Abstract
Parkinson’s disease (PD) is an intractable progressive neurodegenerative disease with poor prognosis in elder patients, which is largely attribute to the deficiency of ultimate pathogenesis. Longitudinal studies have indicated a pivotal role of natural killer (NK) cells, yet the systematic and detailed information of circulating NK cells in PD is largely unknowable. To explore the role of NK cells in PD, we isolated mononuclear cells from peripheral blood (PBMCs) by Ficoll-based density gradient centrifugation, and detected the content of total resident NK cells in healthy donors (HD-NKs) and PD patients (PD-NKs) and the concomitant subsets by flow cytometry (FCM) assay. Then, we took advantage of our well-established “3ILs”-based strategy for ex vivo NK cell expansion and activation, and followed by cellular viability and cytotoxicity assessment. By conducting RNA-sequencing (RNA-SEQ) and multifaceted bioinformatics analyses, we compared the transcriptomic signatures of expanded HD-NKs and PD-NKs. Compared to HD-NKs, PD-NKs showed increase in resident NK cells but minimal differences in expanded NK cells, together with diversity in the subpopulations of NK cells (CD16+, NKG2D+, NKp46+). Interestingly, PD-NKs revealed a moderate higher percentage of apoptotic cells and cytotoxicity upon the co-cultured Nalm6 and U937 tumor cell lines. Despite with high conservations in gene expression pattern and genetic variations, PD-NKs revealed multifaceted diversity in gene set-associated immune response and metabolism. Overall, our data revealed the multidimensional biological and transcriptomic signatures of resident and expanded NK cells generated from peripheral blood of PD patients and HDs. Our findings would provide new references for the further development of NK cell-based biomarkers for PD diagnosis and novel immunotherapy for neurodegenerative diseases.
Similar content being viewed by others
Introduction
Parkinson’s disease (PD), characterized by bradykinesia and tremor, has been recognized as the second most common neurodegenerative disease worldwide1,2. The incidence of PD is increasing as population aging, and the risk is twice in men than women3. The main pathological features of PD are progressive loss of dopaminergic neurons in the substantia nigra and other brain regions, abnormal aggregation of α-synuclein (α-syn) in residual intraneuronal and intra-axonal (Lewy bodies and Lewy Neurites), and neuroinflammation induced by activation of microglia in PD patients4.The condition is characterized by motor symptoms, including bradykinesia, rigidity and tremor, and a plethora of non-motor disturbances. There are neither clinical biomarkers for the early detection, diagnosis, or prognostic evaluation of PD, nor a cure for the disease, and at this stage the mainstay of symptom relief is through anti-PD medications. Although dopamine replacement therapy can rescue key neurochemical deficits in motor symptoms such as bradykinesia, rigidity, and tremor through exogenous supplementation to improve motor symptoms, there are currently no available therapies to slow disease progression or prevent long-term disabling complications such as postural instability, falls, and dementia, which are major causes of disability and poor quality of life for people with PD5. Therefore, the exploration of biomarkers related to PD diagnosis and disease-modifying therapies to slow or stop disease progression may be an urgent medical need in the field of PD in future.
Longitudinal studies have shown that genetic factors play an important role in the development of PD. More than 100 susceptibility genes and SNP loci have been reported to be associated with the development of PD6, and patients with PD often harbor the coexistence of multiple common high-risk variants and rare or de novo variants, and PD is often facilitated by the combination of multiple minimally effective genes. The pathogenesis of PD is a series of pathophysiological changes under the combined effect of genetic predisposition factors and environmental risk factors1,7. State-of-the-art renewal has indicated the involvement of immune system responses and neuroinflammation in the pathogenesis of PD, and immunoinflammatory modulation therapy provides a possible target for alleviating the progression of PD8,9. Meanwhile, PD patients showed decline in humoral immune responses and cellular immune responses10,11. For instance, differ from those with anti-inflammatory drug and immunosuppressant treatment, patients with autoimmune disease reveal an increasing risk of PD, which is confirmed by multifaceted bioinformatics analyses including the common molecular pathways and polygenic risk variants, together with specific loci variations and activated microglia in the brains of PD patients12,13.
Natural killer (NK) cells, comprising the cytotoxic CD3-CD56dimCD16high and IFN-γ-producing CD3-CD56brightCD16+ subpopulations, are pivotal members of the innate lymphoid cells (ILCs) family with spontaneous anti-microbial infection, tumor-killing capacity, formation of immunological memory, and endocytosis of extracellular proteins14,15. As the key component of innate immune cells, NK cells play a pivotal role in both innate and adaptive immune responses for the maintenance of spatio-temporal homeostasis and the local microenvironment without presensitization16,17.
In recent years, researchers have focused on the possible link between NK cells and PD for two main reasons. On the one hand, numerous studies have indicated the association of NK cell activity with neurological disorders, especially demyelinating disorders including PD. On the other hand, investigators in the field have demonstrated the interaction between NK cells and α-syn in PD animal models, which facilitats the hypothesis that the neuronal degenerative pathological changes in patients with PD may interact with immune abnormalities18,19. Current literatures have indicated the immunomodulatory and neuroprotective effects of NK cells attribute to their migration to the central nervous system (CNS) and the concomitant suppression of the self-reactive T cell hyper-activation20. Of note, NK cells have been employed as an immunotherapeutic approach for glioblastoma, suggesting their potential as a novel therapeutic regimen for CNS disorders21. Meanwhile, the accumulation of NK cells in the brains and CNS parenchyma of PD patients prior to the degeneration of dopaminergic neurons indicate the involvement of NK cells during PD progression22,23. Encouraging evidence has highlighted the mitigating effect of NK cells upon chronic inflammation like PD when migrating to the brain via the disrupted blood-brain barrier by suppressing brain tissue inflammation and abnormal protein aggregation23. However, the systematic and detailed dissection of the biological and transcriptomic signatures of NK cells in PD are largely obscure.
