Abstract
The Ki67 proliferation index (PI) serves as a crucial prognostic indicator in clinical settings, widely utilized for evaluating breast cancer progression and forecasting chemotherapy efficacy. Nonetheless, conventional manual PI estimation methods are plagued by high subjectivity, time inefficiency, and limited reproducibility. Despite notable advancements in deep learning for Ki67 PI computation, current approaches face challenges in concurrently capturing local cellular morphological features and global tumor proliferation data, particularly when addressing issues like complex backgrounds, staining variability, tumor heterogeneity, and noise in low-resolution images. In response, we introduce Kpi-Net, a multi scale deep learning framework built on U-Net, designed to precisely quantify the Ki67 index in breast cancer pathology images. Initially, we develop a Residual Dilated Multi Scale Module (RDMS Module) that utilizes multi-branch dilated convolutions and residual connections to capture both local and global information, while integrating a Transformer Block to bolster global modeling, effectively mitigating missed and erroneous detections arising from irregular cell distribution. Next, we introduce the High-level Screening-Convolutional Block Attention Module Feature Pyramid Networks (HS-CBAM-FPN), incorporating channel and spatial attention mechanisms alongside the Selective Feature Fusion with CBAM (SFF-CBAM) to facilitate effective integration of multi-level features. Lastly, we apply the watershed algorithm to feature maps, refining cell cluster segmentation via distance transformation and local maximum detection, which improves the precision of Ki67 index computation. Comprehensive experimental results indicate that Kpi-Net surpasses current leading methods in metrics like F1 score and root mean square error (RMSE), highlighting its potential for accurate Ki67 index computation and precise cell detection, thereby offering a dependable tool for accurate breast cancer diagnosis and therapeutic decision-making.
Similar content being viewed by others
Introduction
In recent years, the global incidence of breast cancer has continued to rise, with approximately 2.3 million new cases and 685,000 deaths reported globally in 20221. In light of this challenging scenario, early diagnosis and precise treatment are paramount for improving patient survival rates. The formulation of treatment strategies and prognosis assessment for breast cancer are highly dependent on the accurate detection of key biomarkers2. The Ki67 proliferation index (PI), a core metric for quantifying tumor cell proliferative activity, plays an indispensable role in breast cancer molecular subtyping, therapeutic decision-making, and prognosis prediction3. Extensive research has consistently demonstrated that the Ki67 index is significantly associated with tumor aggressiveness, recurrence risk, and patient survival.
However, in clinical practice, while widely utilized, manual assessment of the Ki67 index presents significant limitations. This process is inherently labor-intensive and time-consuming; more critically, it suffers from poor reproducibility. The intraclass correlation coefficient (ICC) among different pathologists is only 0.6–0.84, and even repeated assessments by the same pathologist may yield inconsistent scores. This variability stems primarily from two complexities: (1) the intrinsic spatial heterogeneity of tumors, leading to variation in cell proliferation status across different tissue regions5, and (2) the non-uniform distribution of Ki67 positive cell expression patterns, where local variations in PI can be as high as ±14%6. Consequently, manual visual counting often fails to provide a comprehensive, objective, and consistent representation of the tumor’s overall proliferative state.
The rapid advancement in whole-slide imaging (WSI) technology offers a breakthrough for this conundrum. The increasing availability of high-resolution digital pathology images provides a robust foundation for the development of automated assessment tools leveraging image analysis, machine learning, and artificial intelligence7. The development of automated Ki67 proliferation index scoring tools holds considerable promise for significantly enhancing pathology workflow efficiency, shortening report turnaround times, reducing inter-observer variability, and ultimately improving diagnostic accuracy
Currently, deep learning (DL)-based methods have significantly enhanced the performance of automated Ki67 PI quantification, demonstrating exceptional results across diverse application scenarios. Shaker et al.8 employed digital image analysis (DIA) to assess Ki67 immunohistochemistry on pancreatic neuroendocrine tumor (PanNET) cell blocks. Their results revealed a Pearson correlation coefficient of 0.9681 and a Cohen’s Kappa coefficient of 0.862 between DIA and manual pathologist scoring, indicating high accuracy in Ki67 index estimation. However, the method remained heavily reliant on pathologist confirmation. Vesterinen et al.9 developed a deep learning-based Ki67 Automatic Index (KAI) algorithm for objective Ki67 index calculation in neuroendocrine tumors (NETs). This approach achieved a concordance level of 0.89 with pathologist assessment, with results fully matching in 46% of the tumors analyzed. Govind et al.10 introduced novel computational tools, the Synaptophysin-KI-Estimator (SKIE) and deep-SKIE, based on dual-staining (DS) for Ki67 and synaptophysin to improve quantification accuracy in gastrointestinal neuroendocrine tumors (GI-NETs). Experimental results demonstrated a 90% grade concordance between SKIE and the Gold Standard (GS). Furthermore, deep-SKIE achieved a test accuracy of 91.0% on DS WSIs, significantly outperforming its performance on single-stained (SS) WSIs. In parallel, Fulawka et al.11 proposed an integrated approach combining deep learning with fuzzy set-based interpretation to approximate the Ki67 PI by detecting hotspot regions and segmenting relevant cells. This methodology culminated in a mean absolute error (MAE) of 0.024 during three-fold cross-validation, significantly enhancing the precision of Ki67 index calculation.
Furthermore, Karol et al.12 proposed a novel two-step concept based on a universal deep learning model. By detecting Ki67/lymphocyte classification on Ki67 stained slides, their approach demonstrated the competitiveness of common architectures in instance segmentation and object detection. Rewcastle et al.13 compared the prognostic value and reproducibility of four automated DIA methods and two manual methods for Ki67 scoring. By analyzing Ki67 scores from 367 patients with hormone receptor-positive, HER2-negative, node-negative breast cancer, they assessed the ICC and prognostic correlation of the different methods. They ultimately found that DIA scoring, using a 14% cutoff, provided superior prediction of distant metastasis-free survival compared to manual scoring, although further research is needed to standardize DIA scoring definitions and clinical cutoffs. Aniq et al.14 introduced a deep learning-based method for calculating the Ki67 proliferation index. Utilizing classical image processing techniques applied to 40\(\times\) magnification tumor region images provided by pathologists, their approach segments and counts positive cells, negative cells, and tumor-infiltrating lymphocytes (TILs), ultimately calculating the percentage of Ki67 labeled cells. Phan et al.15 presented an innovative deep learning-based method utilizing advanced neural network architectures including DeepLabv3+, MobileNet-v2, Xception, DenseNet-121, U-Net, and fully convolutional regression networks. This method analyzes Ki67 protein expression in biopsy samples, precisely distinguishing Ki67 positive and negative tumor cells while also detecting TILs. Rigorously evaluated on the SHIDC-B-Ki-67 dataset, the method achieved an accuracy of 98.8% with a processing time of 13 seconds, thereby significantly enhancing the efficiency of breast cancer detection and the timeliness of clinical decision-making. Zehra et al.16 evaluated the performance of the open-source deep learning cloud platform DeepLIIF for quantifying Ki67 expression in gastrointestinal neuroendocrine tumors (GI-NETs) compared to manual quantification. Their results demonstrated that DeepLIIF achieved a high ICC of 0.885, indicating high accuracy in Ki67 quantification. Furthermore, it significantly reduced analysis time and exhibited strong intraobserver and interobserver variability agreement, offering a reliable tool for Ki67 assessment in high-throughput clinical workflows.
In parallel, Negahbani et al.17 introduced the SHIDC-BC-Ki-67 dataset alongside PathoNet, a novel U-Net framework-based post-processing pipeline, for the automated assessment of Ki67 expression and intratumoral lymphocyte infiltration scores in breast cancer cells. Further advancing methodology, Geread et al.18 proposed piNET, a tool centered on U-Net architecture. By mimicking the pathologist’s workflow and employing weakly supervised learning for background region identification and hotspot analysis, piNET significantly enhanced Ki67 quantification performance, achieving an accuracy rate of 86% and a correlation coefficient of 0.927.
