Abstract
Genistein and daidzein are isoflavonoid phytoalexins that increase rapidly during bean hypersensitive immunity to Pseudomonas savastanoi pv. phaseolicola. To understand how genistein and daidzein affect P. savastanoi pv. phaseolicola, non-targeted metabolomic mass spectrometry was performed on the bacterium treated in vitro. The bacterium catabolized genistein and daidzein and responded by producing several different classes of compounds including auxin-like indoles and cyclic dipeptides. Non-targeted metabolomic investigation of bean leaves infiltrated with the cyclic dipeptides revealed no similarities to auxin-induced metabolic changes, but one cyclic dipeptide, cyclo-Trp-Pro (cWP), induced the accumulation of phytoalexins. This implied that cWP application might make beans more resistant to pathogens. Challenge with Uromyces appendiculatus, a rust fungal pathogen, revealed that beans pretreated with cWP had 90% reductions in disease. Arabidopsis thaliana sprayed with cWP had activated salicylic acid-mediated immune responses. Overall, these results reveal that P. savastanoi pv. phaseolicola is adapted to tolerate bean genistein and daidzein, likely sensing the compounds as host signals and producing cyclic dipeptides in response. In turn, beans respond to at least one cyclic dipeptide, cWP, by producing phytoalexins to increase resistance. cWP may be useful for protecting beans and other plants from microbial disease.
Similar content being viewed by others
Introduction
Phytoalexins are antibiotics deployed by plants during immune responses to pathogens1. Isoflavonoid phytoalexins, e.g. glyceollin, genistein, daidzein, coumestrol, and related stilbenoids like resveratrol, are derived from phenylalanine and are commonly found in legumes2,3. By contrast, the phytoalexin camalexin is an indole derived from tryptophan and is commonly found in crucifers1. Pioneering research established that phytoalexins purified from plants inhibited the replication of cellular pathogens in vitro2. Subsequent genetics and molecular biology studies established the roles of phytoalexins in planta. For example, Arabidopsis thaliana pad3 mutants deficient in camalexin biosynthesis are more susceptible to Alternaria brassicicola4,5. Suppression of isoflavone synthase in soybean led to decreased accumulation of glyceollin and decreased resistance to Phytophthora sojae6. Transgenic tobacco engineered to produce more resveratrol had increased resistance to Botrytis cinerea7, and transgenic alfalfa expressing isoflavone 7-O-methyltransferase produced more phytoalexins and had increased resistance to Phoma medicaginis8. These findings helped codify the phytoalexin theory of plant immunity whereby phytoalexins stop disease progression due to their antibiotic activity.
Genistein and daidzein are the two main isoflavonoid phytoalexins produced in bean leaves after infection with Pseudomonas savastanoi pv. phaseolicola, the bacterium that causes halo blight3. Genistein and daidzein are made from 2,4’,5,7-tetrahydroxyisoflavanone and 2,4’,7-trihydroxyisoflavanone, respectively, and diverse phytoalexins and their glycosylated forms are subsequently produced from genistein and daidzein by divergent enzymatic pathways9. Daidzin, coumestrol, glyceollin, medicarpin, and pisatin, for example, are derived from daidzein while genistin and prunetin are derived from genistein. Chemically, genistein is hydroxylated daidzein10, yet this structural difference could provide unique biochemical activities. In human and mammalian cell systems, genistein acts as a tyrosine kinase inhibitor, but daidzein does not11,12,13,14.
Genistein slows the growth of pathogenic and enteric bacteria of mammals in vitro, but greater concentrations are needed to slow the bean-adapted P. savastanoi pv. phaseolicola3,15,16,17. In our ongoing research to understand how phytoalexins biochemically affect bacteria18,19,20, we found that P. savastanoi pv. phaseolicola can enzymatically degrade genistein21. This likely explains why P. savastanoi pv. phaseolicola suffers fewer detrimental proteomic changes from genistein exposure compared to salicylic acid and resveratrol which are more toxic19,20. But not only does P. savastanoi pv. phaseolicola tolerate genistein by degrading it, the bacterium responds by producing varied indole alkaloids that might exert hormonal functions in beans to favor bacterial pathogenicity21. These observations imply that genistein could be a pathogenic trigger for P. savastanoi pv. phaseolicola infection. Such a possibility would be consistent with the findings that genistein induces nod gene expression to enable rhizobium symbiosis with soybean roots and with the observation that genistein induces P. sojae (an oomycete) pathogenicity of soybean roots22,23. Daidzein also induces nod gene expression and P. sojae pathogenicity despite the hydroxylation difference22,23. Thus, we sought to investigate how P. savastanoi pv. phaseolicola responded metabolically to daidzein versus genistein. We found that both genistein and daidzein were modified and degraded and that exposure to both phytoalexins induced the bacterial production of indoles and cyclic dipeptides. We discovered that one of the cyclic dipeptides increased the production of daidzein, genistein, and other phytoalexins when applied back to beans. The increase in phytoalexins after cyclic dipeptide application was sufficient to enhance bean resistance to fungal infection.
