Abstract
Human alveolar type 2 (AT2) cells play a crucial role in maintaining lung homeostasis and facilitating repair. Researchers are investigating AT2 cells in ex vivo platforms due to their essential roles in lung physiology, disease modeling, and drug testing. Herein, we present a comprehensive workflow for the isolation and culture of primary human AT2 cells, including an efficient enzyme-based lung tissue digestion protocol and automated purification of AT2 cells from both healthy and diseased lungs. By targeting the HTII-280 surface protein on AT2 cells, we employed an automated magnetic-activated cell sorting (MACS) separation strategy using the autoMACS® Pro Separator. AT2 cells from healthy and diseased lungs were grown in 3D culture and formed alveolospheres, elucidating that the isolation strategy did not affect cell functionality. This approach provides high AT2 cell yield and purity, offering scalability to generate sufficient quantities of cells for downstream applications, such as disease modeling and cell-based therapeutics.
Similar content being viewed by others
Introduction
Gas exchange occurs in the alveolar compartment, which is composed of alveolar type 2 (AT2) and alveolar type 1 (AT1) epithelial cells, capillaries, and a myriad of resident mesenchymal cells1. The outer layer of the alveoli is composed of epithelial cells. AT1 cells are large squamous cells that comprise 96% of the epithelial surface area and form the epithelial component of the thin air–blood barrier2. In contrast, AT2 cells are cuboidal pneumocytes that are much smaller and constitute 4% of the alveolar epithelial surface area. However, they represent 60% of alveolar epithelial cells by number2,3. A combination of studies using in vivo clonal lineage analysis and injury models in rodents have established the contribution of AT2 cells to alveolar maintenance and repair, demonstrating that AT2 cells function as progenitor cells in the alveoli4,5,6,7,8. Primary AT2 cells are studied in ex vivo platforms to understand their essential roles in maintaining lung homeostasis, simulating disease processes, and evaluating therapeutic responses. Importantly, AT2 cell transplantation has been proposed as a therapeutic strategy for treating chronic lung diseases such as IPF, with evidence of efficacy following transplantation of AT2 cells in small animal models of IPF9,10. The number of cells typically ranged from 9 × 105 to 2.5 × 106 per dose in mice and rats, respectively9,10,11,12. In preclinical trials, the safety and potential benefits of AT2 cell therapy were also evaluated in patients with varying degrees of IPF. However, the total number of instilled AT2 cells was significantly higher at 1 × 109 per dose, administered across four doses13. Higher cell numbers are required for delivery into human lungs, highlighting the need for isolation protocols capable of generating cells at scale.
While cell-sorting technologies have advanced, the affinity-based sorting technology of fluorescence-activated cell sorting (FACS) remains the gold standard14,15,16,17. FACS enables the analysis and sorting of individual cells based on the relative expression of multiple surface markers, with minimal impact on cell differentiation14,15,16,17,18,19,20. While FACS provides highly pure sorted cell populations, it is comparatively slower, has lower throughput, and takes longer to process than other sorting methods. FACS is often constrained by limited machine availability and the need for trained personnel to operate the system14,15,16,17. Magnetic activated cell sorting (MACS), which utilizes MicroBead Technology (Miltenyi Biotec, San Diego, California, USA), offers a potential alternative to FACS. This affinity-based sorting technology offers high-purity selection and viability, with higher throughput at a fraction of the processing time16,17,21,22,23,24,25. The autoMACS Pro Separator (Miltenyi Biotec) is an automated, multisample, bench-top magnetic cell separator compatible with MACS MicroBeads, designed to isolate and enrich cells through both depletion and enrichment strategies. In depletion mode, unwanted cells are magnetically labeled and removed from the cell suspension using a unique marker for that specific cell type, leaving the target population intact. In contrast, the enrichment mode isolates and amplifies the desired cell population by targeting specific surface markers. To ensure specific cell enrichment, we employed two isolation strategies to selectively target AT2 cells expressing HTII-280, a unique apical plasma membrane protein specific to AT2 cells26. Both workflows utilized the autoMACS Pro Separator: (a) a one-step approach involving direct HTII-280 positive selection and (b) a two-step approach combining initial white blood cell depletion with subsequent HTII-280 positive selection. In this study, AT2 cells are defined by co-expression of the surface markers epithelial cell adhesion molecule (EPCAM) and HTII-280; accordingly, the terms “AT2 cells” and “EPCAM + /HTII-280 + cells” are used interchangeably throughout the text. Comprehensive analyses were conducted to evaluate the viability and purity of AT2 cells at each stage of the isolation process. To further assess their functional capacity, the isolated cells were cultured using established alveolar epithelial cell culture platforms27.
Materials and methods
Preservation of lung biopsies
Biospecimen procurement was supported by Ontario Health (Trillium Gift of Life Network) and the Biosafety Office at the University Health Network. Donor lungs were declined for transplantation either at the time of procurement or after Ex-Vivo Lung Perfusion (EVLP). Lungs used and collected in our clinical program were approved under protocol 06-0283 by the Research Ethics Board of the University Health Network. Donor lungs were preserved in cold low-potassium dextran solution (19811, Perfadex Plus, XVIVO Perfusion, Gothenburg, Västra Götaland County, Sweden) and kept on ice until arrival at the Toronto Lung Transplant Program, Toronto General Hospital. Lung biopsies (donor and recipient explanted diseased lung) were collected and provided from the operating room under protocol 11-0509, approved by the Research Ethics Board of the University Health Network, with the help of the Toronto Lung Transplant Program Biobank. All methods were performed in accordance with the relevant guidelines and regulations, and informed consent was obtained from all subjects and/or their legal guardian(s). EVLP-rejected lungs were flushed with a low-potassium dextran solution and stored inflated at 4 °C after donor lung retrieval. Following procurement, the lung biopsies were typically processed within 12 h of collection. Lung biopsies could be preserved in a lung storage solution (Perfadex Plus supplemented with 2% Antibiotic–Antimycotic (450-115-EL, Wisent, Saint-Jean-Baptiste, QC, Canada)) to extend the processing window. Samples were placed in sterile containers (NCS602-1O, Thermo Fisher Scientific, Waltham, MA, USA), submerged in the storage solution, and overlaid with a sponge (2913, Covidien, Dublin, Leinster, Ireland). Under these storage conditions, flow cytometric analysis confirmed that AT2 cell viability remained above 90% at 36 h post-collection (Figure S1). Patient demographic and clinical information are summarized in Supplementary Table 1A–C.
