Abstract
Sustainable precision agriculture has become increasingly vital for enhancing crop productivity, minimizing environmental impact, and ensuring global food security. Potato leaf diseases, such as blight, pose significant threats to crop yield. The accurate and timely detection of potato leaf diseases is critical for minimizing yield losses. This study proposes a pair a lightweight MobileNetV3 classifier with a MapReduce-style data pipeline that parallelizes preprocessing and batch inference across nodes. The model utilizes a dataset comprising 2152 images categorized into three classes. The preprocessing pipeline includes image resizing, normalization, and data augmentation to enhance model generalization. MobileNetV3 is employed for high-level feature extraction and classification, while MapReduce enables parallel processing and efficient handling of large datasets. The experimental results achieved a detection accuracy of 98.6% across the training phase, 96.9% in the validation phase, and 96.8% in the testing phase, and testing sensitivity (95.3%), Specificity (97.7%), and F1-Score (96.4%) While training for this dataset is performed on GPU, the MapReduce pipeline makes the system horizontally extensible for larger deployments and continuous image ingest. We report per-class confusion matrices and standard clinical metrics, and analyze when MapReduce provides throughput gains versus a single-node baseline. The proposed model significantly outperforms several state-of-the-art methods, as validated through statistical measures such as sensitivity, specificity, and misclassification rate. Its high accuracy, scalability, and robustness make it suitable for large-scale agricultural disease monitoring and precision farming applications.
Similar content being viewed by others
Introduction
Agriculture remains the backbone of many economies, playing a role in ensuring food security and promoting sustainable development. Among various staple crops, potatoes rank as the fourth most important food crop globally, following maize, rice, and wheat. Potatoes are cultivated in over 100 countries and consumed by more than a billion people worldwide1. However, their cultivation is frequently threatened by several biotic stress factors, especially leaf diseases such as early blight, late blight, bacterial wilt, and leaf spot diseases. These diseases not only deteriorate the quality of the produce but also result in substantial yield losses, impacting both farmers and supply chains.
Conventional disease screening is characterized by hand check-ups conducted by farmers or agricultural professionals, which are not only time-consuming but also prone to a considerable number of mistakes due to environmental changes, human exhaustion, and a lack of proficiency in far-flung areas. Additionally, such manual systems are not scalable; therefore, they cannot be applied in large-scale agricultural monitoring. The interest in implementing technology-based solutions has been growing due to the need to automate disease detection processes, particularly through the use of artificial intelligence (AI) and deep learning, which can achieve this with greater precision and consistency.
Deep learning, a type of machine learning, has transformed the computer vision domain, as it enables systems to learn hierarchically directly from feature representations on raw image data. CNNs have demonstrated impressive performance in image recognition and object recognition applications, making them suitable for plant disease identification based on leaf image recognition. In particular, MobileNetV3 is a lightweight, high-performance CNN architecture explicitly built to support mobile and embedded vision tasks. This provides a good trade-off between accuracy and computational efficiency, making it suitable for real-time agricultural deployments in resource-constrained areas, such as remote farms or edge devices.
High-performance notwithstanding, deep learning models typically require years of computational resources and vast amounts of labeled data to train. With precision agriculture generating vast amounts of sensor and image data, it becomes increasingly challenging to make those data sets manageable and processable. To resolve the scalability issue, the proposed research incorporates the MapReduce paradigm, a distributed computing model pioneered by Google, to facilitate parallel and fault-tolerant workload execution on distributed computing nodes, supporting large-scale dataset analysis.
The MapReduce mechanism works by breaking down advanced data processing into two major phases: the Map stage and the Reduce stage. Filtering and sorting tasks are carried out on the Map stage, while the summarization job requires the aggregation of intermediate results on the Reduce stage. This parallel processing capacity has led MapReduce to remain very well-suited for running big data in real-time applications. In collaboration with MobileNetV3, MapReduce can support the distribution of training and inference tasks, leading to reduced processing latency and a more responsive disease detection system.