Results
Genistein and daidzein alter small molecule accumulation in a bacterial pathogen
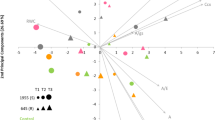
Avirulent P. savastanoi pv. phaseolicola race 5 (R5) induces hypersensitive immunity on Phaseolus vulgaris Black Valentine bean leaves while virulent race 8 (R8) produces water-soaking disease24. At the sites of inoculation with either race, the leaf cells accumulate genistein, daidzein, and other phytoalexins, but greater amounts can accumulate during hypersensitive immunity to R5 3. This is because the avirulent bacterium triggers immune system amplification whereas the virulent bacterium suppresses the immune system. Because R5 and R8 are exposed to many different phytoalexins in addition to genistein and daidzein in bean leaves, it is not possible to know how much any single phytoalexin contributes to immunity. Thus, we have used in vitro, liquid growth assays to assess the individual effects of phytoalexins on P. savastanoi pv. phaseolicola and estimated the minimum inhibitory concentrations of genistein and daidzein to be 400 µM and 500 µM, respectively3. We discovered that in the presence of 400 µM genistein in vitro, R5 degraded genistein and produced indole alkaloids21. To assess whether R5 responds to daidzein similarly, we performed a comparative metabolomic analysis on cell cultures. We lowered the concentration of daidzein to that of genistein rather than use any additional genistein. Thus, R5 liquid cultures were separately exposed to 400 µM genistein, 400 µM daidzein, or an equivalent volume of control dimethyl sulfoxide (DMSO), and the cell extracts were analyzed by non-targeted mass spectrometry. There were 1,719 compound features found in positive ion mode and 2,072 found in negative ion mode. The main compounds from genistein-treated cells with significantly increased accumulation (calculated from compound chromatographic peak areas) were genistein and its degradation products of smaller mass like 2,3-dihydro-4 H-chrome-4-one or its converted products like 5-O-methylgenistein with greater mass (Fig. 1; Supplementary Dataset S1). The main compounds from daidzein-treated cells with significantly increased accumulation were daidzein and its degradation products like dioxybenzone or its converted products like formononetin (Fig. 1; Supplementary Dataset S1). These compounds were identified by tandem mass spectrum spectral library matching for Level 2 identification25,26 (Supplementary Dataset S1). In addition to accumulating these specific degradation products and derivatives of daidzein and genistein, the cells had increased amounts of indole compounds such as N-acetylserotonin and trans-3-indoleacrylic acid (Fig. 1; Supplementary Dataset S1). Some indoles such as 4-indolecarbaldehyde, indole-3-propionitrile, 2-(2-piperidinyl)-1 H-indole, and an isomer of indole-3-acetic acid, as well as several indole beta-carbolines like harmaline, harmol, harmane, and norharman that were previously found in genistein-treated cells21 were induced to a higher extent in daidzein-treated cells (Fig. 1; Supplementary Dataset S1). Daidzein- and genistein-treated P. savastanoi pv. phaseolicola R5 also produced phenylacetoacetonitrile, a derivative of phenylacetonitrile and phenylacetic acid that each have indole-3-acetic acid-like plant hormonal activity (Fig. 1; Supplementary Dataset S1)27,28. Overall, these findings demonstrate that P. savastanoi pv. phaseolicola R5 catabolizes daidzein and genistein and responds by producing indole compounds. We hypothesize that some of the indole compounds could favor bacterial pathogenicity, possibly by modifiying plant host metabolism through auxin-like hormonal pathways in a way that indole-3-acetic acid (auxin) does for P. syringae29.
Compounds found in extracts of P. savastanoi pv. phaseolicola R5 exposed to 400 µM genistein versus 400 µM daidzein (each compared to DMSO controls). The compounds (gray circles) were detected by non-targeted mass spectrometry and their abundances calculated from their chromatographic peak areas. Selected compounds with statistically significant accumulation changes are denoted with blue squares and lettering (FDR < 5%, n = 5). cFP, cyclo-Phe-Pro; cGP, cyclo-Gly-Pro; cLP, cyclo-Leu-Pro; and cWP, cyclo-Trp-Pro.
Another class of compounds induced by daidzein and genistein was cyclic dipeptides. Four of the notable cyclic dipeptides were formed with proline, namely cyclo-Phe-Pro (cFP), cyclo-Gly-Pro (cGP), cyclo-Leu-Pro (cLP), and cyclo-Trp-Pro (cWP) (Fig. 1; Supplementary Dataset S1). Cyclic dipeptides are intriguing because some like cFP, cLP, and cyclo-Tyr-Pro function in bacterial quorum sensing30 while others like cFP, cyclo-Val-Pro, and cyclo-Tyr-Pro may promote auxin-like cell growth in A. thaliana31. Thus, we hypothesize that the bacterial production of cyclic dipeptides in response to genistein or daidzein could support pathogenicity.
Cyclic dipeptides alter small molecule accumulation in bean leaves
To begin assessing any potential roles these bacterial compounds may have as pathogenicity factors, we examined whether they could alter plant cell metabolism to favor bacterial infection. We focused on the cyclic dipeptides because cFP, cyclo-Val-Pro, and cyclo-Tyr-Pro promote weak, auxin-like cell growth in A. thaliana31. We first confirmed the identification of the cyclic dipeptides using pure standards (Fig. 2a–d). We also demonstrated that P. savastanoi pv. phaseolicola race 8 (R8) produces similar amounts of the same cyclic dipeptides as R5 when treated with 400 µM daidzein (Fig. 2e). R5 elicits hypersensitive immunity on Black Valentine beans whereas R8 does not24. Thus, cFP, cGP, cLP, and cWP are made by virulent and avirulent P. savastanoi pv. phaseolicola races.
Cyclic dipeptides from P. savastanoi pv. phaseolicola, and metabolomics after P. vulgaris infiltration. (a–d) Mirror plots of tandem mass spectra for cyclic dipeptides for (a) cFP, (b) cGP, (c) cLP, and (d) cWP produced in genistein- and daidzein-treated P. savastanoi pv. phaseolicola R5 versus chemical standards. The observed spectrum is on top of each plot and the standard is on the bottom. (e) Mean chromatographic peak areas of cFP, cGP, cLP, and cWP in daidzein-treated P. savastanoi pv. phaseolicola R5 and R8 [error bars depict standard deviation (the differences are not significant); n = 5]. (f) Principal component (PC) analysis of compounds detected by mass spectrometry in positive ion mode from P. vulgaris infiltrated with 30 µM auxin, cFP, cGP, cLP, cWP, or water 24 h after infiltration (n = 4). (g–i) Volcano plots of fold changes of compounds from P. vulgaris treated with (g) 30 µM auxin, (h) 30 µM cFP, and (i) 30 µM cWP versus water-treated controls as a function of − log10(p-value) of their chromatographic peak areas. For (g-i), compounds with FDR < 5% (Tukey HSD p-value < 1%) and significant fold-changes are represented as solid color dots, and those without significant fold-changes or with FDR > 5% are represented as gray circles. cFP, cyclo-Phe-Pro; cGP, cyclo-Gly-Pro; cLP, cyclo-Leu-Pro; and cWP, cyclo-Trp-Pro.
