focused on evaluating the pathogenic factors of nine leaf rust isolates collected from four different climates in Iran, using various differential genotypes. The assessment of leaf rust infection types was conducted on 49 durum and bread wheat genotypes, including susceptible control genotypes and 55 differential genotypes at the seedling stages. The results revealed a significant difference among wheat genotypes in their response to all isolates (p ≤ 0.01). Notably, certain genotypes, such as the Italian landrace (P.S. No4), Shabrang, Chamran2, Mehregan, Shosh, and Gonbad, exhibited resistance to all isolates at the seedling stage, indicating the presence of seedling resistance genes. Additionally, we determined the virulence/avirulence patterns for various resistance genes in the differential genotypes by assessing their responses to different isolates and recording the infection types. The findings indicated that all isolates were virulent on the lines carrying the Lr34 and Lr37 genes, whereas none of the isolates displayed a virulence on the lines carrying the Lr19 gene. This research provides valuable insights into the resistance patterns of wheat genotypes against leaf rust isolates in different climates in Iran, contributing to our understanding of the genetic basis of resistance and aiding in the development of effective strategies for disease management in wheat cultivation. The XGBoost (extreme gradient boosting) algorithm generated the most accurate predictions for the variables thousand grain weight and grain yield, while the MARS (multivariate adaptive regression spline) algorithm generated the most accurate predictions for the variables spike weight, number of grains per spike, and grain weight per spike. For each of these variables, GP (Gaussian process), MARS, and XGBoost achieved the lowest RMSE (root mean square error) values, indicating minimal prediction errors, and the highest R² values, signifying a strong correlation between the predicted and observed data. These prediction performances highlighted the robustness and accuracy of the GP, MARS and XGBoost algorithms in modeling wheat disease severity and its effects on yield outcomes.
Similar content being viewed by others
Introduction
Wheat brown rust disease, commonly referred to as leaf rust and caused by Puccinia recondita f. sp. tritici, poses a significant threat to wheat grain yield, resulting in substantial losses1. While earlier studies primarily focused on the effectiveness of resistance genes in controlling leaf rust more recent research has specifically quantified yield losses caused by this disease, which can range from 5 to 15% depending on crop growing season conditions, the severity of the disease epidemic, and the growth stage of the plant2. Notably, leaf rust caused a 14% loss in Kansas winter wheat in 2007, impacting U.S. wheat production significantly, with susceptibility leading to a loss rate exceeding 50% in certain cultivars3. Additionally, the economic impact of a brown rust epidemic in Pakistan in 1987 was estimated at $86 million4. In recent times in Iran, the significance of this disease has surpassed strip rust but remains lower than yellow rust5. Grain yield losses of 10–30% have been reported in Iranian wheat fields. Dadrezaei, et al.3 estimated a brown rust-induced yield loss of 25.2% for all studied wheat genotypes in Ahvaz. The reduction in wheat yield caused by leaf rust infection is attributed to decreased grain per spike and 1000-grain weight3. Importantly, a crop cultivar may exhibit susceptibility during the seedling stage or early growth stages but show resistance in the adult plant stage6. Breeding for resistance, as emphasized by Qi, et al. 7, plays a crucial role in preventing yield losses.
Planting resistant cultivars emerges as the most effective method for disease control. To develop such cultivars, identifying virulent races and phenotypes within a region and those introduced from other countries is crucial. The understanding of resistance to leaf rust has been advanced through the identification of specific genes (Lr genes) with their sequence numbers initiated by Harrington, et al. 8. Johnston and Heyne9 pioneered the development of wheat near-isogenic lines differing by single genes for leaf rust resistance. Subsequently, additional near-isogenic lines were derived from the wheat cultivar ‘Thatcher,’ proving invaluable for scrutinizing virulence variations in the P. triticina population and unraveling the genetics of leaf rust resistance. The study of P. triticina populations among plant pathogens has a rich history, with rust race investigations starting in the United States in 1926, Canada in 193110, and Australia in 192011. Global P. triticina populations exhibit high diversity in virulence phenotypes or races12.
The effectiveness of race-specific resistance genes varies worldwide due to the widespread use of such genes in the United States, resulting in a rapid increase in virulent leaf rust races. In Australia, the effectiveness of race-specific resistance genes persists for a more extended period, likely due to the presence of less susceptible cultivars. New virulence phenotypes have emerged globally in recent years, indicating the continual evolution of P. triticina. For instance, in 1984, a virulence phenotype on Lr27, Lr16, and Lr31 initially appeared in Australia but originated from outside the country13. In 2001, phenotypes of P. triticina emerged as virulent on Mexican durum wheat cultivars that had been resistant for 16 years14. Isolates collected from durum wheat in Europe and South America closely resembled Mexican isolates, suggesting the possibility of continental-scale migration of leaf rust isolates15.
To control the continual evolution of P. triticina, diligent monitoring and surveillance of virulence variations are essential. Utilizing commercial cultivars with leaf rust-resistant genes is considered the most effective, economically viable, and environmentally safer approach. Genetic studies and understanding the function and heritability of genes controlling resistance play a crucial role in breeding techniques. The inheritance of leaf rust resistance has been elucidated in various wheat cultivars16. Two primary breeding strategies for producing leaf rust-resistant cultivars involve pyramiding of major resistance genes (Lr genes) conferring complete resistance and accumulation of minor resistance genes conferring partial resistance17. According to Mapuranga, et al.18 over 100 LR resistance (Lr) genes and 80 YR resistance (Yr) genes have been identified. Effective management of wheat leaf rust relies heavily on breeding resistant cultivars, which requires comprehensive knowledge of the genetic mechanisms underlying resistance at a genome-wide level19. A total of 102 Chinese winter wheat cultivars and advanced lines were evaluated for seedling resistance to 24 Puccinia triticina pathotypes and for field-based slow rusting across two cropping seasons. Fourteen resistance genes Lr1, Lr2a, Lr3bg, Lr3ka, Lr14a, Lr16, Lr17a, Lr18, Lr20, Lr23, Lr24, Lr26, Lr34, and LrZH84 were postulated, with Lr26 being the most prevalent, while 37 genotypes lacked known resistance genes. Fourteen lines consistently exhibited slow rusting, highlighting valuable germplasm for breeding leaf rust–resistant wheat20. A total of 250 common winter wheat breeding lines were evaluated for resistance to Puccinia triticina under field and greenhouse conditions in Bulgaria. Twenty lines exhibited varying levels of resistance, including race-specific, race non-specific, and slow-rusting types. Lines demonstrating stable resistance represent valuable germplasm for breeding wheat cultivars with durable leaf rust resistance21.
A comprehensive review of documented history indicates that resistance controlled by a single gene has rapidly broken down with the emergence of novel virulent races. Consequently, breeders have shifted their focus towards partial resistance, deeming it more sustainable than hypersensitive resistance22. Various wheat cultivars have been identified to possess partial resistance to leaf rust22,23,24. Studies have reported the inheritance of partial resistance to leaf rust, ranging from polygenic to oligogenic, with moderate narrow-sense heritability25. The partial resistance of wheat to leaf rust is characterized by an extended latent period, reduced frequency of infections, and smaller pustule size. The incubation period is considered a critical component of partial resistance in cereal rusts26,27.
