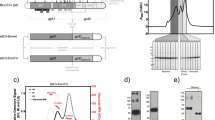
Abstract
Enzootic bovine leukosis (EBL) is an economically important disease of cattle caused by the bovine leukemia virus (BLV). Although BLV-seropositive dairy cattle were previously reported in the Emirate of Dhabi, UAE (EAD), molecular characterization of circulating BLV strains has not been undertaken. Therefore, the objectives of this study were to reassess the seroprevalence along with evaluating the genetic diversity of BLV strains circulating in dairy cattle in the EAD. Sera from 782 dairy cattle distributed across 11 farms were ELISA-screened and RT-qPCR testing of seropositive samples was followed by Sanger sequencing of the partial BLV env-gp51 gene (~ 423 bp) and phylogenetic analysis. The overall BLV herd seroprevalence was 27.3% (CI: 6.03%–61.00%), mean animal seroprevalence 33.5% (CI:30.20%–36.93%), and individual farm seroprevalence 28.00% (CI:19.00%–36.00%), 70.00% (CI:56.00%–84.00%), and 64.00% (59.00%–70.00%) for Farms 7, 11, and 14 respectively. Viral RNA was detected in 107 of 205 (52.2%) seropositive cattle, and phylogenetic analysis revealed a high genetic relatedness (~ 99.3–100.0%) among the BLV strains from the EAD. Additionally, study BLV isolates cluster under BLV-genotype 4 along with strains from Belgium, Russia and Vietnam. BLV infection is confirmed in EAD cattle, with circulating genotype 4 strains closely related to those from Europe and Asia, suggesting potential transboundary connections and underscoring the need for coordinated regional control measures. Future studies should focus on characterizing BLV infection risk factors in dairy cattle farms in the EAD. In the meantime, UAE livestock health authorities should urgently consider developing a national EBL control policy.
Similar content being viewed by others
Introduction
Enzootic bovine leukosis (EBL) is an economically important disease of cattle caused by the bovine leukaemia virus (BLV). The latter is an oncogenic retrovirus that belongs to the Genus Deltaretrovirus, and family Retroviridae along with several other lymphotropic viruses of animals and humans1,2. While majority of the BLV-infected cattle remain asymptomatic, approximately 30–70% of the animals demonstrate lifelong persistent lymphocytosis that is associated with immunosuppression3,4. In addition, about 0.1 to 10% of the infected cattle ultimately develop invariably fatal multiorgan B-cell lymposarcomas3,4. As a result of BLV-induced immunosuppression, BLV-positive cattle herds may show a higher incidence of opportunistic infections including but not limited to mastitis, diarrhoea, and pneumonia5
In respect to disease spread, both vertical and horizontal transmission of BLV have been documented in cattle herds worldwide4,6,7. Of the two modes of spread, horizontal transmission is more prevalent and driven by animal-to-animal contact through direct and indirect exposure to infected body fluids4. The specific risk factors for horizontal virus transmission include the use of contaminated needles8 the use of unsterilized dehorning tools4,9 not changing rectal palpation gloves between animals4,10 presence of biting dipterous flies at the farm 4,9 and the use of BLV-contaminated semen and bovine embryos4. In respect to vertical transmission, in utero transfer of the virus from dam to the unborn calf4 and peripartum acquisition of the virus to neonatal calves through the consumption of colostrum and milk have also been reported11. Presence of BLV-positive cattle in any herd perpetuates virus spread to in-contact cattle12 recent evidence has demonstrated that removal from the herd of animals with a high viral load significantly reduces BLV spread13,14. For this reason, most EBL control programs are focused on eliminating the BLV-infected cattle from the herd.
Across the globe, EBL is associated with significant economic implications for individual farms and the dairy industry at large. In a review paper reported by Kuczewski et al.4, the economic significance of the disease was attributed to the rejection of tumorigenic carcasses during meat inspection, reduced production and lifespan of BLV-infected cattle that frequently justifies premature culling, as well as the quantitative and qualitative changes in milk production. Besides, a number of researchers have reported that BLV-infected cattle suffer immunosuppression leading to increased susceptibility to other infectious agents3,4,5,15,16. This does further exacerbate the economic harm caused by the virus to cattle production. For example, in one study, economic losses associated with increased incidence of subclinical mastitis in BLV-infected cattle was estimated at US $418.59 per cow and total losses of US$ 6,097,255 for the entire prefecture of Hokkaido, Japan16.
Despite the tremendous economic transformation realized over the years in several oil-rich countries in the Middle East, the region still has significant livestock disease challenges, which in part, may be attributed to glaring gaps in disease control efforts at both national and regional levels17. Specifically, these authors have cited bottlenecks in the early detection, diagnosis, and reporting of animal diseases as major challenges of animal disease control in the region. In respect to the UAE, a high burden of infectious livestock diseases remains serious challenges to profitable dairy cattle production. Despite this, there remains limited livestock disease research in the country which is reflected by a critical paucity of infectious livestock disease research data in the literature. The status quo is complicating policy formulation needed to guide livestock disease surveillance and home-grown animal disease control efforts in the country. As such, there currently exists in the UAE a list of neglected livestock diseases among which EBL/BLV is becoming quite apparent! Despite previous serological evidence of EBL in the UAE18, there remains no statutory mandates for EBL biosecurity protocols, nor presence of nationally or regionally driven EBL control/eradication programs in the country. As a result of these gaps, the otherwise vibrant and rapidly growing dairy sector is being endangered thus jeopardizing the national food security aspirations originally inspired by Sheikh Zayed bin Sultan Al Nahyan the founding father of modern nation of the UAE.
