Abstract
Acute lung injury (ALI) commonly occurs as a complication of sepsis, significantly increasing mortality rates in septic patients. Macrophages play a critical role in sepsis pathogenesis. KDM6B, a histone demethylase, has been reported to regulate macrophage death in ALI. In this study, we investigated the impact of KDM6B on macrophage processes during sepsis-induced ALI and elucidated the underlying molecular mechanisms. A cecal ligation and puncture (CLP)-induced septic mouse model was established to assess KDM6B’s effects on lung injury severity and survival outcomes. Raw264.7 macrophages were stimulated with lipopolysaccharide (LPS) and interferon-γ (IFN-γ) to create an in vitro cell injury model, followed by flow cytometric analysis for apoptosis detection, DCFH-DA staining for oxidative stress assessment, and western blot analysis for mitophagy evaluation. Chromatin immunoprecipitation assays were performed to determine KDM6B’s regulatory effects on histone H3 lysine 27 (H3K27) methylation status in the MFN1 promoter region. Results demonstrated upregulated KDM6B expression in CLP-induced septic mouse lungs. In vitro experiments revealed that KDM6B knockdown significantly suppressed LPS/IFN-γ-induced apoptosis while promoting mitophagy in macrophages. In vivo analyses showed that KDM6B suppression attenuated lung injury severity, inflammatory responses, and improved survival rates in septic mice. Mechanistically, KDM6B knockdown increased H3K27me3 levels, reduced MFN1 expression, and increased H3K27me3 enrichment specifically at the MFN1 promoter region. These findings demonstrate that KDM6B silencing facilitates H3K27me3 modification at the MFN1 promoter to inhibit its transcription, subsequently suppressing macrophage apoptosis and promoting mitophagy, which collectively mitigates sepsis-induced ALI progression.
Similar content being viewed by others
Introduction
Sepsis is a critical public health concern characterized by an aberrant host response to infection, leading to potentially fatal organ dysfunction1. The lung is the organ most susceptible to sepsis-induced damage, with up to 50% of septic patients in ICU developing acute lung injury (ALI)2. Globally, sepsis-induced ALI carries a mortality rate as high as 40%3. Current therapeutic strategies for sepsis-associated ALI primarily include antibiotic administration, fluid resuscitation, and vasoactive medications. However, these conventional approaches fail to fully resolve the condition and are associated with significant adverse effects4. Despite mounting research efforts that have enhanced our understanding of the pathophysiological mechanisms underlying sepsis-induced ALI, the precise molecular pathways driving its progression remain incompletely elucidated.
A defining characteristic of ALI is the development of acute inflammation that leads to pulmonary interstitial edema and immune cell infiltration into alveolar spaces4. Macrophages, a type of innate immune cell, exhibit dual roles in both promoting and suppressing inflammatory responses, and their functional dysregulation is closely associated with sepsis pathogenesis5;6. Specifically, macrophages modulate inflammatory processes through polarization into pro-inflammatory and anti-inflammatory phenotypes, thereby contributing to tissue damage and repair mechanisms7. Furthermore, alveolar macrophage death amplifies inflammatory signaling by impairing the function of other immune cell populations, ultimately exacerbating the severity of sepsis-induced ALI8. Additionally, excessive production of reactive oxygen species (ROS) in the lungs represents a hallmark of sepsis-induced ALI9. ROS can trigger mitophagy—a cellular process that eliminates damaged mitochondria to maintain mitochondrial and cellular homeostasis10. During sepsis, mitophagy exerts regulatory effects on inflammatory responses and plays a critical role in modulating lung injury progression11. Nevertheless, the precise mechanisms governing macrophage behavior, particularly mitophagy regulation, in the context of sepsis-induced ALI remain poorly understood.
KDM6B, also known as JMJD3, is a histone H3 lysine 27 (H3K27) demethylase that specifically catalyzes the removal of methyl groups from H3K27 trimethylation (H3K27me3) and H3K27 dimethylation (H3K27me2)12. KDM6B has been implicated in various pathological conditions, including cancer, atherosclerosis, and neurodegenerative disorders13,14,15. Notably, its expression is upregulated in septic patients and modulates inflammatory signaling pathways16. Furthermore, KDM6B has been shown to influence macrophage cell death in ALI17. However, the role of KDM6B in sepsis-induced ALI remains to be fully elucidated.
MFN1 is a mitochondrial fusion protein localized to the outer mitochondrial membrane. Its absence in cells results in mitochondrial fragmentation, accompanied by reduced mitochondrial respiration and impaired membrane potential18. MFN1 has been implicated in sepsis-related pathologies, including sepsis-induced acute kidney injury and sepsis-associated encephalopathy19;20. However, the role of MFN1 in sepsis-induced ALI remains unclear.
In the present study, we identified upregulated KDM6B expression in ALI through microarray analysis. Therefore, we investigated the effects of KDM6B on macrophage apoptosis and mitophagy in sepsis-induced ALI. Additionally, we explored whether KDM6B regulates MFN1 through histone modifications. This study uncovers a novel epigenetic mechanism underlying macrophage mitophagy regulation in sepsis-induced ALI, highlighting KDM6B as a potential therapeutic target.