Mechanical and enzymatic digestion of a healthy lung tissue biopsy
This enzymatic digestion protocol was adapted from that described initially by Fujino et al.28, with modifications introduced to enhance the speed and efficiency of AT2 cell extraction. While 10 g of lung tissue were routinely processed per experiment (Figure S2A), our protocol effectively accommodated biopsy samples with dry weights ranging from 0.2 to 18 g. Dispase II (4942078,001, Roche, Basel, Switzerland) solution was prepared at 2 U/mL in 50 mL conical tubes (CLS430829-500EA, Corning, Corning, NY, USA) and then mixed and pre-warmed at 37 °C for 30 min before lung tissue digestion. The tissue samples were placed in a petri dish (431762, Thermo Fisher Scientific) and washed with phosphate-buffered saline (PBS) (Wisent). Cover sponges were gently pressed onto the lung tissue to remove excess solution. 10 mL of dispase II solution was added to a 10 mL syringe (309695, BD Biosciences, San Jose, CA, USA) and injected through a 27 G × 1.5″ needle (301629, BD Biosciences) into multiple distal airways (Figure S2B). Targeted injection through the distal airways resulted in lung inflation, significantly enhancing the extraction of AT2 cells. The specimen was cut into ~ 2-g pieces using operating scissors (7607-A8-114, Almedic, Montreal, QC, Canada) and splinter forceps (7721-A10-624, Almedic). Each piece was then placed in its own 50 mL conical tube for enzymatic digestion. 8 mL of dispase II and 100 µL of 10 mg/mL DNase 1 (DN25-1G, Sigma-Aldrich, St. Louis, MO, USA) were added to each 50 mL conical tube. Lung tissue samples were minced for 1 min using two operating scissors and cut into ~ 1 mm3 pieces. The tubes were placed laterally on a heated shaker (37 °C) for 30 min at 60 revolutions per minute. After rocking, the dispase II was neutralized with cold washing solution (319-005 CL, Dulbecco’s Modified Eagle Medium (DMEM) (Wisent) supplemented with 10% fetal bovine serum (FBS) (12483020, Thermo Fisher Scientific) and 50 μg/mL DNase 1). The lung tissue was minced for 1 min per tube, and tissue samples were passed through a 50 mL syringe (309653, BD Biosciences) with a 16 G × 1.5″ needle (305198, BD Biosciences) attached. This process was performed 2–3 times for each 50 mL conical tube. Next, the solutions were filtered through a 100 µm filter (352360, Corning) into new tubes, using a syringe plunger (BD Biosciences) if needed. The collection was topped with washing solution and centrifuged at 350 g for 10 min. The cell pellet was resuspended in pre-warmed (37 °C) red blood cell (RBC) lysis buffer (11814389,001, Roche). The solution was then incubated for 5 min at room temperature (RT), after which it was resuspended in washing solution and filtered through a 40 µm filter (352340, Corning). Finally, cells were spun down and reconstituted in washing solution for counting using a hemacytometer; only viable cells were counted. Cells were transferred into FACS tubes (352008, Corning) containing 1 mL of flow cytometry buffer (PBS supplemented with 2% FBS) for future analysis.
Mechanical and enzymatic digestion of IPF lung biopsy
The digestion protocol was slightly modified to improve AT2 cell yields from diseased biopsy samples (Figure S2C), resulting in yields comparable to those typically obtained from healthy tissue isolations. Specifically, more lung tissue (> 25 g) was used for processing, and 20 mL of dispase II was injected into multiple distal airways (Figure S2D). After the dispase II injection, each ~ 2-g piece was minced for an additional 3 min before being placed on the heated shaker. A final adjustment to the protocol included two consecutive RBC lysis steps due to increased RBC content, as the diseased tissue had not been flushed during procurement.
Labeling of AT2 cells with MACS MicroBeads
After mechanical and enzymatic digestion of the lung tissue, the cells were counted and prepared for labeling. The autoMACS Pro Separator (Miltenyi Biotec) was used to isolate AT2 cells either by (a) HTII-280 positive selection (1-step) or (b) white blood cell depletion, followed by HTII-280 positive selection (2-step). Biological tissue samples were allocated randomly to either protocol. IPF cell isolations were performed using only the 2-step protocol described below. The amount of CD45 MicroBeads (130-045-801, Miltenyi Biotec) used for labeling white blood cells was determined from the product datasheet (20 µL of MicroBeads per 1 × 107 cells). For example, for 1 × 108 cells, 200 µL of CD45 MicroBeads was required as per the Miltenyi protocol. The cell suspension was incubated with Microbeads and isolation buffer (PBS supplemented with 2% FBS and 2 mM EDTA (AM9261, Thermo Fisher Scientific, Waltham, MA, USA)) at 4 °C for 15 min in a 15 mL conical tube (CLS430791-500EA, Corning). The cell suspension was topped up with isolation buffer and then filtered through a 40 µm filter. This was followed by centrifugation at 300 g for 10 min. After aspiration, the cell pellet was resuspended in 2 mL of isolation buffer and run on the autoMACS Pro Separator using the depletion selection program in sensitive mode. The negative fraction, where CD45-positive cells were removed, was collected and topped with isolation buffer. It was then centrifuged at 300 g for 10 min. Next, the cell pellet was resuspended in 100 µL of solution A (DMEM with 20% FBS) and 100 µL of solution B (DMEM with 10% FBS and 50 μg/mL DNase I) with gentle mixing. Then, 100 µL of undiluted HTII-280 antibody (TB-27AHT2-280, Terrace Biotech, San Francisco, CA, USA) was added to label the AT2 cells, and the mixture was incubated at 4 °C for 20 min. The cell solution was topped up to 10 mL with isolation buffer and centrifuged at 300 g for 10 min. Using the product datasheet, the volume of anti-Mouse IgM MicroBeads (130-047-301, Miltenyi Biotech) was calculated and added based on the viable cell count. This cell-bead mixture was incubated at 4 °C for 15 min. Enrichment of AT2 cells was performed using the autoMACS Pro Separator in sensitive mode with the positive selection program. The positive fraction was collected, and viable cells were counted for seeding. A sample of cells was transferred into FACS tubes containing 1 mL of flow cytometry buffer for future analysis.
Flow cytometry analysis
A total of 2 × 105 cells were analyzed per tube for flow cytometry. Cells were centrifuged at 400 g for 5 min, and dead cells were labeled by incubating with fixable viability stain (1:1000, 562247, BD Biosciences) at RT for 10 min in the dark. Tubes were washed with PBS, centrifuged at 400 g for 5 min, and resuspended in 500 µL of 4% paraformaldehyde (PFA) (PI28908, Thermo Fisher Scientific, diluted in PBS) for 15 min at RT. After washing with PBS, the cells were stained using surface markers: EPCAM – PE-Cy7 (1:100, 324,222, BioLegend, San Diego, CA, USA), CD45 – BV605 (1:100, 564047, BD Biosciences), and HTII-280 (1:100). After washing, the secondary anti-Mouse IgM, Alexa Fluor 488 (1:100, A21042, Thermo Fisher Scientific), was added to bind the unconjugated primary HTII-280 antibody. All incubation steps were performed at 4 °C in the dark for 15 min. The cells were washed, centrifuged, and resuspended in 300 µL of PBS for flow cytometry analysis on the BD FACSymphony A3 Cell Analyzer (BD Biosciences). Data were analyzed using FlowJo software v10.10 (Tree Star, San Carlos, CA, USA).
Maintenance of AT2 cells in 3D culture
24-well plates (142475, Thermo Fisher Scientific) were pre-warmed overnight before use. 5 × 103 AT2 cells were resuspended in a 20 μL droplet of 100% growth factor-reduced (GFR) Matrigel (356231, Corning) and allowed to solidify in a 37 °C incubator for 20 min. 500 µL of pre-warmed 3D alveolosphere medium was added to each well to submerge the droplet completely. The media was changed every 2–3 days. This media was adapted for 3D droplet culture from Youk et al.27: Advanced DMEM/F12 (12634010, Thermo Fisher Scientific) supplemented with 10 mM HEPES (H0887-100ML, Sigma-Aldrich), 1X Penicillin Streptomycin (450-201-EL, Wisent), 10 mM Nicotinamide (N0636-100G, Sigma-Aldrich), 1 mM N-Acetylcysteine (A9165-5G, Sigma-Aldrich), 100 µg/mL Primocin (InvivoGen, San Diego, CA, USA), 1X B27 (17504044, Thermo Fisher Scientific), 10% R-Spondin 1 (The Princess Margaret Living Biobank, manually produced), 100 ng/mL human Noggin (120-10C-250UG, Peprotech, Cranbury, NJ, USA), 50 ng/mL human EGF (AF-100-15-100UG, Peprotech), 100 ng/mL human FGF-10 (100-26-250UG, Peprotech), 100 ng/mL human FGF-7 (100-19-250UG, Peprotech), and 10 µM SB431542 (616464-5MG, Millipore Sigma). Additionally, 3 µM of CHIR 99021 (4423/10, Tocris, Bristol, United Kingdom) was added fresh at each media change. For the first 48 h of culture, Y-27632, a ROCK inhibitor (72302, StemCell Technologies, Vancouver, BC, Canada), was added to promote cellular survival. Organoid cultures were maintained under standard cell culture conditions (37 °C, 5% CO2). After one month of culture, alveolospheres were dissolved from Matrigel using Cell Recovery Solution (CRS) (354253, Corning).