While a previous survey detected BLV-seropositive dairy cattle in the Emirate of Abu Dhabi, UAE, the serological data remain uncorroborated by molecular testing to date. As such, there is an urgent need for definitive molecular confirmation of BLV infection in the national cattle herd ahead of developing an evidence-based control strategy for the disease in the UAE. Elsewhere, the molecular characterization of BLV strains has involved techniques such as RFLP or “restriction fragment length polymorphism”, sequencing of the env-gp51 gene, or sequencing of the entire BLV genome2,19. On the basis of env-gp51 gene sequences, 12 BLV genotypes have been reported in cattle worldwide2,19. Of the 12 BLV genotypes reported across the globe, genotype 1 (G1) is the most prevalent and has been in reported in 17 countries particularly in the Americas, Europe, and Australia20. Worldwide, the env-gp51 gene has been targeted for the diagnosis, genotyping and molecular characterization of BLV isolates21,22,23,24.
The specific objectives of the present study were to re-assess the seroprevalence of BLV at commercial dairy cattle farms in the Emirate of Abu Dhabi, definitively confirm BLV in cattle samples obtained from the study area through testing by RT-qPCR technique, and attempt molecular characterization of the BLV isolates of UAE origin by application Sanger sequencing and phylogenetic analysis of the partial sequences of the env-gp51 gene of BLV. Achieving this goal is anticipated to awaken the interest in EBL from the regional and federal livestock health agencies responsible for the control of animal diseases in the country.
Materials and methods
Study area, sample size calculation and blood sample collection
The Emirate of Abu Dhabi (EAD) has 17 commercial dairy farms, primarily concentrated in the Eastern region, with a total of 41,581 high-grade Holstein-Friesians raised for milk production. In a joint collaboration, a team of researchers from the Department of Veterinary Medicine, United Arab Emirates University (UAEU), and the Abu Dhabi Agriculture & Food Safety Authority (ADAFSA), collected a total of 782 bovine blood samples from 11 commercial dairy cattle farms spatially distributed across the Emirate of Abu Dhabi; these samples were screened by indirect ELISA followed by RT-qPCR-testing of the and BLV-seropositive samples. For privacy reasons communicated through ADAFSA by the participating dairy farms, however, the blood sampling was totally blinded to the UAEU research team. As such, the blood sample collection was done by ADAFSA veterinarians who availed the specimens to the UAEU research team for ELISA screening and subsequent testing by RT-qPCR. The RT-qPCR-positive blood samples were availed to the ADAFSA team who performed the Sanger sequencing and phylogenetic analysis of the BLV isolates.
The sample size for each farm was based on a 2% margin of error and 98% confidence level, with an assumed estimated proportion of 0.5 for each stratum (farm) according to a conservative scenario. Specifically, an expected prevalence of 50% (0.5) was used due to the absence of prior large-scale data on BLV seroprevalence in dairy cattle in the study region. A 98% confidence level was selected to ensure a high degree of confidence in the estimate, and a 2% margin of error was chosen to achieve high precision in estimating the true prevalence. The formula below was used to calculate the respective individual farm sample sizes (Table 1).
where \(N\) is the total population of cattle at the selected study farms put together, \({N}_{h}\) is the total population of cattle at each study farm \(h\), \({p}_{h}\) population proportion at each farm \(\text{h}\), \({\text{q}}_{\text{h}}=1-{\text{p}}_{\text{h}}\), \(\text{D}=\frac{{\upbeta }^{2}}{{\text{z}}^{2}}\), \(\upbeta =\text{margin of error},\text{ z}=1.96 (95\text{\% Confidence level})\). Based on this, the sample size for each farm (\(h\)) is given as:
Blocking enzyme-linked immunosorbent assay for the detection anti-bovine leukemia virus antibody
The sera samples separated by centrifugation from the blood samples collected into plain vacutainers were tested for BLV antibodies using a blocking ELISA per the kit manufacturer’s instructions (IDEXX Laboratories, Switzerland). Briefly, the test sera and the positive and negative controls were diluted at 1:2 in the kit wash buffer solution, 100 μL of diluted sera pipetted into the microplate wells precoated with BLV gp51 antigen and the rest of the ELISA protocol done in accordance with the instructions of the kit manufacturer. Once the 3,3′,5,5′-tetramethylbenzidine substrate (or TMB) was added and the reaction ultimately stopped, the plates were read at 450 nm using spectrophotometer (Multiskan Sky Thermo Scientific, Singapore). The results were calculated as a percentage of the ratio of the OD450 of the test samples to the OD450 value of the negative control (S/N %). Based on this calculation, and as indicated in the kit manual, any sample that showed an S/N % < 40% was determined to be positive while one with an S/N % value ≥ 40% was deemed negative.