Next, we infiltrated 30 µM concentrations of cFP, cGP, cLP, and cWP into bean leaves. We chose these concentrations because cFP induced auxin-like growth activity in A. thaliana at the same concentration31. Speculating these cyclic dipeptides may incite auxin-like activity in beans, especially cWP which has an indole functional group, we tested 30 µM indole-3-acetic acid (auxin) for comparison. Principle Component analysis of 4,274 positive ion mode compound features resolved by mass spectrometry performed on infiltrated leaf extracts demonstrated that only cWP-treatment compound features clustered separately from the others (Fig. 2f). Statistical analysis of compound peak areas revealed that auxin treatment led to the fewest significant changes in abundance (67), the main identifiable compound being a conjugated, inactivated form of indole-3-acetic acid (Fig. 2g; Supplementary Dataset S2). cLP treatment produced 171 significant changes in compound abundance whereas cGP produced 207 (Supplementary Dataset S2). The infiltrated cLP and cGP were the main compounds identified in each case, but there were few other identifiable compounds that we could link to a plausible metabolic response model. cFP treatment led to 543 significant changes including significant increases in 12-oxo phytodienoic acid and 9(S)-HpOTrE, each of the alpha-linolenic acid pathway (Fig. 2h; Supplementary Dataset S2). Phytoprostane A1, also part of the alpha-linolenic acid pathway, decreased (Fig. 2h; Supplementary Dataset S2). These findings imply that cFP may have affected parts of fatty acid biosynthesis, especially the alpha-linolenic acid pathway that leads to jasmonic acid biosynthesis. Prior research has linked cFP to alterations in auxin-like hormonal growth responses in A. thaliana31, but we did not observe responses with cFP consistent with auxin treatment. Consequently, the results for cFP are difficult to interpret regarding pathology and may warrant deeper investigations in beans.
Compared to these cyclic dipeptides and auxin, cWP induced the most compound abundance changes (686; Supplementary Dataset S2). Amongst them were increases in daidzein and genistein and their glycosylated derivatives daidzin, genistin, and 6”-O-malonylgenistin (Fig. 2i). Several structurally similar flavonoids and glycosylated isoflavonoids also were induced (Fig. 2i; Supplementary Dataset S2). We repeated bean leaf infiltrations with 30 µM cWP. Again, we observed drastic increases in daidzein, genistein and their derivatives, and greater increases in other flavonoids and isoflavonoids including daidzein-derived coumestrol (Supplementary Dataset S3). These findings reveal that cWP, but not cFP, cGP, or cLP, elicits phytoalexin production in beans.
cWP induces disease resistance in beans
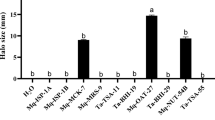
We tested whether phytoalexin increases induced by cWP treatment affected disease resistance. Experimental challenges with virulent P. savastanoi pv. phaseolicola R8 on bean leaves pretreated 24 h in advance with cWP revealed no reduction in bacterial spread and no delay in appearance of bacteria compared to water-pretreated controls as visualized by bacterial autofluorescence (Fig. 3a,b). We speculated that R8 might be adapted to overcoming the phytoalexin response initiated by cWP. So instead, we tested Uromyces appendiculatus race 41, a rust fungus virulent on P. vulgaris variety Black Valentine. The asexual rust uredospores applied to the surface of the leaf will germinate in dew, and their hyphae will actively enter the leaf apoplast through leaf stomata based on thigmotropic recognition32. The number of resulting leaf pustules, the asexual fruiting bodies, is directly proportional to the number of successful fungal infections. Preliminary tests with rust on beans treated with 30 µM cWP revealed a potential 50% reduction in pustules. When we increased the concentration of cWP to 300 µM, we observed 75–90% reductions in numbers of pustules when cWP was applied to primary bean leaves 24 h prior to inoculation with rust spores (Fig. 3c,d). To see if cWP was directly inhibiting rust spore germination, we sprayed glass microscope slides with 300 µM cWP and then sprayed rust spores on the slides as we would for leaf inoculation. After overnight incubation in a dew chamber, spores exposed to cWP germinated normally and had elongated hyphae that appeared to be the same as water-treated slide controls (Fig. 3e,f). By contrast, there was near-complete germination inhibition on slides sprayed with 5% cycloheximide, a natural fungicide produced by Streptomyces griseus (Fig. 3g). These results reveal that cWP is not fungicidal, per se, in its efficacy toward U. appendiculatus but rather functions by increasing the amounts of phytoalexins in leaves.
Effects of cWP on P. vulgaris variety Black Valentine, U. appendiculatus (rust) spores, and A. thaliana. (a) Autofluorescence (transillumination 366 nm) of P. savastanoi pv. phaseolicola race 8 (R8) 4 days after inoculation on Black Valentine bean leaves sprayed with water 24 h prior to inoculation. (b) Autofluorescence (transillumination 366 nm) of R8 4 days after inoculation on Black Valentine sprayed with 300 µM cWP 24 h prior to inoculation. [There is no perceptible difference in spread of R8 between (a) and (b)]. (c) Average number of U. appendiculatus race 41 pustules on water-treated or 300 µM cWP-treated Black Valentine. Leaves were inoculated 24 h after cWP or water treatment. Three trials were performed (p-values are for t-test comparisons of numbers of pustules; error bars represent standard deviation; n = 16). (d) Signs of U. appendiculatus race 41 disease (rust pustules) on primary leaves of Black Valentine treated with water or 300 µM cWP. (e) U. appendiculatus race 41 spore germlings on a glass slide sprayed with 300 µM cWP. (f) U. appendiculatus race 41 spore germlings on a glass slide sprayed with water. (g) Lack of U. appendiculatus race 41 spore germlings on a glass slide sprayed with 5% cycloheximide. (h) Statistically significant Log2 normalized transcript abundance counts of genes in Black Valentine leaves 24 h after 300 µM cWP or water treatment. FC is fold-change for cWP vs. water (n = 4). Genes with corresponding proteins induced by benzothiadiazole treatment of Black Valentine leaves are marked with an asterisk. (i) Mean chromatographic peak areas of compounds found by tandem mass spectrometry in A. thaliana treated with 300 µM cWP or water (p-values are for t-test comparisons of normalized peak areas; error bars represent standard deviation; n = 6).
