Abstract
Esophageal squamous cell carcinoma (ESCC) is an aggressive malignancy with limited therapies and poor prognosis. While exhausted T cell (Tex) heterogeneity under chronic antigen stimulation is recognized, the role of Tex subsets in shaping the tumor microenvironment (TME) and influencing ESCC outcomes remains unclear. We integrated single-cell RNA sequencing, TCR sequencing, microarray, and bulk RNA sequencing to dissect Tex cells. We identified a novel proliferative Tex (prolif Tex) subset across discovery and validation cohorts, with high infiltration correlating significantly with an improved patient survival. Pseudotime trajectories suggested prolif Tex originated from Tex cells, while TCR sequencing revealed clonal expansion and shared receptor repertoires between Tex and prolif Tex. Neoadjuvant chemo-immunotherapy reduced prolif Tex proportions and differentiation potential in ESCC samples. A machine learning-derived prognostic model was developed using prolif Tex subset-specific genes and validated via qRT-PCR using in-house ESCC samples. Experimental validation confirmed differential expression of candidate genes in tumors versus normal tissues, with high ESCO2 expression linked to prolonged survival. Our findings unveil prolif Tex cells as a novel TME subset in ESCC, where its abundance predicts favorable outcomes. The prolif Tex-based prognostic model demonstrates strong prognostic value and validated hub genes offer potential biomarkers and therapeutic targets. Our findings underscore the potential of prolif Tex cells as a biomarker and therapeutic target in ESCC.
Similar content being viewed by others
Introduction
Esophageal cancer (EC) is the eighth most common cancer and the sixth leading cause of cancer-related mortality worldwide. It ranks eighth globally in terms of diagnosed cases (604,100 cases, 3.1%) and sixth in terms of deaths (544,076 cases, 5.5%)1. EC is generally classified into two subtypes: esophageal adenocarcinoma (EAC) and esophageal squamous cell carcinoma (ESCC)2. In North America and Northern Europe, EAC is diagnosed twice as frequently as ESCC, whereas in East Asia, ESCC diagnoses outnumber EAC by more than tenfold3. The high prevalence of ESCC is largely attributed to unhealthy lifestyle factors, such as smoking and excessive alcohol consumption, and dietary habits that include the consumption of hot food, hot drinks, and pickled vegetables4. In the early stages of ESCC, patients do not exhibit obvious symptoms, and as a result, the majority are diagnosed at advanced stages (III-IV), when surgical resection is no longer possible, and non-invasive treatments are typically employed4,5. Currently, the primary treatment options in China include chemotherapy and radiotherapy. However, the 5-year survival rate for ESCC patients remains low, at approximately 30%6,7. Neoadjuvant chemo-immunotherapy (NAT) followed by surgery has been established as the cornerstone of current clinical guidelines in ESCC8. However, recurrence and metastasis rates after surgery remain high, with approximately 42.5%−47.8% of patients experiencing treatment failure within a median follow-up period of 12–18 months9. Notably, while this enhanced treatment approach increases pathological complete response rates, it also significantly raises the incidence of grade ≥ III adverse events10. Despite advancements in treatment modalities, including targeted therapies and immunotherapy11, the clinical outcomes for ESCC patients remain unsatisfactory2, which can be attributed to the complex tumor microenvironment12,13. Thus, defining the characteristics of the ESCC tumor microenvironment is crucial for identifying new targets or optimizing treatment strategies.
The tumor microenvironment (TME) is a complex and highly structured ecosystem consisting of tumor cells surrounded by various nonmalignant cells. This highly vascularized extracellular matrix system includes nontumor cells such as immune cells, cancer-associated fibroblasts (CAFs), endothelial cells, mesenchymal cells, etc. These nonmalignant cells can be directly or indirectly influenced by tumor cells, leading to alterations in their original functions or the secretion of chemokines and exosomes. These changes, in turn, contribute to cancer cell occurrence, development, and migration, while also impairing the normal functioning of the immune system12,13. T cells are a critical cellular component of the TME. T cell exhaustion is a broad term that describes the dysfunctional state of T cells resulting from persistent antigenic stimulation. The term was initially introduced in the context of chronic viral infections and has recently gained significant attention in studies on the TME14. At the molecular level, Tex cells are characterized by the high expression of inhibitory receptors, reduced cytotoxic activity, and diminished proliferative potential. Pathologically, exhaustion is associated with a significant decline in the cytokine secretion capacity of effector T cells (Teff). These cells represent the terminally differentiated state of Teff cells, undergoing abnormal differentiation due to prolonged antigenic stimulation14,15. Some autoreactive T cell subsets have also been reported to exhibit exhaustion-like phenotypes in autoimmune conditions. While this process may contribute to maintain immune tolerance by restraining excessive immune responses, it also compromises immune surveillance against pathogens and tumors, thereby facilitating chronic infections and tumor immune escape16. Emerging evidence demonstrates dynamic transitions between Tex subtypes under specific conditions. These cells maintain functional flexibility by balancing self-renewal and differentiation into effector states, directly strengthening antitumor responses17. T cell exhaustion serves as a protective mechanism, controlling overactivation and functional decline in T cells persistently exposed to chronic antigens18. It defines a distinct differentiation state that diverges from naive, effector, and memory CD8 + T cell lineages. Recent studies have demonstrated progressive substantial heterogeneity within Tex populations, with precursor-derived subsets emerging as exhaustion evolves, revealing layered heterogeneity in their functional states and differentiation trajectories18. While Tex cells are generally regarded as T cells that express high levels of immune checkpoint receptors and contribute to immune escape in the tumor microenvironment19, emerging evidence suggests that these cells can lead to favorable clinical outcomes in certain contexts20,21. However, the heterogeneity of Tex cells in ESCC and their prognostic implications remain insufficiently delineated, necessitating the application of emerging multi-omics approaches to resolve this biological enigma.
Next-generation sequencing has revolutionized transcriptome analysis, with high-throughput transcriptome sequencing (bulk RNA-seq) capturing averaged gene expression trends across cell populations but obscuring rare subpopulations. In contrast, single-cell RNA sequencing (scRNA-seq) provides single-cell resolution to uncover cellular heterogeneity and interactions, offering deeper insights into tumor complexity to solve the critical limitation that bulk RNA-seq has not been able to address22. Existing studies on ESCC employing bulk RNA sequencing and microarray analysis have delineated population-level transcriptomic patterns23,24. But they are limited in detecting rare Tex subsets due to their inability to handle cellular heterogeneity. Conversely, scRNA-seq facilitates high-resolution mapping of Tex precursor-progenitor hierarchies and functional states25,26,27. However, its application in ESCC research is limited by small cohort sizes and a lack of clinical survival data22,25. Moreover, current multi-omics studies predominantly focus on epithelial cells, leaving Tex-specific regulatory networks and their clinical implications insufficiently explored28,29,30. Therefore, integrating data from bulk RNA-seq, microarray, scRNA-seq, and T cell receptor sequencing (TCR-seq) allows for a more comprehensive understanding to identify the key subset of Tex and core genes that play a role in ESCC. Further integration with patient survival data enables the construction of prognostic models to predict the disease progression of ESCC patients and identify potential therapeutic targets.
In this study, we employed a comprehensive multi-omics approach to investigate the characteristics of prolif Tex cells in the ESCC TME. Through integrated analyses of scRNA-seq, TCR-seq, microarray data, and RNA-seq in ESCC, a novel subset of exhausted T cells termed prolif Tex cells was identified. This subset was found to be highly infiltrated and associated with improved patient survival. The prolif Tex cells originated from conventional Tex and exhibited enhanced differentiation potential. Using machine learning, we developed a prolif Tex-based risk model, validated its prognostic value across multiple cohorts, and confirmed the expression of hub genes through qRT-PCR. These findings highlight the potential of prolif Tex cells as both prognostic biomarkers and therapeutic targets.
Materials and methods
Study design
The study design is presented in Fig. 1.