RNA extraction and reverse transcriptase polymerase chain reaction
RNA extraction
The viral RNA needed for RT-PCR was extracted from BLV-ELISA-positive blood samples using the GPSpin Viral RNA Kit (Genetic PCR Solutions, Spain). As whole blood samples were not available, the viral RNA was extracted from homogenates of “left over blood clot” saved after separation of the serum samples. Briefly, 200 μL of each one of the homogenized blood clot samples were placed into a microtube, 400 μL of the kit lysis buffer added, the microtubes vortexed for 20 s, and then incubated at RT for 15 min. Following the incubation step, 30 μL of the kit protein precipitation buffer were added to each tube, the mixture further vortexed for 5 s, and the microtubes incubated for 1 min at RT. After centrifugation at 11,000 rpm for 3 min, the supernatants were transferred into new microtubes, and 350 μL of 100% ethanol added. Following this step, half of the lysates were then loaded into the reservoir of combined microspin column-collection tube assembly units and centrifuged at 8000 rpm for 30 s; and the other half of the cell lysates were similarly processed in the columns and centrifuged. The viral RNA specimens that remained bound to the microspin columns were subjected to a series of washes and additional centrifugation steps to eliminate any contaminating cellular components. Finally, the viral RNA samples were eluted from the columns using the kit elution buffer after and then stored at − 80 °C until testing by RT-PCR.
Reverse transcriptase real-time polymerase chain reaction (RT-qPCR)
For the RT-qPCR assay, molecular analysis of the extracted RNA samples utilized the BLV MONODOSE dtec-qPCR kit (Genetic PCR Solutions, Orihuela, Alicante, Spain), which consists of pre-packaged tubes containing all necessary components for the detection of BLV (specific primer details are confidential per the manufacturer). According to the kit instructions, 7 µL of sample was added to each tube, along with 13 µL of DNase/RNase-free water, reaching a total qPCR volume of 20 µL. The tubes were labeled, sealed, vortexed thoroughly, pulse-spun, and then placed in the thermal cycler. Amplification was carried out using the CFX 96 Real-time detection system (BIO-RAD, Hercules, CA, USA) under the following thermal conditions: 10 min of retrotranscription at 50 °C, followed by 2 min of activation at 95 °C, and then 40 cycles consisting of 5 secs of denaturation at 95 °C and 20 s of hybridization/extension and data collection at 60 °C. Fluorogenic signals were collected during the hybridization/extension and data collection phase, utilizing the FAM channel for the target sample and the HEX channel for the internal control. The software automatically calculated the results (Cts) for the negative control, positive control, and samples from both channels (FAM and HEX).
PCR, Sanger Sequencing, and BLAST analysis
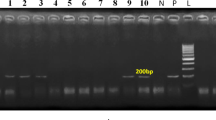
Viral DNA was extracted from blood homogenized blood clots that previously tested positive for BLV antibodies and BLV-RNA by ELISA and RT-qPCR respectively. In total, seven samples, representing farms 11 and 14, were selected for subsequent genotyping based on the Ct values that ranged between 23.31 to 27.66. Amplification of partial sequence of env-gp51 gene of BLV was done using AmpliTaq Gold® 197 360 (Applied Biosystems, Waltham, MA, USA) according to the kit manufacturer’s instructions. The PCR primers and cycling condition were described earlier1. Briefly, the thermal cycling conditions were set: initial incubation at 94 °C for 4 min, followed by repeated 30 cycles: denaturation 95 °C for 60 s, annealing 55 °C 60 s and extension by polymerase at 72 °C 60 s and a final extension step was run at 72 °C for 10 min. The amplicon of env-gp51 gene was purified with the ExoSAP-ITTM PCR Product Cleanup Reagent (Thermo Fisher Scientific), and Sanger sequencing reaction was prepared suing the BigDyeTM Terminator v3.1 Cycle Sequencing kit (Applied Biosystems, MA, USA) and the same primers utilized to perform the PCR. The Cycle Sequencing reaction mixture totaling 20 μL comprised 9 µL of water, 3.5 µL of 5 × Sequencing Buffer, 1 µL of the BigDyeTM Terminator v3.1, 1 µL of primers (3.2 pmol/µL), and 5.5 µL of DNA. Next, the mixture was purified using the BigDyeTM XTerminatorTM Purification kit (Applied Biosystems, MA, USA) according to the manufacturer’s instructions. A bi-directional sequencing was carried out on a SeqStudio Genetic Analyzer (Applied Biosystems, MA, USA) using the ‘MediumSeq BDX’ run module. The obtained sequences were trimmed and assembled using CLC Genomic Workbench v.20 (Qiagen, Aarhus, Denmark) followed by BLAST analysis using the BLAST Tool at NCBI GenBank (Basic Local Alignment Search Tools) (http://www.ncbi.nlm.nih.gov/blast) to confirm the sequence identity. The BLV env-gp51 gene (~ 423 bp) sequence of the representative BLV UAE’s strains identified in the present study were submitted to GenBank databases under accession numbers: PV796100 (BankIt2970609 Farm_11_s7), PV796101 (BankIt2970609 Farm_11_s10), PV796102 (BankIt2970609 Farm_11_s12), PV796103 (BankIt2970609 Farm_11_s14), PV796104 (BankIt2970609 Farm_11_s20), PV796105 (BankIt2970609 Farm_11_s24), and PV796106 (BankIt2970609 Farm_14_s1).