Single-cell RNA sequencing (scRNA-seq) analysis
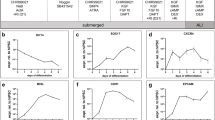
For single EPCAM + /HTII-280 + cells isolated by MACS, single-cell capture and downstream library construction were performed using the Chromium Next GEM Single Cell 3′ Gene Expression v3.1 kit (10 × Genomics, Pleasanton, CA, USA) according to the manufacturer’s instructions. The libraries were sequenced on an Illumina NovaSeq 6000 (Illumina, San Diego, CA, USA). Cell Ranger (v7.0.0, 10 × Genomics) was used to process barcodes and count single-cell 3′ unique molecular identifiers (UMIs), aligning reads to the human reference genome GRCh38.2020 to generate gene–barcode count matrices. Each matrix was processed using the Seurat package (v5.3.0) in R and underwent quality control. In brief, cells with high mitochondrial content (greater than 15%) and small or excessively high library sizes (gene number lower than 340 or higher than 7190; UMI count lower than 1000 or higher than 30,000) were excluded. After filtration, the UMI counts were normalized using Seurat’s SCTransform. Principal component (PC) analysis (PCA) was conducted using the Seurat function RunPCA on the top 2000 variable genes identified by FindVariableFeatures. Cells were clustered using Seurat’s FindClusters function with the top 30 PCs at a resolution of 0.5 and embedded using Uniform Manifold Approximation and Projection (UMAP) dimensionality reduction with the Seurat function RunUMAP on the same 30 PCs. The relative expression levels of canonical AT2 cell markers were visualized using FeaturePlot from the Seurat and scCustomize (v3.1.3) packages in R.
Real-time quantitative PCR (qPCR) analysis
Pipette tips and conical tubes were dip-coated in 1% BSA (A7906-100G, Sigma-Aldrich) to maximize the recovery of alveolospheres from plastic consumables. Matrigel droplets from the 24-well plate were washed with 500 µL of cold PBS, then 500 µL of CRS was added. The plate was placed in an ice box and rocked slowly for an hour in a cold room set to 4 °C. The well contents were pooled into a 15 mL conical tube and centrifuged at 400 g for 5 min. Once aspirated, the alveolospheres were treated with pre-warmed (37 °C) TrypLE Express Enzyme (12604021, Thermo Fisher Scientific). The mixture was pipetted thoroughly and incubated at 37 °C for 10 min. After incubation, the washing solution was added at a 1:1 ratio and mixed thoroughly. Broken-up alveolospheres were centrifuged at 400 g for 5 min. The supernatant was aspirated, and the pellet was resuspended in 1 mL of washing solution for cell counting. Single cells were then collected in an Eppendorf tube (30119401, Millipore Sigma) and centrifuged at 800 g for 5 min. The supernatant was aspirated, and the cell pellets were frozen at − 80 °C until RNA extraction.
Total RNA was extracted using the RNeasy Micro kit (74004, Qiagen, Hilden, Germany) according to the manufacturer’s protocol. mRNA was converted to cDNA using qScript Ultra SuperMix (95217-500, Quantabio, Beverly, MA, USA). Reverse transcription was performed on a C1000 Touch thermal cycler (Bio-Rad, Hercules, CA, USA). qPCR was performed using the SsoAdvanced Universal SYBR Green Supermix (1725271, Bio-Rad) according to the manufacturer’s instructions using a CFX384 qPCR Detection System (Bio-Rad). The protocol involved 40 cycles of a two-step PCR procedure: 95 °C for 20 s (melting) and 60 °C for 1 min (annealing and extension), preceded by an initial hot start at 95 °C for 2 min. Primers used for qPCR included surfactant protein C (SFTPC), tumor protein p63 (TP63), keratin 5 (KRT5), keratin 17 (KRT17), secretoglobin family 1A member 1 (SCGB1A1), forkhead box J1 (FOXJ1), and mucin 5AC (MUC5AC) (Sigma-Aldrich). The oligonucleotide sequences for the qPCR primers are listed in Supplementary Table 2.
Immunofluorescent staining of alveolospheres
Alveolospheres were carefully extracted from Matrigel for more efficient antibody labeling. This protocol was adapted from Dekkers et al.28. Recovered alveolospheres were spun down at 70 g for 3 min, resuspended in 2 mL of 4% PFA, and fixed for 45 min at 4 °C, with mixing halfway. After alveolosphere fixation, 9 mL of cold PBS-T (0.1% Tween 20 (P9416-50ML, Sigma-Aldrich), diluted with PBS) was added, gently mixed, and incubated at 4 °C for 10 min. Alveolospheres were then spun down at 70 g for 3 min, and the supernatant was aspirated. Two milliliters of a 30% sucrose solution (S0389-500G, Millipore Sigma) was added to the alveolospheres for overnight incubation at 4 °C. The next day, the supernatant was carefully removed, and the alveolospheres were resuspended in Frozen Section Compound (FSC) (95057-838, VWR, Radnor, PA, USA) for 30 min at RT. The solution was pipetted into a cryomold (22-363-552, Thermo Fisher Scientific) and placed on dry ice for 30 min to solidify. Afterward, it was stored at − 80 °C. Cryomold blocks were cut at 6 µm thickness, and four sections were attached per microscope slide (22037246, Thermo Fisher Scientific). Slides were kept at − 80 °C until use.
Slides were thawed at RT for 30 min, and a hydrophobic barrier was drawn around each alveolosphere section using a barrier pen (H-4000; Vector Laboratories, Newark, CA, USA). Next, the sections were fixed in 4% PFA for 30 min. After washing with PBS, alveolosphere sections were permeabilized (0.15% Triton X-100 (X100-1L, Sigma-Aldrich), diluted in PBS) for 10 min. After washing, the sections were blocked with blocking solution (PBS with 4% goat serum (PCN5000, Thermo Fisher Scientific, diluted in PBS) and 0.1% Triton X-100) at RT for 20 min. The sections were immediately stained with primary antibodies: epithelial cadherin (ECAD) (1:200, 14472, Cell Signalling Technology, Danvers, MA, USA), HTII-280 (1:100), pro-surfactant protein C (Pro-SPC) (1:500, AB3786, Sigma-Aldrich), aquaporin 5 (AQP5) (sc-514022, 1:200, Santa Cruz, Dallas, TX, USA), KRT5 (1:500, 71536, Cell Signaling Technology), and marker of proliferation Ki-67 (KI67) (1:200, 14-5698-82, Thermo Fisher Scientific), at 4 °C overnight in a dark, humidified chamber. Slides were covered with parafilm (PM996, Millipore Sigma) throughout the staining protocol to prevent evaporation. The following day, the sections were washed with PBS and stained with the corresponding secondary antibodies: anti-mouse IgG1, Alexa Fluor 555 (1:1000, A21127, Thermo Fisher Scientific), anti-rabbit IgG, Alexa Fluor 488 (1:1000, A11008, Thermo Fisher Scientific), anti-rabbit IgG, Alexa Fluor 647 (1:1000, A21244, Thermo Fisher Scientific) and anti-mouse IgM, Alexa Fluor 488 (1:100), were added for 1 h at RT in a dark, humidified chamber. After washing, the nuclei were stained with DAPI (1:500, 10236276001, Roche) for 10 min and ProLong Gold Antifade Mountant (P36930, Thermo Fisher Scientific) was applied to each slide, followed by a coverslip (48393-241, Thermo Fisher Scientific). Prepared slides were stored at 4 °C until imaging. Images were taken using the SP8 Confocal Laser-Scanning Microscope.