Despite significant advancements in automated Ki67 quantification methodologies, their application to breast cancer histopathology images remains challenged by several critical issues. Firstly, the inherent polymorphism of tumor cells renders conventional convolutional operations with fixed receptive fields inadequate for adaptive modeling. Secondly, variations in immunohistochemical (IHC) staining protocols induce fluctuations in positive signal intensity, significantly compromising the robustness of feature extraction. Thirdly, the spatially interwoven distribution of highly proliferative zones and quiescent regions necessitates concurrent model capability for capturing both local cellular details and global tissue architecture patterns. Fourthly, cell cluster adhesion is prevalent in > 40% of invasive carcinoma samples, resulting in ambiguous instance boundaries. This leads to decreased precision in overlapping cell boundary prediction and increased calculation bias for the PI. To address these limitations, we propose Kpi-Net, a novel deep learning model based on a U-Net framework. Kpi-Net integrates four key components: a RDMS Module to enhance multi-receptive field feature capture; Transformer Blocks to augment global contextual modeling capacity; an improved HS-CBAM-FPN attention fusion mechanism for dynamic screening of critical channel and spatial information; and an optimized watershed algorithm to improve segmentation accuracy of adherent cells. Collectively, these innovations are designed to overcome the deficiencies observed in existing methods. The primary contributions of this work are summarized as follows:
-
Innovative Residual Dilated Multi Scale Module: We innovatively design a RDMS Module. This module organizes a multi-branch dilated convolutional architecture, where each branch covers distinct receptive fields ranging from fine local structures to broader regions. It integrates residual connections to effectively mitigate training difficulties. Furthermore, lightweight Transformer Blocks are incorporated to enhance the model’s capacity for global contextual modeling of complex spatial dependencies among cells across the entire histopathology whole-slide image. This integrated approach effectively mitigates problems of missed detections and false positives arising from heterogeneous cell distributions.
-
Proposed High-level Screening-Convolutional Block Attention Module Feature Pyramid Networks: Building upon the hierarchical structure of HS-FPN, we propose the HS-CBAM-FPN. Its core innovation lies in the integration of Convolutional Block Attention Modules (CBAM), comprising channel attention and spatial attention mechanisms. This integration enables the dynamic selection and amplification of critical feature channels and spatial regions highly relevant for Ki67 cell recognition during the feature fusion process. Concurrently, this mechanism synergizes with a Selective Feature Fusion strategy (SFF-CBAM) to facilitate smarter and more efficient refinement and fusion of multi scale, multi-level features extracted by the encoder at different stages. This process yields more discriminative feature representations for subsequent segmentation tasks.
-
Optimized Watershed Post-processing for Cell Instance Segmentation: Following the model’s output of a cell probability map, we integrate an optimized watershed algorithm as a post-processing step for cell instance segmentation and counting. Utilizing distance transformation and local maxima detection, the watershed algorithm enables significantly more accurate delineation of boundaries between adherent or overlapping cells. This substantially mitigates mis-segmentation issues prevalent in densely packed cell regions and overlapping clusters. Ultimately, this ensures more precise separation and counting of individual Ki67 positive and negative cells, thereby enhancing the calculation accuracy of the Ki67 index and reducing resultant errors.
-
Comprehensive Validation on Diverse Public Datasets: We conduct extensive and rigorous experimental evaluations using publicly available breast cancer histopathology datasets–SHIDC-BC-Ki-67, BCData, and DeepSlides–specifically curated to encompass diverse resolutions and IHC staining conditions. The experimental results robustly validate Kpi-Net’s high applicability in real-world clinical pathology analysis scenarios, its strong robustness against variances, and its superior performance under complex challenges.
Related work
Within the field of artificial intelligence, deep neural networks, particularly Convolutional Neural Networks (CNNs), have demonstrated immense potential in medical image processing, achieving significant advances specifically in the automated analysis of pathological images. U-Net19 stands as the most widely adopted network architecture for medical image segmentation. Its encoder-decoder structure effectively captures image features. However, as task complexity and demands for precision increase, U-Net’s successive pooling and convolution operations inevitably lead to the partial loss of spatial information. Consequently, numerous researchers have focused on improving the encoder-decoder framework. Gu et al.20 proposed the Context Encoder Network (CE-Net) to capture richer high-level information while preserving spatial details for 2D medical image segmentation, thereby enhancing the model’s semantic representation capability. Zhou et al.21 developed the Nested U-Net (U-Net++), a structure for medical image segmentation that incorporates ResNet architecture and dense skip connections. By linking each decoder layer’s output to all higher-level features in the encoder, the decoder gains more comprehensive semantic information from the encoder, leading to improved segmentation accuracy and finer detail delineation.
Furthermore, attention mechanisms and multi scale feature fusion techniques have been extensively applied in medical image segmentation tasks. The Convolutional Block Attention Module (CBAM)22 integrates channel attention with spatial attention. When integrated with CNN models, it provides more comprehensive and effective feature extraction capabilities. DeepLabV3+23, inspired by Spatial Pyramid Pooling (SPP), innovates the Atrous Spatial Pyramid Pooling (ASPP) module. It incorporates an Inception-like structure containing Atrous Convolutions with different dilation rates before pixel classification, strengthening the model’s capacity to extract multi scale information. To overcome the limitations of traditional methods when processing images with scarce features and large scale variations, Chen et al.24 proposed the MFDS-DETR method, which includes a High-order Screened Feature Pyramid Network (HS-FPN). This approach enhances global feature extraction through multi-level feature fusion and multi scale deformable self-attention modules.
Liu et al.25 introduced a multi scale Fusion Gene Recognition Network (MultiGeneNet). By leveraging multi scale information within pathological images, this network improves the accuracy of predicting isocitrate dehydrogenase 1 (IDH1) gene mutation status. Evaluated on a dataset comprising hematoxylin and HE -stained pathological slides from 296 patients, it achieved an accuracy of 83.575% and an AUC of 0.886. Eren et al.26 developed DeepCAN, a modular deep learning system for automated cell counting and viability analysis. The system integrates three neural network modules: the Parallel Segmenter, Cluster CNN, and Viability CNN, designed for initial segmentation, cluster separation, and viability classification, respectively. Experimental results demonstrated that DeepCAN attained high precision in cell segmentation and viability analysis across multiple cell types. Finally, Zhao et al.27 presented a novel medical image segmentation method called the multi scale Subtraction Network (\(M^2\)SNet). By designing fundamental Subtraction Units (SUs) and enabling multi-level, multi scale feature aggregation, \(M^2\)SNet effectively reduces redundant information while enhancing feature complementarity. Simultaneously, it utilizes a training-free LossNet module for comprehensive supervision. The method demonstrates superior performance across diverse medical imaging modalities and segmentation tasks.
In recent years, Transformers have demonstrated robust capabilities within the field of image processing, particularly excelling at capturing global context and modeling long-range dependencies. This advancement has provided novel solutions for medical image analysis. Building upon this foundation, Dosovitskiy et al.28 successfully adapted the Vision Transformer (ViT) for image classification tasks, establishing its potential for medical image analysis. Further advancing this development, Chen et al.29 integrated the Transformer into medical image segmentation, proposing TransUNet. This model substantially improved the global context modeling capability inherent in medical image segmentation tasks.
TransAttUnet30, another network based on the Transformer framework, incorporates a multi-level guided attention mechanism and multi scale skip connections. This architecture effectively learns non-local interactions among encoder features and aggregates upsampled features across different semantic scales, resulting in significant performance improvements for medical image segmentation. However, a key limitation of the Transformer architecture remains its reliance on global self-attention mechanisms, which incur substantial memory consumption and computational cost.
To address challenges in biomedical analysis, Prangemeier et al.31 proposed Cell-DETR, a novel attention-based Transformer for direct end-to-end instance segmentation. Cell-DETR is particularly suited for object instance detection and segmentation tasks in biomedical applications, such as lesion detection, tumor identification, and single-cell information extraction from microscopic images. Complementing these developments, Xia et al.32 introduced ViT-CoMer, a streamlined, pre-training-free, and feature-enhanced Vision Transformer backbone network. By incorporating convolutional multi scale feature interactions, ViT-CoMer bridges the gap between CNNs and Transformers, establishing a new backbone architecture suitable for dense prediction tasks.
Building upon the aforementioned research, existing approaches for breast cancer Ki67 quantification still exhibit limitations, including inadequate utilization of multi scale features, constrained global context modeling capability, and poor adaptability to complex staining variations. To address these shortcomings, this paper proposes the Kpi-Net model, achieving performance breakthroughs through multi-dimensional innovations.
