Abstract
Global biodiversity is declining at alarming rates, necessitating efficient and scalable biodiversity monitoring methods. This study evaluated the efficacy and efficiency of using mosquito blood meal analysis (MBA) as a tool to survey terrestrial vertebrate assemblages during wet and dry seasons in a subtropical landscape of central Florida. We compared the performance of MBA against that of a suite of conventional sampling methods for birds, mammals, amphibians and reptiles. As a single method, MBA detected the highest number of species. Nonetheless, our results indicated that during the dry season, conventional methods were more effective and efficient than MBA, particularly for detecting small mammals and ectotherms. Similarly, in the wet season, conventional methods outperformed MBA in assessing the diversity of individual guilds. Nonetheless, during this season, MBA performance was comparable to that of the conventional techniques when the goal was to assess the entire vertebrate diversity. This is likely due to the higher abundance of adult mosquitoes in the landscape during the wet season. A community similarity analysis showed that MBA and conventional methods obtained different profiles of the vertebrate assemblages. Each method seemed to perform better for dissimilar sets of species. These findings highlight the potential of MBA as a scalable biodiversity monitoring tool, particularly as complementary to conventional methods, or stand-alone method under some environmental conditions. These results could also aid in identifying vertebrate species suitable to be surveyed using MBA.
Similar content being viewed by others
Introduction
Global biodiversity is declining at rates comparable to past mass extinctions1 and researchers, managers and policy makers require ecological data at both coarse and fine spatiotemporal resolutions to inform timely, science-based actions to reverse this trend2. Assessments of species richness, community composition, and demographic trends are key to evaluating anthropogenic impacts on biodiversity and the outcomes of conservation actions3,4,5. The urgent need for such data is emphasized in the Kunming-Montreal Global Biodiversity Framework, with its implementation relying on the capacity to measure diversity at various scales6,7. Conventional biodiversity monitoring methods are time and resource intensive, requiring an array of taxon-specific techniques and experienced field personnel8,9. Such costs drive the development of more efficient technological methods to rapidly assess biodiversity. Among these methods, the identification of species or individuals from genetic material available in the substrate, water or air (environmental DNA: eDNA), and the use of vertebrate DNA ingested by invertebrates that feed on vertebrate blood, tissue, or excrement (invertebrate-derived DNA: iDNA) offer great potential10,11.
Numerous terrestrial vertebrate species are threatened and endangered by habitat conversion to human uses and a rapidly changing climate. Other vertebrates have become invasive and ecological destructive outside their native ranges. As a result, monitoring and managing vertebrate taxa, their population and community health, distributions, impacts, and shifting ecological functions largely defines biodiversity conservation12,13,14,15. In terrestrial ecosystems, a diverse array of invertebrates feed on the blood, tissue, and feces of vertebrates; consequently, iDNA is a promising tool for surveying vertebrate assemblages. To date, iDNA from carrion and biting flies, terrestrial leeches, dung beetles and mosquitoes has been used to detect species of vertebrates under diverse ecological conditions16. Although initial approaches focused on detection of specific species, iDNA is now used to model species richness, abundance and occupancy, highlighting the potential of this approach to inform diverse ecological, conservation and management questions10,17. In terrestrial ecosystems, most of these approaches have focused on large mammalian species. Here, we explore the capacity of the tool to survey assemblages of terrestrial vertebrates and test the use of iDNA against personnel-intensive conventional vertebrate survey techniques16,18.
Mosquitoes (Diptera: Culicidae) are a family of hematophagous flies that are particularly well-suited to survey vertebrate assemblages due to the diverse feeding ecologies displayed by this group. For example, a typical community of mosquitoes in a subtropical landscape of central Florida can include species in the genus Uranotaenia, some of which feed on amphibians19, Culex subgenus Melanoconion of which some species are specialized on reptiles20, Anopheles which are specialists on mammals21, and Culiseta some species of which are specialists on birds22. Given a thorough sampling of a diverse mosquito community, it is feasible that the blood meals of these mosquitoes would contain a representative sample of species richness and composition of vertebrate communities. In order to test this hypothesis, ground truthing and direct comparison with conventional methods are required. The inability of a recent meta-analysis16 to draw general conclusions on the efficiency of iDNA to assess the diversity of terrestrial vertebrate groups highlights the need for additional ground truthing of iDNA approaches.
To this end, we evaluated the performance of mosquito blood meal analysis (MBA) as a tool to inventory the entire terrestrial vertebrate assemblage of a subtropical landscape in central Florida, United States. We compared the ability of unique sets of conventional sampling methods designed to survey 4 guilds of vertebrates (birds, herpetofauna, small mammals, and medium and large mammals) against mosquito blood meal analysis to estimate the diversity and quantify species composition of the terrestrial vertebrate assemblage in the study area (Fig. 1). Our specific objectives included: i) Creating accumulation curves to compare the efficacy and efficiency of conventional survey methods and MBA; ii) comparing the effort required to detect each species by both methods, with an emphasis on species of conservation or management interest; and iii) assessing the similarity of the community profiles generated by each method. Given that the life cycle and abundance of mosquitoes is highly influenced by climatic conditions, we performed these comparisons separately during dry and wet seasons.
Research design overview. Mosquitoes were collected employing resting shelters (A) and aspirators (B) in a mesh bag (C); at the lab each mosquito was identified to species and the blood from engorged individuals was extracted on an FTA card employing a sterile pipette tip (D) for PCR amplification of COI sequencing and sequence comparison (E). This protocol yielded information on all vertebrates in the study area. Simultaneously, a suit of conventional methods was used to target each vertebrate guild: point counts (F), drift fences (G), camera traps (H) and Sherman trapping (I).
Methods
Study area
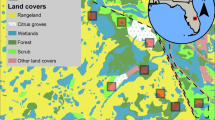
This study was conducted at the University of Florida, Institute of Food and Agricultural Sciences, DeLuca Preserve, located in central Florida (− 80.883°, 27.654°) and occupying 110 km2 of the Eastern Florida Flatwoods Ecoregion23. Habitats included upland and mixed coniferous forest, Florida scrubland, wetlands, and citrus groves, immersed in a matrix of pastureland (Fig. 2A). Sections of the study area were regularly burned as part of a prescribed fire management regime. The area experiences an average annual precipitation of 1380 mm, concentrated from June to September, and average monthly temperatures of 22.9 °C24. Nonetheless, rainfall patterns were atypical during our study period (January to October 2022), with lower-than-average rainfall from June to August, and higher precipitation in the period of September to October (25; Fig. S1). Hence, we conducted separate analyses for the drier period of our study (January to August; dry season henceforth), and the period after the atypical precipitation (October; wet season henceforth).
Study sites and deployment of vertebrate diversity surveys. Pane A displays the location of the University of Florida, Institute of Food and Agricultural Sciences, DeLuca Preserve, and the distribution of the sites surveyed through the dry season and the targeted area for a rapid survey during the wet season. Pane B shows an example of the arrangement of the conventional vertebrate sampling techniques and mosquito surveys at one of the eight sites surveyed during the dry season. Pane C shows the deployment of the rapid assessment performed during the wet season. Maps created using ArcGIS Pro. Land cover data obtained from https://geodata.dep.state.fl.us/datasets/FDEP::statewide-land-use-land-cover/. Satellite images obtained from ArcGIS basemaps (Credits: Esri, Maxar, Earthstar Geographics, and the GIS User Community).