The development of diseases in plants may be a gradual process; nevertheless, it is of utmost importance to promptly diagnose and manage them. Symptoms usually appear as changes in the structure or function of plant tissues, such as variations in color, texture, and general appearance. Early detection and swift action are essential to reduce damage and control the spread of the disease28. Precise detection and quantification of visual symptoms are essential for both research and therapeutic purposes, as accurate assessment helps in formulating effective treatment strategies and understanding disease progression29. Severity, often measured as the percentage of symptomatic area, is a key factor for assessing treatment outcomes and forecasting crop damage29. A multitude of studies in the existing body of research have focused on using machine learning methods to forecast the severity of diseases30,31,32. Machine learning (ML), defined as the capacity of computers to acquire knowledge from data and generate predictions, offers a significant advantage in addressing complex issues, such as the identification and control of plant diseases33,34. ML techniques can analyze large datasets to identify patterns and correlations that may not be apparent through traditional methods, making it a powerful tool in agricultural research and practice35. Despite the advancements in using ML for disease severity prediction, there is a notable gap in the literature regarding the assessment of product losses following the identification of disease severity. Furthermore, there is a lack of comprehensive studies that examine the effects of host-pathogen interactions on the physiological processes and yield of the plant. Additionally, no studies have explored machine learning approaches from a broad perspective, incorporating multiple variables and complex interactions. This highlights the need for further research to develop integrated models that can not only predict disease severity but also provide insights into the broader impacts on crop yield and plant health.
Limited data are available on the inheritance of leaf rust resistance in Iran, with previous studies primarily focusing on cultivar resistance under greenhouse conditions. Therefore, the objective of this study was twofold: first, to assess the resistance of selected bread and durum wheat genotypes to P. triticina isolates at the seedling stage; and second, to integrate these resistance responses with agro-morphological and yield traits obtained under field conditions to develop and evaluate machine learning models capable of predicting the impact of leaf rust severity on wheat growth and productivity.
Results
Response of differential wheat genotypes and virulence comparison of P. triticina isolates
The response of 55 differential wheat genotypes to 9 studied isolates of Puccinia triticina was used. Notably, plants carrying the Lr19 gene exhibited resistance to all fungal isolates, while those with Lr22a, Lr34, and Lr37 genes were susceptible to all isolates. Genotypes carrying Lr28 and Lr46 genes demonstrated resistance to all isolates except for susceptibility to Ahvaz isolates (FSTPR) and Mashhad Toroq (MFHPJ), respectively. Among the studied genes, Lr2a, Lr35, Lr28, and Lr10 + displayed virulence, with Lr2a and Lr28 genes having an 11% virulence frequency, Lr35 gene with a 19% virulence frequency, and Lr10 + gene with a 22% virulence frequency. Despite their virulence, these genes could serve as effective sources of resistance if complete plant resistance is confirmed. Plants carrying the Lr9 gene exhibited resistance to Mashhad Toroq, Ahvaz (FSTPR), East Azerbaijan, Ardebil, Dezful, Neyshabur, and tetraploid isolates. However, they were susceptible to Ahvaz (FSRRS) and Karaj gene bank isolates. For the Lr10 + gene, East Azerbaijan, Ardebil, Dezful, Neyshabur, and Tetraploid isolates were avirulent, along with isolates from Ahvaz. Pavon 76 demonstrated avirulence against Ahvaz, East Azerbaijan, Ardebil, Dezful, Neyshabur, Karaj Gene Bank, and tetraploid isolates. Additionally, Ahvaz regional isolates (FSRRS), Karaj gene bank, East Azerbaijan, Dezful, Neyshabur, and Tetraploid isolates were avirulent for Gatcher, Anahuac 75, and Babax 1 genotypes. Noroeste was found to be resistant to Mashhad Toroq, Ahvaz, Karaj Gene Bank, East Azerbaijan, Neyshabur, and Tetraploid isolates. The Lr25 gene displayed resistance to Mashhad Toroq, Ahvaz, Karaj, Ardebil, Dezful, and tetraploid isolates. Similarly, the Parula genotype showed resistance to Mashhad Toroq, Ahvaz, Karaj Gene Bank, Neyshabur, and tetraploid isolates. These findings provide valuable insights into the interaction between different wheat genotypes and P. triticina isolates, contributing to our understanding of resistance patterns.
Plants carrying the Lr25, Lr32, and Lr23 + genes demonstrated resistance to 5 isolates, while those carrying Lr2c, Lr10, Lr14a, Lr17, and Lr36 genes exhibited resistance to 4 isolates. Additionally, plants with Lr1, Lr3bg, and Lr29 genes displayed resistance to 3 isolates from different regions. However, the frequency of virulence for other genes is relatively high, and their utilization should consider their virulence or avirulence in various regions of the country. For instance, Lr12, Lr14b, Lr15, Lr18, Lr20, Lr26, and Lr35 genes exhibited resistance to only two isolates, with virulence observed in other isolates. The Karaj Gene Bank (BRTHS) isolate has the lowest number of virulence factors, while the Ardebil isolate (PKTSS) possesses the highest number of virulence factors or the most virulence genes. Isolates exhibited differences in virulence or avirulence for one or more genes. Consequently, the Ardebil isolate, with a 78% frequency of virulence genes on the studied differential genotypes, displayed the highest virulence. In contrast, the Karaj Gene Bank isolate, with 50.9%, exhibited the lowest virulence. Various isolates demonstrated virulence against several studied genes, with the degree of virulence closely aligned among these isolates, emphasizing differences in one or more genes. The Ardebil isolate (PKTSS), with 43 virulence factors, had the highest virulence, while the Karaj Gene Bank (BRTHS) isolate, with 28 factors, had the lowest virulence factors or the lowest virulence.
Reaction of wheat genotypes and virulence comparison of P. triticina isolates
The outcomes of the data analysis of variance revealed significant differences among the isolates and genotypes, with a significance level of p ≤ 0.01% (Table 1). This analysis further indicated a significant interaction between genotypes and isolates (p ≤ 0.01%), suggesting a specific interaction influenced by genetic variations in the host genotypes. The observed distinctions emphasize the importance of understanding the specific interactions between wheat genotypes and P. triticina isolates, shedding light on the potential genetic factors contributing to these differences.
The study investigated the response of 49 wheat genotypes to 9 isolates of Puccinia triticina. Notably, certain genotypes exhibited significant resistance to multiple isolates. The genotypes from Italy (P.S. No4), Shabrang, Chamran2, Mehregan, Shosh, and Gonbad demonstrated remarkable resistance to all 9 isolates. Additionally, the Karkhe genotype and the landrace from Kermanshah (Kc-3642) displayed race-specific resistance against 5 isolates, while Yavaros and Behrang genotypes exhibited race-specific resistance against seven isolates. However, it is noteworthy that some genotypes, including the susceptible control cultivar (Boolani), and landraces from Austeria (Wc-46224), Morvarid, Sirvan, Baharan, Alvand, Chamran, and Sorkhtokhm, were found to be susceptible to all leaf rust isolates. Specifically, 16.3% of genotypes (8 genotypes) were susceptible to all fungal isolates, 12.2% (6 genotypes) demonstrated complete resistance, and 71.4% (35 genotypes) exhibited race-specific resistance. The assessment of the virulence pattern of P. triticina isolates on wheat genotypes revealed significant differences among isolates, with none of them being virulent or avirulent on all genotypes. The Mashhad Toroq isolates (MFHPJ) displayed the highest virulence at 83.3%, while the Dezful isolate (PGRQS) exhibited the lowest virulence at 59.1%. These findings underscore the diverse interactions between wheat genotypes and P. triticina isolates, providing valuable insights into resistance and susceptibility patterns.