Within Kpi-Net, the RDMS employs multi-branch dilated convolutions to dynamically adjust the receptive field. This is combined with residual connections to mitigate the vanishing gradient problem. The RDMS effectively overcomes the difficulties faced by existing U-Net variants in balancing the extraction of local cellular morphology and global tissue patterns, thereby reducing the missing detection rate. Concurrently, Kpi-Net incorporates a lightweight Transformer Block. Utilizing multi-head self-attention, this block efficiently models spatial dependencies among cells. Crucially, it achieves an optimal balance between parameter size and performance: it alleviates the locality bias limitations of traditional CNNs for localizing sparsely distributed cells while circumventing the high computational cost inherent to standard Transformers. The enhanced HS-CBAM-FPN performs multi scale feature fusion. It dynamically filters key feature channels relevant to Ki67 cells and integrates the CBAM to focus on positive cell regions. Its SFF-CBAM further effectively suppresses interference caused by staining heterogeneity. Moreover, to tackle the challenge of cell cluster adhesion, Kpi-Net incorporates an optimized watershed algorithm. Leveraging precise distance transforms and local maxima detection, this component minimizes prediction errors arising from overlapping cell boundaries. Experimental results demonstrate that Kpi-Net achieves superior overall performance metrics compared to mainstream models like PathoNet and TransAttUnet on public datasets, exhibiting particularly enhanced robustness in complex pathological scenarios.
Methodology
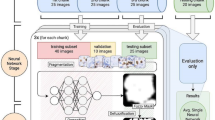
Kpi-Net is a multi scale deep learning model specifically designed for Ki67 quantification tasks in breast cancer histopathological images. Figure 1 illustrates its framework for automated Ki67 PI computation. In clinical practice, histopathological slides are first digitized into WSIs. Pathologists then select several regions of interest (ROI) from Ki67-stained WSIs to calculate the positive rate of tumor cells. Following the pathologist’s ROI selection, Kpi-Net facilitates the automated detection of Ki67 positive (\(Ki67^+\)) and Ki67 negative (\(Ki67^-\)) cells, thereby enabling fully automated PI computation.
Automated proliferation index framework of Kpi-Net. The workflow initiates with the selection of ROI from digitized WSI. These ROI undergo feature extraction and cellular detection by Kpi-Net, enabling quantification of both \(Ki67^+\) and \(Ki67^-\) cells. This process culminates in the automated calculation of the PI.
Kpi-Net network architecture
The overall architecture of Kpi-Net consists of three integral components: an encoder, a decoder, and a post-processing module, culminating in automated PI computation. Through multi-level feature extraction and fusion, the model progressively extracts rich semantic information from input images, outputs cellular segmentation and classification results, with subsequent post-processing refining the final PI calculation. Figure 2 schematically illustrates the network architecture of Kpi-Net, while detailed functionalities of each constituent module are elaborated in subsequent sections.
Kpi-Net model architecture.
Encoder
The input image is initially processed through two consecutive 3\(\times\)3 convolutional layers, each followed by BatchNorm and a LeakyReLU activation function, to extract preliminary low-level features. Subsequently, a 2\(\times\)2 max-pooling layer is applied for downsampling, reducing the spatial dimensions of the feature maps. The core of the encoder comprises four RDMS Modules. Each module extracts multi scale features through parallel dilated convolutional branches while preserving original input information via residual connections.
The output feature map of each RDMS Module is flattened into a sequence and fed into a Transformer Block. This block employs a multi-head self-attention mechanism to capture long-range dependencies within the feature sequence, with features further enhanced through a feed-forward neural network (FFN). The processed sequence is then reshaped via max-pooling into a feature map and transferred to the subsequent convolutional layer. In the final encoder stage, feature maps from all hierarchical levels (Stage 2 to Stage 5) are input to the HS-CBAM-FPN module. This module first utilizes the CBAM to screen and enhance features (P2–P5). Through the SFF-CBAM mechanism, it fuses low-level and high-level features to generate multi scale fused features (N2–N5), thereby providing rich semantic information for the decoder.
Decoder
Following multi-level feature extraction in the encoder, the decoder progressively restores image resolution through upsampling and feature fusion, ultimately generating a per-pixel categorical probability map. At each upsampling level, features are concatenated with their corresponding-level feature maps from the encoder and multi scale features (N2–N5) derived from the HS-CBAM-FPN module. This integration preserves fine-grained spatial details while enriching semantic context. The concatenated feature maps then undergo further feature extraction and fusion via the RDMS Module, enabling simultaneous capture of local cellular details and global contextual information. In the final decoder stage, after iterative upsampling and feature refinement, the feature map undergoes three successive 3\(\times\)3 convolutional layers for detail enhancement. This culminates in a 1\(\times\)1 convolutional layer that outputs the per-pixel categorical probability map. A Softmax activation function is applied to the output layer, guaranteeing that the sum of category probabilities for each pixel equals 1.
Post-processing stage
In the post-processing stage, we employed a watershed algorithm to refine cell segmentation and localization. First, Gaussian filtering and morphological operations were applied to the class probability map to eliminate noise and fill holes. Subsequently, the Euclidean distance transform (EDT) was computed to generate a distance map, followed by local maxima detection to identify target centers. These seed points were used to create marker maps, which–combined with the distance map and probability map–served as inputs for the watershed algorithm to achieve precise segmentation. Post-segmentation, small regions were filtered out, and the minimum enclosing circle was calculated for each remaining region. The final output included the cell’s centroid coordinates (x, y) and class label (ch), where (x, y) denotes the geometric center of the cell and ch represents its classification (\(Ki67^+\) and \(Ki67^-\)).
Residual dilated multi scale module
Accurate feature extraction in cell detection, classification, and counting tasks presents significant challenges due to complex morphological characteristics, highly heterogeneous staining patterns, and frequent cell overlapping. While capturing multiscale contextual information is crucial for enhancing pathological image analysis performance, conventional CNNs suffer from inherent limitations: their fixed receptive fields struggle to simultaneously preserve local details and global structures, while progressive spatial resolution degradation with network depth leads to critical detail loss.
To address these issues, we designed the RDMS Module, which employs a multi-branch dilated convolution architecture. The core innovation of RDMS lies in its four parallel convolutional branches, each configured with specific dilation rates(d) to capture distinct-scale features: (1) Branch A1 uses standard 3\(\times\)3 convolution (d = 1, 3\(\times\)3 receptive field) for local cellular morphology; (2) Branch A4 (d = 4, 9\(\times\)9 receptive field) captures inter-cellular relationships; (3) Branch A6 (d = 6, 13\(\times\)13 receptive field) perceives cell cluster structures; and (4) Branch A12 (d = 12, 25\(\times\)25 receptive field) models global tissue distribution. This multiscale design, while inspired by ASPP, optimizes the dilation rate combination. As shown in Fig. 3, RDMS adopts a stepped dilation scheme that avoids receptive field overlap from small dilations (1-3) while balancing local-global coverage and computational efficiency. Ablation studies presents ablation studies validating this configuration. In the encoder pathway (Fig. 3a), RDMS implements feature reuse through a four-branch parallel structure (d = 1/4/6/12). For the decoder pathway (Fig. 3b), RDMS employs feature compression: concatenated branch outputs undergo 1\(\times\)1 convolution for channel reduction. This strategy maintains computational efficiency while ensuring effective feature fusion and progressive spatial resolution recovery.
Architecture of the residual dilated multi scale module. (a) Encoder path: Input features are directly concatenated with outputs from four parallel branches. (b) Decoder path: Concatenated multi-branch outputs undergo dimensionality reduction via 1\(\times\)1 convolution.
Transformer block
Histopathological images exhibit pronounced spatial heterogeneity in cellular distribution, with cell overlap rates reaching up to 65% in dense regions while dropping below 5% in sparse areas. Although the RDMS module extracts local features through multiscale convolutions, it still suffers from an 18% false-negative rate in low-density regions. To address this limitation, we integrate the Transformer Block - a core component of Transformer architecture - at each encoder stage as an effective attention mechanism to enhance global context modeling.
As illustrated in Fig. 4, the incorporated Transformer Block establishes long-range dependencies among cells through multi-head self-attention, effectively capturing contextual information across the entire input sequence. The subsequent FFN further refines these features through representation learning. While Transformer architectures typically incur substantial computational overhead, Ablation Studies provides comprehensive analysis of the performance-resource trade-off. Experimental results demonstrate that the performance gains justify the module’s inclusion, with the achieved algorithmic improvement validating both the necessity and cost-effectiveness of this design.
Transformer block.
The input to the Transformer Block is a feature sequence \(X \in \mathbb {R}^{N \times D}\), where the feature sequence first undergoes linear transformations to generate query (Q), key (K), and value (V) matrices. Each attention head computes the attention weights independently and generates the corresponding output headi through a weighted sum, as shown in Eq. (1).
