Abstract
SKP2 (S-phase kinase-associated protein 2) plays an important role in the regulation of the cell cycle, tumorigenesis, progression, and metastasis. However, its functional role in gliomas is currently unclear. This study aimed to investigate the relationship between SKP2 and the progression of gliomas, and its effects on the glioma cell cycle. The relationship between SKP2 expression levels and clinical characteristics of gliomas was explored in our datasets. Kaplan–Meier analysis and COX regression were conducted to evaluate the prognostic role of SKP2. Differentially expressed genes were identified between high and low SKP2 groups. The function and related mechanisms of SKP2 were evaluated based on Gene Ontology and Gene Set Enrichment Analysis. LASSO-Cox was used to screen the variables, and the prediction model was established by nomogram analysis. The effects of SKP2 on the glioma cell cycle were examined in cellular experiments. The SKP2 expression level was positively correlated with adverse clinical features of gliomas, and overall survival with high SKP2 expression was significantly shortened. Gene enrichment analysis showed that the differential genes in the SKP2 high-expression group were mainly enriched in cell cycle and mitosis. In vitro experiments confirmed that silencing of SKP2 arrested the glioma cell cycle at the G1 phase in both U87 and U251 cells. In conclusion, SKP2 may serve as a potential prognostic biomarker for glioma patients and is associated with poor prognosis. Additionally, SKP2 is closely related to the glioma cell cycle, suggesting that SKP2 may serve as a potential therapeutic target. However, further prospective studies and external cohort validations are needed to confirm its clinical applicability.
Similar content being viewed by others
Introduction
Glioma is the most common primary occupying tumor of the central nervous system in adults. The fifth edition of the World Health Organization’s Classification of Tumors of the Central Nervous System, revised in 2021, classifies gliomas into four grades1, of which the prognosis of low-grade gliomas is relatively good, while high-grade gliomas are fast-growing, short-course and have a poor prognosis2. The treatment of glioma is mainly a combination of surgical resection, radiotherapy and chemotherapy3. However, it is extremely difficult to completely remove the tumor without damaging normal brain tissue. Since the genome of malignant glioma cells is very unstable, it is easy to develop drug resistance, which greatly reduces the effectiveness of radiotherapy and chemotherapy treatments4.
Tumor formation is multifactorial, multistep and multistage. From a biological point of view, the growth of tumor cells is mainly regulated by the cell cycle. The cell cycle can be divided into five periods, G1, S, G2, M, and G0, and is regulated by the cell cycle protein cyclin, cyclin dependent kinase (CDK), CDK-activating kinase (CAK), and CDK inhibitor (CKI)5,6. Among them, CDK is the most central protein, which can activate different cell cycle proteins-CDK complexes and phosphorylate their target proteins at specific stages of the cell cycle7. There are already cell cycle inhibitors that use the target of CDK as an entry point, the most successful being the inhibitors of CDK4 and CDK68, which have made breakthroughs in the treatment of breast cancer9. Despite the success of CDK4/6 inhibitors, tumor therapy targeting cell cycle proteins is still in its infancy. The role of cell cycle proteins within tumor cells and in the anti-tumor immune response will be the subject of intensive research for tumor therapy in the future. Therefore, we need more studies to explore the mechanisms that regulate the cell cycle of gliomas and to find possible points of intervention to improve the efficacy of glioma treatment.
The SKP2 gene is closely related to cell cycle regulation, and many cell cycle regulators such as p27, p21, p53, and Cyclin E are ubiquitin proteasome pathway. SKP2 can specifically recognize phosphorylated substrates and mediate their ubiquitination degradation. SKP2 is also involved in the ubiquitination degradation rate of smad4, an important protein factor downstream of the TGF-β signaling pathway10. The degradation of the transcription factor E2F-1 in the S-G2 phase is related to SKP2, and the interaction of SKP2 with E2F-1 can prevent E2F-1 from degrading in the S-G2 phase11. Myc, as an oncoprotein transcription factor, is associated with cell growth, differentiation, proliferation and tumorigenesis, and SKP2 both enhances C-myc-induced cellular S-phase transition and activates C-myc target genes, and also participates in Myc-induced transcription12.
Although the oncogenicity of SKP2 has been demonstrated in many other tumors, there is a lack of comprehensive analysis of SKP2 in gliomas. In this study, we aimed to investigate the expression profiles of SKP2 in different gliomas and further explored whether SKP2 expression was associated with clinicopathological features and survival of glioma patients, and investigate the relationship between SKP2 and cell cycle regulation as well as tumorigenesis, progression and prognosis based on the Chinese Glioma Genome Atlas (CGGA) and The Cancer Genome Atlas (TCGA) databases. In addition, we verified the role of SKP2 in the gliomas’ cell cycle, and our results provide powerful evidence for the potential therapeutic role of SKP2 regarding glioma.
Data and methodologies
Patients and datasets
Our study used data from two publicly available databases to analyze gliomas: the Cancer Genome Atlas (TCGA) and the Chinese Glioma Genome Atlas (CGGA). The TCGA database provided RNA-seq data and clinical information on 702 patients, and CGGA served as the validation cohort. The CGGA database has two RNA-seq datasets as well as relevant clinical information (CGGA693 data for cohort 1 and CGGA325 data for cohort 2).
Relationship between SKP2 and clinical characteristics
To analyze the correlation between SKP2 gene expression levels and typical prognostic features, World Health Organization classification (grade 1–4), IDH wild type/mutant type, primary/recurrent, 1p/19q codeletion/non-codeletion, age, gender, and MGMT promoter methylation were compared with SKP2 expression. Furthermore, glioma patients were stratified into high- and low-expression groups based on the median SKP2 expression level, and overall survival was assessed using Kaplan–Meier analysis.
Functional enrichment analysis
Spearman correlation analysis was used to screen genes related to SKP2 (|Spearman R|> 0.5 and p < 0.05). A list of the most relevant genes or cell clusters associated with SKP2 was uploaded to Metascape, using official gene symbols as identifiers and restricting the species to human. Functional enrichment was then performed, including Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analyses13,14. Pathway data were obtained from the KEGG database (Kanehisa Laboratories). In addition, Gene and Gene Set Enrichment Analysis (GSEA) was applied to assess gene expression patterns at the gene set level, providing more comprehensive insights into enriched pathways.
