Abstract
In the present work, qRT-PCR was used to detect miR-204-5p level in the normal oral epithelial cells (HaCat cell line) and OSCC cells (SCC-15 and SCC-25 cell lines). We explored the impact of miR-204-5p on OSCC through overexpression of miR-204-5p in OSCC cells by transient transfection. CCK-8, EdU, and clone formation assays were applied to estimate how miR-204-5p acts on OSCC proliferation in vitro. Wound healing and transwell assays were performed to evaluate the in vitro effect of miR-204-5p on OSCC migration. Flow cytometry, Dual luciferase report assay and western blotting were used to explore the mechanism. The in vivo effect of miR-204-5p was evaluated by xenograft models. Compared with normal cells, OSCC cells showed lower expression levels of miR-204-5p. miR-204-5p was evidently elevated in OSCC after transfection. In CCK-8, clone formation and Edu assays, miR-204-5p inhibited OSCC proliferation. Meanwhile, miR-204-5p attenuated OSCC migration in wound healing and transwell assays. Moreover, The levels of N-cadherin, E-cadherin, and vimentin were lowered. In flow cytometry and western blotting, miR-204-5p promoted the apoptosis rate and decreased Bcl-2/Bax ratio. Moreover, elevated LC3-Ⅱ/LC3-Ⅰ ratio suggested miR-204-5p induced autophagy in OSCC. Interestingly, autophagy inhibitor 3-MA partially restored the OSCC’s resistance to apoptosis. Further, miR-204-5p inactivated the PI3K/Akt/mTOR pathway, whereas dual luciferase assay demonstrated PIK3CB was a direct target of miR-204-5p. In vivo, miR-204-5p hindered OSCC growth. This study demonstrated that miR-204-5p inhibited OSCC progression through inducing autophagy-promoted apoptosis by targeting PIK3CB.
Similar content being viewed by others
Introduction
Oral squamous cell carcinoma (OSCC)1 is one of the most common malignant tumors of the head and neck region, contributing to a majority of cancer deaths in Southeast Asian countries1. Surgical resection of primary lesions combined with radiotherapy and chemotherapy are primary therapeutic options in clinical practices. However, OSCC is prone to early metastasis and late diagnosis, leading to poor prognosis in patients with advanced-stage OSCC2. Therefore, exploration of the mechanism behind OSCC and effective biomarkers are urgently required. Non-coding RNA (ncRNA) plays a critical role in physiological development process and diseases, especially in cancer. The research on ncRNAs has considerable prospects in OSCC.
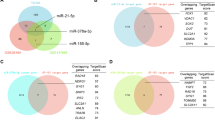
MicroRNA (miRNA), a class of noncoding RNA (19–22 nucleotides in length), suppresses its target genes by inhibiting mRNA translation3. Increasing data suggest that miRNAs regulate cancer behaviors, including proliferation, migration, drug resistance, and apoptosis by interacting with downstream target genes4. In our preliminary study, we identified 20 miRNAs with differential expression in OSCC. Subsequent regression analysis revealed a significant correlation between miR-204-5p and patient prognosis, which led us to select it as a candidate molecule for in-depth investigation in OSCC5.Studies have shown that miR-204-5p accelerates apoptosis and hampers migration by targeting HER-2 in gastric cancer6. In addition, miR-204-5p has similar antitumor effects in breast cancer and esophageal squamous cell carcinoma7,8These aforementioned studies have suggested that miR-204-5p functions as an anti-oncogene.We were intrigued by the unexplored function of miR-204-5p in OSCC. We subsequently characterized a risk model, and bioinformatic analyses (GO and KEGG) predicted phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit beta (PIK3CB) as a potential target gene of miR-204-5p5.The PI3K family is divided into three categories according to the structure and substrate preference. Dysregulation of phosphoinositide 3-kinase (PI3K) and its downstream signals is a common phenomenon in cancer9. PIK3CB is a subtype of class I PI3K that positively regulates PI3K/protein kinase B (AKT)/mammalian target of rapamycin (mTOR) pathway. The PI3K/Akt/mTOR was a classical autophagy pathway which regulates tumor growth and metabolism10,11. In esophageal cancer, PIK3CB acted as an oncogene and promoted the proliferation of cancer cells via stimulating the PI3K/AKT/mTOR pathway12. However, the relationship between miR-204-5p and PI3K/Akt/mTOR axis in OSCC needs further exploration.
Notably, evidence suggests that miRNAs modulate autophagy in cancers13. The indispensable role of autophagy in human diseases has been recognized, particularly in aging and cancer14. Autophagy is a biological process with dual effects—protecting cells and killing cells. Autophagy prevents carcinogenesis by eliminating damaged organelles and misfolded proteins and induces cell death by interacting with multiple biological processes such as apoptosis15. In contrast, autophagy endows cancer cells with the ability to resist various microenvironmental stresses such as hypoxia and energy deficiency, thus protecting cancer cells from chemotherapy16. Autophagy controlled by miRNA has been proved to be a possible breakthrough point in cancer treatment17. Besides, research comfirmed that autophagy induced by inactivation of PI3K/Akt/mTOR pathway could inhibit the malignant progression of ovarian cancer18. However, the effect of autophagy in OSCC needs further exploration. We hypothesized that miR-204-5p inactivates the PI3K/Akt/mTOR pathway by targeting PIK3CB, thus inducing tumor suppressive autophagy to prevent the malignant features of OSCC.
In the present study, we demonstrated miR-204-5p inhibited OSCC proliferation and migration, whereas miR-204-5p promoted apoptosis through inducing autophagy by inactivating PIK3CB-mediated PI3K/Akt/mTOR signal pathway, indicating the therapeutic potential of miR-204-5p in OSCC.
Materials and methods
Cell culture
SCC-15 and SCC-25 cells (the American Type Culture Collection, Manassas, United States) were cultured in DEME-F12 medium (Gibco, NY, United States), and Hacat cells (the Cell Bank of the Chinese Academy of Sciences, Shanghai, China) were cultured in the RPMI-1640 medium (Hyclone, UT, United States). All the cell lines supplemented with 10% fetal bovine serum (FBS), 100 U/mL penicillin, and 0.1 mg/mL streptomycin were cultured in 100-mm culture dishes (JET BIOFIL, China) at 37 °C and in the presence of 5% CO2. Cells were subcultured when they reached 85%–90% confluency, and fourth-generation cells were used for the following experiments.
