Abstract
To establish a procedure for screening compounds that inhibit ligand–receptor binding, we employed a multidimensional virtual-system coupled molecular dynamics (mD-VcMD), a generalized ensemble method recently developed by our group. In this approach, each compound is initially placed far from the receptor. Both receptor and compound were fully flexible in explicit solvent during sampling. The mD-VcMD generated a free-energy landscape of the compound–receptor interactions, in which a probability of existence was assigned to each sampled conformation. We examined four compounds that bind to the papain-like protease (PLpro) of SARS-CoV-2. The resultant free-energy landscapes were funnel-like for all compounds. The probabilities assigned to the free-energy basins correlated well with the measured dissociation constants. Furthermore, structural clustering revealed two types of binding modes within the free-energy basin. The probabilities assigned to the binding modes correlated well with the measured enzyme inhibitory activity. These results suggest that the proposed procedure is effective for selecting candidate inhibitors among the examined compounds.
Similar content being viewed by others
Introduction
Since the 1990s, drug discovery research using computers has progressed rapidly. This trend coincides with significant improvements in computer performance. Simultaneously, the development of molecular dynamics simulations (MD) and the advancement of related application technologies have accelerated dramatically. For example, multicanonical MD-based dynamic docking simulations have elucidated the binding mechanism of a receptor and a ligand (i.e., a compound binding to the receptor). Moreover, the calculated binding constants are consistent with experimentally measured values1,2.
Progress has advanced rapidly in the development of classical in-silico screening methods by which, after various ligands are arranged on the surface of the receptor, the plausibility of these complexes is judged using an empirical evaluation function without performing the heavy calculations necessary for MD simulation3. Nevertheless, it remains difficult to calculate the multidimensional free-energy landscape in a wide conformational space that includes the binding and dissociation states of two flexible molecules in an explicit solvent. Efficient calculation and utilization of the free-energy landscape are expected to be useful for elucidating the binding process. Classical screening methods are often inadequate when molecules (receptors and/or ligands) are in high motion and when the solvent effects on the complex formation are not negligible.
Various enhanced sampling methods have been used widely for such efficient computational sampling4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20. In fact, the multi-dimensional virtual-system coupled molecular dynamics (mD-VcMD) method used for the present study is one such method7,8,19. As a concrete example, we investigated the free energy landscape of complex formation and the most stable complex structure between the membrane protein GPCR and a drug molecule: bosentan21. The simulation was initiated from an unbound conformation, with GPCR and bosentan spatially separated. As a result of an extensive structural search, the state with the lowest free energy (the most thermodynamically stable state) coincided with the experimentally.
determined complex structure. By analyzing the free energy landscape, we were able to elucidate the binding mechanism in the complex formation21. We obtained a similar result from an mD-VcMD simulation of binding between globular protein 14-3-3 and its associated intrinsically disordered protein (MLF1)9.
The mD-VcMD method has also been applied for drug discovery. It has been demonstrated by experimentation that a compound that inhibits the complex formation of the mammalian SIN3 transcription corepressor isoform B (mSin3B) and repressor element 1 silencing transcription factor (REST) suppresses germination22. mD-VcMD simulation4 of several compounds binding to mSin3B has reported that a compound that binds firmly to the cleft of mSin3B in the simulation has greater inhibitory activity, measured by experimentation, than those of other compounds22. This finding suggests that mD-VcMD is applicable to drug discovery.
In addition, the free-energy landscape obtained from the mD-VcMD method is useful not only for finding the most stable complex structure, but also for identifying the following quantities: the positions of the most stable and metastable complexes (e.g., encounter complexes) in the landscape, the paths of structural changes among these stable complexes, and the free-energy barriers among the complexes4,6,9,21. Consequently, the free-energy landscape provides comprehensive understanding of the complex formation process. When mD-VcMD is applied to different compounds, the obtained free-energy landscapes provide differences in complex structure formation among the compounds4,6,9,21.
We note that the mD-VcMD method was developed based on statistical mechanics and physical chemistry, as explained in Refs. 4, 6, 8, 9, 12, 15, 17, 18, 21, 43, and 44. This method assigns thermodynamic weights to all sampled snapshots, enabling the construction of a free energy landscape that includes both bound and dissociated states. By sampling diverse conformations, we account for entropic contributions from molecular motions and conformational flexibility, as well as enthalpic contributions from inter- and intra-molecular interactions. Accurate evaluation of complex stability requires consideration of both states and their statistical weights.
The study described herein targets SARS-CoV-2 PLpro, the papain-like protease of SARS-CoV-2 which has caused a global pandemic. This protease was discovered next to SARS-CoV-2 3CLpro. For this study, SARS-CoV-2 PLpro is designated simply as PLpro. Below is a biological explanation of PLpro. The papain-like protease, including PLpro, is an essential enzyme of coronaviruses that is required for the processing of viral polyproteins to form functional replicase complexes and to enable viral spread. Also, PLpro is involved in cleaving proteinaceous post-translational modifications to host proteins as an evasion mechanism against the host’s antiviral immune response. PLpro selectively cleaves ubiquitin-like interferon-stimulated gene 15 (ISG15) proteins. Crystallographic analysis of PLpro in complex with ISG15 shows unique interaction with the amino-terminal ubiquitin-like domain of ISG15 with high affinity and specificity. Furthermore, PLpro is involved in the cleavage of ISG15 from interferon responsive factor 3 (IRF3) during infection, attenuating the type I interferon response. Specifically, inhibition of PLpro with GRL0617 in infected cells suppresses virus-induced cytopathic effects, maintains the antiviral interferon pathway, and reduces viral replication23,24. Consequently, targeting PLpro is a promising therapeutic strategy that combines the two aspects of suppressing SARS-CoV-2 infection and promoting antiviral immunity, enabling more efficient drug discovery.
For drug discovery targeting the proteases 3CLpro and PLpro, experimentation has already been conducted with SARS-CoV-2, such as structural analyses of proteases25,26. Moreover, computational studies of proteases, including in silico screening using machine learning27, in silico screening using MD28, functional dynamics research covering homologous proteins29, inhibitor binding research on pathway dynamics30,31 and in silico screening that combines information spectral method ISM and MD calculations32 are being conducted actively. Understanding of these important topics is deepening.
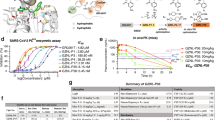
In the present study, we selected four complexes of PLpro with compounds from the ligand database designed by Shen et al.: GRL0617, XR8-24, XR8-23, and ZN-3-56 (Fig. 1a)33. By applying the mD-VcMD method to these compounds binding to PLpro, we went beyond simple screening to elucidate details of the inhibition mechanism. Furthermore, we investigated the correlation between the computed stabilities of the compound–PLpro complexes and the experimentally obtained dissociation constants and IC50 values33. We also applied clustering methods of two types34,35 to classify the numerous complex structures obtained from the simulations.
(a) Chemical structures of four compounds used for mD-VcMD. Broken line circles are “common regions” that are common among the four compounds. This region is used for setting the atom group \(\:{G}_{B}^{\left(RC2\right)}\) to define \(\:RC2\). (b) Two atom groups \(\:{G}_{A}^{\left(RC1\right)}\) (orange segment) and \(\:{G}_{B}^{\left(RC1\right)}\) (green segment) define \(\:RC1\); \(\:{G}_{A}^{\left(RC2\right)}\) and \(\:{G}_{B}^{\left(RC2\right)}\) define \(\:RC2\). Table S2 presents the atom groups explicitly. Supplementary Sect. 2 presents the procedure for setting reaction coordinates \(\:RC1\) and \(\:RC2\).
Materials and methods
To study binding of compounds to the receptor PLpro, we conducted the mD-VcMD simulation starting from an unbound conformation. Each system consists of PLpro and one of four compounds: GRL0617, XR8-24, XR8-23, or ZN-3-56 (Fig. 1a). We respectively designated the systems as PLpro–GRL0617, PLpro–XR8-24, PLpro–XR8-23, and PLpro–ZN-3-56.
The mD-VcMD simulation is a generalized ensemble method. For it, three variants have been developed to date: the original, subzone-based, and GA-guided mD-VcMD methods8. For this study, we used the subzone-based mD-VcMD, although we call this variant simply “mD-VcMD”. The mD-VcMD enhances the conformational sampling in a space (RC space) constructed by multiple reaction coordinates (RCs), which are set in advance. The mD-VcMD simulation generates an ensemble of conformations (i.e., snapshots) in the RC space. After the simulation, a statistical weight (i.e., a thermodynamically equilibrated weight at the simulation temperature) is assigned to each snapshot stored in the ensemble8. Therefore, this ensemble is regarded as a canonical ensemble (i.e., thermodynamically equilibrated ensemble), from which various physical quantities at equilibrium are computed.
Below, after first explaining the simulation systems, we introduce two RCs. Therefore, mD-VcMD is “2D”-VcMD in this study. Third, the mD-VcMD procedure is explained briefly. Fourth, we present a method to compute a physical quantity from the resultant ensemble. Subsequently, the principal component analysis (PCA) method is explained, by which the receptor -ligand binding binding affinity is computed. Finally, we present a procedure to compute structural clusters from the ensemble.
Molecular systems and initial conformations of the simulation
The simulation systems were generated as explained hereinafter. First, the crystallographic structure of the PLpro–GRL0617 complex was taken from PDB (PDB ID: 7CJM; 3.2 \(\:\hbox{\AA}\:\) resolution)36. Then, we removed the compound GRL0617 from the structure. The 111-th amino-acid residue of PLpro in the crystallographic structure is serine, whereas it is cysteine in the wild-type PLpro. Therefore, we back-mutated it to cysteine. Acetyl and N-methyl groups were put respectively to the N-terminal and C-termini of PLpro.
PLpro was put at the center of a periodic boundary box filled by water molecules: Box size = 100 \(\:\hbox{\AA}\:\) × 100 \(\:\hbox{\AA}\:\) × 100 \(\:\hbox{\AA}\:\). One can recall that GRL0617 was removed from the crystallographic structure described above. Then, we put one of the four compounds (GRL0617, XR8-24, XR8-23, and ZN-3-56) at a position distant from the ligand-binding site of PLpro. In this stage, therefore, the system comprises PLpro, one of four compounds, and water molecules. When putting PLpro and the compound in the box, water molecules overlapping to PLpro or the compound were removed. Furthermore, \(\:{Na}^{+}\) and \(\:{Cl}^{-}\) ions were put randomly in the solvent region. Water molecules overlapping the introduced ions were removed. The quantities of ions were set so that the NaCl concentration was 150 mM and so that the net charge of the system was neutralized. Table S1 of Supporting Information (SI) presents the quantities of constituent atoms of the generated systems. Before initiating the mD-VcMD calculations, we examined various box sizes. Larger boxes require longer computation times, and if the ligand and receptor are too far apart, informative snapshots cannot be obtained. Notably, two spatially distant molecules contribute to the system’s entropy, but such entropic contributions cancel out when comparing different systems. Conversely, smaller boxes reduce computation time but fail to capture structures in which the ligand and receptor are fully dissociated. To obtain the free-energy landscape of complex formation, it is essential to sufficiently incorporate dissociated states into the ensemble. Ultimately, we found that a 100 × 100 × 100 Å box allows complete dissociation of the ligand from the receptor in all systems, and thus we adopted this box size.