2D culture and immunofluorescent staining of AT2 cells
MACS-isolated cells were cultured in 2D using 8-well chamber slides (80841, IBIDI, Gräfelfing, Germany), as each chamber could be treated individually. Briefly, chamber slides were coated with Collagen 1 (354236, Corning) to promote AT2 cell attachment. AT2 cells were cultured in chamber slides in SAGM BulletKit (CC-3118, Lonza, Basel, Switzerland) supplemented with 1% FBS and 100 µg/mL Primocin. On Day 2, slides were briefly rinsed with PBS to remove the media, and cells were fixed with 4% PFA for 10 min at RT. Cells were washed with PBS to remove excess fixative, then permeabilized (0.15% Triton X-100) for 5 min at RT. Cells were washed with PBS and blocked with IT-FX Signal Enhancer (136933, Thermo Fisher Scientific) for 20 min at RT. Following blocking, the slides were stained with Pro-SPC antibody (1:500) overnight at 4 °C in a dark, humidified chamber. The next day, chamber slides were washed with PBS and stained with anti-rabbit IgG, Alexa Fluor 488 secondary antibody (1:1000) in a dark, humidified chamber for 1 h at RT. Next, chamber slides were washed with PBS and then incubated with DAPI (1:500) for 10 min in a dark, humidified chamber at RT. After removing the chamber divider, the slides were treated with ProLong Gold Antifade Mountant, and a coverslip was placed on top. Prepared slides were stored at 4°C until imaging. Images were taken using the SP8 Confocal Laser-Scanning Microscope.
Statistical analysis
Paired t-tests were conducted to compare yields before and after isolating each donor’s healthy or IPF lung cells. To compare the 1- and 2-step protocols for viability and purity, Welch’s t-tests were performed (to account for the difference in sample size). ANCOVA was also performed on the output purities of the 1- vs. 2-step protocols, with the random difference in input purities between the two groups as a covariate. Data represent mean ± SD (standard deviation) unless otherwise indicated. Statistical analyses were performed on GraphPad Prism v. 10.3.1 (La Jolla, CA, USA).
Results
Efficient and reproducible isolation of AT2 cells from healthy and diseased lung tissue
Using a combined approach of enzymatic digestion, mechanical agitation, and targeted antibody-based affinity purification against HTII-280, we successfully isolated AT2 cells with over 85% EPCAM + /HTII-280 + purity for downstream experiments (Fig. 1A and B). These cells were subsequently used for 3D culture and characterization (Fig. 1C). To perform the isolation, whole lung tissue was enzymatically and mechanically dissociated into single-cell suspensions, yielding an average of 23.9 × 106 cells per gram (cells/gram) of tissue (Fig. 2A). There is inherent donor-to-donor variability in the number of cells isolated per gram of healthy lung tissue. After automated purification, the AT2 cell purity increased from an average of 17% to 85% (± 7%) in the 1-step isolation protocol and from 23 to 95% (± 3%) in the 2-step isolation protocol (Fig. 2B). The 2-step protocol yielded higher EPCAM + /HTII-280 + cell purity with reduced variability, providing a more consistent enrichment outcome than the shorter 1-step approach. The cell viability following gentle enzymatic digestion and mechanical agitation of lung tissue was approximately 95%. Cellular viability remained consistently high throughout the workflow, averaging approximately 92% (± 6%) for both protocols after AT2 cell isolation (Fig. 2C). AT2 cell yield per gram of input lung tissue was higher with the 1-step protocol, yielding 2.5 × 106 (± 1.4 × 106) cells/gram, compared to 2.0 × 106 (± 1.4 × 106) cells/gram with the 2-step protocol. However, the difference was not statistically significant (Fig. 2D). An example of the AT2 cell enrichment from healthy lung tissue is shown in Fig. 2E.
Schematic overview of the lung digestion and cell isolation process, with alveolosphere culture and downstream analyses. (A) AT2 cells were extracted from lung tissue using enzymatic and mechanical dissociation, followed by mincing and filtering steps to achieve single cells for counting (the pre-MACS sample). (B) Using either a 1-step (HTII-280 positive enrichment) protocol or a 2-step (CD45 depletion, followed by HTII-280 positive enrichment) protocol, AT2 cells were labeled for MACS separation based on surface antigen HTII-280 and isolated using the autoMACS Pro Separator. (C) Following HTII-280 enrichment, healthy and IPF-diseased AT2 cells were counted, mixed with Matrigel, and seeded as droplets for 3D culture. After one month of growth, mature alveolospheres were collected for downstream analyses. The alveolospheres were either prepared for immunostaining and imaging or dissociated into single cells for real-time quantitative PCR (qPCR) to assess gene expression.
Analysis for healthy AT2 cells using the autoMACS Pro Separator. (A) Cells/gram of healthy lung tissue after enzymatic and mechanical digestion (n = 67). (B, C) Comparison of the 1-step (HTII-280 positive enrichment only, n = 40) and 2-step (CD45 depletion and HTII-280 positive enrichment, n = 27) isolation protocols. (B) Purity of AT2 cells (% of EPCAM + /HTII-280 + cells) before and after cell separation. The 1- and 2-step protocols increased purity by 68 and 71% on average, respectively (p < 0.0001 for both protocols). The 2-step protocol demonstrated higher output purities with less variability than the 1-step protocol (p < 0.0001 by Welch’s t-test, not significant [ns] by ANCOVA). (C) Cell viability (% of live cells/total cells) before and after the 1-step or 2-step protocols. Both protocols showed a slight, expected decrease in viability of approximately 2–3% (p = 0.04 and p = 0.0003). The average viability after isolation did not differ significantly between the 1- and 2-step protocols. (D) AT2 cell yield per gram of input lung tissue was higher with the 1-step protocol (2.5 × 106 cells/gram) compared to the 2-step protocol (2.0 × 106 cells/gram); however, this difference was not statistically significant. (E) An example of the flow cytometry analysis for healthy AT2 cell isolation using the 2-step protocol. By enzymatic and mechanical digestion, 15.8% of EPCAM + /HTII-280 + AT2 cells were extracted from lung tissue. More than 90% of the cells were located in the EPCAM-/HTII-280- quadrant and were CD45-positive. CD45 depletion using the autoMACS Pro Separator increased the EPCAM + /HTII-280 + AT2 population to 91.5%. HTII-280 positive enrichment using the autoMACS Pro Separator resulted in a final purity of 99% EPCAM + /HTII-280 + AT2 cells. FSC = forward scatter, SSC = side scatter, and the green dashed box indicates AT2 cells. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001 by Welch’s two-tailed t-test (1- vs. 2-step protocols) or paired two-tailed t-test (before vs. after separation).