Flow chart illustrating the study design.
Data collection
This study utilized several public datasets for analysis. The scRNA-seq data for ESCC were obtained from the Gene Expression Omnibus (GEO, http://www.ncbi.nlm.nih.gov/geo): GSE160269 (n = 64)25, GSE145370 (n = 28)31, and the Genome Sequence Archive (GSA, https://ngdc.cncb.ac.cn/gsa-human): OMIX005710 (n = 46)32. The RNA-seq data for bulk ESCC samples were obtained from The Cancer Genome Atlas (TCGA), including 86 tumor samples and 13 normal samples. Microarray data were obtained from the GEO dataset: GSE53622 (n = 60), GSE53624 (n = 119) and GSE53625 (n = 179)24.
Single-cell data processing
Quality control of scRNA-seq data was performed using Seurat (version 4.1.0)33. Cells with fewer than 500 or more than 7,500 detected genes, or those with over 20% mitochondrial gene expression, were excluded. Genes expressed in fewer than three cells were also removed. Data normalization was performed using the NormalizeData function, and doublets were identified and excluded using the DoubletFinder package (version 2.0.4) with an expected doublet rate of 8%. Batch effects across datasets were corrected using the Harmony R package (version 1.2.0). Dimensionality reduction was conducted via Principal Component Analysis (PCA), followed by Uniform Manifold Approximation and Projection (UMAP) for visualization34.
RNA-seq data processing
RNA-seq gene expression data were log2-transformed and standardized to z-scores. The TCGA-ESCC cohort was used for training, and the clinical data were integrated with the gene expression data to form the final matrix for survival analysis. Samples with survival times of less than 50 days were excluded to ensure data reliability.
Cell annotation
Cell clustering and differential expression analysis were performed using the Seurat R package. Clustering was conducted via the FindNeighbors and FindClusters functions (dim = 30, resolution = 1.0). Differentially expressed genes (DEGs) were identified using the FindAllMarkers function. Cell annotations were based on canonical cell marker genes. Chromosomal copy number variation (CNV) scores were evaluated using the inferCNV package (version 1.18.1)35. T cells were reclustered using the same Seurat pipeline, and cell annotations for subsets of T cells were conducted using differentially expressed genes of each cluster and canonical cell marker genes.
T cell pathway enrichment analysis
T cell proliferation, exhaustion, and cytotoxicity scores were calculated using gene sets from Cheng et al.36. Cell cycle gene sets were obtained from the MsigDB database (https://www.gsea-msigdb.org/gsea/msigdb). Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses were conducted using the enrichGO and enrichKEGG functions within the clusterProfiler R package (version 4.10.1)37.
T cell receptor analysis
TCR clonality and expansion were assessed for the GSE160269 dataset using the scRepertoire R package (version 2.2.1)38. The Morisita overlap index was used to quantify TCR similarity across T cell subsets39.
Trajectory analysis of T cells
T cell pseudotime trajectories were modeled using the Monocle2 R package (version 2.30.1)40. The DDRTree algorithm was applied to embed cells in a lower-dimensional space. Genes with sufficient expression levels were selected after filtering, and dimensionality reduction was performed using the reduceDimension function. Cells were ordered using the orderCells function to construct pseudotime trajectories. The CytoTRACE2 R package (version 1.0.0)41 was used to assess differentiation potential.
T cell infiltration and survival analysis
The single-sample Gene Set Enrichment Analysis (ssGSEA) method in the GSVA R package (version 1.50.5)42 was applied to assess T cell subtype infiltration in bulk RNA-seq samples. Gene Set Enrichment Analysis (GSEA) was also conducted to examine cell cycle gene enrichment. Kaplan-Meier plots were generated using the survfit function from the survival R package (version 3.7.0) and the ggsurvplot function from the survminer (version 0.4.9) R package. Survival differences were evaluated using log-rank tests. The optimal cutoff for classifying high- and low-risk groups was determined using the ‘surv_cutpoint’ function. The TCGA-ESCC dataset served as training set, while the GEO datasets were used for independent validation.
Construction of Prolif Tex-associated prognostic model
DEGs were identified between normal and tumor samples in the TCGA-ESCC dataset using the Limma package (version 3.58.1) and among T cell clusters using the FindAllMarkers function, with thresholds of adjusted P < 0.05 and |avg_log2 fold change| > log2(1.2). Univariate Cox regression analysis was performed to identify prognostic genes. Overlapping DEGs of prolif Tex cells across GSE160269, GSE145370, and OMIX005710 datasets were used to construct the prognostic model. A prolif Tex-based risk signature was developed using machine learning, selecting the optimal algorithm from Ridge, Least Absolute Shrinkage and Selection Operator (Lasso), and Elastic Net (Enet). Risk groups were stratified, and survival outcomes were evaluated via Kaplan-Meier analysis.
Nomogram construction based on the risk signature
Univariate and multivariate Cox regression were used to analyze risk signatures and clinicopathological features. A nomogram was constructed to predict overall survival (OS) in ESCC patients, incorporating clinical features and risk scores. Variables with P < 0.05 in multivariate analysis were included. Calibration curves were generated to assess the nomogram’s accuracy and clinical relevance.
Immune infiltration analysis
The correlation between the risk signature and the tumor immune microenvironment (TIME) were analyzed using the CIBERSORT, ESTIMATE, and TcellSI algorithms. Stromal, immune, and ESTIMATE scores were calculated using the ESTIMATE R package (version 1.0.13)43. CIBERSORT (version 0.1.0)44 was used to estimate the proportions of 28 immune cell subtypes, and correlations between model genes and immune scores were analyzed to assess immune function. The T cell states scores were calculated using the TCellSI R package (version 0.1.0)45.
Immune therapy response analysis
The association between the risk model and immune checkpoint inhibitor (ICI) response was evaluated using transcriptomic and clinical data from ICI-treated patients. Data from the IMvigor210 cohort (anti-PD-L1 therapy), GSE78220 (melanoma, anti-PD-1 therapy), GSE67501 (renal cell carcinoma, anti-PD-1 therapy), and GSE165252 (ESCC, combining a PD-L1 inhibitor with neoadjuvant chemoradiotherapy) were used to assess the model’s predictive ability for therapeutic response. In all external cohorts, we applied the same prognostic model constructed from the prolif Tex-derived genes in TCGA-ESCC without incorporating clinicopathological covariates.
Drug sensitivity analysis
The Connectivity Map (CMap) database (http://www.broadinstitute.org) was used to identify potential compounds associated with the risk groups defined by the PTex model46. Specifically, differentially expressed genes between the high- and low-risk groups were used as input, and compounds predicted to reverse the high-risk transcriptional profile were prioritized. The top 50 drugs with the strongest predicted reversal effects were selected for mechanism of action (MoA) analysis. Drug sensitivity analysis was conducted using the Genomics of Drug Sensitivity in Cancer (GDSC) database47, with IC50 values predicted using the OncoPredict R package (version 1.2)48.
Quantitative RT-PCR validation
Primary tumor and adjacent normal tissue samples from 15 ESCC patients who received chemoradiotherapy were collected after surgical resection at Fudan University Shanghai Cancer Center (FUSCC). Informed consent was obtained, and the study was approved by the FUSCC Ethical Review Committee. Total RNA was extracted, and qRT-PCR was performed via SGExcel FastSYBR Mixture (Sangon) on a LightCycler 480 II System (Roche). Gene expression levels were normalized to GAPDH and calculated using the 2-ΔΔCT method. Primer sequences are listed in supplementary material 1: Table S1. Since the RNA was extracted from preserved Formalin-Fixed Paraffin-Embedded samples, the amount of available RNA was insufficient for assaying all genes of the PTex model. Therefore, six candidate genes (ESCO2, CORO1A, DBF4, NDUFB11, RAB8A, and TNFSF10) from the PTex model were selected for validation using bulk RNA extracted from paired tumor and normal tissues, as these genes have previously been reported to be associated with cancer. Additionally, it is worth noting that the use of bulk RNA to measure the expression of these six representative genes may dilute cell type-specific signals, such as ESCO2 expression derived from proliferating Tex cells.