RNA sequencing of cWP-treated beans confirmed increased expression of genes for enzymes that catalyze key steps of isoflavonoid biosynthesis including phenylalanine ammonia lyase, chalcone synthase, and 2-hydroxyisoflavonone synthase genes (Fig. 3h; Supplementary Dataset S4). Furthermore, there was increased expression of leucine-rich repeat and cysteine-rich receptor kinases and other candidate immune receptors and signaling proteins (Fig. 3h; Supplementary Dataset S4). A few of these genes had increased abundances of proteins in beans sprayed with benzothiadiazole, an unrelated synthetic chemical that induces immunity to rust33 (Fig. 3h; Supplementary Dataset S4). These results revealed that not only did cWP induce phytoalexin production, but it also primed the bean immune system for the detection of pathogens and for concomitant immune system signaling. This attribute also was observed in cWP-sprayed A. thaliana which subsequently accumulated six times more salicylic acid and pipecolic acid, regulators of acquired and systemic acquired resistance34,35 (Fig. 3i; Supplementary Dataset S5). Camalexin, a tryptophan-derived phytoalexin controlled by the salicylic acid defense pathway36, increased by 32-fold, and bactericidal glucosinolates glucoerucin and glucoberteroin increased by more than 15-fold (Fig. 3i; Supplementary Dataset S5)37.
Discussion
Over the last few decades we have learned exquisite details about the interactions between plants and their pathogens and how this leads to disease susceptibility or immunity38. The immune response includes physiological, transcriptional, translational, post-translational, enzymatic, and metabolic responses. Specific metabolic responses include the production of phytoalexins, small molecules with antibiotic potential that accumulate at the site of infection to stop disease progression. The accumulation of a variety of phytoalexins and other toxic compounds could explain why induced immunity kills most invading bacteria39, although additional mechanisms of immunity could lead to the starvation, dehydration, or immobilization of bacterial invaders39,40,41,42. Nevertheless, how plant immunity output, including phytoalexin toxicity, acts upon bacterial pathogens and how bacterial pathogens respond to plant immune system output remain among some of the crucial but understudied topics in the field.
We have reopened investigation into these topics and demonstrate here that P. savastanoi pv. phaseolicola, a bacterial pathogen of beans, metabolically degrades genistein and daidzein, the two major isoflavonoid phytoalexins of beans, and responds by producing indoles and cyclic dipeptides (Fig. 1). These indoles and cyclic dipeptides contain nitrogen whereas genistein and daidzein do not. Thus, they are produced in response to, not from, genistein and daidzein. These findings reveal an intriguing possiblity whereby this adapted pathogen of beans perceives genistein and daidzein, isoflavonoids that differ by a hydroxylation moiety10, as a signals to drive infection. This is consistent with observations that symbiotic rhizobia of soybean and pathogenic P. sojae tolerate genistein and daidzein23,43, and that genistein and daidzein exudates from roots serve as signals to direct rhizobia symbiosis and P. sojae infections in soybean, a legume related to bean22.
These observations prompted us to investigate whether the induced bacterial compounds assisted infection by modifying plant cell metabolism. In particular, we focussed on the potential biological activity of the cyclic dipeptides. While it is known that bacteria use cyclic dipeptides like cLP, cFP, cyclo-Tyr-Pro, and cyclo-Val-Pro for bacterial quorum sensing30,31,44, only a few studies have shown the effects of cyclic dipeptides on plants. In A. thaliana, cFP, cyclo-Tyr-Pro and cyclo-Val-Pro induced weak auxin-like effects31. In Nicotiana benthamiana, cyclo-Pro-Pro (but not cFP) induced local acquired resistance to Phytophthora nicotianae45. Our metabolomic investigation revealed that neither cFP, cGP, cLP, nor cWP elicited a metabolic response similar to a herbicidal concentration of indole-3-acetic acid (Fig. 2g–i). Rather, we found that cWP elicited phytoalexin production in beans and primed immune system gene expression (Figs. 2i and 3h). These changes likely contributed to increased resistance to rust fungal infection (Fig. 3c,d). Because cWP also elicited phytoalexin and glucosinolate production in A. thaliana (Fig. 3i), it is possible that other plants will respond to cWP in a similar way, and that cWP, a natural compound, can be used to deter multiple diseases in numerous crops.
Limitations and future directions
Whether cyclic dipeptides assist P. savastanoi pv. phaseolicola quorum sensing or are produced to serve other bacterial functions remains to be determined. Nevetheless, our discovery of the bacterial production of cWP upon treatment with plant phytoalexins and the induction of phytoalexins by cWP implies there is a hypothetical biochemical feedback loop between the host and its pathogen. Although P. savastanoi pv. phaseolicola encounters amplified amounts of genistein and daidzein during an immune reaction in beans3 and increases its cWP production in the presence of an inhibitory amount of genistein and daidzein in vitro (Fig. 1), we do not know if bacterial cWP is made at the site of infection or if bacterial cWP induces bean immune signaling during infection. A recent report revealing that A. thaliana produces cyclo-His-Pro during abiotic stress responses opens the door to the possibility that beans could also make cWP or other cyclic dipeptides46. If so, does this mean that plant cWP is regulated by the plant immune system? Is there a cWP plant receptor that can recognize bacterial cWP? Does the amount of bacterial cWP overcome a plant cWP threshold to induce biochemical immunity feedback? These intriguing questions deserve future attention.