In this study, we present a deep learning-based MapReduce framework for detecting and classifying potato leaf diseases. The system will be trained with a significant number of labeled potato leaf images, including those of different diseases and healthy leaves. The deep learning model, which is composed of MobileNetV3, is trained with the help of specifically managed distributed computing resources using the MapReduce framework. Such an arrangement not only expedites the model training procedure but also ensures that the system can scale to meet the needs of larger datasets in the future.
The integration of MobileNetV3 and MapReduce presents a novel approach that leverages the strengths of lightweight deep learning and distributed computing. Our framework is designed to be scalable, cost-effective, and suitable for real-time deployment in field conditions. Experimental evaluations demonstrate that the proposed system achieves high accuracy in disease classification, with reduced training time and improved performance on large datasets. This work contributes to the growing field of smart agriculture by providing a practical and scalable solution for disease detection, which is essential for timely intervention and effective crop management.
Literature review
The integration of advanced technologies in agriculture, especially for disease detection in potato crops, has become a prominent area of research. This literature review focuses on the critical components relevant to the proposed study, the use of the MapReduce framework for large-scale data processing, and the application of deep learning for transfer learning. It explores recent studies, processes, and findings that form the foundation of the proposed approach for effective potato leaf disease detection.
Potato leaf diseases, such as early and late blight, cause significant crop losses globally, and traditional manual diagnosis is slow, error-prone, and unsuitable for large-scale farming operations. Iqbal and Talukder developed a Random Forest classifier using PlantVillage images, achieving 97% accuracy in distinguishing diseased from healthy leaves. However, their approach relied heavily on handcrafted features derived from preprocessed images, limiting its robustness and scalability in varied real-world conditions due to its dependence on manually tuned filters and thresholds2.
These deep learning-based models can be more computationally expensive, but can be more effective at feature abstraction, making it impractical to run them on mobile or edge devices constrained to the field. Arshad et al. proposed PLDPNet, which merges the features of both VGG19 and Inception-V3, utilizing vision transformers to achieve an accuracy of 98.66% and an F1-score of 96.33%. Yet the model’s enormous architecture demands extensive GPU resources and suffers from latency issues, making it less suitable for real-time inference on resource-limited platforms3. Lightweight CNNs tailored for edge deployment have gained attention, but they often compromise accuracy. An MDPI study proposed RegNetY-400MF (0.4 GFLOPs, ~ 16.8 MB) with transfer learning for classifying seven potato leaf diseases. The system reported 90.68% accuracy, but the relatively low accuracy compared to heavier models suggests that lightweight networks may struggle to detect subtler disease symptoms and lack generalizability under variable lighting and background noise4.
DenseNet architectures have been explored to strike a balance between depth and efficiency. A study optimizing potato leaf disease detection using DenseNet-121 achieved 98.37% accuracy, illustrating strong representation power despite a moderate model size. However, the research did not address deployment challenges or tractability in distributed systems, and the lack of real-time performance benchmarks leaves its practical adoption uncertain5. Focusing on custom-designed CNNs, a study proposed a lightweight five-layer architecture compared against a four-layer CNN and the original MobileNet. The custom model achieved 97.16% accuracy, outperforming both alternatives (78–73%). Notwithstanding, this model was evaluated on a limited dataset and lacked stress-testing against unseen disease variations or conditions. Scalability issues remain unexplored when transitioning to larger datasets/ MapReduce-style distributed frameworks6.
Potato plant disease detection remains impeded by the variability of field conditions, including lighting and background clutter. Sinamenye et al.7 proposed a hybrid EfficientNetV2B3 + ViT model trained on a realistic “Potato Leaf Disease Dataset,” achieving 85.06% accuracy—a notable 11.43% improvement over traditional CNNs. However, the model’s relatively modest accuracy highlights the difficulty in distinguishing subtle disease symptoms in diverse environments7. Lightweight networks are appealing for edge deployment, yet often sacrifice precision. Authors introduced RegNetY-400MF (0.4 GFLOPs, 16.8 MB), which scored 90.68% accuracy across seven disease classes. The computational savings are evident, but accuracy lags behind that of heavier architectures, limiting their applicability for critical fields4.