Statistical analysis
All the statistical analyses were performed using R software (version 4.3.3). Group comparisons were performed using the Wilcoxon test or Student’s t-test, depending on the data distribution. Pearson and Spearman correlation coefficients were used for correlation analysis. Survival analysis was conducted using the Kaplan-Meier method, with survival differences evaluated by log-rank tests. Statistical significance was defined as P < 0.05, with levels of significance set at * P < 0.05, ** P < 0.01, and *** P < 0.001.
Results
Identification of prolif Tex cells in ESCC by scRNA-seq
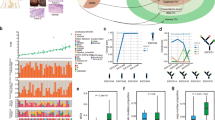
To explore the cellular heterogeneity within the TME of ESCC, we processed and integrated scRNA-seq data from the GSE160269 cohort, obtaining a total of 208,125 cells. Based on DEGs in each cluster, we annotated cell types and identified immune cells (CD45+), including proliferative T cells, T cells, NK cells, B cells, and myeloid cells, as well as non-immune cells (CD45-), such as epithelial cells, endothelial cells, fibroblasts, and pericytes (Fig. 2A). Proliferative T cells were characterized by the expression of canonical T cell markers (CD3D, CD3E, CD3G) and cell cycle-associated genes (MKI67, TYMS, and RRM2) (Fig. 2B,C). Proliferative T cells were found to be enriched in tumor samples compared to normal tissues (Fig. 2D). To better distinguish tumor cells from normal epithelial cells, we utilized inferCNV to analyze copy number variations (CNVs), categorizing cells with significant CNVs as tumor cells (Fig. S1A).
To provide a comprehensive characterization of T cell subtypes in ESCC, we focused on T cells and proliferative T cells, identifying multiple CD4 + T cell clusters (including CD4 T Naive, Tfh1, Tfh2, TmemCD4, and Treg) and CD8 + T cell clusters (including MAIT, TmemCD8, NKT, Teff and Tex). Treg are effective suppressors of inflammatory immune responses and are essential in all tissues to prevent destructive immunity49. NKT cells are recently discovered T cells that express both NK cell surface markers and T cell surface receptors50. Proliferative T cells were further classified into proliferative Treg, prolif Tex cells, and proliferative NKT cells (Fig. 2E). Cell cycle analysis revealed that proliferative T cells primarily entered the G2/M and S phases, indicating active proliferation (Fig. 2F). To investigate the expression variations among the T cells, we explored the expression of several marker genes associated with different cell states including Naïve, Effector, Exhaustion, and Proliferation (Fig. 2G). Both prolif Tex and prolif Treg cells exhibited high expression of proliferation markers (MKI67, UBE2C, TOP2A), and prolif Tex cells expressed high levels of exhaustion markers (LAG3, PDCD1, CTLA4), while prolif Treg exhibited high expression of CD4 cell markers (FOXP3, CD4, IL2RA). Furthermore, prolif Tex cells were divided into two primary subtypes based on the dominant expression of either CDK4 or MKI67 (Fig. 2H). Based on these distinctive expression patterns of prolif Tex cells, we further analyzed the correlation of this subset with clinical prognosis in TCGA-ESCC bulk RNA-seq data.
Single-cell atlas of ESCC cohort (GSE160269). (A) UMAP visualization of the 208,125 cells (64 ESCC samples: 60 tumor and 4 normal), colored by cell type. (B) Dot plot of the expression of marker genes for cell types defined in (A). Dot size corresponds to the percentage of cells expressing the marker gene, and dot color indicates the average expression. (C) Feature plot illustrating the distribution of selected marker genes of T cells. (D) Box plot of cell-type fractions identified in tumor and normal tissues, values are presented as mean ± SD. (E) UMAP visualization of T cells, colored by cell type. (F) Distribution of T cell cycle phases (G1, S, and G2/M). (G) Dot plot illustrating the expression levels and percentage of cells expressing selected T cell function-associated genes across different T cell clusters. (H) Kernel density estimation plot showing the distribution of MKI67 and CDK4 gene expression of T cells. (I) Infiltration scores of T cell subtypes between tumor and normal samples in the TCGA-ESCC dataset. (J) Survival analysis stratifying ESCC patients by T cell infiltration scores in the TCGA-ESCC dataset.
High expression of representative gene set from prolif Tex cells correlated with better prognosis
To investigate the functional role of prolif Tex cells within ESCC, we performed immune infiltration analysis and survival analyses. We selected the top 50 upregulated DEGs in the GSE160269 single-cell dataset as representative gene sets characterizing these subtypes. Subsequently, the ssGSEA was used to calculate the infiltration scores for T cells in the TCGA-ESCC cohort. A comparison of the infiltration scores between normal and tumor tissue revealed that Treg, prolif Treg, Tex and prolif Tex cells exhibited significantly higher infiltration scores in tumor tissue compared to normal tissues, indicating the enrichment of prolif Tex cells in ESCC (Fig. 2I). The infiltration of prolif Tex cells showed an increasing trend with tumor progression, though the difference across tumor stages was not statistically significant (Fig. S1B). To further evaluate the prognostic impact of prolif Tex cells on survival outcomes, patients were divided into high- and low-risk groups based on prolif Tex cell infiltration levels. The analysis revealed that high infiltration levels of most T cell subtypes were associated with poor patient outcomes. However, high infiltration levels of prolif Tex cells were significantly correlated with improved survival in ESCC patients (Fig. 2J). These findings suggest that prolif Tex cells might play a supportive role in antitumor immune responses within ESCC.
Functional enrichment analysis revealed high exhaustion and proliferative characteristics of prolif Tex cells
To further investigate the expression characteristics of prolif Tex cells, we performed GSEA using cell cycle gene sets from MSigDB. The analysis revealed significant enrichment of cell cycle-related genes in prolif Tex cells, which was not observed in Tex or Teff cells (Fig. 3A). GO and KEGG enrichment analyses were conducted on the DEGs of prolif Tex cells, Tex and Teff cells (Fig. 3B). These analyses demonstrated that prolif Tex cells were significantly enriched in pathways associated with cell cycle, DNA replication and microtubule-associated binding, suggesting their predominant role in tumor-associated proliferation. Furthermore, proliferation, exhaustion, and cytotoxicity scores were calculated using the gene sets provided by Cheng et al.36. The results revealed that prolif Tex cells exhibited significantly higher proliferation scores compared to non-proliferative T cells (Fig. 3C). Moreover, the exhaustion and cytotoxicity scores of prolif Tex cells and Tex cells were significantly higher than Teff cells, suggesting the high proliferative capacity and potential cytotoxic function of prolif Tex cells.
Characteristics of prolif T cells. (A) GSEA of cell cycle-associated gene sets for Teff, Tex, and prolif Tex cells. (B) GO and KEGG pathway enrichment analysis of DEGs in Teff, Tex, and prolif Tex cells. The KEGG database was used under permission from Kanehisa Laboratories (www.kegg.jp)51,52. (C) Violin plots comparing proliferation score, exhaustion score, and cytotoxicity score for Teff, Tex, and prolif Tex cells, values are presented as mean ± SD. (D) UMAP visualization of T cell receptor clonal populations. (E) Heatmap showing T cell subtype TCR overlap indices using the Morisita overlap index. (F) CytoTRACE analysis and visualization of the degree of differentiation of T cells, where higher scores indicate greater stemness. (G) Differentiation trajectory of the CD8 + T cells, colored by cell subtype (left) and pseudotime (right). Each point indicates a single cell.