Materials and methods
Bioinformatic analysis
Differentially expressed genes were identified using the GSE3037 dataset from the GEO online database (https://www.ncbi.nlm.nih.gov/geo/). Data analysis was performed using the “Limma” R package. Results were visualized using a volcano plot. Genes with |Log2(fold change)| > 0.6 and P < 0.05 were defined as differentially expressed genes and further displayed in a heatmap using the “heatmap” R package.
Animal experiments
C57BL/6 mice (male, 10–12 weeks old, 23–25 g) were obtained from Charles River (Beijing, China). Animals were housed under a 12 h light/dark cycle at 22 °C with ad libitum access to food and water. The animal experiments were conducted according to the guide for the care and use of laboratory animals. The animal study was approved by the Laboratory Animals Ethics and Welfare Committee of MDKN Biotechnology Co., Lt (No. MDKN-2024-287).
Mice were randomly assigned to four groups: sham, cecal ligation and puncture (CLP), CLP + short hairpin RNA targeting KDM6B (shKDM6B), and CLP + short hairpin RNA negative control (shNC). The septic mouse model was established via CLP surgery21. Briefly, mice were anesthetized with isoflurane. A 1-cm ventral midline incision was made, and the cecum was isolated. The proximal 1/3 of the cecum was ligated using 4 − 0 sutures and punctured twice with a 22-gauge needle to release a small volume of fecal material into the peritoneal cavity. The cecum was then returned to the abdominal cavity, and the incision was sutured. Subcutaneous fluid resuscitation (50 mL/kg normal saline) was administered, followed by recovery in standard housing conditions. Sham-operated mice underwent identical anesthesia and surgical procedures without cecal ligation or puncture. Lentivirus carrying shKDM6B and shNC were synthesized by GenePharma (Shanghai, China). Mice in the CLP + shNC and CLP + shKDM6B groups received intravenous injections of 2 × 107 TU lentivirus via the tail vein 5 days prior to CLP surgery22. For survival analysis, eight mice per group were monitored, with survival status recorded every 12 h. Kaplan-Meier survival curves and log-rank tests were performed to evaluate survival outcomes. Six mice per group were euthanized 24 h post-surgery via isoflurane inhalation (Cat. No. 792632; Sigma-Aldrich, St. Louis, MO, USA). The peripheral blood, bronchial alveolar lavage fluid (BALF), and lungs were collected. The upper and middle lobes of the right lung were weighed, while remaining lung tissue was used for histopathological examination and protein expression analysis.
Cell culture
Raw264.7 cells (Cat. No. TIB-71; ATCC, Manassas, VA, USA) were cultured in Dulbecco’s Modified Eagle’s Medium (DMEM; Cat. No. 30-2002; ATCC) supplemented with 10% fetal bovine serum (FBS; Cat. No. 30-2020; ATCC) at 37 °C in a 5% CO2 humidified incubator.
To establish a macrophage injury model, Raw264.7 cells were treated with 100 ng/mL lipopolysaccharide (LPS; Cat. No. SMB00610; Sigma-Aldrich) and 20 ng/mL interferon-γ (IFN-γ; Cat. No. I4777; Sigma-Aldrich) for 24 h.
Cell transfection
Short hairpin RNA targeting KDM6B (shKDM6B), its negative control (shNC), MFN1 overexpression plasmids (oeMFN1), and the empty vector (oeNC) were purchased from Sangon Biotech (Shanghai, China). For transfection, Raw264.7 cells were seeded into 6-well plates at a density of 2 × 105 cells per well. The plasmids were transfected using Lipofectamine 3000 (Cat. No. L3000150; Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s protocol. After 48 h of incubation, cells were harvested for subsequent experiments.
Flow cytometry
Apoptosis was assessed using the Annexin V-PE/7-AAD Apoptosis Detection Kit (Cat. No. A213-01; Vazyme Biotech Co., Ltd., Nanjing, China). Raw264.7 cells (2 × 105 cells) were digested with trypsin, harvested, and resuspended in 100 µL of binding buffer. The cells were incubated with 5 µL Annexin V-PE and 5 µL 7-AAD staining solution at room temperature for 10 min in the dark. Following incubation, 400 µL of binding buffer was added to the cell suspension, and apoptotic cells were analyzed by flow cytometry.
Determination of ROS levels
ROS levels were quantified using the ROS Assay Kit (Cat. No. E004-1-1; Jiancheng, Nanjing, China). The DCFH-DA staining solution was diluted in serum-free medium to a final concentration of 10 µM. Raw264.7 cells were seeded into 6-well plates and incubated with the DCFH-DA solution at 37 °C for 30 min. Following incubation, the cells were resuspended in serum-free medium and washed twice. Fluorescence intensity was measured using a fluorescence microplate reader.
Quantitative real-time polymerase chain reaction (qPCR)
Total RNA was extracted from mouse lung tissue and Raw264.7 cells using TRIzol reagent (Invitrogen). RNA purity was assessed by measuring the A260/280 nm ratio, and cDNA was synthesized via reverse transcription using the HiScript III 1st Strand cDNA Synthesis Kit (+ gDNA wiper) (Cat. No. R312-02; Vazyme Biotech Co., Ltd.). Subsequently, qPCR was performed with the Taq Pro Universal SYBR qPCR Master Mix (Cat. No. Q712-02; Vazyme Biotech Co., Ltd.). Relative mRNA expression levels were calculated using the 2−∆∆Ct method.