Study design
During the dry season of this study, we sampled eight sites within the DeLuca Preserve covering the dominant habitat types of the ecoregion. Two sites were sampled for each of the following habitats: upland forest, Florida scrub, wetland edges, and citrus groves (Fig. 2A). The sites occurred in the northeastern portion of the preserve, with an average spacing of 1.7 km (minimum 1.3 km) between sites. During each of nine sampling bouts, every site was simultaneously surveyed by wildlife biologists employing unique conventional sampling methods for each of four guilds of vertebrates (birds, herpetofauna, small mammals, medium and large mammals), and entomologists collecting blood-fed mosquitoes (Figs. 1, 2B). The sampling bouts were spaced between two and four weeks, and each one consisted of five days of sampling.
Given the atypical rainfall pattern of the study year, and to assess the performance of each survey approach under different climatic conditions, we had one sampling bout of five days on October 10–14 of 2022, after intense rains in September. During this sampling bout a different area of the DeLuca Preserve was surveyed (Fig. 2C), and the sampling methods were modified to cover diverse habitat types in a short amount of time (see below).
Mosquito sampling and bloodmeal analysis
Two techniques were employed to collect blood fed mosquitoes: resting shelters and wide diameter aspirators. We focused our surveys, using both methods, to areas within our sampling sites where mosquitoes were expected to rest to maximize the proportion of blood-engorged female mosquitoes collected26,27.
The resting shelters consisted of a self-expanding cylindrical waste receptacle (48 cm × 61 cm), constructed of spiral metal spring covered by a dark polyester tarp material or thick black plastic bag (Campers Choice Sequim, Washington, USA). Resting shelters were installed in shaded areas and mosquitoes were collected from each shelter by placing a lid with a collection cup over the shelter opening and compressing the shelter several times to force the mosquitoes inside into the collection cup28. The wide diameter aspirator consisted of a custom-made device constructed using an automotive radiator fan, powered by a 12-V battery, and attached to a wire field fencing frame covered in polyester tarp26,29. A mesh bag was attached inside the aspirator to capture the mosquitoes sucked into the aspirator intake. During the dry season, two to three shelters per site were sampled daily during the sampling bouts. Aspiration was performed daily for ten minutes during each bout at each site, focusing on microhabitats where mosquitoes are expected to rest (leaf litter, undergrowth, woody debris, and recently disturbed areas). During the wet season sampling bout, six sites were selected covering diverse habitats and surveyed daily by aspirators only using the methods described above and the same effort for five days.
Captured mosquitoes were killed in an insulated cooler with dry ice, transferred into labeled vials, and kept on dry ice for transportation back to the lab. Mosquitoes were stored at − 20 °C or − 80 °C until processing. In the laboratory, individual mosquitoes were taken to room temperature and immediately identified using the morphological keys in Darsie and Ward (2005)30, afterwards blood-engorged females were separated for blood meal analysis. Mosquito blood meals were preserved by placing an engorged female onto a Whatman Flinders Technology Associates (FTA) Classic Card, and contents of the abdomen smeared onto the card using a sterile pipette tip31. FTA cards with preserved blood meals were stored at room temperature.
Two punches of 1 mm were taken from each blood spot on the FTA card and placed into 1.0 mL microcentrifuge tube. DNA was extracted from the punches using the Hot Sodium Hydroxide and Tris (HotSHOT) method32. A fragment of the Cytochrome C Oxidase Subunit I (COI) gene was amplified from the extracted DNA using vertebrate-specific primers33 in a Polymerase Chain Reaction (PCR). The PCR products were visualized on an agarose gel and positive amplicons were sent to Eurofins Genomics (Louisville, KY) for chain termination sequencing in one direction34. The returned DNA sequence files were edited using Geneious Prime 2023.2.1 and submitted to the Barcode of Life Data System (BOLD) Identification Engine for comparison to reference sequences35. Any matches with more than 98% similarity were considered a host detection, with some exceptions36. Representative sequences of the detected vertebrate species are available in GenBank database via accession numbers PV786364–PV786446.
During the dry season a field effort of 211 h and laboratory effort of 390 h was invested in MBA for a total effort of 601 h. During the wet season, 22.5 h of fieldwork and 178.6 h of laboratory work were invested in MBA for a total effort of 201.1 (Supplementary Materials S1).
Conventional vertebrate surveys – dry and wet seasons
We sampled differently in wet and dry seasons owing to wet season access issues. In dry season at all eight sites, four guilds of vertebrates were sampled through conventional methods: birds, herpetofauna (amphibians and reptiles), medium-to-large-mammals, and small non-volant mammals (Fig. 2B). An overview of the methods employed to survey each group is provided below, see Botero-Cañola et al. (2024)37 for further details. To survey the southwestern end of the DeLuca Preserve during the wet season, we adapted the Rapid Assessment Project biodiversity protocol38 to cover as much habitat variation and detect as many species as possible in one sampling bout of five days throughout the area (Fig. 2C).
In the dry season, birds were sampled using point counts39. At each site, three points spaced an average of 255 m (min. 150 m) were sampled once each field bout, except during one bout in July when logistical constraints precluded bird sampling. The protocol at each point was conducted by a single skilled observer who stood quietly after arriving at the point center for 1–2 min, then conducted five minutes of observation while recording the species and number of birds detected by sight and sound. In the wet season, birds were surveyed using two transects, each consisting of five point-counts spaced 250 m apart (Fig. 2C). Birds were recorded for ten minutes at each point count following the protocol described above, as well as while walking the distance between points at an average speed of 1 km/h. The transects were laid out to cover as much habitat variation as possible (wetlands, Florida scrub, mesic and hammock forest), and were surveyed for four days. During both seasons bird sampling was typically conducted in the mornings, from 5:55 to 10:00 h. All detected birds were recorded regardless of their distance to the observer given that the objective was to quantify the diversity of the entire study area.
Amphibians and reptiles were surveyed using two approaches in dry season: drift fences and opportunistic counts. As both taxa were surveyed employing the same methods, they were pooled together for all analyses and are referred hereafter as Herpetofauna. The first approach consisted of the installation of drift fences composed of three arms, with each 7.6 m long arm radiating from the center at 120° angles. Pitfall traps were installed at the center and the three ends of the arms, while sets of four double-sided funnel traps were installed along the fence in each arm40. To protect the captured animals, the funnel traps were kept covered by a shade cloth, and the pitfall traps included a section of pool noodle or a sponge in the bottom to provide shade and a flotation device in case of flooding. Each drift fence was complemented by six cover boards (1 × 1 m) and four ground-level PVC pipes (1 m long, 5.1 cm diameter) installed in the vicinity41. During the dry season we deployed two drift fence arrays at each site installed at an average of 200 m apart for a total of 16 drift fences in the dry season study area. Each drift fence was checked twice daily, while the cover boards and pipes were checked once each bout. During one bout in July, only four drift fences were active due to logistic restrictions. In addition, during the dry season we surveyed herpetofauna opportunistically by recording any species of this group observed (sight or sound) when walking within 400 m of the centroid of each site.
