Abstract
Building on previous studies, we identified mathematical patterns in HIV-1 and SARS-CoV-2 genomes using transfer learning and explainability with a pre-trained CNN on genomic spectrograms. These patterns seemed to define viral characteristics, leading us to hypothesize that inherent mathematical patterns in a virus’s genome determine its features. To explore this further, we focused on SARS-CoV-2 variant classification, designing a methodology with genomic spectrograms, a two-stage transfer learning approach, and two-step explainability. This approach identified genomic regions and nucleotide frequency patterns that characterize specific variants, revealing clear, distinguishable patterns for each category. The distinct and consistent total regions of high activation for each variant highlight the significance of the genomic region from the beginning of S gene to the end of 3’UTR in identifying the variants under study. The frequencies \(f = 1/9\) and particularly \(f = 1/3\) within this region appeared to play a key role in their identification. The shared prominence of \(f = 1/3\) in the final segment of the genome for both pre-VOC and Omicron (despite different pattern shapes) may hint at a phylogenetic connection in SARS-CoV-2 or even suggest that Omicron evolved from a pre-VOC lineage. The confirmation that mathematical patterns are associated with variant classification represents a step forward in demonstrating that these patterns play a role in viral characterization, suggesting the existence of an additional layer of genomic information that may enable virus characterization in a low-computing, and efficient manner compared to traditional methodologies.
Similar content being viewed by others
Introduction
Genome frequency-domain analysis offers a promising graphical approach for identifying biological patterns1,2,3. The application of signal processing tools to genomic sequences across different organisms has revealed significant relationships between nucleotide periodicity and various genomic features4. For example, it has provided insights into the connection between coding regions and sensitivity at frequency \(f = 1/3\)5, the localization of non-coding RNA molecules6, and the detection of GpC islands and micro-satellites7.
One of the advantages of these methods is that they bypass the need for Multiple Sequence Alignment (MSA), thus reducing computational costs8. Moreover, spectrogram imaging, which utilizes Fast Fourier Transform (FFT), is increasingly being applied in genomic analysis9, offering an efficient and scalable approach to understanding complex biological data.
Convolutional neural networks (CNNs) have several important advantages:
-
They have demonstrated solid robustness in pattern identification in images, which has remained over time despite the emergence of new architectures10.
-
They are suitable for large-scale data analysis in terms of computational efficiency11.
-
There are multiple pre-trained models that are highly powerful, with their performance extensively demonstrated in the existing literature12. These models enable robust implementation of transfer learning, a deep learning paradigm where a pre-trained model, having learned hierarchical feature representations from a source task, can be repurposed or fine-tuned for a related target task. Instead of constructing models from scratch for each application, transfer learning leverages these pre-existing feature hierarchies, optimizing convergence efficiency and mitigating data scarcity constraints. This approach significantly enhances generalization and accelerates model adaptation, especially in low-data regimes13.
-
They allow for reliable and visually understandable application of explainability tools14,15.
Gradient-weighted Class Activation Mapping (Grad-CAM) is a technique for detecting the most important areas of an image so that the deep learning architecture can classify it into a certain category. It uses gradients of the classification score with respect to the final convolutional feature map16,17. Grad-CAM is a highly visual tool, making it intuitive to interpret, though it may sacrifice some precision in pinpointing the exact regions of high activation18,19.
Applying a methodology based on transfer learning using a pre-trained CNN (specifically VGG-1620) with a dataset of genomic spectrogram images, and subsequently applying Grad-CAM as an explainability tool, we accurately identified the genomic regions where mathematical patterns related to the recombinant feature are located. This approach builds upon previous studies21,22, where we combined double transfer learning and a three-step explainability framework to detect such patterns in the genomic spectrogram of full-length severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) sequences, leveraging a CNN originally trained to identify the recombinant feature in human immunodeficiency virus type 1 (HIV-1).
In the case of HIV-1, we identified a key mathematical pattern associated with the recombinant feature, located at \(f = 1/3\) in 5’ and 3’ LTR, with nucleotides A and T being the most relevant in relation to this characteristic.
In the case of SARS-CoV-2, we detected a similar mathematical pattern, clearly located at \(f = 1/6\) in the S gene.
These findings revealed a clear connection between mathematical patterns related to nucleotide periodicity in the genome and the recombinant feature in viruses as phylogenetically distant as HIV-1 and SARS-CoV-2.
Based on the results obtained, we hypothesized that inherent mathematical patterns in a virus’s genome may determine viral characteristics.
Therefore, we aimed to verify the existence of mathematical patterns that determine the categorization of a complete SARS-CoV-2 sequence into one of its main variants.
Results
Optimal hyperparameters
The processing of 1,116,523 images in each of the two subsamples imposed significant computational demands, which were crucial in selecting the optimal hyperparameters. The hyperparameters chosen were:
-
Learning Rate = 0.0001
-
Batch Size = 52
-
Epochs = 1
When the Learning Rate was set to 0.0100 and/or the Batch Size to 128, the system did not have sufficient capacity to complete the required computations, causing training interruptions due to collapse. For the number of epochs, the minimum value was chosen.
Starting from the second epoch, the model reached 100% accuracy on both the training and validation sets. However, the validation loss exhibited a progressive increase, which revealed a mismatch between the accuracy metric and the probabilistic quality of the predictions. This phenomenon corresponded to an early stage of overfitting: the model memorized the available data patterns with excessive confidence without improving its generalization capacity, producing less calibrated probability distributions despite maintaining a perfect classification rate.
Figures 1a, b show the training curves for the two subsamplings, labeled VARIANT SUBSAMPLING 01 and VARIANT SUBSAMPLING 02, respectively. Both curves followed a similar pattern.
As shown, the training times were substantial. For Fig. 1a, training required 1190 min and 10 s–almost 20 h per epoch. For Fig. 1b, it took 1626 min and 16 s, equivalent to 27 h per epoch.
Notably, the training curves stabilized within roughly one-third of an epoch, with both datasets achieving a Validation Accuracy close to 99.5% (Fig.1a: 99.42%; Fig. 1b: 99.52%).
Overall, the graphs were similar both in shape and in Validation Accuracy values.
The use of just one epoch for training can be attributed to the large volume of data. With 669,914 sequences in the Training Set and 223,305 sequences in the Validation Set, the extensive dataset allowed for a thorough representation within just a single epoch.
Training curves for both subsamplings. The upper panel in each subfigure depicts Accuracy (%) for the Training (blue) and Validation (black) sets, and the lower panel illustrates Loss for the Training (orange) and Validation (black) sets.