For the purpose, we isolated PBMCs from HDs and PD patients, and compared the cytophenotypic characteristics of resident HD-NKs and PD-NKs. Meanwhile, with the aid of our well-established “3ILs”-based strategy, we further verified the biological and transcriptomic signatures of expanded HD-NKs and PD-NKs. Taken together, our data indicated the multifaceted conservations and variations of HD-NKs and PD-NKs both at the cellular and molecular levels, including the contents of subpopulations, cellular viability, ex vivo activation and cytotoxicity, gene expression pattern, and genetic variations. Collectively, our findings provided novel references for further dissecting the features of HD-NKs and PD-NKs in future.
Materials and methods
Subjects and samples
Peripheral blood samples were collected from 7 PD patients (male, 2; female, 5; age, 54-75 years) and 5 HDs (male, 3; female, 2; age, 53-74 years). All individuals signed informed consents according to Declaration of Helsinki. The study was approved by the Ethical Committee of The Second Hospital of Shandong University (approval no.: KYLL-2023LW050). The detailed information of PD patients and HDs was listed in Supplementary Information: Supplementary Table S1-S2.
PBMC isolation and NK cell culture
PBMCs in PD patients and HDs were isolated from peripheral blood by Ficoll-based density gradient centrifugation (Tianjin Haoyang Biological Products Technology Co. LTD, China) as we recently reported24,25. For ex vivo NK cell expansion and activation, the PBMCs were cultured in NK MACS Medium (Miltenyi Biotec, Germany) supplemented with 1% NK MACS Supplement (Miltenyi Biotec, Germany), 5% human AB serum (Sigma-Aldrich, St Louis, MO, USA), 1% penicillin/streptomycin (Thermo Fisher Scientific, USA), 1000 U/ml rhIL-2 (PeproTech Inc, USA), 10 ng/ml rhIL-15 (PeproTech Inc, USA) and 50 ng/ml rhIL-18 (R&D Systems, USA) at 37 ℃, 5% CO2. The cells were cultured in a 37 ℃, 5% CO2 incubator for 14 days. The culture medium was replaced every 2-3 days, and the cells were gently mixed by pipetting with a micropipette24,26. Cell counts were performed for each well at Day 0, Day 7, and Day 14 of the culture, and the cell plates were photographed under a microscope to record the cell morphology. The list of the indicated cytokines was available in Supplementary Information: Supplementary Table S3.
Flow cytometry (FCM) analysis
FCM assay was conducted as we recently described25,27.The resident NK cells in peripheral blood and expanded NK cells after the 14-day’s ex vivo stimulation were harvested and washed with 1×PBS (Solarbio, China) for twice. Then, the cells were incubated with fluorescence-conjugated antibodies in dark for 30 min. Finally, the cells at the indicated time potints were washed with 1× PBS (Solarbio, China) for twice, and detected by Canto II and FlowJo V10.0 (BD, USA) and analyzed by FlowJo V10.0 (BD, USA). The information of the antibodies was listed in Supplementary Information: Supplementary Table S4.
Cell counting of NK cells
Total cell counting was accomplished by utilizing trypan blue staining (Sigma-Aldrich, StLouis, MO, USA) under the phase contrast microscope (Thermo FisherScientific Company, USA). The percentages of total NK cells (CD3-CD56+) and total activated NK cells (CD3-CD56+CD16+) were analyzed by Canto II and FlowJo V10.0 (BD, USA). The number of NK cells was calculated according to the formula: Total number of NK cells = total cell number × the percentage of NK cells, Fold change of NK cells = total NK cells at Day 14 / total NK cells at Day 0 × 100%.
Apoptotic analysis of NK cells
Cell apoptosis of HD-NKs and PD-NKs was performed as we recently described with serveral modifications25,27. In brief, 1×106 cells were washed with precooled (Solarbio, China) for twice and incubated in 100 μL 1 × Binding Buffer (Solarbio, China). Then, the cells were incubated in Annexin V-FITC (Solarbio, China) for 10 min and 7-AAD solution (Solarbio, China) for 5 min in dark, respectively. Finally, the percentages of apoptotic cells in HD-NKs and PD-NKs were measured by FACS Canto II (BD Biosci, USA) and FlowJo 10.0 software (Tree Star, USA).
Cell cycle assessment of NK cells
Cell cycle of HD-NKs and PD-NKs was verified as we described before26,27. Briefly, HD-NKs and PD-NKs were precooled in 70% (v/v) ethanol, and then fixed overnight at 4 ℃. Then, the cells were washed with 1 × PBS (Solarbio, China) at 4 ℃ for twice. Then, the cells were incubated with Propidium Iodide (PI) staining solution for 30 min at 37 ℃, and turned to BD LSR II (BD Biosci, USA) and ModFit software (Verity Software House Co. Ltd, USA) for FCM analysis.