In the wet season, herpetofauna were sampled through visual encounter and audio surveys at five sites (Fig. 2C). Each survey was constrained to one hour of search covering at least 100 linear meters. The five sites were surveyed once during the day and once during the night by 2–3 researchers who recorded all detections of visually or acoustically identified species. Given the short duration of the wet season surveys, no drift fences were installed during these surveys.
Medium to large mammals were surveyed using camera traps (Reconyx Hyperfire) installed at game trails and roads and mounted approximately 50 cm above the ground and angled towards the nearest trail or road42. In the dry season three camera traps were placed at each site with an average distance of 475 m (235–813 m) between cameras. In dry season, one of the three cameras was baited for carnivores, using dry cat food, tuna, or sardines as attractants. Cameras were continuously active during the study and checked every three months. In the wet season, we installed six camera-traps throughout the study area for five days for a total of 30 camera/nights (Fig. 2C). During this season, cameras were installed with a minimum distance of 1 km on game trails and roads and no bait was used. Photos were processed using the Wildlife Insights platform (Project Id 2004061), where blank pictures were automatically flagged for removal, and species were manually identified.
In dry season, small non-volant mammals were surveyed at each site with a transect of 30 Sherman aluminum folding traps (7.62 × 8.89 × 22.86 cm; H.B. Sherman Traps, Tallahassee, Florida) spaced ten meters. The Sherman traps were baited with oats and cotton was placed inside the traps to act as bedding for the captured rodents. Traps were checked and closed each morning and opened near sunset every sampling day. Each bout consisted of four trapping nights except when logistical constraints (e.g. high temperatures or prescribed fires) or the activity of mesopredators forced us to halt trapping at certain sites/bouts. Sampling at forested sites was complemented with the installation of ten Sherman traps on trees or understory vegetation. In wet season, we deployed two transects consisting of two Sherman traps every ten meters for 300 m (60 traps per transect). Traps were baited with oats and checked following the dry season protocol. Captured individuals were marked, measured and identified to species using morphological characters.
A total effort of 1,805 h was invested in surveying vertebrate diversity employing conventional methods during the dry season. On the other hand, 96 h were invested in conventional surveys during the wet season.
Statistical analysis
To compare the efficacy and efficiency of MBA with that of conventional methods, we created species accumulation curves of both methods for each vertebrate guild, as well as for the entire vertebrate assemblage. These curves displayed the accumulation of unique species as the number of observations or captures increased through time and by effort. To assess the accumulation of species with effort, the x axis of these curves represented the number of personnel hours accumulated in the field and lab (sampling effort). To allow for robust comparisons between the methods, we used the Chao and Jost (2012)43 non-parametric method of rarefaction and extrapolation of the accumulation curve to predict the expected number of species detected at every amount of sampling effort. In addition, for each value of sampling effort, we estimated the sample coverage, defined as the estimated proportion of individuals in the assemblage represented by the set of detected species. Sample coverage (SC) represents a robust way of standardizing samples for comparison, especially when comparing communities (or methods) where the same amount of effort can result in different numbers of detections44,45.
The efficiency (i.e., how well a community was sampled in relation to the invested effort), and efficacy (i.e., the proportion of the community that could be detected) of each method can be compared using the interpolation and extrapolation of species accumulation curves. We compared two parameters related to efficiency, which determines how fast species accumulate as sampling increases. First, we compared the number of detected species at the lowest effort achieved by either conventional or MBA sampling by interpolating the species richness value to the smallest effort in each guild (i.e., rarefaction comparison). Subsequently, we estimated the amount of effort required by each method to achieve a sample coverage of 0.98 (SC = 0.98), a desirable target for biodiversity sampling45. On the other hand, efficacy of each method determines the number of species at which the accumulation curve reaches its asymptote (no further effort will result in new species detection). Given that the asymptotic value could present high uncertainty44, we compared the estimated species richness as the curve approaches its asymptote at a SC = 0.98 (i.e. the number of species that represent 98% of the samples of the assemblage-asymptotic richness hereafter).
We used the function iNext in the r package iNext46 to create the accumulation curves and estimate the abovementioned performance parameters. We used as input the incidence frequency list with the number of effort units (hours) in which each species was detected and used 100 bootstrapped replicates of the model to estimate the upper and lower limits of richness estimates at 95% confidence. To estimate the confidence limits of our second measure of efficiency (required effort to achieve SC = 98%), we used the standard deviation of the effort values for which the estimated SC confidence limits included 0.98. We report as significantly different the comparisons of MBA and conventional methods when there was no overlap in their confidence limits. This is a conservative assumption as overlapping intervals might not preclude statistical significance44,47. These analyses were done for each vertebrate guild and the entire assemblage for the dry and wet seasons. We also analyzed these data taking only into account the effort invested in field sampling and these results are presented in the Supplementary Materials S1.
To evaluate the effort required to detect each species by conventional methods and MBA method, we calculated the accumulated effort required to obtain the first detection of every species. We report this parameter for every species and discuss these in the light of species of management importance defined as invasive species and species listed as of conservation concern at the state, national and global scales48. This analysis was also done separately for the dry and wet seasons.
Finally, we compared the community composition portrayed by each method for both dry and wet seasons. We compared the assemblages portrayed by the identity of the detected species in each method using the Sorensen Index, as well as by the relative frequency of detections of these species using the Bray–Curtis Index. Both the Sorensen and Bray–Curtis indices give a score of 0 to 1, where 1 are completely dissimilar and 0 completely similar assemblages in terms of species composition, and community structure, respectively. These indices were estimated using the function vegdist of the Vegan R package. The relative frequency was calculated as the number of detections divided by the respective effort and was standardized using the Wisconsin method49. A cluster analysis was performed using each dissimilarity matrix and employing the average method. To graphically display the differences in community structure and highlight the species driving these differences, we performed a Non-metric Multidimensional Scaling (NMDS) analysis using the function metaNMDS from the Vegan R package and employing the Bray–Curtis dissimilarity matrix. In the NMDS we plotted the species accounting for the highest loadings in each vertebrate guild (0.85 percentile).
The formatted dataset employed to conduct the abovementioned analyses is included in Supplementary materials 2.
Ethical statement
We ensured that all animal trapping and handling was carried out in accordance with the taxon-specific relevant guidelines, reported in accordance with ARRIVE guidelines, and in compliance with the laws and regulations of the state of Florida and the U.S. Our methods to capture and process vertebrates were approved by the Institutional Animal Care and Use Committee (IACUC) of the University of Florida (IACUC202100000040).