Western blot
Raw264.7 cells and mouse lung tissues were lysed using radioimmunoprecipitation assay (RIPA) lysis buffer. After centrifugation at 14,000×g for 15 min, the supernatant was collected. A portion of the supernatant was used to determine protein concentration using the bicinchoninic acid Protein Assay Kit (Cat. No. P0012; Beyotime Biotechnology, Shanghai, China). The remaining supernatant was separated via sodium dodecyl sulfate-polyacrylamide gel electrophoresis. Proteins were transferred onto polyvinylidene difluoride membranes (Cat. No. E801-01; Vazyme Biotech Co., Ltd.). The membranes were blocked with 5% skim milk in Tris-buffered saline with Tween 20 for 1 h. Membranes were then incubated with primary antibodies against H3K27me3 (ab192985, Abcam, Cambridge, MA, USA), H3 (17168-1-AP, Proteintech, Wuhan, China), MFN1 (13798-1-AP, Proteintech), MFN2 (12186-1-AP, Proteintech), Drp1 (12957-1-AP, Proteintech), ATG7 (10088-2-AP, Proteintech), PINK1 (23274-1-AP, Proteintech), Parkin (14060-1-AP, Proteintech), TOM20 (11802-1-AP, Proteintech), LC3 (14600-1-AP, Proteintech), and GAPDH (10494-1-AP, Proteintech) at 4 °C overnight. Subsequently, membranes were incubated with horseradish peroxidase (HRP)-conjugated goat anti-rabbit IgG secondary antibody (ab6721, Abcam) at room temperature for 1 h. Protein bands were visualized using a BeyoECL chemiluminescence detection system (Cat. No. P0018S; Beyotime Biotechnology).
Immunofluorescence (IF)
Raw264.7 cells were fixed with 4% paraformaldehyde (Cat. No. P0099; Beyotime Biotechnology) for 20 min and subsequently permeabilized with PBS containing 0.1% Triton X-100 (Cat. No. T8787; Sigma-Aldrich) for 10 min. After PBS washing, the cells were incubated with anti-H3K27me3 (ab192985, Abcam) at 4 °C overnight, followed by incubation with Alexa Fluor 488-conjugated goat anti-rabbit IgG (ab150077, Abcam) at room temperature for 1 h in the dark. Cell nuclei were visualized with 4′,6-diamidino-2-phenylindole (Cat. No. C1006; Beyotime Biotechnology) staining prior to fluorescence microscopy observation.
Chromatin Immunoprecipitation (CHIP)
The effect of KDM6B on the interaction between H3K27me3 and the MFN1 promoter was analyzed using a ChIP kit (Cat. No. ABS50034; Absin, Shanghai, China). Raw264.7 cells (1 × 106 cells) were fixed with 1% paraformaldehyde for 10 min, and the reaction was terminated by adding glycine. After being washed with PBS containing protease inhibitors, the cells were collected, centrifuged at 800×g for 5 min, and sonicated for 30 min to fragment nuclei. A 50 µL aliquot of the sample was mixed with 450 µL ChIP dilution buffer and incubated with anti-H3K27me3 (ab192985, Abcam) or normal IgG (A7016, Beyotime Biotechnology) at 4 °C for 4 h. Subsequently, 20 µL of protein A/G magnetic beads was added, and the mixture was incubated at 4 °C for 2 h. After washing, the sample was incubated with ChIP elution buffer containing proteinase K at 65 °C for 2 h, followed by incubation at 95 °C for 10 min. Following cooling and centrifugation at 10,000×g for 10 s, the magnetic beads were removed, and the supernatant was collected. The eluted DNA was subjected to qPCR to quantify enrichment of the MFN1 promoter region.
Lung wet/dry (W/D) weight ratio determination
The upper and middle lobes of the right lung were weighed immediately after collection. Following drying at 60 °C for 48 h, the lobes were reweighed. The W/D weight ratio was calculated as the ratio of wet weight to dry weight.
Hematoxylin and Eosin (H&E) staining
Lung tissues from mice were fixed in 4% paraformaldehyde for 48 h. The tissues were embedded in paraffin, sectioned into 5-µm slices, and mounted on slides. After dewaxing with xylene and rehydration through graded ethanol solutions, the sections were stained with hematoxylin (Cat. No. H3136; Sigma-Aldrich) and eosin (Cat. No. 230251; Sigma-Aldrich) consecutively. Stained sections were visualized under light microscopy.
Histopathology was scored as previously described23. Five key parameters of the sections were evaluated, including alveolar congestion, hemorrhage, neutrophil infiltration, alveolar wall thickness, and hyaline membrane formation. The final score ranged from 0 to 4 points. In detail, 0 points: no injury, 1 point: mild injury, 2 points: moderate injury, 3 points: severe injury, and 4 points: maximal injury. Each section was scored blindly by investigators who were unaware of the grouping and experimental details.
