Abstract
Tau tubulin kinase 2 (TTBK2) is a ubiquitous serine-threonine protein kinase implicated in diverse cellular processes, including microtubule regulation, ciliogenesis, synaptic signaling, and the phosphorylation of key proteins like TDP-43. Despite its relevance, many aspects of TTBK2 function in both physiological and pathological conditions remain poorly understood. Truncating variants in TTBK2 gene cause spinocerebellar ataxia type 11 (SCA11), a rare form of autosomal dominant cerebellar ataxia. However, the functional consequences and pathogenic potential of missense variants have yet to be elucidated. In this study, we developed a CRISPR/Cas9 knock-in cell model harboring a missense variant in TTBK2 kinase domain (NM_173500.4:c.625 C > T; p.Leu209Phe) to evaluate its impact on TTBK2 expression, associated protein levels, and phosphoproteomic profiles. TTBK2 missense variant (TTBK2-L209F) was associated with reduced TTBK2 protein levels, altered levels of cytoskeleton-related proteins, and impaired kinase activity, namely toward TDP-43. Phosphoproteomic analyses identified dysregulation in pathways linked to gene regulation, protein degradation, cytoskeletal organization, and TGF-β signaling. These findings provide valuable insights into the biological roles of TTBK2 in cellular signaling. Moreover, this study underscores the importance of functional studies to better understand the consequences of TTBK2 missense variants, particularly those affecting the kinase domain, and their potential contribution to disease.
Similar content being viewed by others
Introduction
Tau-tubulin kinase 2 (TTBK2) is a serine-threonine protein kinase that belongs to the casein kinase superfamily. Unlike TTBK1, the other member of theTTBK family, which is neuron-specific, TTBK2 is ubiquitously expressed across multiple tissues . TTBK2 was initially identified due to its ability tophosphorylate the microtubule-associated protein tau and β-tubulin[ 2,3,,3] . Since then, TTBK2 has been implicated in various cellular functions, including: 1initiation of ciliogenesis, being a key player in the cilium assembly pathway4,5 ; (2) microtubule plus-end tracking protein (+TIP), regulating themicrotubule-depolymerizing activity of kinesin family member 2A (KIF2A) 6,7;3 regulation of neuronal membrane transporters and receptors, importantfor synaptic function and neuronal signaling ; 4 phosphorylation of the transactive response DNA-binding protein 43 kDa (TDP-43)11,12 whichaccumulates in cytoplasmic inclusions in amyotrophic lateral sclerosis (ALS) and frontotemporal lobar dementia (FTLD-TDP) ; and 5 maintenance ofconnectivity and viability of Purkinje cells . Nevertheless, only a few functions have been described in detail, and many aspects of TTBK2 biological rolesare not fully characterized.Pathogenic variants in TTBK2 gene have been linked to spinocerebellar ataxia type 11 (SCA11), an autosomal dominant disease with adult onset, mainly characterized by slowly progressive cerebellar ataxia. Patients can present with limb and gait imbalance, dysarthria, dysphagia, nystagmus, and hyperreflexia, among other symptoms14,15,,16. Life span is thought to be normal, but most patients require a wheelchair decades after disease onset15,17. The prevalence of SCA11 is unknown; however, it is extremely rare, likely accounting for less than 1% of autosomal dominant ataxias in Europe17.
All described cases of SCA11 have shown small deletions or insertions in TTBK2, resulting in frameshifts and truncated proteins15,16,17,18,19. Additional studies have reported TTBK2 missense variants in patients with cerebellar ataxia20,21,22,23,24,25,26,27 but, to our knowledge, none has been definitively linked to SCA1128. Furthermore, the precise disease mechanism underlying TTBK2 variants remains poorly understood and is primarily associated with truncating variants. Houlden et al. reported that mutated mRNA is partially degraded by nonsense-mediated decay (NMD)15, while other studies showed that truncated TTBK2 exhibits decreased kinase activity8,9,29 and may mislocalize from the cytosol to the nucleus29. Additionally, truncated TTBK2 appears to dominantly disrupt TTBK2 function, impairing cilia formation and signaling4,30,31.
In this study, we investigated the deleterious effects of a missense variant in the kinase domain of TTBK2 (c.625 C > T; p.Leu209Phe). Functional analyses using a CRISPR/Cas9 knock-in cell model demonstrated a loss of function mechanism for this variant, leading to impaired phosphorylation of both known and new potential TTBK2 targets. Ultimately, these results underscore the importance of investigating the effect of TTBK2 missense variants, particularly those in the kinase domain.
Materials and methods
Variants in Silico analysis
The deleterious effect of TTBK2 missense variants was accessed by in silico predictions using the Variant Effect Predictor (VEP) tool (Ensembl release 113; https://www.ensembl.org/Tools/VEP), which include the following: MutationTaster, Mutation Assessor, MutPred, FATHMM, FATHMM-MKL, FATHMM-XF, LRT, Deogen2, Eigen/Eigen PC, SIFT, Provean, MVP, Revel, Primate AI, MetaSVM/MetaLR, M-CAP, PolyPhen, LoFtool, Condel, BayesDel (addAF and noAF), ClinPred, LIST-S2, VEST, DANN and CADD. The pathogenic potential of the missense variant was further evaluated resorting to the minor allele frequency in gnomAD (v4.1.0; https://gnomad.broadinstitute.org/), 1000 Genomes Project (Phase 3; https://www.internationalgenome.org/), and dbSNP (build 157; https://www.ncbi.nlm.nih.gov/snp/) databases; and the nucleotide conservation given by PhyloP, PhastCons, Integrated fitCons and GERP++, all from the VEP tool (Ensembl release 113; https://www.ensembl.org/Tools/VEP).
The prediction of protein stability changes of TTBK2:c.625 C>T variant (p.Leu209Phe) was accessed through DynaMut32 (http://biosig.unimelb.edu.au/dynamut/) and DynaMut232 (http://biosig.unimelb.edu.au/dynamut2/) web servers, obtaining six different predictions of changes in folding free energy (ΔΔG): DynaMut2, DynaMut, ENCoM, mCSM, SDM, and DUET, recurring to the partial or complete 3D structure available in the Protein Data Bank (PDB) (6VRF; https://www.rcsb.org/). The DynaMut2 web server was also used to generate images of interatomic interactions.
Expression vectors and antibodies
The pEGFP-C1-TTBK2-WT plasmid (MRC PPU, Dundee, Scotland) was modified by site-directed mutagenesis using the QuikChange II kit (Agilent, Santa Clara, CA, USA) to introduce TTBK2:c.625 C > T variant. The following primers pairs were used to introduce the variant: forward primer
5′-GAAATGGGAAGACATGATGACTTTTGGTCCTTATTCTACATGT-3′
and reverse primer 5′-ACATGTAGAATAAGGACCAAAAGTCATCATGTCTTCCCATTTC-3′, respectively.
Antibodies used for Western blot and immunofluorescence studies are described in Table S1.
Cell culture and CRISPR/Cas9 gene editing
HEK293T (ATCC; kindly provided by Elsa Logarinho, IBMC/i3S, Porto) were grown in DMEM high glucose GlutaMAX™ supplemented with 10% FBS and 1% antibiotic-antimycotic (Gibco, ThermoFisher Scientific, Waltham, MA, USA) at 37 °C in a humidified 5% CO2 atmosphere. For overexpression studies, cells were transiently transfected for 24 h with each plasmid using Lipofectamine™ 2000 (Invitrogen, ThermoFisher Scientific, Waltham, MA, USA) according to the manufacturer’s protocol.