Results
Efficacy and efficiency of vertebrate surveys
During the 211 h of field work and 390 h of lab work of the dry season, 3,508 blood-engorged mosquitoes belonging to 21 species were collected. Of these, 2,075 yielded identifiable DNA sequences, with 85% of these blood meals representing four mosquito species: Culex nigripalpus (49%), Aedes infirmatus (14%), Culex pilosus (14%), and Psorophora columbiae (8%). After removing the sequences not identified to species, and those corresponding to humans or domestic animals, our sample consisted of 1,965 vertebrate detections. For more details about the mosquito community and host associations during the dry season see Atsma (2023)50 and Atsma et al. (in. review)36. This collection of mosquitoes resulted in the detection of 56 species of birds, 15 species of herpetofauna (6 amphibians and 9 reptiles), nine species of medium to large mammals and one species of small terrestrial mammal, for a total of 81 species (Table 1). On the other hand, 1,805 h of effort using conventional sampling techniques during the dry season resulted in the detection of 74 species of birds, 40 species of herpetofauna (15 amphibians and 25 reptiles), 14 species of medium to large mammals and six species of small terrestrial mammals, for a total of 134 species (Table 1).
The conventional methods were more effective and efficient than MBA in detecting vertebrate diversity during the dry season (Fig. 3; Table 1). For all individual guilds conventional methods were projected to detect more species than MBA under similar sampling efforts (rarefaction richness). The maximum number of species potentially detected (Asymptotic richness) was also higher for conventional methods in all guilds except for birds, which did not display a significant difference in this parameter. Approaching this asymptotic richness required a higher effort using MBA for birds, medium-to-large mammals and small mammals, while the herpetofauna guild seemed to require lower efforts by MBA, with a very narrow overlap between the estimates of both methods. Similarly, when the goal was to survey the entire vertebrate community, conventional methods were projected to detect more species under similar sampling efforts, and included more species at the asymptote of the sampling curve (52% higher species richness detected). On the other hand, MBA could reach this lower asymptotic richness with almost half effort required by conventional methods to reach their estimated asymptotic richness (Fig. 3; Table 1). As a single method, MBA detected more species than any individual conventional method (Table 1). When only field effort was taken into account to estimate the performance metrics, conventional methods still outperformed MBA for individual guilds; but for assessing the entire vertebrate community, both methods presented similar asymptotic richness and conventional methods presented a higher asymptotic richness that requires more effort to detect (Supplementary materials 2; Figure S2).
Accumulation curves of species detected using mosquito blood meal analysis (MBA, red) and a suite of conventional vertebrate survey methods (blue). The curves present results for birds (A and B), amphibians and reptiles (C and D), medium to large mammals (E and F), small mammals (G and H), and the entire vertebrate community (I and J). Accumulation curves are shown for dry (left) and wet (right) seasons sampling. Interpolated estimates are displayed as continuous lines and extrapolated estimates as dotted lines. Shaded areas represent 95% confidence limits of the estimations. Vertical dotted lines indicate the effort value at which richness was compared for the rarefaction metric.
During the wet season, 2,482 blood-fed mosquitoes belonging to 15 species were collected and their bloodmeal sequenced through 22 h of fieldwork and 179 h of lab work. Of these, 1,494 yielded identifiable DNA sequences, with 90% of these blood meals representing three mosquito species: Culex nigripalpus (73%), Culex pilosus (12%) and Aedes infirmatus (6%). After removing the sequences corresponding to humans or domestic animals, our sample was 1,342 detections. This MBA sample resulted in the detection of 36 species of birds, 21 species of herpetofauna (9 amphibians and 12 reptiles), and nine species of medium to large mammals for a total of 66 species (no species of small terrestrial mammal were detected; Table 1). The 96 h of effort employing conventional methods during this season resulted in the detection of 39 species of birds, 20 species of herpetofauna (12 amphibians and 8 reptiles), six species of medium to large mammals and two species of small terrestrial mammals, for a total of 67 species (Table 1). For birds, the lower vocalizations produced in the wet season may have lessened their detection probabilities compared to the dry and help explain why wet season counts by people were less than during the dry season.
Similar to our results for the dry season, during the wet season the rarefaction richness was higher using conventional survey methods for all groups except for medium-to-large mammals which did not present differences in this metric with MBA (Fig. 3; Table 1). On the other hand, the asymptotic richness was similar between both methods during this season, presenting significant differences only for small mammals. Effort tended to be higher for each individual guild using MBA with the exception of medium-to-large mammals which did not present significant differences. Although the rarefaction richness of the entire vertebrate assemblage was higher for conventional methods, asymptotic richness and effort did not differ between these and MBA for the entire assemblage. Again, the single method with the highest diversity of detected species was the MBA, followed by point counts (Table 1). When only field effort was considered to estimate the performance metrics during this season, there were few differences in the performance of both methods for individual guilds; and MBA showed a higher efficiency in its rarefied richness and effort to achieve the asymptote (Supplementary materials 2; Fig. S2).
Effort and species detection
During the dry season, 17 species of vertebrates were detected only by MBA, 70 species were detected only by conventional sampling, and 64 species were detected by both methods. Of these jointly detected species, 51 species were detected with a lower sampling effort using conventional methods, while 13 were detected with a lower effort using MBA (Figs. 4 and 5; Supplementary Materials 2). During this season, 27 species of management concern were detected, of which five species, corresponding to birds, were detected only by MBA; 11 species only by conventional sampling; and 11 by both methods (Figs. 4 and 5). In three of these species, MBA resulted in more rapid detections. Of these cases, two corresponded to invasive herpetofauna: brown anole (Anolis sagrei) and Cuban tree frog (Osteopilus septentrionalis), which were detected remarkably faster with MBA than with conventional sampling (55 vs. 136 h and 225 vs. 577 h, respectively). In line with the results from the species accumulation curves, a considerable proportion of the species of vertebrates was not detected by MBA. This was particularly evident for small non-volant mammals for which only one species was detected (Supplementary materials 2).
Effort attained until the first detection of each species of bird during the dry and wet seasons of sampling using conventional methods (blue) and mosquito blood meal analysis (red). Species of conservation concern are highlighted in green and their corresponding bar identified with an asterisk.
Effort attained until the first detection of each species of reptile (A), amphibian (B) and mammal (C) during the dry and wet seasons of sampling using conventional methods (blue) and mosquito blood meal analysis (red). Species of conservation concern are highlighted in green and invasive species in red. The bars corresponding to these species of management concern are indicated with an asterisk.
During the wet season 29 species of vertebrates were detected only by MBA, 30 species were only detected by conventional sampling, and 37 species were detected by both methods. Of these jointly detected species, 31 species were detected with a lower sampling effort using conventional methods, while five were detected with a lower effort using MBA, and the white-tailed deer (Odocoileus virginianus), was detected with equal effort (Fig. 5). During this season, 19 species of conservation concern were detected, of which six species were detected only by MBA, five species were detected only by conventional sampling and eight by both methods (Figs. 4 and 5). In two of these species, MBA resulted in more rapid detections, while another six species were more rapidly detected by conventional sampling. Interestingly, despite the performance of MBA for all other guilds, this method did not detect any small non-volant mammal species (Supplementary materials 2).