The assessment of ex vivo NK cell cytotoxicity
The cytotoxicity of HD-NKs and PD-NKs was assessed as we reported28,29. The tumor cell lines (K562, Nalm6, U937) were cultured in RPMI-1640 basal medium (Gibco, USA) supplemented with 10% FBS (Gibco, USA) at 37 ℃, 5% CO2 as we recently described26,28. For the assessment of NK cell cytotoxicity, the indicated tumor cells were labelled with CellTrace Violet (tested with BV421 laser channel, Invitrogen, USA), and then solely (negative control, NC) or co-cultured with PD-NKs or HD-NKs at the indicated effector-to-target ratio (E: T=1:5). 8 hr later, the cells were incubated with the indicated antibodies (anti-CD3-FITC, ant-CD56-APC, anti-CD-107-PE and 7-AAD). Finally, the cells were resuspended in 200 μL 1 × PBS (Solarbio, China) with 5 μL Precision Count Beads (BioLegend, USA), and then turned to FACS Canto II (BD Biosci, USA) and FlowJo 7.0 software (Tree Star, USA) for analysis. Cytotoxic activity of NK cells was calculated according to the formula: Cytotoxicity = (1-N1/N0) × 100%. N1 and N0 represent the total number of living tumor cells in the experimental group and in the control group, respectively.
RNA-SEQ and bioinformatics analyses
Total mRNAs in HD-NKs and PD-NKs were prepared as we reported before by utilizing the TRIzol reagent (Invitrogen, USA) according to the manufacturer’s instructions25,30. Then, the mRNAs were qualified by NanoDrop (Thermo) and sequenced by Novogene (Tianjin, China). For the comparison of the gene expression pattern and genetic variation between HD-NKs and PD-NKs, multifaceted bioinformatics analyses were conducted as we reported before30,31,32. KEGG analysis is conducted by using the commercial BGI system (https://biosys.bgi.com/#/report/login) rather than the KEGG software from the Kanehisa laboratory (www.kegg.jp/feedback/copyright.html). The differentially expressed genes (DEGs) between HD-NKs and PD-NKs were listed in Supplementary file 2: Supplementary Table S5.
Statistical analysis
Statistical analysis was performed with GraphPad Prism 6.0 (GraphPad Software, USA) software as we reported before25,33,34. One-way ANOVA assay was performed for the comparison among three groups, while two different unpaired groups was conducted with unpaired t test. All data were indicated as means ± SEM (N=3 independent experiments), and significant statistical difference was considered only when P<0.05. *, P<0.05; **, P<0.01; ***, P<0.001; ****, P<0.0001; NS, not significant.
Results
Resident PD-NKs in PBMCs revealed diversity in the content of subpopulations
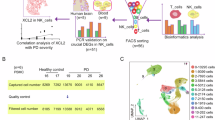
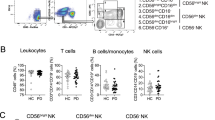
To verify the cytophentypoic characteristics between HD-NKs and PD-NKs, we enriched mononuclear cells from the corresponding peripheral blood (PBMCs) by utilizing the Ficoll-based density gradient centrifugation. Then, with the aid of flow cytometry (FCM) and statistical analysis, we found resident PD-NKs (PD-NKs (d0)) revealed higher proportion over HD-NKs (HD-NKs (d0)) (Figure 1A-1B). Furthermore, we found the percentage of total activated CD3-CD56+CD16+ PD-NKs and the indicated subpopulations (NKG2D+, NKp46+, CD16+ ) were higher than that in the HD-NKs, whereas the subsets of NKp44+ PD-NKs showed moderate decline instead (Figure 1C-1D). Meanwhile, we also observed the higher percentage of CD3+CD8+ T cells in the PD-NK group, whereas the decline in the proportion of CD3+CD4+ T cells (Figure 1E). Additionally, minimal differences were observed in the percentages of CD3+ total T cells and CD3+CD56+ NKT-like cells between the indicated groups (Figure 1E). Taken together, our data indicated the diversity in the content of NK cells as well as relative immune cells in the peripheral blood of PD patients.
Comparison of the phenotypic features of resident HD-NKs and PD-NKs (A-B) Representative FCM diagram (A) and statistical analysis (B) of total resident CD3-CD56+ NKs in PBMCs of PD patients and healthy donor (HD). (C-D) Representative FCM diagram (C) and statistical analysis (D) of the subpopulations (CD16+, NKG2D+, CD25+, NKp44+, NKp46+) in resident HD-NKs (d0) and PD-NKs (d0). (E) statistical analysis of total CD3+ T cells, CD3+CD56+ NKT cells, CD3+CD4+ T cells, and CD3+CD8+ T cells in PBMCs of PD patients and HD. All data were shown as Mean ± SEM (N=3 independent experiments). *, P<0.05; **, P<0.01; NS, not significant.
Expanded PD-NKs showed moderate decline in content and diversity in the subpopulations
To further dissect the potential variations of PD-NKs, we took advantage of our well-established “3ILs”-based strategy for ex vivo NK cell expansion and activation, and observed the increase in the emergency of cell aggregates and unicellular number in the PD-NK group (Figure 2A-2B). With the aid of FCM analysis, we found the percentage of total expanded PD-NKs (PD-NK (d14)) and HD-NKs (HD-NK (d14)) showed minimal differences in content (Figure 2C-2D). Differ from the total activated CD3-CD56+CD16+ NK cells, we found the percentage of the indicated subpopulations (NKG2D+, CD16+) in the PD-NK (d14) group was higher than that in the HD-NK (d14) group, the percentage of the indicated subpopulations (NKp44+, NKp46+) in these two groups showed minimal differences in content . Whereas the percentage of the indicated subpopulations (NKG2D+, CD16+,NKp44+, NKp46+) in the expanded-NK (d14) group was higher than that in the resident-NK (d14) group, respectively (Figure 2E-2G). Collectively, these data showed the variations in the content of total NKs and the indicated subsets between HD-NKs and PD-NKs after ex vivo expansion and activation.