Spatial maps generated by the Geographical Information System (GIS)
The spatial coordinates for the study commercial dairy cattle farms located in the Emirate of Abu Dhabi, provided by the Animal Health Division of the Abu Dhabi Agriculture & Food Safety Authority were mapped using “Esri. (2025). ArcGIS Pro, Version 3.4; Redlands, CA: Environmental Systems Research Institute”, as well as ArcGIS Online. The seroprevalence data were projected in the WGS 1984 coordinate system and the farms that tested negative for BLV were denoted with grey, while BLV-positive farms were marked in red symbols. Graduated color intensity and proportional symbols were applied to represent varying seroprevalence levels. The GIS map showing the spatial distribution of the BLV-positive (red symbols) and BLV-negative (grey symbols) dairy cattle farms across the Emirate of Abu Dhabi is presented in Fig. 2.
Sequence Alignment and Phylogenetic Analysis
The env-gp51 gene of the BLV-UAE strains was aligned using the ClustalW program in MEGA 1125. This alignment included sequences of both the BLV isolates from the UAE and corresponding sequences of reference strains from 12 genotypes of the BLV lineages obtained from the GenBank of National Center for Biotechnology Information (NCBI) or previous publications4,19,26 totaling 95 sequences. A phylogenetic tree was constructed using the Maximum Likelihood technique and the Kimura 2-parameter model27 within the MEGA 11 program28. The reliability of phylogenetic relationships was evaluated using bootstrap analysis with 1000 replicates. The pairwise nucleotide sequence similarity within the BLV-UAE env-gp51 gene was then determined.
Data analysis
In this study, summary statistical methods were employed to estimate the BLV seroprevalence in dairy cattle herds across the EAD in the UAE. To this end, the proportion of seropositive cattle was the primary measure derived by dividing the number of seropositive animals by the total number of cattle sampled within each herd. To quantify the uncertainty associated with these estimates, the corresponding 95% confidence intervals were calculated. Additionally, we examined the statistical significance of differences among BLV-seropositive dairy cattle farms using a “generalized linear model” (GLM) featuring a binomial distribution and a logit link function. In this analysis, the response variable was defined as the number of seropositive cattle out of the total number of cattle tested on each farm, while the predictor was a factor with several levels representing cattle farms. The GLM framework allowed the modeling of the proportions of seropositive animals by transforming the probabilities of seropositivity into log-odds through the logit link function. This transformation ensured that the predicted probabilities remained within the [0, 1] range, which was essential for accurate interpretation. By including a categorical variable representing the different seropositive cattle farms, the GLM facilitated the comparison of seroprevalence among these farms. The model estimates coefficients for each farm indicating how the likelihood of seropositivity varies across different locations (farms). In respect to molecular characterization of the BLV strains from the study area of the UAE, the sequences obtained by amplification of partial sequence of env-gp51 gene were trimmed and assembled using CLC Genomic Workbench v.20 (Qiagen, Aarhus, Denmark). To confirm the sequence identity, this was followed by BLAST analysis using the BLAST Tool at NCBI GenBank (Basic Local Alignment Search Tools: http://www.ncbi.nlm.nih.gov/blast). The BLV env-gp51 gene (~ 423 bp) sequence of the representative BLV strains from the study area of the UAE were submitted to GenBank databases. For the sequence alignment and phylogenetic analysis, the env-gp51 gene of the BLV-UAE strains was done using the ClustalW program in MEGA 1125. This alignment also included corresponding sequences of reference BLV strains from 12 genotypes of the BLV lineages obtained from the GenBank of National Center for Biotechnology Information (NCBI) or previous publications4,19,26 totaling 95 sequences. A phylogenetic tree was constructed using the Maximum Likelihood technique and the Kimura 2-parameter model27 within the MEGA 11 program28 The reliability of phylogenetic relationships was evaluated using bootstrap analysis with 1000 replicates after which the pairwise nucleotide sequence similarity within the BLV-UAE env-gp51 gene was determined.
Ethical approvals
Before undertaking these studies, approval of the research protocol was obtained from the UAEU Animal Ethics Committee, Research Ethics Sub-committee under approval Certificate No ERA_2023_2611 dated 20th March 2023. In addition, all field and laboratory methods used in the present study were performed in accordance with the relevant guidelines and regulations. Furthermore, permission was obtained from the participating dairy farms before the ADAFSA veterinarians performed the blood sample collections.
Results
Herd and animal seroprevalence of enzootic bovine leukosis in the Abu Dhabi Emirate
Of the 11 study farms, only three (Farms #7, 11, and 14) were confirmed to be BLV-positive by ELISA and RT-qPCR. Thus, the herd prevalence of BLV was 27.3% (95% confidence interval, CI: 6.03%–61.00%). Although only three farms were positive, the seroprevalence of BVL appeared to increase as the herd size increased. During the study, 782 cattle blood samples were tested by ELISA (Table 1) giving an overall animal BLV seroprevalence of 33.50% (95% CI: 30.20% to 36.93%) with individual animal seroprevalences of 28.00%, 70.00%, and 64.00% for farms 7, 11, and 14 respectively (Table 1). All the blood samples collected from farms 1, 2, 4, 6, 9, 10, 12, and 13 totaling to 520 were seronegative.