HEK293T cells were genetically modified using CRISPR/Cas9 system to insert a N-terminal 3xFLAG-tag in both alleles of TTBK2 (establishment of FLAG-TTBK2-WT cell line), according to methods already described34. A single-chain guide RNA (sgRNA) was selected using the CRISPOR online tool35 (v4.97; https://crispor.gi.ucsc.edu/); 5’-ACACATGCATCAGGTTTAGG-3’) and cloned in BbsI (ThermoFisher Scientific, Waltham, MA, USA) restriction sites into the mammalian expression construct pSpCas9(BB)−2 A-Puro (PX459) V2.0 (Addgene, Watertown, MA, USA), bearing both sgRNA scaffold backbone and S. pyogenes Cas9. A single-stranded oligodeoxynucleotide (ssODN) with 50 bp homology arms
(5´-GGGAACTGGATGCCTGTGTAGCTGTTCTACCATATCAGT
GTATTGCAATGGACTACAAAGACCATGA
CGGTGATTATAAAGATCATGACATCGATTACAAGGATGACGATGACA
AGGGTGGTGGTGGATCCAGTGGGGGAGGAGAGCAGCTGG
ATATCCTGAGTGTTGGAATCCTAGTGAA-3´)
was designed to serve as a template for homology-directed repair (HDR). A silent point mutation within the PAM sequence was introduced in the ssODN. The Cas9/sgRNA (2 µg) plasmid was co-transfected along with the ssODN (10 µM), in the presence of an inhibitor of non-homologous end joining (NHEJ) repair - NU702634 (10 µM; Sigma-Aldrich, Darmstadt, Germany), for 24 h using the DreamFect Gold reagent (OZ Biosciences, Marseille, France), according to the manufacturer’s protocol. Following transfection, clonal cell lines were isolated through fluorescence-activated cell sorting (FACS) and further genotyped by PCR (forward primer 5′-TATTAGTAGTGCTTCCTGATGCTG-3′ and reverse primer 5′- CTCTCTGAGATGTACACTCACCAC-3′), using Ranger Mix (Bioline, London, UK). PCR products with the expected molecular weight were purified with Exo/SAP (GRiSP, Porto, Portugal) and further analysed by Sanger sequencing using BigDye Terminator Cycle Sequencing v1.1 (Applied Biosystems, Foster City, CA, USA) in an ABI 3130xl Genetic Analyzer (Applied Biosystems, Foster City, CA, USA).
Then, homozygous knock-in of TTBK2:c.625C > T variant was performed on the FLAG-TTBK2-WT cell line described above (establishment of the FLAG-TTBK2-L209F cell line). Following the same methodology, one sgRNA (5’-ACATGTAGAATAAGGACCAA-3’) was cloned into the pSpCas9(BB)−2 A-Puro (PX459) V2.0. The ssODN with 50 bp homology arms
(5´-TGTGACTGGATTCTGTTTTGTTTATGAAGGAAATGGGAAGA
CATGATGACTTTTGGTCATTATTCTACATGTTGGTGGAGTTT
GTGGTTGGTCAGCTGCCCTGGAGAAA-3´) contained a silent point mutation to disrupt the AvaII restriction site, allowing genotyping of cell clones; the PAM sequence was disrupted when HDR on the variant site occurred. Transfection and establishment of clonal cells was performed as described above. Cells were genotyped through PCR (forward primer 5′-ACACATTAACTAATGGGCTGTT-3′ and reverse primer 5′-TCTTCTACCACAATTGACATCT-3′) followed by AvaII (New England Biolabs, Ipswich, MA, USA) digestion. Potential hits were sequenced by Sanger sequencing (as described above).
Off-targets of both sgRNAs were predicted using CRISPOR35 (v4.97; https://crispor.gi.ucsc.edu/) and CRISPR Finder37 (https://wge.stemcell.sanger.ac.uk) tools and checked for alterations using Sanger sequencing (as described above) in the selected clonal cell lines. Alteration in off-targets with four or fewer mismatches were excluded by sequencing (Table S2). Also, all TTBK2 exons and flanking intronic regions (Table S3) were checked for potential off-target effects.
Cell viability, proteasome, and autophagy assays
The CellTiter-Glo® Luminescent Cell Viability Assay (Promega, Madison, WI, USA) was used to determine the number of viable or metabolically active cells in the cell lines created, according to the manufacturer’s protocol. Luminescence was measured using the Synergy Mx Microplate Reader (Agilent, Santa Clara, CA, USA).
The Proteasome-Glo™ Chymotrypsin-Like, Trypsin-Like and Caspase-Like Cell-Based Assays (Promega, Madison, WI, USA) were used to measure the protease activities associated with the proteasome complex in the cell lines, according to the manufacturer’s protocol. Cells were treated with 5 µM MG132 (proteasome inhibitor) for 3 h as a control condition and to help define the activity attributable to the proteasome. The Synergy Mx Microplate Reader (Agilent, Santa Clara, CA, USA) was used to measure luminescence.
To monitor the autophagy flux, the cell lines were incubated with the autophagy inhibitor chloroquine (10 µM for 18 h; Sigma-Aldrich, Darmstadt, Germany) or control vehicle DMEM (Gibco, ThermoFisher Scientific, Waltham, MA, USA). Protein lysates were extracted from cells and the expression of autophagy protein markers was analyzed by Western blotting, as described below.
qPCR
Total RNA was isolated from HEK293T, FLAG-TTBK2-WT and -L209F cells, using the NZY Total RNA Isolation kit (Nzytech, Lisbon, Portugal), as per the manufacturer’s recommendations. RNA quantification was performed on the NanoDrop 2000 Spectrophotometer (ThermoFisher Scientific, Waltham, MA, USA). cDNA was synthesized by reverse transcription-PCR of 3 µg total RNA with oligo(dT), using the NZY First-Strand cDNA Synthesis Kit (Nzytech, Lisbon, Portugal), according to the manufacturer’s protocol. qPCR was performed using PowerUp™ SYBR™ Green Master Mix (Applied Biosystems, Foster City, CA, USA), along with respective primers (Table S4). Human ACTB was used as endogenous control. All reactions were performed in triplicate and replicated three times, using the Applied Biosystems 7500 Fast Real-Time PCR (Applied Biosystems, Foster City, CA, USA). The analysis was carried out using 7500 Software (version 2.0.6; Applied Biosystems, Foster City, CA, USA), and mRNA expression was calculated by the 2−ΔΔCT method.
Western blot analysis
HEK293T, FLAG-TTBK2-WT and -L209F cells were collected in RIPA buffer (150 mM NaCl, 1.0% IGEPAL® CA-630, 0.5% sodium deoxycholate, 0.1% SDS, 50 mM Tris, pH 8.0; Sigma-Aldrich, Darmstadt, Germany) supplemented with cOmplete Protease Inhibitor Cocktail (Roche, Basel, Switzerland) and then sonicated for 10 s intermittently with a power of 60 W (Branson sonifier 250, ThermoFisher Scientific, Waltham, MA, USA). Samples analyzed for protein phosphorylation were also supplemented with 1 mM Na3VO4 and 5 mM NaF. Total protein concentration was measured with the Pierce BCA protein assay kit (ThermoFisher Scientific, Waltham, MA, USA) according to the manufacturer’s instructions. Samples (30–50 µg of total protein) were separated on SDS-PAGE (Mini-PROTEAN; Bio-Rad, Hercules, CA, USA) and electrophoretically transferred onto PVDF membranes (Merck-Milipore, Darmstadt, Germany) using wet (Mini Trans-Blot electrophoretic transfer cell; Bio-Rad, Hercules, CA, USA) or semi-dry (SemiPhor transfer unit; Hoefer, Holliston, MA, USA) transfer systems. Membranes were firstly stained with Ponceau S and then blocked in 5% non-fat dry milk or bovine serum albumin (BSA) in Tris buffer saline/0.1% Tween-20 (TBS-T) for 1 h with agitation, at room temperature (RT), and subsequently incubated with specific primary antibodies as indicated (diluted in 5% BSA in TBS-T and 1% sodium azide), overnight at 4 °C with agitation. The membranes were washed with TBS-T and then incubated with HRP-conjugated secondary antibody (diluted in 3% non-fat dry milk or BSA in TBS-T) for 1 h with agitation at RT. Following washes with TBS-T, detection was achieved using WesternBright Sirius or WesternBright Standard ECL-HRP substrates (Advansta, San Jose, CA, USA), and chemiluminescence detected with a ChemiDoc XRS + Imaging System (Bio-Rad, Hercules, CA, USA). Ponceau S staining (used as loading control) and protein bands were quantified using the ImageLab software (version 6.1; Bio-Rad, Hercules, CA, USA). Quantitative comparisons between samples of each experiment were always performed on the same blot. When necessary, membranes were stripped by incubation in a stripping solution (62.5 mM Tris-HCl pH 6.7, 2% SDS, 100 mM β-mercaptoethanol) at 50 °C for 30 min, with gentle agitation.
Phosphoprotein enrichment and precipitation
Phosphoprotein enrichment was performed in FLAG-TTBK2-WT and -L209F cells (three independent experiments) using the phosphate metal affinity chromatography columns (TALON® PMAC Phosphoprotein Enrichment Kit; Clontech, Takara Bio, Mountain View, CA, USA), according to the manufacturer’s instructions. The second and third phospho-enriched fractions, which had the highest protein amount, were concentrated using a speed vacuum concentrator (SpeedVac, ThermoFisher Scientific, Waltham, MA, USA) and proteins further precipitated using the ProteoExtract Protein Precipitation Kit (Merck-Millipore, Darmstadt, Germany), according to the manufacturer’s instructions.