RNA extraction and quantitative real-time polymerase chain reaction
Under the guidance of the kit manual, total RNA was extracted from cells by TRIzol (Ambion, TX, United States). We used the reverse transcription kit (Sangon Biotech, China) to reverse transcribe the extracted RNA into cDNA. Quantitative PCR was carried out using SYBR green quantitative kit (Sangon Biotech, China). U6 was selected as the internal control to normalize miRNA expressions. The primer sequences were as follows: U6-RT, 5’-GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACAAAATA-3’; U6-F, 5’-AGAGAAGATTAGCATGGCCCCTG-3’; U6-R, 5’-CAGTGCAGGGTCCGAGGT-3’; miR-204-5p-RT, 5’-GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACAGGCAT-3’; miR-204-5p-F, 5’-CGCGTTCCCTTTGTCATCCT-3’; and miR-204-5p-R, 5’-AGTGCAGGGTCCGAGGTATT-3’.The number of biological replicates for each experimental group was three (n = 3), and the relative expression levels were calculated using the 2^(-ΔΔCt) method.
Cell transfection
miR-204-5p mimic (5’-UUCCCUUUGUCAUCCUAUGCCU-3’) and NC (5’-UUUGUACUACACAAAAGUACUG-3’) was designed and synthetized (HiPro Biotechnology Co., Ltd, China). SCC-15 and SCC-25 cell lines were seeded into 6-well plates (Jet Biofil, China) at a density of 2 × 105/well overnight before transfection. miR-204-5p mimic and NC (5 µL/well) were transfected using Lipofectamine 3000 at a concentration of 5 µL/well (Invitrogen, CA, United States) when cell confluence reached 70%. Subsequent experiments were performed 24 h after transfection. miR-204-5p expression after transfection was detected by quantitative real-time polymerase chain reaction (qRT-PCR), and the primer sequences were as previously described. The efficiency of transfection was estimated by qRT-PCR. To ensure that each experiment was performed under basically the same transfection efficiency, we added Cy5 red fluorescence (HiPro Biotechnology Co., Ltd, China) to the mimic and NC. The following experiments were conducted after confirming the transfection efficiency under the microscope.
CCK-8 assay
The transfected cells were seeded into 96-well plates (Jet Biofil, China) at a density of 4 × 103/well. After cell adherence was observed using a microscope, CCK-8 reagent (ApexBio, TX, United States) was added to each well (10 µL/well) and incubated at 37 °C for 2 h. A microplate reader was used to detect OD value at 450 nm. The first results were marked on day 0, and the following on days 1, 3, and 5; the aforementioned operations were repeated to obtain data of cell proliferation capacity at different time points. The final data were visualized by drawing a line chart with GraphPad Prism 7.
Clone formation assay
Cells that had been transfected for 24 h were digested and seeded into a 6-well plate with a density of 5 × 102/well. The culture medium were changed every 3 days. The culture was ended after about 2 weeks. Cells were gently cleaned twice with PBS, and fixed in 4% paraformaldehyde solution for 20 min, stained with crystal violet for 30 min. The excess dye was washed by PBS. The individual colonies was pictured and analysed under a microscope.
Edu assay
The transfected cells were seeded into 6-well plates at a density of 2 × 105/well and cultured overnight until the cell confluence reached 70%. The cells were mixed with diluted 1× Edu (Beyotime Biotechnology, China) working solution and incubated at 37 °C for 4 h. Afterward, the cells were fixed with 4% paraformaldehyde solution for 20 min according to the manufacturer’s instructions, permeabilized with 0.3% Triton X-100 for 30 min, incubated with the configured click reaction solution in the dark for 30 min, and stained with Hochest. After washing the excess dye with PBS, the sample was placed under an inverted fluorescence microscope (Zeiss, Germany) for visualization and determining statistics.Quantitative analysis was performed using ImageJ software. First, total nuclei were segmented based on the Hoechst staining channel. Subsequently, EdU-positive nuclei were identified in the corresponding EdU signal channel using a consistent fluorescence intensity threshold. The cell proliferation rate was calculated as the percentage of EdU-positive nuclei relative to the total number of nuclei (EdU-positive nuclei / total nuclei × 100%).
Wound healing assay
The transfected cells were seeded into 6-well plates at a density of 4 × 105/well and cultured overnight until the cell confluence reached 90%. Wounds were created using 200-µL tips perpendicular to the 6-well plates. The wound condition was visualized under an inverted microscope and recorded as 0 h. Cells were cultured at 37 °C in the presence of 5% CO2 for 24 h, and the cell migration status at the same sites was visualized and recorded.The images were analyzed using ImageJ software, where the wound region was manually outlined and its area was measured. The wound healing rate was calculated using the formula: [1 - (Area at 24 h / Area at 0 h)] × 100%.
Transwell assay
Sterile Transwell chambers (Corning, NY, United States) were placed in 24-well plates, and 100 µL of serum-free medium was added to the upper chamber to activate the filter membrane for 10 min. The transfected cells were digested and diluted by serum-free medium after counting to adjust the cell density to 4 × 104/100 µL. Next, 100 µL of cell suspension was added to the upper chamber, and 800 µL of medium containing 10% FBS was added to the lower chamber. Cells were cultured at 37 °C in the presence of 5% CO2 for 24 h. After fixing the cells with 4% paraformaldehyde, the cells were subjected to crystal violet staining for 15 min. The adhered cells in the upper chamber were gently wiped off with a cotton swab. Cell migration was visualized using an inverted microscope and the number of cells was counted.