Performance measurement ratios
Table 1 presents the performance metrics for the two VARIANT SUBSAMPLING 01 and 02 case.
“Test Accuracy” for each of the four categories is expressed as percentage of correctly classified sequences (Correct Predictions per Category / Total Number of Samples in that Category), as well as the “Test Accuracy Global” (Total Correct Predictions / Total Number of Samples).
The training process required a total of 19.84 hours, primarily due to the substantial computational demands of the large dataset and the limited hardware resources. Despite the extended training time, the accuracy values achieved for both the Validation and Test Sets were exemplary and nearly identical. The overall Test Accuracy was only 0.02% lower than the Validation Accuracy.
The performance measurement results for the VARIANT SUBSAMPLING 02 followed the same framework as in VARIANT SUBSAMPLING 01.
For VARIANT SUBSAMPLING 02, the training time amounted to 27.10 h, reflecting a 36.64% increase compared to the training time for VARIANT SUBSAMPLING 01. Nevertheless, both training sessions required significant computational time.
Regarding accuracy, the performance was outstanding across the board, with a slight improvement in the overall results compared to VARIANT SUBSAMPLING 01. However, this improvement was marginal and not significant.
In conclusion, the results from both subsamplings were excellent and closely aligned.
The impressive results observed could be attributed to the large number of samples in the dataset. The extensive dataset likely provided a comprehensive and varied representation of the phenomena, leading to stronger validation and more thorough pattern recognition. This may have allowed for near-optimal model training.
In both subsamplings, a slight decline in Test Accuracy was noted for the pre-VOC category. This could be explained by the fact that the pre-VOC category is not a distinct variant but rather encompasses all SARS-CoV-2 lineages existing prior to the emergence of Alpha, the first variant officially classified as a VOC.
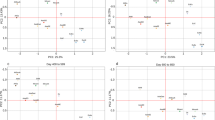
The confusion matrix results for the test set of VARIANT SUBSAMPLING 01 and VARIANT SUBSAMPLING 02 are presented in Fig. 2a, b, respectively.
The correct predictions are highlighted in blue on the grid, with deep blue indicating a high accuracy rate and light blue representing a medium to low accuracy. Misclassifications are marked in orange, with very light orange reflecting a low or very low error rate and intense orange indicating a medium to high error rate. Due to the high accuracy achieved, the blues are predominantly dark, while the oranges remain light.
Confusion matrices for the two subsamplings variants. 4\(\times\)4 confusion matrix for the classification problem with four classes. Rows correspond to the true classes and columns to the model’s predictions. Diagonal values indicate correct classifications (true positives), while off-diagonal values represent misclassifications.
In both subsamplings, the accuracy rates were outstanding across all categories, with classification errors being almost anecdotal.
These results were even more relevant considering that the complexity of this experiment increased due to two changes introduced in the second phase of Transfer Learning. Not only was the virus under investigation changed (from HIV-1 to SARS-CoV-2), but the detection target was also modified. Instead of classifying the genomic spectrograms as recombinant or non-recombinant, the task shifted to classifying the genomic spectrograms into four distinct variants.
Omicron exhibited the highest accuracy, followed by Delta and Alpha, while pre-VOC had the lowest accuracy rates, even though pre-VOC’s accuracy was still very high. A possible factor influencing these accuracy levels could be the sample size and uniformity within each variant. Omicron, being the most prevalent variant, had a larger and more consistent dataset, whereas Alpha had fewer complete genomes sequenced. Additionally, pre-VOC represents a collection of lineages prior to the emergence of Alpha, which may lead to greater variability compared to the more homogeneous Omicron.
Nevertheless, the accuracy ratios were very positive across all categories, with similar performance observed in both subsamplings. These results support that the CNN effectively detected distinguishing patterns within each of the four categories.
Total regions of high activation per variant
The term “region of high activation” refers to the areas the CNN focuses on when classifying a genomic spectrogram into one of the four SARS-CoV-2 variants. All total regions of high activation were computed by summing the scoremaps of the sequences in the Test Set.
As discussed previously, the high Test Accuracy for each category was the initial indicator that the CNN effectively learned the distinguishing patterns of each SARS-CoV-2 variant. By calculating the Grad-CAM total regions of high activation, we were able to pinpoint the specific genomic regions the CNN used to determine the variant to which each sequence belongs.
Total regions of high activation per variant (VARIANT_SUBSAMPLING_01): (a) pre-VOC, (b) Alpha, (c) Delta and (d) Omicron). The x-axis represents the nucleotide position. Below each Grad-CAM image, a scaled representation of the complete SARS-CoV-2 genome scheme is provided to locate each region of high activation within the genome itself. The y-axis represents the frequency range (nucleotide periodicity from 0 to 0.5 Hz). The frequencies of interest are indicated directly on the axis to facilitate the identification of regions of high activation. The z-axis is represented bidimensionally as a jet colormap, where the color scale transitions from blue (low activation) to red (high activation).
In the case of VARIANT SUBSAMPLING 01 (Fig. 3), the total regions of high activation for the pre-VOC compilation (Fig. 3a) were varied and distributed throughout the entire genomic spectrogram. The most prominent regions of high activation were observed in the low-frequency range, particularly from the latter part of Open Reading Frame 1ab (ORF1ab) to the 3’ Untranslated Region (3’UTR). However, the highest intensity was noted from the final quarter of ORF1ab extending toward the end of the S gene. Another significant regions of high activation was clearly located at \(f = 1/3\), spanning from the start of ORF3a to the 3’UTR. The most intense region, however, was found from the N gene to the end of the sequence (end of 3’UTR).
Alongside these primary regions, a secondary region of high activation was detected in the low-frequency range, extending roughly over the first half of ORF1ab, up to around nucleotide 10,200. The focal point of this zone was observed near nucleotide 5000.
Two secondary total regions of high activation were observed at \(f = 1/6\). The first encompassed the 5’UTR and an initial portion of ORF1ab, extending to roughly nucleotide 2500. The second region of high activation, also at \(f = 1/6\), was situated around the S gene region. In previous research, we identified a separate region of high activation in the S gene region that was associated with sequences classified as recombinant, also at \(f = 1/6\). The observed overlap between these regions of high activation may suggest that pre-VOC evolutionary branches could harbor previously undetected genetic recombination events. Further studies focusing on the exact location and characteristics of these sequences are warranted to investigate this hypothesis.
Another secondary region of high activation in the total regions of high activation corresponding to pre-VOC was identified in the high-frequency range (0.5 Hz) to \(f = 1/3\), around nucleotide 7500, within ORF1ab. The final secondary region of high activation also corresponded to the frequency range from \(f = 1/3\) to 0.5, covering the genomic region from S region to the end of the sequence.