Community composition similarity
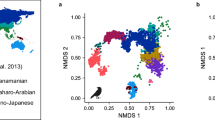
The samples of the vertebrate community obtained by each method and in each season were different both in terms of species identity as well as regarding their relative abundances (Supplementary materials 2). The Jaccard Index (based on species presence/absence) ranged between 0.54 between the communities detected by MBA in the dry and wet seasons, and 0.69 between the community detected by MBA during the wet season and that by conventional methods during the dry season (Table 2). The Bray–Curtis Index suggested even larger differences in terms of relative abundance of the species in the community samples. This index ranged between 0.74 between the assemblages portrayed by the conventional methods in the dry and wet season, to 0.9 for the profiles provided by the conventional wet and MBA dry. Using both distance metrics, the communities were clustered together by sampling method (Fig. S2). The NMDS shows a similar picture, with communities obtained with the same methods being closer. The NMDS showed a clear separation by methods on the first axis of ordination, with abundances of various bird and rodent species increasing in communities portrayed by conventional methods, while turkeys (Meleagris gallopavo), marsh rabbit (Sylvilagus palustris), invasive herpetofauna, the barred owl (Strix varia) and the barking tree frog (Hyla gratiosa) being more abundant in communities characterized by MBA. The second axis of ordination of the NMDS separated dry from wet surveys for both methods, with bald eagle (Haliaeetus leucocephalus) and rattlesnakes (Crotalus adamanteus) being more abundant in the dry season, and the great blue heron (Ardea herodias), southern leopard frog (Rana sphenocephala), black racer snake (Coluber constrictor), black rat snake (Pantherophis obsoletus), and box turtle (Terrapene Carolina) being more abundant during the wet season (Fig. 6).
NMDS analysis of the community samples obtained by mosquito blood meal analysis (MBA) and conventional methods for each season at the vertebrate assemblage of the DeLuca Preserve based on Bray–Curtis dissimilarity matrix estimated from relative abundance data. The species presenting the highest loads from each guild are also plotted. The vectors are scaled to 0.7 for display.
Discussion
Our results show that for the semi-tropical ecosystem of south central Florida, conventional sampling methods remain essential to effectively and efficiently survey vertebrate assemblages, especially when targeting particular guilds. This conclusion was supported by the fact that for most guilds, conventional methods outperformed MBA in at least two of the calculated metrics and agreed with a recent meta-analysis focused on the use of eDNA for mammal inventories18. However, this work also points to the utility of MBA in vertebrate biodiversity sampling as a complementary method that maximizes the ability of surveys to detect vertebrate species, and, in certain circumstances, as a standalone method. For example, in the wet season, MBA achieved similar asymptotic richness for the entire vertebrate community with comparable effort to the suite of conventional methods. Furthermore, when the goal was to reduce time spent in the field, MBA outperformed conventional methods during the wet season. As ever, the selection of vertebrate survey methods should be carefully decided based on the research objectives, environmental conditions, available personnel, techniques and funding. Overall, this study contributes to the much-needed empirical data on the performance of novel iDNA methods16.
Effectiveness and efficiency of MBA to survey vertebrate communities
Results from previous studies comparing mosquitoes against other dipterans or dung beetles as iDNA samplers show mixed results, with mosquitoes presenting lower capture rates and blood-meal diversity than other samplers in most cases10,51,52, and outperforming sandflies in one study53. The only one of these studies that compared MBA to conventional methods, showed that the use of camera traps outperformed MBA10. However, these studies used collection methods such as CDC light traps that do not target blood-fed mosquitoes and could present bias towards certain species of mosquitoes54. Our mosquito collection protocol, targeting blood-fed mosquitoes has been the most successful in terms of the total number of vertebrate species detected, highlighting the need to adjust methods for collecting blood-engorged mosquitoes36. Similar to our results, studies assessing the performance of other iDNA samplers have found that despite potential, the results are mixed and, in all cases, iDNA seems to be a complement rather than replacement of conventional sampling techniques18,55. Thus, comparisons in different environmental contexts are of paramount importance before a particular approach to iDNA or eDNA can be routinely applied and scaled56.
To our knowledge, this is the first study to compare vertebrate diversity recovered by MBA with that recovered by a sampling design targeting all guilds of terrestrial vertebrates. Our work highlights the fact that when the time required to sort, preserve, extract, sequence and identify mosquito bloodmeals is taken into account, the effort required to achieve a satisfactory sampling completeness can be higher with MBA than conventional methods. Thus, more studies assessing iDNA and taking into account the complete required effort are needed to further understand the conditions under which iDNA is most efficient. It remains to be seen if the effect of pooling mosquito samples and excluding identification from the workflow can influence the efficiency of the method. On the other hand, we found that when only the field effort is considered, MBA can outperform conventional methods, particularly during the wet season. This is important information for cases when logistic or security constraints limits time available in the field.
Wet versus dry season effectiveness of MBA
In tropical and subtropical regions, mosquito communities reach their peak of richness and abundance during or after wet seasons, when species can readily breed57,58,59. Thus, the enhanced performance of MBA in wet seasons was not surprising. These increases in diversity and abundance expanded the number and diversity of vertebrate taxa on which the mosquito community fed, and the abundance of mosquitoes captured. Evidence of this lies in the fact that five days of sampling during the wet season yielded three quarters of the sample size obtained during the 45 days of sampling in the dry season (2,075 vs. 1,494 identifiable sequences from the dry and wet season, respectively). The most striking outcome of these seasonal sampling differences is the fact that MBA turns out to be an effective and efficient method to survey the entire vertebrate community during the wet season, approaching the asymptote of the species accumulation curve with the same effort required to do so by conventional sampling methods. By providing an efficient way to sample vertebrates during the wet season, MBA can fill a general sampling gap, resulting from logistical or accessibility constraints, in which a larger amount of survey effort has been invested in dry season for some vertebrate guilds60.
Although we attribute this difference in the performance of MBA to seasons, we cannot completely rule out the effect of location as dry and wet season surveys occurred on different areas of the preserve. Nonetheless, the land cover types among the sites surveyed during the dry and wet season were similar, suggesting the findings are due to a season effect. Similarly, differences in the communities characterized by the conventional methods could partially be explained by methodological differences, especially the use of active search of herpetofauna during the wet seasons as opposed to the use of drift fences during the dry season61,62.
When specific species are of interest
The MBA showed higher or equal efficiency in detecting certain species of management importance such as invasive brown anoles, Cuban tree frogs and invasive wild pigs (Sus scrofa), or taxa of conservation concern such as the American woodcock (Scolopax minor) or the Kentucky warbler (Geothlypis formosa). These are species with current monitoring needs, and for which new efficient detection techniques could prove useful63,64,65,66. During the wet season MBA and conventional methods appear to be complementary, with around 30 species (~ 30%) uniquely detected by each method. During this season, mosquito bloodmeal analysis was faster at detecting wetland birds, as well as certain reptiles of management interest such as the gopher tortoise or the brown anole. This differential capacity by MBA and conventional methods suggests that MBA can be included as a tool to detect certain species regardless of its efficiency in quantifying overall vertebrate diversity. For example, MBA was as efficient in detecting invasive wild pigs in the landscape during the dry season and required less effort than camera trapping to detect this species during the wet season. Furthermore, MBA sampling has the additional advantage of providing a biological sample with a verifiable species identification that could be used to identify individuals, assess genetic diversity, or even assess the occurrence of pathogens67,68,69.