Comparison of the phenotypic features of expanded HD-NKs and PD-NKs (A) Representative morphology of PBMCs-derived NKs at the indicated time points (day 0, 7, 14) during the 14-days’ ex vivo induction. Scale bar=100 μm. (B) Fold change of NK cells in the HD-NK group and the PD-NK group. (C-D) Representative FCM diagram (C) and statistical analysis (D) of total expanded CD3-CD56+ NKs after the 14-day’s induction in the indicated groups. (E-F) Representative FCM diagram (E) and statistical analysis (F) of the subpopulations (CD16+, NKG2D+, CD25+, NKp44+, NKp46+) in the expanded HD-NKs (d14) and PD-NKs (d14). (G) Statistical analysis of resident (d0) and expanded (d14) NK cells in the HD-NK group and the PD-NK group. All data were shown as mean ± SEM (N=3 independent experiments). NS, not significant; *, P<0.05; **, P<0.01; ***, P<0.001; ****, P<0.0001.
PD‑NKs revealed moderate increase in apoptosis and similarity in cytotoxicity with HD-NKs
To verify the similarities and differences in cellular viability, we turned to apoptotic analysis and found that the expanded PD-NKs (PD-NKs (d14)) showed increase in the percentage of 7-AAD-Annexin Ⅴ+ and Annexin Ⅴ+ apoptotic NK cells compared to that in HD-NKs, whereas with minimal differences in the percentage of 7-AAD+Annexin Ⅴ+ apoptotic NK cells (Figure 3A-3B). As shown by the FCM diagrams, the percentage of expanded PD-NKs in the S and G2/M sub-stages of cell cycle was moderately higher than that in the HD-NK (d14) group, whereas those in the G0/G1 sub-stage showed an opposite pattern (Figure 3C-3D).
Comparison of the cellular viability and the ex vivo cytotoxicity of expanded HD-NKs and PD-NKs (A-B) Representative FCM diagrams (A) and statistical analysis (B) of the percentages of apoptotic NKs in expanded HD-NKs and PD-NKs at day 14. (C-D) The representative distribution (C) and statistical analysis (D) of expanded HD-NKs and PD-NKs in the indicated sub-stages of cell cycle. (E-F) Representative FCM diagram (E) and statistical analysis (F) of CD107a+ expanded HD-NKs and PD-NKs in co-culturing with the indicated tumor cell lines (K562, Nalm6, U937) at the effector-to-target ratios (E: T=1:5). (G-H) Comparison of the ex vivo cytotoxicity of the expanded HD-NKs and PD-NKs against the indicated tumor cell line on the basis of BV-421+ cells (G) and living of tumor cells (H). All data were shown as mean ± SEM (N=3 independent experiments). NS, not significant; *, P<0.05; **, P<0.01; ***, P<0.001.
Subsequently, we tried to verify the in vitro cytotoxicity of the expanded PD-NKs and HD-NKs by co-culturing with the indicated tumor cell lines. As shown by Figure 3E-3F, the proportion of PD-NKs with CD107a expression were higher than those in the HD-NK group against Nalm6 cell line, whereas with minimal differences in the expression of CD107a against K562 and U937 cell lines. Notably, we found the percentages of living tumor cells (K562, Nalm6 and U937 cells) in the HD-NK (d14) group were higher than those in the PD-NK (d14) group (Figure 3G). Consequently, compared to the HD-NK (d14) group, the PD-NK (d14) group revealed moderate higher cytotoxicity against K562, Nalm6 and U937 cells (Figure 3H). Collectively, our data indicated the diversity of HD-NKs and PD-NKs in the distributions of sub-stages of cell cycle and apoptotic cells, together with the ex vivo cytotoxicity.
PD‑NKs and HD-NKs exhibited diversity in gene expression profiling
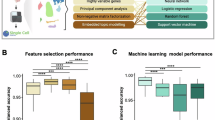
Having illuminated the multifaceted biological properties, we next turn to dissect the gene expression pattern between HD-NKs and PD-NKs. By conducting RNA-sequencing and bioinformatics analyses, we noticed the conservation in the distribution of gene expression pattern between the HD-NK (HD-NK-1, HD-NK-2, HD-NK-3) and PD-NK groups (PD-NK-1, PD-NK-2, PD-NK-3) according to the accumulation map and box plot of gene expression (Figure 4A-4B). With the aid of principal component analysis (PCA) and correlation analysis, we observed the differences in affinity between HD-NKs and PD-NKs (Figure 4C-4D).
Gene expression profiling of expanded HD-NKs and PD-NKs (A-B) The accumulation map (A) and box plot of gene numbers with the corresponding expression levels based on TPM values in expanded HD-NK (HD-NK-1, HD-NK-2, HD-NK-3) and PD-NK (PD-NK-1, PD-NK-2, PD-NK-3). (C-D) The PCA diagram (C) and correlation analysis (D) of the expanded HD-NK and PD-NK based on FPKM values. (E-F) Volcano Plot (E) and hierarchical cluster analysis (F) of the differentially expressed genes (DEGs) in the expanded HD-NK and PD-NK. (G-H) GOBP analysis (G) and KEGG analysis (H) of the aforementioned DEGs between the expanded HD-NK and PD-NK.