Table 2 presents the parameter estimates of GLM fitted for the three farms (Farms 7, 11 and 14) at which BLV seropositive cattle were detected with Farm 7 used as a reference or control level for this analysis. It is clear that all of the estimated coefficients (log-odds) for Farm 11 and Farm 14 are significantly higher than those for Farm 7 (p-value < 0.01, Table 2) meaning that the chance of an animal being BLV-seropositive is higher in Farm 11 than in Farm 7 and in Farm 14 than in Farm 7. By exponentiating these coefficients, the odd ratios were obtained and interpreted as follows: For Farm 11, the odds ratio is (e^1.8141 = 6.1), implying that the odds of being seropositive in Farm 11 are more than 6 times the odds of being seropositive in Farm 7. Similarly, the odds of being seropositive in Farm 14 (e^1.5634 = 4.8) are approximately 5 times the odds of being seropositive in Farm 7. Given the high variability in the estimated BLV seroprevalence, the interpretation of these observations is cautiously considered.
Detection of viral RNA in bovine leukaemia virus-seropositive cattle
RNA samples extracted from homogenized blood clots originally taken from the BLV-ELISA-positive blood samples from Farms 7, 11, and 14 were tested by RT-qPCR. It is noted that for Farm 7, only 11 out of the 25 (44.00%) ELISA-positive samples tested positive by RT-qPCR (Table 3). By comparison, BLV was detected in 13 out of 23 (56.52%) ELISA-positive blood samples from Farm 11, and in 83 out of 157 ELISA-positive samples (40.49%) from Farm 14 (Table 3). In summary, a total of 107 animals out of a total of 205 that were ELISA-positive for BLV ultimately tested positive for the BLV viral RNA by the RT-qPCR test (Fig. 1; Table 3) while 98 out of 205 were negative. The RT-qPCR negative samples include 14/25 (56.00%) from Farm 7, 10/23 (43.48%) from Farm 11, and 74/157 (59.51%) from Farm 14.
GIS map showing the spatial distribution of BLV-positive and BLV-negative farms across the Emirate of Abu Dhabi, United Arab Emirates (Map created using ArcGIS® software Version 3.4 by Esri. Basemap and data sources: Esri, TomTom, Garmin, FAO, NOAA, USGS, OpenStreetMap contributors, & GIS User Community. (n.d.). Streets [Web map]. ArcGIS Online. Retrieved July 23, 2025, from https://www.arcgis.com/home/item.html?id=e3e6df1d2f6a485d8a70f28fdd3ce19e).
Spatial distribution of BLV-positive farms
As presented in Fig. 2 below, the three BLV-positive dairy cattle properties including Farms 7, 11, and 14 clustered in a small area of the Al Ain Region, Emirate of Abu Dhabi. It is noteworthy that despite the different spatial locations, Farms 7 and 11 belong to the same proprietor while Farm 14 belongs to a different entity altogether.
A maximum likelihood (ML) phylogenetic tree was constructed using nucleotide sequences of the partial BLV env-gp51 gene (~ 423 bp) from UAE strains alongside representative strains of all known BLV genotypes. The BLV strains identified in this study are highlighted in bold red text and marked with red star (*), while the remaining strains are labeled with their respective country names and GenBank accession numbers. Genotypes are labeled with the symbol “G.” Bootstrap support values for the ML analysis are displayed at the branch nodes.
Phylogenetic analysis of env-gp51 gene sequences
The BLAST analysis indicated the BLV sequence from UAE share 99.00–100.00% sequence identity to the BLV sequences available in the GenBank nucleotide dataset (Fig. 2). Phylogenetic analysis of the partial (~ 423 bp) BLV env-gp51 gene indicated that the BLV strains from the UAE originated from Farm 11 and farm 14 both located in the Al Ain Region of the Emirate of Abu Dhabi. These strains were clustered within genotype 4, and share 99.30–100.00% identity within all the sequences of the UAE-BLV strains. When the spatial distribution of the BLV-positive farms is assessed against the genetic relatedness of the BLV strains, the BLV-stricken farms appear to cluster in one small geographical area of the Emirate of Abu Dhabi.