Flow cytometry
The transfected cells were seeded into 6-well plates at a density of 1 × 106/well and cultured for 24 h. According to the manufacturer’s instructions, the cells were incubated in Annexin V-fluorescein isothiocyanate/propidium iodide in the dark by using the apoptosis detection kit (4 A Biotech, China). The prepared samples were added to the flow cytometer (Beckman Coulter Diagnostics, CA, United States) for analysis. The obtained data were processed in CytExpert software to calculate the early and late apoptosis rates.
Western blotting
Proteins were isolated from cells using the RIPA kit (Beyotime, China). Sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) was used to separate proteins with different molecular weights. After electrophoresis, the methanol-activated polyvinylidene fluoride (PVDF) membrane (Millipore, MA, United States) was used for protein transfer. Sponge-filter paper-glue-PVDF membrane-filter paper-sponge was placed in order from the negative electrode to the positive electrode. Protein-loaded PVDF membranes were blocked in milk for 2 h at room temperature and then incubated overnight at 4 °C with specific primary antibodies. The PVDF membrane was washed thrice with TBST solution the next morning (10 min each time) and incubated with the corresponding secondary antibody for 1 h at room temperature. The PVDF membrane was washed thrice again with TBST (10 min each time) to remove excess antibodies. The protein content was detected using the luminescent solution kit (4 A Biotech, China) and the luminometer (Bio-Rad Laboratories, CA, United States). Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) was selected as the internal control to calculate protein expression levels.Quantification was performed using ImageJ software. First, local background correction was applied to each band, and then its integrated density value was measured. The expression level of the target protein was normalized by calculating the ratio of its integrated density value to that of the corresponding loading control protein (GAPDH). Using the normalized average value of the control group as the baseline, the relative expression levels of each experimental group were calculated.
Dual luciferase report assay
Wild-type (WT) and mutant (MUT) plasmids designed according to the binding site of miR-204-5p and PIK3CB were constructed (SyngenTech Co., Ltd, China). The plasmids were co-transfected into SCC-15 and SCC-25 cells with the miR-204-5p mimic and NC by using Lipofectamine 3000 (the specific transfection method is as described previously). After 48 h of transfection, the cells were lysed, and fluorescent signals were detected using the dual luciferase report detection kit (Vazyme, China). The sea kidney luciferase reaction intensity was selected as the internal reference for calculation and statistical analysis.
Tumor model
Four-week-old female nude mice were divided into two groups, with six mice in each group. miR-204-5p mimic and NC were transfected into SCC-15 cells, whereas nude mice were subjected to adaptive feeding for a week. Then, 1 × 107 transfected SCC-15 cells were injected into the left flank of each nude mouse. The nude mice were euthanized by intraperitoneal injection of an overdose of pentobarbital sodium (150 mg/kg) 16 days later, and the tumor volumes were measured. Tumors were subsequently used to extract samples for western blotting, qRT-PCR, and immunohistochemistry. The tumor volume was calculated using the following formula: V = ab2/2 (where a is the long axis and b is the short axis).
Statistical analysis
Image J software was used for image analysis in this study. For data analysis, GraphPad Prism software was used for testing the significant difference and plotting figures. The data were expressed as the mean ± standard deviation and charts were drawn. In this study, a p value < 0.05 was considered to be statistically significant. Adobe Illustrator was used for splicing the figures and charts.
Results
miR-204-5p was downregulated in OSCC
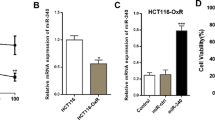
qRT-PCR was used to detect miR-204-5p level in different cell lines. Hacat cell line represented normal epithelial cells that showed an apparently higher miR-204-5p expression than SCC-15 and SCC-25 OSCC cell lines (Fig. 1A). Because of the low expression of miR-204-5p in OSCC cells, we artificially elevated miR-204-5p expression to explore its impact on OSCC. miR-204-5p mimic and NC were transfected into SCC-15 and SCC-25 using Lipofectamine 3000 for 24 h. The transfection efficiency was determined by qRT-PCR (Fig. 1B). To ensure that the cells were effectively transfected, we added red fluorescence to the NC and mimic. Only cells that achieved transfection efficiency (the group successfully transfected with miR-204-5p mimic in the following content is referred to as the mimic group for short) were used for experiments (Figure S1).
LThe miR-204-5p level in different cell lines. A. miR-204-5p was downregulated in the SCC-15 and SCC-25 (OSCC cell lines) when compared with Hacat (normal epithelial cell line). B. miR-204-5p was apparently elevated in SCC-15 and SCC-25 after transfection.
miR-204-5p inhibited proliferation of OSCC in vitro
The impact of miR-204-5p on OSCC proliferation was determined through the CCK-8 assay, clone formation assay, and EdU assay. As shown in Fig. 2A, the SCC-15 cells transfected with miR-204-5p mimic displayed lower viability on 5th day than those transfected with NC. Similarly, On the 3rd and 5th days, the SCC-25 cell viability of the mimic group was significantly lower than that of the NC group. In the clone formation assay, the number of individual clones formed within 2 weeks in the mimic group was also significantly lower than that in the NC group (Fig. 2B). This suggested that miR-204-5p could weaken the proliferation ability of OSCC individual cells. In the Edu assay, Edu dyes with green fluorescence could be embedded in DNA during cell division. Our results revealed that the combination rate with EdU was significantly lower in the cells treated with the mimic, indicating miR-204-5p impaired cell division capacity of OSCC (Fig. 2C). According to the results, artificially elevated miR-204-5p expression obviously weakened the proliferation ability of OSCC.
miR-204-5p inhibited proliferation of OSCC in vitro. A. CCK-8 showed miR-204-5p impaired cell viability of OSCC. B. Clone formation assay showed miR-204-5p significantly lower the number of individual clones formed by OSCC cells. C. Edu assay demonstrated miR-204-5p hampered proliferation of OSCC cells.