Simultaneously, numerous differentially expressed genes (DEGs) were observed between HD-NKs and PD-NKs according to the volcano plot and hierarchical cluster analysis (Figure 4E-4F). As shown by gene ontology biological process (GOBP) analysis, the DEGs were mainly involved in diverse bioprocesses such as angiogenesis, immunoregulation, metabolism and extracellular matrix organization (Figure 4G). KEGG analysis is accomplished by utilizing the BGI system (https://biosys.bgi.com/#/report/login). As suggested by high-throughput histology KEGG analysis, we found that DEGs are potentially involved in signaling pathways related to actin cytoskeleton regulation, cytokine-cytokine receptor interactions, Th17 cell differentiation, and antigen processing and presentation, which is consistent with our clinical observations, but due to the small sample size, we did not see a statistically significant difference (Q > 0.05), which can be followed up with increased sample size to further validate (Figure 4H). Therefore, these data indicated the multifaceted signatures of PD-NKs and HD-NKs in gene expression pattern.
PD‑NKs exhibited diverse genetic variations with HD-NKs
To further elucidate the potential similarities and differences at transcriptomic level, we turned to GSEA and observed that HD-NKs and PD-NKs showed significant differences in gene sets associated with immune response (e.g., INF-α, IFN-γ and complement), metabolism (e.g., bile acid metabolism, fatty acid metabolism) (P<0.05) (Figure 5A-5B). However, no statistical significances were observed in gene sets such as P53 pathway and KRAS signaling between HD-NKs and PD-NKs (P>0.05) (Figure 5C).
Comparison of gene sets and the variations of SNP spectrum in expanded HD-NKs and PD-NKs (A-C) GSEA diagrams showed the immune response- (A) and metabolism- (B) associated gene sets and signaling pathway-associated gene sets (C) between the expanded HD-NK and PD-NK. (D) KDA analysis revealed the XXXX of the indicated genes (associated genes, raw genes, KDA) between the expanded HD-NK and PD-NK. (E-F) The histogram exhibited the proportion of genes with variable shear event (VSE) (E) and differentially VSEs (DVSEs) between the expanded HD-NK and PD-NK. (G) Circos diagrams showed the distribution of fusion genes with the indicated genetic variations between the expanded HD-NK and PD-NK.
From the view of key derive gene analysis (KDA), we intuitively noticed the spatial correlations among representative KDR genes (e.g., PRRX1, COL1A1, CD44, VEGFA, and CDH1) between HD-NKs and PD-NKs (Figure 5D). As shown by Figure 5E, both HD-NKs and PD-NKs showed high conservations in the distribution of variable shear events (VSEs) including as_a3ss, as_a5ss, as_mxe, as_ri and as_se (Figure 5E). Among them, only a small number of differentially VSEs (DVSE) were observed, and as_se occupied the dominating proportion (Figure 5F). Finally, with the aid of Circos diagrams, we could intuitively observe the loci regional distribution of genes with the indicated somatic variations (e.g., SNPs, INDELs, gene fusion events and FPKM values) (Figure 5G). Taken together, these data exhibited the diversity in gene subsets and VSE alterations, together with the conservations in genetic modifications.
Discussion
For the past three decades, Parkinson’s disease (PD) has increased in prevalence and affects 1-2 per 1000 of the population globally35,36. As an intractable progressive neurodegenerative disorder, PD has been extensively explored to verify the pathogenesis (e.g., family history, aging, pesticide exposure, environmental chemicals) whereas the ultimate causes are still unknowable35. Current literatures have indicated the paramount role of immune response and neuroinflammation during the progress of neuroimmune disease, yet the pathogenesis and neuroprotective effect of NK cells in PD remain largely obscure8,35. In this study, we verified the multidimensional biological variations between HD-NKs and PD-NKs, and in particular, the subpopulations in resident NKs from peripheral blood and expanded counterparts after ex vivo culture. With the aid of RNA-SEQ analysis, we further illuminated the landscape of gene expression profiling and genetic variations between HD-NKs and PD-NKs. Collectively, our data revealed the cellular and molecular signatures of PD-NKs, which would benefit the further exploration of the pathogenesis of PD in future.
For decades, peripheral blood and the concomitant PBMCs have been considered as alternative sources for autologous or allogeneic NK cell preparation, which largely attribute to the low expression of MHC-Ⅰ and the convenient ex vivo preparation14,37,38. However, the inherent defects (e.g., donor variations, low amplification efficiency) largely hinder the large-scale clinical application14,39. Therewith, we and other investigators have been committed to explore diverse alternative sources for NK cell preparation such as perinatal blood (e.g., cord blood, placental blood), NK cell lines (e.g., NK-92MI, YT), and stem cells (e.g., hematopoietic stem cells, pluripotent stem cells)14,16,29,40,41. Of note, we recently reported the high-efficient generation of NK cells within 2 weeks by utilizing the “3ILs”-based cytokine cocktails (IL-2, IL-15, IL-18), which would collectively facilitate the further optimization of ex vivo NK cell preparation25,28,29.
State-of-the-art literature has indicated the involvement of NK cells in the pathogenesis of PD. For instance, Earls and the colleagues found that depletion of NK cells could increase motor symptoms, disease incidence, pathological aggregation of α-syn and neuroinflammation in a PD mouse model, which suggested the potential neuroprotective effect of NK cells upon PD and relative neurodegenerative diseases23. In this study, we identified both resident and expanded NK cells from peripheral blood of PD patients, and conducted systematic and detailed dissection of the biological and transcriptomic properties, including the content of subpopulations, cellular viability, ex vivo cytotoxicity, gene expression pattern and genetic variations.