Here,\(Q_i \in \mathbb {R}^{N \times d_k}\) represents the query matrix for the i-th attention head, \(K_i \in \mathbb {R}^{N \times d_k}\) denotes the key matrix for the i-th attention head, \(V_i \in \mathbb {R}^{N \times d_v}\) signifies the value matrix for the i-th attention head, N indicates the length of the input sequence, \(d_k\) is the dimensionality of queries and keys, and \(d_v\) is the dimensionality of values.\(Q_i K_i^T \in \mathbb {R}^{N \times N}\) represents the dot product of the query matrix and key matrix, which is then normalized by the softmax function into a probability distribution to produce the attention weight matrix. The outputs from multiple attention heads are concatenated and subjected to a linear transformation to derive the final output of the multi-head attention mechanism, as illustrated in Eq. (2).
The output of the multi-head attention mechanism is combined with the input feature sequence through residual connections, and then layer normalization is applied to yield \(Y_{\text {att}}\). \(Y_{\text {att}}\), after layer normalization, is passed through a two-layer fully connected network for non-linear transformation, resulting in FFN(\(Y_{\text {att}}\)). The output of the feed-forward neural network is combined with \(Y_{\text {att}}\) through residual connections, followed by layer normalization. The final output is the feature sequence \(Y \in \mathbb {R}^{N \times D}\), enriched through global context modeling and feature enhancement. The complete workflow is depicted in Eqs. (3)–(5).
High-level screening-convolutional block attention module feature pyramid networks
For cell detection tasks, conventional feature pyramid networks (e.g., FPN) often integrate multi scale features via basic upsampling and element-wise summation. Nevertheless, this method exhibits notable drawbacks in handling cell images: firstly, the feature information in cell images tends to be sparse, hindering the model’s ability to precisely identify targets;secondly, the size differences between various cell types are significant, and cells of the same type may display varying sizes under different microscopes, exacerbating detection challenges due to their multi scale characteristics. To tackle these challenges, this study adopts the design principles of the HS-FPN module. HS-FPN improves the model’s feature representation by incorporating a channel attention mechanism to select low-level features and integrating them with high-level features. Given that the Ki67 cell counting task in this research shares similar technical challenges with cell detection–both involving multi scale features and complex background information in pathological images–we integrate the HS-FPN module into our model, utilizing its multi scale feature fusion strategy to achieve precise detection and classification of pathological images. To further boost model performance, we refine the original HS-FPN module by substituting the CA attention module with the superior CBAM, introducing the improved HS-CBAM-FPN. The CBAM module, which integrates both channel attention and spatial attention mechanisms, enables a more thorough extraction of critical feature information from cell images.
As shown in Fig. 5, the HS-CBAM-FPN model comprises two primary parts: the feature selection module, which filters and selects multi scale features. The Top–down feature selection module integrates the filtered features.
High-level screening-convolutional block attention module feature pyramid networks.
Feature selection HS-CBAM-FPN takes feature maps from each encoder layer as input (S2–S5), with \(fin \in \mathbb {R}^{C \times H \times W}\), where C denotes the number of channels, H represents the height, and W signifies the width of the feature map. The feature map is initially processed by the CBAM module, comprising channel attention and spatial attention components. Initially, the channel attention operation is executed, applying global average pooling and global max pooling to the input feature map fin to derive two feature vectors. These feature vectors are processed by a shared multilayer perceptron (MLP) and activated using the Sigmoid function to produce channel weights \(M_c \in \mathbb {R}^{C \times 1 \times 1}\). The channel weights \(M_c\) are element-wise multiplied with the input feature map fin to yield the channel attention-enhanced feature map \(f_{\text {channel}}\). Subsequently, the spatial attention operation is conducted, applying average pooling and max pooling to the channel attention-enhanced feature map \(f_{\text {channel}}\) across the channel dimension to generate two feature maps. These two feature maps are combined and processed by a 7\(\times\)7 convolutional layer to produce spatial attention weights \(M_s \in \mathbb {R}^{1 \times H \times W}\).
The spatial attention weights \(M_s\) are multiplied element-wise with the channel attention-enhanced feature map \(f_{\text {channel }}\) to derive the final feature map \(f_{\text {CBAM}}\). To align the feature map’s channel count with the dimensions required for subsequent operations, a 1\(\times\)1 convolution is performed on the CBAM module’s output \(f_{\text {CBAM}}\), producing the feature map \(f_{1 \times 1} \in \mathbb {R}^{256 \times H \times W}\). To retain original feature information and prevent gradient vanishing, a residual connection is incorporated, adding the 1\(\times\)1 convolution output \(f_{1 \times 1}\) element-wise to the input feature map fin to yield the final feature map \(f_{\text {out}}\). The final output feature map \(f_{\text {out}} \in \mathbb {R}^{256 \times H \times W}\) encompasses the selected critical feature information while preserving the original feature details. These feature maps (P2–P5) are suitable for further multi scale feature integration.
Top–down feature selection The Top–down feature selection’s main objective is to combine the selected low-level features with high-level features, producing multi scale fused features. Conventional feature fusion approaches often involve direct element-wise summation of low-level and high-level features. Although straightforward, this method lacks the ability to effectively select and combine critical feature information. To tackle this challenge, this research enhances the SFF module by incorporating the CBAM attention mechanism, resulting in an advanced Top–down feature selection named SFF-CBAM. The central concept of the SFF-CBAM module is to leverage the semantic information from high-level features as weights, enabling dynamic selection of crucial information from low-level features. In detail, high-level features contain abundant semantic information but exhibit lower spatial resolution, whereas low-level features offer higher spatial resolution but limited semantic information. By utilizing the CBAM attention mechanism, the SFF-CBAM module produces channel attention and spatial attention weights, dynamically modulating the weights of low-level features to preserve task-critical information and filter out irrelevant details, leading to enhanced feature fusion.
Initially, the high-level features \(f_{\text {high}}\) are upsampled to match the spatial resolution of the low-level features \(f_{\text {low}}\). Upsampling is performed using transposed convolution and bilinear interpolation, yielding the upsampled high-level features \(f_{\text {att}}\). The upsampled high-level features \(f_{\text {att}}\) are processed by the CBAM module to produce attention weights, selecting significant high-level feature information and resulting in \(f_{\text {att}}^{\text {CBAM}}\). The CBAM-generated attention weights are applied to the low-level features flow to obtain the filtered low-level features \(f_{\text {filtered}}\). The filtered low-level features \(f_{\text {filtered}}\) are combined with the high-level features \(f_{\text {att}}^{\text {CBAM}}\) through element-wise addition, with \(f_{\text {att}}^{\text {CBAM}}\) contributing rich semantic information and \(f_{\text {filtered}}\) supplying detailed information. The resulting multi scale fused features \(f_{\text {final}}\) are generated. The final output feature map \(f_{\text {final}} \in \mathbb {R}^{C' \times H_1 \times W_1}\) encompasses multi scale feature information, integrated with the upsampled results in the encoder to improve feature representation. The complete workflow is outlined in Eqs. (6)–(9).
Here, BL denotes the bilinear interpolation operation, T-Conv represents the transposed convolution operation, \(W_{\text {channel}}\) indicates the channel attention weights, and \(W_{\text {spatial}}\) signifies the spatial attention weights.
Within the decoder, the HS-CBAM-FPN output feature map (N2–N5) \(f_{\text {final}}\) undergoes stepwise upsampling and is concatenated with the corresponding encoder-level feature maps, aiding in the recovery of spatial resolution and detail information, which improves the model’s detection and classification capabilities.
Watershed algorithm
Following the model’s final output, a post-processing step utilizing the Watershed Algorithm is implemented to enhance cell segmentation and localization. The Watershed Algorithm, a morphological image segmentation technique, is adept at managing overlapping objects and intricate backgrounds in images. The following outlines the detailed post-processing workflow.
The model’s output is \(\text {pred} \in \mathbb {R}^{H \times W \times 2}\), with H and W denoting the image’s height and width, and the two channels representing probability maps for distinct classes.
The first channel (channel 0) represents the probability map for \(Ki67^+\), while the second channel (channel 1) encodes the probability map for \(Ki67^-\). These dual-channel probability maps facilitate precise quantification of cellular proliferation states through subsequent pixel-wise classification and counting operations.
Initially, Gaussian filtering is performed on each channel’s probability map to mitigate noise. Subsequently, morphological operations are employed to eliminate noise and fill gaps: opening eliminates minor noise, and closing addresses small holes within target areas.