The conformation of each system generated above was energy-minimized. An NVT simulation (300 K temperature) was done to relax the structural frustrations originating from the PDB structure. Then, an NPT simulation (1 atm pressure; 300 K temperature) was conducted to relax the box size at the pressure. Table S1 of SI presents the resultant box size for each system. The resultant conformations are portrayed in Figure S1 of SI, which indicates that the compound and PLpro are mutually separated in solvent for all systems. We used these unbound conformations for the initial conformations of the mD-VcMD simulations.
The four systems generated above did not involve a zinc (Zn) ion. By contrast, the crystallographic complex structures for PLpro–GRL0617 (PDB ID: 7CJM; Figures S2a of SI) and PLpro–XR8-24 (PDB ID: 7LBS; Figures S2b of SI) respectively contain three and one Zn ions. No crystallographic structure is reported for the other two systems, PLpro–XR8-23 and PLpro–ZN-3-56, in PDB (Aug. 2023). In the structure of 7CJM (Figure S2a) the distances from the N2 atom of GRL0617 to the three Zn ions are 22.1 \(\:\hbox{\AA}\:\), 30.9 \(\:\hbox{\AA}\:\), and 39.4 \(\:\hbox{\AA}\:\). Similarly, the distance from the N03 atom of XR8-24 to the Zn ion is 30.9 \(\:\hbox{\AA}\:\) in 7LBS (Figure S2b). Figure S2c of SI presents a ligand-free (apo-form) of PLpro (PDB ID: 6XG3), where the Zn ion is also distant from the ligand-binding site of PLpro. Furthermore, we searched other crystallographic structures containing PLpro in PDB, which revealed that most Zn ions are far from the ligand-binding site, as shown in Supplementary Sect. 1 of SI. Therefore, it is likely that the direct effect of Zn ions on the compound-binding is small. For that reason, no Zn ion was involved in the systems.
The force field parameters for PLpro, and the ions (\(\:{Na}^{+}\) and \(\:{Cl}^{-}\)) were obtained respectively from AMBER99SB-ILDN37 and the Joung–Cheatham parameters38. The TIP3P potential39 was used for water. Force field parameters for the four compounds were modeled by us. First, the atomic partial charges of the compounds were calculated quantum-chemically using Gaussian1640 at the HF/6-31G* level, with subsequent RESP fitting41. Then, the obtained atomic partial charges were incorporated into a general amber force field 2 (GAFF2)42, which was designed to be compatible with conventional AMBER force-fields.
Two RCs introduced to control system motion
The mD-VcMD simulation enhances the conformational sampling in a space constructed by multiple reaction coordinates (RCs)8. The RCs are usually set before the simulation. The \(\:i\)-th RC, denoted herein as \(\:RCi\), was defined by the inter-centroid distance between two atom groups introduced into the system: \(\:{G}_{A}^{\left(RCi\right)}\)and \(\:{G}_{B}^{\left(RCi\right)}\)(Supplementary Sect. 2 of SI, Figure S3). As explained earlier, the space constructed by the multiple RCs is designated as an RC space. Because we introduced two RCs in this study, the RC space is two-dimensional.
\(\:\text{F}\text{i}\text{g}\text{u}\text{r}\text{e}\:1\text{b}\:\text{s}\text{h}\text{o}\text{w}\text{s}\:RC1\) and \(\:RC2\) for the PLpro–GRL0617 system. \(\:RC1\) is the inter-centroid distance between the atom groups \(\:{G}_{A}^{\left(RC1\right)}\)and \(\:{G}_{B}^{\left(RC1\right)}\), both of which are set in PLpro. The atom group \(\:{G}_{A}^{\left(RC1\right)}\) is the orange segment in the figure, which is a loop region (residues G266-N267-Y268-Q269-C270-G271). Also,\(\:\:{G}_{A}^{\left(RC1\right)}\) is the green segment, which is a part of a helix (residues V165-R166-E167-T168). The side-chains of the segments are eliminated from the definition of the atom groups. Also, \(\:{G}_{A}^{\left(RC1\right)}\)and \(\:{G}_{B}^{\left(RC1\right)}\)form the cleft of the ligand–binding site of PLpro. \(\:RC2\) is the inter-centroid distance separating \(\:{G}_{A}^{\left(RC2\right)}\)from \(\:{G}_{B}^{\left(RC2\right)}\). Atom group \(\:{G}_{A}^{\left(RC2\right)}\) is set to the combined region of \(\:{G}_{A}^{\left(RC1\right)}\)and \(\:{G}_{B}^{\left(RC1\right)}\) (i.e., \(\:{G}_{A}^{\left(RC2\right)}={G}_{A}^{\left(RC1\right)}+{G}_{B}^{\left(RC1\right)}\)). Atom groups \(\:{G}_{A}^{\left(RC1\right)}\), \(\:{G}_{B}^{\left(RC1\right)}\), and \(\:{G}_{A}^{\left(RC2\right)}\) are the same for all systems. Atom group \(\:{G}_{B}^{\left(RC2\right)}\) is set to a region of the compound GRL0617. One might consider that \(\:{G}_{B}^{\left(RC2\right)}\) depends on the system because \(\:{G}_{B}^{\left(RC2\right)}\) is set to the different compounds. However, the compounds share a common region (Fig. 1a). Then we set \(\:{G}_{B}^{\left(RC2\right)}\) to the common region. Consequently, \(\:{G}_{B}^{\left(RC2\right)}\) is the same among the four systems. Table S2 of SI presents details for the atom groups. The benefit of using the same atom groups for all systems is that the free-energy landscape can be investigated from universal points of view.
Figure 1b shows that \(\:RC1\) quantifies the cleft width of the ligand-binding site of PLpro and that \(\:RC2\) quantifies the distance from the ligand-binding site to the compound. Therefore, \(\:RC1\) controls opening (or closing) of the cleft, and \(\:RC2\) controls the compound approach to (or departure from) the binding site. We have used similar RCs to study ligand-binding to the binding site of a receptor4,6,21,43,44.
During an mD-VcMD simulation, the conformation is confined to the range \(\:[RC{i}_{min},RC{i}_{max}]\) along each of the \(\:RCi\) axes, where \(\:RC{i}_{min}\) and \(\:RC{i}_{max}\) respectively represent the minimum and maximum boundaries for \(\:RCi\). To ensure that mD-VcMD samples the 2D-RC space widely, we set the 2D-RC range so that the conformation can adopt various structures. So that \(\:RCi\) does not protrude from the range, the confinement is achieved by application of a restoring force to the conformation12 only when \(\:RCi\) is flying outside the range. For the present study, these 2D ranges involve both bound and unbound conformations. Table S3 of SI presents the actual values for the ranges. The smallest RCs are set as \(\:RC{i}_{min}=0.0\:\hbox{\AA}\:\) (\(\:i=1\) and \(\:2\)), which is the smallest value in principle because a negative distance does not exist in the real space. The value of \(\:25.4890\:\hbox{\AA}\:\) for \(\:RC{1}_{max}\) is sufficiently large to open the binding cleft. The value of 49.2801 for \(\:RC{2}_{max}\) is also sufficiently large to allow the compound to move in the unbound state. We show in Results and Discussion that various complex and unbound conformations are sampled during the simulation.
Division of RC axes into small zones and execution of mD-VcMD
With mD-VcMD, each RC axis is divided into small zones, which are set in the range of \(\:[RC{i}_{min},RC{i}_{max}]\). The conformation moves in a zone (intra-zone movement) for a time interval and transitions to another zone (inter-zone transition) occasionally, as explained later. The numbers of zones set along the \(\:RCi\) axis are set as \(\:{n}_{vs}^{\left(RCi\right)}\): \(\:{n}_{vs}^{\left(RC1\right)}=34\) and \(\:{n}_{vs}^{\left(RC2\right)}=48\) (Table S3 of SI). The lower and upper boundaries for the \(\:k\)-th zone of the axis \(\:RCi\) are denoted respectively as \(\:RC{i}_{low}\left(k\right)\) and \(\:RC{i}_{up}\left(k\right)\). Their actual values are given in Table S4 of SI.
The simulation is assumed to be at time \(\:t\) at present. The conformation at this time is assumed as in the \(\:l\)-th and \(\:m\)-th zones, respectively, along the \(\:RC1\) the \(\:RC2\) axes. In other words, the conformation is in the 2D zone \(\:(l,m)\) at \(\:t\). We designated this 2D zone as the “current zone”. In mD-VcMD, the conformation is confined in the current zone for a short interval \(\:\varDelta\:{\tau\:}_{trns}\) where the restoring force12 towards the current zone is applied to the system only when the conformation is flying to outside the current zone. Alternatively, the conformation moves without the restoring force when it is inside the current zone. At a time-passage of \(\:\varDelta\:{\tau\:}_{trns}\) (time \(\:=\:t+\varDelta\:{\tau\:}_{trns}\)), the conformation transitions from \(\:(l,m)\) to another zone \(\:(l{\prime\:},m{\prime\:})\) using an inter-zone transition probability. The protocol to execute the transition and a method to set the transition probability are explained in Supplementary Sect. 3 and Figure S4 of SI. The zone \(\:(l{\prime\:},m{\prime\:})\) is one of zones surrounding the current zone \(\:(l,m)\) or the current zone itself (self-transition). The current zone is updated from \(\:(l,m)\) to\(\:(l{\prime\:},m{\prime\:})\). We set \(\:\varDelta\:{\tau\:}_{trns}=2\:ps\), which corresponds to \(\:1\times\:{10}^{3}\) simulation steps in this study (simulation time-step \(\:=2\:fs\)).
Repeating this procedure, the conformation moves among many 2D zones during the simulation. However, because the RC space \(\:[RC{i}_{min},RC{i}_{max}]\) is too large to be sampled in a single run, the mD-VcMD simulation is conducted iteratively. When the \(\:J\)-th iteration has finished, we calculate a probability distribution function \(\:{Q}_{cano}^{\left[J\right]}(l,m)\). The probability of existence of the system assigned to a 2D RC zone \(\:(l,m)\) is calculated using simulation data from the fourth to \(\:J\)-th iterations8. As explained later, we conducted 20 iterations for each system in this study. Data from the first to third iterations were discarded to eliminate an error resulted from a structural distortion in the initial conformation of simulation, although we presume that such error was small in this study because the simulation starts from the unbound conformation (Figure S1).
When convergence of \(\:{Q}_{cano}^{\left[J\right]}\) has been realized in the 2D-RC space, \(\:{Q}_{cano}^{\left[J\right]}(l,m)\) is regarded as the system’s canonical distribution (i.e., a thermally equilibrated distribution at the 300 K simulation temperature). Then, we simply denote \(\:{Q}_{cano}^{\left[J\right]}(l,m)\) as \(\:{Q}_{cano}(l,m)\) and assign a thermodynamic weight at \(\:T\) to each snapshot8. The convergence check is explained in Supplementary Sect. 4. If the convergence has not yet been obtained in iteration \(\:J\), then the inter-zone transition probability is calculated; also, iteration \(\:J+1\) is started.