Construction of a predictive risk scoring model
Differentially Expressed Genes (DEGs) were compared between the SKP2 high and low expression groups using the “samr” package of the R software, and DEGs were filtered by |Log2 FC|> 2 and p < 0.05. If the DRGs identified in the training set TCGA could not be verified in the validation set CGGA, only common DRGs expressed in both TCGA and CGGA datasets were selected for survival analysis. Univariate Cox proportional regression analyses were performed to calculate the relationship between the expression of the differential genes and the overall survival (OS) in the TCGA dataset. Next, we performed the Least Absolute Shrinkage and Selection Operator (LASSO) and selected nine prognostic genes to construct a prognostic risk model. Finally, risk score was calculated using the following formula (Risk score = (β1 × HOXC10) + (β2 × OR51E1) + (β3 × HOXC6) + (β4 × HOXD11) + (β5 × HMGA2) + (β6 × HOXA3) + (β7 × GALNT5) + (β8 × HOXA5) + (β9 × SHOX2)).
Constructing the nomogram
Nomogram analysis was performed using the “rms” package in R software. As shown in Fig. 8, the upper panels represent the scoring system and the lower panels display the prediction system. The 1-, 3-, and 5-year survival probabilities for glioma patients can be predicted from the individual factor scores and the total score. Calibration curves and the concordance index (C-index) were used to evaluate the accuracy of the survival predictions.
Cell culture
U87 and U251 cell lines were obtained from the Chinese Academy of Medical Science (China, Beijing). The U87 cell line is glioblastoma of unknown origin and has been identified as ATCC type by STR profiling. The cells were cultured in Dulbecco’s Modified Eagle’s Medium (DMEM, Thermo Fisher Scientific, Rockford, IL, USA) medium with the addition of 10% fetal bovine serum (FBS, Thermo Fisher Scientific) and 1% penicillin and streptomycin, and stored in a humidified incubator at 37 °C with 5% CO2.
Transient transfection
SKP2 siRNA1/2 or empty vector plasmid were purchased from Shanghai Jimma Biotechnology Co. U87 and U251 cells were then transfected with Lipofectamine 3000 according to the instructions. The medium was changed 6 h after transfection. The cells were collected 48 h after transfection for subsequent experiments. The SKP2 siRNA sequences are as following:
(si-SKP2-1) Sense: 5’-GGA GUG ACA AAG ACU UUG UTT-3’;
Antisense: 5’-ACA AAG UCU UUG UCA CUC CTT-3’.
(si-SKP2-2) Sense: 5’-GGA UGA GGA GAG UUU UGG UUU-3’;
Antisense: 5’-AAA CCA AAA CUC UCU CUC AUU-3’.
Western blot assay
Glioma cells were lysed with RIPA lysis extraction buffer (Thermo Fisher Scientific) and protease inhibitor mixture (Thermo Fisher Scientific). Protein concentrations were determined using the Pierce BCA Protein Assay Kit (Thermo Fisher Scientific). Cell lysates containing 30 μg of protein were separated by sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) and transferred to a polyvinylidene fluoride (PVDF) membrane. The membrane was blocked in 5% fetal bovine serum and incubated overnight at 4 °C with the following primary antibodies: SKP2 (Rabbit mAb, Abconal, A4046, 1:1000), CDK2 (Proteintech, 10,122–1-AP, 1:2000), Cyclin E1 (Affinity, AF0144, 1:1000), and β-actin (Proteintech, 20,536–1-AP, 1:5000). After washing, membranes were incubated with HRP-conjugated goat anti-rabbit IgG (H + L) secondary antibody (Abconal, AS014, 1:10,000) for 1.5 h at room temperature. Finally, protein bands were visualized using the Clarity Max Western ECL Substrate (Bio-Rad), and densitometric analysis was performed with Quantity One Software (Bio-Rad). All Western blot experiments were performed in at least three independent biological replicates.
Cell cycle analysis by PI/RNase staining
Cell cycle distribution was analyzed using a PI/RNase cell cycle assay kit (4A Biotech). Briefly, 1 × 106 U87 or U251 cells were harvested, washed twice with pre-cooled PBS, and fixed in 70% cold ethanol at 4 °C overnight. The fixed cells were then washed with PBS and incubated with RNase A (100 μg/mL) at 37 °C for 30 min to remove RNA. Subsequently, cells were stained with propidium iodide (PI, 50 μg/mL) for 30 min at room temperature in the dark. DNA content was measured by flow cytometry within 1 h, and the percentages of cells in G0/G1, S, and G2/M phases were analyzed using FlowJo 10.6.2 software.
Lentivirus transduction
The short hairpin RNA (shRNA) targeting human SKP2 (sh-SKP2) and the negative control shRNA (sh-NC) were designed and synthesized by Shanghai Jimma Biotechnology Co., Ltd. The sh-SKP2 sequence was designed based on the most efficient siRNA sequence (si-SKP2-1). All lentiviral particles were packaged in the pLenti-GIII-CMV-GFP-2A-Puro vector (LV, Shanghai Jimma Biotechnology Co., Ltd.), which also carries the luciferase (LUC) reporter gene.
In vivo experiments
Male BALB/c nude mice (5-weeks-old) were purchased from SPF (Beijing) Biotechnology Co., Ltd. (license: SCXK (Jing) 2024–0001). Ethical clearance for all animal experiments was obtained from the Ethics Committee of Peking University People’s Hospital (Approval No. 2021-PHE072). All procedures strictly followed the Guide for the Care and Use of Laboratory Animals (National Institutes of Health, USA). U87 glioma cells were stably transduced with either NC-LUC (sh-NC) (negative control lentivirus carrying luciferase) or sh-SKP2-LUC (sh-SKP2 lentivirus carrying luciferase). The sh-SKP2 construct was designed based on the si-SKP2-1 sequence, which showed the highest knockdown efficiency in vitro. A total of 5 × 106 cells were injected subcutaneously into the flanks of each mouse (n = 6). Tumor growth was monitored by in vivo fluorescence imaging (IVIS system) on day 25. Mice were anesthetized with 2% isoflurane prior to imaging. At the end of the experiments, animals were euthanized by intraperitoneal injection of an overdose of sodium pentobarbital (≥ 150 mg/kg body weight) under deep anesthesia, in accordance with institutional guidelines. Death was confirmed before tissue collection. Tumors were then excised for measurement of volume and weight, as well as western blot analysis of SKP2, Cyclin E, and CDK2 expression. All animal experiments were conducted and reported in accordance with the ARRIVE guidelines (https://arriveguidelines.org)15.