Following this, the EDT is applied to each channel’s probability map to produce the distance map D. The distance transform assigns each pixel a value based on its distance to the nearest background pixel, emphasizing the central areas of targets. For subsequent processing, the distance map D is normalized to a range of [0, 255]. Local maxima points, often representing target centers, are identified on the normalized distance map. The density of local maxima is regulated by tuning the \(\text {min\_distance}\) parameter, preventing over-segmentation. The detected local maxima points serve as initial markers, and connected component analysis is utilized to create the marker map markers. The Watershed Algorithm is employed for segmentation, using the negative distance map D as input, combined with the marker map markers and the probability map gray as masks. By simulating water flow from high to low elevations, the Watershed Algorithm segments the image into distinct regions, treating it as a topographic map.
Post-segmentation, the labels produced by the Watershed Algorithm are processed to extract each independent region. Regions with areas below a threshold (e.g., 50 pixels) are filtered to eliminate noise or segmentation inaccuracies. The minimum enclosing circle is computed for each retained region, with its center coordinates designated as the cell’s center. The output includes the center coordinates and category of each cell in the format [x, y, ch], with x and y indicating the center and ch representing the class.
Experiments and results analysis
Datasets
This study employed three breast cancer histopathology image datasets for model validation: SHIDC-BC-Ki-67, BCData, and DeepSlides.
The SHIDC-BC-Ki-6717 dataset, meticulously annotated by pathologists, comprises 1,656 training images and 701 test images collected between 2017 and 2020. All samples represent invasive ductal carcinoma cases with a resolution of 256\(\times\)256 pixels, containing annotations for three cell types: Ki67-positive tumor cells, Ki67-negative tumor cells, and TILs. For the purpose of Ki67 index evaluation, only the first two categories were utilized in our experiments.
The BCData33 dataset contains 936 training images and 402 test images (640\(\times\)640 pixels resolution) with 181,074 annotated cells. This dataset exclusively classifies cells into positive and negative tumor cell categories, presenting significant challenges including pronounced cell clustering/overlapping and substantial staining condition variations, thereby providing a more rigorous test environment for model training.
The DeepSlides34 dataset was extracted from whole-slide images of 32 breast cancer patients, consisting of 649 Ki67 region-of-interest images (513\(\times\)513 pixels resolution). While lacking annotation files, this dataset served to validate the model’s effectiveness in Ki67 index calculation.
To provide a clear comparison of the distinct characteristics and complementary nature of these three datasets, a summary is presented in Table 1. The integration of these three datasets with distinct characteristics enabled comprehensive evaluation of the model’s performance across varying resolutions and staining conditions for cell classification and tumor grading tasks, thereby effectively verifying the model’s generalizability and robustness.
Experimental environment
This study uses Python 3.10 and the PyTorch 2.1.2 deep learning framework to implement the construction, training, and testing of the Kpi-Net model. Experiments were conducted on a high-performance workstation equipped with a 16 vCPU Intel Xeon Gold 6430, an NVIDIA RTX 4090 GPU (24 GB VRAM), and 120 GB of RAM, running the Ubuntu 22.04 operating system. The batch size was set to 8, the backbone learning rate to \(1 \times 10^{-3}\), and the CosineAnnealingLR scheduler was used to gradually decrease the learning rate from the initial value of \(1 \times 10^{-3}\) to a minimum of \(1 \times 10^{-5}\) following a cosine function. The Adam optimizer was used, with the momentum (\(\beta _1\)) and second-order momentum (\(\beta _2\)) set to 0.9 and 0.999, respectively. The training process ran for 100 epochs, and the best model was saved at the end of each epoch based on the loss value for subsequent segmentation testing.
Evaluation metrics
This study adopted six evaluation metrics to comprehensively validate the nuclear detection performance and Ki67 PI quantification accuracy of the Kpi-Net model. For nuclear detection performance, three metrics were used: Precision, Recall, and F1 Score, as shown in Eqs. (10)–(12), where TP, FP, and FN denote true positives, false positives, and false negatives, respectively.
The Precision metric primarily evaluates the proportion of correctly predicted positive samples among all predicted positives, reflecting the model’s accuracy in predicting positive-class samples. Recall measures the proportion of actual positive samples correctly identified by the model, indicating its ability to capture positive samples. The F1 Score, as the harmonic mean of Precision and Recall, balances the trade-off between these two metrics. It is particularly suitable for nuclear detection tasks, where sensitivity and precision must be balanced. A higher F1 Score indicates better overall performance in nuclear detection tasks.
Additionally, this study employed RMSE, Pearson correlation coefficient, and PI classification accuracy as metrics to assess the precision of PI estimation. In this process, PI values were categorized into low, medium, and high ranges. The classification accuracy across these different intervals was calculated to evaluate the accuracy of PI estimation results, as shown in Eq. (13).
Among these metrics, RMSE was utilized to quantify the discrepancy between predicted and actual values. A smaller RMSE value indicates higher prediction accuracy. The mathematical formulation is presented in Eq. (14), where \(PI_{true}\) represents the actual PI value, \(PI_{pred}\) denotes the predicted PI value, and N refers to the sample size.
The Pearson correlation coefficient was employed to evaluate the linear relationship between predicted and actual values, where \(\overline{\text {PI}_{\text {pred}}}\) represents the mean of predicted PI values and \(\overline{\text {PI}_{\text {true}}}\) denotes the mean of actual PI values. An R-value closer to 1 indicates stronger agreement between model predictions and ground truth data, as specified in Eq. (15).
In summary, this study evaluated the model not only through precision, recall, and F1 score, but also further validated its performance using RMSE, R, and PI classification accuracy. While the F1 score primarily assesses the raw performance of nuclear detection, it can be significantly affected by scenarios with limited tumor nuclei counts or outlier presence. Therefore, additional metrics including PI classification accuracy and Pearson correlation coefficient provide a more comprehensive reflection of the model’s accuracy and robustness in practical applications.
The SHIDC-BC-Ki-67 dataset explicitly defines PI ranges as: low proliferation (Ki67 score < 16%), intermediate proliferation (16–30%), and high proliferation (> 30%). For BCData and DeepSlides datasets, the PI ranges were defined according to literature35 as low (< 10%), intermediate (10–30%), and high (> 30%). Based on this analysis, these evaluation metrics will be collectively analyzed throughout the paper to ensure a comprehensive assessment of model performance.
Comparison with other classical methods
To assess the effectiveness and generalizability of the Kpi-Net model, this study conducted training and validation on the SHIDC-BC-Ki-67 and BCData datasets, followed by independent testing on DeepSlides to evaluate its generalization capability. This approach enables comprehensive assessment of the model’s adaptability across different data distributions, ensuring not only its validity on training data but also rigorously testing its adaptability and robustness under varying data conditions, thereby achieving a complete validation of Kpi-Net universal applicability.
Experimental results on the SHIDC-BC-Ki-67 dataset
Table 2 presents the performance comparison between the Kpi-Net model and existing mainstream models U-Net19, UNet++21, PathoNet17, TransUNet29, and TransAttUnet30 on the SHIDC-BC-Ki-67 dataset. The results demonstrate that Kpi-Net outperforms all comparative models across various evaluation metrics for both Ki67 positive and negative cell classification tasks, validating its superiority in Ki67 counting applications.
For Ki67+ classification, Kpi-Net achieved superior performance with 89.45% accuracy, 90.95% recall, and 90.19% F1-score, surpassing all comparison models. Specifically, compared to TransAttUnet, Kpi-Net showed improvements of 3.26% in accuracy, 1.85% in recall, and 2.46% in F1-score. In Ki67- classification tasks, the model attained 83.67% accuracy, 85.23% recall, and 84.99% F1-score, representing a 9.79% accuracy improvement and 6.29% F1-score enhancement over the baseline U-Net model. Notably, Kpi-Net maintained a 2.13% F1-score advantage over the robust TransAttUnet model, confirming its enhanced capability in identifying highly proliferative cells. Regarding overall performance, Kpi-Net achieved an average F1-score of 85.79%, exceeding all existing mainstream models. This represents a 7.16% improvement over the baseline U-Net and a 6.83% enhancement compared to its advanced variant UNet++. Furthermore, Kpi-Net maintained performance advantages of 1.32% and 2.10% over the state-of-the-art Transformer-based models TransAttUnet and TransUNet, respectively. These results substantiate that the proposed multi scale fusion strategy effectively integrates feature information while reducing both false positives and false negatives in cell classification.