Prolif Tex and Tex exhibit expanded clonal proliferation in high tumor stage samples
T cell receptor VDJ sequencing data enables the analysis of T cell clonal expansion based on homologous TCRs. Paired TCR-seq and scRNA-seq analyses revealed that CD8 + T cells (Teff cells, Tex, prolif Tex cells, and T memCD8) exhibited highly expanded TCR clones, with Tex and prolif Tex cells predominantly consisting of large and medium clones (Figs. 2E and 3D). We analyzed the clonal composition of T cell subtypes in tumor samples of different stages (Fig. S1C). Large TCR clones were nearly absent in normal tissues. However, large clones of Tex and prolif Tex cells were significantly abundant in stage II/III tumor samples compared to stage I samples, while no significant differences were observed in Teff cells. These results suggest that the clonal size of prolif Tex and Tex cells increases with tumor stage progression.
Overlapping TCR clones provide insights into associations across the T cell differentiation trajectory53. A comparison of the TCR sequences revealed a high degree of overlap among Teff cells, Tex, prolif Tex cells, and T memCD8 cells. Notably, the TCR similarity between Tex and prolif Tex cells was as high as 0.79, indicating a close lineage relationship between these subtypes (Fig. 3E). These findings highlight the pronounced clonal expansion and high TCR similarity between Tex and prolif Tex cells, emphasizing their pivotal role in the progression of ESCC.
Differentiation potential and pseudotime trajectory of prolif Tex cells
To investigate the differentiation potential of prolif Tex cells, CytoTRACE2 differentiation scores were calculated, revealing that prolif Tex cells exhibited lower differentiation plasticity, indicative of a more defined cell fate (Fig. 3F). To further explore the differentiation trajectory of prolif Tex cells, pseudotime analysis was performed based on transcriptional similarity in CD8 + T cells, focusing on Teff cells, Tex, and prolif Tex cells. The pseudotime trajectory began with the Teff cells cluster and demonstrated a gradual differentiation process, transitioning from Teff cells to Tex cells and finally to prolif Tex cells (Fig. 3G). Taken together, the prolif Tex cells exhibit a higher differentiation potential, share partial TCR similarity with the Tex cells, and are positioned at the terminally differentiated state of the differentiation trajectory originating from Teff cells and progressing toward Tex cells.
We aimed to characterize Tex subtypes in ESCC, we performed reclustering analysis (Fig. S2A). The progenitor Tex subset displayed high expression of the stem-like marker IL7R, consistent with its precursor-like features, whereas prolif Tex cells showed elevated expression of proliferation- and cell cycle–related genes (Fig. S2B). TCR analysis revealed that prolif Tex cells exhibited the greatest clonal similarity with CXCL13 + Tex cells, while their overlap with progenitor Tex was relatively low (Morisita index = 0.221) (Fig. S2C). In pseudotime analysis, prolif Tex cells were positioned at the terminal end of the Tex differentiation axis, distinct from progenitor Tex (Fig. S2D and E). These findings indicate that prolif Tex and progenitor Tex represent two transcriptionally and developmentally distinct subsets with divergent molecular features.
To further validate the functional characteristics of prolif Tex cells, the GSE145370 ESCC single-cell dataset was analyzed for independent verification. Using the same approach as GSE160269, the data were processed to identify cell clusters (Fig. S3A), followed by T cell annotation (Fig. S3B). Consistent with the findings from GSE160269, prolif Tex cells were identified, co-expressing proliferation and exhaustion markers (Fig. S3C). Based on MKI67 and CDK4 expression, prolif Tex cells were classified into two main subtypes (Fig. S3D), predominantly located in the G2/M and S phases and exhibiting higher differentiation potential scores (Fig. S3E and F). Furthermore, prolif Tex cells were predominantly positioned at the terminal stage of the CD8 + cell pseudotime trajectory (Fig. S3G). Overall, these suggest that prolif Tex cells occupy the terminal position in the developmental trajectory of the CD8 + T cells and exhibit unique differentiation potential. This conclusion was consistently supported by both our discovery and validation cohorts.
Reduced differentiation potential and decreased proportion of prolif Tex cells after NAT
To investigate the effects of NAT on the differentiation potential and abundance of prolif Tex cells within the TME, we analyzed the OMIX005710 dataset, a recent single-cell dataset of ESCC that includes samples collected before and after NAT. Using the same data processing method, we obtained 166,200 cells from OMIX005710. Cell type annotation was performed based on the differentially expressed genes in each cell cluster (Fig. 4A), with each cell type exhibiting high expression of its characteristic marker genes (Fig. 4B). Subsequently, T cells were extracted, reclustered, and reannotated, resulting in the identification of prolif Tex cells (Fig. 4C). These cells showed high expression of proliferation- and exhaustion-related genes (Fig. 4D) and were further divided into two clusters based on CDK4 and MKI67 expression patterns (Fig. 4E). The cell cycle analysis revealed that prolif Tex cells predominantly occupied the G2/M phase (Fig. 4F). Differentiation potential analysis revealed significantly reduced differentiation potential in prolif Tex cells after NAT treatment (Fig. 4G, S4A–C). Additionally, the proportion of prolif Tex cells decreased after NAT treatment, while Naive-like and Teff cells increased (Fig. 4H and S4C). Pseudotime analysis further revealed that prolif Tex cells primarily appeared at the terminal branch of the CD8 + T-cell differentiation trajectory, branching from conventional Tex cells (Fig. 4I). Notably, the differentiation pathway differed before and after treatment: Teff cells after treatment formed two branches along the main differentiation trajectory, one of which did not originate from Naive-like cells, whereas pretreatment samples exhibited only a single branch. These observations suggest that prolif Tex cells may differentiate into alternative functional states or exhibit increased sensitivity to treatment, leading to their exhaustion.
To investigate the prognostic impact of prolif Tex cells in NAT treatment, We compared their proportions across the three treatment response groups. Although patients with better responses tended to have a higher proportion of prolif Tex cells, this difference was not statistically significant (Fig. S4D). Survival analysis conducted in TCGA-ESCC cohort, based on the infiltration levels of various cell types from the OMIX005710 and GSE145370 datasets, revealed that high infiltration of prolif Tex cells was significantly associated with improved survival outcomes (Fig. S4E), consistent with findings from the GSE160269 cohort. Collectively, NAT treatment significantly reduced both the differentiation potential and the proportion of prolif Tex cells, potentially due to either their differentiation into alternative functional states or increased sensitivity to immunotherapy.
Single-cell atlas of ESCC neoadjuvant chemo-immunotherapy cohort (OMIX005710). (A) UMAP visualization of 6,679 cells (46 ESCC samples: 22 pre-treatment and 24 post-treatment), colored by cell type. (B) Dot plot of the expression of marker genes for cell types defined in (A). Dot size corresponds to the percentage of cells expressing the marker gene, and dot color indicates the average expression. (C) UMAP visualization of T cells, colored by cell type. (D) Dot plot showing the average expression of canonical marker genes across T cells. (E) Kernel density estimation plot showing the distribution of MKI67 and CDK4 gene expression in T cells. (F) Distribution of cell cycle phases of T cells. (G) Differentiation potential scores in proliferative Tex cells before and after NAT treatment. (H) Alluvial plot (left) and boxplots (right) comparing the proportions of prolif Tex cells in samples before and after NAT. (I) Differentiation trajectory of T cells according to pseudotime and sample groups (before and after neoadjuvant chemo-immunotherapy), colored by cell subtype (top left), pseudotime (top right), and treatment group (bottom). Each point indicates a single cell.