The total regions of high activation for Alpha variant were more focused and precise (Fig. 3b). The primary region of high activation was clearly located at \(f = 1/3\), extending to 0.5 Hz, from the S gene to the end of the 3’UTR. The epicenter of this region of high activation stretched from N region to the end of 3’UTR. A secondary region of high activation was found at \(f = 1/9\), covering the region from approximately E region to the end of 3’UTR.
It was noteworthy that the primary region of high activation of Delta variant (Fig. 3c) exhibited similarities to the secondary region of high activation of Alpha, both in terms of location and geometric structure. This observation warrants further investigation, particularly to assess whether this overlap could be linked to shared evolutionary origins between the two variants. For Delta, the primary region of high activation was identified spanning from ORF3a to the end of the 3’UTR, covering a range from low frequencies to \(f = 1/9\). Another significant region of high activation was detected in the high-frequency range, from S gene to the end of the 3’UTR region. However, its epicenter appeared to be located in the 3’UTR, extending toward approximately M.
For Omicron (Fig. 3d), the primary region of high activation was distinct and well-defined, positioned at \(f = 1/3\), extending from S gene to the end of the 3’UTR. The epicenter of this region of high activation was predominantly found in the 3’UTR, with its influence extending toward ORF8. Additionally, a secondary region of high activation was detected in the low-frequency range, with its center in the 3’UTR, radiating toward approximately M.
The frequency \(f = 1/3\) in the 3’UTR region was a key factor in classifying sequences as pre-VOC, Alpha, and Omicron. However, for Delta, this frequency did not correspond to either a primary or secondary region of high activation. This presents an intriguing phenomenon that warrants further study. It would be valuable to identify the common characteristics shared by the Alpha, pre-VOC, and Omicron variants, and then analyze the differences within these shared features. The objective is to pinpoint the unique elements within these overlapping regions that guide the CNN’s classification toward a specific variant.
Delta was the only variant that did not show \(f = 1/3\) in the 3’UTR region, not even as a minor secondary zone. Being the only variant exhibiting this distinct behavior in the CNN, it would be valuable to further investigate the differences between the 3’UTR region in Delta and the other three variants to understand why this particular genomic region does not reveal a pattern that affects Delta and what sets it apart from the others.
Figure 4 presents the total regions of high activation for each category in the case of VARIANT SUBSAMPLING 02.
Total regions of high activation per variant (VARIANT SUBSAMPLING 02): (a) pre-VOC, (b) Alpha, (c) Delta and (d) Omicron). The x-axis represents the nucleotide position. Below each Grad-CAM image, a scaled representation of the complete SARS-CoV-2 genome scheme is provided to locate each region of high activation within the genome itself. The y-axis represents the frequency range (nucleotide periodicity from 0 to 0.5 Hz). The frequencies of interest are indicated directly on the axis to facilitate the identification of regions of high activation. The z-axis is represented bidimensionally as a jet colormap, where the color scale transitions from blue (low activation) to red (high activation).
For the pre-VOC variant (Fig. 4a), there was a single prominent and intense primary region of high activation in the low-frequency range, whose epicenter was distinctly located around the S gene, extending toward nucleotide 17,500 (ORF1ab) on one side and to the end of the 3’UTR on the other.
The remaining zones were secondary, aligning exactly with those identified in VARIANT SUBSAMPLING 01, but with significantly lower intensity. The first secondary region of high activation was positioned at \(f = 1/3\), spanning from the start of ORF3a to the 3’UTR. Another secondary region of high activation, in the low-frequency range, covered approximately the first half of ORF1ab, reaching about nucleotide 10,200.
At \(f = 1/6\), two secondary total regions of high activation were identified. The first of these was located near the S gene, similarly to what was observed in VARIANT SUBSAMPLING 01, a secondary total region of high activation was also found at \(f = 1/6\) in the S region. As previously reported, this is a characteristic feature of recombinant SARS-CoV-2 sequences22. This phenomenon appeared in the pre-VOC category across both subsamplings, highlighting the importance of further investigation into this occurrence and exploring the hypothesis that hidden recombination events might exist within certain pre-VOC lineages.
The second secondary region of high activation at \(f = 1/6\) was faintly visible and spanned the 5’UTR and the initial segment of ORF1ab, reaching up to approximately nucleotide 2500.
Another subtle secondary region of high activation in the pre-VOC category was observed in the high-frequency range (0.5 Hz) to \(f = 1/3\), around nucleotide 7,500 in ORF1ab. The final secondary region of high activation also corresponded to the frequency range from \(f = 1/3\) to 0.5, covering the genomic region from S region to the end of the sequence.
In the case of Alpha (Fig. 4b), the most prominent region of high activation had its epicenter at \(f = 1/3\) in 3’UTR, extending to N. It radiated towards S gene and reached into the high-frequency region.
Another significant region of high activation for Alpha was found around \(f = 1/9\), stretching from approximately ORF3a to the end of 3’UTR.
For Delta (Fig. 4c), the primary region of high activation was located in the low-frequency range up to \(f = 1/9\), spanning from S region to the end of 3’UTR. The epicenter, however, was in 3’UTR, with the highest intensity reaching up to ORF3a.
A secondary region of high activation appeared in the high-frequency range, extending from 3’UTR to ORF3a, with its epicenter also in 3’UTR.
In Omicron’s case (Fig. 4d), the main region of high activation was sharply defined and positioned at \(f = 1/3\), covering the region from S region to the end of 3’UTR. The epicenter of this region of high activation was clearly located in 3’UTR, with its influence radiating up to ORF8.
A secondary zone in the low-frequency range was identified, with its epicenter located in 3’UTR and extending up to M or possibly E genes.
As observed in VARIANT SUBSAMPLING 01, Delta was the only variant where \(f = 1/3\) in 3’UTR showed no significant relevance for classification.
Discussion
To facilitate a more direct comparison between the total region of high activation results from VARIANT SUBSAMPLING 01 and VARIANT SUBSAMPLING 02, Table 2 presents a summary of the findings for both subsamplings, categorized accordingly.
The most striking observation was the difference in the total regions of high activation for the pre-VOC category across the two subsamplings. In VARIANT SUBSAMPLING 01, the zones were more diffuse and extended, whereas in VARIANT SUBSAMPLING 02, the primary total region of high activation was more concentrated and better defined in the low-frequency range around the S gene. Although the other zones in VARIANT SUBSAMPLING 02 were located roughly in the same regions and frequencies as in VARIANT SUBSAMPLING 01, their power was considerably diminished.