miR-204-5p inhibited migration of OSCC in vitro
Because the poor prognosis of OSCC is largely related to lymph node metastasis, we further evaluated the impact of miR-204-5p on OSCC migration capacity. The wound healing assay revealed that miR-204-5p evidently slowed down the wound healing process compared with NC in both SCC-15 and SCC-25 cell lines (Fig. 3A). In the transwell assay, cells treated with miR-204-5p exhibited a lower membrane passing rate in the transwell chamber, indicating a weaker migration ability (Fig. 3B). We also used western blotting simultaneously to detect molecular changes in the NC and mimic groups. The epithelial–mesenchymal transition (EMT) markers N-cadherin and vimentin were downregulated, whereas E-cadherin was upregulated after miR-204-5p transfection (Fig. 3C). These results proved that the increased miR-204-5p expression could inhibit the migration of OSCC cells.
miR-204-5p inhibited migration of OSCC in vitro. A. Wound healing assay revealed miR-204-5p evidently weakened the wound healing process in OSCC. B. Transwell assay showed miR-204-5p hindered OSCC cells to cross the transwell chamber membrane. C. Western blotting showed miR-204-5p lowered N-cadherin/E-cadherin ratio and Vimentin level in OSCC, which indicated damaged EMT.
miR-204-5p inactivated the PI3K/Akt/mTOR pathway by targeting PIK3CB
Accumulating evidence suggests that the miRNA sponges mRNA of target gene through the mechanism of competitive endogenous RNA (ceRNA). We predicted the binding sites of miR-204-5p and PIK3CB through the TargetScan online database (Fig. 4A). To further prove the underlying mechanism of miR-204-5p inhibiting PIK3CB in OSCC, the dual luciferase report assay was performed. The WT contained the presumed binding site of miR-204-5p while MUT didn’t. WT and MUT were cloned into the report plasmids, and were co-transfected into OSCC cells with the NC and mimic. The luciferase activity in the WT + miR-204-5p group was obviously reduced compared to the WT + NC group, whereas that in the MUT + miR-204-5p group did not show any significant change compared to the MUT + NC group (Fig. 4B). This result confirmed that miR-204-5p directly bound to PIK3CB through the sites in Fig. 4A, thereby inhibiting PIK3CB expression. Further, we determined the expression changes of PI3K/Akt/mTOR signal molecules after transfection through western blotting. The expressions of p-PI3K, p-Akt, and p-mTOR were suppressed in the mimic group, whereas PI3K, Akt and mTOR in both groups showed similar expressions, representing inhibition of the PI3K/Akt/mTOR pathway (Fig. 4C). According to these results, we concluded that miR-204-5p inactivated PI3K/Akt/mTOR pathway by directly targeting PIK3CB in OSCC.
miR-204-5p inactivated PI3K/Akt/mTOR pathway by targeting PIK3CB. A. The binding site of miR-204-5p in the 3’-UTR of PIK3CB predicted by TargetScan. B. The dual luciferase report assay showed the relative luciferase activitie of OSCC cells co-transfected with different reporter plasmids (WT or MUT) and NC or mimic. C. miR-204-5p inactivated PIK3/AKT/mTOR pathway in OSCC.
miR-204-5p promoted apoptosis in OSCC by inducing autophagy
PI3K/Akt/mTOR is a critical pathway modulating autophagy, and the crosstalk between autophagy and apoptosis has great impact on cancer development. We further explored the effect of miR-204-5p on autophagy and apoptosis in OSCC. The levels of autophagy in the NC and mimic groups were measured by western blotting. The p62 level was decreased while microtubule associated protein 1 light chain 3 beta-Ⅱ (LC3-Ⅱ)/microtubule associated protein 1 light chain 3 beta-Ⅰ (LC3-Ⅰ) ratio was increased significantly in the mimic group (Fig. 5A), indicating that miR-204-5p enhanced autophagy in OSCC. Further, we assessed the apoptosis status in SCC-15 and SCC-25 cell lines to interpret the anticancer role of miR-204-5p. Flow cytometry showed that the apoptosis rate of OSCC cells was significantly increased after miR-204-5p overexpressing (Fig. 5B). Moreover, the expression level of B-cell lymphoma/leukemia-2 (Bcl-2) was markedly lowered, whereas that of Bcl-2 Associated X (Bax) was increased in the mimic group, showing that apoptosis increased after miR-204-5p upregulation (Fig. 5A). Accordingly, the tumor-suppressive role of miR-204-5p might be correlated with enhanced autophagy and apoptosis via inactivation of the PI3K/Akt/mTOR pathway.
miR-204-5p promoted apoptosis in OSCC by inducing autophagy. A, B. miR-204-5p promoted autophagy and apoptosis in both OSCC cell lines. C, D. The autophagy inhibitor 3-MA blocked the autophagy promoted by miR-204-5p, and partially restored the apoptosis tolerance of OSCC cells.
To determine whether miR-204-5p-induced apoptosis was related to autophagy, the autophagy inhibitor 3-methyladenine (3-MA) was introduced. 3-MA with the final concentration of 1 mmol was added to the mimic group after transfection for 24 h. Western blotting assay showed that 3-MA could effectively inhibit miR-204-5p-induced autophagy as the p62 level was restored while the LC3-Ⅱ/LC3-Ⅰ ratio was downregulated in the mimic + 3-MA group compared with those in the mimic group (Fig. 5C). Interestingly, the apoptosis marker Bcl-2/Bax ratio was also recovered, suggesting that 3-MA weakened the miR-204-5p-induced apoptosis effect (Fig. 5C). Moreover, flow cytometry intuitively showed that miR-204-5p-induced apoptosis was significantly attenuated after the addition of 3-MA (Fig. 5D). Thus, our results showed that miR-204-5p promoted apoptosis in OSCC via enhanced autophagy that was regulated by the PI3K/Akt/mTOR pathway.