Mass spectrometry analysis
Sample preparation, protein identification, and quantitation, as well as data acquisition and processing, were performed according to the procedure described by Osório et al.38. The acquired raw data were processed using the Proteome Discoverer software (version 2.5.0.400; ThermoFisher Scientific, Waltham, MA, USA). Protein identification analysis was performed with the data available in the UniProt protein sequence database for the Reviewed Homo sapiens Proteome 2021_03 with 20,371 entries and a common contaminant database from MaxQuant (version 1.6.2.6, Max Planck Institute of Biochemistry). Additionally, phosphorylation of serine (Ser), threonine (Thr), and tyrosine (Tyr) residues was defined as variable modification and the ptmRS node was used for analysis and mapping of phosphorylation sites. Mass spectrometry data have been deposited to the ProteomeXchange Consortium via PRIDE39 repository with the identifier PXD056662.
Phosphoprotein network and functional enrichment analysis
Phosphoproteins were considered for the analysis of differentially expressed proteins (DEP) if detected in 2 out of 3 samples per experimental group with at least 50% of samples with protein-related peptides sequenced by MS/MS. Proteins identified with more than two unique peptides or with only one unique peptide but an MW lower than 30 kDa were included for analysis. For the selection of upregulated and downregulated proteins the L209F/WT ratio was set at ≥ 1.60 and ≤ 0.625, respectively. p-value was adjusted using Benjamini-Hochberg correction (FDR ≤ 0.05).
The protein-protein interaction (PPI) network from the significant DEP list (UniProt accession numbers) was achieved using the StringAPP plugin version 1.7.0 (minimum confidence 0.4) in the Cytoscape software40 (version 3.9.1; https://cytoscape.org/). Further analysis was performed using the NetworkAnalyzer plugin version 4.4.8. In the Cytoscape software, the nodes were labeled according to the relative expression [log2 (L209/WT ratio) values] from red/upregulated to blue/downregulated. The node size and edge width were matched to the node degree of distribution and STRING score retrieved to highlight the most relevant and confident interactions inside the network. The AutoAnnotate plugin version 1.3.5 allowed to cluster (GLay algorithm, clusterMaker2 v2.2) the protein-protein interactions and annotate them according to the word frequencies of Gene Ontology (GO) terms from the Biological Process category (previously attributed in the Proteome Discoverer software). The Over Representation Analysis (ORA) and Gene Set Enrichment Analysis (GSEA) methods were performed with the WebGestalt tool (2019 version; https://www.webgestalt.org/) according to default parameters. The weighted set cover redundancy reduction method was included as a post-processing step, allowing to identify the most representative and significant sets.
The phosphorylation sites identified in the DEP were analyzed using PhosphositePlus (v6.7.1.1; https://www.phosphosite.org/), Phospho.ELM (v.9.0; http://phospho.elm.eu.org/index.html), and PhosphoNet (2019 version; http://www.phosphonet.ca/) databases.
Statistical analysis
All quantitative data are expressed as mean ± standard deviation (SD) of at least three independent experiments. Statistical significance analysis was conducted using one-way analysis of variance (ANOVA) with Tukey’s post-hoc tests, when comparing means of more than two groups, or a two-tailed Student’s t-test to compare means between two groups. Non-parametric tests (Mann-Whitney U) were used when normality (Shapiro-Wilk test) and/or homogeneity of the variances (Levene test) were not observed. The level of statistical significance was set at a p-value ≤ 0.05. The statistical analysis was performed using the IBM SPSS Statistics software (version 25.0; IBM, Armonk, NY, EUA).
Results
In Silico analysis of TTBK2 missense variants
Truncating variants have been definitively linked to SCA11, whereas missense variants (Fig. 1a) have been identified in patients with cerebellar ataxia, but their pathogenicity has not been clearly established. Missense variants in the conserved kinase domain of TTBK2 have the potential to disrupt its activity and function. To further explore this, we conducted an in silico analysis of previously reported missense variants within TTBK2 kinase domain (Table S5). Among these, a novel variant (NM_173500.4:c.625 C > T; p.Leu209Phe) identified in a Portuguese family with progressive cerebellar ataxia41 (Supplementary material and methods), was notable for pathogenicity potential across multiple software tools and a higher CADD score. This variant has only three heterozygous allele counts and a total minor allele frequency of 1,859 × 10− 06 according to the gnomAD v4.1.0 database. It is reported in dbSNP as rs1595936604 and classified in ClinVar (VCV001978434.2) as a variant of unknown significance (one submission).
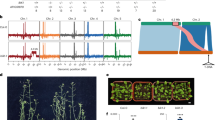
TTBK2 variants associated with cerebellar ataxia. (a) Schematic representation of the TTBK2 protein showing the localization of the missense variant of this study in red (L209F), and truncating and missense variants identified in other studies depicted in black. Sequence alignment of the flanking residues of Leu 209 of human TTBK2 kinase domain against other species is also shown (using Clustal Omega program). (b) Protein models of TTBK2 showing altered residue interactions upon amino acid change leucine to phenylalanine in position 209. On the left, the entire kinase domain is shown; on the right, a close-up view highlights the Leu209/Phe209 residues at the center, displaying their interactions with nearby residues in different colors. The models were performed in DynaMut2 (http://biosig.unimelb.edu.au/dynamut2/; PDB: 6VRF). A table displaying the protein stability prediction scores is also shown.
This TTBK2 variant, hereafter referred to as L209F, affects a highly conserved amino acid (Fig. 1a) located within an alpha helix of the kinase domain. It results in the substitution of leucine in position 209 by a phenylalanine, a residue buried within the core of the kinase domain. Protein stability analysis predicted a destabilizing effect of this missense variant (Fig. 1b), namely an increase in aromatic and hydrophobic contacts with the neighboring residues.
TTBK2-L209F affects protein expression
To study the pathogenic mechanisms triggered by L209F, we first created an endogenous HEK293T knock-in cell model expressing TTBK2 in fusion with an N-terminal 3xFLAG-tag (FLAG-TTBK2-WT cell line) through CRISPR/Cas9 editing. The introduction of the 3xFLAG allowed the detection of TTBK2 using specific FLAG-tag antibodies, overcoming the limitations of the commercial options. Next, we introduced the L209F variant into the FLAG-tagged cell line (FLAG-TTBK2-L209F cell line), using the same methodology (Fig. 2a). The newly established FLAG-TTBK2-L209F and the control FLAG-TTBK2-WT CRISPR/Cas9-modified cell lines (Fig. 2a) were then used for subsequent experiments.
The TTBK2-L209F variant caused a reduction in TTBK2, α-tubulin acetylation and KIF2A levels. (a) Electropherogram showing the variant TTBK2:c.625 C > T (boxed in red) introduced by CRISPR-Cas9 in the FLAG-TTBK2-L209F cell line. These cells also carry a single silent point mutation that disrupts AvaII restriction site, enabling initial screening of cell clones. Electropherogram of the control cell line (TTBK2-WT) is also shown for comparison. (b) Analysis of TTBK2 protein expression in HEK293T, FLAG-TTBK2-WT (WT) and FLAG-TTBK2-L209F (L209F) cells by immunoblotting with an anti-FLAG antibody. Ponceau S staining was used as loading control. Ψ identifies an unspecific band. Quantification data is presented in percentage as mean ± SD of three independent experiments; *P ≤ 0.05 compared with WT cell line (t-test). (c) Analysis of TTBK2 mRNA levels by qPCR. Primers targeting exon 4 or exon 8 of TTBK2 were used. Quantification data are presented in fold change as mean ± SD of three independent experiments; non-significant (ns) compared with HEK293T (non-modified cell line; one-way ANOVA/Tukey). (d) Analysis of acetylated α-tubulin, α-tubulin, HDAC6, and KIF2A expression levels in WT and L209F cells by immunoblotting with specific antibodies. Ponceau S staining was used as the loading control. Quantification data are presented in percentage as mean ± SD of at least three independent experiments; non-significant (ns), *P ≤ 0.05 and **P ≤ 0.01 (t-test).
Expression analysis revealed a significant 60% decrease (p = 0.023) in TTBK2 protein levels in the FLAG-TTBK2-L209F cell line compared to FLAG-TTBK2-WT cells (Fig. 2b). qPCR showed no significant reduction of TTBK2 mRNA levels in FLAG-TTBK2-L209F cells, indicating that the variant does not cause mRNA reduction or instability (Fig. 2c). Thus, TTBK2-L209F reduced levels may result from reduced protein stability, as supported by in silico predictions (Fig. 1b). In addition, we showed that introducing the 3xFLAG-tag did not affect TTBK2 mRNA expression, as shown by comparable mRNA levels between HEK293T and FLAG-TTBK2-WT cells (Fig. 2c).