A disadvantage of MBA is that species detection through mosquito blood meal analysis is only as good as the DNA reference databases used to identify host sequences (e.g., BOLD, National Center for Biotechnology Information GenBank). Taxonomic coverage of such databases is incomplete for many ecosystems and vertebrate species that do not yet have reference DNA sequences in databases cannot be identified. This might be a particularly important problem for biodiversity hotspots of the global south70,71. For example, Fahmy et al. (2024)72 survey of vertebrate diversity in Madagascar employing leech iDNA analysis resulted in a large proportion of detections being identified only to family level.
Vertebrate survey methods are biased
An important issue raised by our analysis is that these methods can portray different communities by displaying higher detection rates for different species. There were differences among the four community samples from the DeLuca preserve (dry-MBA, wet-MBA, dry-conv., wet-conv.) and the largest differences were between the samples obtained by MBA and those obtained by conventional methods regardless of the season. These differences were driven by MBA and conventional methods more efficiently detecting different species. For example, the lack of representation of small mammals, and larger representation of wetland birds in the MBA samples influenced this difference. Thus, great care should be exercised when comparing communities sampled with different methods including emerging eDNA and iDNA tools73,74,75. In this line, also great care should be exercised when interpreting relative abundances obtained from MBA given that besides hosts abundance, other factors such as nutritional value of the blood or chemical cues might significantly influence host choice76,77. These results also suggest potential benefits of simultaneously using conventional methods and MBA to survey communities as these will target different species. It remains to be determined if larger sampling effort by both methods, and under wet conditions could result in converging samples of the community.
Future prospects
Scalable and more efficient ways to measure biodiversity are urgently needed. This is especially true if the idea of selling biodiversity credits to fund the conservation budget gap is to take off. One of the most important challenges to these initiatives is to efficiently and reliably measure biodiversity improvement from investment78, and here, MBA could provide a timely, streamlined and uniform way to gather information on entire vertebrate communities. Nonetheless, we show that in some circumstances, conventional methods can be more efficient than even the most successful MBA36, and for this reason iDNA methods validation against conventional sampling under different conditions is of paramount importance. On the other hand, our work shows that with as little as 20 h of field work, relevant samples of a community can be obtained through MBA under the right conditions, a positive indication of the scalability of this technique. The efficiency of mosquito blood meal analysis could be further improved by implementing a metabarcoding approach to identify the hosts of bulk samples of mosquito blood meals or refine the tool to target the mosquito species feeding on the highest diversity of hosts. Such an approach may reduce laboratory times by eliminating the time-consuming steps associated with individually preserving blood meals and individually extracting DNA. Metabarcoding might not only increase efficiency, but also the sample size by detecting multiple blood meals per individual hematophagous insect. Furthermore, recent work by Mwakasungula et al. (2022)79 and Johnson et al. (2024)80 provide interesting examples of how a metabarcoding approach could be extended to identify vertebrate hosts, the pathogens they carry, and the vectors involved to advance surveillance and our understanding of disease systems. Finally, another application of MBA that deserves further assessment is the use of MBA to target species of management or conservation importance while obtaining biological samples for downstream analysis such as individual genotyping for abundance estimation through mark-recapture studies.
Data availability
A detailed description of the effort employed by each method is provided in the Supplementary Materials 1 file. The data on the number of detections, incidence and time to first detection of each species by each method-season combination are included in the Supplementary Materials 2 file. In addition, the Cytochrome C oxidase subunit I sequence data for detected vertebrate hosts are available on the National Center for Biotechnology Information GenBank database via accession numbers: PV786364 – PV786446. Finally, the raw data employed in this study, including both the mosquito blood meal analyses (collection method, site, date, mosquito identification, and vertebrate species identified) and the results from the conventional methods surveys (records obtained by each method, site and date) is available at the following figshare repository: https://figshare.com/articles/dataset/Vertebrate_species_detection_by_mosquito_bloodmeal_analysis_and_conventional_methods_at_the_DeLuca_Preserve_in_Central_Florida/29847341/1.
References
Ceballos, G. et al. Accelerated modern human–induced species losses: Entering the sixth mass extinction. Sci. Adv. 1, e1400253 (2015).
O’Connor, B., Bojinski, S., Röösli, C. & Schaepman, M. E. Monitoring global changes in biodiversity and climate essential as ecological crisis intensifies. Eco. Inform. 55, 101033 (2020).
Pereira, H. M. & Cooper, H. D. Towards the global monitoring of biodiversity change. Trends Ecol. Evol. 21, 123–129 (2006).
Xu, H. et al. Ensuring effective implementation of the post-2020 global biodiversity targets. Nat. Ecol. Evol. 5, 411–418 (2021).
Gonzalez, A. et al. A global biodiversity observing system to unite monitoring and guide action. Nat. Ecol. Evol. 7, 1947–1952 (2023).
Hughes, A. C. The Post-2020 Global Biodiversity Framework: How did we get here, and where do we go next?. Integr. Conserv. 2, 1–9 (2023).
Ford, H. V. et al. A technological biodiversity monitoring toolkit for biocredits. J. Appl. Ecol. 61, 2007–2019 (2024).
Sutherland, W. J. (ed.) Ecological Census Techniques: A Handbook (Cambridge University Press, 2006).
Perkins, G. C., Kutt, A. S., Vanderduys, E. P. & Perry, J. J. Evaluating the costs and sampling adequacy of a vertebrate monitoring program. Aust. Zool. 36, 373–380 (2013).
Massey, A. L. et al. Invertebrates for vertebrate biodiversity monitoring: Comparisons using three insect taxa as iDNA samplers. Mol. Ecol. Resour. 22, 962–977 (2022).
Pascher, K., Švara, V. & Jungmeier, M. Environmental DNA-based methods in biodiversity monitoring of protected areas: Application range, limitations, and needs. Diversity 14, 463 (2022).
Schulze, C. H. et al. Biodiversity indicator groups of tropical land-use systems: Comparing plants, birds, and insects. Ecol. Appl. 14, 1321–1333 (2004).
Zhang, J. et al. Trophic interactions among vertebrate guilds and plants shape global patterns in species diversity. Proc. R. Soc. B Biol. Sci. 285, 20180949 (2018).
Ceballos, G., Ehrlich, P. R. & Raven, P. H. Vertebrates on the brink as indicators of biological annihilation and the sixth mass extinction. Proc. Natl. Acad. Sci. 117, 13596–13602 (2020).
Luedtke, J. A. et al. Ongoing declines for the world’s amphibians in the face of emerging threats. Nature 622, 308–314 (2023).
Carvalho, C. S., De Oliveira, M. E., Rodriguez-Castro, K. G., Saranholi, B. H. & Galetti, P. M. Jr. Efficiency of eDNA and iDNA in assessing vertebrate diversity and its abundance. Mol. Ecol. Resour. 22, 1262–1273 (2022).
Abrams, J. F. et al. Shifting up a gear with iDNA: From mammal detection events to standardised surveys. J. Appl. Ecol. 56, 1637–1648 (2019).
Van Leeuwen, P. & Michaux, J. Using eDNA for mammal inventories still needs naturalist expertise, a meta-analysis. Ecol. Evol. 13, e10788 (2023).