CD45 depletion using the 2-step protocol was essential for eliminating contaminating white blood cells (WBCs). In Figure S3, the positive fraction consisted mainly of CD45-positive cells, demonstrating the effectiveness of the depletion step using the autoMACS Pro Separator. Fewer than 1% of cells in this fraction were EPCAM + /HTII-280 +, indicating that epithelial cell recovery was largely preserved. In contrast, the negative fraction, which contained the enriched EPCAM + /HTII-280 + cells, showed no detectable CD45-positive cells, indicating the successful removal of WBCs and the efficient enrichment of epithelial cells. The purified negative fraction obtained after CD45 depletion was subsequently used for positive selection of HTII-280 + AT2 cells in the next enrichment step. To further verify the identity of the enriched EPCAM + /HTII-280 + population after isolation, we employed several complementary methods in conjunction with flow cytometry. Firstly, scRNA-seq of the HTII-280 positive population showed robust expression of canonical AT2 markers, including SFTPC, SFTPB, SFTPA1, and NAPSA (Figure S4A), confirming successful enrichment of functional AT2 cells. qPCR analysis also revealed high expression of SFTPC in the HTII-280 (positive) fraction, confirming the selective enrichment of AT2 cells (Figure S4B). In contrast, the HTII-280 negative fraction showed almost no detectable expression. Finally, after seeding MACS-sorted cells in 2D culture, immunostaining at Day 2 demonstrated Pro-SPC-positive cuboidal cells, consistent with AT2 morphology (Figure S4C). We also applied the 2-step protocol to isolate AT2 cells from IPF lungs. Despite IPF lung biopsies containing fewer cells within each gram of lung tissue, we achieved an average yield of 8.8 × 106 cells/gram of tissue (Fig. 3A). Following automated purification, the AT2 cell purity increased from an average of 9% to 90% (± 7%) (Fig. 3B). The cell viability remained high at 96% for collected cells before automated separation, indicating that key tissue digestion parameters optimized for healthy lungs are still applicable and practical in fibrotic lung tissue. The post-isolation viability averaged 87% (± 7%) (Fig. 3C). The AT2 cell yield was 2.2 × 105 (± 2.1 × 105) cells/gram of input fibrotic lung tissue (Fig. 3D), which is lower than yields from healthy tissue but consistent with prior reports, reflecting the reduced abundance of AT2 cells in fibrotic lungs29,30. An example of the AT2 cell enrichment from diseased lung tissue is shown in Fig. 3E.
Analysis for IPF-diseased AT2 cells using the autoMACS Pro Separator (2-step only). (A) Cells/gram of IPF lung tissue after enzymatic and mechanical digestion (n = 11). (B) Purity of AT2 cells (% of EPCAM + /HTII-280 + cells) before and after the 2-step protocol. The purity increased by 81% after cell separation (p < 0.0001). (C) Cell viability (% of live cells/total cells) before and after cell separation. Cellular viability decreased on average by 9% during isolation, from 87 to 78% (p = 0.0016). (D) An average of 2.2 × 105 AT2 cells were obtained per gram of input lung tissue. (E) An example of the flow cytometry analysis for IPF-diseased AT2 cell isolation using the 2-step protocol. 6.2% EPCAM + /HTII-280 + AT2 cells were extracted from the lung tissue by enzymatic and mechanical digestion. 87% of the cells in the EPCAM-/HTII-280- fraction were CD45 positive. CD45 depletion using the autoMACS Pro Separator increased the EPCAM + /HTII-280 + AT2 population to 31.1%. HTII-280-positive enrichment using the autoMACS Pro Separator yielded a final purity of 92% EPCAM + /HTII-280 + AT2 cells. FSC = forward scatter, SSC = side scatter, and the green dashed box indicates AT2 cells. **p < 0.01, ****p < 0.0001 by two-tailed paired t-test.
3D culture of isolated AT2 cells
Here, we present the results of 3D alveolosphere cultures to demonstrate that upstream cell separation did not impair AT2 cell functionality. Isolated EPCAM + /HTII-280 + cells were seeded in Matrigel to investigate whether growth efficiency or differentiation potential differs between the alveolospheres formed from healthy and IPF AT2 cells. Alveolospheres images from Day 14 were quantified to determine colony-forming efficiency (CFE) using the “Analyze Particle” tool in the ImageJ software (Fig. 4A). IPF-derived alveolospheres formed much smaller colonies, suggesting a diminished regenerative potential and impaired self-renewal capacity, consistent with stem cell exhaustion and defective epithelial repair in IPF31. Fold expansion analysis performed at Day 28 demonstrated that IPF AT2-derived alveolospheres exhibited markedly reduced growth potential compared with those derived from healthy AT2 cells, indicating reduced proliferative capacity (Fig. 4B).
IPF alveolospheres demonstrate reduced self-renewal and proliferative potential in 3D culture. (A) Quantification of colony-forming efficiency (CFE) from Day 14 alveolospheres shows a significant reduction in IPF-derived cultures (n = 13) compared with healthy controls (n = 13) (p < 0.0001). (B) Day 28 alveolospheres were dissociated into single cells, and fold expansion analysis revealed significantly reduced growth of IPF alveolospheres (n = 5) compared with healthy controls (n = 5) after one month of 3D culture (p = 0.024). Scale bar = 500 μm. *p < 0.05, ****p < 0.0001 by Mann–Whitney two-tailed test.
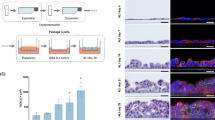
Consistent with previous findings, bright-field images of alveolospheres revealed notable morphological heterogeneity, commonly appearing as either cystic or folded/dense structures (Fig. 5A and B)27,32. Interestingly, a subset of alveolospheres cultured from IPF-derived AT2 cells exhibited characteristics not observed in healthy cells (Fig. 5C). These alveolospheres were defined by a combination of cystic architecture and folded structures. Healthy alveolospheres showed active proliferation, as indicated by KI67-positive staining (Fig. 5D). In contrast, IPF-derived alveolospheres (displayed in Fig. 5E and F) lacked KI67 expression, which further confirms the reduced fold expansion found in Fig. 4B of IPF AT2 cells in 3D culture. Healthy alveolospheres expressed multiple distal lung markers, including Pro-SPC, AQP5, and HTII-280 (Fig. 5G)33,34. IPF-derived alveolospheres had diminished Pro-SPC expression and weakened HTII-280 expression, compared to healthy alveolospheres (Fig. 5H). Moreover, IPF-derived alveolospheres exhibited co-expression of proximal (KRT5) and distal (AQP5) markers within the same structure (Fig. 5I). These alveolospheres displayed a basal-out epithelial polarity, with distal epithelial markers localized toward the lumen and proximal markers oriented outward. This dual expression pattern indicates an aberrant phenotype, likely arising from disordered differentiation in IPF AT2 cells33,34. To further characterize and quantify these differences, gene expression analysis was performed after one month of 3D culture to assess the expression of markers specific to alveolar and airway epithelial cells. Consistent with immunostaining, IPF-derived alveolospheres showed a marked loss of surfactant-producing identity, with significantly reduced Pro-SPC expression (Fig. 5J), and exhibited a proximalized epithelial phenotype (Fig. 5K).
IPF-derived AT2 cells form morphologically aberrant alveolospheres, characterized by the loss of distal identity and the acquisition of proximal features. (A–C) Alveolosphere morphology was broadly classified into cystic and densely folded structures. (A) Example of a cystic alveolosphere characterized by a hollow, lumen-like interior. Scale bar = 200 μm. (B) Example of a densely folded alveolosphere exhibiting compact epithelial organization. Scale bar = 200 μm. (C) IPF-derived alveolospheres displayed a unique hybrid morphology combining cystic and dense epithelial features, which was not observed in healthy AT2 3D cultures. Scale bar = 200 μm. (D) After one month of culture, healthy alveolospheres showed active proliferation, as indicated by KI67-positive staining. (E–F) In contrast, IPF-derived alveolospheres lacked KI67 expression. (G) Healthy donor alveolospheres expressed distal epithelial markers HTII-280, AQP5, and Pro-SPC. (H) IPF-derived alveolospheres exhibited weak HTII-280 expression and were Pro-SPC negative compared with healthy controls. (I) IPF-derived alveolospheres with hybrid morphology (from panel C) co-expressed proximal (KRT5) and distal (AQP5) markers within the same structure, indicating an aberrant mixed phenotype. (J) qPCR analysis of lineage markers demonstrated reduced expression of the distal marker SFTPC in IPF-derived alveolospheres (n = 9) compared with healthy controls (n = 3). (K) IPF-derived alveolospheres also showed increased expression of proximal airway-associated genes (n = 9) relative to healthy controls (n = 3), normalized to GAPDH. *p < 0.05, **p < 0.01 by Mann–Whitney two-tailed test.