To understand if L209F affects cell viability and/or induces apoptosis, we performed a cell viability assay and analyzed caspase-3 expression and cleavage. We found no significant changes in both assays between the cell lines (Fig. S1). Also, no cleaved fragments of caspase-3 were detected, indicating no measurable activation of apoptosis (Fig. S1b).
Redistribution of TTBK2 truncating proteins to the nucleus has been previously reported29. To address this, we performed fluorescence microscopy and subcellular fractionation to determine TTBK2-L209F subcellular localization. In both cell lines, TTBK2 seemed to be distributed throughout the cytoplasm, without noticeable changes in the localization of TTBK2-L209F compared to TTBK2-WT (Fig. S2; Supplementary material and methods).
TTBK2-L209F influences microtubule stability
TTBK2 was reported to participate in microtubule regulation by associating with tau, tubulin and KIF2A2,3,7, and by acting as a + TIP6,7. Therefore, we started by analyzing the expression of acetylated α-tubulin, an indicator of microtubule stability. Immunoblotting analysis showed that acetylated α-tubulin levels were significantly reduced by approximately 50% (p = 0.031) in FLAG-TTBK2-L209F cells compared to WT cells, which was not a consequence of alterations in total α-tubulin levels (Fig. 2d). Reversible deacetylation of α-tubulin is mediated by histone deacetylases, namely HDAC634. Thus, we measured HDAC6 levels and found that FLAG-TTBK2-L209F cells display nearly twice as much HDAC6 compared to WT cells (p = 0.004; Fig. 2d), in agreement with reduced α-tubulin acetylation. Furthermore, KIF2A was expressed at lower levels in FLAG-TTBK2-L209F cells (around 50% less than WT cells; p = 0.035; Fig. 2d), even though KIF2A mRNA levels remained unchanged (Fig. S3), confirming that KIF2A transcription is not affected but rather protein expression or stability. These results indicate dysregulation of TTBK2 cytoskeletal targets, likely resulting in microtubule instability in FLAG-TBK2-L209F cells.
TTBK2-L209F impairs kinase activity against TDP-43
TDP-43 is a well-known target of TTBK2 protein kinase11. To investigate the effect of the L209F variant on TTBK2 kinase activity against TDP-43, we analyzed the phosphorylation state of TDP-43 at Ser409/410 in our cell lines. In FLAG-TTBK2-L209F cells, the levels of phosphorylated TDP-43 (pTDP-43; bands above 74 kDa) were significantly reduced by 36% (p = 0.041; Fig. 3a), while total TDP-43 levels remained unaffected. Transfection of cells with EGFP-TTBK2-WT confirmed that TTBK2 can augment pTDP-43 levels (Fig. 3a), in agreement with previous results. Moreover, we showed that pTDP-43 levels increased with overexpression of EGFP-TTBK2-WT in both cell lines (around 70% and 170%, p = 0.025 and 0.007, respectively; Fig. 3b) but not following transfection with EGFP-TTBK2-L209F. Similar to the effects observed with EGFP-TTBK2-L209F overexpression, a kinase-dead and a SCA11 (R444fs*5) mutant did not alter pTDP-43 levels in HEK293 cells. In contrast, the F287S mutant (variant outside the kinase domain) led to an increase in pTDP-43 levels (Fig. S4). These results support that TTBK2-L209F has impaired kinase activity toward TDP-43.
TTBK2-L209F variant impaired TDP-43 phosphorylation. (a) Analysis of TDP-43 Ser409/410 protein levels (pTDP-43) in FLAG-TTBK2-WT (WT) and FLAG-TTBK2-L209F (L209F) cells. Total TDP-43 was used to normalize pTDP-43 levels, while Ponceau S was used to normalize total TDP-43 levels. (b) WT and L209F cells were transfected with the empty vector EGFP, EGFP-TTBK2-WT or EGFP-TTBK2-L209F, followed by immunoblotting with anti-TDP-43 Ser409/410, anti-TDP-43 and anti-EGFP antibodies. Total TDP-43 levels were used to normalize pTDP-43 levels. Quantification data are presented in percentage as mean ± SD of three independent experiments; non-significant (ns), *P ≤ 0.05 and **P ≤ 0.01 compared with the WT cell line (Mann-Whitney U test in (a) and one-way ANOVA/Tukey in (b)).
TTBK2-L209F is linked to abnormal protein phosphorylation
Phosphoprotein enrichment followed by MS/MS analysis was performed to identify and quantify phosphorylated proteins (Fig. S5a), in the context of TTBK2 impaired kinase activity. After filtering the data, we identified 4429 proteins amongst the phospho-enriched fractions of six independent samples: three of FLAG-TTBK2-WT cells (control condition) and three of FLAG-TTBK2-L209F cells (impaired kinase activity). Using a threshold of L209/WT ratio ≥ 1.60 for upregulated proteins and ≤ 0.625 for downregulated proteins, a total of 110 differentially expressed phosphoproteins (DEPs) were identified: 50 upregulated and 60 downregulated in FLAG-TTBK2-L209F cells (Table 1; Fig.S5b).In addition, eighteen phosphorylation sites across ten DEP (Table 2) were also identified. These include AMOT, HTATSF1, RBM20, SQSTM1 (from nowon referred as p62), SMAD2, CTIF, NEDD4L, TNIK, IRS4, and BAG3. Using publicly available databases, we found that half of the identifiedphosphorylated residues had also been found in other mammalian studies. Additionally, the kinases responsible for the phosphorylation of these residueshave previously been described. For example, p62 [Ser272] is phosphorylated by cyclin-dependent kinase 1 (CDK1) and mitogen-activated protein kinase(MAPK) 13; SMAD2 [Thr8] by MAPK3; and NEDD4L [Ser448] by protein kinase cAMP-activated catalytic subunit alpha (PKACA) andserum/glucocorticoid regulated kinase 1 (SGK1) (Table 2).
Phosphoproteins interactions analysis
To explore potential interactions and functional consequences of DEPs, a PPI network was generated (Fig. 4) along with associated Gene Ontology (GO) Biological Process terms. In this network, eight major clusters were identified (Fig. 4), with central nodes corresponding to either upregulated phosphoproteins – SMAD2, p62, AMOT, and NARS1 or downregulated phosphoproteins – EEF2, ILK, MAP1B, and KIF18A, in FLAG-TTBK2-L209F cells. Some phosphoproteins were found exclusively in FLAG-TTBK2-L209F cells (“exclusive”; ANXA1, FOXM1 and ULK3), while others were identified only in FLAG-TTBK2-WT cells (“lost”; PTBP3) (Fig. 4; triangular and rectangular shaped nodes, respectively).
Protein interactions and biological processes shared by phosphoproteins significantly dysregulated in FLAG-TTBK2-L209F cells. Nodes gradient colors represent the relative expression according to log2 (L209/WT ratio) values. Upregulated phosphoproteins are in reddish colors and the ones recovered only in FLAG-TTBK2-L209F cells (“exclusive”) have triangularly shaped nodes. Downregulated phosphoproteins are in bluish colors and phosphoproteins not recovered in FLAG-TTBK2-L209F cells (“lost”) are shown as rectangular shaped nodes. The node size and edge width are according to the node degree of distribution and STRING score retrieved in the network analysis. Phosphoproteins with identified phosphorylation sites in our data are highlighted with a yellow margin. The protein-protein interaction map was performed using stringAPP from Cytoscape 3.9.1. Network clustering (GLay) and annotations based on GO: Biological Process terms were performed with the AutoAnnotate plugin version 1.3.5.
The largest cluster, linked to cell signaling and metabolism, centered on the upregulated phosphoproteins SMAD2, SMAD4 and NEDD4L, all involved in the TGF-β signaling pathway43. Conversely, downregulated factors at the periphery of this cluster, namely SATB1, ZBTB7A, and PTBP3, may regulate transcription through the TGF-β signaling pathway44,45,46. The p62 cluster highlighted the stress response process, presenting several autophagy-related proteins, with BAG3 and ATG7 downregulated, while ULK3 and p62 itself were upregulated. The MAP1B cluster, related to cell organization and biogenesis, included NOP53 and WDR74, both involved in ribosomal biogenesis and RNA metabolism47. The ILK cluster associated with cell adhesion comprised proteins of the PINCH-ILK-Parvin (PIP) complex, including ILK, LIMS1, PARVA, and PARVB (all downregulated), which provide critical links between integrins and the actin cytoskeleton48. In the EEF2 cluster associated with protein processes, most phosphoproteins were downregulated and involved in translation. The AMOT cluster was engaged in cell organization and RNA transcription and contained mostly upregulated phosphoproteins. In addition, downregulated TRIP10 and PACSIN3 play a role in actin cytoskeleton reorganization during vesicle formation and endocytosis49,50. The NARS1 cluster was associated with RNA and protein metabolism, while the small KIF18A cluster was associated with cell cycle and cell proliferation (Fig. 4).