Enzyme-linked immunosorbent assay (ELISA)
The levels of interleukin-6 (IL-6) and tumor necrosis factor-α (TNF-α) in the serum and BALF of mice were detected using the mouse IL-6 ELISA kit (Cat. No. J24111; JILID Biotechnology, Wuhan, China) and mouse TNF-α ELISA kit (Cat. No. J24082; JILID Biotechnology) according to the manufacturer’s instructions.
Immunohistochemistry (IHC)
Paraffin-embedded lung tissue sections were deparaffinized and rehydrated through graded ethanol solutions. Antigen retrieval was performed using a universal antigen retrieval solution (Cat. No. P0088; Beyotime Biotechnology). Endogenous peroxidase activity was blocked by incubation with 3% H2O2 for 15 min. The sections were incubated with primary antibodies against KDM6B (ab38113, Abcam) and MFN1 (13798-1-AP, Proteintech) at 4 °C overnight, followed by incubation with HRP-conjugated goat anti-rabbit IgG (ab6721, Abcam) at 37 °C for 30 min. Subsequently, the sections were stained with 3,3′-diaminobenzidine (DAB; Cat. No. D8001; Sigma-Aldrich) and visualized under light microscopy.
Statistical analysis
Results were expressed as mean ± standard deviation. Comparisons between two groups were analyzed using Student’s t test, while differences among three or more groups were evaluated using one-way ANOVA with Tukey’s post-hoc test. Data analysis was performed using GraphPad Prism 8 software. A P value < 0.05 was considered statistically significant.
Results
KDM6B expression is elevated in CLP-induced mice
To identify potential genes involved in sepsis-induced ALI, we conducted a microarray analysis using the GSE3037 dataset. A total of 45 genes were upregulated and 11 genes were downregulated in ALI compared to the control group (Fig. 1A, B). Among these, KDM6B expression was significantly higher in ALI samples than in the control group. Subsequently, we established a CLP-induced mouse model of sepsis. Lung tissues were collected, and KDM6B expression was assessed via qPCR and IHC. The results demonstrated that KDM6B expression was elevated in the lungs of CLP mice compared to sham controls (Fig. 1C, D). These findings suggest that KDM6B may play a role in sepsis-induced ALI.
KDM6B expression is elevated in CLP-induced mice. A The heatmap and B the volcano plot showed the differentially expressed genes between the ALI and control groups. C KDM6B expression in the lungs of mice after CLP surgery was detected using qPCR. D KDM6B expression in the lungs of mice after CLP surgery was detected using IHC, and IHC results were quantified. ***P < 0.001 vs. the sham group.
Knockdown of KDM6B inhibits apoptosis and promotes mitophagy of macrophages
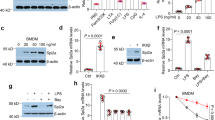
To simulate sepsis-induced macrophage injury in the lungs, Raw264.7 cells were treated with LPS and IFN-γ to establish an in vitro model. First, shKDM6B was transfected into Raw264.7 cells, and KDM6B expression was observed to be reduced in the shKDM6B group compared to the shNC group (Fig. 2A). The effects of KDM6B on cellular biological behaviors were then analyzed. Flow cytometry revealed that LPS and IFN-γ treatment induced apoptosis in Raw264.7 cells compared to the control group, whereas KDM6B knockdown attenuated the increase in apoptosis (Fig. 2B, C). Additionally, ROS levels were elevated in LPS and IFN-γ-treated macrophages, but this elevation was counteracted by KDM6B silencing (Fig. 2D). Furthermore, LPS and IFN-γ treatment downregulated the expression of PINK1, Parkin, and LC3II/LC3I while upregulating TOM20 levels in Raw264.7 cells; however, KDM6B silencing reversed this downregulation (Fig. 2E). Collectively, KDM6B knockdown inhibits apoptosis and promotes mitophagy in LPS and IFN-γ-treated Raw264.7 cells.
Knockdown of KDM6B inhibits apoptosis and promotes mitophagy of macrophages. A KDM6B expression was measured using qPCR after shNC and shKDM6B transfection. Raw264.7 cells were transfected with shNC and shKDM6B, followed by stimulation of LPS and IFN-γ, and cellular behaviors were evaluated. B, C Cell apoptosis was evaluated using flow cytometry. D ROS level was measured using DCFH-DA. E Protein levels of mitophagy markers were examined using western blot. ***P < 0.001 vs. the shNC or control group. ###P < 0.001 vs. the LPS + IFN-γ + shNC group.
Knockdown of KDM6B promotes the enrichment of H3K27me3 in MFN1 promoter
KDM6B is a histone demethylase that removes H3K27me3 and H3K27me2 marks, thereby regulating gene expression24. In this study, we focused on H3K27me3 specifically. We found that H3K27me3 levels were lower in the lungs of CLP mice than those in sham mice (Fig. 3A). In LPS and IFN-γ-treated Raw264.7 cells, knockdown of KDM6B resulted in increased H3K27me3 levels (Fig. 3B, C). To identify genes potentially regulated by H3K27me3, we reviewed published studies and selected candidate genes associated with mitochondrial dysfunction in ALI25,26,27. qPCR results indicated that KDM6B knockdown reduced MFN1 and MFN2 expression but did not significantly alter Drp1 or ATG7 levels (Fig. 3D). Western blot analysis confirmed that KDM6B silencing downregulated MFN1 protein expression without affecting MFN2, Drp1, or ATG7 (Fig. 3D). Therefore, MFN1 was selected for further investigation. ChIP assays revealed that KDM6B knockdown enhanced H3K27me3 enrichment in the MFN1 promoter region (Fig. 3E). Collectively, KDM6B inhibits H3K27me3 enrichment in the MFN1 promoter, thereby promoting MFN1 expression.