The pre-VOC category emerged as the least similar among the four analyzed. Investigating the relationship between the sequences in both subsamplings would be valuable to determine whether there is a connection between these sequences and the differences observed between the two subsamplings. It would also be insightful to explore the underlying causes that lead to the sharpness observed in VARIANT SUBSAMPLING 02 and the variability present in VARIANT SUBSAMPLING 01.
In contrast, homology between the Alpha, Delta, and especially Omicron variants was greater between both subsamplings, showing a high degree of similarity and consistency across the two sets.
For Alpha, the main region of high activation around \(f = 1/3\) in the final genomic region was highly comparable between subsamplings. However, in VARIANT SUBSAMPLING 02, the expansion from \(f = 1/3\) to \(f = 1/2\) (high frequencies) displayed more power.
Furthermore, the secondary region found at \(f = 1/9\) in the final genomic stretch in VARIANT SUBSAMPLING 01 exhibited more power in VARIANT SUBSAMPLING 02.
We observed an opposite trend in the case of Delta. In this category, VARIANT SUBSAMPLING 01 showed slightly more power in the very secondary region of high activation of VARIANT SUBSAMPLING 02, located in the high frequencies towards the final genomic stretch. The main region of high activation, however, was extremely sharp, well-centered, and highly similar between both subsamplings. It was consistently located in the range from low frequencies up to \(f = 1/9\), spanning from the end of S region to the end of 3’UTR.
Omicron, on the other hand, exhibited the highest degree of homology between both subsamplings. The total regions of high activation were almost identical. The primary region of high activation, located at \(f = 1/3\) in the final genomic region, was nearly identical in both subsamplings, though it might have been slightly more expanded towards S gene in VARIANT SUBSAMPLING 01 compared to VARIANT SUBSAMPLING 02. The secondary zone, situated in the low frequency range in the lower right corner of the genomic spectrogram, was virtually identical in both subsamplings, with only a barely noticeable increase in power in VARIANT SUBSAMPLING 02.
After quantifying the high accuracy achieved by the CNN in both subsamplings and calculating the total regions of high activation, it became clear that the high degree of similarity in the total regions of high activation across both subsamplings (particularly in Alpha, Delta, and Omicron, compared to pre-VOC) suggested that the CNN’s performance was robust and consistent in identifying the intrinsic and distinguishing patterns of each of the four categories analyzed.
The results concerning the total regions of high activation for the pre-VOC category displayed two key distinctions when compared to the other variants. First, the regions were more varied and less concentrated on specific areas. Second, the regions between the two subsamplings showed notable differences, not necessarily in their positioning but in their intensity. This variability might be related to the fact that the pre-VOC category is not a distinct variant in itself but rather a compilation of sequences prior to the emergence of Alpha, the first VOC. At first sight, this observation could appear to contradict previous findings reporting that, during the period from the onset of the pandemic to the appearance of Alpha, the genetic variability of SARS-CoV-2 was limited23. However, these two perspectives are not mutually exclusive. The apparent diversity in the Grad-CAM regions of high activation may reflect the heterogeneity inherent in grouping multiple early lineages together under the “pre-VOC” label, as well as noise introduced by uneven sampling. In fact, some studies have identified the co-existence of at least two distinct lineages in Wuhan during the early stages of the virus’s spread24, which supports the idea that small but meaningful differences across sequences could still result in perceptible variation in the regions of high activation.
Interestingly, the pre-VOC category seemed to include a characteristic region of high activation associated with the Omicron variant. This observation coincides with the fact that the Omicron variant (B.1.1.529) of SARS-CoV-2 did not directly evolve from the Alpha (B.1.1.7) or Delta (B.1.617.2) variants, according to multiple studies. Instead, it is classified as part of a separate evolutionary lineage with distinct genetic traits. Several theories about its origin have been proposed, with three standing out as particularly plausible25,26,27.
First, Omicron may have circulated and evolved within a concealed population. One possibility is that Omicron, or its ancestral form, developed in a remote region where nucleic acid testing is infrequent. It is even possible that early traces of Omicron’s ancestor were detected in South Africa through nucleic acid tests28.
Second, Omicron may have emerged from a prolonged “cat-and-mouse” interaction between the virus and the immune system in certain immunocompromised individuals, such as AIDS patients infected with SARS-CoV-229.
Third, it is a plausible hypothesis that Omicron has emerged through adaptation within animal reservoirs, potentially rodents, before being transmitted back to humans30.
Further investigation is required to clarify the potential causes behind the observed variability in the mathematical signatures linked to the pre-VOC category, as well as the possible evolutionary connections between pre-VOC and Omicron.
Nevertheless, it is remarkable that distinct and clearly identifiable mathematical patterns exist for each of the four categories analyzed.
The repeated detection of frequency patterns related to \(f = 1/3\) and its multiples indicates that the mathematical signatures associated with the variant to which the sequence belongs are encoded every 3 nucleotides (or multiples of 3). The unequivocal identification of these mathematical signatures could confirm a new layer of genome reading based on nucleotide periodicity, in line with previous studies in this regard31.
This would enable the characterization of a virus in a low-computing and faster manner compared to traditional methods, which include multiple alignments with manual adjustments or protein sequence alignments.
Conclusions
Training the model for just one epoch delivered promising results across all performance metrics, with particularly high Test Accuracy in both subsamplings. The substantial size of the dataset, comprising 1,116,523 images in total for each subsampling, provided the CNN with a large number of training examples, which likely enabled it to effectively learn the patterns in the data even with a single epoch.
In this experiment, we implemented a dual modification to the two-stage transfer learning approach. We applied a CNN trained to detect genetic recombination in HIV-1 to identify variants in SARS-CoV-2, shifting not only to a new virus but also a different focus of study. This double shift resulted in excellent and robust outcomes, with Test Accuracies of 99.40% in VARIANT SUBSAMPLING 01 and 99.48% in VARIANT SUBSAMPLING 02. The Test Accuracies per category ranged from 97.73% to 99.88%.
The total regions of high activation for each category displayed clear, robust, and highly distinguishable patterns, with pre-VOC showing high homology and Alpha, Delta, and Omicron exhibiting very high homology in both subsamplings.
For pre-VOC, the differences between subsamplings were related to a decrease in power in VARIANT SUBSAMPLING 02 compared to VARIANT SUBSAMPLING 01, though the physical location of the total regions of high activation remained consistent.
The Omicron variant exhibited nearly identical total regions of high activation in both subsamplings.