As demonstrated in Table 3, Kpi-Net achieves superior performance in Ki67 index prediction tasks, with the RMSE value reduced by 1.13% compared to TransAttUnet, while maintaining a high Pearson correlation coefficient (R = 0.965) and PI classification accuracy (95.66%), comparable to PathoNet and TransUNet. Although all models exhibit PI accuracy above 95% with a maximum deviation of only 0.1%, significant variations exist in RMSE and R values. For instance, U-Net shows merely a 0.1% difference in PI accuracy compared to Kpi-Net, yet its RMSE is 11.03% higher.
This phenomenon stems from the ratio-based nature of PI (PI = number of positive cells / total cells). When prediction biases across different models exhibit directional consistency (systematically under- or over-estimating positive cells), the resulting PI ratios may remain similar. However, Kpi-Net’s multi scale feature fusion strategy not only maintains high PI accuracy but also reduces RMSE to 0.04632 and improves R to 0.965. Compared to models optimizing solely for overall ratio accuracy, Kpi-Net demonstrates dual advantages in both quantitative precision and reliability.
This enhancement in quantitative precision and reliability holds significant clinical relevance. A lower RMSE indicates more accurate PI value prediction for individual cases, thereby reducing the risk of misclassifying patients whose PI values lie near critical clinical decision thresholds . A higher R signifies better consistency between the model’s predictions and pathologist-derived ground truth assessments across a variety of cases. Most importantly, the improved segmentation quality and model robustness directly translate to enhanced reproducibility and reliability ineal-world pathological workflows. This mitigates the inter-observer and intra-observer variability inherent in manual assessment, increasing confidence in the automated Ki-67 index for guiding treatment decisions and prognostic evaluation.
To further assess the robustness of Kpi-Net, multiple experiments were conducted on the SHIDC-BC-Ki-67 dataset, yielding the box plot of Average F1 Score and violin plot of RMSE shown in Fig. 6. The box plot demonstrates that Kpi-Net consistently maintains the highest F1 scores, with experimental results concentrated between 84.56% and 85.79%, showing a minimal fluctuation range of only 1.23%, indicating strong stability. In comparison, TransAttUnet exhibits significantly poorer stability, with its lowest score (83.01%) being 4.14% below Kpi-Net’s minimum value. TransUNet shows an overall performance shift downward by 1.5%, with F1 scores ranging from 82.81% to 84.23%. PathoNet demonstrates a performance bottleneck at the 80% level, where its maximum value falls below Kpi-Net’s minimum. U-Net displays its most stable range between 77.35% and 78.41%, nearly 7% lower than Kpi-Net’s results.
The violin plot reveals that Kpi-Net achieves the most concentrated and lowest overall RMSE distribution among all models. While TransAttUnet, as the second-best model, shows a relatively low median error, its distribution exhibits significant heavy-tailed characteristics, with an upper bound reaching 0.04879 - representing a 63.9% greater maximum fluctuation amplitude compd to Kpi-Net. PathoNet demonstrates relatively concentrated RMSE distribution but slightly inferior error control compared to Kpi-Net. Other models display more dispersed RMSE distributions with generally higher values, indicating greater variability and higher prediction errors. This robustness difference holds critical value in clinical scenarios. When processing challenging samples with weak positive cell proportions >15%, Kpi-Net’s narrow fluctuation band ensures grading consistency, whereas TransAttUnet’s variability could lead to misclassification of samples near the 20% cutoff threshold.
Box plots of average F1 score and violin plots of RMSE. Comparative analysis of stability and accuracy for different models in Ki67 PI quantification tasks through multiple experiments on the SHIDC-BC-Ki-67 dataset.
Experimental results on the BCData dataset
To assess the generalization capability of Kpi-Net, multiple repeated experiments were conducted on the BCData dataset. As shown in Table 4, Kpi-Net achieved an F1-score of 89.56% for positive cell detection on this more challenging dataset, representing a 1.23% improvement over TransAttUnet. The model demonstrated 80.86% precision in negative cell identification, effectively controlling false positive risks.
With an average F1-score of 84.25%, Kpi-Net outperformed PathoNet (82.92%) and U-Net (78.05%) by 1.33% and 6.20%, respectively, validating the effectiveness of its multi scale fusion mechanism. Notably, the model reduced the miss rate for weakly positive cells by 28.7% compared to PathoNet, while maintaining a low false positive rate of 15.65% for negative cells. Despite significant dataset heterogeneity, Kpi-Net showed only a 0.54% performance degradation compared to its results on SHIDC-BC-Ki-67, substantially lower than PathoNet’s 1.38% decrease, demonstrating superior clinical generalization potential.
Tables 5 Comparative performance of Kpi-Net versus benchmark models on the BCData dataset. Kpi-Net achieves optimal comprehensive performance with minimal prediction error (RMSE = 0.04854), strongest prediction consistency (R = 0.962), and highest PI accuracy (95.58%). The RMSE value demonstrates 1.24% reduction compared to the suboptimal TransAttUnet model and 9.51% improvement over baseline U-Net. While PI accuracy differences among models remain marginal, Kpi-Net simultaneously maintains global proportion accuracy while minimizing single-cell recognition errors, thereby validating the breakthrough contribution of the multi scale fusion strategy in cellular-level precision.
Figures 7 Generalization performance evaluation on BCData dataset. Box plots demonstrate Kpi-Net’s superior stability with peak F1-score reaching 84.25%, all experimental results exceeding the 82.87% high baseline, and minimal intra-group fluctuation (1.38%). Notably, its minimum value surpasses PathoNet’s entire score range, confirming consistent performance in complex pathological scenarios.
Violin plots show Kpi-Net maintains the lowest RMSE range (0.04854–0.05012) with median 0.04904, outperforming all comparators - 3.56% lower than suboptimal TransAttUnet and demonstrating exceptional stability (range = 0.00158). While TransAttUnet/TransUNet surpass traditional CNNs, they exhibit significant fluctuations. PathoNet/UNet++ show concentrated but elevated distributions (PathoNet median 3.7% higher than Kpi-Net). Crucially, Kpi-Net’s RMSE increase from SHIDC-BC-Ki-67 to BCData (4.1%) remains substantially lower than U-Net’s 8.9%, evidencing the multi scale fusion strategy’s strong adaptability across heterogeneous pathological data sources while minimizing retraining requirements.
Box plots of average F1 score and violin plots of RMSE. Comparative analysis of stability and accuracy for different models in Ki67 PI quantification tasks through multiple experiments on the BCData dataset.
Visual analysis
Visualization results on SHIDC-BC-Ki-67 and BCData test sets
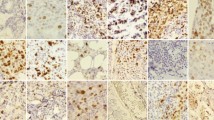
Figures 8 and 9 present comparative visualization results of different models on the SHIDC-BC-Ki-67 and BCData test sets, respectively, demonstrating Kpi-Net’s superior performance in breast cancer histopathology image analysis. In these visualizations, red annotations indicate Ki67 positive cells while green denotes negative cells.
The U-Net architecture, while capable of capturing global cellular features, exhibits significant limitations in processing cell-dense regions due to its fixed receptive field, manifesting as missed detections and false positives when handling cell overlapping and boundary ambiguity. UNet++, an improved variant incorporating dense connections for enhanced feature propagation, still fails to accurately delineate boundaries between positive and negative cells in complex scenarios involving cell adhesion and staining heterogeneity. PathoNet demonstrates relative advantages in positive cell detection but shows pronounced false positives and missed detections for negative cells, particularly in highly heterogeneous regions. TransUNet integrates Transformer modules for global context modeling but suffers from insufficient local feature extraction, leading to missed weakly positive cells in cases of morphological variation or sparse distribution. TransAttUnet, despite its optimized attention mechanism, struggles to suppress noise interference in densely overlapping cell regions, resulting in persistent misclassification.
In contrast, Kpi-Net dynamically adjusts receptive fields through RDMS while strengthening global dependency modeling via Transformer Blocks, combined with HS-CBAM-FPN for critical feature selection. This integrated approach effectively addresses challenges including uneven cell distribution and staining variability. Visualization results confirm Kpi-Net precise boundary delineation in cell-dense regions and accurate differentiation between positive/negative cells, demonstrating balanced global-local feature integration, minimized detection errors, and enhanced clinical applicability and robustness in complex scenarios.
Visualization results of different models on the SHIDC-BC-Ki-67 test set. Red annotations indicate Ki67 positive cells, while green annotations denote negative cells.
Visualization results of different models on the BCData test set. Red annotations indicate Ki67 positive cells, while green annotations denote negative cells.