Discussion
The primary objective of this study was to provide researchers with a comprehensive workflow, including an efficient protocol for enzyme-based lung tissue digestion and the automated purification of AT2 cells from both healthy and diseased lungs. The detailed methodology is intended to guide users in processing healthy and diseased lung tissue effectively, while clearly defined steps promote consistency and reproducibility across experiments. To enhance operator flexibility, we incorporated tissue storage in a preservation solution. Extending the processing window up to 36 h of delayed tissue processing without compromising cell quality. This enabled the initiation of digestion and downstream cell isolation to be done more conveniently. Building on the protocol described by Fujino et al.35, we present a new protocol with several modifications to the digestion procedure. The number of enzymes was minimized, using only dispase II and DNase; dispase has been widely reported in mouse and human AT2 cell extraction studies20,23,35,36,37,38,39,40,41,42,43,44. To enhance tissue processing, the lung tissue was injected through multiple distal airways and inflated to improve access and recovery of epithelial cells. Fine scissors were used for mechanical mincing both before and after enzymatic digestion, and the total digestion time was significantly reduced to a single 30-min round.
Rapid and gentle cell enrichment was achieved using the autoMACS Pro Separator through either a one-step or two-step protocol. The 2-step protocol is recommended for applications such as next-generation sequencing that require maximum cell purity. In contrast, the 1-step protocol may be preferred to conserve time and resources. Our results align with the growing trend towards purifying strategies that utilize either MACS Technologies and/or FACS, rather than older techniques using adherence culture, which usually yield around 80% pure AT2 cells45,46,47. Previous protocols have demonstrated that AT2 cell isolation using FACS, based on HTII-280 surface marker expression, achieves purities ranging from 89 to 100% with limited viability data19,20. Cell separation using MACS Technology offers higher throughput than FACS since multiple samples can be processed in parallel. However, these protocols often require additional optimization steps beyond the manufacturer’s specifications to ensure greater purity16,23. In a smaller cohort of samples, we compared MACS vs. FACS AT2 cell isolation. The table in Figure S5 compares AT2 cell isolation efficiency between MACS and FACS. Both techniques achieved comparable purity (95 ± 4%), while MACS showed slightly lower viability (92 ± 3%) than FACS (98 ± 1%). The difference in viability is likely due to the inclusion of a live-cell selection marker and the subsequent gating strategy applied to viable cells in the FACS protocol. This potentially inflated the apparent FACS viability, rather than indicating a true reduction in cell health within the MACS protocol. We also noticed considerable differences in terms of sorting duration. FACS consistently required over 3 h of continuous operation, and still could not process the entire input fraction, resulting in fewer cells collected per gram. In contrast, MACS isolation was completed in approximately 1 h, with the whole-cell input successfully processed, yielding substantially higher cell recovery.
In IPF-diseased lungs, AT2 cell numbers are reduced, reflecting both the loss of alveolar structures and the expansion of fibrotic regions. These areas are characterized by a dense extracellular matrix and excessive fibrillar collagen deposition48,49, making AT2 cell extraction more challenging. Only a handful of studies have attempted to isolate alveolar epithelial cells from IPF lungs30,50,51,52,53,54. Our literature review revealed that FACS sorting is commonly used to isolate cells from IPF tissue. While purity is often emphasized, many protocols lack data on cell viability or yield from the isolation process. Despite the isolation challenges presented by IPF tissue, our protocol successfully yielded high-purity AT2 cells through automated magnetic separation. Based on our viability and purity measurements, we can be confident in the robustness and reproducibility of this isolation strategy. Purified cells were expanded in an established 3D culture platform to ensure that the separation process did not affect cellular growth and phenotype27. The 3D culture conditions reduce the variability of signaling factors inherent to organotypic co-cultures, enabling a more controlled investigation of alveolar epithelial cell homeostasis6,7,55. Quantification of the CFE at Day 14 revealed that IPF-derived AT2 cells formed significantly fewer colonies, indicating impaired self-renewal and diminished growth compared to those from healthy donors. Consistent with this, fold expansion analysis at Day 28 demonstrated a reduction in proliferative potential, reflecting stem cell exhaustion and defective epithelial repair mechanisms characteristic of IPF56. The AT2 cells isolated using this workflow readily established alveolospheres and maintained their phenotypic characteristics in 3D culture. Morphological assessment revealed that the alveolospheres derived from healthy AT2 cells typically formed cystic or densely folded epithelial structures. In contrast, IPF-derived alveolospheres frequently displayed hybrid morphologies combining both features, an aberrant organization not observed in healthy cultures.
To investigate differentiation potential, we examined marker expression at the protein and transcript levels after one month of 3D culture. Immunostaining revealed that healthy alveolospheres expressed the distal epithelial markers HTII-280, AQP5, and Pro-SPC, whereas IPF-derived alveolospheres showed reduced Pro-SPC and weak HTII-280 expression. A subset of IPF alveolospheres co-expressed proximal (KRT5) and distal (AQP5) markers within the same structure, displaying a basal-out epithelial polarity with proximal markers oriented outward and distal markers localized luminally. Gene expression analysis further confirmed the decreased expression of distal genes, such as SFTPC, and the increased expression of proximal airway-associated genes in IPF-derived cultures. Of note, KRT17 has been reported to be present in a transitional, KRT17-positive state, where cells in fibrotic lungs become arrested in differentiation and acquire basal or airway-like features34,57,58. This shift in gene signature supports the conclusion that IPF AT2 cells engage in an aberrant repair program, which is consistent with defective epithelial regeneration in pulmonary fibrosis. The aberrant co-expression of proximal and distal markers suggests that IPF AT2 cells enter a transitional state, failing to differentiate into fully functional surfactant-producing cells. Organoids usually display marker profiles corresponding to their lung region of origin, expressing either proximal or distal markers. This co-expression might indicate an aberrant phenotype, likely due to disordered differentiation in IPF AT2 cells30,34,54,56. Such disordered epithelial remodeling likely compromises alveolar integrity and repair, contributing to the dysfunctional regenerative environment observed in fibrotic lungs.
In conclusion, our protocol establishes a reliable workflow for isolating primary human AT2 cells with high cell yield and purity, including from fibrotic lung tissue, offering scalability to generate sufficient quantities of cells. Beyond the technical optimization for both tissue sources, we also investigated the disease-associated differences by culturing healthy and IPF-derived AT2 cells in 3D alveolosphere systems. These studies revealed functional impairments in the regenerative and proliferative capacities, as well as disrupted differentiation patterns, in IPF-derived AT2 cells. Collectively, this workflow not only enables the large-scale generation of AT2 cells suitable for downstream molecular and functional assays but also provides a platform for modeling epithelial dysfunction and testing regenerative strategies in fibrotic lung disease.
Limitations of the study
Variability in the AT2 cell yield may reflect differences in the proportion of AT2 cells inherently present within the tissue, which can be influenced by factors such as donor variability, disease state, and the specific lung region sampled. Lung resections obtained in the operating room can differ considerably in anatomical location, size, and the proportion of the alveolar regions, all of which affect the abundance and accessibility of AT2 cells for MACS isolation. Furthermore, the proportion of AT2 cells within the bulk heterogeneous cell population before MACS labeling could not be determined, as data collection was conducted retrospectively. To help minimize this variability, we recommend collecting biopsies from the most distal regions of the lung, preferably the upper lobes, and shaping them into circular pieces to promote consistent inflation and enzymatic digestion.
Data availability
The datasets generated during this study are available from the corresponding author upon reasonable request.
References
Franks, T. J. et al. Resident cellular components of the human lung current knowledge and goals for research on cell phenotyping and function. Proc. Am. Thorac. Soc. 5 (7), 763–766. https://doi.org/10.1513/pats.200803-025HR (2008).
Castranova, V., Rabovsky, J., Tucker, J. H. & Miles, P. R. The alveolar type II epithelial cell: A multifunctional pneumocyte. Toxicol. Appl. Pharmacol. 93, 472–483 (1988).