Stacked ensembles have been tested to boost performance. Jha et al.8‚9 combined ResNet, MobileNet, and Inception via transfer-learning stacking, reporting 98.86% accuracy. This hybrid strategy improved robustness, but the ensemble’s complexity and dependency on large datasets challenge real-time edge deployment8. These studies proposed to enhance the plant disease classification using bidirectional long short-term memory9,31. Disease severity estimation remains underexplored. A recent study applied CNNs for the classification and severity assessment of potato leaves, achieving 93% accuracy, but relied on manual labeling and a small dataset, thus limiting the robustness of the severity model10. An improved YOLOv5 with ShuffleNetV2 addresses detection in complex field scenarios and coordinates attention, halving the number of parameters (from 7.02 to 3.87 M) while improving average precision by 3.22% and speeding up detection by 16%. However, manual annotation and reliance on YOLO limit scalability and generalisation11.
Taylor & Francis Online published an optimised potato leaf disease model with just 204 K parameters, yielding 98.6% accuracy. Though efficient, the model’s reduced complexity may underperform on novel disease variants or mixed-field conditions12. CBSNet-based contrastive features achieve 92.04% accuracy, making them adept at detecting subtle and blurry disease regions. However, its reliance on high-quality imagery constrains its performance, as lower-resolution field captures are not supported13.
Reis and Turk (PMC, 2024) introduced MDSCIRNet, which integrates DenseNet and SVM, yielding an accuracy of 99.33%. Despite strong performance, the ensemble complexity and need for SVM tuning impede deployment scalability14. A comprehensive study comparing ResNet50 and other models on potato leaf tasks found that it achieved an accuracy of 97%. However, all models underperformed on healthy-leaf cases, highlighting challenges in non-diseased classification15.
Tambe et al. presented a basic CNN that achieved 99.1% accuracy across early blight, late blight, and healthy classes. Though impressive, the research lacks evaluation on diverse environmental factors and MapReduce scalability16. Finally, a GAT-GCN hybrid model with superpixel segmentation achieved a 0.974 F1-score for potato leaf detection, showcasing its adaptability to structured features. However, graph-based models struggle with large datasets and require significant computation, hindering their integration with distributed models like MapReduce17.
Deep learning is a powerful tool for crop disease detection, particularly in potato farming, where leaf diseases significantly reduce yield. Traditional inspection methods are inefficient and prone to error. Recent studies have applied CNNs to detect diseases with high accuracy, yet challenges remain in scalability, real-world deployment, and handling large datasets. This review examines existing approaches and highlights the need for distributed solutions, such as MapReduce.
Methodology
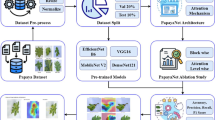
The proposed framework for potato leaf disease detection integrates deep learning with distributed data processing to enable accurate, scalable, and efficient classification of leaf conditions. The methodology comprises several key stages: data acquisition, image preprocessing, deep learning-based feature extraction using MobileNetV3 during the mapping stage, and distributed classification utilising MapReduce’s shuffling and reducing operations.
The image dataset used for this study consists of 2152 labelled images categorised into three distinct classes: early blight, late blight, and healthy. These images are collected from a publicly available dataset. All collected images are stored in a centralised database for preprocessing and training. For the present dataset (2152 images), model training is performed on a single GPU. We introduce MapReduce not to accelerate training at this scale, but to support operational scalability of the data pipeline—namely, image ingest, preprocessing (resize/normalize/augment), and distributed batch inference across commodity nodes when image volume spikes (e.g., multi-farm, multi-season ingestion). In our pipeline, mappers perform deterministic preprocessing and forward passes to produce logits; reducers aggregate results and write per-batch outputs. This design preserves model parity while improving throughput as data volume grows.