To increase the sampling efficiency further, we conducted \(\:{N}_{run}\) (= 1,000) runs of the mD-VcMD simulation in parallel in each iteration, where the runs started from different conformations. Conformations from the \(\:{N}_{run}\) runs construct a canonical ensemble, in theory20,45. If assuming that the length of each run is \(\:\varDelta\:{\tau\:}_{run}\), and that the runs are initiated from various conformations, then the \(\:{N}_{run}\) trajectories cover a conformational space more widely than a long single run of length \(\:{N}_{run}\times\:\varDelta\:{\tau\:}_{run}\) does20. Exceptionally, the \(\:{N}_{run}\) runs for the first iteration start from the single conformation (Figure S1). Then, the sampled region increases with progress of the simulation to the second and third iterations. The \(\:{N}_{run}\) initial conformations for the fourth iteration are the last conformations of the third iteration. When the \(\:J\)-th iteration has finished (\(\:J\ge\:4\)), we have snapshots sampled from the fourth to \(\:J\)-th iterations. Then, the \(\:{N}_{run}\) initial conformations for the \(\:(J+1)\)-th iteration are selected from those snapshots so that they are distributed as evenly as possible in the 2D-RC space.
We set the simulation length \(\:\varDelta\:{\tau\:}_{run}\) of each of the \(\:{N}_{run}\) runs to 2.0 ns (\(\:1\times\:{10}^{6}\) steps) for each of the four systems. Consequently, the total length for an iteration was \(\:2.0\:{\upmu\:}\text{s}\) (\(\:2.0\:\text{n}\text{s}\times\:\text{1,000}\:\text{r}\text{u}\text{n}\text{s}\)). As we described above, after doing 20 iterations, we used the trajectories from the fourth to the twentieth iterations for analyses, for which the simulation length was \(\:2.0\:\text{n}\text{s}\times\:\text{1,000}\times\:17\:\text{i}\text{t}\text{e}\text{r}\text{a}\text{t}\text{i}\text{o}\text{n}\text{s}=34.0\:{\upmu\:}\text{s}\). We saved a snapshot at every \(\:5\times\:1{0}^{4}\) steps (\(\:0.1\:\text{n}\text{s}\)) in each run. The number of snapshots saved for the analyses was therefore \(\:3.4\times\:1{0}^{5}\) (=\(\:34\:{\upmu\:}\text{s}\:/\:0.1\:\text{n}\text{s}\)) for each system. The integrated trajectories produce a canonical ensemble when the \(\:{N}_{run}\) simulations have been performed repeatedly, starting from different initial conformations. These \(\:{N}_{run}\) runs are executed in parallel20,45. Therefore, the total trajectory length does not represent a continuous 34.0 µs simulation, but rather an ensemble-based sampling approach20,45.
The mD-VcMD algorithm was first implemented to an MD simulation program omegagene/myPresto19 and next to GROMACS46 with the PLUMED plug-in47. The present simulation was done using the GROMACS version. The simulation conditions were the following: LINCS48 to constrain the covalent-bond lengths related to hydrogen atoms, the Nosé–Hoover thermostat49,50 to control the simulation temperature, and the zero-dipole summation method51,52 to compute the long-range electrostatic interactions. No structural restraint was applied to the system: All molecules were flexible in the simulation except for the hydrogen-related covalent bonds constrained by LINCS, as explained above. We set the time-step of simulation to 2 fs (\(\:\varDelta\:t=2\) fs) and the simulation temperature to \(\:300\:\text{K}\). All simulations were conducted using the TSUBAME4.0 supercomputer at the Institute of Science Tokyo using GP-GPU. Other analyses were done at Ritsumeikan University and the Research Center for Computational Science, Okazaki, Japan. The force field parameters used for the present study were explained earlier herein.
One might infer that confinement of the system’s motions in a zone for an interval of \(\:\varDelta\:{\tau\:}_{trns}\) might affect the resultant conformational ensemble. This criticism is valid for a conventional MD simulation that uses an artificial potential. For mD-VcMD, however, the bias effect is removed completely from the resultant ensemble by a reweighting technique8,21. The resultant thermodynamic weight assigned to a snapshot is a quantity without the conformational confinement in zones. In fact, mD-VcMD differs from umbrella sampling53, whereas the conformation is confined in a 2D-RC zone for the short time interval \(\:\varDelta\:{\tau\:}_{trns}\), as explained in our previous study44.
Some physical quantities to study binding mechanisms
One can calculate the thermodynamic average (expected value) of a physical quantity \(\:{\Omega\:}\) at the simulation temperature \(\:T\) (\(\:300\:\text{K}\)) using the canonical ensemble of snapshots:
where \(\:{w}_{i}\) denotes the statistical weight (thermodynamic weight) assigned to snapshot \(\:i\), \(\:{{\Omega\:}}_{i}\) represents the value of \(\:{\Omega\:}\) for the snapshot, and \(\:{N}_{snp}\) (=\(\:3.4\times\:1{0}^{5}\)) stands for the number of snapshots stored in the ensemble of a single system. Here, \(\:{\Omega\:}\) can be either a scalar or vector. If one aims to restrict the ensemble so that the snapshots satisfy a condition, then Eq. 1 is redefined as
where
The role of \(\:{D}_{i}\) is simple: If a snapshot does not satisfy the condition, then the snapshot is eliminated from the analysis. The standard deviation of \(\:{\Omega\:}\) is defined as
Principal component analysis (PCA)
Because a molecular conformation is specified by many atomic coordinates, the snapshot conformation is represented by a point in a high-dimensional space, and many snapshots obtained from a simulation distribute in the high-dimensional space. PCA has been used to visualize the distribution by projecting the points to a low- dimensional space54,55,56,57. We applied PCA to the ensembles of the snapshots according to our earlier study57, the methodological details of which are explained in Supplementary Sect. 5. Below we briefly present an outline of the PCA done for this study.
The variance–covariance matrix \(\:A\) is calculated from atom-pair distances between the whole Cα atoms of PLpro (316 Cα atoms) and the heavy atoms (eight atoms) in a core region of the compound (Figure S5 of SI). The element \(\:{A}_{i,j}\) of \(\:A\) is defined in Equation S7 of SI. Adoption of all Cα atoms is done to detect contacts between the compound and the entire surface of PLpro. Alternatively, selection of only the core region is done to compare the PCA results among the four systems from the same perspective: the atoms taken for PCA are the same when the core region is used. The core region is a ring part of the common region of the compounds (Fig. 1a). Both the crystallographic structures for the PLpro–GRL0617 complex (PDB ID: 7CJM) and the PLpro–XR8-24 complex (PDB ID: 7LBS) demonstrate that this ring binds to the binding site of PLpro firmly with the same orientation. This point is explained further in the last paragraph of Supplementary Sect. 5.
We denote two eigenvectors with the largest and second largest eigenvalues respectively as \(\:PC1\) and \(\:PC2\). The contribution ratios \(\:{CR}_{1}\) and \(\:{CR}_{2}\) (Equation S11) to the standard deviation of the entire distribution were \(\:{CR}_{1}=0.647\)and \(\:{CR}_{2}=0.190\) and \(\:CR\left(1;2\right)=0.837\) (Equation S12). This large \(\:CR\left(1;2\right)\) indicates that the 2D-PC space constructed by \(\:PC1\) and \(\:PC2\) is a conformational space that can capture important structural features of the system’s distribution. Then, we computed a distribution function \(\:\rho\:(PC1,PC2)\) for each system in the 2D-PC space. Because \(\:\rho\:(PC1,PC2)\) is computed from the ensemble at 300 K, this function is the thermally equilibrated distribution at 300 K. A free-energy landscape (potential of mean force) at 300 K in 2D-PC space is defined as
where \(\:R\) stands for the gas constant, and where the unit of \(\:PMF\) is kilocalories per mole. Because we intend to compare \(\:PMF\) with an equal weight among the four systems, \(\:\rho\:(PC1,PC2)\) is normalized for each system as \(\:\iint\:\rho\:(PC1,PC2)dPC1\times\:dPC2=1.0\).
Clustering of complex conformations
PCA is a convenient method to view the conformational distribution (the free-energy landscape; \(\:PMF\left(PC1,PC2\right)\)) and to identify a region occupied by thermodynamically stable conformations (i.e., the region with a low \(\:PMF\)) in the conformational space. However, it does not mean that PCA can pick the compound’s conformations which have a particular structural feature merely by viewing the free-energy landscape. Therefore, structural clustering is important to classify the sampled conformations and to assign a binding mode to each of the resultant clusters.
Then, to elucidate the structural features of the sampled conformations in the entire conformational ensemble, we used the t-distributed stochastic neighbor embedding (t-SNE) method34 and the density-based spatial clustering of applications with noise (DBSCAN) method35 sequentially. We explain those procedures below.
A quantity that characterizes the compound–receptor complex structure is the pattern of atomic contacts between the two molecules. Then we quantified the heavy-atomic contacts between PLpro and the core region of each compound. As described earlier, the core region was also used for PCA. Discarding of the non-core region is done to eliminate unimportant inter-molecular contacts from the analysis. As discussed in the preceding subsection, the crystallographic structures for the PLpro–GRL0617 and the PLpro–XR8-24 complexes demonstrate that the core region binds firmly to the ligand-binding site of PLpro with the same orientation.
We represented the atomic distance between the \(\:i\)-th C\(\:\alpha\:\) atom of PLpro and the \(\:j\)-th heavy-atom of the compound’s core region of snapshot \(\:k\) as \(\:{l}_{i,j}^{\left(k\right)}\), where the ordering of \(\:i\) and \(\:j\) is arbitrary. Then, we digitized \(\:{l}_{i,j}^{\left(k\right)}\) as
where \(\:{l}_{lim}^{\left(cnt\right)}\) is a contact-limit parameter. Atoms \(\:i\) and \(\:j\) are contacting when \(\:{c}_{i,j}^{\left(k\right)}=1\). Also, \(\:{l}_{lim}^{\left(cnt\right)}\) is set as \(\:8\:\hbox{\AA}\:\) for this study. To define the contact between two heavy atoms more rigorously, the criteria \(\:{l}_{lim}^{\left(cnt\right)}=7\:\hbox{\AA}\:\) might be better than \(\:8\:\hbox{\AA}\:\): Assuming that the radius of a heavy atom and the diameter of a water molecule are, respectively, approximately \(\:2\:\hbox{\AA}\:\) and \(\:3\:\hbox{\AA}\:\), the water molecule only slightly passes through the space between two heavy atoms when the distance between the two heavy atoms is \(\:7\:\hbox{\AA}\:\) (=\(\:2\:\hbox{\AA}\:+2\:\hbox{\AA}\:+3\:\hbox{\AA}\:\)). However, to conserve computer memory resources, we used only a \(\:C\alpha\:\) atom as a representative atom for each amino-acid residue. Therefore, we increased \(\:{l}_{lim}^{\left(cnt\right)}\) from \(\:7\:\hbox{\AA}\:\) to \(\:8\:\hbox{\AA}\:\) in this study.