miR-204-5p inhibited tumorigenesis of OSCC in vivo
We constructed a xenograft model in nude mice to determine whether miR-204-5p could inhibit OSCC tumorigenesis in vivo. SCC-15 cells transfected with NC or mimic were injected into the left flank of the nude mice. The tumor size (volume) in the mimic group was significantly lower than that in the NC group (Fig. 6A, B). Specifically, the tumor volumes in the NC group were 260.38, 243.71, 244.43, 228.99, 225.06, and 209.93 mm³, whereas those in the mimic group were 147.19, 169.85, 141.58, 136.13, 116.61, and 123.81 mm³. Next, we used qRT-PCR to detect miR-204-5p expression in the two groups. Tumors in the NC group exhibited lower miR-204-5p expression (Fig. 6C). Moreover, immunohistochemical and western blotting assays were used to compare the PIK3CB level. In the mimic group, PIK3CB was downregulated, re-verifying the inhibitory effect of miR-204-5p on PIK3CB in OSCC (Fig. 6D, E). In ki-67 staining, the staining degree of tumor tissue in the mimic group was lower, indicating that miR-204-5p decreased the proliferation index of OSCC (Fig. 6F). In summary, our data showed that miR-204-5p could inhibit OSCC tumorigenesis in vivo.
miR-204-5p inhibited tumorigenesis of OSCC in vivo. A, B. Inhibitory effect of miR-204-5p on OSCC tumor volume. C. The relative miR-204-5p expression level was determined by qRT-PCR. D, E. Western blotting and immunohistochemistry showed PIK3CB was downregualted in the OSCC tumor tissues transfected with miR-204-5p. F. After miR-204-5p transfection, OSCC tumor tissues exhibited a lower ki-67 staining degree.
Discussion
OSCC is one of the largest contributors to cancer deaths in Southeast Asian countries. Satisfactory outcomes are rarely achieved with the current treatment options. Therefore, it is essential to identify effective biomarkers for new treatment methods or drugs to improve the outcomes of patients with OSCC.
miRNA is a distinctive regulator in cancers19. ncRNAs, including long non-coding RNA (lncRNA), circular RNA (circRNA), and miRNA share a special signaling axis to regulate the biological behaviors of cancer, in which lncRNA and circRNA sponge miRNA, whereas miRNA inhibits the expression of its target gene by blocking mRNA translation. Such regulation is called ceRNA20. Previous bioinformatics studies have filtered ncRNAs that might be the prognostic targets in OSCC; however, their practical roles remain to be experimentally validated21. In addition, Changes of biomarkers in saliva have been helpful in head and neck squamous carcinoma (HNSCC)22,23. Determining the level of miR-204-5p in saliva might be a noninvasive method for early screening of OSCC and predicting the prognosis of patients with OSCC.
miR-204-5p was considered to be a tumor suppressor in previous cancer researches. For example, miR-204-5p suppressed proliferation and enhanced apoptosis and chemosensitivity in the HCT116 cell line24. Moreover, miR-204-5p restrained bone metastasis in prostate cancer by inactivating the nuclear factor kappa-B (NF-kb) axis25. However, the role of miR-204-5p in OSCC has not been sufficiently studied. To examine the potential mechanism of miR-204-5p in OSCC, GO and KEGG enrichment analyses were performed8. GO results showed that apoptotic process, cell migration, and mRNA binding were the significantly enriched biological processes, indicating that miR-204-5p possibly modulates OSCC through modulate apoptosis, migration, and downstream target genes. The PI3K/Akt/mTOR pathway and autophagy were significantly enriched KEGG26,27 terms, a strong signal suggesting that miR-204-5p controlled autophagy mediated by the PI3K/Akt/mTOR pathway in OSCC. Besides, PIK3CB (a member of the PI3K family) was predicted to be a target gene of miR-204-5p in three miRNA databases. Because bioinformatics analysis and qRT-PCR results showed that miR-204-5p expression was low in OSCC, we artificially overexpressed miR-204-5p by transfecting with the mimic to explore its role in OSCC.
Uncontrolled proliferation primarily accounts for the rapid progression of OSCC28. In our study, CCK-8, Edu, and clone formation assays showed that miR-204-5p significantly inhibited the proliferation of OSCC cells in vitro. Moreover, the in vivo xenograft model confirmed that the tumorigenesis of OSCC cells was impaired by miR-204-5p, as the tumor volumes and degree of ki-67 staining in the mimic group were evidently lower than those in the NC group. These data indicated that an artificial increase in miR-204-5p expression may slow down the progression of OSCC. EMT endows epithelial cells with reduced adhesion and increased mobility, thus making cancer cells more metastatic. In this process, the level of the epithelial cell marker E-cadherin decreases, whereas those of the mesenchymal cell markers N-cadherin and vimentin increase29. Metastasis promoted by EMT often results in poor OSCC prognosis30. In this study, the wound healing ability of OSCC cells with high miR-204-5p expression was significantly attenuated in the wound healing assay. Moreover, in the transwell assay, the permeability of OSCC cells to the chamber membrane was apparently repressed with miR-204-5p transfection. Concurrently, low N-cadherin and vimentin levels and high E-cadherin level indicated that the EMT level decreased with the upregulation of miR-204-5p in OSCC. These findings verified that miR-204-5p weakened EMT to inhibit OSCC, and miR-204-5p might be the key factor in preventing OSCC from early metastasis. Achieving apoptosis, a programmed cell death process, is the ultimate goal of many cancer treatments. Unfortunately, tolerance to apoptosis leads to unfavorable results in cancers with drug-resistant cells31. Recent research revealed that miRNAs could both promote and inhibit apoptosis in OSCC32,33. In the present study, flow cytometry and western blotting assays were used to evaluate how miR-204-5p affects apoptosis in OSCC. According to the results, elevated miR-204-5p promoted the apoptosis rate of OSCC cells, and the apoptosis-related protein Bcl-2/Bax ratio was decreased. Therefore, we concluded that miR-204-5p could promote apoptosis in OSCC. Overall, the aforementioned data demonstrated that miR-204-5p inhibited OSCC progression by inducing apoptosis, suppressing both proliferation and EMT. This outcome provides us information about the therapeutic value of miR-204-5p in OSCC.