Knockdown of KDM6B promotes the enrichment of H3K27me3 in MFN1 promoter. A H3K27me3 levels in the lungs of mice in the sham and CLP group were measured using western blot, and H3 served as the internal control. B After KDM6B knockdown, H3K27me3 levels in Raw264.7 cells were examined by western blot, and H3 acted as the internal control. C H3K27me3 levels were observed using IF after KDM6B knockdown in Raw264.7 cells. D The mRNA expression levels of MFN1, MFN2, Drp1, and ATG7, as well as their protein levels were measured after KDM6B knockdown using qPCR and western blot, respectively. E The effect of KDM6B knockdown on the enrichment of H3K27me3 in the promoter of the MFN1 gene was evaluated using CHIP. ***P < 0.001 vs. the shNC group. ns: no significance vs. the shNC group.
Overexpression of MFN1 counteracts the effect on macrophage apoptosis and autophagy induced by KDM6B silence
To further investigate the interplay between KDM6B and MFN1 in regulating macrophage apoptosis and mitophagy, we conducted rescue experiments. MFN1 overexpression in Raw264.7 cells was achieved via oeMFN1 transfection, as confirmed by comparison to oeNC transfection (Fig. 4A). Cellular behaviors were assessed in Raw264.7 cells treated with LPS and IFN-γ. KDM6B knockdown suppressed apoptosis, whereas MFN1 overexpression reversed this anti-apoptotic effect (Fig. 4B and C). ROS levels were reduced following KDM6B knockdown, but this decrease was counteracted by MFN1 overexpression (Fig. 4D). Western blot analysis revealed that KDM6B silencing upregulated PINK1, Parkin, and LC3II/LC3I while downregulating TOM20; however, MFN1 overexpression reversed these protein expression changes induced by KDM6B knockdown (Fig. 4E). Collectively, KDM6B silencing inhibits apoptosis and promotes mitophagy in macrophages by downregulating MFN1 expression.
Overexpression of MFN1 counteracts the effect on macrophage apoptosis and mitophagy induced by KDM6B silence. A MFN1 expression was measured using qPCR after oeNC and oeMFN1 transfection. Raw264.7 cells were transfected with shNC, shKDM6B, oeNC, and oeMFN1, followed by stimulation of LPS and IFN-γ, and cellular behaviors were evaluated. B, C Cell apoptosis was evaluated using flow cytometry. D ROS level was measured using DCFH-DA. E Protein levels of mitophagy markers were examined using western blot. ***P < 0.001 vs. the oeNC or shNC group. ###P < 0.001 and ##P < 0.01 vs. the shKDM6B + oeNC group.
Knockdown of KDM6B improves survival rate and attenuates lung damage in CLP mice by regulating MFN1 expression
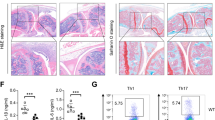
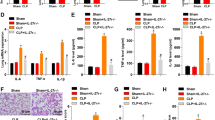
The role of KDM6B in lung injury was investigated in CLP-induced sepsis models. Mice were injected with lentiviral vectors carrying shKDM6B or shNC prior to CLP model establishment. Log-rank test results demonstrated that the survival rate of CLP mice was significantly lower than that of sham-operated mice, whereas KDM6B knockdown improved survival in CLP mice (Fig. 5A). Lung tissues were collected for histopathological analysis. Histological examination revealed exacerbated lung injury in CLP mice, but KDM6B knockdown significantly attenuated pathological damage (Fig. 5B). Consistently, the lung injury score results showed that compared to the sham group, CLP induced severe or maximal lung injury, while knockdown of KDM6B reduced lung injury score in CLP mice (Fig. 5C). The lung wet/dry (W/D) weight ratio, a marker of pulmonary edema, was elevated in CLP mice and reversed by KDM6B silencing (Fig. 5D). ELISA results showed that the levels of TNF-α and IL-6 were elevated in the serum and BALF of CLP mice, while knockdown of KDM6B reversed this elevation (Fig. 5E–H). MFN1 expression was upregulated in the lungs of CLP mice, and this increase was counteracted by KDM6B knockdown (Fig. 5I and J). In summary, targeting the KDM6B/MFN1 axis through KDM6B silencing enhances survival and alleviates lung injury in septic mice.
Knockdown of KDM6B improves survival rate and attenuates lung damage in CLP mice by regulating MFN1 expression. Mice were injected with lentivirus carrying shNC and shKDM6B, and underwent CLP surgery. A Survival rate of mice was monitored every 12 h and analyzed using Log-rank test. B Lung injury was evaluated using H&E staining. C Lung injury scores of mice in each group. D The wet (W) weight and dry (D) weight of the lungs were detected, and W/D weight ratio was quantified. E TNF-α and F IL-6 levels in the serum, as well as G TNF-α and H IL-6 levels in the BALF were measured using ELISA. I The expression of MFN1 in the lungs was detected using IHC, and J IHC results were quantified. ***P < 0.001 and **P < 0.01 vs. the sham group. ###P < 0.001 and ##P < 0.01 vs. the CLP + shNC group.