It is highly interesting to examine the links between the phylogenetic relationships and evolutionary variations of the pre-VOC, Alpha, Delta, and Omicron variants, as well as the mathematical signatures that influence the CNN’s classification of these variants.
Although each variant’s total regions of high activation are clearly distinct, certain similarities open intriguing possibilities for exploring phylogenetic relationships. For example, the shared prominence of \(f = 1/3\) in the final genome segment for both pre-VOC and Omicron (despite different pattern shapes) may suggest a possible phylogenetic connection, but this remains a hypothesis based on observational data. Further research is required to test this idea.
The distinct and consistent total regions of high activation for each variant highlight the significance of the genomic region from the beginning of S gene to the end of 3’UTR in identifying the variants under study. The frequencies \(f = 1/9\) and particularly \(f = 1/3\) within this region seemed to play a key role in their identification. Further research is needed to explore the precise relationship between the nucleotide periodicities corresponding to these frequencies and the variant to which they belong.
Compared to traditional methods that involve multiple alignments and manual adjustments, the designed methodology is lower-computing and enables faster large-scale characterization.
Similar to the process of identifying recombinant sequences21,22, this study demonstrates a mathematical signature linked to the pre-VOC, Alpha, Delta, and Omicron variants. Consequently, it is plausible that other mathematical signatures related to different viral traits may also exist.
The results of our research provide compelling evidence of a new layer of genome reading based on mathematical signatures related to nucleotide periodicity, demonstrating their existence with high accuracy rates and shorter response times compared to traditional methods.
Limitations
In this study, we focused on the most abundant SARS-CoV-2 variants and excluded those with a very limited number of sequences. This decision allowed us to train models with sufficient information per class and achieve robust performance; however, it also limits the generalization of our results to rare or underrepresented variants. For the Omicron variant, we adjusted the number of genomic spectrograms through random subsampling, with the aim of efficiently leveraging the available data for model training while maintaining biological plausibility.
Future work
In light of the results obtained and from a computation-focused perspective, an important line of future research is to rigorously evaluate the contribution of the two-stage transfer learning approach. While pretraining the CNN on HIV-1 genetic recombination detection was designed to capture generalizable patterns in genomic spectrograms, HIV-1 recombination and SARS-CoV-2 variant classification are biologically unrelated. To assess the actual benefit of this pretraining, ablation experiments could be conducted comparing (i) a CNN trained directly on SARS-CoV-2 data, (ii) transfer learning from ImageNet only, and (iii) the proposed two-stage transfer learning. These studies would help determine whether pretraining on unrelated genomic data provides measurable improvements in model performance.
One of our future research directions involves seeking complementarities with other already successful methods, with the aim of moving toward the exact mathematical formulation of the patterns identified in this article. Of particular interest are DNA-based identification tools such as varKoder32 and varKoding33, as well as other genomic sequence-to-image conversions, especially the widely used Frequency Chaos Game Representation34,35,36,37,38,39,40.
Although it would be possible to apply synthetic oversampling techniques to address class imbalance, such as SMOTE41 and its variants (Borderline-SMOTE42 or ADASYN43), or weighted loss functions, we are concerned that generating synthetic examples for minority classes could compromise biological interpretability, given the current limited knowledge about genome structure. For this reason, in the present study we chose to remove very rare variants and adjust the number of spectrograms for the most abundant classes through subsampling, while ensuring the use of as much data as possible for model training. Future work could explore the incorporation of these techniques, carefully assessing their impact on the biological validity of the results.
We also consider it essential to develop methods, techniques, and/or new architectures that effectively enable the inclusion of minority variants in the learning process, with the aim of extending classification to a broader and more representative set of SARS-CoV-2 variants. This would allow for a more refined identification of variants, enhancing the applicability of the model in biosanitary settings.
Furthermore, validation across temporal and geographic dimensions constitutes an important line of our future research. These analyses are particularly relevant for establishing phylogenetic relationships complementary to our results, with greater clarity/precision in identifying such relationships and additional supporting information. Future work could explore more robust temporal and geographic holdout experiments, as well as training with sequences from early pandemic phases or specific regions and testing with later or geographically distinct samples, in order to evaluate the model’s ability to generalize to emerging variants over time and space.
Finally, although nucleotides encode in triplets (codons) that ultimately correspond to amino acids, the present analysis does not aim to establish direct links between the highlighted regions and specific effects at the protein level. Future work could explore these potential functional implications by investigating how the observed genomic patterns relate to protein structure and function, mutational hotspots, conserved motifs, or other genomic factors.
Methods
Equipment
All experiments were run in this equipment:
-
Processing Unit: Intel(R) Core(TM) i7-4770K CPU. 3.5 GHz.
-
Installed RAM: 32 GB usable.
-
Operative System: Windows 10 Education. Version: 22H2.
-
GPU: NVIDIA GeForce RTX 3090. Total memory: 40 GB.
Dataset of SARS–CoV-2 variants
The complete collection of 1,539,728 SARS-CoV-2 sequences, compiled from the NCBI Virus Database44 around March 2023, includes the number of sequences per variant listed in Table 3. Variants were organized according to their estimated emergence date, using data from the GISAID Initiative45. The ’Variant’ column represents the WHO-designated name for each SARS-CoV-2 variant46.
Notably, only pre-VOC, Alpha, Delta, and Omicron variants contributed each more than 10% of the total. None of the other variants individually accounted for more than 1.3% of the total complete sequences, most falling below 1%.
This compilation reveals a significant number disparity between variants that did not persist, such as Beta, Gamma, Epsilon, Eta, Iota, Kappa, Lambda, Mu, Theta, and Zeta; and those that dominated, including pre-VOC, Alpha, Delta, and Omicron. The latter four exhibited evolutionary advantages that enabled them to outcompete the less successful variants (Table 4).
Omicron accounts for nearly half of the dataset, followed by Delta, while Alpha and pre-VOC (compendium of all lineages before the appearance of Alpha) have similar sequence counts. Minority variants (Beta, Gamma, Epsilon, Eta, Iota, Kappa, Lambda, Mu, Theta, and Zeta), total fewer than 50,000 sequences (3.18%). This group also exhibits high variability due to the diversity among its 10 variants.
Data imbalance can hinder Deep Learning models by causing biases toward majority classes, leading to several issues: - Overfitting to majority classes: the model may generalize poorly to minority classes due to insufficient data.
- Decreased accuracy: high overall accuracy may be misleading, as performance on minority classes remains low.
- Sensitivity to decision thresholds: imbalance can distort classification thresholds, affecting predictions.
- Difficulty in identifying patterns: limited data for minority classes makes it harder to learn distinguishing features.