To further explore the mechanism behind miR-204-5p inhibiting OSCC, we predicted and attested the downstream target gene PIK3CB and the related PI3K/Akt/mTOR pathway. The PI3K/Akt/mTOR pathway, a crucial signaling pathway in human cancers, has also been studied in OSCC. For instance, MingBo Wei et al. proved that genipin inhibited OSCC by inactivating the PI3K/Akt/mTOR pathway34. Similarly, PI3K inhibitors including PI-103, PI-828, and PX-866 inhibited OSCC proliferation and invasion, providing us with new insights into OSCC therapy35. Inactivation of the PI3K/Akt/mTOR pathway may be an effective strategy in OSCC treatment, yet the mechanism of action of specific miRNA on the PI3K/Akt/mTOR pathway in OSCC is poorly understood. PIK3CB, an initiator of the PI3K/Akt/mTOR pathway36, was a direct target of miR-204-5p in our bioinformatics analysis. Evidence suggests that PIK3CB functions as an oncogene in multiple human cancers12,37, which might be a potentially critical link between miR-204-5p and the PI3K/Akt/mTOR pathway in OSCC. Our western blotting results showed that in the OSCC cell lines, p-PI3K/PI3K, p-Akt/Akt, and p-mTOR/mTOR ratios evidently declined after miR-204-5p transfection. The decrease in the degree of phosphorylation of each component suggested that miR-204-5p inhibited the PI3K/Akt/mTOR pathway in OSCC. To prove that miR-204-5p directly inhibits PIK3CB, we performed the dual luciferase report assay. The luciferase activity of WT and mimic co-transfection group was significantly lower, whereas there was no significant reduction in luciferase activity in the MUT and mimic co-transfection groups, implying that miR-204-5p directly inhibited PIK3CB via binding to the 3’-UTR of PIK3CB. This is consistent with findings from Hong, B. S. et al., who, using a luciferase reporter assay, identified PIK3CB as a direct target of miR-204-5p. They further demonstrated that ectopic expression of this miRNA reduced both PIK3CB mRNA and protein levels, thereby suppressing PI3K/Akt signaling in breast cancer cells38. Moreover, in western blotting, PIK3CB level was significantly downregulated in the mimic group. Further, we applied qRT-PCR, western blotting, and immunohistochemistry in the xenograft model. A high miR-204-5p expression accompanied by a low PIK3CB level was detected in the mimic group, manifesting that miR-204-5p inhibited PIK3CB in vivo. Our results showed that miR-204-5p inhibited OSCC via inactivation of the PI3K/Akt/mTOR pathway by targeting PIK3CB. In recent years, autophagy has become a hotspot in cancer research. Autophagy regulated by non-coding RNAs in OSCC has also been studied. Upregulation of lncRNA cancer susceptibility candidate 9 (CASC9) accelerated OSCC progression by blocking autophagy-mediated apoptosis via the Akt/mTOR pathway39. In contrast, under hypoxia environment, the circRNA circCDR1as endowed OSCC cells with reinforced survival ability by inducing autophagy via the Akt/extracellular signal-regulated kinase 1/2 (ERK1/2)/mTOR axis40. These contrary results impelled us to explore whether miR-204-5p could change the autophagy level of OSCC by regulating the PI3K/Akt/mTOR pathway, thus affecting OSCC progression. The results showed that after the overexpression of miR-204-5p, the inhibition of the PI3K/Akt/mTOR pathway was accompanied by a decrease in the p62 level and an increase in the LC3-Ⅱ/LC3-Ⅰ ratio, indicating the elevation of autophagy. We hypothesized that miR-204-5p inhibited OSCC progression via inducing autophagy mediated by the PI3K/Akt/mTOR pathway. For further confirmation, we treated cells with 3-MA for 24 h. The autophagic flux activated by miR-204-5p was blocked, as the p62 level recovered and the LC3-Ⅱ/LC3-Ⅰ ratio decreased. We further evaluated the apoptosis status of OSCC cells through western blotting and flow cytometry. Of note, when 3-MA inhibited autophagy, the resistance of OSCC cells to apoptosis was proved to be partially restored, as Bcl2/Bax rebounded and the overall apoptosis rate decreased. These data supported the view that miR-204-5p inhibited OSCC by promoting apoptosis of OSCC cells through autophagy activation mediated by the PI3K/Akt/mTOR pathway.
In summary, we identified miR-204-5p and miR-503-3p as critical diagnostic and prognostic biomarkers in OSCC and provided a prognostic signature. More importantly, we demonstrated the therapeutic value of miR-204-5p in promoting apoptosis and inhibiting proliferation and EMT-induced migration in OSCC. Additionally, miR-204-5p promoted apoptosis through inducing PI3K/Akt/mTOR-mediated autophagy by directly targeting PIK3CB, which may enable early detection, clinical management, and development of new therapies for OSCC.This study, however, has two main limitations. First, although we have obtained strong evidence from the luciferase reporter assay and pathway analysis, we lack rescue experiments to confirm PIK3CB’s role as the key mediator. Second, direct evidence from RNA pull-down assays or loss-of-function studies using a miR-204-5p inhibitor is absent. Thus, these verifications represent the primary aim of our future investigations.
Data availability
The datasets generated during and/or analyzed during this study are available from the corresponding author upon reasonable request.
Abbreviations
- 3-MA:
-
3-Methyladenine
- AUC:
-
Area under the ROC curve
- Bcl-2:
-
B cell lymphoma/leukemia-2
- Bax:
-
Bcl-2 Associated X
- CASC9:
-
Cancer susceptibility candidate 9
- circRNA:
-
Circular RNA
- CeRNA:
-
Competitive endogenous RNA
- ERK1/2:
-
Extracellular signal-regulated kinase 1/2
- FBS:
-
Fetal bovine serum
- GO:
-
Gene Ontology
- GAPDH:
-
Glyceraldehyde-3-phosphate dehydrogenase
- HNSCC:
-
Head and neck squamous carcinoma
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- lncRNA:
-
Long non-coding RNA
- Mtor:
-
Mammalian target of rapamycin
- miRNA:
-
MicroRNA
- LC3-I:
-
Microtubule associated protein 1 light chain 3 beta-I
- LC3-II:
-
Microtubule associated protein 1 light chain 3 beta-II
- MAPK:
-
Mitogen-activated protein kinase
- MUT:
-
Mutant
- ncRNA:
-
Non-coding RNA
- NF-kb:
-
Nuclear factor kappa-B
- OSCC:
-
Oral squamous cell carcinoma
- OS:
-
Overall survival
- PIK3CB:
-
Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit-beta
- PI3K:
-
Phosphoinositide 3-kinase
- PVDF:
-
Polyvinylidene fluoride
- AKT:
-
Protein kinase B
- qRT-PCR:
-
Quantitative real-time polymerase chain reaction
- Ras:
-
Rat sarcoma
- ROC:
-
Receiver operating characteristic
- SDS-PAGE:
-
Sodium dodecyl sulphate polyacrylamide gel electrophoresis
- TCGA:
-
The Cancer Genome Atlas
- WT:
-
Wild-type
References
Almangush, A. et al. Staging and grading of oral squamous cell carcinoma: an update. Oral Oncol. 107, 104799. https://doi.org/10.1016/j.oraloncology.2020.104799 (2020).