Previous investigations have demonstrated a significant increase in CD3-CD56+ NK cells in PD patients compared to controls42,43. The current research has identified a higher proportion of resting CD3-CD56+ NK cells in the PD group compared to HD-NKs (d0), which is in line with the outcomes of previous studies. Although prior investigations have documented an increase in NK cells in PD, yet these studies mainly center on advanced stages of the disease43. Instead, we dissected the cellular and transcriptomic signatures of resident and expanded NK cells in the early stages of PD. For example, our data indicated the increase in resident PD-NKs (PD-NK (d0)) and the indicated subpopulations (NKG2D+, NKp44+, NKp46+) in expanded PD-NKs (PD-NK (d14)). In consistence, Holbrook et al reported the early stage PD cohort with elevated CD3-CD56+ total NK cells and CD3-CD16+CD56dim subset44. The NKG2D receptor on NK cells can be triggered by multiple ligands, leading to augmented NK cytotoxicity upon in vitro stimulation45. Therewith, these data collectively demonstrated the hyperactivation of NK cells during the early phase of PD, which further indicated the involvement in neuroinflammatory response during the initial phases of PD. We also put forward the potentiality for PD treatment by modulating the homeostasis of disease-associated NK cell subpopulations (e.g., CD16+, NKp44+, NKp46+) in future. However, the certain limitations of this study (e.g., the small number of cases due to the inclusion criteria requiring early and newly diagnosed PD patients) should be further validated by the following large-scale clinical investigations (e.g., large-sample, multi-level, multi-center prospective studies), which will enhance the statistical power and prevent the significant statistical association between NK cells and the pathogenesis of PD.
To date, diverse strategies have been formulated for NK cell-mediated immunotherapy upon tumor-related diseases and infectious diseases. As to neurological disorders, it’s of great interesting to illuminate the interactions between NK cells and microglia activation, leukocyte infiltration in the central nervous system, which will offer novel directions for formulating NK cell-targeted therapeutic strategies for PD and related diseases. For instance, epidemiological studies have demonstrated that PD patients with a lower incidence of tumors, which indicate the potential association of increased NK cytotoxic effect in enabling resistance to tumor cell infiltration46. In this study, by conducting coculture with tumor cell lines, we further verified the enhanced cytotoxicity of PD-NKs upon tumor cell lines (K562, U937). The residual living Nalm6 tumor cells line were further declined in the PD-NK group compared to those in the HD-NK group, which indicated the moderate reinforcement in in vitro tumor-killing capacity and cytotoxicity. Additionally, a higher proportion of NK cells in the S and G2/M phases of the cell cycle was observed in PD-NKs, thereby indicated the increase in NK cell proliferative activity in early PD patients.
PD is acknowledged as a multi-system disorder characterized by significant neuroinflammatory and immune dysfunctions. For instance, Earls et al reported the efficient internalization and degradation of α-syn aggregates by NK cells via modulating the endosomal/lysosomal pathway47. The midbrain region of PD patients revealed higher levels of IFN-γ, accompanied by significant expression of α-synuclein48. Immunodysregulation within the peripheral immune system can result in the upregulation of inflammatory factors and the hyperactivation of a cascade of pro-inflammatory signals associated with PD development. Consistently, in this study, higher expression levels of gene sets related to immune responses (IFN-γ, INF-α, and complement) and metabolism were observed in PD-NKs according to the RNA-SEQ data, which indicated the IFN-γ-associated immune-inflammatory response is hyperactive in PD patients at the early stage of primary diagnosis. Collectively, our findings would supply new references to the NK cell-mediated pathogenesis of PD and facilitate the identification of novel biomarkers for PD diagnosis alongside the clinical examinations (e.g., α-syn, bradykinesia, resting tremor or rigidity).
Conclusions
Overall, in this study, we compared the multifaceted biological and transcriptomic signatures of both resident NK cells from PBMCs and expanded NK cells after ex vivo culture. Our findings outlined the similarities and differences of HD-NKs and PD-NKs both at the cellular and molecular levels, and in particular, the subpopulations and the landscape of gene expression pattern, which would benefit the further dissection of the pathogenesis of PD and the novel therapeutic strategies in future.
Data availability
Data is provided within the manuscript or supplementary information files. The datasets in the study are available from the corresponding author on reasonable request.
Abbreviations
- PD:
-
Parkinson’s disease
- NKs:
-
Natural killer cells
- GOBP:
-
Gene ontology biological process
- KEGG:
-
Kyoto encyclopedia of genes and genomes
- GSEA:
-
Gene set enrichment analysis
- PCA:
-
Principal component analysis
- PBMCs:
-
Peripheral blood-derived mononuclear cells
- HD-NKs:
-
Healthy donor-derived NK cells
- PD-NKs:
-
PD patient-derived NK cells
- RNA-SEQ:
-
RNA-sequencing
- rPD-NKs:
-
resident PD-NKs
- rHD-NKs:
-
resident HD-NKs
- PCA:
-
principal component analysis
- DEGs:
-
Differentially expressed genes
- VSEs:
-
Variable shear events
References
Tolosa, E., Garrido, A., Scholz, S. W. & Poewe, W. Challenges in the diagnosis of Parkinson’s disease. Lancet Neurol. 20(5), 385–397 (2021).
Samii, A., Nutt, J. G. & Ransom, B. R. Parkinson’s disease. Lancet (London, England) 363(9423), 1783–1793 (2004).
Cerri, S., Mus, L. & Blandini, F. Parkinson’s disease in women and men: what’s the difference?. J. Parkinson’s Dis. 9(3), 501–515 (2019).
Zhang, Y.-N. et al. Metabotropic glutamate receptor 5 inhibits α-synuclein-induced microglia inflammation to protect from neurotoxicity in Parkinson’s disease. J. Neuroinflammation 18(1), 23 (2021).