Heatmap clustering analysis
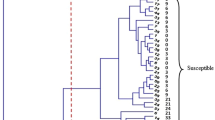
The response of differential genotypes to isolates is illustrated in Fig. 1, where two distinct clusters are observed (Group A and group B). The heat map clearly illustrates the distinct difference between the 19 genotypes in group A and the genotypes in group B, owing to their robust resistance against the 9 isolates used. Group A is partitioned into two subgroups, namely Group C and Group D. The genotypes in group C showed better resistance against 9 isolates compared to the genotypes in group D. These genotypes are TRANSFER/6*TC, TC*6/WEBSTER, TC*7/TR, CS2D-2 M, and TC*7/TRANSEC, respectively (Fig. 1). The genotypes possess disease resistance genes Lr9, Lr2a, Lr19, Lr28, and Lr25, accordingly. Furthermore, it was found that the CS2D-2 M genotype, which has the resistance gene known as Lr28, was able to effectively demonstrate resistance in six (Isolate 4, 7, 1, 8, 6, 2, and 3) out of nine isolates.
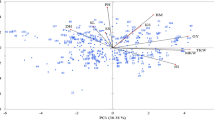
Furthermore, to shed light on the percentage of variation, a correlation-based biplot analysis was used. This study revealed the nature of the relationship between differential genotypes and isolates, as well as the level of the variation that exists between them. For instance, the C and D groups that we observed in the heatmap graphic (Fig. 1) are significantly divided from the isolates. Additionally, there were two components that were found to have eigenvalues that were larger than 1. As can be seen in Fig. 2, the overall variance is 54.13%, which is accounted for by the combined contribution of two components, namely F1 (38.82%) and F2 (15.31%).
Heatmap clustering of differential wheat genotypes using WARD method.
The superior genotypes against leaf rust caused by P. triticina isolates.
To visually represent, analyze the relationship, and provide a clear understanding of the morphological observations considered for the different genotypes, we conducted heatmap clustering (Fig. 3) and biplot (Fig. 4). Cluster analysis separated a total of 49 genotypes into two groups, designated as Group A and Group B. Group A genotypes exhibited a higher level of resistance to nine isolates compared to those in Group B. Subgroup C consisted of the most susceptible genotypes (Morvarid, Alavnd, Sirvan, Baharan, Chamran, Sorkhtokhm, and Wc-4624) in group B, including the control genotype (Boolani). These genotypes exhibit a low infection type and a high incubation period. Group D and Group E are the two subgroups that are part of Group A. Genotypes belonging to subgroup D exhibited a higher level of resistance against all isolates compared to genotypes belonging to subgroup E. The genotypes (Yavaros, Behrang, Gonbad, P.S. No4, Chamran2, Shabrang, Mehregan, and Shosh) grouped together in subgroup D exhibited a higher level of resistance, particularly against Isolate 2, Isolate 5, Isolate 7, and Isolate 9. These genotypes display a high infection type and a low incubation period.
To elucidate the percentage of variation, a correlation-based biplot analysis was used to unveil the nature of the link and extent of variation between genotypes and isolates. For instance, the D and E groups that we discovered in the heatmap picture (Fig. 3) are distinctly segregated from the isolates. The G49 (Boolani) genotype, which serves as a control, is closely associated with the isolates and hence is the genotype that is most impacted by the disease in comparison to other genotypes. Furthermore, two components with eigenvalues greater than 1 were detected. The combined contribution of two components, F1 (54.02%) and F2 (9.92%), accounts for a total variance of 63.94% as shown in Fig. 2.
Heatmap clustering of wheat genotypes and isolates using WARD method.
Biplot analysis of morphological observations of wheat genotypes.
The other objective of this study was to determine whether there is a connection between two separate variable groups of disease resistance, namely genotypes and differential genotypes. Both Figs. 3 and 4 provide the results of differential genotypes’ resistance tests against nine distinct isolates. Moreover, it was found that the genotypes Yavaros, Behrang, Gonbad, P.S. No4, Chamran2, Shabrang, Mehregan, and Shosh, which are shown in Figs. 3 and 4, were much more efficient against these nine isolates. It is possible that the genes that belong to differential genotypes are shared by genotypes Yavaros, Behrang, Gonbad, P.S.No4, Chamran2, Shabrang, Mehregan, and Shosh, which are shown in Figs. 3 and 4. As a result of the fact that the resistance genes (Lr9, Lr2a, Lr19, Lr28, and Lr25) of five distinct genotypes (TRANSFER/6*TC, TC*6/WEBSTER, TC*7/TR, CS2D-2 M, and TC*7/TRANSEC) that we analyzed among the differential genotypes (Figs. 1 and 2) by employing nine different isolates could also be associated to the Yavaros, Behrang, Gonbad, P.S.No4, Chamran2, Shabrang, Mehregan, and Shosh genotypes.
Agro-morphological results and machine learning (ML) analysis
Table 2 presents descriptive data related to the explanatory and outcome factors in the research that aimed to predict morphological and yield findings from the leaf rust responses of wheat genotypes for each Puccinia triticina strain.
Table 3 presents the detailed results of the ML analysis. In the results of ML analysis for train data, the XGBoost algorithm consistently showed superior performance in all studied variables, including DHE, DMA, PH, PED, PEDE, LL, LW, SL, NOS, SW, NOGS, GW, TKW, and GY, compared to other ML models (SVM, RF, GP, and MARS).
For the algorithms trained with train data, their prediction performances were evaluated using test data. The GP algorithm exhibited strong predictive performance for the variables DHE, DMA, PH, PED, PEDE, LL, LW, SL, and NOS, as evidenced by the low RMSE, MAPE, and MAD values as well as the high R2 values. For the DHE variable, the GP model achieved an RMSE of 2.191, indicating a low level of error between the predicted and actual values. The MAPE of 0.459 reflects a high accuracy in percentage terms, while the MAD of 0.897 underscores the consistency of the predictions. The R2 value of 0.949 suggests that the model explains a substantial portion of the variance in DHE. For DMA, the GP model also performed well, with an RMSE of 2.299 and a MAPE of 0.423. The MAD value was 0.957, further indicating the model’s precision. The R2 value of 0.901 demonstrates that the GP model accounts for a large proportion of the variance in DMA. For the PH variable, the GP model produced an RMSE of 2.594, with a MAPE of 0.755. The MAD stood at 0.910, highlighting the model’s reliable predictions. The R2 value of 0.958 confirms the model’s strong explanatory power for this variable. Regarding the PED variable, the GP model’s performance was notable, with an RMSE of 3.027 and a MAPE of 2.872. The MAD was 1.368, which is relatively low, suggesting the model’s predictions were consistent. The R2 value of 0.839 indicates a good fit of the model to the data. For the PEDE variable, the GP model yielded an RMSE of 2.707, a MAPE of 6.555, and a MAD of 1.453. These metrics reflect a moderate level of accuracy and error, with an R2 value of 0.821, indicating that the model captured a substantial portion of the variance in this trait. For the LL variable, the GP model demonstrated strong predictive performance, with an RMSE of 14.695 and a MAPE of 2.861. The MAD was 8.198, reflecting consistent predictions, and the R2 value of 0.917 suggests a very good fit to the observed data. For the LW variable, the GP model exhibited excellent performance, as shown by an RMSE of 0.879, a MAPE of 1.794, and a MAD of 0.381. The R2 value of 0.916 confirms that the model accurately captures the variance in LW. For the SL variable, the GP model produced an RMSE of 0.560 and a MAPE of 2.282, with a MAD of 0.254, indicating precise and consistent predictions. The R2 value of 0.966 further underscores the model’s strong predictive capability for this variable. Lastly, for the NOS variable, the GP model achieved an RMSE of 0.941, a MAPE of 2.077, and a MAD of 0.486, demonstrating high accuracy. The R2 value of 0.943 indicates that the model effectively explains the variance in NOS, making it a reliable predictor for this trait.