PLpro and the core region of a compound respectively contain 316 residues and 8 heavy atoms. Therefore, a set of the intermolecular contacts for the \(\:k\)-th snapshot is expressed as a vector of \(\:\text{2,528}\:(=316\times\:8)\) elements:
One can imagine that the \(\:{N}_{snp}\) snapshots \(\:\{{\varvec{c}}_{1},\dots\:,{\varvec{c}}_{{N}_{snp}}\)} for each system are distributed in the \(\:\text{2,528}\)-dimensional space, where \(\:{\varvec{c}}_{k}\) denotes the position of the \(\:k\)-th snapshot in the space. As stated earlier, \(\:{N}_{snp}\) is the number of snapshots saved for each system. We designate this space as a “contact space”. Because all \(\:C\alpha\:\) atoms of PLpro are involved in Eq. 7, not only the intermolecular native contacts but also non-native ones are counted in this equation. Next, we reduced the dimensionality of the contact space from \(\:\text{2,528}\) to \(\:2\) using t-SNE. Accordingly, the \(\:{N}_{snp}\) data points were translated as
where \(\:{\varvec{c}}_{k}^{\left(2D\right)}\) represents the position of the \(\:k\)-th snapshot in the 2D contact space. Clustering of the 2D data points was done using DBSCAN.
We denote the \(\:m\)-th cluster for the system \(\:\alpha\:\) (\(\:\alpha\:=\) PLpro–GRL0617, PLpro–XR8-24, PLpro–XR8-23 or PLpro–ZN-3-56) as \(\:{Cls}_{m}^{\left(\alpha\:\right)}\). Then, the probability assigned to \(\:{Cls}_{m}^{\left(\alpha\:\right)}\) for each system is computed as
where \(\:{D}_{i}^{\left({Cls}_{m}^{\left(\alpha\:\right)}\right)}\) is a specific case of \(\:{D}_{i}\) (Eq. 3) defined as
Results and discussion
Conformational distribution in the 2D-RC space
We performed 20 iterations of the mD-VcMD simulation for each of the four systems starting from the unbound conformations (Figure S1). Figure 2 presents the probability distribution \(\:{Q}_{cano}(l,m)\) computed from iterations 4–20 with discarding of iterations 1–3 as explained in the Materials and Methods section. Apparently, a small-\(\:RC2\) region was characterized by a large probability for all systems, which means that the compound binding to the ligand-binding site of PLpro was more stable than those in the other regions. The eight sites of \(\:{c}_{1},\dots\:,{c}_{8}\) are marked in Fig. 2. Later, the conformations at these eight sites are shown. First, however, we assess the simulation data quality.
Spatial patterns of the distribution \(\:{Q}_{cano}(l,m)\) for (a) PLpro–GRL0617, (b) PLpro–XR8-24, (c) PLpro–XR8-23, and (d) PLpro–ZN-3-56. The X-axis represents zone index \(\:l\) for \(\:RC1\). The Y-axis shows the zone index \(\:m\) for \(\:RC2\). The main values of panel (a) are zone indices \(\:l\) and \(\:m\). Values in parentheses \(\:RC1\) and \(\:RC2\) respectively correspond to \(\:l\) and \(\:m\). Correspondence between zone index \(\:k\) (\(\:k=l\:\text{o}\text{r}\:m\)) and \(\:RCi\) (\(\:i=1\:\text{o}\text{r}\:2\)) is given as \(\:RCi=\left[RC{i}_{low}\left(k\right)+RC{i}_{up}\left(k\right)\right]/2\). The RC values for the other panels are omitted. Table S4 of SI presents the values of \(\:RC{i}_{low}\left(k\right)\) and \(\:RC{i}_{up}\left(k\right)\). The value of \(\:\text{l}\text{o}{\text{g}}_{10}\left[{Q}_{cano}\right]\) at 2D zone \(\:(l,m)\) is shown by the color scale.
Convergence of simulation data was checked using the correlation coefficient \(\:{C}_{cor}^{[J,20]}\) of the potential of mean force \(\:{F}_{J}\left(l,m\right)\) (Equation S5) between the \(\:J\)-th and 20-th iterations (Equation S6.1 of SI). Figure S6 of SI demonstrates that \(\:{F}_{J}\left(l,m\right)\) approached \(\:{F}_{20}\left(l,m\right)\) quickly in an iteration range of \(\:1\le\:J\le\:10\) and slowly in \(\:11<J\ge\:16\). Finally, the coefficient almost reached a plateau for \(\:J\ge\:17\) in all systems. Therefore, we judged that the simulation provided an equilibrated ensemble approximately. As described in Materials and Methods, we discarded iterations 1–3 from the analyses. Figure S6 shows that the correlation coefficient increased quickly in the three iterations. Therefore, the \(\:{N}_{run}\) (= 1,000) trajectories spread quickly from the single conformation (Figure S1) in the 2D-RC space during the three iterations.
Although the convergence was obtained well in the iterations, Fig. 2 depicts that the data scattering along the \(\:RC1\) direction (i.e., the degree of the binding-site cleft opening) differs among the systems. To check this difference of distribution quantitatively, we calculated the fluctuation of \(\:RC1\) (i.e., \(\:S{D}_{\text{R}\text{C}1}\)) at various values of \(\:RC2\). Details for \(\:S{D}_{\text{R}\text{C}1}\) are presented in Supplementary Sect. 6 of SI. Figure S7 of SI shows that \(\:S{D}_{\text{R}\text{C}1}\) values were similar among the four systems in the large \(\:RC2\) range. In fact, snapshots with a very small thermodynamic weight \(\:{w}_{i}\) contribute less to \(\:S{D}_{\text{R}\text{C}1}\) (Equation S13). Figure 2 is depicted with a log scale of \(\:{Q}_{cano}\).
As explained earlier, \(\:RC1\) was introduced to enhance the motion of the binding-site cleft of PLpro (Fig. 1b). In fact, Figure S7 shows that \(\:RC1\) moved greatly. By contrast, Figure S7 shows that \(\:{SD}_{\text{R}\text{C}1}\) in a small-\(\:RC2\) range has some variety, which is natural because direct interaction between PLpro and the component differs among the systems when the compound and PLpro mutually contact (i.e., when \(\:RC1\) is small).
Figure 3 displays the conformations of PLpro–GRL0617 picked from the sites \(\:{c}_{1},\dots\:,{c}_{8}\) marked in Fig. 2a. The conformations \(\:{c}_{1}\) and \(\:{c}_{2}\), which are from the lowest free-energy region of the PLpro–GRL0617 system, were complex structures (Fig. 3\(\:{\text{c}}_{1}\) and \(\:{\text{c}}_{2}\)), where GRL0617 bound to the ligand-binding site of PLpro. Conformations \(\:{c}_{3}\) and \(\:{c}_{4}\) were those for which the compound bound to a surface of PLpro deviating from the ligand-binding site (Fig. 3\(\:{\text{c}}_{3}\) and \(\:{\text{c}}_{4}\)). Conformations \(\:{c}_{5}\) and \(\:{c}_{6}\) showed that the compound contacted to various surfaces of PLpro (Fig. 3\(\:{\text{c}}_{5}\) is an example of the compound binding to PLpro) or that it was rarely dissociated from PLpro as in Fig. 3\(\:{\text{c}}_{6}\). Conformations in a large-RC2 region (\(\:{c}_{7}\) and \(\:{c}_{8}\) in Fig. 2a) were unbound conformations (Fig. 3\(\:{\text{c}}_{7}\) and \(\:{\text{c}}_{8}\)). This positioning of the compound in the large-RC2 region is natural because the upper bound of \(\:RC2\) (\(\:RC{2}_{max}=49.2801\:\hbox{\AA}\:\); Table S3) was set so that the compound could move freely without touching PLpro. A similar situation was found in the other systems, as shown in Figures S8–10 of SI.
Conformations of the PLpro–GRL0617 system selected from sites \(\:{c}_{1},\dots\:,{c}_{8}\) defined in (Fig. 2a). Panel names of this figure correspond to site names \(\:{c}_{1},\dots\:,{c}_{8}\) of (Fig. 2a). Purple molecules are GRL0617. Orange and green segments are atom groups used to define reaction coordinate \(\:RC1\) (Fig. 1b): Residues 266–271 and 165–168. The brown broken line represents the position of the ligand-binding site of PLpro. Molecules in panels are drawn from different directions to clarify the positions of the ligand-binding site and compound.
It is noteworthy that Fig. 2 depicts a high free energy (i.e., a low probability) assigned to unbound conformations or non-native complex structures. Therefore, we infer that the free-energy landscape of the four systems is funnel-like in 2D-RC space.
PCA
PCA was applied to the resultant conformational ensemble of the four systems, for which details are given in Supplementary Sect. 5 of SI. Figure 4 presents \(\:PMF\left(PC1,PC2\right)\) at 300 K in the 2D-PC space (Eq. 5). The spatial patterns of \(\:PMF\) were similar among the four systems: a deep free-energy basin existed at nearly the same position in the space; \(\:PMF\) increased with departure from the basin. In other words, the free-energy landscape is funnel like. As explained in Materials And Methods, the contribution \(\:CR\left(1;2\right)\) from \(\:PC1\) and \(\:PC2\) (Equation S12) to the entire conformational variety was large: \(\:CR\left(1;2\right)=0.837\). Therefore, this funnel-like feature is likely to be the main feature in the full-dimensional space.
Spatial pattern of \(\:PMF\left(PC1,PC2\right)\) in 2D-PC space for (a) PLpro–GRL0617, (b) PLpro–XR8-24, (c) PLpro–XR8-23, and (d) PLpro–ZN-3-56.
Figure 5 portrays conformations of the four systems picked randomly from the free-energy basin (i.e., region of \(\:PMF\left(PC1,PC2\right)\le\:1.0\:\text{k}\text{c}\text{a}\text{l}/\text{m}\text{o}\text{l}\)) of Fig. 4. All the conformations were complex structures in that the compound bound to the ligand-binding site of PLpro, whereas the conformation of the compound showed some variation in the binding site. Because these conformations were characterized by the low \(\:PMF\) value, it is likely that they are related to the dissociation constant \(\:{K}_{d}\).
Conformations picked randomly from free-energy basin (i.e., region of with \(\:PMF\left(PC1,PC2\right)\le\:1.0\) kcal / mol of Fig. 4 for (a) PLpro–GRL0617, (b) PLpro–XR8-24, (c) PLpro–XR8-23, and (d) PLpro–ZN-3-56.
However, the existence of a low \(\:PMF\) region does not necessarily mean that the complex is thermodynamically stable. If the low \(\:PMF\) region is narrow in the PC space, then the complex might be unstable. Therefore, we computed the probability assigned to the low \(\:PMF\) region as
Therein, \(\:\epsilon\:\) is a threshold specifying the low \(\:PMF\) region. We examined not only \(\:1.0\:\text{k}\text{c}\text{a}\text{l}/\text{m}\text{o}\text{l}\) but also \(\:0.3\:\text{k}\text{c}\text{a}\text{l}/\text{m}\text{o}\text{l}\) and \(\:0.5\:\text{k}\text{c}\text{a}\text{l}/\text{m}\text{o}\text{l}\) for \(\:\epsilon\:\).