Functional enrichment analysis
We used the web tool WebGestalt to conduct over-representation analysis (ORA) in our set of phosphoproteins (Table S6). We found seven statistically significant upregulated pathways in FLAG-TTBK2-L209F cells: embryonic development, negative transcriptional regulation, negative regulation of organelle organization, and cellular responses to stress. Interestingly, SMAD4 and SMAD2 were present in more than half of these enriched sets, while SLIT2, PIK3R3, and histone proteins (H3-3A/B and H3C13/14/15) were identified in more than one enriched set. Furthermore, we identified one downregulated pathway “Cell-extracellular matrix interactions” that includes the ILK and parvin proteins (Table S6).
Additionally, gene set enrichment analysis (GSEA) was performed to obtain more insights into the biological alterations caused directly or indirectly by L209F (Table S7). In FLAG-TTBK2-L209F cells, among the enriched pathways [with positive normalized enriched score (NES)], we highlight the sets related to cytoskeleton regulation, such as “Ciliary part”, “Regulation of Actin Cytoskeleton” and “Regulation of Cytoskeleton Organization”. In addition, several enriched pathways included several members of the histone clusters (e.g., WP2369). Furthermore, the FLAG-TTBK2-L209F cells were found to lack phosphoproteins involved in translation and transcription pathways. Among these negatively enriched pathways, there were several ribosomal and mitochondrial ribosomal proteins, as well as several members of the eukaryotic initiation factor 3 (eIF3) multiprotein complex. In this data, we also observed negative NES in sets related to gene expression and mitochondrial activity. Interestingly, a decrease in Parkinson Disease (PD)-associated phosphoproteins, such as PARK7 and tau, was observed (Table S7).
Potential TTBK2 targets in transcription and autophagy
Among the proteins potentially affected by the L209F variant in the MS/MS data, many were associated with transcription and autophagy/protein degradation pathways. This prompted us to investigate several proteins involved in these biological processes in more detail. Unfortunately, due to the lack of available antibodies, we were unable to consistently confirm the phosphorylation state or specific phosphorylation sites of all targets.
We started by analyzing the transcription factor SMAD2 (upregulated phosphoprotein) and the state of SMAD2 phosphorylation at Ser465/467, which is crucial for SMAD2-mediated TGF-β signaling51. We confirmed that the levels of SMAD2 phosphorylated at Ser465/467 were significantly increased by 43% (p = 0.015), accompanied by a simultaneous increase in SMAD2 protein expression (89%, p = 0.003) in FLAG-TTBK2-L209F cells (Fig. 5). But there were no significant differences between cell lines when the phosphorylated levels were normalized to total protein levels. Nevertheless, SMAD2 high levels suggest transcriptional dysregulation in FLAG-TTBK2-L209F cells.
SMAD2 protein expression was elevated in FLAG-TTBK2-L209F cells. Analysis of SMAD2 Ser465/467 and total SMAD2 protein levels in FLAG-TTBK2-WT (WT) and FLAG-TTBK2-L209F (L209F) by immunoblotting using specific antibodies. SMAD2 Ser465/467 and total SMAD2 levels were normalized to Ponceau S staining (loading control). SMAD2 Ser465/467 phosphorylation levels normalized to total SMAD2 levels are also shown. Quantification data are presented in percentage as mean ± SD of three independent experiments; non-significant (ns), *P ≤ 0.05 and **P ≤ 0.01 compared with the WT cell line (t-test).
Another phosphoprotein – p62 – was also upregulated in FLAG-TTBK2-L209F cells (about a threefold increase compared to non-treated WT cells, p ≤ 0.001); Fig. 6a). Since p62 accumulation has been linked to impaired protein degradation by the autophagic system52, we analyzed the levels of other autophagy-related proteins and phosphorylated forms that are critical for autophagy induction. Phosphorylated levels of raptor (Ser792), ULK1 (Ser555), and beclin-1 (Ser93) were significantly increased (approximately 60, 100 and 40%; p = 0.042, 0.01 and 0.041, respectively) in FLAG-TTBK2-L209F cells, accompanied by an increase in the total expression levels of the respective proteins (Fig. 6b), although only ULK1 increase reached statistical significance (increase of 92%, p = 0.016). However, no significant differences were observed between cell lines when the phosphorylated levels were normalized to total protein levels.
The expression of several autophagy-related proteins is increased in FLAG-TTBK2-L209F cells but the autophagic flux seems unaffected. (a) Protein expression analysis of p62, LC3B-I, LC3B-II and LC3B-II/LC3B-I ratio in non-treated FLAG-TTBK2-WT (WT) and FLAG-TTBK2-L209F (L209F) cells (incubated with the control vehicle DMEM) and in cells incubated with 10 µM chloroquine (CQ) for autophagy blockage. Immunoblotting was performed using specific antibodies and Ponceau S staining was used as loading control. (b) Protein expression analysis of the upstream autophagy-related phosphoproteins raptor, ULK1 and beclin-1 in WT and L209F cells by immunoblotting. Raptor Ser792, ULK1 Ser555 and Beclin-1 Ser93 phosphorylated levels, as well as total raptor, ULK1 and beclin-1 levels were normalized to Ponceau S staining (loading control). Phosphorylated levels of raptor, ULK1 and beclin-1 were also normalized to the respective total protein levels. Quantification data are presented in percentage as mean ± SD of at least three independent experiments; non-significant (ns), *P ≤ 0.05, **P ≤ 0.01 and ***P ≤ 0.001 compared with the WT cell line and with non-treated cells in panel (a) (one-way ANOVA/Tukey); non-significant (ns), *P ≤ 0.05 and **P ≤ 0.01 compared with the WT cell line in panel (b) (t-test).
To investigate further if TTBK2-L209F cells had an altered autophagic flux, we treated cells with chloroquine, an inhibitor of autophagic flux by decreasing autophagosome-lysosome fusion52. We did not find statistically significant differences in either LC3B-I, LC3B-II or LC3B-II/LC3B-I ratio between non-treated FLAG-TTBK2-WT and L209F cells (Fig. 6a). Nevertheless, the levels of LC3B-II increased approximately five and eight times in FLAG-TTBK2-WT and L209F cells, respectively, following drug treatment as expected (p = 0.029 and p = 0.006; Fig. 6a). In accordance, the LC3B-II/LC3B-I ratio increased approximately three times in both cell lines treated with chloroquine (p = 0.006 and 0.019; Fig. 6a). Similarly, p62 levels increased approximately four and two times after drug treatment in FLAG-TTBK2-WT and -L209F cells, respectively (p ≤ 0.001; Fig. 6a). Together these results suggested that autophagic flux was not impaired in the presence of the L209F variant.
Furthermore, we measured the levels of the mRNAs encoding these proteins to ascertain whether elevated protein levels were caused by transcriptional activation rather than dysfunctional protein degradation. The mRNA levels of SMAD2, p62, LC3B and beclin-1 were significantly increased (3.4, 0.5, 0.8 and 1.7 fold change; p = 0.012, 0.006, 0.05 and 0.021 respectively) while raptor and ULK1 mRNAs levels tended to decrease in FLAG-TTBK2-L209F cells (Fig. S6), suggesting that expression changes may be, at least in part, due to transcription regulation for most of these proteins.
Additionally, since proteasome inhibition can induce p62 synthesis52, we assessed the protease activities associated with the proteasome complex. Chymotrypsin, trypsin, and caspase-like protease activities were slightly increased in FLAG-TTBK2-L209F cells compared to WT cells, but the differences were not statistically significant (Fig. S7), indicating normal proteasome function.
Discussion
In this study, we demonstrated for the first time the detrimental impact of a missense variant in TTBK2 (c.625 C > T, p.Leu209Phe; Fig. 1). While all confirmed SCA11 cases to date have been linked to truncating variants in TTBK2, our findings highlight the potential for missense variants to disrupt TTBK2 function, particularly those within the kinase domain. The TTBK2-L209F variant described here is extremely rare, with only three heterozygous alleles identified in the European non-Finnish population. Thereby, incomplete disease penetrance or underdiagnosis until later stages of disease progression cannot be ruled out.