Methods
Bacterial strains and genistein and daidzein treatment
P. savastanoi pv. phaseolicola isolates race 5 (R5) and 8 (R8) were maintained on King Agar B solid medium (Sigma-Aldrich, St. Louis, MO) supplemented with glycerol. The genome sequences for R5 and R8 are reported47. Genistein and daidzein were obtained from Sigma-Aldrich and were solubilized in dimethyl sulfoxide (DMSO). R5 and R8 were cultured in 10 mL Luria broth with 400 µM genistein, 400 µM daidzein, or without but with 50 µL DMSO at 28 °C at 200 rpm on a shaking platform until the control cultures reached late log-phase growth measured at an optical density of 0.75–0.90 at 600 nm wavelength of visible light.
Mass spectrometry of metabolites
Five replicate R5 cultures of each treatment were centrifuged at 5,000 x g for 10 min, washed once in 1 mL phosphate buffered saline, centrifuged at 5,000 x g for 10 min, resuspended in 1 mL of 1:1:1:1 acetone/acetonitrile/methanol/water and 1,250 pmol prednisone, and pulverized with 0.5 mm glass beads with a Qiagen PowerLyzer 24 bead mill (Qiagen, Hilden, Germany) 10 times at 5,000 rpm for 20 s (cooled on ice for 2 min between cycles). Two matrix blank samples were prepared the same way starting with 10 mL Luria broth (without R5 and without genistein or daidzein). The milled extracts were centrifuged for 20 min at 12,000 x g. The supernatants were transferred to fresh tubes and centrifuged again. The supernatants were transferred to glass vials and dried under vacuum. The residues were resuspended in 115 µL 50% methanol/0.1% formic acid. Fifteen µL of each sample was pooled to create a quality control (QC) sample. Five µL of the samples were separated on a 150 × 2.1 mm Hypersil GOLD VANQUISH HPLC column with 1.9 μm particles (Thermo Fisher Scientific) coupled to a Vanquish HPLC pump (Thermo Fisher Scientific) controlling a 10-min linear gradient from 0% to 95% acetonitrile and 0.1% formic acid at a flow rate of 0.2 mL per min. Eluent was electrosprayed at 3.5 kV positive polarity into an Exploris 240 mass spectrometer (Thermo Fisher Scientific) using an internal mass calibrant. Sheath gas was 35, auxiliary gas was 7, and sweep gas was 1 (arbitrary units). The ion transfer tube temperature was 325 °C, and the vaporizer temperature was 275 °C. Advanced peak determination, mild trapping, and internal mass calibration was enabled. Default charge state was 1, and the expected peak width was 6 s. AcquireX software was used to create a background ion exclusion list from the matrix blank sample and an inclusion ion list from the QC sample48. Five subsequent injections of the QC were performed to generate tandem mass spectra (MS2). After each QC injection, the resolved ions were appended to the exclusion list. Survey scans were recorded in the Orbitrap at 60,000 resolution over a range of m/z 70–800. The RF lens was 70%. Monoisotopic precursor selection was enabled, the minimum intensity was 5,000, and charge states were filtered to 1. Precursors selected within a 1.0 Da isolation window were fragmented by high energy collision-induced dissociation (30%, 50%, 70% normalized stepped collision energy), and fragment ions were resolved in the Orbitrap at 30,000 resolution. Subsequently, all test samples were analyzed alongside QC and matrix blank samples in the following order: Matrix blank 1, QC 1, sample replicate set 1, Blank 2, QC 2, sample replicate set 2, etc. Survey scans were recorded in the Orbitrap at 120,000 resolution over a range of m/z 70–800. The RF lens was 70%. The samples also were analyzed in negative ion mode at -2,500 V but with the same other settings.
Result raw files were analyzed with Compound Discoverer version 3.3 (Thermo Fisher Scientific). The ChromAlign node was used to align chromatographic peaks in all files to a QC replicate. The Detect Compounds node was used with 2 ppm mass tolerance, 10,000 minimum peak intensity, at least 5 scans per peak, 0.25 min maximum peak width, and compound detection of [M + H] + 1 ions for positive mode data and [M-H]-1 ions for negative mode data to create peak areas for resolved ions. The Group Compounds node was used with 2 ppm mass tolerance, 0.25 min retention time (RT) tolerance, and a peak rating threshold of 4 for a minimum of 6 files. The Fill Gaps node was used with 2 ppm mass tolerance, the SERRF QC Correction node was used with 80% QC coverage to normalize the peak area results, and the Mark Background Components node was enabled. The Search mzCloud node was used to compare MS2 spectra to all compound classes at a precursor mass tolerance of 5 ppm and fragment mass tolerance of 5 ppm. The Search mzVault node was used to compare MS2 spectra to the NIST_2020_MSMS High Resolution library at fragment mass tolerances of 10 ppm. The Assign Compound Annotations, Map to KEGG Pathways, Search ChemSpider, and Predict Compositions nodes were used with 2 ppm mass tolerances. A one-way ANOVA with a Tukey post-hoc test was used to estimate compound peak area statistical differences. The Benjamini-Hochberg method was used to adjust the p-values. Filtering of results was performed to limit background ions, to require MS2 supporting spectra, for the false discovery rate (FDR) to be less than 5%, and mzCloud or NIST2020_MSMS scores ≥ 70 (Supplementary Dataset S1).