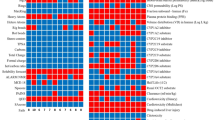
Table 1 presents \(\:{Q}_{PC}\left(\epsilon\:\right)\) of the four systems together with their dissociation constants \(\:{K}_{d}\). The dissociation constant \(\:{K}_{d}\) was reported from a study conducted by experimentation33. Table 2 presents the correlation coefficient between \(\:{Q}_{PC}\left(\epsilon\:\right)\) and \(\:{K}_{d}\) at each value of \(\:\epsilon\:\). It is particularly interesting that the correlation coefficient increased concomitantly with increasing \(\:\epsilon\:\) and that the strongest correlation was − 0.93 from \(\:\epsilon\:=1.0\:\text{k}\text{c}\text{a}\text{l}/\text{m}\text{o}\text{l}\). In general, the structural variety of the compound in the complex increases concomitantly with increasing \(\:\epsilon\:\). Consequently, the conformations with \(\:\epsilon\:=1.0\:\text{k}\text{c}\text{a}\text{l}/\text{m}\text{o}\text{l}\) have the greatest structural diversity among the three values of \(\:\epsilon\:\) examined. Those with \(\:\epsilon\:=0.3\:\text{k}\text{c}\text{a}\text{l}/\text{m}\text{o}\text{l}\) have the least diversity. Therefore, we infer that the reported dissociation constant found through experimentation is contributed by stably formed complex conformations, even when those conformations involve some structural diversity.
The correlation coefficient of -0.93 found for \(\:{Q}_{PC}\left(\epsilon\:=1.0\:\text{k}\text{c}\text{a}\text{l}/\text{m}\text{o}\text{l}\right)\) does not mean that the current study reproduced the absolute experimentally obtained value for \(\:{K}_{d}\). Instead, the current study shows that the order of \(\:{K}_{d}\) for the four compounds was reproduced well. Therefore, we infer that the current method is useful to select good candidate compounds among the computed compounds.
In fact, the periodic boundary box of the current study is smaller than that to reproduce the absolute value of \(\:{K}_{d}\). The box–volume is expected to involve a sufficient margin where the compound fluctuates without any effect from PLpro. A simple way to define the thermodynamic stability of a complex is to calculate the ratio \(\:\varDelta\:{Q}_{bound}/\varDelta\:{Q}_{unbound}\), where \(\:\varDelta\:{Q}_{bound}\) is the probability assigned to the bound state (the stable complex state) and \(\:\varDelta\:{Q}_{unbound}\) is that assigned to the unbound state. If the margin–volume is not counted accurately, then \(\:\varDelta\:{Q}_{unbound}\) cannot be estimated accurately. It is noteworthy that this argument omits correction by a huge volume of bulk solvent, which exists outside the simulation box in a real system, and omits correction by the ligand and receptor concentrations in the solution. We emphasize that the high correlation (-0.93) between \(\:{Q}_{PC}\left(1.0\:\text{k}\text{c}\text{a}\text{l}/\text{m}\text{o}\text{l}\right)\) and \(\:{K}_{d}\) suggests that the error resulted from the margin region was cancelled among the four systems. Finally, we conclude that the current method is useful to screen good candidates for use as inhibitors.
Structural clusters
As shown in the preceding subsection, the complex structures from the free-energy basin exhibited some structural variety (Fig. 5). This variety suggests that the stable conformations are mixtures of some binding modes. Consequently, the structural clustering might help to identify the binding modes of the compound and to discuss whether these modes have inhibitory activity to PLpro, or not.
We applied t-SNE and DBSCAN to the conformational ensemble for the structural clustering, which is based on the atom-contact patterns between the compounds and PLpro (Eq. 6). Because it is difficult to set the optimal parameters for t-SNE and DBSCAN in general, we conducted clustering with three sets of input parameters to search the conformational clusters comprehensively: Set 1, Set 2, and Set 3.58,59,60 Parameters of Set 1 are \(\:perplexity=100\), \(\:init=pca\), and \(\:learning\_rate=800\) for t-SNE, and \(\:eps=2.0\) and \(\:minpts=14000\) for DBSCAN. Those of Set 2 are \(\:perplexity=150\), \(\:init=pca\), and \(\:learning\_rate=800\) for t-SNE, and eps=1.5 and \(\:minpts=15000\) for DBSCAN. Those of Set 3 are \(\:perplexity=150\), \(\:init=pca\), and \(\:learning\_rate=800\) for t-SNE, and \(\:eps=2.5\) and \(\:minpts=15000\) for DBSCAN. Table 3 presents the resultant clusters \(\:{Cls}_{m}^{\left({\upalpha\:}\right)}\) and their probabilities \(\:{P}_{{Cls}_{m}^{\left(\alpha\:\right)}}\) (Eq. 9) for each system: \(\:{\upalpha\:}=\) PLpro–GRL0617, PLpro–XR8-24, PLpro–XR8-23 or PLpro–ZN-3-56. To specify that a cluster \(\:{Cls}_{m}^{\left({\upalpha\:}\right)}\) resulted from parameter Set \(\:i\) (\(\:i=1,\:2\:\text{o}\text{r}\:3\)), we designate the cluster as \(\:{Cls}_{m}^{\left({\upalpha\:}\right)}\)(Set \(\:i\)) if necessary. In Table 3, snapshots in which the compound did not contact to PLpro were classified in a single cluster \(\:{Cls}_{D}^{\left({\upalpha\:}\right)}\). Those in which only the non-core region of the compound contacted to PLpro were also assigned to \(\:{Cls}_{D}^{\left({\upalpha\:}\right)}\) because such a contact was not counted in Eq. 6 (\(\:{c}_{i,j}^{\left(k\right)}=0\)). “Outliers” in Table 3 are conformations that were not assigned to a cluster (i.e., singletons). A large majority of the outliers were conformations for which the core region contacted to a surface of PLpro other than the ligand-binding site. Because these contacts were less stable than the contacts formed between the compound’s core region and the PLpro’s ligand-binding site, no cluster was formed by the conformations in the outliers. Finally, a few (1, 2 or 3) clusters were identified from the clustering.
The intermolecular contact pattern in each cluster from parameter Set 1 is shown in Figs. 6, 7, 8 and 9. Those from Set 2 and Set 3 are presented respectively in Supplementary Figures S11 and S12. In all clusters, the compounds contacted to the ligand-binding site (\(\:{G}_{A}^{\left(RC1\right)}\) and \(\:{G}_{B}^{\left(RC1\right)}\); Fig. 1b) of PLpro, irrespective of the compound type. Although Figs. 6, 7, 8 and 9, S11 and S12 clearly characterize the contact patterns of the clusters, we cannot imagine the tertiary structures of the clusters from the figures.
Contact map between PLpro (X-axis) and GRL0617 (Y axis) for each cluster with clustering parameter Set 1 (Table 3). The contact probability, which is normalized in each cluster, is presented by color scale. Panels (a–c) are for clusters \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\), \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) and \(\:{Cls}_{3}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) of \(\:{\upalpha\:}=\) PLpro–GRL0617 system. Only the names of atoms in the core region are displayed on the Y axis (Fig. S5 of SI). Small green rings denote the position of atom group \(\:{G}_{A}^{\left(RC1\right)}\) (residues 266–271) and \(\:{G}_{B}^{\left(RC1\right)}\) (residues 165–168) (Fig. 1b). Space between \(\:{G}_{A}^{\left(RC1\right)}\) and \(\:{G}_{B}^{\left(RC1\right)}\) corresponds to the ligand-binding site. The green rectangle represents C111 of PLpro contacting the core-region of the compounds.
Contact map between PLpro (X-axis) and XR8-24 (Y axis) for each cluster with clustering parameter Set 1 (Table 3). The contact probability, which is normalized in each cluster, is presented by color scale. This figure is for clusters \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) of \(\:{\upalpha\:}=\) PLpro–XR8-24 system. Only the names of atoms in the core region are displayed on the Y axis (Fig. S5 of SI). Small green rings denote the position of atom group \(\:{G}_{A}^{\left(RC1\right)}\) (residues 266–271) and \(\:{G}_{B}^{\left(RC1\right)}\) (residues 165–168) (Fig. 1b). Space between \(\:{G}_{A}^{\left(RC1\right)}\) and \(\:{G}_{B}^{\left(RC1\right)}\) corresponds to the ligand-binding site.
Contact map between PLpro (X-axis) and XR8-23 (Y axis) for each cluster with clustering parameter Set 1 (Table 3). The contact probability, which is normalized in each cluster, is presented by color scale. Panels (a–c) are for clusters \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\), \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) and \(\:{Cls}_{3}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) of \(\:{\upalpha\:}=\) PLpro–XR8-23 system. Only the names of atoms in the core region are displayed on the Y axis (Fig. S5 of SI). Small green rings denote the position of atom group \(\:{G}_{A}^{\left(RC1\right)}\) (residues 266–271) and \(\:{G}_{B}^{\left(RC1\right)}\) (residues 165–168) (Fig. 1b). Space between \(\:{G}_{A}^{\left(RC1\right)}\) and \(\:{G}_{B}^{\left(RC1\right)}\) corresponds to the ligand-binding site. The green rectangle represents C111 of PLpro contacting the core-region of the compounds.
Contact map between PLpro (X-axis) and ZN-3-56 (Y axis) for each cluster with clustering parameter Set 1 (Table 3). The contact probability, which is normalized in each cluster, is presented by color scale. Panels (a–c) are for clusters \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\), \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) and \(\:{Cls}_{3}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) of \(\:{\upalpha\:}=\) PLpro–ZN-3-56 system. Only the names of atoms in the core region are displayed on the Y axis (Fig. S5 of SI). Small green rings denote the position of atom group \(\:{G}_{A}^{\left(RC1\right)}\) (residues 266–271) and \(\:{G}_{B}^{\left(RC1\right)}\) (residues 165–168) (Fig. 1b). Space between \(\:{G}_{A}^{\left(RC1\right)}\) and \(\:{G}_{B}^{\left(RC1\right)}\) corresponds to the ligand-binding site. The green rectangle represents C111 of PLpro contacting the core-region of the compounds.
The complex structures from the clusters of the four systems from parameter Set 1 are displayed in Figs. 10, 11, 12 and 13. Those from Set 2 and Set 3 are shown respectively in Supplementary Figures S13 and S14. These complex structures were typical of the clusters to which the structures belonged. The figures again show that the core-region of the compounds bound to the ligand-binding site of PLpro.
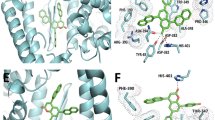
Complex structure picked from each cluster obtained from clustering parameter Set 1 of \(\:{\upalpha\:}=\) PLpro–GRL0617 system (Table 3). Panels (a), (b) and (c) are respectively for clusters \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\), \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) and \(\:{Cls}_{3}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\).The orange segment is the loop region (\(\:{G}_{A}^{\left(RC1\right)}\); residues 266–271). The green one is a part of helix (\(\:{G}_{B}^{\left(RC1\right)}\) ; residues 165–168): Fig. 1b shows details for these segments. Side-chains of two amino-acid residues Y268 and Q269 are labeled. Cornflower blue molecules are compounds, with broken-line circles representing their core regions (Fig. S5 of SI).
