Abstract
Approximately 30% of women of reproductive age are affected by uterine leiomyomas (fibroids). In women of reproductive age, these benign tumours have been associated with a higher risk of developing type II diabetes and infertility, which can lead to significant mental stress. Cytogenic abnormalities and somatic mutations—particularly in transcription factors such as GATA2 and FOXPI—are being considered as potential biomarkers for early detection and management. There is a pressing need for safer and more effective therapeutic options with fewer adverse effects. Therefore, this study evaluated the antioxidant, α-amylase, and α-glucosidase enzymes inhibitory and ameliorative effects of Crassocephalum crepidiodes on monosodium glutamate (MSG)-induced uterine leiomyoma in albino rats and proposed their mechanisms of action using an in silico approach. The NO radical scavenging efficacy of the control used, quercetin, is comparable to that of an aqueous extract of C. crepidiodes leaves. The extract’s ability to chelate iron (Fe2+) is lower than quercetin, the used control. The extract’s α-amylase inhibitory activity is lower than the control’s. The IC50 value for AChE inhibition by the extract was 0.113 ± 0.010 µg/mL, which is approximately 1.8-fold higher than that of the standard inhibitor, donepezil (0.061 ± 0.009 µg/mL), indicating considerable AChE inhibition. Similarly, the extract significantly inhibited MAO activity in a dose-dependent manner. Although the MSG-treated group and the C. crepidiodes-treated group had non-comparable levels of luteinizing hormone (LH) levels, the MSG-induced uterine leiomyoma rats also had noticeably higher levels of testosterone. The extract-treatment group’s reduced levels of testosterone and oestradiol relative to the MSG-induced fibrotic group demonstrated that C. crepidiodes effectively corrected the fibrotic albino rats’ hormonal abnormalities. C. crepidiodes extract’s HPLC-identified components, chlorogenic acid and ellagic acid, demonstrated high binding profile and stability with GATA2 and FOXP1 targets, supporting the in vivo antifibrotic activity of this plant. Hence, C. crepidiodes may have intricate effects on the molecular pathways linked to uterine fibroids, such as re-establishing normal ovarian morphology, hormone balance, and insulin sensitivity.
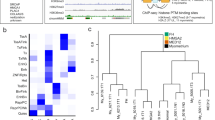
Similar content being viewed by others
Introduction
The most prevalent type of gynecologic neoplasms is uterine leiomyomas (uterine fibroids), which are tumours originating from uterine smooth muscle cells1. Uterine leiomyomas affect around 30% of women who are of reproductive age. Menorrhagia, dysmenorrhea, anemia, infertility, and miscarriage are all brought on by uterine leiomyomas, even though they are benign1,2. As a result, women with uterine leiomyomas have a far lower quality of life. Age, family history, ethnic origin, obesity, nutrition, lack of exercise, some chemicals (such as monosodium glutamate), and certain prescription medications that may raise total protein, cholesterol, and oestradiol levels are risk factors for uterine leiomyoma3. Young women with leiomyomas were associated with a high risk of incident type II diabetes and infertility, which can result in mental stress4. Multiple leiomyomas were present in 79.9% of South-West Nigerian women. However, nearly half of the patients detected in the North had multiple leiomyomas, with 48.8% of the women exhibiting these conditions5. Inconsistent diagnostic features make diagnosis challenging. Therefore, researchers have looked for leiomyoma-specific biomarkers to clarify the molecular pathophysiology of uterine leiomyomas.
Uterine leiomyoma is a monoclonal tumor that originates from a single cell6,7. Somatic mutations and cytogenetic abnormalities are therefore likely to develop into valuable biomarkers8. GATA2 is part of the GATA family of transcription factors, which play a crucial role in the regulation of gene expression during cell development and differentiation. In particular, GATA2 plays a role in endothelial function, haematopoiesis (the production of blood cells), and the control of cell growth and survival. GATA2 dysregulation has been linked to several malignancies and medical disorders, including fibroids. The function of transcription factors like GATA2 in controlling the growth and development of fibroids is one of the most promising directions for research on fibroid treatments. A transcription factor called GATA2 is involved in differentiation, survival, and proliferation, among other cellular functions. Knowing how GATA2 affects fibroid growth offers a promising target for treatment9. Research indicates that via interacting with the oestrogen and progesterone pathways, GATA2 may potentially contribute to the hormonal sensitivity of fibroids, hence promoting tumour growth10.