Discussion
The present study elucidates a previously unrecognized role of KDM6B in exacerbating sepsis-induced ALI by epigenetically regulating MFN1 to disrupt mitochondrial homeostasis in macrophages. These findings reveal a novel regulatory axis involving KDM6B in sepsis-induced ALI.
The involvement of KDM6B in pathological processes has been reported in prior studies. Samanta et al. 17 demonstrated that a combination of 5-azacytidine and trichostatin A protects against endotoxemia-induced ALI by inhibiting the activation of the STAT3/KDM6B pathway. Peng et al. 28 showed that KDM6B deficiency attenuates LPS-induced ALI by suppressing ferroptosis in alveolar epithelial cells. However, the role of KDM6B in sepsis-induced ALI has remained unexplored. In this study, KDM6B expression was elevated in the lungs of CLP-induced septic mice, and its knockdown contributed to improving survival rates. Moreover, we found that knockdown of KDM6B attenuated lung injury in CP mice, and we observed a significant reduction in lung injury scores, indicating that KDM6B silencing can preserve alveolar structure and reduce hallmarks of injury such as alveolar wall thickening, hemorrhage, and neutrophil infiltration. Concurrently, the attenuation of the systemic inflammatory response was evidenced by markedly decreased levels of TNF-α and IL-6 in the serum and BALF, indicating that KDM6B silence is crucial for suppressing systemic and local lung inflammation. This study directly links the downregulation of KDM6B to the alleviation of uncontrolled inflammation, a central driver of organ dysfunction and mortality in sepsis. These findings collectively demonstrate that targeting KDM6B not only improves survival but also confers concrete histological and immunological benefits in sepsis-induced ALI, suggesting that KDM6B represents a promising therapeutic target for sepsis-induced ALI.
To explore how KDM6B affects lung injury, we focused on the functions of macrophages. Macrophages play a critical role in inflammatory regulation during sepsis. Macrophage infiltration is a hallmark of ALI and is closely associated with cell apoptosis29. Several previous studies have investigated how KDM6B regulates macrophage behaviors in various diseases, primarily focusing on mechanisms such as macrophage polarization, pyroptosis, and inflammatory infiltration30,31,32. In ALI, KDM6B promotes pyroptosis of bone marrow-derived macrophages17. In the present study, our findings demonstrate that KDM6B knockdown reduces apoptosis in LPS/IFN-γ-treated macrophages, suggesting that the protective effect of KDM6B silencing on lung injury partially relies on the inhibition of macrophage apoptosis.
Mitochondrial damage disrupts cellular homeostasis, increases ROS production, induces oxidative stress, and activates inflammatory pathways10. Thus, mitophagy restores cellular homeostasis by eliminating damaged mitochondria. However, inhibiting mitophagy paradoxically promotes macrophage activation, thereby increasing survival rates in septic mice33. Moreover, PINK1-mediated mitophagy mitigates macrophage injury and reduces CLP-induced ALI34. Zhu et al. 22 demonstrated that PRKCA overexpression attenuates sepsis-induced lung injury by enhancing mitophagy while simultaneously reducing inflammatory responses and ROS production in macrophages. Li et al. 35 reported that Kahweol alleviates septic ALI by promoting mitophagy, facilitating the transition of mitochondria from fission to fusion, and suppressing oxidative stress. Despite these findings, the regulatory mechanisms governing mitophagy in macrophages during septic ALI remain poorly understood. The role of KDM6B in modulating mitophagy is yet to be elucidated. Our results revealed that LPS/IFN-γ treatment elevated ROS levels and suppressed mitophagy in macrophages, aligning with a prior study33. Furthermore, KDM6B knockdown enhanced mitophagy and decreased ROS levels in LPS/IFN-γ-stimulated macrophages, indicating that KDM6B silencing mitigates sepsis-induced ALI through mitophagy promotion.
As a histone demethylase, KDM6B typically removes repressive H3K27me3 and permissive H3K27me2 marks from histones. Histone methylation patterns at gene promoters are well-characterized. Specifically, H3K27me3 is associated with gene repression, whereas H3K27me2 is linked to transcriptional activation36. Our findings demonstrated that KDM6B knockdown increased H3K27me3 levels, consistent with prior reports37. To determine which genes are regulated by KDM6B-mediated H3K27me3 modification, we identified MFN1 as a novel downstream target of KDM6B. Silencing KDM6B reduced MFN1 expression in macrophages in vitro. Furthermore, KDM6B knockdown enhanced H3K27me3 enrichment at the MFN1 promoter, suggesting that KDM6B-mediated removal of H3K27me3 relieves repression of MFN1 transcription. MFN1, a mitochondrial fusion-related protein, plays a critical role in regulating mitophagy in macrophages38. Its involvement in sepsis has been previously investigated. For instance, CaMKIV-mediated downregulation of MFN1 attenuates oxidative stress-induced kidney injury19, and MFN1 overexpression ameliorates cognitive dysfunction in sepsis-associated encephalopathy by mitigating mitochondrial dysfunction20. However, the role of MFN1 in sepsis-induced ALI remains unclear. A single prior study reported that elevated MFN1 expression alleviates endotoxin-induced lung injury25. In our experiments, overexpression of MFN1 reversed the inhibition of apoptosis and the promotion of mitophagy induced by KDM6B knockdown, indicating that KDM6B silencing modulates macrophage apoptosis and mitophagy by suppressing MFN1 expression through H3K27me3 modification. Moreover, IHC results showed that MFN1 levels were elevated in the lungs of CLP mice, while knockdown of KDM6B reversed this elevation, which is consistent with the mechanism axis we have observed in vitro. The downregulation of MFN1, resulting from KDM6B silencing, serves as the critical missing link that connects the upstream epigenetic mechanism to the downstream improvement in clinical phenotypes—reduced inflammation, attenuated histological damage, and ultimately, enhanced survival. We considered that MFN1 is functionally significant in the complex pathophysiology of sepsis-induced ALI in vivo. Nevertheless, the direct impact of MFN1 on lung injury in vivo has yet to be elucidated and warrants further investigation.