To reduce noise from minority classes, unrepresentative variants were removed, as shown in Table 5. This adjustment redistributes the dataset, with Omicron comprising 50%, Delta 20%, and pre-VOC and Alpha under 15%.
Omicron is clearly the most widespread variant. Initially detected in Botswana in November 202147, it quickly became the dominant circulating variant. As of September 2022, it accounts for 100% of sequenced cases48, effectively eliminating all other variants from the epidemic phase.
The Omicron dataset is more than twice the size of Delta’s and 3.5 times larger than pre-VOC and Alpha. To address this imbalance, we split Omicron into two equally sized random subsamples, ensuring no overlap between them. Each subsample included all samples from Alpha, Delta, and pre-VOC, plus half of Omicron, creating a more balanced dataset as shown in Table 6. This approach helped reduce bias and improved generalization.
Although Delta and Omicron still predominated over Alpha and pre-VOC, the large number of samples across all categories provided enough data for effective model learning, minimizing the impact of sample variability.
For dataset organization, 60% of the total sequences were allocated to the Training Set, while the Validation Set and Test Set each accounted for 20%.
The composition of the dataset for each of the two subsamples is shown in Table 7.
Spectrogram generation
The initial step in generating the spectrogram of a sequence involved converting the sequence into four digital signals (\(U_a\), \(U_g\), \(U_c\), \(U_t\)), each corresponding to the presence of a specific nucleotide (A, G, C, T) at every position.
The following step involved calculating the spectrogram for each of the four binary signals derived from this decomposition.
The FFT was applied to each signal, which is an efficient computational method for performing the Discrete Fourier Transform (DFT)49.
Where:
FFT algorithms break down an N-point DFT into smaller DFTs50 by utilizing sliding windows.
Calculating the spectrogram of a genomic sequence provides a graphical representation of the importance of each periodicity (frequency) at every position within the genome, treating position as analogous to time.
The x-axis represents the position of each nucleotide, the y-axis indicates the frequency range, so the range of the x-axis corresponds to the full genome length of SARS-CoV-2. The maximum value of the y-axis range (frequency) corresponds to 0.5 Hz. The z-axis shows the FFT value. The spectrogram is a two-dimensional representation created by replacing the z-axis with a color palette that corresponds to the amplitude of the FFT at each nucleotide position (x-axis) for a specific frequency (y-axis)51.
In this research, we used the representation called Superposed Spectrogram. In this representation, the z-axis is the arithmetic sum of the values along the z-axis for each of the four nucleotide types:
This spectrogram representation helps to more clearly define the influential regions of the genome.
For two-dimensional spectrograms, the z-axis is represented by a “jet” color palette (Fig. 5). Blue color corresponds to lowest sensitivities and red to the highest ones52.
Using the matplotlib.pyplot.pcolormesh function, we assigned the minimum FFT value to deep blue (0) and the maximum value to deep red (1), with intermediate values automatically mapped to corresponding colors. By default, the data range is scaled linearly to the colorbar range.
Jet color map. Commonly employed color scale in scientific visualization, encoding low values in dark blue, progressing through cyan, green, and yellow, and representing high values in bright red.
Each spectrogram is labeled with the reference of its corresponding sequence.
We generated spectrograms for both datasets using Python and the Scipy library’s scipy.signal.spectrogram function. To prevent biases and ensure proper CNN training, we removed axes, margins, and any other elements that could interfere with the network’s performance. As a result, the image focuses solely on the spectrogram53,54.
The parameters used to generate the genomic spectrograms were set to their default values55.
Two-stage transfer learning
We conducted a two-stage transfer learning using MATLAB 2021b App Deep Learning Designer.
We used the HIV-1 case study as the first Transfer Learning phase for genetic recombination21. Figure 6 illustrates the Transfer Learning methodology applied for the classification of SARS-CoV-2 variants.
A pre-trained VGG-16 model was used with the ImageNET dataset. In Phase 1, we applied Transfer Learning to the genomic spectrogram dataset of complete HIV-1 sequences to detect the recombinant feature. In Phase 2, we applied Transfer Learning to the resulting network from Step 1 (VGG-16 HIV-1 GENETIC RECOMBINATION) using a genomic spectrogram dataset of complete SARS-CoV-2 sequences to detect the variant to which each SARS-CoV-2 sequence belongs (VGG-16 SARS-CoV-2 VARIANTS).
Two-stage transfer learning methodology variant detection. The first stage trains a pre-trained VGG-16 network, initially trained on ImageNet, to detect patterns in genomic spectrograms, in this case classifying complete HIV-1 sequences as recombinant or non-recombinant. In the second stage, this specialized VGG-16 is retrained, this time to recognize patterns related to the variant to which a complete SARS-CoV-2 sequence belongs.
As shown in Fig. 6, the first phase of Transfer Learning remained the same. However, in the second phase, the goal shifted from detecting the recombinant feature in SARS-CoV-2 to classifying complete SARS-CoV-2 sequences into one of the following four variants: pre-VOC, Alpha, Delta, or Omicron.
Performance metrics
The performance metrics used in this study were:
- Validation accuracy: this metric was used to assess the model’s performance during the validation phase, helping to adjust hyperparameters and optimize performance during training.
- Training time: this metric measured the computational cost associated with training the model.
- Confusion matrix: a 4x4 confusion matrix was used for this classification problem with four distinct classes. Each row represented the true class of instances from the dataset, while each column represented the predicted class. Diagonal elements indicated true positives (correctly classified instances), while off-diagonal elements showed classification errors.
- Test accuracy: this metric was used to evaluate the model’s real-world performance on previously unseen data. It was computed both for each individual class and overall.
Explainability results generation
We used Grad-CAM (grad-CAM MATLAB function) to pinpoint the critical regions of high activation for classifying a SARS-CoV-2 sequence into its respective variant. In addition, image processing techniques were employed to calculate the total regions of high activation for each variant.
The explainability score maps were processed in two steps. A schematic summary of the explainability processing methodology is shown in Fig. 7.
The first step consisted of 223,305 images, corresponding to the total number of data points in the Test Set. The second step involved eight images, with four categories divided into two subsamples.
Two-step explainability. In the first step, individual Grad-CAM activation maps are computed for each sequence. Once calculated, in the second step, the activation maps for each category are summed, obtaining the total activation map and, consequently, the regions of high activation for the four analyzed categories.
In the first step, we calculated the score maps for each sequence from the four categories in the Test Set and generated the corresponding Grad-CAM images using a jetmap color scale (Fig. 5). This process was repeated for each of the two subsamples.