The side effects and temporary symptomatic cure of the conventional pharmaceuticals necessitate a better treatment option with reduced risk and adverse effects on the individual, as well as one that is widely available and relatively less expensive11,12. This involves the use of nutraceuticals, or food-based medicinal plants, such as fruits and vegetables13,14. Crassocephalum crepidioides (Asteraceae) Benth. S. Moore is an edible vegetable found in tropical and sub-tropical regions, including Nigeria. In Southwest Nigeria, it is called “ebolo,” Akwa Ibom and Edo, South-Southern Nigeria, call it “mkpafit” and “Obuinenawa” respectively15,16. Several ethnomedicinal applications of C. crepidioides have been documented, including the treatment of acute hepatitis, boils, edema, fever, wounds, indigestion, and stomach ulcers17,18,19,20. Numerous biological characteristics, including anti-bacterial, anti-helminthic, anti-inflammatory, antioxidant, and anti-coagulant properties, as well as cytoprotective, cancer chemopreventive, and anti-tumor activity in mice by inducing the production of nitric oxide in vivo, have been reported for C. crepidioides15,17,18,19,20,21,22,23,24,25,26,27,28. Therefore, this study evaluated the antioxidant, α-amylase, and α-glucosidase enzymes inhibitory and ameliorative effects of C. crepidioides on monosodium glutamate (MSG)-induced uterine leiomyoma in albino rats. In addition, an in silico approach was used to propose their mechanisms of action.
Materials and methods
Plant collection and extraction
Crassocephalum crepidiodes, the green vegetable leaves (GR), devoid of dirt, were collected from Oluponna, Osun State, Nigeria. Identification and authentication were done by Dr Idowu Obisesan at Bowen University Herbarium, Iwo, which provided the plant with its voucher number (BUH057). For 72 h at room temperature and with sporadic stirring, freshly pulverized leaves were extracted with distilled water. After filtering the extract, the filtrate was concentrated using a freeze-drier. The filtrate was further concentrated using Buchi Rotavapor in vacuo at 40 °C. The dark green extract was kept in the refrigerator until further use.
In vitro antioxidant assays
2,2-Diphenyl-1-picrylhydrazyl (DPPH) scavenging ability
With minor adjustments, the procedure outlined by Fadogba et al.29 was utilized to conduct the DPPH radical scavenging assay using 2,2-diphenyl-1-picryl-hydrazyl. Labels were applied to four test tubes: 10 mg/mL control, 20 mg/mL, 30 mg/mL, and 40 mg/mL test tubes. 1 mL of DPPH solution was added to each of the other test tubes, but 2 mL was added to the control test tube. Each test tube received 1 mL of the test samples; the control tube received nothing. The absorbance of each tube was measured at 516 nm after being kept in a pitch-black chamber for half an hour. This assay will be repeated for all other samples in triplicate. This procedure was repeated three times.
Ferric reducing antioxidant power (FRAP) potential
An adjustment to the procedure outlined by Ogunlakin et al.30 was used to ascertain the reducing power of the extract. The following four test tubes were labeled thus: control (10 mg/mL), 20 mg/mL, 30 mg/mL, and 40 mg/mL. A volume of 200 µL was transferred to each test tube, followed by the addition of 2 mL of pH 6.6 phosphate buffer and 2 mL of 1% K3[Fe(CN)4]. The test tubes were then submerged in a water bath and allowed to incubate at 50 °C for ten minutes. Following the incubation period, 2 mL of 10% TCA was added to each test tube. The mixture was then allowed to stand for fifteen minutes, and 2 mL of the supernatant was moved to another test tube. Each supernatant received 500 µL of 0.1% added to it, and a spectrophotometer was used to measure the solution’s absorbance at 700 nm.