Cyclic dipeptides and application to bean and Arabidopsis thaliana leaves
Cyclo-L-phenylalanine-L-proline (cFP) was obtained from Toronto Research Chemicals (Toronto, ON, Canada). Cyclo-L-leucine-L-proline (cLP) and cyclo-L-tryptophan-L-proline (cWP) were obtained from Ambeed (Arlington Hts, IL). Cyclo-glycine-L-proline (cGP) was obtained from Accela (San Diego, CA). Approximately 1 nmol of each cyclic dipeptide was analyzed for RT on the reverse phase column and to produce a reference tandem mass spectrum under the mass spectrometry conditions noted above.
Primary leaves of 10-day-old P. vulgaris (common bean) variety Black Valentine plants were infiltrated with 30 µM cFP, cGP, cLP, cWP, or auxin (indole-3-acetic acid, Sigma Aldrich), or water (control) using a flat-nosed 1 mL syringe. We chose these concentrations because cFP induced auxin-like growth activity in A. thaliana at the same concentration31. The two primary leaves on each plant were each infiltrated 8 times (4 times on each half leaf) with a single compound. Infiltration was performed on the abaxial side without puncturing the leaf, and each infiltration produced a 1 cm2 water-soaked area. After 24 h, the leaf tissue at the site of infiltration was sampled with a 1 cm diameter cork-borer. Ten to twelve sites were collected from each plant to reach approximately 170 mg total tissue which was extracted in 1 mL 100% methanol using glass beads as described above. Four separate plants of each treatment were used to generate four replicates. A one-way ANOVA with a Tukey post-hoc test was used to estimate compound peak area statistical differences (Supplementary Dataset S2). In a later, focused experiment to compare cWP to water, seven replicates were used, and the tissues were collected at 24 h after infiltration. Non-targeted mass spectrometry was performed the same as above, but reference library searches included the cyclic dipeptide reference spectra. An empirical Bayes statistics test was used in the LIMMA package in the R programming environment to assess compound normalized peak area statistical differences and to estimate the FDR (the Benjamini-Hochberg method). Filtering of results was performed to limit background ions, to require MS2 supporting spectra, for the FDR to be less than 5%, and mzCloud or NIST2020_MSMS scores ≥ 70 (Supplementary Dataset S3).
R8 was cultured on King Agar B solid medium supplemented with glycerol, resuspended in water, and adjusted to an optical density at 600 nm = 2. The abaxial sides of two primary leaves of 10-day-old P. vulgaris variety Black Valentine were each inoculated 10 times using a Master airbrush at 50 psi at a 1- to 2-cm distance. Water was sprayed as a control. Bacterial infection and spread was assessed by bacterial autofluorescence visualized by transillumination at 366 nm.
Two primary leaves from 8, 10-day-old P. vulgaris variety Black Valentine plants were sprayed with a liquid suspension of 300 µM cWP or water. A day later, the leaves were inoculated with a liquid suspension of uredospores of U. appendiculatus race 41 and placed in an 18 °C dew chamber for 12 h and then moved to a 23 °C growth chamber with fluorescent lighting and 50% relative humidity49. Inoculum was adjusted to produce 400–500 pustules per leaf. Pustules were enumerated on all plants when they appeared on water-treated controls inoculated with race 41. Sixteen leaves were examined per treatment, and three biological replicate experiments were evaluated. A two-tailed Student’s t-test was used to assess differences in numbers of pustules.
To assess spore germination, glass slides were sprayed with 300 µM cWP, water, or 5% cycloheximide (germination inhibition positive control). When the slides dried, they were sprayed with a liquid suspension of uredospores of U. appendiculatus race 41 and placed in an 18 °C dew chamber. Spore germination was assessed by light microscopy 18 h later.
Primary leaves from four Black Valentine plants were sprayed with a liquid suspension of 300 µM cWP or water. 24 h later, RNA was purified from the leaves using the Qiagen RNeasy Plant Mini Kit. mRNA sequencing via polyA selection was performed by Genewiz (South Plainfield, NJ) with an Illumina 2 × 150 bp platform. The overall project yielded 164 × 106 reads, and 49 × 103 Mbases with a mean quality score of 38.02. Sequence reads were trimmed to remove adapter sequences and nucleotides with poor quality using Trimmomatic v.0.36. The trimmed reads were mapped to the Phaseolus_vulgaris_2_0 reference genome available on ENSEMBL using STAR aligner v.2.5.2b. Unique gene hit counts were calculated by using featureCounts from the Subread package v.1.5.2. The hit counts were summarized and reported using the gene_id feature in the annotation file. Only unique reads that fell within exon regions were counted. Comparisons of gene expression between water and cWP-treated groups of samples were performed using DESeq2. The Wald test was used to generate p-values and log2 fold changes. Genes (1,341) with an adjusted p-value < 0.05 and absolute log2 fold change > 1 were called as differentially expressed genes. Gene names were linked to corresponding JGI accession numbers, A. thaliana gene orthologs, Gene Ontology process and function terms, and differentially accumulating proteins from benzothiadiazole-treated Black Valentine33 (Supplementary Dataset S4).
Leaves of 3-week-old A. thaliana ecotype Columbia-0 were sprayed with 300 µM cWP or water. After 24 h, approximately 150 mg of leaf tissue was collected from 7 to 10 plants and pooled to form a replicate. This was performed six times for each treatment. The samples were prepared and analyzed by mass spectrometry as described above. A two-tailed Student’s t-test was used to assess compound normalized peak area statistical differences and p-values were adjusted by the Benjamini-Hochberg method (Supplementary Dataset S5).
Data availability
Mass spectrometry data files can be retrieved from https://massive.ucsd.edu/. (1) Pseudomonas savastanoi pv. phaseolicola race 5 treated with genistein, daidzein, and DMSO. (MSV000096150). (2) Phaseolus vulgaris infiltrated with cyclic dipeptides and auxin. (MSV000096164). (3) Phaseolus vulgaris infiltrated with cWP and water. (MSV000096165). (4) Arabidopsis thaliana sprayed with cWP or water. (MSV000096166).