Kalia, L. V. & Lang, A. E. Parkinson’s disease. Lancet (London, England) 386(9996), 896–912 (2015).
Blauwendraat, C., Nalls, M. A. & Singleton, A. B. The genetic architecture of Parkinson’s disease. Lancet Neurol. 19(2), 170–178 (2020).
Ascherio, A. & Schwarzschild, M. A. The epidemiology of Parkinson’s disease: risk factors and prevention. Lancet Neurol. 15(12), 1257–1272 (2016).
Qi, C. & Liu, Q. Natural killer cells in aging and age-related diseases. Neurobiol Dis. 183, 106156 (2023).
Zhang, L., Zhang, Y. & Fan, D. Pathological role of natural killer cells in parkinson’s disease: a systematic review. Front. Aging Neurosci. 14, 890816 (2022).
Vavilova, J. D. et al. Reduced Immunosenescence of peripheral blood T cells in parkinson’s disease with CMV infection background. Int. J. Mol. Sci. 22(23), 13119 (2021).
Zhou, C. et al. Reduction of peripheral blood iNKT and γδT cells in patients with parkinson’s disease: an observational study. Front. Immunol. 11, 1329 (2020).
Tan, E. K. et al. Parkinson disease and the immune system - associations, mechanisms and therapeutics. Nat. Rev. Neurol. 16(6), 303–318 (2020).
Ye, H., Robak, L. A., Yu, M., Cykowski, M. & Shulman, J. M. Genetics and pathogenesis of parkinson’s syndrome. Ann. Rev. Pathol. 18, 95–121 (2023).
Zhang, L. et al. Natural killer cells: of-the-shelf cytotherapy for cancer immunosurveillance. Am. J. Cancer Res. 11(4), 1770–1791 (2021).
Nikzad, R. et al. Human natural killer cells mediate adaptive immunity to viral antigens. Sci. Immunol. 4(35), eaat8116 (2019).
Zhang, L., Meng, Y., Feng, X. & Han, Z. CAR-NK cells for cancer immunotherapy: from bench to bedside. Biomark. Res. 10(1), 12 (2022).
Cong, J. Metabolism of natural killer cells and other innate lymphoid cells. Front. Immunol. 2020, 11 (1989).
Earls RH, Menees KB, Chung J, Gutekunst CA, Lee HJ, Hazim MG, Rada B, Wood LB, Lee JK: NK cells clear α-synuclein and the depletion of NK cells exacerbates synuclein pathology in a mouse model of α-synucleinopathy. Proc. of the National Academy of Sciences of the United States of America 117(3):1762-1771 (2020)
Bokor, M., Faragó, A., Garam, T., Malatinszky, G. & Schnabel, R. Antibody-dependent cell-mediated cytotoxicity (ADCC) in Parkinson’s disease. J. Neurol. Sci. 115(1), 47–50 (1993).
Zhang, C. & Tian, Z. NK cell subsets in autoimmune diseases. J. Autoimmunity 83, 22–30 (2017).
Genssler, S. et al. Dual targeting of glioblastoma with chimeric antigen receptor-engineered natural killer cells overcomes heterogeneity of target antigen expression and enhances antitumor activity and survival. Oncoimmunology 5(4), e1119354 (2016).
Earls, R. H. et al. Intrastriatal injection of preformed alpha-synuclein fibrils alters central and peripheral immune cell profiles in non-transgenic mice. J. Neuroinflammation 16(1), 250 (2019).
Earls RH, Menees KB, Chung J, Gutekunst CA, Lee HJ, Hazim MG, Rada B, Wood LB, Lee JK: NK cells clear alpha-synuclein and the depletion of NK cells exacerbates synuclein pathology in a mouse model of alpha-synucleinopathy. Proc. of the National Academy of Sciences of the United States of America 117(3):1762-1771 (2020)
Zhang, L. et al. Characterization of the biological and transcriptomic signatures of natural killer cells derived from cord blood and peripheral blood. Am. J. Cancer Res. 13(8), 3531–3546 (2023).
Zhang, L. et al. Decoding the multidimensional signatures of resident and expanded natural killer cells generated from perinatal blood. Am. J. Cancer Res. 12(5), 2132–2145 (2022).
Liu, M., Meng, Y., Zhang, L., Han, Z. & Feng, X. High-efficient generation of natural killer cells from peripheral blood with preferable cell vitality and enhanced cytotoxicity by combination of IL-2, IL-15 and IL-18. Biochem. Biophys. Res. Commun. 534, 149–156 (2021).
Wei, Y. et al. High-efficient generation of VCAM-1+ mesenchymal stem cells with multidimensional superiorities in signatures and efficacy on aplastic anaemia mice. Cell Prolif. 53(8), e12862 (2020).
Liu, M., Meng, Y., Zhang, L., Han, Z. & Feng, X. High-efficient generation of natural killer cells from peripheral blood with preferable cell vitality and enhanced cytotoxicity by combination of IL-2, IL-15 and IL-18. Biochem. Biophys. Res. Commun. 534, 149–156 (2021).
Gao, H., Liu, M., Zhang, Y., Zhang, L. & Xie, B. Multifaceted characterization of the biological and transcriptomic signatures of natural killer cells derived from cord blood and placental blood. Cancer Cell Int. 22(1), 291 (2022).
Zhang, L. et al. MSX2 Initiates and accelerates mesenchymal stem/stromal cell specification of hPSCs by regulating TWIST1 and PRAME. Stem Cell Rep. 11(2), 497–513 (2018).