The performance of the MARS algorithm on the SW, NOGS, and GW variables revealed notable outcomes. For the SW variable, the MARS model showed a RMSE of 0.492, indicating a relatively low level of prediction error. The MAPE stood at 7.858, which was a reasonable error rate, suggesting that the model’s predictions were fairly accurate. Additionally, the MAD for SW was 0.258, further reinforcing the model’s precision. The R² was 0.849, demonstrated that the model explained approximately 85% of the variability in the SW data. For the NOGS variable, the MARS algorithm yielded an RMSE of 4.161, while higher than that for SW, still indicated a decent predictive capability. The MAPE was 6.403, signifying a moderate prediction error, and the MAD was 3.096, reflecting the model’s absolute prediction accuracy. The R² value for NOGS was 0.910, revealing that the model captured 91% of the variance in the NOGS data, indicating a strong predictive performance. Finally, the GW variable results showed an RMSE of 0.265, highlighting a very low prediction error, which was ideal for precise modeling. The MAPE was 8.383, slightly higher than those for SW and NOGS, but still within an acceptable range. The MAD for GW was 0.154, demonstrated the model’s strong ability to predict with minimal deviation. The R² for GW was 0.923, indicating that the MARS algorithm accounts for 92.3% of the variability in the GW data, underscoring its effectiveness in modeling this variable.
The XGBoost algorithm’s performance for the TKW and GY variables demonstrated strong predictive capabilities with distinct outcomes for each variable. For the TKW variable, the XGBoost model had a RMSE of 2.711, an MAPE of 3.357, and a MAD of 1.513. The R² was 0.949. For the GY variable, the XGBoost model had an RMSE of 10.899, a MAPE of 9.281, and a MAD of 5.602. The R² value was 0.919.
Figure 5 presents scatter plots comparing the predicted values from the models that predicted each variable with the highest accuracy and the true values of the variables. The R² values, which indicate the goodness of fit for each model, are displayed in each plot along with the corresponding regression equations. The findings showed that the models performed well on the variables, the R² values were consistently high and the predicted values closely matched the actual values. This showed that the GP, MARS and XGBoost models were highly effective in predicting the outcomes of the analyzed features.
Scatter plot for variables predicted with high accuracy by GP, MARS and XGBoost models.
Discussion
Leaf rust disease, caused by Puccinia triticina Eriksson, was first reported in Iran by Esfandiari in 19481. Utilizing genetic resistance in improved cultivars is a crucial method for effectively controlling leaf rust disease and minimizing its associated losses. The statistical analysis revealed that Morvarid, Sirvan, Baharan, Alvand, Chamran, Sorkhtokhm, and Wc-4624 genotypes were susceptible to all leaf rust isolates in this study, recommending against the cultivation of these genotypes. Dadrezaei, et al. 36 reported that Alvand and Chamran cultivars showed susceptibility to 4 and 3 isolates, respectively. Seedling infection plays a significant role in yield loss, with 1000-seed weight identified as the most crucial reduced component by Kolmer37. Leaf rust resistance genes can be categorized into two groups: seedling resistance genes and adult plant resistance genes19. Seedling resistance refers to resistance from the emergence of the first leaf of the seedling until the end stage of growth, and it can be hypersensitive, immune, or moderate resistance38. This type of resistance is usually monogenic with a large gene effect39. Yavaros, Behrang, P.S. No4, Shabrang, Chamran 2, Mehregan, Shosh, and Gonabad genotypes exhibited resistance, possibly attributed to seedling resistance genes against leaf rust in these genotypes40. In a study by Mohajervatan, et al.41, Sorkhtokhm and Gonabad genotypes also demonstrated susceptibility and resistance to all leaf rust isolates, respectively.
In this study, we assessed wheat genotypes with prevalent leaf rust races in the country under greenhouse conditions to explore the resistance genetics and virulence of leaf rust in interaction with wheat. After doing our research, we concluded that the differential genotypes TRANSFER/6*TC, TC*6/WEBSTER, TC*7/TR, CS2D-2 M, and TC*7/TRANSEC, which had resistance genes Lr9, Lr2a, Lr19, Lr28, and Lr25, demonstrated a greater level of resistance against the nine isolates that were used. In their study, Tomar and Menon42 reported that some Lr genes, including Lr9, Lr19, Lr24, Lr28, and Lr32, provide full protection against leaf rust pathotypes in wheat-growing areas of Asia. Kokhmetova, et al.43 reported that the Lr25 and Lr24 genes had the highest efficacy in Kazakhstan. They observed that isogenic lines carrying these genes were immune to both pathotypes. The virulence of the Lr19 gene is scarcely reported worldwide44. Despite its effectiveness in most regions of the world45, the international use of this gene has been limited due to its association with the gene responsible for producing a yellow color in wheat46,47. Torabi, et al.48 conducted a study on isolates from various parts of Iran, reporting the absence of virulence in lines carrying the Lr19 gene. Our findings align with their results, as well as those of Wang, et al. 48 and Draz, et al.49. Afshari, et al.50 also noted a virulence in plants harboring Lr9, Lr19, Lr25, and Lr28 genes, consistent with Afshari5 observations on the a virulence of the Lr19 gene and Rafiei, et al.51 identification of virulence factors for the Lr22a gene. Comparing our study with previous reports, it can be inferred that variations in the genetic structure of the pathogen have been highly limited in the last eight years. Worldwide, virulence for the Lr9 gene is uncommon, primarily reported in the United States, Brazil, Argentina, Italy, Africa, Pakistan, and Mexico. The presence of a virulence factor for this gene in Iran has not been confirmed, at least from 1993 (the initiation of research in this field at the Karaj Research Institute) to the present. In our study, the genotype CS2D-2 M, which has the Lr28 gene, exhibited the greatest resistance to all isolates, except for the Milan isolate (MSNPB) and the Neyshabur isolate (TSTPD). Nemati, et al.52 reported similar findings of virulence for the Lr9 gene. It’s crucial to note that the Lr9 gene is considered a race-specific resistance gene. Huerta-Espino, et al. 2 observed that Pakistani pathotypes were able to overcome this gene, indicating the need to combine it with other genes that confer resistance to leaf rust. Therefore, widespread, and solitary cultivation of genotypes with this gene may lead to the development of virulent races and potentially render the resistance ineffective in the short term. It is important to note that the genes studied in this research are seedling resistance genes44. In our study, the Lr10 gene exhibited resistance to four isolates, aligning with the findings reported by Wang, et al.48. Torabi, et al.53 reported resistance to the Lr13 gene in the Gachsaran region and for the Lr34 gene in the Ahvaz region, consistent with our results. Additionally, Draz, et al.49 observed resistance to the Lr34 gene in another study. Regarding the Lr34, Lr16, Lr13, and Lr1 genes, our results are in line with Afshari5 study, which identified virulence on lines containing these genes. Torabi, et al.53 and Rahmatov, et al.54 also reported virulence on lines containing the Lr16 gene in their respective studies. This consistency in findings across studies contributes to a more comprehensive understanding of the resistance and virulence patterns associated with these specific genes in different regions.