Construction of prognostic model based on prolif Tex cells
Using the previously analyzed three single-cell datasets, we identified shared DEGs across prolif Tex cells (supplementary material 2: Table S2). Through univariate Cox regression analysis, 42 genes significantly correlated with prognosis were selected (Fig. 5A). Utilizing these prognostic DEGs, machine learning techniques were employed to construct a comprehensive predictive model. Ultimately, a prognostic model using the Enet (α = 0.4) was chosen (Fig. 5B and C). This model consisted of nineteen genes, including 12 risk genes and 7 protective genes (Fig. 5D). Risk scores for each sample were calculated and patients were divided into high- and low- risk groups. Kaplan-Meier survival analysis revealed that patients in the high-risk group had significantly poorer prognoses than those in the low-risk group (Fig. 5E). The distributions of patient survival status and risk scores in the TCGA and GEO cohorts are shown in Fig. S5A. To further validate the model, we analyzed the expression distribution of the 19 genes in the model within the single-cell dataset GSE160269. The results indicated that the majority of these genes were highly expressed in the prolif Tex cells (Fig. 5F). Additionally, ssGSEA based on the DEGs of prolif Tex cells was applied to evaluate the infiltration of prolif Tex cells in TCGA-ESCC cohort. The findings revealed that patients in the low-risk group exhibited greater infiltration levels of prolif Tex cells (Fig. 5G), aligning with our previous findings that the high infiltration of prolif Tex cells is associated with improved survival outcomes in ESCC patients.
Construction and evaluation of a prognostic risk model. (A) Volcano plot of 42 prognosis-correlated genes obtained via univariate Cox regression analysis (P < 0.05), labeled with genes retained in the final model. (B) Trajectories of variables for lambda selection (optimal λ = 0.061). (C) Distributions of independent variables at optimal λ. (D) Enet regression coefficients for the 19 genes in the risk signature. (E) Kaplan-Meier survival analysis of overall survival based on the prognostic model in the TCGA-ESCC, GSE53622, GSE53624, and GSE53625 cohorts (TCGA-ESCC: n = 86, GSE53622: n = 60, GSE53624: n = 119, GSE53625: n = 179). (F) Expression of the 19 prognostic genes across cell types in scRNA-seq dataset GSE160269. (G) ssGSEA-based infiltration scores of high- and low-risk groups in TCGA-ESCC. (H, I) Univariate (left, RiskScore HR = 6.536, P = 1.15 × 10−8) and multivariate (right, RiskScore HR = 7.263, P = 1.45 × 10−8) cox regression analysis based on the risk score and clinicopathological features. (J) Nomogram integrating gender, T-stage, N-stage, clinical stage, and risk score for OS prediction. (K) Calibration curves for 1-year and 2-year OS predictions.
Independent risk factors investigation and nomogram construction
To identify independent prognostic factors and develop a predictive nomogram, we integrated clinicopathological features with risk scores. Both univariate and multivariate Cox regression analyses confirmed that risk scores were an independent prognostic factor (P < 0.001) (Fig. 5H and I). Using multivariate Cox regression, a predictive nomogram was constructed by incorporating risk scores along with gender, T-stage, and N-stage (Fig. 5J). The calibration plot results demonstrated that the nomogram exhibited strong predictive accuracy for actual survival outcomes (Fig. 5K).
Immune infiltration characteristics of the prognostic model
To further elucidate the immunological mechanisms underlying the prognostic significance of our model, we analyzed immune infiltration features by comparing immune and stromal cell compositions between the high- and low-risk groups in the TCGA-ESCC cohort. CIBERSORT showed that high-risk patients showed increased proportion of CD8 + T cells, M1 and M2 macrophages, and Treg cells, while the low-risk group showed increased proportions of resting NK cells and resting memory CD4 + T cells (Fig. 6A). Further analysis of immune and stromal cell infiltration within the tumor microenvironment revealed that the ESTIMATE scores, immune scores, and stromal scores were significantly elevated in the high-risk group compared to the low-risk group (Fig. 6B). Despite higher tumor purity in the low-risk group, the levels of immune and stromal infiltration were decreased, suggesting pronounced immunosuppressive characteristics in the TME of the low-risk group. Additionally, CORO1A and TNFSF10 were positively correlated with immune scores, while ESCO2 and DBF4 showed negative correlations (Fig. S5B).
Using the TcellSI R package, we analyzed T cell-related immune state and found that, except for the resting state, T cell functional activity was significantly higher in the high-risk group compared to the low-risk group (Fig. 6C). Patients in the high-risk group exhibited significantly higher levels of immune and stromal cell infiltration (Fig. 6D). Additionally, protective genes such as NUDT11 and DBF4 were inversely correlated with stromal scores, immune scores, and ESTIMATE scores (Fig. 6F). Risk genes including CORO1A, PSMB8, and TNFSF10 were significantly associated with higher infiltration of most immune cells, whereas protective genes such as DBF4, ESCO2, and RBBPB were correlated with decreased infiltration levels (Fig. 6E and S5C). Taken together, patients in the high-risk group exhibited enhanced immune infiltration and activity, characterized by elevated immune scores, contrasting with the immunosuppressive tumor microenvironment observed in low-risk group tumors. These findings suggest that the low-risk patients in TCGA-ESCC may be more likely to benefit from immunotherapy, potentially improving survival time.
Immune infiltration analysis. (A) Distribution of 28 immune cell types in high- and low-risk groups (TCGA-ESCC). (B) ESTIMATE, stromal, and immune scores between risk groups (TCGA-ESCC). (C) T cell state scores calculated via TcellSI algorithm (TCGA-ESCC). (D) Heatmap of immune cell infiltration in the TME of ESCC. (E-F) Correlations of prognostic model signatures with immune-related cell proportions, Immune, Stromal, and ESTIMATE scores.
Prediction of immunotherapy response based on risk signature
To evaluate the ability of our prognostic model to predict immunotherapy response, we analyzed two independent datasets, GSE78220 and IMvigor210. The IMvigor210 cohort, comprising 348 melanoma patients treated with anti-PD-L1 antibodies, was classified into complete responses (CR), partial responses (PR), stable disease (SD), and progressive disease (PD) groups based on treatment outcomes. Similarly, the GSE78220 dataset includes patients with melanoma who received anti-PD-1 immune checkpoint inhibition therapy. In both datasets, patients in the high-risk score groups exhibited significantly worse OS. Moreover, the proportion of PD/SD patients in the high-risk group was significantly higher than in the low-risk group, whereas CR/PR rate was notably higher in the low-risk group. These findings suggest that the high-risk group is associated with a poorer response to immune checkpoint inhibition (Fig. 7A and B). To validate these results, we further analyzed two additional independent datasets, GSE67501 and GSE165252 (Fig. 7C and D). Consistent with previous findings, CR/PR patients in both cohorts presented significantly low risk scores, whereas PD/SD responders were more likely to have high risk scores. Collectively, these results indicate that immunotherapy outcomes were significantly worse in the high-risk group than in the low-risk group.
Prediction of immunotherapy response via the risk signature across multiple public cohorts. (A) Kaplan-Meier plot and treatment response distribution in GSE78220. (B) Kaplan-Meier plot and treatment response distribution in IMvigor210. (C) Immunotherapy response rates by risk group in GSE67501. (D) Immunotherapy response rates by risk group in GSE165252.
Drug sensitivity analysis
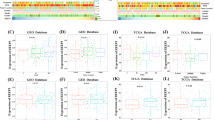
Considering the prognostic differences between the high- and low-risk groups, we aimed to identify potential therapeutic drugs by conducting a mechanism of action (MoA) analysis through the CMap database. Fig. S6A highlights the top 50 drugs predicted to be the most effective for ESCC treatment, along with their associated pathways. The OncoPredict package in R was used to construct a ridge regression model and to predict drug sensitivity, generating IC50 values for 198 drugs. A comparison of IC50 values between high- and low-risk groups revealed significant differences in drug efficacy, with 34 drugs showing statistically significant variations (Fig. S6B).
Further correlation analysis between IC50 values and risk scores revealed significant associations between genes involved in cell cycle regulation and drug resistance or sensitivity (Fig. S6C). The risk score was negatively correlated with the IC50 values of most drugs, indicating their higher effectiveness in high-risk groups. Furthermore, based on the average IC50 values across samples, we identified the top 20 candidate drugs with the lowest IC50 values (Fig. S6D), which may serve as promising therapeutic options for ESCC.