Joseph, J. P., Harishankar, M. K., Pillai, A. A. & Devi, A. Hypoxia induced EMT: a review on the mechanism of tumor progression and metastasis in OSCC. Oral Oncol. 80, 23–32. https://doi.org/10.1016/j.oraloncology.2018.03.004 (2018).
Fang, T. et al. Tumor-derived Exosomal miR-1247-3p induces cancer-associated fibroblast activation to foster lung metastasis of liver cancer. Nat. Commun. 9, 191. https://doi.org/10.1038/s41467-017-02583-0 (2018).
Pan, G., Liu, Y., Shang, L., Zhou, F. & Yang, S. EMT-associated MicroRNAs and their roles in cancer stemness and drug resistance. Cancer Commun. 41, 199–217. https://doi.org/10.1002/cac2.12138 (2021).
Gou, Q. T. et al. Bioinformatic analysis and experimental verification of prognostic MiRNA in oral squamous cell carcinoma. Mod. Oncol. 31, 2988–2995. https://doi.org/10.3969/j.issn.1672-4992.2023.16.008 (2023).
Yang, S. et al. miR-204-5p promotes apoptosis and inhibits migration of gastric cancer cells by targeting HER-2. Mol. Med. Rep. 22, 2645–2654. https://doi.org/10.3892/mmr.2020.11367 (2020).
Su, Q., Shen, H., Gu, B. & Zhu, N. miR-204-5p hampers breast cancer malignancy and affects the cell cycle by targeting PRR11. Comput. Math. Methods Med. 4010947. https://doi.org/10.1155/2022/4010947(2022) (2022).
Luo, H. et al. miR-204-5p inhibits cell proliferation and induces cell apoptosis in esophageal squamous cell carcinoma by regulating Nestin. Int. J. Med. Sci. 19, 472–483. https://doi.org/10.7150/ijms.67286 (2022).
Yang, J. et al. Targeting PI3K in cancer: mechanisms and advances in clinical trials. Mol Cancer 18, 26, https://doi.org/10.1186/s12943-019-0954-x(2019).
Yang, H. et al. Histocompatibility minor 13 (HM13), targeted by miR-760, exerts oncogenic role in breast cancer by suppressing autophagy and activating PI3K-AKT-mTOR pathway. Cell Death Dis 13, 728, https://doi.org/10.1038/s41419-022-05154-4(2022).
Liao, S. et al. CD38 is involved in cell energy metabolism via activating the PI3K/AKT/mTOR signaling pathway in cervical cancer cells. Int. J. Oncol. 57, 338–354. https://doi.org/10.3892/ijo.2020.5040 (2020).
Xu, W., Wang, Z., Zhang, Z., Xu, J. & Jiang, Y. PIK3CB promotes oesophageal cancer proliferation through the PI3K/AKT/mTOR signalling axis. Cell. Biol. Int. 46, 1399–1408. https://doi.org/10.1002/cbin.11847 (2022).
Long, J. et al. The effect of MiRNA and autophagy on colorectal cancer. Cell. Prolif. 53, e12900. https://doi.org/10.1111/cpr.12900 (2020).
Levine, B. (ed, G.) Biological functions of autophagy genes: a disease perspective. Cell 176 11–42 https://doi.org/10.1016/j.cell.2018.09.048 (2019).
Song, S., Tan, J., Miao, Y., Li, M. & Zhang, Q. Crosstalk of autophagy and apoptosis: involvement of the dual role of autophagy under ER stress. J. Cell. Physiol. 232, 2977–2984. https://doi.org/10.1002/jcp.25785 (2017).
Lin, C. et al. Encoding gene RAB3B exists in linear chromosomal and circular extrachromosomal DNA and contributes to cisplatin resistance of hypopharyngeal squamous cell carcinoma via inducing autophagy. Cell Death Dis 13, 171, https://doi.org/10.1038/s41419-022-04627-w(2022).
Wu, H. et al. MIR145-3p promotes autophagy and enhances bortezomib sensitivity in multiple myeloma by targeting HDAC4. Autophagy 16, 683–697. https://doi.org/10.1080/15548627.2019.1635380 (2020).
Zhou, J., Jiang, Y. Y., Chen, H., Wu, Y. C. & Zhang, L. Tanshinone I attenuates the malignant biological properties of ovarian cancer by inducing apoptosis and autophagy via the inactivation of PI3K/AKT/mTOR pathway. Cell. Prolif. 53, e12739. https://doi.org/10.1111/cpr.12739 (2020).
Hill, M. & Tran, N. MiRNA interplay: mechanisms and consequences in cancer. Dis. Model. Mech. 14, dmm049079. https://doi.org/10.1242/dmm.047662 (2021).
Han, T. S., Hur, K., Cho, H. S. & Ban, H. S. Epigenetic associations between lncRNA/circRNA and MiRNA in hepatocellular carcinoma. Cancers 12, 3292. https://doi.org/10.3390/cancers12092622 (2020).
Liu, Z. et al. Multiomics immune-related LncRNA analysis of oral squamous cell carcinoma and its correlation with prognosis. Dis. Markers. 6106503. https://doi.org/10.1155/2022/6106503(2022) (2022).