Identifying the influence that plant diseases have on agricultural output and yield loss has been a topic of study for a considerable amount of time55. There are several published models that attempt to establish a connection between the incidence or severity of disease and crop yield56. Especially in the context of pathogen-plant health investigations, machine learning approaches have not been investigated when viewed from a holistic viewpoint, taking into consideration a multiplicity of factors and complicated interactions. The application of a variety of machine learning (ML) algorithm models in the field of plant biology is a relatively new and expanding area of investigation that is designed to predict and optimize variables within complex biological systems29,57. ML is a paradigm of data science that addresses complex challenges across a variety of scientific disciplines58,59. This method transcends traditional unidirectional analyses, enabling a precise interpretation and nuanced comprehension of the results33,34. In the literature, many studies have focused on predicting disease severity using machine learning techniques. Chung, et al. 60 used the support vector machine (SVM) algorithm to categorize rice seedlings as either healthy or infected with Bakanae disease. This method proved to be more time-efficient compared to visually inspecting the seedlings. Moshou, et al.61 explored the creation of a system that automatically distinguished between water-stressed Septoria tritici infected and healthy winter wheat canopies. They used optical multisensor fusion in conjunction with a least squares (LS)-support vector machine (SVM) classifier. Ahmad, et al.62 concentrated on developing an automated system using machine learning methodologies to identify the presence of cotton leaf curl virus (CLCuV) in cotton plants. Their findings indicate that the random forest classifier had the most efficacy in identifying Cotton Leaf Curl Virus (CLCuV), with an accuracy rate of 87.542%. Feng, et al.63 used remote sensing data to assess the magnitude of powdery mildew disease in wheat. To do this, the researchers used four distinct spectrum transformation techniques (mean centralization, multivariate scattering correction, standard normal variate transform, and original reflectivity) and developed a prediction model by using machine learning algorithms (PLSR: partial least square regression, SVR: support vector regression, and RF: random forest regression) to interpret the resulting data. The researchers reported that they achieved the optimal prediction model by combining the RF algorithm with the mean centralization preprocessing approach. In addition, they reported the R² value of the RF model was found to be in the range of 0.849–0.852.
Our research focused on the yield loss in wheat that was caused by the severity of the disease produced by various strains. This was done in connection with the ML method, which pushed the link between the severity of the disease and the morphology of the plant one step further. Through the process of comparing several models, we were able to determine that the XGBoost, GP, and MARS models were able to accurately predict morphological data with a remarkable accuracy of over 90%. It has also been underlined in several studies that have used these models that provide a high level of prediction accuracy when compared to other models57,64,65,66,67. Once identifying the severity of the disease, a model can be developed by concentrating on plant yield losses, as shown in our research. With accurate predictions, a corrective intervention can be made to yield losses in the early phases of the plant.
Conclusions
The study on the pathotypes and physiological races of leaf rust disease in wheat, along with its distribution in Iran, highlights the substantial diversity within the population of leaf rust pathogens across the country. The findings emphasize that only a limited number of resistance genes exhibit effective resistance to all pathotypes, attributing this to the high diversity of races in Iran. The overreliance on single-gene resistances, such as Lr22a, Lr34, and Lr37, can lead to the selection of rust pathotypes capable of overcoming these specific resistances. These studies indicated that the cultivation of genotypes carrying a single resistance gene can result in the increased selection and prevalence of virulence for that gene. Recognizing the limitations of relying solely on single-gene resistance, the study suggests that the sustainability of resistance in a genotype can be enhanced by combining or “pyramiding” two or more resistance genes. This approach aims to reduce the risk of breakdown due to pathogen mutations or the emergence of new races in the region. As well as collaboration with international research centers such as CIMMYT and ICARDA, it is recommended to leverage practical achievements and advance cereal breeding programs. We advise the wheat breeders to be encouraged to incorporate gradual rust resistance genes alongside race-specific resistance genes in breeding programs. In dealing with genes for which virulence has been observed, the study underscores the importance of better managing their use in breeding programs. Previous research has indicated a lack of variety in resistant genes and low diversity in Iranian wheat germplasm used in cereal research. To address this, the study advocates for continuous and extensive research to identify rust-resistance genes in the rich collection of wheat germplasm and its relatives in Iran. Previous studies have focused on investigating disease severity using ML algorithms and have generally not investigated the direct impact of the disease on crop health and yield loss. However, this study represents a new approach by not only examining the severity of the disease but also predicting its morphological and yield-related effects on wheat genotypes. Based on extensive analysis using various machine learning algorithms, the GP, MARS and XGBoost models were shown in this study to exhibit superior predictive capabilities across a range of variables. The models demonstrated excellent accuracy for prediction with high R² values, reflecting their robustness in modeling complex biological data. These findings underscore the potential of machine learning models to accurately predict key agronomic traits and thus provide valuable tools for advancing crop breeding programs. The results of this study highlight the effectiveness of these models in capturing complex relationships between variables that can significantly improve decision-making in agricultural research and development.
Materials and methods
Herbal materials
In this study, the seeds utilized encompassed three distinct types of genetic materials. These included 13 cultivars of bread wheat (Triticum aestivum L., 2n = 6x = 42), 25 genotypes of durum wheat (Triticum durum L., 2n = 4x = 28), comprising both Iranian varieties and landraces from various countries. Among these, 8 accessions originate from 8 countries: Austria, Afghanistan, Portugal, France, Türkiye, Argentina, Bulgaria, and Italy. Additionally, 10 cultivars of durum wheat, one susceptible control (Boolani), and 55 wheat differential genotypes are included (Tables 4 and 5). The foreign durum wheat accessions and seeds for the 49 wheat genotypes were obtained from the gene bank of the Cereal Research Department of the Seed and Plant Research Improvement Institute in Karaj, Iran. All experimental procedures and field studies involving plants, including the collection of plant materials, were conducted in accordance with applicable institutional, national, and international guidelines and regulations.
Seedling assessment
Leaf rust-infected samples were gathered from multiple wheat fields displaying symptoms across diverse regions of Iran during the spring of 2016. These samples were then dispatched to the Seed and Plant Research Improvement Institute, specifically the Cereal Pathology Unit in Karaj, for further analysis. Following purification, nine isolates were chosen for greenhouse experiments, originating from Ahvaz (two isolates), Ardebil, Neyshabur, Mashhad Toroq, Dezful, Tetraploid, East Azerbaijan, and Karaj Gene Bank (Table 6). Each collected sample was individually inoculated onto the first leaf of seedlings belonging to the susceptible cultivar Boolani. This process is aimed at spore reproduction and purification. The initial purification involved the use of the single-pustule technique. Subsequently, the leaves, cut to an appropriate size, were placed on moistened filter paper in petri dishes for 48 h at 4 °C in a refrigerator. After inducing sporulation, the generated spores were inoculated onto Boolani cultivar leaves using an ear cleaner. The inoculated seedlings were kept in a darkroom for 24 h at 18 ± 2 °C with saturated humidity and were then transferred to a greenhouse with conditions set at 24 ± 2 °C and a relative humidity greater than 50%. Approximately 12–14 days later, the single-pustule spores were individually removed using the single-pustule technique and inoculated on susceptible wheat leaves. For further reproduction, the purified spores were inoculated and cultured again on the susceptible cultivar. In this process, the collected spores were mixed with talc powder or Saltrol mineral oil at a spore to oil volume ratio of 1:20 and inoculated on many Boolani pots. After collecting, the spores were placed into a petri dish and stored in a desiccator containing silica gel dehumidifier for 48 h. Once the spores attained a moisture content of approximately 20–30%, they were transferred to plastic vials and stored in the freezer at -20 °C. When needed, the plastic vials were immersed in distilled water for 5 min to reactivate the spores stored in the freezer. Spores from each isolate were individually cultured on the first leaf of seedlings, with the seeds of each cultivar sown in pots containing a mixture of sterilized soil, sand, and rotten animal manure (1:1:1). The inoculation with rust spores occurred 8 days after snowing.