Experimental validation of risk feature genes
To further validate the prolif Tex-based model and candidate genes, six genes included in the risk signature were selected for experimental validation in in-house ESCC patients. TNFSF10, ESCO2, and DBF4 had significantly higher expression levels in tumor tissues compared to normal tissues (Fig. 8A), indicating that prolif Tex cells are enriched in tumor tissues. Interestingly, although ESCO2 and DBF4 were defined as protective genes in our risk model, their expression levels were elevated in tumor tissues. Additionally, certain risk genes such as CORO1A and RAB8A exhibited a trend of increased expression in tumor samples, while others, like NDUFB11, displayed relatively low expression levels in tumor tissues (Fig. S7A), despite the differences not being statistically significant. These observations suggest that the functional regulation of these genes in tumors may involve more complex mechanisms.
Survival analysis was conducted to assess the association between gene expression levels and clinical outcomes. Kaplan-Meier survival curves were constructed for disease-free survival (DFS) and OS (Fig. 8B). Patients in the higher expression group of ESCO2 exhibited a significantly longer DFS, with a similar but non-significant trend observed for OS, suggesting that ESCO2 may serve as a potential biomarker for early ESCC diagnosis. KM curves for the other candidate genes are presented in Fig. S7B and C. Correlation analysis of the clinical variables revealed that T-stage and N-stage were significantly negatively correlated with survival time, indicating that patients with advanced-stage patients had shorter survival periods (Fig. 8C). In summary, experimental validation demonstrated tumor upregulation of protective genes ESCO2 and DBF4, with high ESCO2 expression associated with improved survival outcomes.
qRT-PCR validation of prolif Tex cell hub genes in ESCC. (A) DBF4, ESCO2 and TNFSF10 expression in normal esophageal tissue and ESCC tissue from patients. Student’s t-test was used to compare gene expression between normal and tumor tissues. (B) Kaplan-Meier survival analysis comparing DFS and OS in ESCC patients stratified by ESCO2 expression levels. (C) Pearson’s correlation analysis of clinical factors in patients with ESCC.
Discussion
This study provides a comprehensive multi-omics characterization of tumor-infiltrating T cells in ESCC. Through analysis of over 208,125 single-cell transcriptomes, we identified a novel subset of prolif Tex cells characterized by the co-expression of exhaustion and proliferation-related genes. Further investigation revealed that prolif Tex cells not only exhibit high expression of cell cycle-related genes but also have a significantly higher infiltration in tumor samples compared to normal samples. Integrative analysis of TCR-seq and pseudotime trajectories demonstrated that prolif Tex cells originated from Tex. Moreover, greater infiltration of prolif Tex cells was correlated with improved survival, underscoring their critical role in tumor immunity. Notably, in ESCC the common paradigm that high immune infiltration predicts favorable prognosis does not always hold true. Several studies have demonstrated that patients with elevated immune or stromal scores often show reduced tumor purity and significantly worse survival outcomes54,55,56,57. Consistent with these findings, our analysis revealed that the low-risk group with high PTex infiltration exhibited higher tumor purity and lower overall immune levels, but still, the precise immune functions of prolif Tex in ESCC remain to be elucidated and will require further functional assays.
Increasing evidence suggests that T cell exhaustion is marked by heterogeneity and characterized by a well-established three-stage differentiation trajectory: progenitor-transition-terminal58,59. Consistent with this model, we found that prolif Tex cells are distinct from progenitor Tex cells. Progenitor Tex were enriched for stem-like markers such as IL7R and located at the root of the Tex trajectory, whereas prolif Tex showed high expression of cell cycle-related genes (MKI67, TOP2A, UBE2C) and occupied the terminal branch. TCR analysis further revealed that prolif Tex cells shared greater clonal similarity with CXCL13 + Tex than with progenitor Tex, underscoring their separate developmental state. Similar prolif Tex cluster has been identified in a study on head and neck squamous cell carcinoma (HNSCC)36, where their enrichment correlated with improved prognosis in HPV + patients via prolonged survival and enhanced cytotoxicity. Consistent with these findings, our study shows that the enrichment of prolif Tex cells is associated with significantly better prognosis in ESCC patients. Another recent study in ESCC also identified a similar subtype of exhausted T cells in a cycling state60. This subtype was found to express cell cycle-related genes and was enriched in metastatic lymph nodes, suggesting that the proportion of the subset of Tex with characteristics of proliferation may increase as tumor progression advances. Additionally, pseudotime analysis revealed that this Cycling-Tex subtype occupies the terminal position in the CD8 + T cell pseudotime trajectory. These findings align with our results on prolif Tex cells, indicating that they likely belong to the same subset. However, our study represents the first identification of prolif Tex cells specifically in ESCC.
Among the candidate genes in our prognostic model derived from prolif Tex cells, three proliferation-related genes (ESCO2, DBF4, and TNFSF10) have previously been reported to be associated with cancer progression. ESCO2, a cell division-related gene, plays a crucial role in sister chromatid cohesion during mitosis61. High expression of ESCO2 has varying prognostic implications across different cancer types62,63,64. DBF4, in cooperation with CDC7 kinase, participates in initiating DNA replication and in the G1/S transition of the cell cycle65. High expression of DBF4 is frequently linked to poor prognosis in cancers66. TNFSF10, a member of the tumor necrosis factor superfamily, induces apoptosis in tumor cells by binding to death receptors67. While TNFSF10 expression may contribute to enhancing the cytotoxicity of prolif Tex cells, its excessive expression could contribute to immune suppression, worsening patient prognosis. In this study, transcriptomic analysis and quantitative qRT-PCR validation demonstrated that the expression levels of these three genes (ESCO2, DBF4, TNFSF10) were significantly higher in ESCC tumor tissues compared to adjacent non-tumor tissues. Analysis of in-house clinical samples revealed a trend toward better prognosis associated with high expression of these genes, although only ESCO2 showed a statistically significant difference in the DFS of patients. The lack of significant differences for the other genes may be due to the limited sample size, and further validation with an expanded cohort is planned. These findings highlight the clinical potential of the prolif Tex prognostic model and its candidate genes, particularly ESCO2, warranting further investigation in future studies.
A previous study demonstrated that neoadjuvant radiotherapy enhanced CD8 + T effector cell infiltration in the TME of ESCC32. In our study, it is worth noting that NAT led to a significant decrease in the proportion of prolif Tex cells within T cells, whereas the proportion of Teff cells increased. Additionally, in untreated samples, prolif Tex cells were predominantly localized in the region associated with unipotent differentiation potential. However, following NAT treatment, prolif Tex cells were more frequently observed in regions associated with the oligopotent differentiation potential. These findings indicate that prolif Tex cells are highly responsive to NAT treatment and hold potential as a predictive biomarker for treatment response.
There are also several limitations in this study. First, the in-house ESCC cohort included a limited number of patients, which may reduce the robustness of prognostic gene selection, and larger cohorts will be required for more comprehensive modeling in the future. While the transcriptome-level analysis and TCR-seq data have provided insights into the characteristics of prolif Tex cells, the integration of additional modalities, such as spatial transcriptomics and ATAC-seq, is absent. Incorporating these modalities could facilitate the exploration of the spatial distribution of prolif Tex cells in ESCC and the identification of transcriptional regulatory factors driving their differentiation. Future research should prioritize leveraging publicly available multi-modal omics data or collecting new samples for experimental data acquisition. Furthermore, whether prolif Tex cells are broadly present in other cancers and whether they hold prognostic significance remain unresolved questions that require comprehensive pan-cancer analyses. Finally, although a correlation between the presence of prolif Tex cells and improved prognosis has been observed in this study, the biological functions and mechanisms underlying this phenomenon remain unclear due to the absence of cellular, molecular, and animal model studies. Addressing these gaps will be an important focus for future investigations.
In brief, this study is the first to identify prolif Tex cells in ESCC, demonstrating that their infiltration is associated with improved survival in ESCC patients. These findings offer new insights into the role of exhausted T cell heterogeneity in tumor immunity.