Wang, Y. et al. Pulmonary alveolar type I cell population consists of two distinct subtypes that differ in cell fate. Proc. Natl. Acad. Sci. U S A. 115 (10), 2407–2412. https://doi.org/10.1073/pnas.1719474115 (2018).
Zacharias, W. J. et al. Regeneration of the lung alveolus by an evolutionarily conserved epithelial progenitor. Nature 555 (7695), 251–255. https://doi.org/10.1038/nature25786 (2018).
Olajuyin, A. M., Zhang, X. & Ji, H. L. Alveolar type 2 progenitor cells for lung injury repair. Cell. Death Discov. 5 (1), 63. https://doi.org/10.1038/s41420-019-0147-9 (2019).
Barkauskas, C. E. et al. Type 2 alveolar cells are stem cells in adult lung. J. Clin. Invest. 123 (7), 3025–3036. https://doi.org/10.1172/JCI68782 (2013).
Nabhan, A. N., Brownfield, D. G., Harbury, P. B., Krasnow, M. A. & Desai, T. J. Single-cell Wnt signaling niches maintain stemness of alveolar type 2 cells. Science 359 (6380), 1118–1123 (2018).
Zhao, F., Ma, Q., Yue, Q. & Chen, H. SARS-CoV-2 infection and lung regeneration. Clin. Microbiol. Rev. 35 (2), 1–25 (2022).
Serrano-Mollar, A. et al. Intratracheal transplantation of alveolar type II cells reverses Bleomycin-Induced lung fibrosis. Am. J. Respir Crit. Care Med. 176 (12), 1261–1268. https://doi.org/10.1164/rccm.200610-1491OC (2007).
Lopez-Rodriguez, E., Gay-Jordi, G., Knudsen, L., Ochs, M. & Serrano-Mollar, A. Improved alveolar dynamics and structure after alveolar epithelial type II cell transplantation in bleomycin induced lung fibrosis. Front. Med. (Lausanne) https://doi.org/10.3389/fmed.2021.640020 (2021).
Weiner, A. I. et al. Mesenchyme-free expansion and transplantation of adult alveolar progenitor cells: Steps toward cell-based regenerative therapies. NPJ Regen. Med. https://doi.org/10.1038/s41536-019-0080-9 (2019).
Guillamat-Prats, R., Gay-Jordi, G., Xaubet, A., Peinado, V. I. & Serrano-Mollar, A. Alveolar type II cell transplantation restores pulmonary surfactant protein levels in lung fibrosis. J. Heart Lung Transpl. 33 (7), 758–765. https://doi.org/10.1016/j.healun.2014.03.008 (2014).
Serrano-Mollar, A. et al. Safety and tolerability of alveolar type II cell transplantation in idiopathic pulmonary fibrosis. Chest 150 (3), 533–543. https://doi.org/10.1016/j.chest.2016.03.021 (2016).
González-González, M., Vázquez-Villegas, P., García-Salinas, C. & Rito-Palomares, M. Current strategies and challenges for the purification of stem cells. J. Chem. Technol. Biotechnol. 87 (1), 2–10. https://doi.org/10.1002/jctb.2723 (2012).
Zhu, B. & Murthy, S. K. Stem cell separation technologies. Curr. Opin. Chem. Eng. 2 (1), 3–7. https://doi.org/10.1016/j.coche.2012.11.002 (2013).
Sutermaster, B. A. & Darling, E. M. Considerations for high-yield, high-throughput cell enrichment: Fluorescence versus magnetic sorting. Sci. Rep. https://doi.org/10.1038/s41598-018-36698-1 (2019).
Bacon, K. et al. Present, and future of affinity-based cell separation technologies. Acta Biomater. 112, 29–51. https://doi.org/10.1016/j.actbio.2020.05.004 (2020).
Thiel, A., Scheffold, A. & Radbruch, A. Immunomagnetic cell sorting-pushing the limits. Immunotechnology 4, 89–96 (1998).
Pan, J. & Wan, J. Methodological comparison of FACS and MACS isolation of enriched microglia and astrocytes from mouse brain. J. Immunol. Methods https://doi.org/10.1016/j.jim.2020.112834 (2020).
Hasegawa, K. et al. Fraction of MHCII and EpCAM expression characterizes distal lung epithelial cells for alveolar type 2 cell isolation. Respir. Res. https://doi.org/10.1186/s12931-017-0635-5 (2017).
Corti, M., Brody, A. R. & Harrison, J. H. Isolation and primary culture of murine alveolar type II cells. Am. J. Respir Cell. Mol. Biol. 14 (4), 309–315. https://doi.org/10.21769/bioprotoc.2288 (1996).
Rice, W. R. et al. Maintenance of the mouse type II cell phenotype in vitro. Am. J. Physiol. Lung Cell. Mol. Physiol. 283 (2), 256–264. https://doi.org/10.1152/ajplung (2002).
Janas, P. P. et al. Cold dispase digestion of murine lungs improves recovery and culture of airway epithelial cells. PLoS ONE https://doi.org/10.1371/journal.pone.0297585 (2024).
Marsh, L. M. et al. Surface expression of CD74 by type II alveolar epithelial cells: A potential mechanism for macrophage migration inhibitory Factor-Induced epithelial repair. Am. J. Physiol. Lung Cell. Mol. Physiol. 296, 442–452. https://doi.org/10.1152/ajplung.00525.2007.-Macrophage (2009).
Messier, E. M., Mason, R. J. & Kosmider, B. Efficient and rapid isolation and purification of mouse alveolar type II epithelial cells. Exp. Lung Res. 38 (7), 363–373. https://doi.org/10.3109/01902148.2012.713077 (2012).
Gonzalez, R. F., Allen, L., Gonzales, L., Ballard, P. L. & Dobbs, L. G. HTII-280, a biomarker specific to the apical plasma membrane of human lung alveolar type II cells. J. Histochem. Cytochem. 58 (10), 891–901. https://doi.org/10.1369/jhc.2010.956433 (2010).
Youk, J. et al. Three-Dimensional human alveolar stem cell culture models reveal infection response to SARS-CoV-2. Cell. Stem Cell. 27 (6), 905–919 (2020).
Dekkers, J. F. et al. High-Resolution 3D imaging of fixed and cleared organoids. Nat. Protoc. 14 (6), 1756–1771. https://doi.org/10.1038/s41596-019-0160-8 (2019).
Liang, J. et al. Hyaluronan and TLR4 promote Surfactant-Protein-C-Positive alveolar progenitor cell renewal and prevent severe pulmonary fibrosis in mice. Nat. Med. 22 (11), 1285–1293. https://doi.org/10.1038/nm.4192 (2016).
Xu, Y. et al. Single-cell RNA sequencing identifies diverse roles of epithelial cells in idiopathic pulmonary fibrosis. JCI Insight https://doi.org/10.1172/jci.insight.90558 (2016).
Parimon, T., Yao, C., Stripp, B. R., Noble, P. W. & Chen, P. Alveolar epithelial type II cells as drivers of lung fibrosis in idiopathic pulmonary fibrosis. Int. J. Mol. Sci. https://doi.org/10.3390/ijms21072269 (2020).
Hoffmann, K. et al. Human alveolar progenitors generate dual lineage bronchioalveolar organoids. Commun. Biol. https://doi.org/10.1038/s42003-022-03828-5 (2022).
Marmai, C. et al. Alveolar epithelial cells express mesenchymal proteins in patients with idiopathic pulmonary fibrosis. Am. J. Physiology- Lung Cell. Mol. Physiol. 301, L71–L78. https://doi.org/10.1152/ajplung.00212.2010.-Prior (2011).
Choi, H. K. et al. Regenerative capacity of alveolar type 2 cells is proportionally reduced following disease progression in idiopathic pulmonary fibrosis-derived organoid cultures. Tuberc. Respir. Dis. (Seoul). 88 (1), 130–137. https://doi.org/10.4046/trd.2024.0094 (2025).