Statistical analysis
All statistical analyses were performed using R software (version 4.0.0), GraphPad Prism 9.0 (GraphPad Inc., San Diego, CA, USA) or SPSS 18.0 (IBM, Chicago, IL, USA). Data are presented as mean ± SEM from at least three independent biological replicates unless otherwise specified. Unpaired t-tests were used to assess differences between two groups, and one-way ANOVA followed by Tukey’s post-hoc test was applied for comparisons among more than two groups. Spearman correlation analysis was used to evaluate associations between variables, with correlation coefficients (r) and p-values reported. For Kaplan–Meier survival analysis, patients were stratified into high and low expression groups using the median SKP2 expression level as the cutoff value. The log-rank test was performed to assess differences in overall survival between groups. Univariate and multivariate Cox regression analyses were conducted to identify prognostic factors. To ensure model validity, variance inflation factor (VIF) analysis was performed to confirm the absence of significant multicollinearity among covariates (all VIF values < 5). Hazard ratios (HR) with 95% confidence intervals (CI) and exact p-values are reported. For all statistical analyses, p-values are indicated in the figures with the following thresholds for statistical significance: *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. Asterisks in the figures correspond to these p-value thresholds to indicate statistical significance. P-values less than 0.05 were considered statistically significant.
Results
SKP2 expression in different tumors
Using the online analysis tool GEPIA, we compared SKP2 expression between tumor and paired normal tissues across TCGA and GTEx datasets. The results showed that SKP2 expression was generally elevated in multiple tumor types compared with paired normal tissues (Fig. 1A). In gliomas, SKP2 mRNA expression was significantly upregulated in GBM compared with paired normal brain tissues (GBM, n = 163; normal brain, n = 207; p < 0.05), whereas no significant difference was observed in LGG (LGG, n = 518; normal brain, n = 207; Fig. 1B).
Expression profile of SKP2 in different tumor samples and normal tissues. (A) GEPIA analysis of TCGA and GTEx datasets showed that SKP2 expression was generally elevated in multiple tumor types compared with normal tissues. (B) In glioma, SKP2 mRNA levels were significantly higher in GBM compared with normal brain tissues, while the difference in LGG was not statistically significant.*p < 0.05.
SKP2 expression is closely associated with clinical features of gliomas
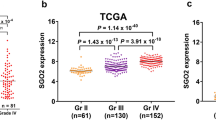
Patients with different SKP2 expression levels exhibited different clinical and pathological features. SKP2 expression, MGMT promoter methylation status, 1p/19q codeletion status, IDH mutation status, WHO grading, and increased histological diagnosis showed an asymmetric distribution in the CGGA and TCGA datasets (Fig. 2A, B). Comparing the different subtypes of patients separately, in the TCGA database, SKP2 was more highly expressed in samples with high-grade gliomas (Fig. 2C), IDH -wild-type gliomas (Fig. 2D), without 1p/19q codeletion (Fig. 2E), and MGMT-unmethylated (Fig. 2F). These results were validated in the CGGA database, but the difference between MGMT and non-MGMT in CGGA was not statistically significant, probably due to the earlier year of patient inclusion in the TCGA database (Fig. 2G, J; Supplementary Fig. 1). These results suggest that more malignant gliomas are enriched for SKP2, that SKP2 is statistically correlated with conventional molecular pathological factors in gliomas, and that specific molecular pathological features are presented in high-grade gliomas.
Relationship between SKP2 and clinicopathologic features and prognosis of glioma. (A) Characteristics of the overall distribution of SKP2-related clinicopathological features of glioma in The Cancer Genome Atlas (TCGA) dataset. (B) The overall distribution characteristics of glioma SKP2-related clinicopathological features in the Chinese Glioma Genome Atlas (CGGA) dataset. (C) and (G), SKP2 was significantly elevated in high-grade gliomas in the TCGA and CGGA Cohort1 datasets. (D) and (H), in the TCGA and CGGA Cohort1 datasets, SKP2 was significantly elevated in gliomas without isocitrate dehydrogenase (IDH) mutations. (E) and (I), SKP2 was significantly increased in gliomas without 1p/19q codeletion in the TCGA and CGGA Cohort1 datasets. (F) and (J), SKP2 is elevated in gliomas unmethylated with the O6-methylguanine-DNA methyltransferase (MGMT) promoter. This difference was statistically significant in the TCGA dataset but not in the CGGA Cohort1 dataset.
High SKP2 expression levels correlate with poor prognosis in gliomas
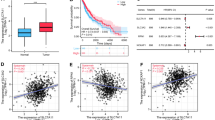
To investigate the prognostic value of SKP2 in glioma patients, we performed Kaplan–Meier analysis based on CGGA and TCGA databases. In the TCGA database, the overall survival of patients with high SKP2 expression was significantly shorter than those with low SKP2 expression (Fig. 3A), and the results of CGGA cohort 1 and CGGA cohort 2 were consistent with those of TCGA (Fig. 3D; Supplementary Fig. 2A). In the univariate analysis of the TCGA database, MGMT status, 1p/19q status, IDH status, age, WHO grade, and SKP2 were all associated with patient prognosis (Fig. 3B), and this result was verified in the CGGA database (Fig. 3E; Supplementary Fig. 2B). In the multivariate analysis of the TCGA database, MGMT status, 1p/19q status, IDH status, age, WHO grade, and SKP2 were included in the model, and the p-value for SKP2 remained < 0.05, suggesting that SKP2 is an independent prognostic factor (Fig. 3C). Similar findings were validated in the CGGA cohort 1 and CGGA325 cohort 2 (Fig. 3F; Supplementary Fig. 1C) (Supplementary Tables 1, 2 and 3).