To ascertain the potential pathogenicity of TTBK2-L209F, we conducted functional studies using a HEK293T cell model genetically modified by CRISPR/Cas9 to express the TTBK2 missense variant in homozygosity (Fig. 2a). While this model does not fully replicate the disease context, it provided a reliable system to investigate the variant’s impact on TTBK2 expression and molecular functions.
Our results provided evidence for a loss of function mechanism underlying the pathogenesis of TTBK2-L209F, as shown by reduced TTBK2 expression and kinase activity. The variant caused reduced TTBK2 protein levels while maintaining normal mRNA expression, likely due to decreased protein stability (Fig. 2b and c). In contrast, previous studies of TTBK2 truncating variants reported a partial reduction in mRNA levels, suggestive of premature degradation by NMD15. Consistent with prior findings8,9,29, our data showed impaired kinase activity of mutated TTBK2. Specifically, analysis of the phosphorylation state of TDP-43 at Ser409/410, a known target of TTBK211,12, showed that TTBK2-L209F has a reduced capacity to phosphorylate TDP-43 (Fig. 3). Furthermore, these experiments did not support a dominant negative effect, since TTBK2-L209F was not able to interfere with the ability of TTBK2-WT to phosphorylate TDP-43 (Fig. 3b). Although, this mechanism cannot be entirely ruled out, as the levels of both endogenous and overexpressed TTBK2-L209F proteins were lower in comparison with WT proteins, potentially skewing the results.
Cytoskeleton abnormalities were identified in the FLAG-TTBK2-L209F cell line (Fig. 2d), specifically reduced tubulin acetylation and KIF2A levels. Future studies are required to determine if these effects were a direct result of impaired TTBK2 kinase activity against its cytoskeletal targets or were indirectly influenced by other mechanisms. Nevertheless, we hypothesize that reduced TTBK2 expression or kinase activity could lead to microtubule dynamic instability, resulting in reduced tubulin acetylation, as TTBK2 functions as a + TIP6,7and phosphorylates tubulin and other microtubule-associated proteins (MAPs)2,3. Another plausible scenario is that TTBK2 could indirectly influence tubulin acetylation by phosphorylating TDP-43. TDP-43 is known to influence neurite outgrowth by regulating tubulin deacetylase HDAC6 levels53, which were elevated in FLAG-TTBK2-L209F cells (Fig. 2). Deficits in tubulin acetylation can alter the MAP landscape, leading to impairment of axonal transport, abnormal polarization and migration in neurons, all hallmarks of many neurodegenerative diseases54. For example, increasing microtubule acetylation has been shown to rescue axonal transport and locomotor deficits in Huntington’s disease (HD) and PD, respectively, and to revert axonal loss in Charcot-Marie-Tooth disease54. Regarding KIF2A, it is known that phosphorylation at Ser135 by TTBK2 inhibits KIF2A interaction with microtubules and decreases its microtubule depolymerizing activity7. It remains to be determined whether reduced KIF2A levels correlate with decreased phosphorylation of KIF2A at Ser135, as well as whether these changes are regulated by TTBK2 or are a consequence of cytoskeleton instability. Interestingly, pathogenic variants in KIF2A have been linked to cortical dysplasia55, and KIF2A deficiency has disrupted neurogenesis and axonal transport, leading to neurodegeneration in mice56. Moreover, pathogenic variants that disrupt the functions of other kinesins (e.g., KIF1C, KIF7, KIF26B) have been also associated with cerebellar ataxia57.
Our analysis of protein phosphorylation in FLAG-TTBK2-L209F cells (Table 1) revealed potential alterations in the phosphorylation of other microtubule-associated proteins (e.g., KIF18A and MAP1B, both downregulated). Interestingly, several actin-binding proteins (e.g., PARVA/B/parvin alpha and beta, PPP1R9A/neurabin 1, and PFN2/profilin 2) were also identified in this dataset, as well as intermediate filament-associated proteins (e.g., EPPK1/epiplakin 1 and INA/alpha-internexin), suggesting that other cytoskeletal components can also be disrupted in FLAG-TTBK2-L209F cells.
Phosphoproteome data also drew our attention towards SMAD2, which was present in several enriched sets in the ORA analysis (Table S6) and emerged as a central node in the PPI network (Fig. 4). Notably, SMAD2 was highly phosphorylated at Thr8 in FLAG-TTBK2-L209F cells (Table 2). This residue is phosphorylated by MAPK3, leading to increased SMAD2 protein levels and complex formation with SMAD4, thereby enhancing transcriptional activity58. SMAD4 was also identified as an upregulated phosphoprotein in our MS/MS data, along with NEDD4L (Table 1), which targets SMAD proteins for degradation, modulating the TGF-β pathway43. NEDD4L was phosphorylated at Ser488 in our data (Table 2). Dephosphorylation of this residue has been reported to inhibit the activity of NEDD4L E3 ubiquitin ligase by reducing NEDD4L protein levels and regulating neuronal excitability59. Therefore, we hypothesize that increased phosphorylation at Ser488 may correlate with elevated NEDD4L protein levels. Additionally, we analyzed the phosphorylation state of SMAD2 at Ser465/467, which was elevated in FLAG-TTBK2-L209F cells, along with total SMAD2 levels (Fig. 5). Phosphorylation of Ser465/467 enables SMAD2 association with SMAD4 and facilitates interaction with other co-factors to regulate gene expression and mediate TGF-β signaling43,51. Therefore, TGF-β signaling may be affected in FLAG-TTBK2-L209F cells. Curiously, downregulation of several genes involved in TGF-β signaling, including daf-14 (the orthologue of SMAD2), partially rescued worm incoordination with an unc-2 (orthologue of CACNA1A; genetic cause of SCA6 and episodic ataxia type 2) truncating variant60.
Additional upregulated phosphoproteins involved in transcription and gene expression were identified in our MS/MS data, including various histone proteins, PIAS2/protein inhibitor of activated STAT 2, and histone-lysine N-methyltransferase MECOM (Fig. 4). The precise mechanisms by which reduced TTBK2 expression and/or kinase activity led to the upregulation of phosphoproteins associated with transcription and gene expression remain to be fully elucidated. Intriguingly, several phosphoproteins involved in translation and transcription pathways (Table S6) were downregulated in FLAG-TTBK2-L209F cells, including ribosomal proteins, members of the eIF3 multiprotein complex, and EEF2/elongation factor 2. A pathogenic variant in EEF2 was found to cause SCA26, by impairing translation and increasing susceptibility to proteostatic disruption61. In addition to EEF2, two other downregulated phosphoproteins associated with hereditary cerebellar ataxias were identified in FLAG-TTBK2-L209F cells (Table 1): SLC1A3/excitatory amino acid transporter 1, linked with episodic ataxia type 6 through decreased glutamate uptake62; and ATG7, a ubiquitin-activating enzyme E1, implicated in a recessive form of cerebellar ataxia and impaired autophagic flux63.
Beyond ATG7, other phosphoproteins involved in protein degradation pathways were significantly dysregulated in FLAG-TTBK2-L209F cells. Among them, p62 may be particularly relevant, since autophagy and proteasome degradation are often impaired in SCAs64 and dysregulation of p62 levels has been observed in several neurodegenerative diseases65. Phosphorylation of p62 at Ser272 was elevated in FLAG-TTBK2-L209F cells (Table 2). This residue was previously reported to be phosphorylated upon nocodazole treatment during mitosis by CDK134. Also, inflammation-related protein mitogen-activated protein kinase kinase kinase 7 (MAP3K7) phosphorylates p62 at different residues, potentially including Ser272, inhibiting the autophagic degradation of p62 and promoting p62-mediated cellular signaling67. Total p62 protein levels were increased in FLAG-TTBK2-L209F cells (Fig. 6a), suggesting autophagy impairment52. However, protein levels of several autophagy-associated proteins were also increased, along with its phosphorylated forms raptor (Ser792), beclin-1 (Ser93) and ULK1 (Ser555) (Fig. 6) related to autophagy induction52,68,69. Treatment with chloroquine, an autophagic inhibitor that blocks autophagosome-lysosome fusion52, resulted in increased LC3B-II and p62 levels, along with an elevated LC3B-II/LC3-I ratio in both cell lines (Fig. 6a), indicating a normal LC3B-II turnover, autophagosome formation, and autophagic flux52. Furthermore, mRNA levels of p62 and beclin-1 were also upregulated in FLAG-TTBK2-L209F cells (Fig. S6), indicating that expression changes may be due to transcriptional activation. However, this was not the case for all autophagy proteins, as mRNA levels of ULK1 and raptor tended to decrease in FLAG-TTBK2-L209F cells. It is known that increased protein levels, at least for ULK1, do not always correlate with its mRNA levels under certain conditions70. While this study did not confirm whether TGF-β signaling drives these expression changes, previous research has shown that TGF-β can upregulate the mRNA levels of some autophagy-related proteins and activate autophagy in specific cancer cells71.