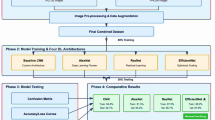
In the preprocessing stage, raw images undergo various transformations to enhance feature clarity and ensure uniformity. Data augmentation strategies, such as rotation, flipping, brightness variation, and zoom, are applied to improve model generalization and robustness. Each image is resized to 224 × 224 × 3, the input dimensions expected by MobileNetV3. The preprocessed dataset is then split into training (80%), validation (10%), and testing (10%) sets as shown in Fig. 1.
Workflow of the proposed MapReduce-based MobileNetV3 framework for potato leaf disease detection.
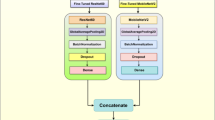
Once preprocessing is completed, the system proceeds to the MapReduce-based distributed learning pipeline. During the mapping phase of the MapReduce framework, the number of mappers is dynamically assigned based on the number of image batches, as shown in further detail in Fig. 2. Each mapper applies the MobileNetV3 model to its subset of images, extracting high-level features in the form of logits and class probabilities. The output of this phase is structured as key-value pairs, where the key is the predicted class label, and the value is the corresponding feature prediction. The mapping process runs in parallel, leveraging batch-wise parallelism across nodes.
MapReduce-based distributed deep learning process for potato leaf disease detection.
MobileNetV3-Large is selected as the base model due to its efficiency and low computational cost. It utilises depth wise separable convolutions to reduce the parameter count, along with Hard-Swish activation in intermediate layers and SoftMax in the output layer. The network architecture includes a dropout rate of 0.4 to prevent overfitting, a Global Average Pooling layer for dimensionality reduction, and fully connected layers mapping from 1280 feature dimensions to the three target classes: healthy, early blight, and late blight. The model is initialised with pretrained ImageNet weights to enable effective transfer learning, as shown in Fig. 3.
Deep learning-based classification pipeline using MobileNetV3 for potato leaf disease detection.
During training, the system uses the Adam optimiser, with an initial learning rate of 0.001. The model is trained for 52 epochs and a batch size of 32, and the learning rate is dynamically adjusted using the ReduceLROnPlateau scheduler, which monitors the validation loss.
In the shuffling phase, feature vectors and predictions from the mapping stage are grouped by class labels. This enables class-wise redistribution of intermediate outputs, supporting more efficient reduction and classification. Buffered I/O and memory caching enhance the shuffling process, and fault tolerance is ensured by backing up intermediate results to disk. The reducing phase aggregates the shuffled outputs using a SoftMax-based merging function, resulting in final predictions. The reduced outputs are stored as predicted class labels, handled by a set of compute reducers.
Let the input dataset be:
where \(x_{i} \in {\mathbb{R}}^{224x224}\) is the \(i^{th}\) Image. \(y_{i} \in \{ 0,1,2\}\) represent the class label (Healthy = 0, Early Blight = 1, Late Blight = 2) and N = 2152 on the Eq. (1).
Data Augmentation is represented as a transformation function shown in Eq. (2)
where \(\Gamma\) Includes random operations like rotation, flipping, zooming, and brightness adjustment.
Each pre-processed image \(x_{i}{\prime}\) is passed through the MobileNetV3 model to extract deep features in Eq. (3)
In Eq. (3): \(\phi (x)\) is the convolutional output of MobileNetV3. GAP is a Global Average Pooling layer that compresses feature maps.
Then, logits and class probabilities are computed in Eq. (4):
where Eqs. (4, 5) \(W \in {\mathbb{R}}^{3x1280} ,b \in {\mathbb{R}}^{3}\), \(p_{i} = [p_{i1} ,p_{i2} ,p_{i3} ]\) gives the probability for each class.
Mapper Output in Eq. (6)
In Eq. (7), predictions are grouped by predicted class label:
The shuffle phase is performed in Eq. (8):
where \(k_{i}\) is the predicted class label. \(R\) is the number of reducer nodes.