In the experiments, a total of 55 differential genotypes and 49 wheat cultivars were inoculated, and each experiment was conducted in three replications. To ensure adequate infection, a susceptible cultivar was placed next to the pots in each replication. Infection types were recorded 12–14 days after inoculation following the criteria outlined by McIntosh et al.44 on a scale ranging from 0 to 4. In this scale, infection types 0–2 were categorized as a virulence or resistant (R), while infection types 3–4 were categorized as virulence or susceptible (S). To assess the virulence frequency in leaf rust isolates, the percentage of virulence factors in each isolate relative to the total number of resistance genes was calculated. Additionally, the percentage of isolates with the virulence gene was determined to calculate the frequency of virulence for each resistance gene. This comprehensive analysis aimed to provide insights into the prevalence and distribution of virulence factors and resistance genes in the studied leaf rust isolates.
Agro-morphological evaluations
In this study, 48 genotypes from the Grain Department of the Seedling and Seed Breeding Research Institute, originating from various regions of the country and sourced from other countries, were selected under irrigated conditions at the Karaj station. Three spikes from each genotype were selected and evaluated through an observational study with four control cultivars (Vee/Nac, Soisson, Dena, Zardak) planted in 144 rows, excluding the controls. These genotypes were chosen from a preliminary screening of local Iranian genotypes, as well as advanced durum wheat lines from the National Plant Gene Bank of Iran and the Cereals Research Department of the Plant and Seed Breeding Research Institute. Each pure line was sown in a 0.5-meter row. Three spikes collected from each genotype in the previous year were shown separately but adjacent to one another. The sowing was conducted systematically, with four durum wheat control lines placed before and after every six durum lines. The germplasm under study was cultivated at the research farm of the Seedling and Seed Breeding Institute located in Karaj (51°10’ E longitude, 35°48’ N latitude, and 1321 m above sea level). Throughout the growing season, these lines were assessed for various traits, including days to heading (DHE), days to maturity (DMA), plant height (PH), peduncle length (PL), peduncle extrusion length (PEDe), flag leaf length (LL), flag leaf width (LW), spike length (SL), spikelet number per spike (NoS), spike weight (SW), number of grain per spike (NoGS), grain weight per spike (GW), thousand grain weight (TKW) and grain yield (GY). In this context, the study not only examined seedling-stage resistance under greenhouse conditions but also evaluated agro-morphological and yield-related traits from field-grown adult plants, ensuring that both early resistance responses and adult-plant performance were considered when training and testing the machine learning models.
Statistical analysis
Data entry and initial descriptive statistics were performed using Microsoft Excel (Microsoft Corp., USA). XLSTAT (Addinsoft, version 2023.1.3) was employed for conducting ANOVA and post-hoc comparisons due to its user-friendly interface and reliability for agricultural datasets. PAST (version 3.14) was used for multivariate analyses such as cluster analysis and biplot visualization, as it provides robust tools for morphological and yield trait data. Prior to performing ANOVA, assumption checking was carried out: normality of residuals was tested using the Shapiro–Wilk test, and homogeneity of variances was tested using Levene’s test. The results of these tests confirmed that the assumptions were met, and therefore, ANOVA was conducted.
Process of modeling via the use of machine learning
In this study, machine learning algorithms were employed specifically to predict and model the impact of leaf rust infection on wheat morphology and yield under disease pressure. As input variables, we used the seedling-stage resistance responses of wheat genotypes to nine Puccinia triticina isolates obtained from the greenhouse evaluations. As output variables, we included the agro-morphological and yield traits recorded from field-grown plants, which reflect the adult-plant performance under varying levels of disease severity. In this way, the ML models were designed not to classify infection types, but to integrate early resistance responses with field-based performance data in order to quantify the broader impact of leaf rust on wheat productivity. Under the scope of the research, six ML algorithms were employed: Support vector machine (SVM), random forest (RF), extreme gradient boosting (XGBoost), Gaussian process (GP), and multivariate adaptive regression spline (MARS). The evaluation of these methods was conducted by considering the maximum R² (Eq. 1) values and the minimum root mean square error (Eq. 2), mean absolute percentage error (Eq. 3), and mean absolute deviation (Eq. 4) criteria. Before doing the study, the dataset was divided into two parts: 70% for training data and 30% for testing data. This division enabled the algorithms to be trained using the training data and then assessed using the testing data to make accurate predictions. The optimum hyperparameters for each machine learning model were obtained using Eq. 5. The use of the Grid Search Cross-Validation (GCV) optimization method facilitated this achievement. The computer language R (version 4.3.2) was used to compute performance metrics and implement machine learning methods57,65.
The symbol n symbolizes the quantity of the training and testing sample in the dataset. The actual value to be measured is denoted as \(\:{y}_{i}\), while the predicted value is represented as \(\:{y}_{ip}\). The mean of the measured values is denoted as \(\:\stackrel{-}{y}\). The penalty function that is associated with the complexity of the model that includes λ terms is denoted by M(λ).
Support vector machine (SVM)
Support vector machines algorithm tries to find the most appropriate hyperplane to separate data points between two classes in classification problems68. For this purpose, the SVM algorithm provides the maximum margin. Equation 1 is the basic mathematical formulation of SVM. In the formula, \(\:y\) refers to the predicted value (class label). \(\:{a}_{i}\) and \(\:{a}_{i}^{*}\) are lagrange multipliers, these multipliers correspond to support vectors. \(\:k\left(x,\left.{x}_{i}\right)\right.\) is the kernel function, which represents the inner product of the data points \(\:x\) and \(\:{x}_{i}\) in the feature space. \(\:{x}_{i}\) is the \(\:i\) -th data point in the training data. \(\:b\) is a constant term (bias). This formula solves nonlinear classification problems without transforming data into a high-dimensional space using the kernel trick (Eq. 6)68.
Random forest (RF)
One method of ensemble learning is known as the random forest, which forms a learning model by integrating several decision trees into a single structure. During the training process, each tree is trained on a randomly chosen section of the dataset and makes use of a randomly selected subset of different characteristics69. Because of this, the projections from each tree are combined to get more accuracy and consistency in the predictions. The RF approach, which may be used in our investigation because the desired output data is comprised of numerical data, may be formulated as follows:
Here, \(\:T\) denotes the total quantity of trees. The forecasts generated by each decision tree, represented as \(\:{h}_{t}\left(x\right)\), are the continuous values outputted by the t-th tree. To get the final forecast \(\:\widehat{y}\), the average of all the forecasts produced by the trees is used.
Extreme gradient boosting (XGBoost)
The XGBoost model is a variant of the gradient boosting decision tree algorithm that has been optimized for both speed and performance70. The optimization is achieved by using a gradient boosting framework. The ability of XGBoost to learn from mistakes and decrease error rates via iterative processes is a notable attribute that distinguishes it from other optimization approaches71. The equation below is the mathematical formula that calculates the output of the XGBoost model:
The symbol \(\:{\widehat{y}}_{i}\) is used to represent the value that the model predicts for the i-th observation. The total number of decision trees in existence is denoted by the letter \(\:K\). The k-th decision tree is represented by the symbol \(\:{f}_{k}\). The feature vector \(\:{x}_{i}\) corresponds to the i-th observation.
Gaussian process (GP)
The random variable distribution is described using the Gaussian probability density function72. The GP approach was considered in the investigation, as indicated by the following statistical formula:
The average function of the Gaussian process is represented by the symbol \(\:m\left(x\right)\), while the kernel function is represented by \(\:k\left(x\right.,{x}^{{\prime\:}})\). The covariance function (kernel) is employed to determine the similarities between data points. The mean function is frequently assigned a constant value, such as zero or the average of the training dataset72.
Multivariate adaptive regression spline (MARS)
MARS is a non-parametric regression method that estimates the associations between input variables and output variable(s) by fitting piecewise linear regressions. It is especially advantageous when working with intricate datasets where conventional linear regression models may not provide satisfactory results (Okumuş et al., 2024).