Abbreviations
- DMSO:
-
Dimethyl sulfoxide
- QC:
-
Quality control
- MS2 :
-
Tandem mass spectra
- RT:
-
Retention time
- FDR:
-
False discovery rate
- cFP:
-
Cyclo-L-phenylalanine-L-proline
- cLP:
-
Cyclo-L-leucine-L-proline
- cWP:
-
Cyclo-L-tryptophan-L-proline
- cGP:
-
Cyclo-glycine-L-proline
- PC:
-
Principal component
References
Piasecka, A., Jedrzejczak-Rey, N. & Bednarek, P. Secondary metabolites in plant innate immunity: conserved function of divergent chemicals. New Phytol. 206, 948–964 (2015).
Kuc, J. Phytoalexins, stress metabolism, and disease resistance in plants. Annu. Rev. Phytopathol. 33, 275–297 (1995).
Cooper, B., Campbell, K. B. & Garrett, W. M. Salicylic acid and phytoalexin induction by a bacterium that causes halo blight in beans. Phytopathology 112, 1766–1775 (2022).
Thomma, B. P. H. J., Nelissen, I., Eggermont, K. & Broekaert, W. F. Deficiency in phytoalexin production causes enhanced susceptibility of Arabidopsis Thaliana to the fungus alternaria brassicicola. Plant J. 19, 163–171 (1999).
Glazebrook, J. & Ausubel, F. M. Isolation of phytoalexin-deficient mutants of Arabidopsis Thaliana and characterization of their interactions with bacterial pathogens. Proc. Natl. Acad. Sci. U S A. 91, 8955–8959 (1994).
Graham, T. L., Graham, M. Y., Subramanian, S. & Yu, O. RNAi Silencing of genes for elicitation or biosynthesis of 5-deoxyisoflavonoids suppresses race-specific resistance and hypersensitive cell death in phytophthora Sojae infected tissues. Plant. Physiol. 144, 728–740 (2007).
Hain, R. et al. Disease resistance results from foreign phytoalexin expression in a novel plant. Nature 361, 153–156 (1993).
He, X. Z. & Dixon, R. A. Genetic manipulation of isoflavone 7-O-methyltransferase enhances biosynthesis of 4’-O-methylated isoflavonoid phytoalexins and disease resistance in alfalfa. Plant. Cell. 12, 1689–1702 (2000).
Ahuja, I., Kissen, R. & Bones, A. M. Phytoalexins in defense against pathogens. Trends Plant. Sci. 17, 73–90 (2012).
Kim, M., Han, J. & Kim, S. U. Isoflavone daidzein: chemistry and bacterial metabolism. J. Appl. Biol. Chem. 51, 253–261 (2008).
Ogawara, H., Akiyama, T., Ishida, J., Watanabe, S. & Suzuki, K. A specific inhibitor for tyrosine protein kinase from Pseudomonas. J. Antibiot. 39, 606–608 (1986).
Akiyama, T. et al. Genistein, a specific inhibitor of tyrosine-specific protein kinases. J. Biol. Chem. 262, 5592–5595 (1987).
Xagorari, A. et al. Luteolin inhibits an endotoxin-stimulated phosphorylation cascade and Proinflammatory cytokine production in macrophages. J. Pharmacol. Exp. Ther. 296, 181–187 (2001).
Nakashima, S., Koike, T. & Nozawa, Y. Genistein, a protein tyrosine kinase inhibitor, inhibits thromboxane A2-mediated human platelet responses. Mol. Pharmacol. 39, 475–480 (1991).
Hong, H., Landauer, M. R., Foriska, M. A. & Ledney, G. D. Antibacterial activity of the soy isoflavone genistein. J. Basic Microbiol. 46, 329–335 (2006).
Wells, C. L., Jechorek, R. P., Kinneberg, K. M., Debol, S. M. & Erlandsen, S. L. The isoflavone genistein inhibits internalization of enteric bacteria by cultured Caco-2 and HT-29 enterocytes. J. Nutr. 129, 634–640 (1999).
Verdrengh, M., Collins, L. V., Bergin, P. & Tarkowski, A. Phytoestrogen genistein as an anti-staphylococcal agent. Microbes Infect. 6, 86–92 (2004).
Cooper, B. et al. Quantitative proteomic analysis of Staphylococcus aureus treated with Punicalagin, a natural antibiotic from pomegranate that disrupts iron homeostasis and induces SOS. Proteomics 18, e1700461 (2018).
Cooper, B. The detriment of Salicylic acid to the Pseudomonas Savastanoi pv. phaseolicola proteome. Mol. Plant. Microbe Interact. 35, 814–824 (2022).
Cooper, B. Disruptive effects of Resveratrol on a bacterial pathogen of beans. J. Proteome Res. 22, 204–214 (2023).
Cooper, B., Yang, R. & Campbell, K. B. Indole alkaloid production by the halo blight bacterium treated with the phytoalexin genistein. Phytopathology 114, 1196–1205 (2024).
Kosslak, R. M., Bookland, R., Barkei, J., Paaren, H. E. & Appelbaum, E. R. Induction of Bradyrhizobium Japonicum common Nod genes by isoflavones isolated from Glycine max. Proc. Natl. Acad. Sci. U S A. 84, 7428–7432 (1987).
Morris, P. F., Bone, E. & Tyler, B. M. Chemotropic and contact responses of phytophthora Sojae hyphae to soybean isoflavonoids and artificial substrates. Plant. Physiol. 117, 1171–1178 (1998).
Cooper, B., Campbell, K. B., Beard, H. S., Garrett, W. M. & Ferreira, M. E. The proteomics of resistance to halo blight in common bean. Mol. Plant. Microbe Interact. 33, 1161–1175 (2020).
Schymanski, E. L. et al. Identifying small molecules via high resolution mass spectrometry: communicating confidence. Environ. Sci. Technol. 48, 2097–2098 (2014).
Sumner, L. W. et al. Proposed minimum reporting standards for chemical analysis chemical analysis working group (CAWG) metabolomics standards initiative (MSI). Metabolomics: Official J. Metabolomic Soc. 3, 211–221 (2007).