Despite our findings that highlight the crucial role of KDM6B in sepsis-induced ALI, this study has several limitations. First, we only investigated the effect of KDM6B on lung injury, while its influence on other physiological processes, such as immune cell differentiation, inflammation resolution, and tissue repair remains unclear. In addition, our in vivo approach utilized shRNA lentivirus strategy. The potential for off-target effects and the long-term safety profile of modulating KDM6B activity require further rigorous investigation using conditional knockout models or cell-specific delivery systems in future studies. These limitations underscore the necessity for future research to comprehensively evaluate the safety and specificity of KDM6B-targeted therapies.
In conclusion, KDM6B silencing inhibits macrophage apoptosis and promotes mitophagy by reducing MFN1 expression, thereby alleviating lung injury. Mechanistically, KDM6B knockdown enhances H3K27me3 modification at the MFN1 promoter, leading to transcriptional repression of MFN1. This study elucidates a novel mechanism underlying the pathogenesis of sepsis-induced ALI and provides a theoretical foundation for developing therapeutic strategies targeting KDM6B.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Zheng, Y. et al. A diagnostic model for sepsis-induced acute lung injury using a consensus machine learning approach and its therapeutic implications. J. Transl. Med. 21, 620 (2023).
Sevransky, J. E. et al. Mortality in sepsis versus non-sepsis induced acute lung injury. Crit. Care 13, R150 (2009).
Sun, B. et al. Acute lung injury caused by sepsis: how does it happen? Front. Med. 10, 1289194 (2023).
Xu, Y. et al. Mechanisms of Sepsis-Induced acute lung injury and advancements of natural small molecules in its treatment. Pharmaceuticals 17, 472 (2024).
Qiu, Z. et al. Bidirectional effects of moxifloxacin on the pro–inflammatory response in lipopolysaccharide-stimulated mouse peritoneal macrophages. Mol. Med. Rep. 18, 5399–5408 (2018).
Gan, Z. et al. Excitatory amino acid transporter supports inflammatory macrophage responses. Sci. Bull. 69, 2405–2419 (2024).
Wang, Z. & Wang, Z. The role of macrophages polarization in sepsis-induced acute lung injury. Front. Immunol. 14, 1209438 (2023).
Fan, E. & Fan, J. Regulation of alveolar macrophage death in acute lung inflammation. Respir. Res. 19, 50 (2018).
Liu, B. et al. Microrna-1224-5P Aggravates Sepsis-Related Acute Lung Injury in Mice. Oxid. Med. Cell. Longev. 9493710 (2022).
Cheng, H., Wang, X., Yao, J., Yang, C. & Liu, J. Mitophagy and ferroptosis in sepsis-induced Ali/Ards: molecular mechanisms, interactions and therapeutic prospects of medicinal plants. J. Inflamm. Res. 17, 7819–7835 (2024).
Mohsin, M., Tabassum, G., Ahmad, S., Ali, S. & Ali, S. M. The role of mitophagy in pulmonary sepsis. Mitochondrion 59, 63–75 (2021).
Williams, K. et al. The histone lysine demethylase Jmjd3/Kdm6B is recruited to P53 bound promoters and enhancer elements in a P53 dependent manner. Plos One 9, e96545 (2014).
Lagunas-Rangel, F. A. Kdm6B (Jmjd3) and its dual role in cancer. Biochimie 184, 63–71 (2021).
Neele, A. E. et al. Myeloid Kdm6B deficiency results in advanced atherosclerosis. Atherosclerosis 275, 156–165 (2018).
Burchfield, J. S., Li, Q., Wang, H. Y. & Wang, R. F. Jmjd3 as an epigenetic regulator in development and disease. Int. J. Biochem. Cell. Biol. 67, 148–157 (2015).
Chen, Y. et al. Jmjd3 is involved in neutrophil membrane proteinase 3 overexpression during the hyperinflammatory response in early sepsis. Int. Immunopharmacol. 59, 40–46 (2018).
Samanta, S. et al. Dnmt and Hdac inhibitors together abrogate endotoxemia mediated macrophage death by Stat3-Jmjd3 signaling. Int. J. Biochem. Cell. Biol. 102, 117–127 (2018).