Relationship between SKP2 and prognosis of glioma. (A) and (D), Kaplan–Meier analysis of SKP2 expression in TCGA and CGGA Cohort1 datasets. (B), (C), (E) and (F), Forest plot showing univariate and multivariate Cox regression analysis of SKP2 and clinicopathological features in TCGA and CGGA Cohort1 datasets.
SKP2 expression is closely related to the cell cycle
To further analyze the potential functions and pathways of the obtained candidate genes, functional enrichment analysis was conducted. GO enrichment analysis revealed the most significant enriched functions and pathways in TCGA, including “Cell Cycle,” “positive regulation of cell cycle process,” “meiotic cell cycle,” “cell cycle phase transition,” “Transcriptional Regulation by TP53,” “E2F PATHWAY,” etc. (Fig. 4A). GSEA results further confirmed these findings, with "Cell Cycle," "DNA replication," "p53 signaling pathway," and "hallmark E2F targets" pathways being significantly enriched in TCGA (Fig. 4B). Pathways enriched in the CGGA cohort 1 include “Cell Cycle, Mitotic,” “DNA replication,” “VEGFA-VEGFR2 signaling pathway,” “EGF/EGFR signaling pathway,” “regulation of I-kappaB kinase/NF-kappaB signaling,” etc. (Fig. 4C). Similar to TCGA, GSEA analysis of CGGA cohort 1 also showed significant enrichment in pathways related to "Cell Cycle," "DNA replication," "p53 signaling pathway," and "hallmark E2F targets" (Fig. 4D). Pathways enriched in the CGGA cohort 2 include “Cell Cycle, Mitotic,” “DNA metabolic process,” “DNA repair pathways,” etc. (Supplementary Fig. 3A). GSEA results showed that, in addition to the “Cell Cycle” function being significantly enriched, candidate genes in TCGA were significantly enriched in the “CHEMOKINE SIGNALING PATHWAY,” “CYTOKINE RECEPTOR INTERACTION” pathway. In CGGA cohort 1, pathways such as “CELL CYCLE,” “P53 SIGNALING PATHWAY,” “PATHWAYS IN CANCER,” “TGF BETA SIGNALING PATHWAY” were significantly enriched. In CGGA cohort 2, pathways including “CELL CYCLE,” “P53 SIGNALING PATHWAY,” “PATHWAYS IN CANCER,” “SPLICEOSOME” were also enriched (Supplementary Fig. 3B).
GO analysis and GSEA enrichment analysis of SKP2-related genes. SKP2 positively correlated genes (TCGA (A), CGGA Cohort1 (C)) were analyzed by GO. The best enrichment pathways (TCGA (B), CGGA (D)) were identified by GSEA. SKP2 is closely related to the regulation of cell cycle processes. (E) and (F), correlation analysis of SKP2 expression and cell cycle checkpoints in TCGA and CGGA Cohort1 datasets. Pathway data were obtained from the KEGG database (Kanehisa Laboratories).
Since the results of both GO and GSEA analyses specified the cell cycle as the functional pathway with the highest relevance to SKP2, we selected a cluster of seven cell cycle-related metagenes as markers of cell cycle status, including p53, Retinoblastoma1 (RB1), Checkpoint Kinase 1 (CHEK1), Checkpoint Kinase 2 (CHEK2), CDK Inhibitor 1A (CDKN1A), CDK7, and Ataxia Telangiectasia Mutated (ATM). The results showed a significant correlation between SKP2 and these seven cell cycle checkpoints in the TCGA and CGGA databases (Fig. 4E, F; Supplementary Fig. 3C).
Prognostic modeling and validation
To further assess the importance of SKP2 on the prognosis of glioma patients, prognostic markers were established based on SKP2 and DEGs. In Fig. 5A, B, the LASSO regression method revealed the set of nine genes with the best prognostic value, i.e., HOXC10, OR51E1, HOXC6, HOXD11, HMGA2, HOXA3, GALNT5, HOXA5, and SHOX2, and their corresponding coefficients were used to calculate the risk scores (Fig. 5C). Using the median risk score as a threshold, patients were categorized into high-risk and low-risk groups in the TCGA dataset, and the mortality rate of patients with high-risk scores was higher than that of patients with low-risk scores (Fig. 5D). Kaplan–Meier analysis showed that patients with a low SKP2 risk score had significantly higher overall survival than that of patients with a high SKP2 risk score (Fig. 5E, p < 0.0001). In addition, OS at 1, 3, and 5 years was predicted using temporal ROC curve analysis based on the TCGA risk score, with area under the curve (AUC) values of 0.94, 0.92, and 0.88, respectively (Fig. 5F). Risk scores were also calculated in CGGA cohort 1 versus CGGA cohort 2, and patients with high risk scores had a higher mortality rate than those with low risk scores (Fig. 6A; Supplementary Fig. 4A), and a Kaplan–Meier survival analysis (Fig. 6B; Supplementary Fig. 4B), which showed that patients with low SKP2 risk scores survived longer. The ROC curves were used to predict OS at 1, 3, and 5 years, respectively, and the areas under the curve were 0.83, 0.82, and 0.77 for CGGA cohort 1 (Fig. 6C), and 0.85, 0.83, and 0.74 for CGGA cohort 2 (Supplementary Fig. 4C). The above results indicated that there was a significant correlation between the expression of SKP2 and the survival of patients, and we could predict the prognosis of patients by determining the expression of SKP2. These results indicate a significant correlation between SKP2-related risk scores and patient survival. Notably, the nine-gene signature is biologically relevant to SKP2-mediated cell cycle regulation, as several of these genes are involved in pathways such as DNA replication, DNA repair, and mitotic progression, thereby supporting their potential prognostic value in glioma. For example, HOXC6 and SHOX2 have been shown to correlate with glioma progression and poor survival outcomes16,17, and HOXC10 was identified as a survival-associated gene in GBM18. In addition, HMGA2 has been implicated in chromatin remodeling and DNA repair processes that drive tumorigenesis19. Together, these findings reinforce the biological plausibility of our nine-gene risk signature being functionally linked to SKP2-mediated pathways in glioma.