Visualization results of PI Index calculation on SHIDC-BC-Ki-67 test set ROIs
To ensure accurate PI calculation, this study utilized sufficiently large ROIs. During model training, each original high-resolution image was divided into smaller sub-images. For PI accuracy evaluation, sub-images from the same patient in the SHIDC-BC-Ki-67 dataset were reassembled into complete high-resolution images (4912\(\times\)3684 pixels) to simulate WSI analysis in clinical practice and enable comprehensive evaluation of larger tissue structures (Fig. 10).
Figure 10 presents Kpi-Net prediction results on these reassembled images, demonstrating its superior performance in cell detection and counting tasks. The model accurately identifies and classifies both \(Ki67^+\) and \(Ki67^-\) . Following established clinical standards, PI indices were categorized as: low proliferation (<16%), intermediate proliferation (16–30%), and high proliferation (>30%). The predicted PI indices of 5.66%, 4.40%, 20%, and 70.42% exemplify Kpi-Net’s accuracy and reliability across samples with varying proliferation levels.
PI index prediction by Kpi-Net on ROI regions from the SHIDC-BC-Ki-67 test set.
Visualization results on DeepSlides sets
This study comprehensively demonstrates Kpi-Net’s cell segmentation capability on the DeepSlides dataset through pixel-level visualization across low, medium, and high cellular density regions. The results visually confirm the model’s exceptional capacity to integrate local details with global contextual information, effectively circumventing subjective biases inherent in traditional downstream PI value assessments.
As illustrated in Fig. 11, Kpi-Net exhibits differentiated segmentation performance: (1) precise isolation of individual cells in low-density regions provides the foundation for PI calculation; (2) effective resolution of adjacent cell boundaries in medium-density areas through multiscale design; and (3) preservation of overlapping cell morphology integrity in high-density regions via global-local cooperative processing, overcoming adhesion segmentation challenges. Experimental results confirm Kpi-Net’s stable performance in clear-boundary scenarios and superior adaptability to complex tissue architectures, establishing a viable solution for cell segmentation and counting tasks.
Cell segmentation and PI prediction. Demonstration of Kpi-Net segmentation results and PI index calculation on the DeepSlides dataset.
Model evaluation and ablation studies
Model efficiency and performance evaluation
To validate the computational efficiency and performance balance of Kpi-Net’s multi-module integration, comparative experiments were conducted on the SHIDC-BC-Ki-67 dataset. As evidenced in Table 6, Kpi-Net achieves lightweight architecture with 22.2 GFLOPs computational complexity and 15.89 million parameters, representing 27.6% reduction in computations and 10.9% fewer parameters compared to TransAttUnet, while improving average F1-score by 1.32%. With an inference speed of 68 FPS, Kpi-Net demonstrates nearly 3\(\times\) acceleration over TransAttUnet.
Ablation studies
To validate the efficacy and synergistic interactions of core modules within Kpi-Net, systematic ablation studies were conducted on the SHIDC-BC-Ki-67 and BCData datasets. Comparative models were constructed by incrementally integrating modules to quantify the performance contributions of the RDMS Module, Transformer Block, and HS-CBAM-FPN.
Tables 7 demonstrate that the RDMS Module enhanced the Avg.F1 score from a baseline of 79.89% to 80.24% on SHIDC-BC-Ki-67 by leveraging multi scale convolutions to accommodate cellular morphological heterogeneity, confirming its efficacy in establishing adaptive receptive fields for polymorphic cell recognition. On BCData with more pronounced staining heterogeneity, it further boosted F1 by 0.64%, exhibiting notable effectiveness in addressing cell adhesion. The Transformer Block elevated Avg.F1 by approximately 1.8% across both datasets through global self-attention mechanisms modeling long-range spatial dependencies among cells, establishing it as the core solution for mitigating missed detection of sparse cells. HS-CBAM-FPN achieved an Avg.F1 of 81.45% on SHIDC-BC-Ki-67 by employing attention mechanisms to prioritize features robust to staining variations, with its superior adaptability to heterogeneous staining further validated on BCData, thereby enhancing model robustness against staining noise.
Synergistic effects between modules proved critical. The RDMS and Transformer Block combination increased Avg.F1 to 82.89% on SHIDC-BC-Ki-67, a 1.16% gain over the standalone Transformer Block. Subsequent integration of HS-CBAM-FPN for feature refinement elevated the ternary combination’s performance to 85.79%, marking a 5.9% improvement over the baseline and demonstrating strong complementarity between staining-robust features and cross-scale contextual information. This synergy generalized robustly to BCData, where the ternary combination achieved 84.25% Avg.F1, outperforming all binary combinations. Module integration order did not alter performance trends.
In summary, ablation studies delineate distinct functional roles: RDMS provides foundational support for multi scale morphology recognition, Transformer Block addresses sparse cell detection limitations, and HS-CBAM-FPN primarily enhances staining robustness. Critically, their synergistic integration forms a comprehensive processing chain, driving Kpi-Net’s exceptional Avg.F1 exceeding 85% on both datasets and establishing modular design principles for precise quantification in complex histopathological imaging.
Ablation studies on SHIDC-BC-Ki-67 validate the optimal tiered dilation scheme [1,4,6,12] for the RDMS Module, where excessive density in [1,2,3,4] causes receptive field overlap and performance degradation; [3,6,9,12] lacks fine-grained local details with partial overlap; [1,6,12,18] suffers intercellular relationship loss from absent mid-scale coverage; and [8,12,16,24] impairs local detail capture. Conversely, [1,4,6,12] achieves maximal precision-recall equilibrium through complementary scaled receptive fields (3\(\times\)3px for cellular morphology; 9\(\times\)9px for intercellular proximity; 13\(\times\)13px for cell clusters; 25\(\times\)25px for global tissue distribution), eliminating computational redundancy while balancing feature diversity and spatial complexity. Tables 8 confirms this configuration uniquely optimizes multi scale integration across local-to-global hierarchies.
Although ablation studies demonstrate the Transformer Block critical contribution to model performance–particularly in mitigating missed detection of sparse cells through global context modeling–its associated computational overhead warrants evaluation. To comprehensively assess this module’s cost-performance ratio, a controlled experiment was designed, analyzing changes in computational complexity, parameter count, inference speed, and core performance metrics when integrating versus removing the Transformer Block while maintaining identical RDMS and HS-CBAM-FPN module structures.
As quantified in Table 9, introducing the Transformer Block elevated Avg.F1 from 82.89 to 85.79%, confirming its essential role in sparse cell localization and classification accuracy, while concurrently increasing FLOPs from 18.6G to 22.2G (+19.4%), parameters from 14.1 to 15.9M (+12.8%), and reducing inference speed from 78FPS to 68FPS (−12.8%)–all within manageable margins without order-of-magnitude escalation.
This experiment quantifies the efficiency-effectiveness balance of the Transformer Block. Results indicate that despite moderate computational and speed costs, the performance gains for sparse cell detection and complex background handling prove decisive. Its favorable cost-performance ratio underscores its indispensability as a core component in Kpi-Net.
Conclusion
The proposed Kpi-Net model establishes an effective multi-scale deep learning framework by integrating RDMS, Transformer Block, and HS-CBAM-FPN structures to capture and combine both local and global features. Experimental results demonstrate its superior performance in improving cell detection accuracy and Ki-67 index quantification precision. The model outperforms existing approaches in terms of F1-score, RMSE, and PI accuracy across multiple datasets, particularly excelling in cell-dense regions, while exhibiting remarkable generalization capability and stability under challenging conditions including uneven cell distribution, staining inconsistency, and cell overlapping. Its innovative multi-scale feature fusion mechanism, enabled by the RDMS module and HS-CBAM-FPN structure, facilitates precise multi-scale feature extraction and attention-based feature integration optimization, thereby enhancing critical feature recognition.
However, this study has several limitations. First, although the validation was performed on three public datasets, these data sources remain limited. Further multi-center validation on larger and more diverse prospective cohorts is necessary before the model can be deployed in clinical practice to comprehensively demonstrate its generalizability across real-world clinical settings. Second, the evaluation focus of this study was on instance-level detection and counting metrics (e.g., F1-score) and the accuracy of the proliferation index (PI), which are most relevant to the clinical task of Ki-67 quantification. It did not include pixel-level segmentation metrics (such as Dice coefficient or Intersection-over-Union, IoU) to evaluate segmentation boundaries. Future work will incorporate such pixel-wise analyses to provide a more comprehensive evaluation of the segmentation boundaries predicted by the model.
Future research directions will focus on: expanding multi-center and multi-staining protocol datasets to improve model generalization; optimizing computational efficiency through lightweight design and distributed computing for practical deployment; enhancing model interpretability via attention visualization and feature importance analysis to facilitate clinical decision-making; as well as exploring multi-task learning and cross-modal data fusion to extend Kpi-Net’s applications in pathological image analysis.