Son, J. M. et al. Mitofusin 1 and optic atrophy 1 shift metabolism to mitochondrial respiration during aging. Aging Cell 16, 1136–1145 (2017).
Zhang, X. et al. Camkiv regulates mitochondrial dynamics during sepsis. Cell Calcium 92, 102286 (2020).
Duan, H. et al. Nrf2 mitigates sepsis-associated encephalopathy-induced hippocampus ferroptosis via modulating mitochondrial dynamic homeostasis. Int. Immunopharmacol. 143, 113331 (2024).
Reilly, B. et al. Necroptosis-mediated ecirp release in sepsis. J. Inflamm. Res. 15, 4047–4059 (2022).
Zhu, Q. J. et al. Prkca promotes mitophagy through the Mir-15a-5P/Pdk4 axis to relieve sepsis-induced acute lung injury. Infect. Immun. 91, e46522 (2023).
Li, R. et al. 4-Octyl itaconate alleviates endothelial cell inflammation and barrier dysfunction in Lps-Induced sepsis via modulating Tlr4/Mapk/Nf-Kappab signaling: 4-octyl itaconate alleviates endothelial dysfunction. Mol. Med. 31, 240 (2025).
Xiao, Z. G. et al. The roles of histone demethylase Utx and Jmjd3 (Kdm6B) in cancers: current progress and future perspectives. Curr. Med. Chem. 23, 3687–3696 (2016).
Song, K. et al. Dexmedetomidine modulates mitochondrial dynamics to protect against endotoxin-induced lung injury via the protein kinase C-a/Haem Oxygenase-1 signalling pathway. Biomarkers 27, 159–168 (2022).
Shi, J. et al. Pi3K/Akt pathway-mediated Ho-1 induction regulates mitochondrial quality control and attenuates endotoxin-induced acute lung injury. Lab. Investig. 99, 1795–1809 (2019).
Wang, C., Yuan, J. & Du, J. Resveratrol alleviates acute lung injury through regulating Plscr-3-mediated mitochondrial dysfunction and mitophagy in a cecal ligation and puncture model. Eur. J. Pharmacol. 913, 174643 (2021).
Peng, J., Fan, B., Bao, C. & Jing, C. Jmjd3 deficiency alleviates lipopolysaccharide-induced acute lung injury by inhibiting alveolar epithelial ferroptosis in a Nrf2-dependent manner. Mol Med. Rep. 24, 807 (2021).
Cui, H. & Zhang, Q. Dexmedetomidine ameliorates lipopolysaccharide-Induced acute lung injury by inhibiting the Pi3K/Akt/Foxo1 signaling pathway. J. Anesth. 35, 394–404 (2021).
Xun, J. et al. Cancer-derived exosomal Mir-138-5P modulates polarization of tumor-associated macrophages through Inhibition of Kdm6B. Theranostics 11, 6847–6859 (2021).
Chen, L. et al. Dual-targeted nanodiscs revealing the cross-talk between osteogenic differentiation of mesenchymal stem cells and macrophages. Acs Nano 17, 3153–3167 (2023).
Davis, F. M. et al. Inhibition of macrophage histone demethylase Jmjd3 protects against abdominal aortic aneurysms. J Exp. Med. 218, e20201839 (2021).
Patoli, D. et al. Inhibition of mitophagy drives macrophage activation and antibacterial defense during sepsis. J. Clin. Investig. 130, 5858–5874 (2020).
Chen, H. et al. Hydrogen alleviates cell damage and acute lung injury in sepsis via Pink1/parkin-mediated mitophagy. Inflamm. Res. 70, 915–930 (2021).
Li, G. et al. Pretreatment with Kahweol attenuates sepsis-induced acute lung injury via improving mitochondrial homeostasis in a Camkkii/Ampk-dependent pathway. Mol. Nutr. Food Res. 67, e2300083 (2023).
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837 (2007).
He, Z., Wang, H. & Yue, L. Endothelial progenitor cells-secreted extracellular vesicles containing Microrna-93-5P confer protection against sepsis-induced acute kidney injury via the Kdm6B/H3K27Me3/Tnf-Alpha axis. Exp. Cell. Res. 395, 112173 (2020).
Choi, S. H. et al. Intracellular Aibp (Apolipoprotein a-I binding Protein) regulates oxidized Ldl (low-density lipoprotein)-induced mitophagy in macrophages. Arterioscler. Thromb. Vasc. Biol. 41, e82–e96 (2021).
Funding
The authors declare that no funds, grants, or other support were received during the preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
XD conceived the study; HD conducted the experiments; SZ, XS and YG analyzed the data; HD was a major contributor in writing the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
This study was approved by the Laboratory Animals Ethics and Welfare Committee of MDKN Biotechnology Co., Lt. All methods were carried out in accordance with ARRIVE guidelines. All experiments were performed in accordance with relevant guidelines and regulations.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Deng, H., Zhu, S., Song, X. et al. KDM6B induces demethylation of H3K27me3 in MFN1 to modulate aberrant mitophagy in sepsis-induced acute lung injury. Sci Rep 15, 43339 (2025). https://doi.org/10.1038/s41598-025-27199-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-27199-z