The second step of the explainability process involved computing the total score maps for each of the four categories. Each total score map was obtained by summing the individual score maps for each category (matrix summation).
After obtaining the total score map matrix, we generated the Grad-CAM image depicting the overall regions of high activation using the jet map color scale.
This process was applied to both subsamples, resulting in eight images, each representing the four categories across the two generated subsamplings.
With the aim of facilitating the localization of relevant genomic areas throughout the entire SARS-CoV-2 genome, we drew a to-scale representation of it (Fig. 8).
To-scale representation of the complete SARS-CoV-2 genome. Schematic scaled representation of the complete SARS-CoV-2 genome, including a zoom of its final fragment, also scaled for better visualization of the genomic elements in that region.
Data availability
The datasets used and/or analysed during the current study are available from the corresponding author on reasonable request.
Code availability
The codes developed and used in this study are provided as a supplementary document to this article. The datasets and pre-trained models generated and/or analyzed during the current study are available from the corresponding author upon reasonable request.
References
Liew, A. W. C., Yan, H. & Yang, M. Pattern recognition techniques for the emerging field of bioinformatics: A review. Pattern Recognit. 38, 2055–2073 (2005).
Oueslati, A. E., Ellouze, N. & Lachiri, Z. 3D spectrum analysis of DNA sequence: Application to Caenorhabditis elegans genome. In 2007 IEEE 7th International Symposium on BioInformatics and BioEngineering. 864–871 (IEEE, 2007).
Dimitrova, N., Cheung, Y. H. & Zhang, M. Analysis and visualization of DNA spectrograms: Open possibilities for the genome research. In Proceedings of the 14th ACM International Conference on Multimedia. 1017–1024 (2006).
Bucur, A., Van Leeuwen, J., Dimitrova, N. & Mittal, C. Alignment method for spectrograms of DNA sequences. IEEE Trans. Inf. Technol. Biomed. 14, 3–9 (2009).
Tiwari, S., Ramachandran, S., Bhattacharya, A., Bhattacharya, S. & Ramaswamy, R. Prediction of probable genes by Fourier analysis of genomic sequences. Bioinformatics 13, 263–270 (1997).
Vaidyanathan, P. & Yoon, B. J. The role of signal-processing concepts in genomics and proteomics. J. Franklin Inst. 341, 111–135 (2004).
Sussillo, D., Kundaje, A. & Anastassiou, D. Spectrogram analysis of genomes. EURASIP J. Adv. Signal Process. 2004, 1–14 (2004).
Kubicova, V. & Provaznik, I. Use of whole genome DNA spectrograms in bacterial classification. Comput. Biol. Med. 69, 298–307 (2016).
Morales, J. A. et al. Deep learning for the classification of genomic signals. Math. Probl. Eng. 2020 (2020).
Alzubaidi, L. et al. Review of deep learning: Concepts, cnn architectures, challenges, applications, future directions. J. Big Data 8, 1–74 (2021).
Kim, H., Nam, H., Jung, W. & Lee, J. Performance analysis of cnn frameworks for gpus. In 2017 IEEE International Symposium on Performance Analysis of Systems and Software (ISPASS). 55–64 (IEEE, 2017).
Stančić, A., Vyroubal, V. & Slijepčević, V. Classification efficiency of pre-trained deep cnn models on camera trap images. J. Imaging 8, 20 (2022).
Gupta, J., Pathak, S. & Kumar, G. Deep learning (cnn) and transfer learning: A review. J. Phys. Conf. Ser. 2273, 012029 (IOP Publishing, 2022).
Zhang, Q.-S. & Zhu, S.-C. Visual interpretability for deep learning: A survey. Front. Inf. Technol. Electron. Eng. 19, 27–39 (2018).
Chakraborty, S. et al. Interpretability of deep learning models: A survey of results. In 2017 IEEE Smartworld, Ubiquitous Intelligence & Computing, Advanced & Trusted Computed, Scalable Computing & Communications, Cloud & Big Data Computing, Internet of People and Smart City Innovation (Smartworld/SCALCOM/UIC/ATC/CBDcom/IOP/SCI). 1–6 (IEEE, 2017).
Selvaraju, R. R. et al. Grad-cam: Visual explanations from deep networks via gradient-based localization. In Proceedings of the IEEE International Conference on Computer Vision. 618–626 (2017).
MathWorks. Grad-CAM Explains Why (2024). Accessed 22 Oct 2021.
Hamilton, N. et al. Enhancing visualization and explainability of computer vision models with Local Interpretable Model-agnostic Explanations (LIME). In 2022 IEEE Symposium Series on Computational Intelligence (SSCI). 604–611 (IEEE, 2022).
Wargnier-Dauchelle, V., Grenier, T., Durand-Dubief, F., Cotton, F. & Sdika, M. A weakly supervised Gradient Attribution constraint for interpretable classification and anomaly detection. In IEEE Transactions on Medical Imaging (2023).
Gupta, V. & Patel, R. Lungs disease classification using VGG-16 architecture with PCA. In 2023 International Conference on Advancement in Computation & Computer Technologies (InCACCT). 495–500 (IEEE, 2023).
Guerrero-Tamayo, A. et al. Discovering mathematical patterns behind HIV-1 genetic recombination: a new methodology to identify viral features. In IEEE Access (2023).
Guerrero-Tamayo, A. et al. Classification of SARS-COV-2 sequences as recombinants via a pre-trained cnn and identification of a mathematical signature relative to recombinant feature at spike, via interpretability. PLoS ONE (2024).
Markov, P. V. et al. The evolution of SARS-CoV-2. Nat. Rev. Microbiol. 21, 361–379 (2023).
Pagani, I., Ghezzi, S., Alberti, S., Poli, G. & Vicenzi, E. Origin and evolution of SARS-COV-2. Eur. Phys. J. Plus 138, 157 (2023).
Dhama, K. et al. Global emerging omicron variant of SARS-COV-2: Impacts, challenges and strategies. J. Infect. Public Health 16, 4–14 (2023).
Rahmani, S. & Rezaei, N. Omicron (b. 1.1. 529) variant: Development, dissemination, and dominance. J. Med. Virol. 94, 1787 (2022).
Gili, R. & Burioni, R. SARS-COV-2 before and after omicron: Two different viruses and two different diseases?. J. Transl. Med. 21, 251 (2023).
Du, P., Gao, G. F. & Wang, Q. The mysterious origins of the omicron variant of SARS-COV-2. Innovation 3 (2022).
Wei, C. et al. Evidence for a mouse origin of the SARS-COV-2 omicron variant. J. Genet. Genomics 48, 1111–1121 (2021).