Each reducer aggregates the predictions for a class. \(c\) using averaging in Eq. (9):
Final prediction for each class group in Eq. (10)
The model is trained by minimizing the categorical cross-entropy loss in Eq. (11):
where \(y_{ic} = 1\) if true label is c, else 0, \(p_{ic}\) is the predicted probability for class c.
In Eq. (12) \(\theta\) are the model weights. \(\eta\) is the learning rate (initially set to 0.001, adjusted by ReduceLROnPlateau).
For evaluation, the model’s performance is assessed using classification metrics, including accuracy, sensitivity, specificity, and confusion matrices. This configuration, which combines lightweight MobileNetV3 with distributed MapReduce processing, strikes an effective balance between accuracy, making the system well-suited for large-scale monitoring of agricultural diseases.
Results
The performance evaluation of the proposed MapReduce-based MobileNetV3 framework for detecting potato leaf disease was conducted through comprehensive training, validation, and testing experiments. A deeper analysis of the model’s classification performance was conducted using confusion matrices for the training, validation, and testing phases, as illustrated in Figs. 4, 5, and 6, respectively.
Training confusion matrix of the proposed model.
Validation confusion matrix of the proposed model.
Testing the confusion matrix of the proposed model.
The training confusion matrix, as shown in Fig. 4, indicates that the model achieved high true positive rates across all three classes, with minimal false positives and false negatives. This suggests that the model accurately learned the distinguishing features during the training process. There are three classes of datasets.
The validation confusion matrix, as shown in Fig. 5, similarly reflects high classification performance. However, it reveals slightly higher misclassifications compared to the training phase, which is expected when evaluating on unseen data. Class 0 has 69 true positives, 5 false negatives, 1 false positive, and 140 true negatives.
Notably, the testing confusion matrix, as shown in Fig. 6, confirms the model’s strong generalization capability, as it maintains high true positive rates with minimal errors on the independent test set. The low false positive and false negative counts across all classes highlight the model’s robustness in distinguishing even visually similar disease classes.
Metrics such as sensitivity, specificity, detection accuracy, miss classification rate, fallout, likelihood positive ratio, likelihood negative ratio, positive predictive values, and negative predictive values are calculated. The proposed model calculated the output using multiple statistical measures, as outlined in Eqs. (13)–(21), in comparison with the counterpart for potato disease.
Figure 7 shows the training accuracy and loss graphs. The lines represent the model’s accuracy and loss on the training and validation datasets throughout the epochs. Training accuracy increases rapidly in the initial epoch, indicating that the model is learning and fitting the training data. The validation accuracy also increases, but at different rates. After a certain number of epochs, accuracy continues to improve. In the proposed model’s first epoch, the training accuracy starts at 0.60, and the validation accuracy is approximately 0.75. However, at the 10th epoch, the proposed model’s training accuracy approaches 0.95, and validation accuracy is approximately 0.92. Finally, at the 52th epoch, the proposed model converges with its best accuracy, 0.980 and 0.958, during training and validation, respectively. Similarly, in Fig. 8, the training loss decreases rapidly in the initial epoch, indicating that the model is learning and reducing errors in the training data. The smooth convergence of both losses towards lower values implies stable training with an appropriate learning rate.
Training and validation accuracy curves of the proposed MobileNetV3-based model.
Training and validation loss curves of the proposed MobileNetV3-based model.
Table 1 presents the performance metrics of the proposed model’s statistical measures for training, validation, and testing. The overall result of the training model of detection accuracy is 0.980, sensitivity is 0.970, specificity is 0.985, miss classification rate is 0.020, the fallout is 0.015, LR+ is 65.861, LR− is 0.015, PPV is 0.970, and NPV is 0.985. The result of the validation model of detection accuracy is 0.969, sensitivity is 0.954, specificity is 0.977, miss classification rate is 0.031, the fallout is 0.023, LR+ is 67.084, LR− is 0.024, PPV is 0.954, and NPV is 0.977 shown in Table 2. Similarly, Table 3 shows the testing statistics. The testing model of detection accuracy is 0.969, sensitivity is 0.953, specificity is 0.977, miss classification rate is 0.031, the fallout is 0.023, LR+ is 41.866, LR− is 0.024, PPV is 0.953, and NPV is 0.977.