Roi, A. et al. The challenges of OSCC diagnosis: salivary cytokines as potential biomarkers. J. Clin. Med. 9, 2865. https://doi.org/10.3390/jcm9092866 (2020).
Kang, J. W., Eun, Y. G. & Lee, Y. C. Diagnostic value of salivary MiRNA in head and neck squamous cell cancer: systematic review and meta-analysis. Int. J. Mol. Sci. 22, 7026. https://doi.org/10.3390/ijms22137026( (2021).
Yao, S. et al. Exosome-mediated delivery of miR-204-5p inhibits tumor growth and chemoresistance. Cancer Med. 9, 5989–5998. https://doi.org/10.1002/cam4.3248 (2020).
Wa, Q. et al. miR-204-5p represses bone metastasis via inactivating NF-κB signaling in prostate cancer. Mol Ther. Nucleic Acids 18, 567–579, https://doi.org/10.1016/j.omtn.2019.09.008(2019).
Kanehisa, M., Sato, Y., Kawashima, M., Furumichi, M. & Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 44 https://doi.org/10.1093/nar/gkv1070 D457–D462 (2016).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30. https://doi.org/10.1093/nar/28.1.27 (2000).
He, S., Chakraborty, R. & Ranganathan, S. Proliferation and apoptosis pathways and factors in oral squamous cell carcinoma. Int. J. Mol. Sci. 23, 1481. https://doi.org/10.3390/ijms23031562( (2022).
Derynck, R. & Weinberg, R. A. EMT and cancer: more than Meets the eye. Dev Cell 49, 313–316, https://doi.org/10.1016/j.devcel.2019.04.026(2019).
Ling, Z., Cheng, B. & Tao, X. Epithelial-to-mesenchymal transition in oral squamous cell carcinoma: challenges and opportunities. Int. J. Cancer. 148, 1548–1561. https://doi.org/10.1002/ijc.33352 (2021).
Carneiro, B. A. & El-Deiry, W. S. Targeting apoptosis in cancer therapy. Nat Rev. Clin. Oncol 17, 395–417, https://doi.org/10.1038/s41571-020-0341-y(2020).
Wang, J. et al. miR-195 inhibits proliferation and enhances apoptosis of OSCC cells via targeting TLR4. J. Healthc. Eng. 2270777 https://doi.org/10.1155/2022/2270777(2022) (2022).
Bao, W. W. et al. MiR-590-5p regulates cell proliferation, apoptosis, migration and invasion in oral squamous cell carcinoma by targeting RECK. Histol. Histopathol. 36, 355–365. https://doi.org/10.14670/HH-18-306( (2021).
Wei, M., Wu, Y., Liu, H. & Xie, C. Genipin induces autophagy and suppresses cell growth of oral squamous cell carcinoma via PI3K/AKT/MTOR pathway. Drug Des. Devel Ther. 14, 395–405. https://doi.org/10.2147/DDDT.S222694 (2020).
Aggarwal, S. et al. Targeted disruption of PI3K/Akt/mTOR signaling pathway, via PI3K inhibitors, promotes growth inhibitory effects in oral cancer cells. Cancer Chemother. Pharmacol. 83, 451–461. https://doi.org/10.1007/s00280-018-3746-x (2019).
Cen, B. et al. An efficient bivalent Cyclic RGD-PIK3CB SiRNA conjugate for specific targeted therapy against glioblastoma in vitro and in vivo. Mol. Ther. Nucleic Acids. 13, 220–232. https://doi.org/10.1016/j.omtn.2018.09.002 (2018).
Mazloumi Gavgani, F. et al. Class I phosphoinositide 3-kinase PIK3CA/p110α and PIK3CB/p110β isoforms in endometrial cancer. Int. J. Mol. Sci. 19, 3931. https://doi.org/10.3390/ijms19123931 (2018).
Hong, B. S. et al. Tumor suppressor miRNA-204-5p regulates growth, metastasis, and immune microenvironment remodeling in breast cancer. Cancer Res. 79, 1520–1534. https://doi.org/10.1158/0008-5472.CAN-18-0891( (2019).
Yang, Y., Chen, D., Liu, H. & Yang, K. Increased expression of LncRNA CASC9 promotes tumor progression by suppressing autophagy-mediated cell apoptosis via the AKT/mTOR pathway in oral squamous cell carcinoma. Cell. Death Dis. 10 (41). https://doi.org/10.1038/s41419-018-1280-8 (2019).
Gao, L. et al. CircCDR1as upregulates autophagy under hypoxia to promote tumor cell survival via AKT/ERK(½)/mTOR signaling pathways in oral squamous cell carcinomas. Cell Death Dis 10, 745, https://doi.org/10.1038/s41419-019-1971-9(2019).
Acknowledgements
We thank Professor Jingang Xiao and Professor Yandong Mou for the donated cells. We also thank Yanmei Chen and Yiren Yang for their good discussion.
Funding
This work was supported by Luzhou Science and Technology Bureau (Grant No. 2021-JYJ-68) and Southwest Medical University Technology Program (Grant No. 2021ZKMS014).
Author information
Authors and Affiliations
Contributions
Formal analysis, Wenke Jin; Funding acquisition, Haixia Huang; Investigation, Zixiang Li; Methodology, Qiang Xie; Project administration, Haixia Huang; Software, Qiutong Gou; Supervision, Qiang Xie; Validation, Chaopei Chen; Writing – original draft, Chaopei Chen; Writing – review & editing, Minru Liao.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
The study was conducted in accordance with the Declaration of Helsinki.All experimental protocols were approved by the Experimental animal ethics Committee of Southwest Medical University (protocol code 20211118-032, 2021-11-18) and were performed in accordance with the ARRIVE guidelines 2.0 and the National Institutes of Health Guide for the Care and Use of Laboratory Animals.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chen, C., Gou, Q., Jin, W. et al. Upregulated miR-204-5p inhibits oral squamous cell carcinoma progression via inducing autophagy-promoted apoptosis by targeting PIK3CB. Sci Rep 16, 4299 (2026). https://doi.org/10.1038/s41598-025-34428-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-34428-y