Reeves, L. E. et al. Identification of Uranotaenia sapphirina as a specialist of annelids broadens known mosquito host use patterns. Commun. Biol. 1, 92 (2018).
Edman, J. D. Host-feeding patterns of Florida mosquitoes (Diptera: Culicidae) VI. Culex (Melanoconion). J. Med. Entomol. 15, 521–525 (1979).
Edman, J. D. Host-feeding patterns of Florida mosquitoes I. Aedes, anopheles, coquillettidia, Mansonia and Psorophora. J. Med. Entomol. 8, 687–695 (1971).
Edman, J. D., Webber, L. A. & Kale, H. W. Host-feeding patterns of Florida mosquitoes II. Culiseta. J. Med. Entomol. 9, 429–434 (1972).
Omernik, J. M. & Griffith, G. E. Ecoregions of the conterminous United States: Evolution of a hierarchical spatial framework. Environ. Manag. 54, 1249–1266 (2014).
NOAA. U.S. Climate Normals. Kissimmee Prairie Preserve Station. https://www.ncei.noaa.gov/access/us-climate-normals/#dataset=normals-monthly&timeframe=15&location=FL&station=USC00084621 (2024).
FAWN. Florida Automated Weather Network. https://fawn.ifas.ufl.edu/ (2024).
Sloyer, K. E. et al. Evaluating sampling strategies for enzootic Venezuelen equine encephalitis virus vectors in Florida and Panama. PLoS Negl. Trop. Dis. 16, e0010329 (2022).
Burkett-Cadena, N. D., Eubanks, M. D. & Unnasch, T. R. Preference of female mosquitoes for natural and artificial resting sites. J. Am. Mosq. Control Assoc. 24, 228–235 (2008).
Burkett-Cadena, N. D., Hoyer, I., Blosser, E. & Reeves, L. E. Human-powered pop-up resting shelter for sampling cavity-resting mosquitoes. Acta Trop. 190, 288–292 (2019).
Nasci, R. S. A lightweight battery-powered aspirator for collecting resting mosquitoes in the field. Mosq. News 41, 808–811 (1981).
Darsie, R. F. & Ward, R. A. Identification and Geographical Distribution of the Mosquitoes of North America (University Press of Florida, 2005).
Reeves, L. E., Holderman, C. J., Gillett-Kaufman, J. L., Kawahara, A. Y. & Kaufman, P. E. Maintenance of host DNA integrity in field-preserved mosquito (Diptera: Culicidae) blood meals for identification by DNA barcoding. Parasit. Vectors 9, 503 (2016).
Truett, G. E. et al. Preparation of PCR-quality mouse genomic DNA with hot sodium hydroxide and tris (HotSHOT). Biotechniques 1, 52–54 (2000).
Reeves, L. E., Gillett-Kaufman, J. L., Kawahara, A. Y. & Kaufman, P. E. Barcoding blood meals: New vertebrate-specific primer sets for assigning taxonomic identities to host DNA from mosquito blood meals. PLoS Negl. Trop. Dis. 12, e0006767 (2018).
Sanger, F., Nicklen, S. & Coulson, A. R. DNA-sequencing with chain-terminating inhibitors. Proc. Natl. Acad. Sci. 74, 5463–5467 (1977).
Ratsnasingham, S. & Hebert, P. D. N. BOLD: The barcode of life data system (www.barcodinglife.org). Mol. Ecol. Notes 7, 355–364 (2007).
Atsma, H., Burkett-Cadena, N. D., Wisely, S. M., Urlaub, A., Botero-Cañola, S. & Reeves, L. E. In review. Monitoring biodiversity and detection of diverse vertebrate species with mosquito blood meal analysis at the DeLuca Preserve, Florida, USA. Sci. Rep.
Botero-Cañola, S. et al. Acoustic indices track local vertebrate biodiversity in a subtropical landscape. Ecol. Ind. 166, 112292 (2024).
Larsen, T. H. (ed.) Core Standardized Methods for Rapid Biological Field Assessment (Conservation International, 2016).
Drake, A. et al. Combining point counts and autonomous recording units improves avian survey efficacy across elevational gradients on two continents. Ecol. Evol. 11, 8654–8682 (2021).
Enge, K. M. A standardized protocol for drift-fence surveys (Florida Game and Fresh Water Fish Commission, 1997).
Campbell, K. R., Campbell, T. S. & Johnson, S. A. The use of PVC pipe refugia to evaluate spatial and temporal distributions of native and introduced treefrogs. Florida Sci. 73, 78–88 (2010).
Tobler, M. W., Carrillo-Percastegui, S. E., Leite Pitman, R., Mares, R. & Powell, G. An evaluation of camera traps for inventorying large-and medium-sized terrestrial rainforest mammals. Anim. Conserv. 11, 169–178 (2008).
Chao, A. & Jost, L. Coverage-based rarefaction and extrapolation: Standardizing samples by completeness rather than size. Ecology 93, 2533–2547 (2012).
Chao, A. et al. Rarefaction and extrapolation with Hill numbers: A framework for sampling and estimation in species diversity studies. Ecol. Monogr. 84, 45–67 (2014).
Roswell, M., Dushoff, J. & Winfree, R. A conceptual guide to measuring species diversity. Oikos 130, 321–338 (2021).
Hsieh, T. C., Ma, K. & Chao, A. iNEXT: An R package for rarefaction and extrapolation of species diversity (Hill numbers). Methods Ecol. Evol. 7, 1451–1456 (2016).
Gorta, S. B. et al. Multi-taxon biodiversity responses to the 2019–2020 Australian megafires. Glob. Change Biol. 29, 6727–6740 (2023).
Florida Fish and Wildlife Conservation Commission (FWC). Florida’s Wildlife Legacy Initiative: Florida’s State Wildlife Action Plan. Tallahassee, Florida (2019).
Oksanen J., Simpson G., Blanchet F., Kindt R., Legendre P., Minchin P., O'Hara R., Solymos P., Stevens M., Szoecs E., Wagner H., Barbour M., Bedward M., Bolker B., Borcard D., Carvalho G., Chirico M., De Caceres M., Durand S., Evangelista H., FitzJohn R., Friendly M., Furneaux B., Hannigan G., Hill M., Lahti L., McGlinn D., Ouellette M., Ribeiro Cunha E., Smith T., Stier A., Ter Braak C. & Weedon J. 2022. Vegan: Community Ecology Package. R package version 2.6-4. https://CRAN.R-project.org/package=vegan.
Atsma, H. Mosquito bloodmeal analysis for biodiversity monitoring at a Florida preserve. Masters thesis, University of Florida, Vero Beach, FL. https://ufdc.ufl.edu/ufe0060708/00001 (2023).
Danabalan, R. et al. Comparison of mosquito and fly derived DNA as a tool for sampling vertebrate biodiversity in suburban forests in Berlin, Germany. Environ. DNA 5, 476–487 (2023).
Saranholi, B. H. et al. Comparing iDNA from mosquitoes and flies to survey mammals in a semi-controlled Neotropical area. Mol. Ecol. Resour. 23, 1790–1799 (2023).