Complex structure picked from each cluster obtained from clustering parameter Set 1 of \(\:{\upalpha\:}=\) PLpro–XR8-24 system (Table 3). This figure is respectively for clusters \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\).The orange segment is the loop region (\(\:{G}_{A}^{\left(RC1\right)}\); residues 266–271). The green one is a part of helix (\(\:{G}_{B}^{\left(RC1\right)}\) ; residues 165–168): Fig. 1b shows details for these segments. Side-chains of two amino-acid residues Y268 and Q269 are labeled. Cornflower blue molecules are compounds, with broken-line circles representing their core regions (Fig. S5 of SI).
Complex structure picked from each cluster obtained from clustering parameter Set 1 of \(\:{\upalpha\:}=\) PLpro–XR8-23 system (Table 3). Panels (a–c) are respectively for clusters \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\), \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) and \(\:{Cls}_{3}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\).The orange segment is the loop region (\(\:{G}_{A}^{\left(RC1\right)}\); residues 266–271). The green one is a part of helix (\(\:{G}_{B}^{\left(RC1\right)}\) ; residues 165–168): Fig. 1b shows details for these segments. Side-chains of two amino-acid residues Y268 and Q269 are labeled. Cornflower blue molecules are compounds, with broken-line circles representing their core regions (Fig. S5 of SI).
Complex structure picked from each cluster obtained from clustering parameter Set 1 of \(\:{\upalpha\:}=\) PLpro–ZN-3-56 system (Table 3). Panels (a–c) are respectively for clusters \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\), \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) and \(\:{Cls}_{3}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\).The orange segment is the loop region (\(\:{G}_{A}^{\left(RC1\right)}\); residues 266–271). The green one is a part of helix (\(\:{G}_{B}^{\left(RC1\right)}\) ; residues 165–168): Fig. 1b shows details for these segments. Side-chains of two amino-acid residues Y268 and Q269 are labeled. Cornflower blue molecules are compounds, with broken-line circles representing their core regions (Fig. S5 of SI).
For \(\:{\upalpha\:}=\) PLpro–GRL0617, three clusters were obtained from Set 1. The first cluster \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) was similar to the ligand orientations of the native-like complex structures (PDB ID: 7CJM), but the side-chain of Y268 was standing on the surface of PLpro and the side-chain of Q269 covered the core region of GRL0617 (Fig. 10a). The second cluster \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) consisted of the native-like complex structures where the side-chains of Y268 and Q269 covered the core region of GRL0617 (Fig. 10b). In the third cluster \(\:{Cls}_{3}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\), by contrast, the orientation of GRL0617 was greatly different from that in \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) (Fig. 10b) where the side-chains of Y268 and Q269 were standing on the PLpro surface and were in contact with the double-ring part (non-core region) of GRL0617 (Fig. 10c). In parameter Set 2, we again obtained three clusters: \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:2\right)\) (Figure S13a), \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:2\right)\) (Figure S13b) and \(\:{Cls}_{3}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:2\right)\) (Figure S13c), with binding modes that were respectively similar with those of \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\), \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) and \(\:{Cls}_{3}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\). Also, two clusters were identified from Set 3: \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:3\right)\), and \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:3\right)\). The cluster \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:3\right)\) was similar to \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) and \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:2\right)\) (Figure S14a), and \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:3\right)\) was similar to \(\:{Cls}_{3}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) and \(\:{Cls}_{3}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:2\right)\) (Figure S14b).
For \(\:{\upalpha\:}=\) PLpro–XR8-24, only one cluster \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) was obtained from parameter Set 1. XR8-24 had a native-like binding mode (PDB ID: 7LBS) and the side-chains Y268 and Q269 covered XR8-24 (Fig. 11). It is particularly interesting that \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) was divided into two clusters in Set 2: \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:2\right)\) and \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:2\right)\). In \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:2\right)\), the core region of XR8-24 did not contact to C111 of PLpro(Figure S11d), although the core region contacted to C111 of that in \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:2\right)\) (Figure S11e). In Set 3, we found only one cluster \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:3\right)\) (Figure S14c), for which the binding modes were similar, respectively, with those of \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\).
For \(\:{\upalpha\:}=\) PLpro–XR8-23, three clusters were obtained from Set 1. In \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\), the side-chains of Y268 and Q269 covered the core region (Fig. 12a). In \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\), however, the side-chains of Y268 and Q269 were standing on the PLpro surface and the tail part (non-core region) contacted to the side-chains (Fig. 12b). Furthermore, in \(\:{Cls}_{3}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\), the core region of XR8-23 was covered by the side-chains of Y268 and Q269, as in \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\), and also contacted C111 (Figs. 8c and 12c). The parameters Set 2 and Set 3 obtained two clusters each. For Set 2, the binding mode of \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:2\right)\) (Figure S13f) was similar to that of \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\). That of \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:2\right)\) (Figure S13g) was similar to that of \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\). This situation was the same for Set 3. The binding modes of \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:3\right)\) and \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) were similar. Those of \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:3\right)\) and \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) were similar.
For \(\:{\upalpha\:}=\) PLpro–ZN-3-56, three clusters were obtained from Set 1. In the cluster \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\), the side-chain of Y268 was standing on the surface of PLpro and the side chain of Q269 covered the core region of ZN-3-56 (Fig. 13a). In both \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) (Fig. 13b) and \(\:{Cls}_{3}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) (Fig. 13c), the core region of ZN-3-56 was covered by the side chains of Y268 and Q269. In this respect, \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) and \(\:{Cls}_{3}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) were similar. However, in \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\), the ZN-3-56 core region did not contact to C111 of PLpro (Fig. 9b), whereas in \(\:{Cls}_{3}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\), the core region did contact to C111 of that (Fig. 9c). In Set 2, by contrast, we found only one cluster: \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:2\right)\). The binding mode of \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:2\right)\) was similar to that of \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) (Figure S13h). This point is argued in the next paragraph. In Set 3, two clusters \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:3\right)\) and \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:3\right)\) were obtained. The first cluster \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:3\right)\) corresponded to \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) and \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:2\right)\). Also, the second cluster \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:3\right)\) was similar to \(\:{Cls}_{3}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\).
The results of clustering with three parameter sets showed that even though the ligand orientations are similar, they are classified as different clusters due to the different orientations of the Y268 and Q269 side-chains in the ligand binding site. For example, \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) and \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) of α = PLpro-GRL0617 had the same ligand orientation (Fig. 10a and b), but the orientation of the side-chains of Y268 and Q269 were different. In \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\), the side-chain of Y268 was perpendicular to the surface of PLpro, while the side-chain of Q269 was parallel to the surface of PLpro and covered the core region of GRL0617 (Fig. 10a). However, in \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\), both the side-chains of Y268 and Q269 covered the core region of GRL0617 (Fig. 10b). The classification of those binding modes was also similar to the parameter Set 2 (\(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:2\right)\) and \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:2\right)\)) for α = PLpro-GRL0617, Set 1 (\(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) and \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\)) and Set 3 (\(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:3\right)\) and \(\:{Cls}_{3}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:3\right)\)) for α = PLpro-ZN-3-56. As described above, even though the orientation of the ligands were similar, they were classified as different clusters due to the different side-chain orientations of Y268 and Q269 (\(\:{G}_{A}^{\left(RC1\right)}\)) at the ligand-binding site. Furthermore, the clusters similar to \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) and \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:2\right)\) for α = PLpro-GRL0617 were obtained from parameter Set 3. Therefore, this is a concrete case in which the clustering result depends on the parameter set. We infer that it is important to examine some parameter sets to analyze the binding modes comprehensively. Also, depending on the orientation of the ligands, we were able to detect clusters of structures in which the side chains of Y268 and Q269 were parallel or perpendicular to the surface of PLpro. These structural features are used to assign the clusters obtained from the structural clustering to a binding mode below.
The position of the core region relative to the ligand-binding site of PLpro was common for all the clusters. Consequently, the core-region position is not a good identifier for analyzing the binding mode. By contrast, the molecular orientation of the compound relative to the PLpro’s surface can be a good identifier. For instance, the compound’s orientation in \(\:{Cls}_{1}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) and \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) for \(\:{\upalpha\:}=\) PLpro–GRL0617 was approximately parallel to the PLpro surface (Fig. 10a and b). By contrast, in \(\:{Cls}_{3}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\), a part of the compound (usually non-core region) was perpendicular to the surface (Fig. 10c). As another identifier, the side-chain orientation of Y268 and Q269 of PLpro relative to the surface of PLpro exhibited clear differences among the clusters. The side-chains in \(\:{Cls}_{2}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) covered the core region of the compound, whereas those in \(\:{Cls}_{3}^{\left({\upalpha\:}\right)}\left(\text{S}\text{e}\text{t}\:1\right)\) contacted to a part (usually non-core region) of the compound. Therefore, we classified the complex structures into two binding modes using the identifiers above: a parallel mode (Fig. 14a) and a perpendicular mode (Fig. 14b). In the parallel mode, the compound is approximately parallel to the PLpro surface. And, the two side-chains of Y268 and Q269 or the side-chains of Q269 cover the core region of the compound. In the perpendicular mode, a part (mostly the non-core region) of the compound took an orientation approximately perpendicular to the PLpro surface. This part contacts to the side-chains of Y268 and/or Q269, which are also approximately perpendicular to the PLpro surface. Tables 4 and 5 respectively assign clusters to the parallel mode and the perpendicular mode.
Schematical representation of two binding modes: (a) parallel mode and (b) perpendicular mode. The red model is compound, with a hexagon as its core region. The blue vector with the label “perpendicular” in panel (b) shows that the side chains of Y268 and Q269 are standing on the PLpro surface. The main text explains more details related to the modes.
In all clusters, the core region bound to the ligand–binding site of PLpro (Figs. 10, 11, 12 and 13 and S13 and S14). Therefore, we inferred that all clusters obtained here have an enzyme-inhibitory effect. We defined the fraction for these clusters to the entire conformational ensemble as
In this equation, the probabilities for \(\:{Cls}_{D}^{\left({\upalpha\:}\right)}\) and outliers were eliminated from the summation. Table 6 presents the resultant \(\:{Q}_{inhib}^{\left(\alpha\:\right)}\) and the enzyme inhibition \(\:I{C}_{50}\)33 for each system: \(\:I{C}_{50}\) is the concentration of the compound in the solution at which 50% of the enzymatic function of PLpro is inhibited. Table 7 presents the correlation coefficient between \(\:{Q}_{inhib}^{\left(\alpha\:\right)}\) and \(\:I{C}_{50}\). Also, Table 8 presents the correlation coefficient between \(\:{Q}_{inhib}^{\left(\alpha\:\right)}\) and \(\:{K}_{d}\). Those shows that the coefficient is anti-correlative, irrespective of the parameter set. In fact, lower values of \(\:I{C}_{50}\) and \(\:{K}_{d}\) are generally associated with stronger inhibition. Therefore, the anti-correlation reflects that the computed \(\:{Q}_{inhib}^{\left(\alpha\:\right)}\) supports the experimental \(\:I{C}_{50}\) and \(\:{K}_{d}\) for all parameter sets. Findings from the present study suggest that the mD-VcMD simulation can be used effectively for screening promising compounds from others with using \(\:{Q}_{inhib}^{\left({\upalpha\:}\right)}\). Consequently, GRL0617, XR8-24 and XR8-23 are more promising than and ZN-3-56.