Discussion
Based on ELISA and RT-qPCR data, the present study corroborates the presence of BLV infection in dairy cattle in the EAD, UAE, as initially reported by Hassan et al.18. Furthermore, data obtained from the phylogenetic analysis of the partial (~ 423 bp) BLV env-gp51 gene indicate that the BLV strains circulating in the study area exhibit high genetic similarity (99.30–100.00%) and belong to genotype 4. Notably, only three of the eleven dairy cattle farms surveyed in the EAD, UAE, had BLV-seropositive animals, highlighting the heterogeneous distribution of the virus and suggesting that farm-level factors may strongly influence infection risk. While the mean seroprevalence across the 11 farms (14.73%) observed in the present study was lower than the 37.00% previously reported for the same area by Hassan et al.18, individual farm seroprevalence values were particularly high for Farm 11 (70.00%) and Farm 14 (64.00%), and moderate for Farm 7 (28.00%). Notably, whereas the seroprevalence data reported by Hassan et al.18 were not supported by RT-qPCR evidence, the present study complemented serological findings with molecular testing and phylogenetic analysis of the BLV strains. As the present data show however, BLV seroprevalence varied markedly across the surveyed farms, with 95% confidence intervals reflecting the uncertainty around these estimates. Seven of the eleven farms showed no evidence of infection (0.00%), whereas Farms 7, 14, and 11 had seroprevalences of 28.00%, 64.00%, and 70.00%, respectively, highlighting substantial farm-to-farm variability. This pattern suggests that BLV transmission is not uniform across the region and may be influenced by farm-level factors such as herd management, biosecurity practices, and animal movement, underscoring the need for targeted control measures. Such variability is consistent with observations elsewhere and has been, indeed, attributed to differences in farm-level risk factors, including biosecurity practices, sourcing of new stock from herds of unknown BLV status, and purchasing semen from companies with unverified BLV status4,29,30. Additional risk factors in the present context may include the use of unsterilized instruments and failure to change arm-length gloves between animals during rectal palpation4,8,10. Comprehensive epidemiological studies are needed to identify the factors driving farm-to-farm variability in BLV seroprevalence, as this would enable seropositive farms to adopt biosecurity practices from seronegative farms and limit virus transmission. Furthermore, such data could inform government agencies in designing targeted BLV control programs at both regional and national levels. It is noteworthy that the Ministry of Climate Change & Environment (MOCCAE) is the federal agency responsible for livestock health and disease control in the UAE, while ADAFSA holds similar jurisdiction within the EAD. As part of the present EBL study, the leadership of these agencies should utilize the current research findings to develop and implement a BLV control and eradication strategy, protecting the UAE’s dairy sector from the substantial economic and food security risks posed by the disease.
In the study reported by Saushkin et al.31 RT-qPCR and ELISA concurred in 93.40% of the cases consistent with a very high degree of concordance between the two techniques. In the present study, however, 98 out 205 (47.80%) of the BLV-ELISA-positive cattle blood samples ultimately tested negative by the RT-qPCR. The relatively low concordance (52.2%) observed between ELISA and RT-qPCR in the present study is not unexpected, despite the blocking ELISA being reputed for its high diagnostic specificity and sensitivity, typically reported at 97% or higher31,32,33. In it is also possible that the relatively low concordance between the two techniques could be attributed to the testing by RT-qPCR of “left over” blood clots instead of using the buffy coat. Notably, whole blood samples were unavailable, so the research team extracted viral RNA from homogenized blood clots remaining after serum separation. Various researchers have reported that BLV is associated with B-lymphocytes34,35,36 and therefore the highest concentration of virus would have been expected to be in the buffy coat. Future studies should prioritize testing buffy coats instead of blood clots to enhance BLV detection by RT-qPCR. Screening proviral DNA, which offers even greater sensitivity, may also be considered as the preferred molecular detection approach, improving the accuracy of epidemiological assessments and complementing the findings of the present study.
It is noteworthy that phylogenetic analysis of the partial (~ 423 bp) BLV env-gp51 gene revealed that BLV strains from the study area in the UAE belong to genotype 4, exhibiting a high sequence identity of 99.30–100.00%. Such limited genetic variability suggests a relatively recent and possibly single introduction event, followed by local transmission among local farms in the study area. Moreover, the UAE strains clustered with genotype 4 strains previously reported in Russia and Belgium, indicating a potential European origin. This close clustering implies potential epidemiological linkage, likely reflecting historical importation of breeding or dairy stock from Europe—a common source of Holstein–Friesian cattle in the Gulf region. The predominance of genotype 4 in geographically diverse locations also supports its broad distribution and adaptability across bovine populations. To clarify transmission dynamics and introduction routes, future studies should incorporate whole-genome sequencing, combined with detailed trade, movement, and epidemiological data, as well as screening of imported and local animals. Such integrated approaches would help trace the origin of infection, elucidate viral evolution within the UAE, and inform effective biosecurity and control measures. Furthermore, the molecular characterization of BLV isolates from the UAE, as initiated in this study, could facilitate the development of customized vaccines and biologics tailored to the locally circulating strains37,38,39.
Conclusion
The present study has corroborated the presence of BLV infections in UAE cattle and provided data from the initial molecular characterization of locally circulating strains in dairy cattle. Nationwide surveys and targeted investigations of farm-level risk factors are needed to further clarify BLV epidemiology and guide effective control strategies. Once fully done, molecular characterization could also support the development of vaccines and biologics tailored to local BLV strains. In the meantime, livestock health authorities should prioritize BLV control across the dairy sector.
Data availability
The datasets generated and/or analysed during the current study are available in the GenBank of the National Center for Biotechnology Information (NCBI) repository under accession numbers: PV796100 (BankIt2970609 Farm_11_s7), PV796101 (BankIt2970609 Farm_11_s10), PV796102 (BankIt2970609 Farm_11_s12), PV796103 (BankIt2970609 Farm_11_s14), PV796104 (BankIt2970609 Farm_11_s20), PV796105 (BankIt2970609 Farm_11_s24), and PV796106 (BankIt2970609 Farm_14_s1).
References
Moratorio, G. et al. Phylogenetic analysis of bovine leukemia viruses isolated in South America reveals diversification in seven distinct genotypes. Arch. Virol. 155, 481–489. https://doi.org/10.1007/s00705-010-06063 (2010).
Polat, M., Sn, T. & Aida, Y. Epidemiology and genetic diversity of bovine leukemia virus. Virol. J. 14, 209. https://doi.org/10.1186/s12985-017-0876-4 (2017).