(A) LASSO coefficient curves for differentially differentiated genes in a ten-fold cross-validation. (B) Cross-validation of parameter selection in proportional risk model. (C) Risk factors for 9 candidate genes. (D) Risk scores, overall survival (OS) and expression level distribution of 9 genes in the risk profile. (E) Kaplan–Meier survival analysis of risk scores in patients with glioma. (F) Temporal receiver operating characteristic (ROC) curve analysis, predicting 1, 3, and 5-year OS based on the risk score of the TCGA dataset. (G) and (H), the relationship between risk scores of TCGA and CGGA Cohort1 datasets and glioma grade, 1p/19q codeletion status, and IDH mutation status. Risk scores are markedly elevated in high-grade gliomas; The risk score was significantly elevated in gliomas without isocitrate dehydrogenase (IDH) mutations; The risk score was significantly increased in gliomas without 1p/19q codeletion.
GO analysis and GSEA enrichment analysis of risk scores. (A), GO analysis of risk scores of TCGA dataset; (C) GO analysis of risk scores for CGGA Cohort1 dataset. (B) and (D), GSEA enrichment analysis of risk scores in TCGA and CGGA Cohort1 datasets. (E) and (F), Correlation analysis of risk score and cell cycle checkpoint in TCGA and CGGA Cohort1 databases. Pathway data were obtained from the KEGG database (Kanehisa Laboratories).
Glioma patients in the TCGA database were classified according to tumor grade, 1p19q coding, and IDH mutation/wild type, and SKP2 risk scores were calculated for patients under different classifications, respectively, with patients scoring higher risk scores in high-grade tumors, 1p19q non-coding, or IDH wild-type tumors (**** p < 0.0001) (Fig. 5G). Consistent results were obtained in CGGA cohort 1 versus CGGA cohort 2, (Fig. 5H; Supplementary Fig. 4D, F).
Functional enrichment analysis of SKP2 risk scores
Based on the SKP2 risk scores, further functional enrichment analyses were performed using GO and GSEA methods to obtain the potential functions and pathways of the target genes. GO enrichment analysis revealed the most important enriched functions and pathways in TCGA, including “Cell Cycle”, “mitotic cell cycle”, “regulation of cell cycle process”, “positive regulation”, and “cell cycle process”, mitotic cell cycle”, “regulation of cell cycle process”, “positive regulation of cell cycle process”, “cell cycle process”, and “cell cycle process”. The results of GO enrichment analysis of CGGA cohort 1 were “Cell Cycle”, “mitotic cell cycle”, “regulation of cell cycle process”, (Fig. 6A). “, “regulation of cell cycle process”, “positive regulation of cell cycle process “, (Fig. 6C). CGGA enrichment results for column 2 included “mitotic cell cycle”, “regulation of smoothened signaling pathway The results of GO enrichment analysis in TCGA and CGGA can be verified with each other, and both of them are basically consistent with the results of GO enrichment analysis obtained with SKP2 as the main variable. GSEA analysis results showed that the candidate genes in TCGA had “cell cycle” (NES = 1.830), “DNA replication” (NES = 1.770), “p53 signaling pathway” (NES = 1.740), and “hallmark E2F targets” (NES = 2.300) pathways were significantly enriched (Fig. 6B). In CGGA cohort 1, the “cell cycle” (NES = 1.340), “DNA replication” (NES = 1.580), “p53 signaling pathway” (NES = 1.480), and “hallmark E2F targets” (NES = 1.690) pathways were significantly enriched (Fig. 6D). In CGGA cohort 2, “DNA replication” (NES = 2.150), “ECM–receptor interaction” (NES = 2.510), “hallmark E2F targets” (NES = 2.470), and “hallmark G2M checkpoint” (NES = 2.330) were significantly enriched (Supplementary Fig. 5B).
We also analyzed the correlation between SKP2 risk scores and the 7 cell cycle checkpoints and showed that there was a correlation between SKP2 risk scores and all 7 checkpoints, with stronger correlations with CHEK1 and CHEK2 (Fig. 6E, F; Supplementary Fig. 5C).
The SKP2 risk score can help predict the prognosis of patients with glioma
We further evaluated the prognostic value of the SKP2 risk score in the CGGA Cohort1 dataset. Patients with higher risk scores showed significantly worse overall survival, and the predictive accuracy of the model was confirmed by the distribution of risk scores, Kaplan–Meier survival analysis, and time-dependent ROC curve analyses (Fig. 7A–C). When constructing the column line plot for survival analysis of the TCGA database, age, tumor grade, gender, and SKP2 risk score were included as predictors and their corresponding scores were calculated. The total scores obtained can correspond to 1-, 3-, and 5-year survival rates, respectively. The calibration curve can reflect the actual risk compared with the predicted risk, and the closer the curve is to the diagonal line, the better the prediction is. The deviation between the predicted and actual survival rates of patients using this method is small, and the curve basically conforms to the diagonal line (Fig. 8A, B), and the results obtained by plotting the survival analysis column line graphs of patients in CGGA cohort 1 and CGGA cohort 2 using the same method are also in good agreement with the actual results (Fig. 8C, D; Supplementary Fig. 6).
Validation of the SKP2 risk score in the CGGA Cohort1 dataset. (A) Distribution of risk scores, overall survival (OS), and expression levels in the CGGA Cohort1 dataset. (B) Kaplan–Meier survival analysis of risk scores in patients with glioma in the CGGA Cohort1 dataset. (C) Receiver Operating Characteristic (ROC) curve analysis at time, predicting 1, 3, and 5-year OS based on risk scores from the CGGA Cohort1 dataset.
Building personalized prediction models in TCGA and CGGA Cohort1. (A) Nomogram (TCGA). (B) Calibration plot for 1-, 3-, and 5-year OS (TCGA). (C) Nomogram (CGGA Cohort1). (D) Calibration plot for 1-, 3-, and 5-year OS (CGGA Cohort1).