As described earlier, correlation between \(\:{Q}_{PC}\left(\epsilon\:\right)\) (Eq. 11) and \(\:{K}_{d}\) increased concomitantly with increase of threshold \(\:\epsilon\:\). The best correlation was from \(\:\epsilon\:=1.0\) kcal / mol (-0.93; Table 2) among the three examined \(\:\epsilon\:\) values. Therefore, the complex state involves some structural variety. Furthermore, the complex state, which has inhibitory activity, consists of structurally distinct clusters (Fig. 14). These results might represent a concrete case in which both the receptor and the ligand (i.e., the compound in the present study) vary their tertiary structures to search stable complex structures61. Furthermore, the existence of two binding modes suggests that the binding mechanism of the current system is a fuzzy complex formation62 or a multimodal binding mechanism10,16.
We examined four compounds to verify our screening procedure. One suggestion is to apply the procedure to more compounds. We consider that suggestion to be valid. An earlier study33 conducted by experimentation found \(\:{k}_{d}\) and \(\:I{C}_{50}\) values measured for many compounds, which bind to PLpro. We consider that future studies should be done for those compounds to clarify the correlation between computational probabilities (\(\:{Q}_{PC}\left(\epsilon\:\right)\) or \(\:{Q}_{inhib}^{\left({\upalpha\:}\right)}\)) and experimentally obtained \(\:{K}_{d}\) or \(\:I{C}_{50}\) values.
Determination of atom groups \(\:{G}_{A}^{\left(RCi\right)}\) and \(\:{G}_{B}^{\left(RCi\right)}\) is important to set RCs effectively. We presume that an inappropriate RC set requires more computational time than an appropriate one, although any set of RCs can produce a canonical ensemble of the system, in theory. For the present study, the crystallographic structures of the PLpro–GRL0617 complex (PDB ID: 7CJM) and the PLpro–XR8-24 complex (PDB ID: 7LBS) helped the setting of RCs. However, if an essential region for the target compound and the binding site of the receptor is unknown, then another approach is required. Moreover, such an approach must be free from the setting of atom groups. Such a version free from atom-group setting is now under development.
We emphasize that mD-VcMD is a method to compute a free-energy landscape to study biomolecular binding4,6,21,43,44. The study described herein specifically addressed only the free-energy basin of the conformational space because the aim of this study is verification of the correlation between the computational \(\:{Q}_{PC}\left(\epsilon\:\right)\) (or \(\:{Q}_{inhib}^{\left({\upalpha\:}\right)}\)) and the experimentally obtained values of \(\:{K}_{d}\) (or \(\:I{C}_{50}\)). In this last paragraph, it must be stated that the free-energy landscape and the conformational ensemble from mD-VcMD can provide various intermediates (encounter complexes), which might be far from the free-energy basin in the conformational space, and pathways among the various complexes. Such insights from the free-energy landscape might provide important hints supporting drug discovery, along with knowledge related to the basic study of biology.
As Zn ions in PLpro are known to be important for the structure and function61,62,63, we also performed preliminary mD-VcMD simulations including zinc ions. We then calculated the RMSD (root-mean-squared-displacement among conformations) for the entire PLpro and for the binding site residues. The resulting RMSD values ranged between 2 and 3 Å in almost all calculations, regardless of the presence or absence of Zn ions, using the experimental structure (PDB ID: 7CJM) as the reference. Importantly, the experimental binding pose was successfully reproduced, likely because the Zn ions are located far from the ligand-binding site and the overall PLpro structure remained unchanged. The structural integrity of PLpro during the simulation was maintained by setting an upper limit for reaction coordinate RC1 (25.49 Å; see Table S4), which was introduced to effectively enhance ligand–receptor binding. We presume that this upper limit helped prevent structural collapse. Therefore, we conducted detailed mD-VcMD simulations excluding zinc ions in this study and confirmed that the PLpro structure was preserved across all systems. Notably, the binding site of PLpro underwent substantial movement due to the enhancement of RC1 and RC2, without structural collapse, and the conformational space was extensively sampled, as shown in Fig. 2.
Our simulation results are consistent with the experimental results reported by Shen et al.33, as shown in Tables 1, 2 and 6–8, confirming the validity of our computational method.
Data availability
All data generated or examined throughout this study are included in this published article and its Supplementary Information.
References
Bekker, G. J., Fukuda, I., Higo, J. & Kamiya, N. Cryptic-site binding mechanism of medium-sized Bcl-xL inhibiting compounds elucidated by McMD-based dynamic Docking simulations. Sci. Rep. 11, 5046. https://doi.org/10.1038/s41598-021-84488-z (2021).
Bekker, G. J., Fukuda, I., Higo, J. & Kamiya, N. Mutual population-shift driven antibody-peptide binding elucidated by molecular dynamics simulations. Sci. Rep. 10, 1406. https://doi.org/10.1038/s41598-020-58320-z (2020).
Fukunishi, Y., Higo, J. & Kasahara, K. Computer simulation of molecular recognition in biomolecular system: from in-silico screening to generalized ensembles. Biophys. Rev. 14, 1423–1447. https://doi.org/10.1007/s12551-022-01015-8 (2022).
Hayami, T. et al. Difference of binding modes among three ligands to a receptor mSin3B corresponding to their inhibitory activities. Sci. Rep. 11, 6178. https://doi.org/10.1038/s41598-021-85612-9 (2021).
Moritsugu, K. et al. Flexibility and cell permeability of Cyclic Ras-Inhibitor peptides revealed by the coupled Nosé–Hoover equation. J. Chem. Inf. Model. 61, 1921–1930. https://doi.org/10.1021/acs.jcim.0c01427 (2021).
Higo, J., Takashima, H., Fukunishi, Y. & Yoshimori, A. Generalized-ensemble method study: A helix-mimetic compound inhibits protein–protein interaction by long-range and short-range intermolecular interactions. J. Comput. Chem. 42, 956–969. https://doi.org/10.1002/jcc.26516 (2021).
Kasahara, K. et al. myPresto/omegagene 2020: a molecular dynamics simulation engine for virtual-system coupled sampling. Biophys. Physicobiol. 17, 140–146. https://doi.org/10.2142/biophysico.BSJ-2020013 (2020).
Higo, J. et al. GA-guided mD-VcMD: A genetic-algorithm-guided method for multi-dimensional virtual-system coupled molecular dynamics. Biophys. Physicobiology. 17, 161–176. https://doi.org/10.2142/biophysico.BSJ-2020008 (2020).
Higo, J. et al. Molecular interaction mechanism of a 14-3-3 protein with a phosphorylated peptide elucidated by enhanced conformational sampling. J. Chem. Inf. Model. 60, 4867–4880. https://doi.org/10.1021/acs.jcim.0c00551 (2020).
Iida, S. et al. Multimodal structural distribution of the p53 C-terminal domain upon binding to S100B via a generalised ensemble method: from disorder to Extra-Disorder. J. Chem. Theory Comput. 15, 2597–2607. https://doi.org/10.1021/acs.jctc.8b01042 (2019).
Shimato, T., Kasahara, K., Higo, J. & Takahashi, T. Effects of number of parallel runs and frequency of bias-strength replacement in generalized ensemble molecular dynamics simulations. PeerJ Phys. Chem. 1, e4. https://doi.org/10.7717/peerj-pchem.4 (2019).
Hayami, T., Higo, J., Nakamura, H. & Kasahara, K. Multidimensional virtual-system coupled canonical molecular dynamics to compute free-energy landscapes of peptide multimer assembly. J. Comput. Chem. 40, 2453–2463. https://doi.org/10.1002/jcc.26020 (2019).
Kasahara, K., Terazawa, H., Takahashi, T. & Higo, J. Studies on molecular dynamics of intrinsically disordered proteins and their fuzzy complexes: a mini-review. Comput. Struct. Biotech. J. 17, 712–720. https://doi.org/10.1016/j.csbj.2019.06.009 (2019).
Iwai, R., Kasahara, K. & Takahashi, T. Influence of various parameters in the replica-exchange molecular dynamics method: number of replicas, replica-exchange frequency, and thermostat coupling time constant. Biophys. Physicobiol. 15, 165–172. https://doi.org/10.2142/biophysico.15.0_165 (2018).
Hayami, T., Kasahara, K., Nakamura, H. & Higo, J. Molecular dynamics coupled with a virtual system for effective conformational sampling. J. Comput. Chem. 39, 1291–1299. https://doi.org/10.1002/jcc.25196 (2018).
Kasahara, K. et al. Phosphorylation of an intrinsically disordered region of Ets1 shifts a multi-modal interaction ensemble to an out-inhibitory state. Nucleic Acids Res. 46, 2243–2251. https://doi.org/10.1093/nar/gkx1297
Higo, J., Kasahara, K. & Nakamura, H. Multi-dimensional virtual system introduced to enhance canonical sampling. J. Chem. Phys. 147, 134102. https://doi.org/10.1063/1.4986129 (2017).
Higo, J., Kasahara, K., Dasgupta, B. & Nakamura, H. Enhancement of canonical sampling by virtual-state transitions. J. Chem. Phys. 146, 044104. https://doi.org/10.1063/1.4974087 (2017).
Kasahara, K. et al. myPresto/omegagene: a GPU-accelerated molecular dynamics simulator tailored for enhanced conformational sampling methods with a non-Ewald electrostatic scheme. Biophys. Physicobiology. 13, 209–216. https://doi.org/10.2142/biophysico.13.0_209 (2016).
Ikebe, J. et al. Theory for trivial trajectory parallelization of multicanonical molecular dynamics and application to a polypeptide in water. J. Comput. Chem. 32, 1286–1297. https://doi.org/10.1002/jcc.21710 (2011).
Higo, J. et al. Fly casting with ligand sliding and orientational selection supporting complex formation of a GPCR and a middle sized flexible molecule. Sci. Rep. 12, 13792. https://doi.org/10.1038/s41598-022-17920-7 (2022).
Kurita, J. et al. Sertraline, chlorprothixene, and chlorpromazine characteristically interact with the REST-binding site of the corepressor mSin3, showing Medulloblastoma cell growth inhibitory activities. Sci. Rep. 8, 13763. https://doi.org/10.1038/s41598-018-31852-1 (2018).
Shin, D. et al. Papain-like protease regulates SARS-CoV-2 viral spread and innate immunity. Nature 587, 657–662. https://doi.org/10.1038/s41586-020-2601-5 (2020).
Narayanan, A. et al. Identification of SARS-CoV-2 inhibitors targeting Mpro and PLpro using in-cell-protease assay. Commun. Biol. 5, 169. https://doi.org/10.1038/s42003-022-03090-9 (2022).