Frie, M. C. & Coussens, P. M. Bovine leukemia virus: a major silent threat to proper immune responses in cattle. Vet. Immunol. Immunopathol. 163, 103–114. https://doi.org/10.1016/j.vetimm.2014.11.014 (2015) (Epub 2014 Dec 15 PMID: 25554478).
Kuczewski, A. et al. Invited review: Bovine leukemia virus-transmission, control, and eradication. J. Dairy Sci. 104(6), 6358–6375. https://doi.org/10.3168/jds.2020-18925 (2021).
OIE Terrestrial Manual, (2018). https://www.woah.org/fileadmin/Home/eng/Health_standards/tahm/2.04.10_EBL.pdf
Mekata, H. et al. Evaluation of the natural perinatal transmission of bovine leukaemia virus. Vet. Rec. 176, 254. https://doi.org/10.1136/vr.102464 (2015).
Sajiki, Y. et al. Intrauterine infection with bovine leukemia virus in pregnant dam with high viral load. J. Vet. Med. Sci. 79, 2036–2039. https://doi.org/10.1292/jvms.17-0391 (2017).
Ramírez Vásquez, N. F., Villar Argaiz, D. & Fernández Silva, J. A. Seroprevalence and risk factors of several bovine viral diseases in dairy farms of San Pedro de los Milagros, Antioquia, Colombia. CES Med. Vet. Zootec. 11, 15–25 (2016).
Kobayashi, S. et al. Risk factors associated with within-herd transmission of bovine leukemia virus on dairy farms in Japan. BMC Vet. Res. 6, 1. https://doi.org/10.1186/1746-6148-6-1 (2010).
Kohara, J., Konnai, S. & Onuma, M. Experimental transmission of Bovine leukemia virus in cattle via rectal palpation. Jpn J. Vet. Res. 54, 25–30 (2006).
Hopkins, S. G. & DiGiacomo, R. F. Natural transmission of bovine leukemia virus in dairy and beef cattle. Vet. Clin. North Am. Food Anim. Pract 13, 107–128. https://doi.org/10.1016/s0749-0720(15)30367-4 (1997).
Maresca, C. et al. Enzootic bovine leukosis: Report of eradication and surveillance measures in Italy over an 8-year period (2005–2012). Prev. Vet. Med. 119, 222–226. https://doi.org/10.1016/j.prevetmed.2015.02.024 (2015).
Rodríguez, S. M. et al. Preventive and therapeutic strategies for bovine leukemia virus: Lessons for HTLV. Viruses 3, 1210–1248. https://doi.org/10.3390/v3071210 (2011).
Ruggiero, V. J. et al. Controlling bovine leukemia virus in dairy herds by identifying and removing cows with the highest proviral load and lymphocyte counts. J. Dairy Sci. 102, 9165–9175. https://doi.org/10.3168/jds.2018-16186 (2019).
Nakada, S., Fujimoto, Y., Kohara, J., Adachi, Y. & Makita, K. Estimation of economic loss by carcass weight reduction of Japanese dairy cows due to infection with bovine leukemia virus. Prev. Vet. Med. 198(105528), 01675877. https://doi.org/10.1016/j.prevetmed.2021.105528 (2022).
Nakada, S., Fujimoto, Y., Kohara, J. & Makita, K. Economic losses associated with mastitis due to bovine leukemia virus infection. J. Dairy Sci. 106, 576–588. https://doi.org/10.3168/jds.2021-21722 (2023).
Shimshony, A. & Economides, P. Disease prevention and preparedness for animal health emergencies in the Middle East. Rev. Sci. Tech. 25, 253–269. https://doi.org/10.20506/rst.25.1.1667 (2006).
Hassan, N. A. D. et al. Serological evidence of enzootic bovine leukosis in the periurban dairy cattle production system of Al Ain, United Arab Emirates. Trop. Anim. Health Prod. 52, 2327–2332. https://doi.org/10.1007/s11250-020-02262-1 (2020).
Le, D. T. et al. Detection and genotyping of bovine leukemia virus (BLV) in Vietnamese cattle. J. Vet. Med. Sci. 82, 1042–1050. https://doi.org/10.1292/jvms.20-0094 (2020).
Olaya-Galán Nno. Virus de la leucosis bovina (VLB) y evidencias de su potencial zoonótico. (PhD thesis, Universidad del Rosario; 2021) https://doi.org/10.48713/10336_33690
Felmer, R., Munoz, G., Zuniga, J. & Recabal, M. Molecular analysis of a 444 bp fragment of the bovine leukaemia virus gp51 env gene reveals a high frequency of non-silent point mutations and suggests the presence of two subgroups of BLV in Chile. Vet. Microbiol. 108, 39–47 (2005).
Lee, E. et al. Sequencing and phylogenetic analysis of the gp51 gene from Korean bovine leukemia virus isolates. Virol. J. 12, 64. https://doi.org/10.1186/s12985-015-0286-4 (2015).
Lee, E. et al. Molecular epidemiological and serological studies of bovine leukemia virus (BLV) infection in Thailand cattle. Infect. Genet. Evol. 41, 245–254. https://doi.org/10.1016/j.meegid.2016.04.010 (2016).