Sun, Y., Lin, W., Dong, W. & Xu, J. Origin and evolutionary analysis of the SARS-COV-2 omicron variant. J. Biosaf. Biosecur. 4, 33–37 (2022).
Guerrero-Tamayo, A. et al. The third codon nucleotide’s role in genetic recombination within SARS-COV-2 spike protein: A pilot study. In International Conference on Hybrid Artificial Intelligence Systems. 29–40 (Springer, 2024).
Asprino, R. C. et al. A curated benchmark dataset for molecular identification based on genome skimming. Sci. Data 12, 906. https://doi.org/10.1038/s41597-025-05230-2 (2025) (publisher: Nature Publishing Group).
de Medeiros, B. A. S. et al. A composite universal DNA signature for the tree of life. Nat. Ecol. Evolut. 1–15. https://doi.org/10.1038/s41559-025-02752-1 (2025) (publisher: Nature Publishing Group).
Avila Cartes, J., Anand, S., Ciccolella, S., Bonizzoni, P. & Della Vedova, G. Accurate and fast clade assignment via deep learning and frequency chaos game representation. GigaScience 12, giac119. https://doi.org/10.1093/gigascience/giac119 (2022).
Hoang, T., Yin, C. & Yau, S. S. T. Numerical encoding of DNA sequences by chaos game representation with application in similarity comparison. Genomics 108, 134–142. https://doi.org/10.1016/j.ygeno.2016.08.002 (2016).
Deschavanne, P. J., Giron, A., Vilain, J., Fagot, G. & Fertil, B. Genomic signature: Characterization and classification of species assessed by chaos game representation of sequences. Mol. Biol. Evolut. 16, 1391–1399. https://doi.org/10.1093/oxfordjournals.molbev.a026048 (1999).
Arias, P. M., Alipour, F., Hill, K. A. & Kari, L. DeLUCS: Deep learning for (unsupervised) clustering of DNA sequences. bioRxiv 2021.05.13.444008. https://doi.org/10.1101/2021.05.13.444008 (2021) (preprint posted by Cold Spring Harbor Laboratory under a CC BY 4.0 license).
Sengupta, D. C., Hill, M. D., Benton, K. R. & Banerjee, H. N. Similarity studies of corona viruses through chaos game representation. Comput. Mol. Biosci. 10, 61–72. https://doi.org/10.4236/cmb.2020.103004 (2020).
Löchel, H. F. & Heider, D. Chaos game representation and its applications in bioinformatics. Comput. Struct. Biotechnol. J. 19, 6263–6271. https://doi.org/10.1016/j.csbj.2021.11.008 (2021).
He, H., Kari, L. & Arias, P. M. Bridging Chaos Game Representations and \$k\$-mer Frequencies of DNA Sequences. https://doi.org/10.48550/arXiv.2506.22172 (2025). arXiv:2506.22172 [cs].
Chawla, N. V., Bowyer, K. W., Hall, L. O. & Kegelmeyer, W. P. Smote: Synthetic minority over-sampling technique. J. Artif. Intell. Res. 16, 321–357 (2002).
Han, H., Wang, W.-Y. & Mao, B.-H. Borderline-smote: A new over-sampling method in imbalanced data sets learning. In International Conference on Intelligent Computing. 878–887 (Springer, 2005).
He, H., Bai, Y., Garcia, E. A. & Li, S. Adasyn: Adaptive synthetic sampling approach for imbalanced learning. In 2008 IEEE International Joint Conference on Neural Networks (IEEE World Congress on Computational Intelligence). 1322–1328 (IEEE, 2008).
National Library of Medicine (US), N. C. F. B. I. NCBI Virus. https://www.ncbi.nlm.nih.gov/labs/virus/vssi/ (2023). Accessed 11 Sep 2023.
GISAID Initiative. GISAID-Tracking of hCoV-19 Variants. https://gisaid.org/hcov19-variants/ (2023). Accessed 21 Nov 2023.
World Health Organization. WHO COVID-19 Dashboard. https://covid19.who.int/ (2020). Accessed 11 Sep 2023.
Vitiello, A., Ferrara, F., Auti, A. M., Di Domenico, M. & Boccellino, M. Advances in the omicron variant development. J. Intern. Med. 292, 81–90 (2022).
Beesley, L. J. et al. SARS-COV-2 variant transition dynamics are associated with vaccination rates, number of co-circulating variants, and convalescent immunity. EBioMedicine 91 (2023).
Nussbaumer, H. J. The fast Fourier transform. In Fast Fourier Transform and Convolution Algorithms. 80–111 (Springer, 1981).
Proakis, J. G. & Manolakis, D. G. Introduction to Digital Signal Processing (Prentice Hall Professional Technical Reference, 1988).
Santo, E. & Dimitrova, N. Improvement of spectral analysis as a genomic analysis tool. In 2007 IEEE International Workshop on Genomic Signal Processing and Statistics. 1–4 (IEEE, 2007).
Contributors, M. Matplotlib: Visualization with Python. (2024). Accessed 22 March 2024.
Catalá, O. D. T. et al. Bias analysis on public X-ray image datasets of pneumonia and COVID-19 patients. IEEE Access 9, 42370–42383 (2021).
Chen, L. et al. Hidden bias in the DUD-E dataset leads to misleading performance of deep learning in structure-based virtual screening. PloS One 14, e0220113 (2019).
Contributors, S. Documentation of spectrogram function of the Scipy library in python environment. (2024). Accessed 22 March 2024.
Acknowledgements
This work was supported by the Research Training Grants Program - University of Deusto. Ref. FPI UD_2021_10. The funders had no role in the conceptualization, design, data collection, analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
A.G-T. designed the methodology, generated the genomic spectrograms, prepared the datasets, conducted the experiments, contributed to the interpretation of results, and wrote this work. B.S.U. contributed to the design of the methodology, the datasets, and the experimentation, and substantially reviewed the work. M-D.M.T. contributed to the design of the methodology, the interpretation of results, and substantially reviewed the work. I.O. contributed to the design of the methodology, the interpretation of results, and substantially reviewed the work. C.C. contributed to the design of the methodology, the interpretation of results, and substantially reviewed the work. I.P-L. contributed to the design of the methodology, the datasets, and the experimentation, and substantially reviewed the work. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Guerrero-Tamayo, A., Urquijo, B.S., Tosantos, MD.M. et al. Identification of mathematical patterns in genomic spectrograms linked to variant classification in complete SARS-CoV-2 sequences. Sci Rep 15, 43214 (2025). https://doi.org/10.1038/s41598-025-27279-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-27279-0