Zhao, Y. et al. High-throughput screening identifies established drugs as SARS-CoV-2 PLpro inhibitors. Protein Cell. 12, 877–888. https://doi.org/10.1007/s13238-021-00836-9 (2021).
Calleja, D. J., Lessene, G. & Komander, D. Inhibitors of SARS-CoV-2 PLpro. Front. Chem. 10, 876212. https://doi.org/10.3389/fchem.2022.876212 (2022).
Zhang, L. et al. Design of SARS-CoV-2 Mpro, PLpro dual-target inhibitors based on deep reinforcement learning and virtual screening. Future Med. Chem. 14, 393–405. https://doi.org/10.4155/fmc-2021-0269 (2022).
Negru, P. A. et al. Virtual screening of substances used in the treatment of SARS-CoV-2 infection and analysis of compounds with known action on structurally similar proteins from other viruses. Biomed. Pharmacother. 153, 113432. https://doi.org/10.1016/j.biopha.2022.113432 (2022).
Kidera, A., Moritsugu, K., Ekimoto, T. & Ikeguchi, M. Allosteric regulation of 3CL protease of SARS-CoV-2 and SARS-CoV observed in the crystal structure ensemble. J. Mol. Biol. 443, 167324. https://doi.org/10.1016/j.jmb.2021.167324 (2021).
Moritsugu, K., Ekimoto, T., Ikeguchi, M. & Kidera, A. Binding and unbinding pathways of peptide substrates on the SARS-CoV-2 3CL protease. J. Chem. Inf. Model. 63, 240–250. https://doi.org/10.1021/acs.jcim.2c00946 (2023).
Hengphasatporn, K. et al. Promising SARS-CoV-2 main protease inhibitor ligand-binding modes evaluated using LB-PaCS-MD/FMO. Sci. Rep. 12, 17984. https://doi.org/10.1038/s41598-022-22703-1 (2022).
Sencanski, M. et al. Identification of SARS-CoV-2 Papain-like protease (PLpro) inhibitors using combined computational approach. ChemistryOpen 11, e202100248. https://doi.org/10.1002/open.202100248 (2022).
Shen, Z. et al. Design of SARS-CoV-2 PLpro inhibitors for COVID-19 antiviral therapy leveraging binding cooperativity. J. Med. Chem. 65, 2940–2955. https://doi.org/10.1021/acs.jmedchem.1c01307 (2022).
Van der Maaten, L. & Hinton, G. Visualizing data using t-SNE. J. mach. Learn. Res. 9, 2579–2605 (2008).
Ester, M., Kriegel, H. P., Sander, J. & Xu, X. A density-based algorithm for discovering clusters in large spatial databases with noise. In Proceedings of the Second International Conference on Knowledge Discovery and Data Mining (KDD’96). 226–231. (AAAI, 1996).
Fu, Z. et al. The complex structure of GRL0617 and SARS-CoV-2 PLpro reveals a hot spot for antiviral drug discovery. Nat. Commun. 12, 488. https://doi.org/10.1038/s41467-020-20718-8 (2021).
Lindorff-Larsen, K. et al. Improved side-chain torsion potentials for the amber ff99SB protein force field. Proteins: Struct. Funct. Genet. 78, 1950–1958. https://doi.org/10.1002/prot.22711 (2010).
Joung, I. S. & Cheatham, T. E. III Determination of alkali and halide monovalent ion parameters for use in explicitly solvated biomolecular simulations. J. Phys. Chem. B. 112, 9020–9041. https://doi.org/10.1021/jp8001614 (2008).
Jorgensen, W. L. et al. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–936. https://doi.org/10.1063/1.445869 (1983).
Frisch, M. J. et al. Gaussian 16, Revision C.01 Gaussian, Inc., https://gaussian.com/gaussian16/ (2016).
Bayly, C. I., Cieplak, P., Cornell, W. & Kollman, P. A. A well-behaved electrostatic potential based method using charge restraints for deriving atomic charges: the RESP model. J. Phys. Chem. 97, 10269–10280. https://doi.org/10.1021/j100142a004 (1993).
Wang, J. M. et al. Development and testing of a general amber force field. J. Comput. Chem. 25, 1157–1174. https://doi.org/10.1002/jcc.20035 (2004).
Higo, J. et al. Free-energy landscape of molecular interactions between endothelin 1 and human endothelin type B receptor: Fly-casting mechanism. Protein Eng. Des. Sel. 32, 297–308. https://doi.org/10.1093/protein/gzz029 (2019).
Higo, J. et al. Binding free-energy landscapes of small molecule binder and non-binder to FMN riboswitch: All-atom molecular dynamics. Biophys. Physicobiology. 20, e200047. https://doi.org/10.2142/biophysico.bppb-v20.0047 (2023).
Higo, J. et al. Verifying trivial parallelization of multicanonical molecular dynamics for conformational sampling of a polypeptide in explicit water. Chem. Phys. Lett. 473, 326–329. https://doi.org/10.1016/j.cplett.2009.03.077 (2009).
Kutzner, C. et al. More Bang for your buck: improved use of GPU nodes for GROMACS 2018. J. Comput. Chem. 40, 2418–2431. https://doi.org/10.1002/jcc.26011 (2019).
Bonomi, M. et al. PLUMED: A portable plugin for free-energy calculations with molecular dynamics. Comp. Phys. Commun. 180, 1961–1972. https://doi.org/10.1016/j.cpc.2009.05.011 (2009).
Hess, B., Bekker, H., Berendsen, H. J. C. & Fraaije, J. G. E. M. LINCS: A linear constraint solver for molecular simulations. J. Comput. Chem.18, 1463–1472 (1997).
Nosé, S. A unified formulation of the constant temperature molecular dynamics methods. J. Chem. Phys. 81, 511–519. https://doi.org/10.1063/1.447334 (1984).
Hoover, W. G. Canonical dynamics: equilibrium phase-space distributions. Phys. Rev. A. 31, 1695–1697. https://doi.org/10.1103/PhysRevA.31.1695 (1985).
Kamiya, N., Fukuda, I. & Nakamura, H. Application of zero-dipole summation method to molecular dynamics simulations of a membrane protein system. Chem. Phys. Lett. 568–569, 26–32. https://doi.org/10.1016/j.cplett.2013.03.014 (2013).
Fukuda, I., Yonezawa, Y. & Nakamura, H. Molecular dynamics scheme for precise Estimation of electrostatic interaction via zero-dipole summation principle. J. Chem. Phys. 134, 164107. https://doi.org/10.1063/1.3582791 (2011).
Torrie, G. M. & Valleau, J. P. Nonphysical sampling distributions in Monte Carlo free-energy estimation: umbrella sampling. J. Comput. Phys. 23, 187–199. https://doi.org/10.1016/0021-9991(77)90121-8 (1977).
Pearson, K. On lines and planes of closest fit to systems of points in space. Phil. Mag. 2, 559–572. https://doi.org/10.1080/14786440109462720 (1901).
Higo, J., Galzitskaya, O. V., Ono, S. & Nakamura, H. Energy landscape of a β-hairpin peptide in explicit water studied by multicanonical molecular dynamics. Chem. Phys. Lett. 337, 169–175. https://doi.org/10.1016/S0009-2614(01)00118-X (2001).
Higo, J. et al. Energy landscape of a peptide consisting of α-helix, 3_10-helix, β-turn, β-hairpin, and other disordered conformations. Protein Sci. 10, 1160–1171. https://doi.org/10.1110/ps.44901 (2008).
Xie, Q., Kasahara, K., Higo, J. & Takahashi, T. Molecular mechanisms of functional modulation of transcriptional coactivator PC4 via phosphorylation on its intrinsically disordered region. ACS Omega. 8, 14572–14582. https://doi.org/10.1021/acsomega.3c00364 (2023).
Li, M. et al. A novel Traveling-Wave-Based method improved by unsupervised learning for fault location of power cables via sheath current monitoring. Sensors 19, 2083. https://doi.org/10.3390/s19092083 (2019).
Sonnewald, M., Dutkiewicz, S., Hill, C. & Forget, G. Elucidating ecological complexity: unsupervised learning determines global marine eco-provinces. Sci. Adv. 6, eaay4740. https://doi.org/10.1126/sciadv.aay4740 (2020).
Yigin, B. O. & Saygili, G. Effect of distance measures on confidences of t-SNE embeddings and its implications on clustering for scRNA-seq data. Sci. Rep. 13, 6567. https://doi.org/10.1038/s41598-023-32966-x (2023).
Ratia, K. et al. Severe acute respiratory syndrome coronavirus papain-like protease: structure of a viral deubiquitinating enzyme. Proc. Natl. Acad. Sci. U S A. 103, 5717–5722. https://doi.org/10.1073/pnas.0510851103 (2006).
Barretto, N. et al. The papain-like protease of severe acute respiratory syndrome coronavirus has deubiquitinating activity. J. Virol. 79, 15189–15198. https://doi.org/10.1128/jvi.79.24.15189-15198.2005 (2005).
Arya, R., Prashar, V. & Kumar, M. Evaluating stability and activity of SARS-CoV-2 PLpro for high-throughput screening of inhibitors. Mol. Biotechnol. 64, 1–8. https://doi.org/10.1007/s12033-021-00383-y (2022).
Acknowledgements
This work was supported by JSPS KAKENHI Grants No. 21K06052 (J.H.) and was conducted in part under the Cooperative Research Program of the Institute for Protein Research, Osaka University, CR22-02 and CR-23-02. We acknowledge support by the HPCI System Research Project (Project IDs: hp220002, hp220015, hp220022, hp230003, hp230011, hp230013, hp230109, hp240032, hp240167, hp250017, and hp250061). J.H. is supported by the Project Focused on Developing Key Technology for Discovering and Manufacturing Drugs for Next-Generation Treatment and Diagnosis (project ID: 23ae0121028h0003; 2018–2021 and 2021–) from AMED and the Japan Biological Informatics Consortium (JBiC). The mD-VcMD was performed on TSUBAME4.0 supercomputers at the Institute of Science Tokyo. Other analyses were done at Ritsumeikan University and Research Center for Computational Science, Okazaki, Japan (Project: 25-IMS-C175). We used UCSF Chimera ver. 1.15 for drawing molecular structures and gnuplot for graphs. Other analyses were done using programs produced in-house.
Author information
Authors and Affiliations
Contributions
Research project was proposed by Masashi Muramoto and Takuya Takahashi. Computer program preparation was done by Masashi Muramoto, Qilin Xie and Kota Kasahara; MD simulation was performed by Masashi Muramoto. Analyses were performed by Muramoto Masashi, Simon Hikiri, Suzuka Saito and Junichi Higo. Manuscript writing was done by Masashi Muramoto, Simon Hikiri, Suzuka Saito, Junichi Higo and Takuya Takahashi. The entire process of this research was supervised by Takuya Takahashi.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Muramoto, M., Hikiri, S., Saito, S. et al. Affinity of drug candidates binding to SARS-CoV-2 PLpro assessed using a generalized-ensemble method. Sci Rep 16, 4214 (2026). https://doi.org/10.1038/s41598-025-34319-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-34319-2