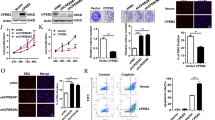
SKP2 knockdown efficiency validation
To verify the knockdown efficiency of SKP2, U87 and U251 cells were transfected with si-NC, si-SKP2-1, or si-SKP2-2. Western blot analysis confirmed that both siRNAs decreased SKP2 protein levels, with si-SKP2-1 showing the strongest effect (Fig. 9A, B). Therefore, si-SKP2-1 was used in subsequent experiments.
Knockdown efficiency of SKP2 in glioma cells. (A) Western blot showing SKP2 expression in U87 cells after transfection with si-NC, si-SKP2-1, or si-SKP2-2. β-actin was used as a loading control. Quantification of relative SKP2 expression is shown below. (B) Western blot analysis of U251 cells with the same treatments. Quantification shows that both siRNAs reduced SKP2 expression, with si-SKP2-1 showing the highest efficiency, which was used for further experiments. Statistical analysis was performed using Student’s t-test (two groups) or one-way ANOVA (multiple groups). ****p < 0.0001.
SKP2 expression significantly affects cell cycle regulation in glioma cells
The effects of SKP2 were experimentally verified in SKP2 knockdown U87 and U251 cells. Real-time fluorescence quantitative PCR and Western blot were used to detect SKP2 expression in the transfected cells. The results showed that the expression of SKP2 was significantly suppressed in the Si group compared with the NC group.
Flow cytometry was performed to detect the cell cycle in the NC and knockdown groups, respectively. The results showed that the cell cycle of SKP2 knockdown U87 and U251 cells was stalled in G1 phase compared with the NC group (Fig. 10A, B). This suggests that SKP2 affects the cell cycle by interfering with key proteins in the G1 phase of the cell cycle. Western Blot was performed to detect changes in the expression levels of key proteins in the G1 phase. The results showed that the expression levels of Cyclin E and CDK2 were significantly reduced in SKP2 knockdown U87 and U251 compared with the NC group (Fig. 10C, D).
Flow cytometry and Western blot assay were used to verify that SKP2 regulates the cell cycle. (A) Flow cytometry to detect the cell cycle. (B) Changes in cell cycle phases after SKP2 knockdown. (C) Western blot experimental results. (D) Changes in key cell cycle proteins after SKP2 knockdown. The data were obtained from three independent experiments. Statistical analysis was performed using Student’s t-test. *p < 0.05.
The above experimental results indicate that the expression level of SKP2 affects cell cycle regulation by influencing the expression of key proteins in the G1 phase, thus influencing the development of glioma cells.
SKP2 promotes glioma tumorigenesis in vivo
To examine the effect of SKP2 in vivo, U87 cells stably transduced with sh-NC-LUC (sh-NC) or sh-SKP2-LUC (sh-SKP2) were injected subcutaneously into nude mice. Bioluminescence imaging at day 25 showed weaker tumor signals in the sh-SKP2 group compared with controls (Fig. 11A, C). Tumors in the sh-SKP2 group were smaller in size (Fig. 11D), with significantly reduced tumor volume and weight (Fig. 11E, F). Western blotting of tumor tissue further confirmed decreased SKP2 expression, along with reduced Cyclin E and CDK2 levels in the sh-SKP2 group (Fig. 11G).
SKP2 knockdown suppresses tumor growth in vivo. (A) Representative bioluminescence images of nude mice bearing subcutaneous xenografts of U87 cells stably expressing sh-NC or sh-SKP2 at day 25 post-injection. The sh-SKP2 group showed markedly reduced luminescence signal. (B) Body weight curves showing no significant difference between groups throughout the experiment. (C) Quantification of bioluminescence intensity demonstrating significantly reduced tumor burden in the sh-SKP2 group. (D) Representative photographs of excised tumors at day 25, showing smaller tumor size in the sh-SKP2 group. (E) Tumor growth curves measured every 5 days showing significantly slower growth in the sh-SKP2 group. (F) Tumor weight at sacrifice was significantly lower in the sh-SKP2 group. (G) Western blot analysis of tumor lysates confirming reduced SKP2 expression and corresponding decreases in Cyclin E and CDK2 levels in sh-SKP2 tumors. β-actin served as loading control. Quantification of three independent tumors per group is shown below. Statistical analysis was performed using Student’s t-test. Error bars represent mean ± SEM. ***p < 0.001, ****p < 0.0001.
Discussion
SKP2 is functionally important in the regulation of the cell cycle, tumorigenesis, and is also required for the functional regulation of core transcription factors, such as p53, following cellular stress response to injury20,21,22. It has been demonstrated that SKP2 is abnormally highly expressed in a variety of tumor tissues and tumor cells and has oncogene properties23,24,25,26, but its mechanism of action as an oncogene in gliomas remains a mystery. This study presents analytic and experimental evidence on the functional role of SKP2 in gliomas. The main findings not only indicates that higher SKP2 expression is associated with poorer prognosis, but also verifies that SKP2 is a key factor in regulating gliomas’ cell cycle, which reflecting the malignance in gliomas’ progression.
Our study analyzed the expression levels of SKP2 in different pathological types and grades of gliomas based on mRNA-seq data from CGGA and TCGA databases. Our results confirmed that SKP2 is enriched in malignant gliomas and is highly expressed with pathologic staging of IDH wildtype, 1p19q non-codel and MGMT-unmethylated gliomas and correlated with poorer survival. Further, we determined the function of SKP2 by GO and GSEA analyses, which showed that SKP2 was mainly associated with cell cycle regulation27, in addition to signaling pathways such as DNA damage repair, chemokine signaling pathway, p53 signaling pathway28, and cytokine interactions. We analyzed the relationship between SKP2 and the classical cell cycle checkpoints p53, RB1, CHEK2, CHEK1, CDKN1A28,29, CDK7, and ATM30, and the results showed that there was a strong correlation between SKP2 and all seven checkpoints.