Conclusions
In this study, we identified a novel subset of prolif Tex cells within ESCC tumor microenvironment, characterized by coexpression of exhaustion and proliferation markers. A higher infiltration of prolif Tex cells was associated with improved patient survival. These cells originated from conventional Tex and were specially enriched in tumor tissue. Furthermore, a prolif Tex cell-based prognostic model demonstrated efficacy in predicting the prognosis of ESCC patients. Our findings underscore the potential of prolif Tex cells as a biomarker and therapeutic target in ESCC, offering valuable insights for risk stratification and predicting responses to immunotherapy.
Data availability
The public datasets used in this study can be searched in the GEO database (https://www.ncbi.nlm.nih.gov/geo), TCGA database (https://www.cancer.gov/tcga/) and GSA database (https://ngdc.cncb.ac.cn/gsa-human).
Abbreviations
- ESCC:
-
Esophageal squamous cell carcinoma
- Tex:
-
Exhausted T cells
- Prolif Tex:
-
Proliferative exhausted T cells
- EC:
-
Esophageal cancer
- EAC:
-
Esophageal adenocarcinoma
- NAT:
-
Neoadjuvant chemo-immunotherapy
- TME:
-
Tumor microenvironment
- CAFs:
-
Cancer-associated fibroblasts
- Teff:
-
Effector T cells
- Bulk RNA-seq:
-
High-throughput transcriptome sequencing
- scRNA-seq:
-
Single-cell RNA-sequencing
- GEO:
-
Gene expression omnibus
- GSA:
-
Genome sequence archive
- TCGA:
-
The cancer genome atlas
- PCA:
-
Principal components analysis
- UMAP:
-
Uniform manifold approximation and projection
- DEGs:
-
Differentially expressed genes
- CNVs:
-
Copy number variations
- GO:
-
Gene ontology
- KEGG:
-
Kyoto encyclopedia of genes and genomes
- TCR:
-
T cell receptor
- ssGSEA:
-
Single-sample gene set enrichment analysis
- GSEA:
-
Gene set enrichment analysis
- Lasso:
-
Least absolute shrinkage and selection operator
- Enet:
-
Elastic net
- OS:
-
Overall survival
- TIME:
-
Tumor immune microenvironment
- ICI:
-
Immune checkpoint inhibitor
- CMap:
-
Connectivity map
- MoA:
-
Mechanism of action
- GDSC:
-
Genomics of drug sensitivity in cancer
- FUSCC:
-
Fudan university shanghai cancer center
- CR:
-
Complete remission
- PR:
-
Partial remission
- SD:
-
Stable disease
- PD:
-
Progressive disease
- DFS:
-
Disease-free survival
- HNSCC:
-
Head and neck squamous cell carcinoma
References
Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin.71 (3), 209–249 (2021).
Yang, J. et al. Understanding esophageal cancer: the challenges and opportunities for the next decade. Front. Oncol.10, 1727 (2020).
Morgan, E. et al. The global landscape of esophageal squamous cell carcinoma and esophageal adenocarcinoma incidence and mortality in 2020 and projections to 2040: new estimates from GLOBOCAN 2020. Gastroenterology163 (3), 649–658 (2022). e2.
Liang, H., Fan, J-H. & Qiao, Y-L. Epidemiology, etiology, and prevention of esophageal squamous cell carcinoma in China. Cancer Biology Med.14 (1), 33 (2017).
Zeng, H. et al. Cancer survival in C hina, 2003–2005: A population-based study. Int. J. Cancer. 136 (8), 1921–1930 (2015).
Wang, X. et al. Quality of life and survival outcomes of patients with inoperable esophageal squamous cell carcinoma after definitive radiation therapy: A multicenter retrospective observational study in China from 2015 to 2016. J. Natl. Cancer Cent.3 (2), 150–158 (2023).
Li, C. et al. Clinical practice and outcome of radiotherapy for advanced esophageal squamous cell carcinoma between 2002 and 2018 in china: the multi-center 3JECROG survey. Acta Oncol.60 (5), 627–634 (2021).
van Hagen, P. et al. Preoperative chemoradiotherapy for esophageal or junctional cancer. N. Engl. J. Med.366 (22), 2074–2084 (2012).
Yang, H. et al. Neoadjuvant chemoradiotherapy followed by surgery versus surgery alone for locally advanced squamous cell carcinoma of the esophagus (NEOCRTEC5010): a phase III multicenter, randomized, open-label clinical trial. J. Clin. Oncol.36 (27), 2796–2803 (2018).
Zhang, Z. & Zhang, H. Impact of neoadjuvant chemotherapy and chemoradiotherapy on postoperative cardiopulmonary complications in patients with esophageal cancer. Dis. Esophagus.https://doi.org/10.1093/dote/dox002 (2017).
Gao, T-T. et al. Comparative efficacy and safety of immunotherapy for patients with advanced or metastatic esophageal squamous cell carcinoma: a systematic review and network Meta-analysis. BMC Cancer. 22 (1), 992 (2022).
Luo, Y. & Liang, H. Single-cell dissection of tumor microenvironmental response and resistance to cancer therapy. Trends Genet.39 (10), 758–772 (2023).
De Visser, K. E. & Joyce, J. A. The evolving tumor microenvironment: from cancer initiation to metastatic outgrowth. Cancer cell.41 (3), 374–403 (2023).
Blank, C. U. et al. Defining ‘T cell exhaustion’. Nat. Rev. Immunol.19 (11), 665–674 (2019).
Thommen, D. S. & Schumacher, T. N. T cell dysfunction in cancer. Cancer cell.33 (4), 547–562 (2018).
Wherry, E. J. T cell exhaustion. Nat. Immunol.12 (6), 492–499 (2011).
Im, S. J. et al. Defining CD8 + T cells that provide the proliferative burst after PD-1 therapy. Nature537 (7620), 417–421 (2016).
Baessler, A. & Vignali, D. A. T cell exhaustion. Annu. Rev. Immunol.42 (1), 179–206 (2024).
Jiang, Y., Li, Y. & Zhu, B. T-cell exhaustion in the tumor microenvironment. Cell Death Dis.6 (6), e1792–e (2015).
Guo, L. et al. Tumoral PD-1hiCD8 + T cells are partially exhausted and predict favorable outcome in triple-negative breast cancer. Clin. Sci.134 (7), 711–726 (2020).
Choo, J. et al. Clinical relevance of PD-1 positive CD8 T-cells in gastric cancer. Gastric Cancer. 26 (3), 393–404 (2023).
Ross, E. M. & Markowetz, F. OncoNEM: inferring tumor evolution from single-cell sequencing data. Genome Biol.17, 1–14 (2016).
Rao, M. S. et al. Comparison of RNA-Seq and microarray gene expression platforms for the toxicogenomic evaluation of liver from short-term rat toxicity studies. Front. Genet.9, 636 (2019).
Li, J. et al. LncRNA profile study reveals a three-lncRNA signature associated with the survival of patients with oesophageal squamous cell carcinoma. Gut63 (11), 1700–1710 (2014).
Zhang, X. et al. Dissecting esophageal squamous-cell carcinoma ecosystem by single-cell transcriptomic analysis. Nat. Commun.12 (1), 5291 (2021).
Kim, N. et al. Single-cell RNA sequencing demonstrates the molecular and cellular reprogramming of metastatic lung adenocarcinoma. Nat. Commun.11 (1), 2285 (2020).
Jovic, D. et al. Single-cell RNA sequencing technologies and applications: A brief overview. Clin. Translational Med.12 (3), e694 (2022).
Grün, D. & van Oudenaarden, A. Design and analysis of single-cell sequencing experiments. Cell163 (4), 799–810 (2015).
Lei, Y. et al. Applications of single-cell sequencing in cancer research: progress and perspectives. J. Hematol. Oncol.14 (1), 91 (2021).