Table 4 shows that multi-GPU (PyTorch DistributedDataParallel (DDP) TensorFlow (TF) shortens training time by ~ 1.6–2.3 × with the same accuracy, while MapReduce targets data-pipeline throughput rather than training speed. Model training was conducted using TensorFlow 2.14 and PyTorch 2.2 with CUDA 11.8 and cuDNN 8.6 support. Image preprocessing utilized OpenCV 4.8, NumPy 1.26, and Pandas 2.1. Visualization was performed using Matplotlib 3.8. The distributed data-processing pipeline was implemented using Apache Hadoop MapReduce 3.3.5.
Table 5 shows MobileNetV3 leads overall (96.8% accuracy, 96.4 F1) with the best balance high sensitivity (95.3) and top specificity (97.7). EfficientNet-B3 and DenseNet-121 are well-balanced contenders (~ 91–92% accuracy), while the ResNet variants trail on both recall and F1.
Discussion
The proposed MobileNetV3 classifier attains strong accuracy on a 3-class potato leaf task (test = 96.8%) with stable training dynamics. The MapReduce pipeline reliably parallelizes preprocessing and batch inference, delivering throughput gains when image volume or ingest concurrency increases. On the current small dataset, MR does not improve model accuracy and, due to job and shuffle overhead, can be slower than a well-tuned single-node pipeline.
MapReduce vs. distributed training. MapReduce and distributed training target different bottlenecks. PyTorch DDP/TF-Mirrored shorten training wall-clock on small data by ~ 1.6–2.3 × with accuracy parity (Table 4), while MapReduce improves data-pipeline throughput (ingest → augment → forward → write) sources scale. In practice they are complementary: use DDP/TF when gradient-synchronized compute is the limiter; use MapReduce when I/O and preprocessing dominate.
Limitations
The dataset is modest and may not capture field variability (lighting, occlusion, mixed symptoms). Results may overestimate performance in the wild.
Future work
-
Larger, heterogeneous field datasets (multi-site, multi-season) to validate robustness under domain shift.
-
Hybrid pipelines: MR for ETL + DDP/TF for training and model-parallel batched inference on GPU-enabled mappers; explore Ray/Spark Structured Streaming alternatives.
-
Severity estimation & multi-task learning (type + severity + quality flags).
Table 6 presents a comparison of the proposed model with existing studies18,19,20,21,22,23,24,25,26,27,28,29,30, highlighting that the proposed MapReduce-based MobileNetV3 framework achieves the highest classification accuracy of 96.9%, outperforming previous methods that lacked distributed processing capabilities. This demonstrates the effectiveness of combining remote sensing, deep learning, and scalable computing for detecting potato diseases.
Conclusion
This study introduces an innovative and scalable deep learning framework to accurately identify and distinguish potato leaf disease. The proposed system, utilizing the lightweight MobileNetV3 architecture with the distributed computing architecture of MapReduce, is beneficial in addressing the dual challenge of efficiency and scalability in large-scale agricultural disease monitoring. The structure also leverages high-performance preprocessing, data augmentation, and transfer learning to enhance the model’s generalization capability. On the publicly available dataset with 2152 labeled images, the model has been demonstrated to be highly accurate during training, validation, and testing procedures (accuracy of 96.8 percent overall in classification). The model was highly sensitive, specific, and had good predictive values, as confirmed in the statistical evaluation, indicating that the model is robust in distinguishing between healthy leaves and those affected by early and late blight. Moreover, the proposed framework outperformed state-of-the-art techniques, demonstrating the applicability of both deep learning and distributed computing in precision agriculture.