To further assess the prognostic role of SKP2 in glioma, we integrated transcriptomic data from TCGA and CGGA cohorts and developed a nine-gene prognostic signature using LASSO-Cox regression. Patients with higher SKP2-associated risk scores consistently exhibited poorer overall survival across independent datasets, and the ROC curves indicated reasonable predictive performance, particularly at 1- and 3-year endpoints. These findings are consistent with previous reports highlighting SKP2 as a marker of poor prognosis in gliomas and other malignancies31,32.
Functional enrichment analyses (GO and GSEA) revealed that SKP2-related risk genes were significantly enriched in pathways regulating the cell cycle, DNA damage repair, and p53 signaling, which are well-recognized hallmarks of glioma biology21,22. Importantly, SKP2 risk scores showed strong correlations with canonical checkpoint regulators such as p53, RB1, CHEK2, and CDKN1A, underscoring the relevance of SKP2 in cell cycle control. While these associations are biologically plausible, further mechanistic studies are needed to delineate whether SKP2 directly regulates these pathways in glioma cells.
We also established a nomogram incorporating SKP2 risk score and clinicopathological variables, which demonstrated good calibration for predicting 1-, 3-, and 5-year survival. This approach highlights the potential clinical value of SKP2-based models in prognostic stratification of glioma patients. After theoretical analysis of the effect of SKP2 on glioma development, specific experiments were conducted to verify the analysis results. We transiently transfected U87 and U251 cell lines with si-SKP2 and verified the knockdown efficiency of SKP2 by Western blot. Results of flow cytometry showed that the cell cycle of si-SKP2–transfected cells was stalled at the G1 phase compared with that of the NC group. To further confirm this finding, we used Western blot to examine the expression of key proteins involved in the G1 phase of the cell cycle, and the results showed a significant reduction in the expression of Cyclin E33,34 and CDK235,36in the SKP2 knockdown group compared with the NC group. These results suggest that SKP2 may influence glioma cell cycle progression, although additional experiments will be required to clarify the detailed molecular mechanisms of SKP2 in tumor progression. Taken together, these results suggest that SKP2 affects cell cycle regulation by influencing the expression of Cyclin E and CDK2, key proteins in the G1 phase of the cell cycle, thereby affecting tumor cell growth. Previously, Liu et al.37 reported that they successfully arrested cell cycle at G0/G1 phrase in lung cancer cells by using SKP2 inhibitor AAA-237. Another piece of evidence provided by Zheng et al.38 showed that FBXO43 can stabilize SKP2 by reducing auto-ubiquitylation, and therefore promotes the cell cycle in lung cancer cells, which illustrates the potential molecular mechanism of SKP2 in tumor progression, suggesting that SKP2 may be a pivotal therapeutic target for malignant tumors in the future. However, further clinical researches are needed to verify these effects of SKP2 in gliomas31,32. Recent evidence further supports the role of SKP2 in glioma progression, particularly in cell cycle regulation and DNA damage responses. Hume et al.39 demonstrated that the NUCKS1–SKP2–p21/p27 pathway is essential for the G1/S transition, while Jing et al.40reported that SKP2 also regulates DNA repair processes in addition to CDKI degradation. Moreover, a recent review by Feng41 emphasized the importance of SKP2 in tumor-associated cell cycle regulation, and Kamel41 highlighted its central role in promoting p21 and p27 degradation through the SKP2–Cks1 complex. This classical mechanism aligns with our bioinformatics findings and supports the biological plausibility of SKP2 in glioma progression.
In conclusion, our study reveals the promotional effect of SKP2 on gliomas. SKP2 affects the progression of the G1 phase of the cell cycle by influencing Cyclin E and CDK2 protein expression in glioma cells, thereby promoting tumor growth. The present study was primarily focused on bioinformatics analysis, with limited wet-lab validation. More detailed mechanistic experiments, such as p21/CDK4 detection, p53 rescue assays, or expanded glioma models, remain important directions for future research.
Data availability
The data during the current study are available from the corresponding author on reasonable request.
References
Louis, D. N. et al. The 2021 WHO Classification of Tumors of the Central Nervous System: a summary. Neuro-Oncol. 23, 1231–1251 (2021).
Ma, P. et al. Advances and challenges in novel drug delivery systems for glioma therapy. Front. Pharmacol. 16, 1655241 (2025).
Barthel, L. et al. Glioma: molecular signature and crossroads with tumor microenvironment. Cancer Metastasis Rev. 41, 53–75 (2022).
Varn, F. S. et al. Glioma progression is shaped by genetic evolution and microenvironment interactions. Cell 185, 2184-2199.e16 (2022).
Stallaert, W. et al. The structure of the human cell cycle. Cell Syst. 13, 230-240.e3 (2022).
Basu, S., Greenwood, J., Jones, A. W. & Nurse, P. Core control principles of the eukaryotic cell cycle. Nature 607, 381–386 (2022).
Li, F., Mladenov, E., Mortoga, S. & Iliakis, G. SCFSKP2 regulates APC/CCDH1-mediated degradation of CTIP to adjust DNA-end resection in G2-phase. Cell Death Dis. 11, 548 (2020).
Xiao, J. et al. CDK4/6 inhibition enhances oncolytic virus efficacy by potentiating tumor-selective cell killing and T-cell activation in refractory glioblastoma. Cancer Res. 82, 3359–3374 (2022).
Palafox, M. et al. High p16 expression and heterozygous RB1 loss are biomarkers for CDK4/6 inhibitor resistance in ER+ breast cancer. Nat. Commun. 13, 5258 (2022).
Pavlides, S. C. et al. TGF-β activates APC through Cdh1 binding for Cks1 and Skp2 proteasomal destruction stabilizing p27kip1 for normal endometrial growth. Cell Cycle Georget. Tex 15, 931–947 (2016).
Chen, H.-Y. et al. A derivate of benzimidazole-isoquinolinone induces SKP2 transcriptional inhibition to exert anti-tumor activity in glioblastoma cells. Mol. Basel Switz. 24, 2722 (2019).
García-Gutiérrez, L. et al. Myc stimulates cell cycle progression through the activation of Cdk1 and phosphorylation of p27. Sci. Rep. 9, 18693 (2019).