Hedlund, E. & Deng, Q. Single-cell RNA sequencing: technical advancements and biological applications. Mol. Aspects Med.59, 36–46 (2018).
Zheng, Y. et al. Immune suppressive landscape in the human esophageal squamous cell carcinoma microenvironment. Nat. Commun.11 (1), 6268 (2020).
Ji, G. et al. Single-cell profiling of response to neoadjuvant chemo-immunotherapy in surgically resectable esophageal squamous cell carcinoma. Genome Med.16 (1), 49 (2024).
Stuart, T. et al. Comprehensive integration of single-Cell data. Cell177 (7), 1888–1902 (2019). e21.
Wu, R. et al. Comprehensive analysis of Spatial architecture in primary liver cancer. Sci. Adv.7 (51), eabg3750 (2021).
Patel, A. P. et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science344 (6190), 1396–1401 (2014).
Cheng, D. et al. Proliferative exhausted CD8 + T cells exacerbate long-lasting anti-tumor effects in human papillomavirus-positive head and neck squamous cell carcinoma. Elife12, e82705 (2023).
Yu, G., Wang, L-G., Han, Y. & He, Q-Y. ClusterProfiler: an R package for comparing biological themes among gene clusters. Omics: J. Integr. Biology. 16 (5), 284–287 (2012).
Borcherding, N., Bormann, N. L. & Kraus, G. scRepertoire: An R-based toolkit for single-cell immune receptor analysis. F1000Researchhttps://doi.org/10.12688/f1000research.22139.2 (2020).
Chiffelle, J. et al. T-cell repertoire analysis and metrics of diversity and clonality. Curr. Opin. Biotechnol.65, 284–295 (2020).
Qiu, X. et al. Single-cell mRNA quantification and differential analysis with census. Nat. Methods. 14 (3), 309–315 (2017).
Kang, M. et al. Mapping single-cell developmental potential in health and disease with interpretable deep learning. BioRxiv 2024:2024.03. 19.585637.
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinform.14, 1–15 (2013).
Yoshihara, K. et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat. Commun.4 (1), 2612 (2013).
Newman, A. M. et al. Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods. 12 (5), 453–457 (2015).
Yang, J. M. et al. TCellSI: A novel method for T cell state assessment and its applications in immune environment prediction. Imeta3 (5), e231 (2024).
Lamb, J. The connectivity map: a new tool for biomedical research. Nat. Rev. Cancer. 7 (1), 54–60 (2007).
Yang, W. et al. Genomics of drug sensitivity in cancer (GDSC): a resource for therapeutic biomarker discovery in cancer cells. Nucleic Acids Res.41 (D1), D955–D61 (2012).
Maeser, D., Gruener, R. F. & Huang, R. S. OncoPredict: an R package for predicting in vivo or cancer patient drug response and biomarkers from cell line screening data. Brief. Bioinform.22 (6), bbab260 (2021).
Sakaguchi, S., Miyara, M., Costantino, C. M. & Hafler, D. A. FOXP3 + regulatory T cells in the human immune system. Nat. Rev. Immunol.10 (7), 490–500 (2010).
Wu, L. & Van Kaer, L. Natural killer T cells in health and disease. Front. Biosci.3, 236 (2011).
Kanehisa, M., Sato, Y., Kawashima, M., Furumichi, M. & Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res.44 (D1), D457–D62 (2016).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res.28 (1), 27–30 (2000).
Aoki, H., Shichino, S., Matsushima, K. & Ueha, S. Revealing clonal responses of tumor-reactive T-cells through T cell receptor repertoire analysis. Front. Immunol.13, 807696 (2022).
Zhang, D., Qian, C., Wei, H. & Qian, X. Identification of the prognostic value of tumor Microenvironment-Related genes in esophageal squamous cell carcinoma. Front. Mol. Biosci.7, 599475 (2020).
Xiong, T. et al. An esophageal squamous cell carcinoma classification system that reveals potential targets for therapy. Oncotarget8 (30), 49851 (2017).
Pang, J. et al. Prognostic value of immune-related multi-IncRNA signatures associated with tumor microenvironment in esophageal cancer. Front. Genet.12, 722601 (2021).
Xiong, T. et al. An esophageal squamous cell carcinoma classification system that reveals potential targets for therapy. Oncotarget8 (30), 49851–49860 (2017).
Beltra, J-C. et al. Developmental relationships of four exhausted CD8 + T cell subsets reveals underlying transcriptional and epigenetic landscape control mechanisms. Immunity52 (5), 825–841 (2020). e8.
Im, S. J. & Ha, S.-J. Re-defining T-cell exhaustion: subset, function, and regulation. Immune Netw.https://doi.org/10.4110/in.2020.20.e2 (2020).
Guo, W. et al. Single-cell RNA sequencing and Spatial transcriptomics of esophageal squamous cell carcinoma with lymph node metastases. Exp. Mol. Med.57 (1), 59–71 (2025).
Alomer, R. M. et al. Esco1 and Esco2 regulate distinct cohesin functions during cell cycle progression. Proc. Natl. Acad. Sci.114(37), 9906–9911 (2017).
Huang, Y. et al. ESCO2’s oncogenic role in human tumors: a pan-cancer analysis and experimental validation. BMC Cancer. 24 (1), 452 (2024).
Gan, W. et al. Prognostic values and underlying regulatory network of cohesin subunits in esophageal carcinoma. J. Cancer. 13 (5), 1588 (2022).
Hu, J. et al. ESCO2 promotes hypopharyngeal carcinoma progression in a STAT1-dependent manner. BMC Cancer. 23 (1), 1114 (2023).
Fang, D. et al. Dbf4 recruitment by forkhead transcription factors defines an upstream rate-limiting step in determining origin firing timing. Genes Dev.31 (23–24), 2405–2415 (2017).
Montagnoli, A. et al. Cdc7 Inhibition reveals a p53-dependent replication checkpoint that is defective in cancer cells. Cancer Res.64 (19), 7110–7116 (2004).
Wiley, S. R. et al. Identification and characterization of a new member of the TNF family that induces apoptosis. Immunity3 (6), 673–682 (1995).
Acknowledgements
We acknowledge the Gene Expression Omnibus (GEO) and The Cancer Genome Atlas (TCGA) databases for providing valuable public data. We also extend our gratitude to National Natural Science Foundation of China (NSFC) and the Fudan DreamLab program for their financial support. Finally, we thank the editors and reviewers for their meticulous reading, constructive feedback, and valuable suggestions.
Funding
This work was supported by the National Natural Science Foundation of China (NSFC) (Grant No. 62472415), the Fudan University Undergraduate Academic Science and Technology Innovation Project Dream Lab and the Shanghai Undergraduate Training Program on Innovation and Entrepreneurship (SUTPLE).
Author information
Authors and Affiliations
Contributions
H Zhou conceptualized and designed the study. C Lam and GL conducted data collection and analysis. GL, C Lam, RC and XW performed the statistical analysis and visualized the data. GL and C Lam drafted the manuscript, which was subsequently revised by H Zhou. H Zhang provided advice with data analysis and edited the manuscript. FF performed surgical operations and provided clinical samples. All authors read and approved the final manuscript. GL and C. Lam contributed equally to this work. H Zhou reviewed the manuscript and supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
All human experiments of in-house cohort in this study were approved by the Ethics Committee of Fudan University Shanghai Cancer Center (FUSCC). Informed consent was obtained from all participants prior to their inclusion in the study. The research was conducted in accordance with the Declaration of Helsinki and received approval from the Ethics Committee of FUSCC (Approval No. 050432-4-1911D).
Consent for publication
All authors have read the manuscript and provided their consent for the submission.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, G., Lam, C.Y., Chen, R. et al. Discovery and validation of proliferative exhausted T cells as a favorable prognostic biomarker in esophageal squamous cell carcinoma. Sci Rep 15, 41619 (2025). https://doi.org/10.1038/s41598-025-25535-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-25535-x