Yu, C. et al. Genotyping bovine leukemia virus in dairy cattle of Heilongjiang, northeastern China. BMC Vet. Res. 15, 1–9 (2019).
Larkin, M. A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2848 (2007).
Mendoza, W., Isaza, J. P., López, L., López-Herrera, A. & Gutiérrez, L. A. Bovine leukemia virus molecular detection and associated factors among dairy herd workers in Antioquia, Colombia. Acta Trop. 256, 107253 (2024).
Kimura, M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol. 16, 111–120 (1980).
Tamura, K., Stecher, G. & Kumar, S. MEGA11: Molecular evolutionary genetics analysis version 11. Mol. Biol. Evol. 38, 3022–3027 (2021).
Bulla-Castañeda, D. M. et al. Seropositivity and risk factors associated with the presentation of bovine leukosis virus in Sotaquirá, Colombia. Vet. World 14, 2212–2218. https://doi.org/10.14202/vetworld.2021.2212-2218 (2021).
Lancheros-Buitrago, D. J., Bulla-Castañeda, D. M., Giraldo-Forero, J. C. & Pulido-Medellin, M. O. Risk factors associated with enzootic bovine leukosis in Boyacá and Cundinamarca municipalities, Colombia. Open Vet. J. 13, 1012–1020. https://doi.org/10.5455/OVJ.2023.v13.i8.7 (2023).
Saushkin, N. Y., Samsonova, J. V., Osipov, A. P. & Kondakov, S. E. Strip-dried blood sampling: Applicability for bovine leukemia virus detection with ELISA and real-time PCR. J. Virol. Methods 263, 101–104. https://doi.org/10.1016/j.jviromet.2018.11 (2019).
John, E. E. et al. Development of a predictive model for bovine leukemia virus proviral load. J. Vet. Intern. Med. 36, 1827–1836. https://doi.org/10.1111/jvim.16506 (2022).
Trono, K. G., Pérez-Filgueira, D. M., Duffy, S., Borca, M. V. & Carrillo, C. Seroprevalence of bovine leukemia virus in dairy cattle in Argentina: Comparison of sensitivity and specificity of different detection methods. Vet. Microbiol. 83, 235–248. https://doi.org/10.1016/s0378-1135(01)00420-5 (2001).
Jensen, W. A., Sheehy, S. E., Fox, M. H., Davis, W. C. & Cockerell, G. L. In vitro expression of bovine leukemia virus in isolated B-lymphocytes of cattle and sheep. Vet. Immunol. Immunopathol. 26, 333–342. https://doi.org/10.1016/0165-2427(90)90117-b (1990).
Paul, P. S. et al. Detection of bovine leukemia virus in B-lymphocytes by the syncytia induction assay. J. Natl. Cancer Inst. 59, 1269–1272. https://doi.org/10.1093/jnci/59.4.1269 (1977).
Paul, P. S. et al. Evidence for the replication of bovine leukemia virus in the B lymphocytes. Am. J. Vet. Res. 38, 873–876 (1977).
Abdala, A. et al. BLV: Lessons on vaccine development. Retrovirology 16, 26. https://doi.org/10.1186/s12977-019-0488-8 (2019).
Gutiérrez, G. et al. Vaccination against δ-retroviruses: The bovine leukemia virus paradigm. Viruses 6, 2416–2427. https://doi.org/10.3390/v6062416 (2014).
Suárez Archilla, G. et al. A safe and effective vaccine against bovine leukemia virus. Front. Immunol. 13, 980514. https://doi.org/10.3389/fimmu.2022.980514 (2022).
Acknowledgements
The authors are grateful to the participating commercial dairy farms for availing their cattle for purposes of sample collection. We also acknowledge the State veterinarians from the Abu Dhabi Agriculture and Food Safety Authority, UAE who, as part of their routine disease surveillance mandate, collected the blood samples.
Funding
The research project was financially supported by the United Arab Emirates University, as part of the SURE Plus funding program, under Grant No G00003833.
Author information
Authors and Affiliations
Contributions
RB was project principal investigator and along with co-authors AA, IMAA and GA conceptualized and wrote the research proposal, and subsequently contributed to data analysis and took a lead in drafting of the manuscript. NAH and AZ are research fellows who did the laboratory testing of the blood samples while co-authors OKB, SAA are State veterinarians who collected the blood samples from the study farms while MS A supervised the State veterinarians who collected blood samples. Co-author HZI, with involvement of MA and AMAS did the phylogenetic analysis. Last but not least, co-authors AM, PT, LS, HAMA, and YMA were undergraduate veterinary students who were mentored during the research project and participated in the laboratory testing of samples as a requirement for the Summer Undergraduate Research Experience (SURE) Plus grant. Co-author AWW generated the GIS maps.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
This research was approved by the United Arab Emirates University Animal Research Ethics Committee No: ERA_2023_2611, titled: “Molecular Studies Of Bovine Leukosis in Commercial Dairy Cattle Herds From the Periurban Region of Al Ain, Abu Dhabi” and dated 20 March 2023.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hassan, N.A., Ishag, H.Z., Anthony, A. et al. Molecular characterization of bovine leukemia virus detected in dairy cattle herds from the Emirate of Abu Dhabi, United Arab Emirates. Sci Rep 15, 44915 (2025). https://doi.org/10.1038/s41598-025-28570-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-28570-w