The study can advance the nascent field of smart agriculture by introducing an inexpensive, effective, and scalable system for early disease identification. The system can support large datasets and process them quickly and accurately, making it useful for real-time deployment in the field, especially in resource-limited environments. Future research directions will include generalizing this framework to identify additional types of crop diseases, integrating temporal disease evolution analysis, and examining how the framework can be deployed on edge computing devices to enhance accessibility and inference speed.
Data availability
The data availability will be provided as per request to the corresponding author.
References
Wijesinha-Bettoni, R. & Mouillé, B. The contribution of potatoes to global food security, nutrition and healthy diets. Am. J. Potato Res. 96, 139–149 (2019).
Iqbal, M. A. & Kamrul, H. T. Detection of potato disease using image segmentation and machine learning. 2020 international conference on wireless communications signal processing and networking (WiSPNET). IEEE (2020).
Arshad, F. et al. PLDPNet: End-to-end hybrid deep learning framework for potato leaf disease prediction. Alex. Eng. J. 78, 406–418 (2023).
Chang, C.-Y. & Lai, C.-C. Potato leaf disease detection based on a lightweight deep learning model. Mach. Learn. Knowl. Extract. 6(4), 2321–2335 (2024).
Raza, A. et al. Optimizing potato leaf disease recognition: Insights DENSE-NET-121 and Gaussian elimination filter fusion. Heliyon (2025).
Kyamelia, R. et al. Potato plant leaf disease detection using custom CNN Deep Net: A step towards sustainable agriculture. 2024 ITU Kaleidoscope: Innovation and digital transformation for a sustainable world (ITU K). IEEE (2024).
Sinamenye, J. H., Ayan, C. & Raju, S. Potato plant disease detection: Leveraging hybrid deep learning models. BMC Plant Biol. 25(1), 1–15 (2025).
Jha, P., Dembla, D. & Dubey, W. Deep learning models for enhancing potato leaf disease prediction: Implementation of transfer learning based stacking ensemble model. Multimedia Tools Appl. 83(13), 37839–37858 (2024).
Rameshkumar, R. et al. Enhanced plant disease classification via wild horse optimizer and convolutional attention-bidirectional long short term memory. SIViP 19(12), 1010 (2025).
Biswas, S. et al. Severity identification of potato late blight disease from crop images captured under uncontrolled environment. 2014 IEEE Canada international humanitarian technology conference-(IHTC). IEEE (2014).
Li, J. et al. Potato late blight leaf detection in complex environments. Sci. Rep. 14(1), 31046 (2024).
Dey, T. K., Jitesh P. & Danish A. K. Optimized potato leaf disease detection with an enhanced convolutional neural network. IETE J. Res., 1–14 (2025).
Chen, Y. & Liu, W. CBSNet: An effective method for potato leaf disease classification. Plants 14(5), 632 (2025).
Reis, H. C. & Turk, V. Potato leaf disease detection with a novel deep learning model based on depthwise separable convolution and transformer networks. Eng. Appl. Artif. Intell. 133, 108307 (2024).
Erlin, I. F. et al. Deep learning approaches for potato leaf disease detection: Evaluating the efficacy of convolutional neural network architectures. Revue d’Intell. Artif. 38(2), 717–727 (2024).
Tambe, U. Y. et al. Potato leaf disease classification using deep learning: A convolutional neural network approach. arXiv preprint arXiv:2311.02338 (2023).
Sundhar, S. et al. Enhancing leaf disease classification using GAT-GCN hybrid model. arXiv preprint arXiv:2504.04764 (2025).
Kussul, N., Lavreniuk, M., Skakun, S. & Shelestov, A. Deep learning classification of land cover and crop types using remote sensing data. IEEE Geosci. Remote Sens. Lett. 14, 778–782 (2017).
Rahnemoonfar, M. & Sheppard, C. Deep count: Fruit counting based on deep simulated learning. Sensors 17, 905 (2017).