The basis function is represented by the mathematical expression \(\:{h}_{\text{km}}\left({X}_{v(k,m)}\right)\), where \(\:{\beta\:}_{0}\) is a constant (intercept) and denotes the coefficient of basis functions. The index of the independent variable in the m-th component of the kth product is denoted by \(\:v(k,m)\). The order of interaction is restricted by the parameter \(\:{K}_{m}\).
Data availability
All data supporting the conclusions of this article are included in this article.
References
Ghasemzadeh, E., Afshari, F., Khodarahmi, M. & Bihamta, M. R. Study on the genetics of resistance to leaf rust in some advanced lines of wheat at seedling stage. Seed Plant. 19, 281–294 (2010).
Huerta-Espino, J. et al. Global status of wheat leaf rust caused by puccinia triticina. Euphytica 179, 143–160 (2011).
Dadrezaei, S. T., Tabatabai, N., Lakzadeh, I., Jafarnezhad, A. & Afshari, F. Hassan Bayat, Z. Evaluation of tolerance to leaf rust disease in some selected bread wheat genotypes. Appl. Entomol. Phytopathol. 86, 29–40 (2018).
Bolton, M. D., Kolmer, J. A. & Garvin, D. F. Wheat leaf rust caused by puccinia triticina. Mol. Plant Pathol. 9, 563–575 (2008).
Afshari, F. in Proceedings of 11th international wheat genet symposium 709–711.
Roelfs, A. P., Singh, R. P. & Saari, E. Rust Diseases of Wheat: Concepts and Methods of Disease Management (Cimmyt, 1992).
Qi, A., Li, X., Shi, L., Liu, D. & Li, Z. Identification of a leaf rust resistance gene in the Chinese wheat line LB0288. Czech J. Genet. Plant. Breed. 51, 43–49 (2015).
Harrington, J., Reitz, L., Worzella, W. & Ausemus, E. Summary of genetic studies in hexaploid and tetraploid wheats. J. Am. Soc. Agron. 38, 1082–1099 (1946).
Johnston, C. & Heyne, E. Wichita wheat back-cross lines for differential hosts in identifying physiologic races of Puccinia recondita. Phytopathology 54, 385–388 (1964).
Johnson, R. Jul. Durable resistance to yellow (stripe) rust in wheat and its implications in plant breeding. Breeding Strategies for Resistance to the Rusts of Wheat. El Batan, Mexico (Mexico) (1987).
Waterhouse, W. in Proceedings of the Linnean Society of New South Wales. Linnean Society of New South Wales 209–258 (1929).
Kolmer, J., Long, D. & Hughes, M. Physiologic specialization of Puccinia triticina on wheat in the united States in 2006. Plant Dis. 92, 1241–1246 (2008).
Park, R. Pathogenic specialisation of Puccinia recondita f. sp. tritici in Australia and new Zealand in 1990 and 1991. Australas. Plant Pathol. 25, 12–17 (1996).
Singh, R., Huerta-Espino, J., Pfeiffer, W. & Figueroa-Lopez, P. Occurrence and impact of a new leaf rust race on durum wheat in Northwestern Mexico from 2001 to 2003. Plant Dis. 88, 703–708 (2004).
Ordoñez, M. & Kolmer, J. Virulence phenotypes of a worldwide collection of Puccinia triticina from durum wheat. Phytopathology 97, 344–351 (2007).
Sharma, P. & Saini, R. Genetics of durable resistance to leaf rust in bread wheat cultivars Capelle Desprez and Pari 73. J. Phytology. 3, 10–15 (2011).
Messmer, M. M. et al. Genetic analysis of durable leaf rust resistance in winter wheat. Theor. Appl. Genet. 100, 419–431 (2000).
Mapuranga, J. et al. Harnessing genetic resistance to rusts in wheat and integrated rust management methods to develop more durable resistant cultivars. Front. Plant Sci. 13, 951095 (2022).
Ren, X. et al. Genetics of resistance to leaf rust in wheat: an overview in a genome-wide level. Sustainability 15, 3247 (2023).
Li, Z. F. et al. Seedling and slow rusting resistance to leaf rust in Chinese wheat cultivars. Plant Dis. 94, 45–53 (2010).
Ivanova, V. Seedling and adult plant resistance to leaf rust in some Bulgarian common wheat lines. Vavilov J. Genet. Breed. 27, 447 (2023).
Broers, L. Influence of development stage and host genotype on three components of partial resistance to leaf rust in spring wheat. Euphytica 44, 187–195 (1989).
Broers, L. & Jacobs, T. The inheritance of host plant effect on latency period of wheat leaf rust in spring wheat. II: number of segregating factors and evidence for trangressive segregation in F3 and F5 generations. Euphytica 44, 207–214 (1989).
Van der Gaag, D. & Jacobs, T. Inheritance of host plant effect on latent period of wheat leaf rust in single-seed descent F8 lines. Euphytica 97, 67–72 (1997).
Das, M. K., Rajaram, S., Mundt, C. C. & Kronstad, W. E. Inheritance of slow-rusting resistance to leaf rust in wheat. Crop Sci. 32, 1452–1456 (1992).
Shaner, G. & Finney, R. E. New sources of slow leaf rusting resistance in wheat. Phytopathology 70, 1183–1186 (1980).
Shaner, G., Roberts, J. & Finney, R. Culture of Puccinia recondita virulent to the wheat cultivar transfer. Plant. Disease Report. 56, 827–830 (1972).
Allen-Sader, C. et al. C.A. An early warning system to predict and mitigate wheat rust diseases in Ethiopia. Environ. Res. Lett. 14, 1150042019 (2019).
Bock, C., Poole, G., Parker, P. & Gottwald, T. Plant disease severity estimated visually, by digital photography and image analysis, and by hyperspectral imaging. CRC. Crit. Rev. Plant Sci. 29, 59–107 (2010).
Khan, H., Haq, I. U., Munsif, M., Mustaqeem, Khan, S. U. & Lee, M. Y. Automated wheat diseases classification framework using advanced machine learning technique. Agriculture 12, 1226 (2022).
Odilbekov, F., Armoniené, R., Henriksson, T. & Chawade, A. Proximal phenotyping and machine learning methods to identify Septoria tritici blotch disease symptoms in wheat. Front. Plant Sci. 9, 685 (2018).
Singh, R., Krishnan, P., Singh, V. K. & Das, B. Estimation of yellow rust severity in wheat using visible and thermal imaging coupled with machine learning models. Geocarto Int. 38, 2160831 (2023).
Türkoğlu, A. et al. Modeling callus induction and regeneration in hypocotyl explant of fodder pea (Pisum sativum var. Arvense L.) using machine learning algorithm method. Agronomy 13, 2835 (2023).
Türkoğlu, A. et al. Machine learning analysis of the impact of silver nitrate and silver nanoparticles on wheat (Triticum aestivum L.): callus induction, plant regeneration, and DNA methylation. Plants 12, 4151 (2023).
Das, S., Pattanayak, S. & Behera, P. R. Application of machine learning: a recent advancement in plant diseases detection. J. Plant. Prot. Res. 122-, 135–122 (2022).
Dadrezaei, S., Afshari, F. & Patpour, M. Evaluation of phenotypic resistance to rusts in some Iranian wheat genotypes in greenhouse and field conditions. Seed Plant. Improv. J. 31, 531–546 (2015).
Kolmer, J. A. Tracking wheat rust on a continental scale. Curr. Opin. Plant. Biol. 8, 441–449 (2005).
Chen, W., Hu, C. & Zhang, S. Analysis of virulence genes of Puccinia recondita f. sp. tritici population in China. Scientia Agricultura Sinica. 25, 17–23 (1993).