Kanehisa, M., Sato, Y., Kawashima, M., Furumichi, M. & Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 44, D457-462 (2016).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Percie du Sert, N. et al. The ARRIVE guidelines 20: updated guidelines for reporting animal research. PLOS Biol. 18, e3000410 (2020).
Ren, P. et al. A novel hypoxia-driven gene signature that can predict the prognosis and drug resistance of gliomas. Front. Genet. 13, 976356 (2022).
Wu, X., Chen, H., You, C. & Peng, Z. A potential immunotherapeutic and prognostic biomarker for multiple tumors including glioma: SHOX2. Hereditas 160, 21 (2023).
Cao, M. et al. A four-gene signature-derived risk score for glioblastoma: prospects for prognostic and response predictive analyses. Cancer Biol. Med. 16, 595–605 (2019).
Mansoori, B. et al. HMGA2 as a critical regulator in cancer development. Genes 12, 269 (2021).
Engeland, K. Cell cycle regulation: p53–p21-RB signaling. Cell Death Differ. 29, 946–960 (2022).
Raali, R. & Suresh, P. K. Unraveling glioblastoma: TME implication and gene therapy advances. Curr. Gene Ther. 25, 497–517 (2025).
Jiang, M. et al. Integrated Analysis of Clinical Outcome of Mesenchymal Stem Cell-related Genes in Pan-cancer. Curr. Genomics 25, 298–315 (2024).
Kanamori, A. et al. Mint3 depletion restricts tumor malignancy of pancreatic cancer cells by decreasing SKP2 expression via HIF-1. Oncogene 39, 6218–6230 (2020).
Wang, G. et al. Her2 promotes early dissemination of breast cancer by suppressing the p38 pathway through Skp2-mediated proteasomal degradation of Tpl2. Oncogene 39, 7034–7050 (2020).
Li, H., Ouyang, J. & Liu, R. Platycodin D suppresses proliferation, migration, and invasion of human glioblastoma cells through regulation of Skp2. Eur. J. Pharmacol. 948, 175697 (2023).
Yu, M. et al. RIOK2 inhibitor NSC139021 exerts anti-tumor effects on glioblastoma via inducing Skp2-mediated cell cycle arrest and apoptosis. Biomedicines 9, 1244 (2021).
Ferrena, A. et al. SKP2 knockout in Rb1/p53-deficient mouse models of osteosarcoma induces immune infiltration and drives a transcriptional program with a favorable prognosis. Mol. Cancer Ther. 23, 223–234 (2024).
Shen, L. et al. NDRG2 facilitates colorectal cancer differentiation through the regulation of Skp2-p21/p27 axis. Oncogene 37, 1759–1774 (2018).
de Queiroz, R. M., Moon, S.-H. & Prives, C. O-GlcNAc transferase regulates p21 protein levels and cell proliferation through the FoxM1-Skp2 axis in a p53-independent manner. J. Biol. Chem. 298, 102289 (2022).
Qi, L. et al. Phosphorylation of BRCA1 by ATM upon double-strand breaks impacts ATM function in end-resection: a potential feedback loop. Iscience 25, 104944 (2022).
Bombino, A., Magnani, M. & Conti, A. A promising breakthrough: the potential of VORASIDENIB in the treatment of low-grade glioma. Curr. Mol. Pharmacol. 17, e18761429290327 (2024).
Zhang, Q., Yu, X., Zheng, Q., He, Y. & Guo, W. A molecular subtype model for liver HBV-related hepatocellular carcinoma patients based on immune-related genes. Front. Oncol. 10, 560229 (2020).
Zeng, J., Hills, S. A., Ozono, E. & Diffley, J. F. X. Cyclin E-induced replicative stress drives p53-dependent whole-genome duplication. Cell 186, 528-542.e14 (2023).
Chu, C., Geng, Y., Zhou, Y. & Sicinski, P. Cyclin E in normal physiology and disease states. Trends Cell Biol. 31, 732–746 (2021).
Arora, M. et al. Rapid adaptation to CDK2 inhibition exposes intrinsic cell-cycle plasticity. Cell 186, 2628-2643.e21 (2023).
Kirova, D. G. et al. A ROS-dependent mechanism promotes CDK2 phosphorylation to drive progression through S phase. Dev. Cell 57, 1712-1727.e9 (2022).
Liu, J. et al. Anti-tumor effects of Skp2 inhibitor AAA-237 on NSCLC by arresting cell cycle at G0/G1 phase and inducing senescence. Pharmacol. Res. 181, 106259 (2022).
Zheng, L. et al. FBXO43 promotes cell cycle progression in cancer cells through stabilizing SKP2. Cancer Lett. 591, 216848 (2024).
Hume, S. et al. The NUCKS1-SKP2-p21/p27 axis controls S phase entry. Nat. Commun. 12, 6959 (2021).
Jing, J. et al. Small-molecule compounds inhibiting S-phase kinase-associated protein 2: A review. Front. Pharmacol. 14, (2023).
Targeting the Skp2–Cks1 protein–protein interaction: structures, assays, and preclinical inhibitors. Eur. J. Pharmacol. 1006, 178129 (2025).
Funding
This study was supported by the National Key R&D Program of China (No.2023YFC2510000).
Author information
Authors and Affiliations
Contributions
L.G: Contributed to conceptualization; methodology; software; formal analysis; visualization; writing—original draft preparation. H.L: Contributed to conceptualization; methodology; software; visualization; writing—original draft preparation. Y.L: Contributed to conceptualization; methodology; software; visualization; writing—review and editing; data curation. J.O: Contributed to investigation; visualization; supervision. C.Y: Contributed to investigation; visualization; software; supervision; F.L: Contributed to conceptualization; supervision; data curation; project administration. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval and consent to participate
All animal procedures were approved by the Ethics Committee of Peking University People’s Hospital (Approval No. 2021-PHE072) and were conducted in accordance with the ARRIVE guidelines and the Guide for the Care and Use of Laboratory Animals (NIH, USA).
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Guan, L., Li, H., Liu, Y. et al. SKP2 protein as a prognostic biomarker for glioma and promotes tumor progression by regulating the cell cycle. Sci Rep 15, 42668 (2025). https://doi.org/10.1038/s41598-025-26722-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-26722-6