Data availability
The datasets employed in this study–namely SHIDC-BC-Ki-67, BCData, and DeepSlides–are publicly available at https://shiraz-hidc.com/ki-67-dataset, https://sites.google.com/view/bcdataset, and https://zenodo.org/record/1184621, respectively. The source code and trained models generated during this study are available from the corresponding author upon reasonable request.
References
Han, B. et al. Cancer incidence and mortality in china, 2022. J. Natl. Cancer Cent. 4, 47–53 (2024).
Giaquinto, A. N. et al. Breast cancer statistics 2024. CA Cancer J. Clin. 74, 477–495 (2024).
Nyqvist-Streng, J. et al. The prognostic value of changes in ki67 following neoadjuvant chemotherapy in residual triple-negative breast cancer: A swedish nationwide registry-based study (Breast Cancer Res, Treat, 2025).
Acs, B. et al. Systematically higher Ki-67 scores on core biopsy samples compared to corresponding resection specimen in breast cancer: A multi-operator and multi-institutional study. Mod. Pathol. 35, 1362–1369 (2022).
Lu, W. et al. Ai-based intra-tumor heterogeneity score of ki67 expression as a prognostic marker for early-stage er+/her2- breast cancer. J. Pathol. Clin. Res. 10, e346 (2024).
Rewcastle, E. et al. The Ki-67 dilemma: Investigating prognostic cut-offs and reproducibility for automated Ki-67 scoring in breast cancer. Breast Cancer Res. Treat. 207, 1–12 (2024).
Luchini, C. et al. Ki-67 assessment of pancreatic neuroendocrine neoplasms: Systematic review and meta-analysis of manual vs. digital pathology scoring. Mod. Pathol. 35, 712–720 (2022).
Shaker, N. et al. Automated imaging analysis of Ki-67 immunohistochemistry on whole slide images of cell blocks from pancreatic neuroendocrine neoplasms. J. Am. Soc. Cytopathol. 13, 205–212 (2024).
Vesterinen, T. et al. Automated assessment of Ki-67 proliferation index in neuroendocrine tumors by deep learning. APMIS 130, 11–20 (2022).
Govind, D. et al. Improving the accuracy of gastrointestinal neuroendocrine tumor grading with deep learning. Sci. Rep. 10, 2045–2322 (2020).
Fulawka, L., Blaszczyk, J., Tabakov, M. & Halon, A. Assessment of Ki-67 proliferation index with deep learning in dcis (ductal carcinoma in situ). Sci. Rep. 12, 3166 (2022).
Karol, M. et al. Deep learning for cancer cell detection: Do we need dedicated models?. Artif. Intell. Rev. 57, 53 (2024).
Rewcastle, E. et al. The ki67 dilemma: Investigating prognostic cut-offs and reproducibility for automated ki67 scoring in breast cancer. Breast Cancer Res. Treat. 207, 1–12 (2024).
Aniq, E., Chakraoui, M. & Mouhni, N. Artificial intelligence in pathological anatomy: Digitization of the calculation of the proliferation index (Ki-67) in breast carcinoma. Artif. Life Robotics 29, 177–186 (2024).
Phan, T. C. & Huu, H. N. Enhancing breast cancer detection with advanced deep learning techniques for Ki-67 nuclear protein analysis. SN Comput. Sci. 5, 663 (2024).
Zehra, T. et al. Use of novel open-source deep learning platform for quantification of Ki-67 in neuroendocrine tumors-analytical validation. Int. J. Gen. Med. 16, 5665–5673 (2023).
Negahbani, F. et al. Pathonet introduced as a deep neural network backend for evaluation of Ki-67 and tumor-infiltrating lymphocytes in breast cancer. Sci. Rep. 11, 8489 (2021).
Geread, R. S. et al. pinet-an automated proliferation index calculator framework for ki67 breast cancer images. Cancers 13, 11 (2021).
Ronneberger, O., Fischer, P. & Brox, T. U-net: Convolutional networks for biomedical image segmentation. In International Conference on Medical Image Computing and Computer-Assisted Intervention(MICCAI) 234–241 (Springer, Munich, Germany, 2015).
Gu, Z. et al. Ce-net: Context encoder network for 2d medical image segmentation. IEEE Trans. Med. Imaging 38, 2281–2292 (2019).
Zhou, Z., Rahman Siddiquee, M. M., Tajbakhsh, N. & Liang, J. Unet++: A nested u-net architecture for medical image segmentation. In Stoyanov, D. et al. (eds.) Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support, vol. 11045 of Lecture Notes in Computer Science, 1–12 (Springer, Cham, 2018).
Woo, S., Park, J., Lee, J. Y. & Kweon, I. S. Cbam: Convolutional block attention module. In (eds. Ferrari, V., Hebert, M., Sminchisescu, C. & Weiss, Y.) Computer Vision – ECCV 2018, vol. 11211 of Lecture Notes in Computer Science (Springer, Cham, 2018).
Yang, Z., Peng, X., Yin, Z. & Yang, Z. Deeplab_v3_plus-net for image semantic segmentation with channel compression. In Proceedings of International Conference on Communication Technology (ICCT) 1320–1324 (2020).
Chen, Y. et al. Accurate leukocyte detection based on deformable-detr and multi-level feature fusion for aiding diagnosis of blood diseases. Comput. Biol. Med. 170, 107917 (2024).
Liu, X. et al. Multi-scale feature fusion for prediction of idh1 mutations in glioma histopathological images. Comput. Methods Programs Biomed. 248, 108116 (2024).
Eren, F. et al. Deepcan: A modular deep learning system for automated cell counting and viability analysis. IEEE J. Biomed. Health Inform. 26, 5575–5583 (2022).
Zhao, X. et al. M2snet: Multi-scale in multi-scale subtraction network for medical image segmentation. arXiv:2303.10894 (2023).
Dosovitskiy, A. et al. An image is worth 16x16 words: Transformers for image recognition at scale. arXiv (2020). arXiv:2010.11929.
Chen, J. et al. Transunet: Transformers make strong encoders for medical image segmentation. arXiv preprint arXiv:2102.04306 (2021).
Chen, B., Liu, Y., Zhang, Z., Lu, G. & Kong, A. W. K. Transattunet: Multi-level attention-guided u-net with transformer for medical image segmentation. IEEE Trans. Emerg. Topics Comput. Intell. 8, 55–68 (2024).
Prangemeier, T., Reich, C. & Koeppl, H. Attention-based transformers for instance segmentation of cells in microstructures. In 2020 IEEE International Conference on Bioinformatics and Biomedicine (BIBM) 700–707 (2020).
Xia, C., Wang, X., Lv, F., Hao, X. & Shi, Y. Vit-comer: Vision transformer with convolutional multi-scale feature interaction for dense predictions. In 2024 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) 5493–5502 (2024).
Huang, Z. et al. BCData: A Large-Scale Dataset and Benchmark for Cell Detection and Counting, vol. 12265 of Lecture Notes in Computer Science 1–12 (Springer, Cham, 2020).
Senaras, C. Deepslides Dataset (CERN, Meyrin, Switzerland, Zenodo, 2018).
Khademi, A. Image analysis solutions for automatic scoring and grading of digital pathology images. Can. J. Pathol. 5, 51–55 (2013).
Acknowledgements
This work was supported in part by the National Talent Introduction Program of China (Grant No. H20250815), the Natural Science Foundation of Hebei Province (H2024403001), the Science and Technology Program of Shijiazhuang (2511301801A), the Science Research Project of Hebei Provincial Department of Education (BJK2024099), and the Shijiazhuang High-level Talents Startup Funding Project (248790067A).
Author information
Authors and Affiliations
Contributions
Author 1 (Qi Liu): Conceived the research, designed the experiments, and drafted the manuscript. Author 2 (Zhenfeng Zhao): Participated in the experimental design, jointly analyzed the data, and reviewed the manuscript. Author 3 (Lei Lou): Collaborated in revising the manuscript and supervised the study. Author 4 (Yuehong Li): Reviewed the experimental results and provided suggestions. Author 5 (Shenwen Wang): Contributed to the interpretation of results and provided feedback on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, Q., Zhao, Z., Lou, L. et al. Multi scale deep learning quantifies Ki67 index in breast cancer histopathology images. Sci Rep 15, 44972 (2025). https://doi.org/10.1038/s41598-025-28734-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-28734-8