Although our study offers valuable insights, some limitations must be acknowledged. First, our study relies on a single WT and a single clonal mutant cell line, so we cannot fully exclude the possibility that some of the observed differences may reflect clonal variability. Future work will be needed to strengthen these findings, for example by analyzing additional independent clones, performing rescue experiments, or even using complementary model systems. Secondly, our approach to assess TTBK2 kinase activity may provide only an indirect assessment. Thus, a kinase assay directly measuring TTBK2 activity would complement our results on TDP-43 phosphorylation. Finally, our data did not definitively establish the role of deregulated phosphoproteins but highlight several critical aspects that should be explored in future research to better understand TTBK2 biological roles in both health and disease. Additionally, it would be valuable to explore in detail the TTBK2 interactome, which remains superficially characterized, and compare those results with our phosphoproteomics data.
This study provided the first evidence linking a TTBK2 missense variant (TTBK2-L209F) to a loss of function mechanism. This variant led to impaired protein phosphorylation, potentially disrupting key cellular processes and molecular pathways, including cytoskeleton dynamics, gene regulation and TGF-β signaling. Future studies investigating these processes will be crucial for unraveling the pathogenic mechanisms of TTBK2 dysfunction and its role in the development of cerebellar ataxia.
Data availability
The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD056662. Sanger sequencing data have been deposited in NCBI’s Sequence Read Archive (SRA) database with the accession number PRJNA1375520. Additional data supporting the findings of this study are available from the corresponding author upon reasonable request.
References
Ikezu, S. & Ikezu, T. Tau-tubulin kinase. Front Mol. Neurosci 7, 33 (2014).
Takahashi, M. et al. Involvement of T protein kinase I in paired helical Filament-Like phosphorylation of the juvenile T in rat brain. J. Neurochem. 64, 1759–1768 (1995).
Tomizawa, K., Omori, A., Ohtake, A., Sato, K. & Takahashi, M. Tau-tubulin kinase phosphorylates Tau at Ser-208 and Ser-210, sites found in paired helical filament-tau. FEBS Lett. 492, 221–227 (2001).
Goetz, S. C., Liem, K. F. & Anderson, K. V. The spinocerebellar ataxia-associated gene Tau tubulin kinase 2 controls the initiation of ciliogenesis. Cell 151, 847–858 (2012).
Bowie, E. & Goetz, S. C. Ttbk2 and primary cilia are essential for the connectivity and survival of cerebellar purkinje neurons. Elife 9, e51166 (2020).
Jiang, K. et al. A Proteome-wide screen for mammalian sxip Motif-Containing microtubule Plus-End tracking proteins. Curr. Biol. 22, 1800–1807 (2012).
Watanabe, T. et al. TTBK2 with EB1/3 regulates microtubule dynamics in migrating cells through KIF2A phosphorylation. J. Cell. Biol. 210, 737–751 (2015).
Almilaji, A., Munoz, C., Hosseinzadeh, Z. & Lang, F. Upregulation of Na,Cl - -coupled betaine/γ-amino-butyric acid transporter BGT1 by Tau tubulin kinase 2. Cell. Physiol. Biochem. 32, 334–343 (2013).
Nieding, K. et al. Tau tubulin kinase TTBK2 sensitivity of glutamate receptor GluK2. Cell. Physiol. Biochem. 39, 1444–1452 (2016).
Zhang, N. et al. Phosphorylation of synaptic vesicle protein 2A at Thr84 by casein kinase 1 family kinases controls the specific retrieval of Synaptotagmin-1. J. Neurosci. 35, 2492–2507 (2015).
Liachko, N. F. et al. The Tau tubulin kinases TTBK1/2 promote accumulation of pathological TDP-43. PLoS Genet 10, e1004803 (2014).
Taylor, L. M. et al. Pathological phosphorylation of Tau and TDP-43 by TTBK1 and TTBK2 drives neurodegeneration. Mol. Neurodegener. 13, 7 (2018).
Prasad, A., Bharathi, V., Sivalingam, V., Girdhar, A. & Patel, B. K. Molecular mechanisms of TDP-43 misfolding and pathology in amyotrophic lateral sclerosis. Front. Mol. Neurosci. 12, 45 (2019).
Worth, P. F. et al. Autosomal dominant cerebellar ataxia type III: linkage in a large British family to a 7.6-cM region on chromosome 15.q14-21.3. Am. J. Hum. Genet. 65, 420–426 (1999).
Houlden, H. et al. Mutations in TTBK2, encoding a kinase implicated in Tau phosphorylation, segregate with spinocerebellar ataxia type 11. Nat. Genet. 39, 1434–1436 (2007).
Lindquist, S. G. et al. A novel TTBK2 de Novo mutation in a Danish family with Early-Onset spinocerebellar ataxia. Cerebellum 16, 268–271 (2017).
Bauer, P. et al. Spinocerebellar ataxia type 11 (SCA11) is an uncommon cause of dominant ataxia among French and German kindreds. J. Neurol. Neurosurg. Psychiatry. 81, 1229–1232 (2010).
Németh, A. H. et al. Next generation sequencing for molecular diagnosis of neurological disorders using ataxias as a model. Brain 136, 3106–3118 (2013).
Coutelier, M. et al. Efficacy of exome-targeted capture sequencing to detect mutations in known cerebellar ataxia genes. JAMA Neurol. 75, 591–599 (2018).
Deng, Y., Fu, J., Zhong, Y. Q., Zhang, M. & Qi, X. First finding of Familial spinal cerebellar Ataxia11 in china: clinical, imaging and genetic features. Neurol. Sci. 41, 155–160 (2020).
Galatolo, D. et al. NGS in hereditary ataxia: when rare becomes frequent. Int. J. Mol. Sci. 22, 8490 (2021).
Edener, U. et al. Missense exchanges in the TTBK2 gene mutated in SCA11. J. Neurol. 256, 1856–1859 (2009).
Jiao, Q. et al. The combination of whole-exome sequencing and copy number variation sequencing enables the diagnosis of rare neurological disorders. Clin. Genet. 96, 140–150 (2019).
Iqbal, Z. et al. Targeted high throughput sequencing in hereditary ataxia and spastic paraplegia. PLoS One. 12, e0174667 (2017).
Fakhro, K. A. et al. Point of care exome sequencing reveals allelic and phenotypic heterogeneity underlying Mendelian disease in Qatar. Hum. Mol. Genet. 28, 3970–3981 (2019).
Coutelier, M. et al. Alteration of ornithine metabolism leads to dominant and recessive hereditary spastic paraplegia. Brain 138, 2191–2205 (2015).
Choi, K. D. et al. Genetic variants associated with episodic ataxia in Korea. Sci. Rep. 7, 13855 (2017).
Felício, D. & Santos, M. Spinocerebellar ataxia type 11 (SCA11): TTBK2 variants, functions and associated disease mechanisms. Cerebellum 23, 678–687 (2023).
Bouskila, M. et al. TTBK2 kinase substrate specificity and the impact of spinocerebellar- ataxia-causing mutations on expression, activity, localization and development. Biochem. J. 437, 157–167 (2011).
Bowie, E., Norris, R., Anderson, K. V. & Goetz, S. C. Spinocerebellar ataxia type 11-associated alleles of Ttbk2 dominantly interfere with ciliogenesis and cilium stability. PLoS Genet 14, e1007844 (2018).
Čajánek, L. & Nigg, E. A. Cep164 triggers ciliogenesis by recruiting Tau tubulin kinase 2 to the mother centriole. Proc. Natl. Acad. Sci. U S A. 111, E2841–E2850 (2014).
Rodrigues, C. H., Pires, D. E. & Ascher, D. B. DynaMut: predicting the impact of mutations on protein conformation, flexibility and stability. Nucleic Acids Res. 46, W350–W355 (2018).
Rodrigues, C. H. M., Pires, D. E. V. & Ascher, D. B. DynaMut2: assessing changes in stability and flexibility upon single and multiple point missense mutations. Protein Sci 30, 60–69 (2021).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Concordet, J. P. & Haeussler, M. CRISPOR: intuitive guide selection for CRISPR/Cas9 genome editing experiments and screens. Nucleic Acids Res. 46, W242–W245 (2018).
Kostyushev, D. et al. Suppressing the NHEJ pathway by DNA-PKcs inhibitor NU7026 prevents degradation of HBV CccDNA cleaved by CRISPR/Cas9. Sci. Rep. 9, 1–11 (2019).