Fujino, N. et al. Isolation of alveolar epithelial type II progenitor cells from adult human lungs. Lab. Invest. 91 (3), 363–378. https://doi.org/10.1038/labinvest.2010.187 (2011).
Quantius, J. et al. Influenza virus infects epithelial stem/progenitor cells of the distal lung: Impact on Fgfr2b-driven epithelial repair. PLoS Pathog. https://doi.org/10.1371/journal.ppat.1005544 (2016).
Kadur Lakshminarasimha Murthy, P. et al. Human distal lung maps and lineage hierarchies reveal a bipotent progenitor. Nature 604 (7904), 111–119. https://doi.org/10.1038/s41586-022-04541-3 (2022).
Warshamana, G. S., Corti, M. & Brody, A. R. TNF-α, PDGF, and TGF-Β1 expression by primary mouse Bronchiolar-Alveolar epithelial and mesenchymal cells: TNF-α induces TGF-Β1. Exp. Mol. Pathol. 71 (1), 13–33. https://doi.org/10.1006/exmp.2001.2376 (2001).
Lv, Z. et al. Alveolar regeneration by airway secretory-cell-derived P63 + Progenitors. Cell. Stem Cell. 31 (11), 1685–1700. https://doi.org/10.1016/j.stem.2024.08.005 (2024).
Brownfield, D. G. et al. Alveolar cell fate selection and lifelong maintenance of AT2 cells by FGF signaling. Nat. Commun. https://doi.org/10.1038/s41467-022-34059-1 (2022).
Naikawadi, R. P. et al. Telomere dysfunction in alveolar epithelial cells causes lung remodeling and fibrosis. JCI Insight https://doi.org/10.1172/jci.insight.86704 (2016).
Bortnick, A. E. et al. Identification and characterization of rodent ABCA1 in isolated type II pneumocytes. Am. J. Physiol. Lung Cell. Mol. Physiol. 285(4), 869–878. https://doi.org/10.1152/ajplung.00077.2003.-ATP-bind (2003).
Chung, K. P. et al. Alveolar epithelial cells mitigate neutrophilic inflammation in lung injury through regulating mitochondrial fatty acid oxidation. Nat. Commun. https://doi.org/10.1038/s41467-024-51683-1 (2024).
Bondonese, A. et al. Impact of enzymatic digestion on single cell suspension yield from peripheral human lung tissue. Cytometry Part. A. 103 (10), 777–785. https://doi.org/10.1002/cyto.a.24777 (2023).
Wang, J. et al. Differentiated human alveolar epithelial cells and reversibility of their phenotype in vitro. Am. J. Respir Cell. Mol. Biol. 36 (6), 661–668. https://doi.org/10.1165/rcmb.2006-0410OC (2007).
Kosmider, B. et al. Nrf2 protects human alveolar epithelial cells against injury induced by influenza A virus. Respir. Res. https://doi.org/10.1186/1465-9921-13-43 (2012).
Moimas, S. et al. MiR-200 family members reduce senescence and restore idiopathic pulmonary fibrosis type II alveolar epithelial cell transdifferentiation. ERJ Open Res. https://doi.org/10.1183/23120541.00138 (2019).
Thannickal, V. J. et al. Matrix biology of idiopathic pulmonary fibrosis: A workshop report of the National Heart, Lung, and blood Institute. Am. J. Pathol. 184 (6), 1643–1651. https://doi.org/10.1016/j.ajpath.2014.02.003 (2014).
Plantier, L. et al. Physiology of the lung in idiopathic pulmonary fibrosis. Eur. Respir. Rev. https://doi.org/10.1183/16000617.0062-2017 (2018).
Newton, D. A., Lottes, R. G., Ryan, R. M., Spyropoulos, D. D. & Baatz, J. E. Dysfunctional lactate metabolism in human alveolar type II cells from idiopathic pulmonary fibrosis lung explant tissue. Respir. Res https://doi.org/10.1186/s12931-021-01866-x (2021).
Liang, J. et al. Lipid deficiency contributes to impaired alveolar progenitor cell function in aging and idiopathic pulmonary fibrosis. Am. J. Respir Cell. Mol. Biol. 71 (2), 242–253. https://doi.org/10.1165/rcmb.2023-0290OC (2024).
Yao, C. et al. Senescence of alveolar type 2 cells drives progressive pulmonary fibrosis. Am. J. Respir. Crit. Care Med. 203 (6), 707–717. https://doi.org/10.1164/rccm.202004-1274OC (2021).
Wu, H. et al. Progressive pulmonary fibrosis is caused by elevated mechanical tension on alveolar stem cells. Cell 180 (1), 107–121. https://doi.org/10.1016/j.cell.2019.11.027 (2020).
Kathiriya, J. J. et al. Human alveolar type 2 epithelium transdifferentiates into metaplastic KRT5 + Basal cells. Nat. Cell. Biol. 24 (1), 10–23. https://doi.org/10.1038/s41556-021-00809-4 (2022).
Liao, D. & Li, H. Dissecting the niche for alveolar type II cells with alveolar organoids. Front. Cell. Dev. Biol. https://doi.org/10.3389/fcell.2020.00419 (2020).
Chakraborty, A., Mastalerz, M., Ansari, M., Schiller, H. B. & Staab-Weijnitz, C. A. Emerging roles of airway epithelial cells in idiopathic pulmonary fibrosis. Cells https://doi.org/10.3390/cells11061050 (2022).
Wang, F. et al. Regulation of epithelial transitional states in murine and human pulmonary fibrosis. J. Clin. Invest. https://doi.org/10.1172/JCI165612 (2023).
Kobayashi, Y. et al. Persistence of a regeneration-associated, transitional alveolar epithelial cell state in pulmonary fibrosis. Nat. Cell. Biol. 22 (8), 934–946. https://doi.org/10.1038/s41556-020-0542-8 (2020).
Acknowledgements
We want to acknowledge the members of the Latner Thoracic Laboratories and the Biobank team of the Toronto Lung Transplant Program for their unwavering support in providing the necessary resources and assisting in the procurement of lung samples. We also wish to acknowledge James Jonkman, Judith Cathcart, and Courtney McIntosh from The Advanced Optical Microscopy Facility at the University Health Network for guidance and support with imaging analysis. We want to sincerely thank Emily Reddy and Bruno Rodrigues from the SickKids-UHN Flow Cytometry Core Facility for their support with the operation of the autoMACS Pro Separator. Finally, we sincerely thank Miltenyi Biotec for providing essential reagents that contributed to the successful completion of this work.
Funding
This work was supported by the Canadian Institutes of Health Research (CIHR), Application Number 407666.
Author information
Authors and Affiliations
Contributions
A.L.E., T.K.W., and G.K. conceived and designed the project. A.L.E. performed the experiments, analyzed the data, and wrote the manuscript. T.K.W. and G.K. supervised the project and guided the analysis. S.K. ensured the study’s alignment with UHN REB protocols and oversaw the collection of biopsies. A.L.E. and M.C. performed the biopsy collection. A.L.E., M.S., J.L., J.I., and D.T. performed lung tissue processing, including AT2 cell separation, using the autoMACS Pro Separator. A.L.E., M.S., and J.L. performed *in vitro* culture of AT2 cells. A.L.E. and J.L. performed immunofluorescent staining of alveolospheres. M.S. assisted with data and statistical analyses. All authors contributed to manuscript preparation. S.K., T.K.W., and G.K. reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Eiliazadeh, A.L., Suma, M.A.V., Li, J. et al. High-throughput isolation of normal and pulmonary fibrosis human alveolar epithelial cells. Sci Rep 16, 757 (2026). https://doi.org/10.1038/s41598-025-30242-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-30242-8