Zhao, Q. et al. Systematic comparison of hUC-MSCs at various passages reveals the variations of signatures and therapeutic effect on acute graft-versus-host disease. Stem Cell Res. Ther. 10(1), 354 (2019).
Huo, J. et al. Multifaceted characterization of the signatures and efficacy of mesenchymal stem/stromal cells in acquired aplastic anemia. Stem Cell Res. Ther. 11(1), 59 (2020).
Zhang, L. et al. Two-step generation of mesenchymal stem/stromal cells from human pluripotent stem cells with reinforced efficacy upon osteoarthritis rabbits by HA hydrogel. Cell Biosci. 11(1), 6 (2021).
Zhang, L. et al. Bone marrow-derived mesenchymal stem/stromal cells in patients with acute myeloid leukemia reveal transcriptome alterations and deficiency in cellular vitality. Stem Cell Res. Ther. 12(1), 365 (2021).
Beitz, J. M. Parkinson’s disease: a review. Front. Biosci. (Scholar edition) 6(1), 65–74 (2014).
Tysnes OB, Storstein A: Epidemiology of Parkinson’s disease. Journal of neural transmission (Vienna, Austria : 1996) 124(8):901-905. 2017
Zhang, L., Meng, Y., Feng, X. & Han, Z. CAR-NK cells for cancer immunotherapy: from bench to bedside. Biomarker Res. 10(1), 12 (2022).
Guillerey, C., Huntington, N. D. & Smyth, M. J. Targeting natural killer cells in cancer immunotherapy. Nat. Immunol. 17(9), 1025–1036 (2016).
Wu, S. Y., Fu, T., Jiang, Y. Z. & Shao, Z. M. Natural killer cells in cancer biology and therapy. Mol. Cancer 19(1), 120 (2020).
Arias, J., Yu, J., Varshney, M., Inzunza, J. & Nalvarte, I. Hematopoietic stem cell- and induced pluripotent stem cell-derived CAR-NK cells as reliable cell-based therapy solutions. Stem Cells Transl. Med. 10(7), 987–995 (2021).
Zhu, H. & Kaufman, D. S. Engineered human pluripotent stem cell-derived natural killer cells: the next frontier for cancer immunotherapy. Blood Sci. 1(1), 4–11 (2019).
Tian, H., Yuan, L., Song, Y., Deng, J. & Qi, X. W. A review of recent advances in nanomaterial-based stem cell therapy and the corresponding risks. Curr. Stem Cell Res. Ther. 17(3), 195–206 (2022).
Konstantin Nissen, S. et al. Changes in CD163+, CD11b+, and CCR2+ peripheral monocytes relate to Parkinson’s disease and cognition. Brain Behav. Immunity 101, 182–193 (2022).
Holbrook, J. et al. Natural killer cells have an activated profile in early Parkinson’s disease. J. Neuroimmunol. 382, 578154 (2023).
Dumitriu, A. et al. Integrative analyses of proteomics and RNA transcriptomics implicate mitochondrial processes, protein folding pathways and GWAS loci in Parkinson disease. BMC Med. Genom. 9, 5 (2016).
Mihara, T. et al. Natural killer cells of Parkinson’s disease patients are set up for activation: a possible role for innate immunity in the pathogenesis of this disease. Parkinsonism Relat. Disord. 14(1), 46–51 (2008).
Vijiaratnam, N., Simuni, T., Bandmann, O., Morris, H. R. & Foltynie, T. Progress towards therapies for disease modification in Parkinson’s disease. Lancet Neurol. 20(7), 559–572 (2021).
Tansey, M. G. et al. Inflammation and immune dysfunction in Parkinson disease. Nat. Rev. Immunol. 22(11), 657–673 (2022).
Acknowledgements
The coauthors thank the doctors and nurses in The Second Hospital & Cheeloo College of Medicine, Shandong Provincial Key Medical and Health Laboratory of Blood Ecology and Biointelligence and Shandong Provincial Youth Health Science and Technology Innovation Team in The Fourth People’s Hospital of Jinan Affiliated to Shandong Second Medical University for their technical support.
Funding
This work was supported by grants from Taishan Scholar Special Funding (tsqnz20240858), Natural Science Foundation of Shandong Province (ZR2025MS1382), The Senior Medical and Healthcare Talent Visiting Program of Shandong Province (202403001), Medical and Health Technology Project of Shandong Province (202402050122), Science and Technology Development Plan of Jinan Municipal Health Commission (2024301008, 2025202005), Natural Science Foundation of Hunan Province (2024JJ5365), Natural Science Foundation of Jiangxi Province (20252BAC210008).
Author information
Authors and Affiliations
Contributions
Xinyi Yue, and Qianwen Hu: designed and performed the experiments, collection and assembly of data, manuscript writing; Haoyang Cheng, Jiuzhou Diao, and Yansheng Du: helped with collection and assembly of data; Pengrui Fu, and Ping Wang: data analysis and interpretation, manuscript writing; Leisheng Zhang, and Qingbo Zhou: conception and design, revision, final approval of manuscript. All the co-authors have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics
All patients and healthy donors involved in this study have signed informed consents. Meanwhile, this study was approved by the Ethical Committee of The Second Hospital of Shandong University (approval no.: KYLL-2023LW050) according to the principle of Declaration of Helsinki.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yue, X., Hu, Q., Cheng, H. et al. Decoding the multifaceted cellular and transcriptomic variations of natural killer cells generated from patients with Parkinson’s disease. Sci Rep 15, 42750 (2025). https://doi.org/10.1038/s41598-025-26756-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-26756-w