Wheeler, A. W. Auxin-Like growth activity of phenylacetonitrile. Ann. Bot-London. 41, 867–872 (1977).
Perez, V. C., Zhao, H., Lin, M., Kim, J. & Occurrence function, and biosynthesis of the natural auxin phenylacetic acid (PAA) in plants. Plants (Basel) 12 (2023).
Djami-Tchatchou, A. T. et al. Dual role of auxin in regulating plant defense and bacterial virulence gene expression during Pseudomonas syringae PtoDC3000 pathogenesis. Mol. plant-microbe Interactions: MPMI. 33, 1059–1071 (2020).
Degrassi, G. et al. Plant growth-promoting Pseudomonas Putida WCS358 produces and secretes four Cyclic dipeptides: cross-talk with quorum sensing bacterial sensors. Curr. Microbiol. 45, 250–254 (2002).
Ortiz-Castro, R. et al. Transkingdom signaling based on bacterial cyclodipeptides with auxin activity in plants. Proc. Natl. Acad. Sci. U S A. 108, 7253–7258 (2011).
Hoch, H. C., Staples, R. C., Whitehead, B., Comeau, J. & Wolf, E. D. Signaling for growth orientation and cell differentiation by surface topography in Uromyces. Science 235, 1659–1662 (1987).
Cooper, B., Beard, H. S., Garrett, W. M. & Campbell, K. B. Benzothiadiazole conditions the bean proteome for immunity to bean rust. Mol. Plant. Microbe Interact. 33, 600–611 (2020).
Bernsdorff, F. et al. Pipecolic acid orchestrates plant systemic acquired resistance and defense priming via Salicylic acid-Dependent and -Independent pathways. Plant. Cell. 28, 102–129 (2016).
Klessig, D. F., Choi, H. W. & Dempsey, D. A. Systemic acquired resistance and Salicylic acid: Past, Present, and future. Mol. Plant. Microbe Interact. 31, 871–888 (2018).
Jirage, D. et al. Constitutive Salicylic acid-dependent signaling in cpr1 and cpr6 mutants requires PAD4. Plant. J. 26, 395–407 (2001).
Wang, W. et al. An Arabidopsis secondary metabolite directly targets expression of the bacterial type III secretion system to inhibit bacterial virulence. Cell. Host Microbe. 27, 601–613e607 (2020).
Ngou, B. P. M., Ding, P. & Jones, J. D. G. Thirty years of resistance: Zig-zag through the plant immune system. Plant. Cell. 34, 1447–1478 (2022).
Cooper, B., Beard, H. S., Yang, R., Garrett, W. M. & Campbell, K. B. Bacterial immobilization and toxicity induced by a bean plant immune system. J. Proteome Res. 20, 3664–3677 (2021).
O’Leary, B. M. et al. Early changes in Apoplast composition associated with defence and disease in interactions between phaseolus vulgaris and the halo blight pathogen Pseudomonas syringae Pv. phaseolicola. Plant. Cell. Environ. 39, 2172–2184 (2016).
Freeman, B. C. & Beattie, G. A. Bacterial growth restriction during host resistance to Pseudomonas syringae is associated with leaf water loss and localized cessation of vascular activity in Arabidopsis Thaliana. Mol. Plant. Microbe Interact. 22, 857–867 (2009).
Yamada, K., Saijo, Y., Nakagami, H. & Takano, Y. Regulation of sugar transporter activity for antibacterial defense in Arabidopsis. Science 354, 1427–1430 (2016).
Parniske, M., Ahlborn, B. & Werner, D. Isoflavonoid-inducible resistance to the phytoalexin Glyceollin in soybean rhizobia. J. Bacteriol. 173, 3432–3439 (1991).
Kapadia, C. et al. Pseudomonas aeruginosa inhibits quorum-sensing mechanisms of soft rot pathogen Lelliottia Amnigena RCE to regulate its virulence factors and biofilm formation. Front. Microbiol. 13, 977669 (2022).
Wu, L., Wu, H., Chen, L., Zhang, H. & Gao, X. Induction of systemic disease resistance in Nicotiana benthamiana by the cyclodipeptides cyclo (l-Pro-l-Pro) and cyclo (d-Pro-d-Pro). Mol. Plant Pathol. 18, 67–74 (2017).
Minen, R. I. et al. Characterization of the cyclic dipeptide cyclo(His-Pro) in arabidopsis. Plant Physiol. 198 (2025).
Cooper, B. & Yang, R. Genomic resources for Pseudomonas Savastanoi pv. phaseolicola races 5 and 8. Phytopathology 111, 893–895 (2021).
Cooper, B. & Yang, R. An assessment of acquirex and compound discoverer software 3.3 for non-targeted metabolomics. Sci. Rep. 14, 4841 (2024).
Lee, J. et al. Quantitative proteomic analysis of bean plants infected by a virulent and avirulent obligate rust fungus. Mol. Cell. Proteom. 8, 19–31 (2009).
Funding
This project was funded by USDA-ARS.
Author information
Authors and Affiliations
Contributions
B.C. conceived the study, cultured cells and prepared extracts, analyzed mass spectrometry data, and wrote the paper. R.Y. performed mass spectrometry and analyzed mass spectrometry data. K.B.C. maintained germplasm, grew bean plants and performed pathological assays.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Permits and permissions
Pseudomonas savastanoi pv. phaseolicola and Uromyces appendiculatus are authorized for use under controlled conditions by permit from USDA-APHIS. All infected plant materials were destroyed after experimentation. Relevant agency and national guidelines are outlined by USDA-ARS research project 8042-21220-261-000-D.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cooper, B., Yang, R. & Campbell, K.B. Untargeted metabolomics identifies a bacterial cyclic dipeptide that induces resistance to a rust fungus of beans. Sci Rep 16, 3606 (2026). https://doi.org/10.1038/s41598-025-33515-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-33515-4