Kocher, A. et al. iDNA screening: Disease vectors as vertebrate samplers. Mol. Ecol. 26, 6478–6486 (2017).
Day, C. A. & Trout Fryxell, R. T. Are they there, how many, and how big? Investigating potential trap biases in the surveillance of La Crosse virus vectors. J. Med. Entomol. 62, 189–198 (2025).
Gogarten, J. F. et al. Fly-derived DNA and camera traps are complementary tools for assessing mammalian biodiversity. Environ. DNA 2, 63–76 (2020).
Cristescu, M. E. & Hebert, P. D. Uses and misuses of environmental DNA in biodiversity science and conservation. Annu. Rev. Ecol. Evol. Syst. 49, 209–230 (2018).
Franklin, D. C. & Whelan, P. I. Tropical mosquito assemblages demonstrate ‘textbook’ annual cycles. PLoS ONE 4, e8296 (2009).
de Melo Freire, R. C. et al. Ecological aspects of mosquitoes (Diptera: Culicidae) in a fragment of seasonal dry tropical forest (Caatinga) in Brazil. J. Arid Environ. 190, 104528 (2021).
Bates, M. Observations on climate and seasonal distribution of mosquitoes in eastern Colombia. J. Anim. Ecol. 14, 17–25 (1945).
Kays, R. et al. An empirical evaluation of camera trap study design: How many, how long and when?. Methods Ecol. Evol. 11, 700–713 (2020).
Ryan, T. J. et al. Monitoring herpetofauna in a managed forest landscape: Effects of habitat types and census techniques. For. Ecol. Manag. 167, 83–90 (2002).
Ali, W. et al. Comparison of different trapping techniques used in herpetofaunal monitoring: A review. Punjab Univ. J. Zool. 33, 57–68 (2018).
Bianchini, K., Frei, B., Cox, A. R. & Roy, C. Evaluating occurrence and abundance of displaying male American woodcock (Scolopax minor) north of the current Singing-Ground Survey range. J. Wildl. Manag. 89, 70018 (2025).
Engeman, R. M. et al. Eradication of feral swine from a barrier island in Florida, USA: An examination of effort and multi-method, multi-species population indexing. NeoBiota 93, 91–116 (2024).
Goldberg, N. Distribution of the Cuban treefrog (Anura, Hylidae, Osteopilus septentrionalis) in Florida using geolocated records from an invasive species monitoring app. Manag. Biol. Invas. 15, 445–454 (2024).
Stewart, D. R. et al. Assessing trends and density of bird species in bottomland hardwood forests and riparian forests using simulation and sample size optimization for surveys. Sci. Rep. 15, 7137 (2025).
Nguyen, T. V. et al. Using terrestrial leeches to assess the genetic diversity of an elusive species: The Annamite striped rabbit Nesolagus timminsi. Environ. DNA 3, 780–791 (2021).
Vieira, C. J. S. et al. Mosquito bloodmeals can be used to determine vertebrate diversity, host preference, and pathogen exposure in humans and wildlife. Sci. Rep. 14, 23203 (2024).
Wisely, S. M., Torhorst, C. W., Atsma, H., Botero-Canola, S. Burkett-Cadena, N. B. & Reeves, L. E. Xenosurveillance of wild pigs using mosquito bloodmeals. In Proceedings of the 31st Vertebrate Pest Conference (2024).
Lim, B. K. Preliminary assessment of Neotropical mammal DNA barcodes: an underestimation of biodiversity. Open Zool. J. 5, 10–17 (2022).
Siziba, V. I. & Willows-Munro, S. An assessment of South African small mammal barcode sequence libraries: Implications for future carnivore diet analyses by DNA. Afr. J. Ecol. 62, e13233 (2024).
Fahmy, M., Andrianoely, D., Wright, P. C. & Hekkala, E. Leech-derived iDNA complements traditional surveying methods, enhancing species detections for rapid biodiversity sampling in the tropics. Environ. DNA 5, 1557–1573 (2023).
Cao, Y., Hawkins, C. P. & Storey, A. W. A method for measuring the comparability of different sampling methods used in biological surveys: Implications for data integration and synthesis. Freshw. Biol. 50, 1105–1115 (2005).
Quilumbaquin, W., Carrera-Gonzalez, A. & Ortega-Andrade, H. M. Environmental DNA and visual encounter surveys for amphibian biomonitoring in aquatic environments of the Ecuadorian Amazon. PeerJ 11, e15455 (2023).
Robinson, K. M. et al. Paired environmental DNA and dive surveys provide distinct but complementary snapshots of marine biodiversity in a temperate fjord. Environ. DNA 5, 597–612 (2023).
Lyimo, I. N. & Ferguson, H. M. Ecological and evolutionary determinants of host species choice in mosquito vectors. Trends Parasitol. 25, 189–196 (2009).
Yan, J. et al. Understanding host utilization by mosquitoes: determinants, challenges and future directions. Biol. Rev. 96, 1367–1385 (2021).
Antonelli, A., Rueda, X., Calcagno, R. & Nantongo Kalunda, P. How biodiversity credits could help to conserve and restore nature. Nature 634, 1045–1049 (2024).
Mwakasungula, S. et al. Using haematophagous fly blood meals to study the diversity of blood-borne pathogens infecting wild mammals. Mol. Ecol. Resour. 22, 2915–2927 (2022).
Johnson, B., Graham M., Panahi E., Vieira C., Mason P., Gleadhill J., Thomas D., Onn M., Shivas M., Shearman D. & Darbro J. An all-in-one metabarcoding approach to mosquito and arbovirus xenosurveillance (2024).
Acknowledgements
We would like to acknowledge the UF/IFAS DeLuca Preserve for hosting us through this work and the UF/IFAS Office of the Dean for Research, DeLuca Preserve/Forest Systems Jumpstart Funding Program for providing the funding for this work. We want to thank Jorge Poll, Brian Beasley and Brent Sellers for their help and support through the field phase of this study. We would also like to thank the help and enthusiasm of the field and lab technicians Marina McCampbell, Chanakya R. Bhosale, Jose Cosme, Delana Gonzales, Maggie Anderson, Alexander Urlaub and Lilyanna Lopez.
Author information
Authors and Affiliations
Contributions
L.E.R., N.D.B-C., S.M.W., K.E.S., and C.R. devised the project. S.M.W., K.E.S., C.R., N.D.B-C. and L.E.R. supervised the project. L.E.R. and S.B-C. supervised field activities. K.W., M.H., E.G. and S.B-C conducted conventional vertebrate surveys. L.E.R, N.D.B-C. and H.A. collected bloodmeals, morphologically identified the mosquitoes, extracted and sequenced DNA and identified the bloodmeals. S.B-C. analyzed the data. S. B-C., S.M.W, K.E.S. and N.D.B-C. drafted the manuscript and designed the figures and tables. All authors edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Botero-Cañola, S., Atsma, H., Wilson, K. et al. A comparison of conventional methods and mosquito blood meal analysis to assess vertebrate diversity. Sci Rep 15, 42398 (2025). https://doi.org/10.1038/s41598-025-26514-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-26514-y