Zhong, L., Hu, L. & Zhou, H. Deep learning based multi-temporal crop classification. Remote Sens. Environ. 221, 430–443 (2019).
Diaz, C. A. M., Castaneda, E. E. M., Vassallo, C. A. M. Deep learning for plant classification in precision agriculture. In: Proceedings of the 2019 International Conference on Computer, Control, Informatics and its Applications (IC3INA), Tangerang, Indonesia (2019).
Burhan, S. A., Minhas, S., Tariq, A. & Hassan, M. N. Comparative study of deep learning algorithms for disease and pest detection in rice crops. In: Proceedings of the 2020 12th International Conference on Electronics, Computers and Artificial Intelligence (ECAI), Bucharest, Romania (2020).
Narvekar, C. & Rao, M. Flower classification using CNN and transfer learning in CNN-agriculture perspective. In: Proceedings of the 2020 3rd International Conference on Intelligent Sustainable Systems (ICISS), Thoothukudi, India (2020).
Yang, S., Gu, L., Li, X., Jiang, T. & Ren, R. Crop classification method based on optimal feature selection and hybrid CNN-RF networks for multi-temporal remote sensing imagery. Remote Sens. 12, 3119 (2020).
Chugh, G., Sharma, A., Choudhary, P. & Khanna, R. Potato leaf disease detection using inception V3. Int. Res. J. Eng. Technol. 7(11), 1363–1366 (2020).
Rozaqi, A. J. & Sunyoto, A. Identification of disease in potato leaves using convolutional neural network (CNN) algorithm. In: Proceedings of the 2020 3rd International Conference on Information and Communications Technology (ICOIACT), Yogyakarta, Indonesia, pp. 72–76 (2020).
Ahmed, A. A. & Reddy, G. H. A mobile-based system for detecting plant leaf diseases using deep learning. AgriEngineering 3, 478–493 (2021).
de Camargo, T., Schirrmann, M., Landwehr, N., Dammer, K. H. & Pflanz, M. Optimized deep learning model as a basis for fast UAV mapping of weed species in winter wheat crops. Remote Sens. 13, 1704 (2021).
Kanmani, R., Muthulakshmi, S., Subitcha, K. S., Sriranjani, M., Radhapoorani, R. & Suagnya, N. Modern irrigation system using convolutional neural network. In: Proceedings of the 2021 7th International Conference on Advanced Computing and Communication Systems, ICACCS 2021, Coimbatore, India, pp. 592–597 (2021).
Vypirailenko, D., Kiseleva, E., Shadrin, D. & Pukalchik, M. Deep learning techniques for enhancement of weeds growth classification. In: Proceedings of the 2021 IEEE International Instrumentation and Measurement Technology Conference (I2MTC), Glasgow, UK (2021).
SenthilPandi, S. et al. Hybrid crossover oppositional firefly optimization for enhanced deep transfer learning in plant leaf disease classification. J. Crop Health 77(5), 1–16 (2025).
Acknowledgements
Not applicable.
Funding
This work was supported by the Deanship of Scientific Research, Vice Presidency for Graduate Studies and Scientific Research, King Faisal University, Saudi Arabia (Grant No. KFU254377).
Author information
Authors and Affiliations
Contributions
M.A., T.A., and S. A., collected data from different resources. T. A., S. I., M.N.A., and A.A.A., contributed to writing—original draft preparation; M.A., T.A., S.I., and J.M.H., contributed to writing—review and editing. S. I., A.A.A., and T.A. supervised the paper. M.A., J.M.H., and M.N.A. drafted pictures and tables. S.I., and T.A., performed revisions and improved the quality of the draft. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Asif, M., Abbas, T., Iqbal, S. et al. MapReduce-based deep learning framework for potato leaf disease detection in sustainable precision agriculture. Sci Rep 16, 1607 (2026). https://doi.org/10.1038/s41598-025-30940-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-30940-3