Hodgkins, A. et al. WGE: a CRISPR database for genome engineering. Bioinformatics 31, 3078–3080 (2015).
Osório, H. et al. Proteomics analysis of gastric cancer patients with diabetes mellitus. J. Clin. Med. 10, 1–14 (2021).
Perez-Riverol, Y. et al. The PRIDE database resources in 2022: a hub for mass spectrometry-based proteomics evidences. Nucleic Acids Res. 50, D543–D552 (2022).
Shannon, P. et al. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Brandão, A. F., Sousa, A. P., Sequeiros, J. & Alonso, I. Ataxia causada Por Nova mutação no gene TTBK2: a Primeira família Portuguesa com ataxia Espinocerebelosa Tipo 11? Sinapse 13, 68 (2013).
Hubbert, C. et al. HDAC6 is a microtubule-associated deacetylase. Nature 417, 455–458 (2002).
Hata, A. & Chen, Y. G. TGF-β signaling from receptors to Smads. Cold Spring Harb Perspect. Biol. 8, a022061 (2016).
Yasui, D., Miyano, M., Cai, S. & Varga-Weisz, P. Kohwi-Shigematsu, T. SATB1 targets chromatin remodelling to regulate genes over long distances. Nature 419, 641–645 (2002).
Yang, Y. et al. Pokemon (FBI-1) interacts with Smad4 to repress TGF-β-induced transcriptional responses. Biochim. Biophys. Acta - Gene Regul. Mech. 1849, 270–281 (2015).
Dong, C. et al. PTBP3 mediates TGF-β-induced EMT and metastasis of lung adenocarcinoma. Cell. Cycle. 21, 1406–1421 (2022).
Lo, Y. H., Romes, E. M., Pillon, M. C., Sobhany, M. & Stanley, R. E. Structural Analysis Reveals Features of Ribosome Assembly Factor Nsa1/WDR74 Important for Localization and Interaction with Rix7/NVL2. Structure 25, 762–772.e4 (2017).
Wu, C. The PINCH–ILK–parvin complexes: assembly, functions and regulation. Biochim. Biophys. Acta - Mol. Cell. Res. 1692, 55–62 (2004).
Modregger, J., Ritter, B., Witter, B., Paulsson, M. & Plomann, M. All three PACSIN isoforms bind to endocytic proteins and inhibit endocytosis. J. Cell. Sci. 113, 4511–4521 (2000).
Tsujita, K. et al. Coordination between the actin cytoskeleton and membrane deformation by a novel membrane tubulation domain of PCH proteins is involved in endocytosis. J. Cell. Biol. 172, 269–279 (2006).
Abdollah, S. et al. TβRI phosphorylation of Smad2 on Ser465 and Ser467 is required for Smad2-Smad4 complex formation and signaling. J. Biol. Chem. 272, 27678–27685 (1997).
Klionsky, D. J. et al. Guidelines for the use and interpretation of assays for monitoring autophagy. Autophagy 8, 445–544 (2012).
Fiesel, F. C., Schurr, C., Weber, S. S. & Kahle, P. J. TDP-43 knockdown impairs neurite outgrowth dependent on its target histone deacetylase 6. Mol. Neurodegener. 6, 64 (2011).
Nekooki-Machida, Y. & Hagiwara, H. Role of tubulin acetylation in cellular functions and diseases. Med. Mol. Morphol. 53, 191–197 (2020).
Poirier, K. et al. Mutations in TUBG1, DYNC1H1, KIF5C and KIF2A cause malformations of cortical development and microcephaly. Nat. Genet. 45, 639–647 (2013).
Ruiz-Reig, N. et al. KIF2A deficiency causes early-onset neurodegeneration. Proc. Natl. Acad. Sci. 119, (2022).
Kalantari, S. & Filges, I. Kinesinopathies’: emerging role of the Kinesin family member genes in birth defects. J. Med. Genet. 57, 797–807 (2020).
Funaba, M., Zimmerman, C. M. & Mathews, L. S. Modulation of Smad2-mediated signaling by extracellular Signal-regulated kinase. J. Biol. Chem. 277, 41361–41368 (2002).
Kim, J. E., Lee, D. S., Kim, M. J. & Kang, T. C. PLPP/CIN-mediated NEDD4-2 S448 dephosphorylation regulates neuronal excitability via GluA1 ubiquitination. Cell. Death Dis. 10, 545 (2019).
Pereira, M. C., Morais, S., Sequeiros, J. & Alonso, I. Large-Scale functional RNAi screen in C. elegans identifies TGF-β and Notch signaling pathways as modifiers of CACNA1A. ASN Neuro. 8, 175909141663702 (2016).
Hekman, K. E. et al. A conserved eEF2 coding variant in SCA26 leads to loss of translational fidelity and increased susceptibility to proteostatic insult. Hum. Mol. Genet. 21, 5472–5483 (2012).
Jen, J. C., Wan, J., Palos, T. P., Howard, B. D. & Baloh, R. W. Mutation in the glutamate transporter EAAT1 causes episodic ataxia, hemiplegia, and seizures. Neurology 65, 529–534 (2005).
Collier, J. J. et al. Developmental consequences of defective ATG7-Mediated autophagy in humans. N Engl. J. Med. 384, 2406–2417 (2021).
Duenas, A. M. Molecular pathogenesis of spinocerebellar ataxias. Brain 129, 1357–1370 (2006).
Ma, S., Attarwala, I. Y. & Xie, X. Q. SQSTM1/p62: A potential target for neurodegenerative disease. ACS Chem. Neurosci. 10, 2094–2114 (2019).
Linares, J. F., Amanchy, R., Diaz-Meco, M. T. & Moscat, J. Phosphorylation of p62 by cdk1 controls the timely transit of cells through mitosis and tumor cell proliferation. Mol Cell. Biol 31, 105–117 (2011).
Kehl, S. R. et al. TAK 1 converts sequestosome 1/p62 from an autophagy receptor to a signaling platform. EMBO Rep 20, e46238 (2019).
Kim, J. et al. Differential regulation of distinct Vps34 complexes by AMPK in nutrient stress and autophagy. Cell 152, 290–303 (2013).
Gwinn, D. M. et al. AMPK phosphorylation of raptor mediates a metabolic checkpoint. Mol. Cell. 30, 214–226 (2008).
Allavena, G. et al. Suppressed translation and ULK1 degradation as potential mechanisms of autophagy limitation under prolonged starvation. Autophagy 12, 2085–2097 (2016).
Kiyono, K. et al. Autophagy is activated by TGF-β and potentiates TGF-β–Mediated growth Inhibition in human hepatocellular carcinoma cells. Cancer Res. 69, 8844–8852 (2009).
Acknowledgements
We would like to thank Isabel Alonso for her support in this study, as well as Elsa Logarinho (IBMC/i3S, Porto) for kindly providing HEK293T cells.
Funding
This work was funded by Ataxia UK – Small grant ZGRACA. It was also supported by national funds through FCT - Fundação para a Ciência e a Tecnologia, I.P., under the projects UIDB/00215/2020 (DOI: https://doi.org/10.54499/UIDB/00215/2020), UIDP/00215/2020 (DOI: https://doi.org/10.54499/UIDP/00215/2020), LA/P/0064/2020 (DOI: https://doi.org/10.54499/LA/P/0064/2020), and UID/215/2025. MS acknowledges funding from FCT through program DL 57/2016 – Norma Transitória. DS is the recipient of a fellowship (UI/BD/154402/2023) funded by FCT. This work had support from the Portuguese Mass Spectrometry Network, integrated in the National Roadmap of Research Infrastructures of Strategic Relevance (ROTEIRO/0028/2013; LISBOA-01-0145-FEDER-022125).
Author information
Authors and Affiliations
Contributions
D. Felício performed most of the experiments with the assistance of M. Santos. H. Osório performed the LC-MS experiments and assisted in data analysis. C. Pereira helped to design and implement CRISPR/Cas9 experiments. A.F. Brandão, J.P. Freixo and I. Carvalho performed the genetic analysis. A.P. Sousa and I. Carvalho identified the Portuguese family and performed clinical data collection and analysis. M. Santos conceived and supervised the study. D. Felício, M. Castro-Caldas, C. Lemos and M. Santos analyzed the data. M. Santos, C. Lemos and J. Sequeiros assisted with material resources and funding. D. Felício and M. Santos wrote the first draft of the manuscript. All authors critically revised and edited the manuscript. All authors approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Felício, D., Osório, H., Pereira, C. et al. Missense variant in TTBK2 kinase domain causes loss of function and impaired protein phosphorylation. Sci Rep 16, 2501 (2026). https://doi.org/10.1038/s41598-025-32288-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-32288-0