Hyde, P. & Elahinia, S. The expression of adult-plant resistance in wheat to Puccinia striiformis. J. Phytopathol. 124, 31–38 (1989).
Knott, D. R. The Wheat rusts—breeding for Resistance Vol. 12, 1–199 (Springer Science & Business Media, 2012).
Mohajervatan, F., Nezhad Ghomi, N., Kalate Arabi, A. A., Dehghan, M. A. & M. & Evaluation of resistance to leaf rust in some wheat cultivars in field and greenhouse conditions. J. Crop Breed. 8, 76–70 (2017).
Tomar, S. & Menon, M. Genes for resistance to rusts and powdery mildew in wheat 152 (2001).
Kokhmetova, A. et al. Evaluation of wheat germplasm for resistance to leaf rust (Puccinia triticina) and identification of the sources of Lr resistance genes using molecular markers. Plants 10, 1484 (2021).
McIntosh, R. A., Wellings, C. R. & Park, R. F. Wheat Rusts: an Atlas of Resistance Genes (CSIRO publishing, 1995).
Martens, J. & Dyck, P. Occurrence and virulence of puccinia recondita in Canada in 1986. Can. J. Plant Pathol. 10, 268–272 (1988).
Torabi, M., Nazari, K. & Afshari, F. Genetic of pathogenicity of Puccinia recondita f. sp. tritici, the causal agent of leaf rust of wheat. Iran. J. Agric. Sci. 32, 625–635 (2001).
Xu, S. et al. Genetic mapping of the wheat leaf rust resistance gene Lr19 and development of translocation lines to break its linkage with yellow pigment. Theor. Appl. Genet. 136, 200 (2023).
Wang, J., Shi, L., Zhu, L., Li, X. & Liu, D. Genetic analysis and molecular mapping of leaf rust resistance genes in the wheat line 5R618. Czech J. Genet. Plant. Breed. 50, 262–267 (2014).
Draz, I. S., Abou-Elseoud, M. S., Kamara, A. E. M. & Alaa-Eldein, O. A. E. El-Bebany, A. F. Screening of wheat genotypes for leaf rust resistance along with grain yield. Annals Agricultural Sci. 60, 29–39 (2015).
Afshari, F. et al. Monitoring of virulence factors of Puccinia triticina Eriksson, the causal agent of wheat leaf rust in Iran during 2002–2004 (2006).
Rafiei, F., Arzani, A., Afshari, F. & Torabi, M. Characterization of leaf rust resistance genes in seedlings of wheat cultivars. Genetic Breed. 36, 19–27 (2007).
Nemati, Z., Mostowfizadeh-Ghalamfarsa, R., Dadkhodaie, A., Mehrabi, R. & Steffenson, B. J. Virulence of leaf rust physiological races in Iran from 2010 to 2017. Plant Dis. 104, 363–372 (2020).
Torabi, M. et al. Virulence genes of Puccinia recondita f. sp. tritici, the causal agent of wheat leaf rust in some regions of Iran during 1995–1999 (2003).
Rahmatov, M. et al. Characterization of stem, Stripe and leaf rust resistance in Tajik bread wheat accessions. Euphytica 215, 1–22 (2019).
Madden, L., Hughes, G. & Irwin, M. Coupling disease-progress-curve and time-of-infection functions for predicting yield loss of crops. Phytopathology 90, 788–800 (2000).
Cerda, R. et al. Primary and secondary yield losses caused by pests and diseases: assessment and modeling in coffee. PloS One. 12, e0169133 (2017).
Demirel, F. et al. Prediction of grain yield in wheat by CHAID and MARS algorithms analyses. Agronomy 13, 1438 (2023).
Keçeci, M., Gökmen, F., Usul, M., Koca, C. & Uygur, V. Prediction of cadmium content using machine learning methods. Environ. Earth Sci. 83, 362 (2024).
Okumuş, O. et al. Using Machine Learning Algorithms to Investigate the Impact of Temperature Treatment and Salt Stress on Four Forage Peas (Pisum sativum var. arvense L.). Horticulturae 10, 656 (2024).
Chung, C. L. et al. Detecting Bakanae disease in rice seedlings by machine vision. Comput. Electron. Agric. 121, 404–411 (2016).
Moshou, D., Pantazi, X. E., Kateris, D. & Gravalos, I. Water stress detection based on optical multisensor fusion with a least squares support vector machine classifier. Biosyst. Eng. 117, 15–22 (2014).
Ahmad, N., Qadri, S., Akhtar, N. & Nawaz, S. A. Multifeature analysis to detect cotton leaf curl virus. J. Comput. Biomedical Inf. 7, 215–223 (2024).
Feng, Z. H. et al. Hyperspectral monitoring of powdery mildew disease severity in wheat based on machine learning. Front. Plant Sci. 13, 828454 (2022).
Bozkurt, T., İnan, S., Dündar, İ., Isak, M. A. & Şimşek, Ö. Optimizing the in vitro propagation of tea plants: A comparative analysis of machine learning models. Horticulturae 10, 721 (2024).
Demirel, F., Uğur, R., Popescu, G. C., Demirel, S. & Popescu, M. Usage of machine learning algorithms for Establishing an effective protocol for the in vitro micropropagation ability of black chokeberry (Aronia melanocarpa (Michx.) Elliott). Horticulturae 9, 1112 (2023).
Şimşek, Ö. Machine learning offers insights into the impact of in vitro drought stress on strawberry cultivars. Agriculture 14, 294 (2024).
Şimşek, Ö. et al. Optimizing micropropagation and rooting protocols for diverse lavender genotypes: A synergistic approach integrating machine learning techniques. Horticulturae 10, 52 (2024).
Noble, W. S. What is a support vector machine? Nat. Biotechnol. 24, 1565–1567 (2006).
Breiman, L. Random forests. Mach. Learn. 45, 5–32 (2001).
Rasmussen, C. E. In Summer School on Machine Learning 63–71 (Springer, 2003).
Chen, T. & Guestrin, C. in Proceedings of the 22nd acm sigkdd international conference on knowledge discovery and data mining. 785–794 (2016).
Rasmussen, C. E. in Seventh Valencia international meeting, dedicated to Dennis V. Lindley 651–659 (Oxford University Press).
Acknowledgements
The present manuscript was financially supported by Seed and Plant Research Improvement Institute, Karaj, Iran.
Funding
Open access funding provided by the Scientific and Technological Research Council of Türkiye (TUBİTAK).
Author information
Authors and Affiliations
Contributions
Conceptualization, T.B., R.M., and B.B.; methodology, T.B., R.M., M.A.S., and F.S.A.; software R. M., T.B., M.A.S., F.S.A., B.B. and J.B.; validation, M.A.S., A.T., K.H., J.B., and B.B.; formal analysis, T.B., R.M., M.A.S., F.S.A., F.D., B.B. and J.B.; investigation, T.B., R. M., M.A.S., F.S.A., B.B. and J.B.; resources, T.B., R. M., M.A.S., F.S.A., B.B. and J.B.; data curation, M.A.S. A.T., K.H., J.B., and B.B.; writing—original draft preparation, M.A.S., A.T., and B.B.; writing—review and editing, T.B., R. M., M.A.S., F.S.A., B.B., K.H., F.D., A.T. and J.B.; visualization, T.B., R.M., M.A.S., supervision, M.A.S.; project administration, M.A.S.; funding acquisition, M.A.S., J.B. and B.B. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Bakhshi, T., Mehrabi, R., Sarbarzeh, M.A. et al. Monitoring wheat leaf rust severity using machine learning techniques. Sci Rep 15, 42951 (2025). https://doi.org/10.1038/s41598-025-24407-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-24407-8