Fenton’s reaction (OH radical scavenging activity)
The ability of the phenolics to prevent Fe2+/H2O2-induced decomposition of deoxyribose was carried out using the method of Olanrewaju et al.31 Briefly, an appropriate test sample was added to a reaction mixture containing 120 µL 20 mM deoxyribose, 400 µL 0.1 M phosphate buffer, 40 µL 20 mM hydrogen peroxide, and 40 µL 500 µM FeSO4, and the volume was made up to 800 µL with distilled water. The reaction mixture was incubated at 37℃ for 30 min, and the reaction was then stopped by the addition of 0.5 mL of 2.8% trichloroacetic acid (TCA); this was followed by the addition of 0.4 mL of 0.6% thiobarbituric acid (TBA) solution. The tubes were subsequently incubated in boiling water for 20 min. The absorbance was measured at 532 nm in a spectrophotometer. The percentage (%) ˙OH radical scavenging ability was subsequently calculated.
Determination of Fe2+ chelating ability
Twenty microliters of 2 milligrams of FeCl2 were mixed with 100 µL of the extract to make 3 mL of ethanol. For 10 min, the mixture was allowed to sit at room temperature after 40 µL of 5 mM ferrozine was added to start the reaction. The sample’s absorbance was measured at 562 nm31.
Nitric oxide scavenging activity
The concept underlying the assay is that NO and/or its oxidative derivatives react with a non-fluorescent chemical to produce a fluorescent result31. The extract and ascorbic acid at different concentrations were combined with 250 µL of 10 mM sodium nitroprusside. After the mixture had been incubated for 180 min at 25 ̊C, 500 µL of Griess reagent was added, and the absorbance at 546 nm was measured. Control samples contain 250 µL of sodium nitroprusside, phosphate-buffered saline, and Griess reagent but no extract or standard.
Inhibition of lipid peroxidation
The female albino rats were anaesthetized by intraperitoneally administration of sodium pentobarbital (40 mg/kg), and the cerebral tissue (whole-brain) and pancreas were rapidly dissected and placed on ice and weighed. This tissue was subsequently homogenized in cold saline (1:10, w/v) with about 10 up-and-down strokes at approximately 1200 rev/min in a Teflon-glass homogenizer. The homogenate was centrifuged for 10 min at 3000 × g to yield a pellet that was discarded, and a low-speed supernatant (S1) was kept for lipid peroxidation assay. The lipid peroxidation assay was carried out using the modified method of Obuh et al.32, briefly, 100 µL S1 fraction was mixed with a reaction mixture containing 30 µL of 0.1 M Tris–HCl buffer (pH 7.4), extract (0–100 µL) and 30 µL of the pro-oxidant solution (7 mM sodium nitroprusside and 15 mM Quinolinic acid). The volume was made up to 300 µL with water before incubation at 37 °C for 1 h. The color reaction was developed by adding 300 µL, 8.1% sodium dodecyl sulphate (SDS) to the reaction mixture containing S1; this was subsequently followed by the addition of 500 µL of acetic acid/HCl (pH 3.4) and 500 µL, 0.8% TBA (thiobarbituric acid). This mixture was incubated at 100 °C for 1 h. TBARS (thiobarbituric acid reactive species) produced were measured at 532 nm, and the absorbance was compared with that of a standard curve using malondialdehyde (MDA).
Acetylcholinesterase (AChE) inhibitory assay
Acetycholinesterase inhibitory study was carried out using the colorimetric method33. The reaction assay mixture consisted 2000 mL 100 mM phosphate butter pH 8.0, 100 mL of test sample stock solution in methanol (a final concentration of 42.5 µg/mL), 100 mL, of enzyme AChE (type VI-S, from electric eel; purchased from Sigma-Aldrich Co., St. Louis, MO, USA) solution at a final concentration of 0.03 U/mL and 0.01 µg/mL, respectively, 100 µL of DTNB (0.3 mM) prepared in 100 M phosphate buffer pH 7.0 containing 120 mM sodium bicarbonate. The reaction mixture was vortexed and then pre-incubated in a water bath at 37 °C for 30 min. The reaction was then initiated by the addition of 100 µL of ATCI or BTCI at a final concentration of 0.5 mM as a negative control. The inhibitor solution was replaced with methanol. The change in absorbance at λmax 412 nm was then measured for a period of 5 min at ambient temperature. All assays were carried out in triplicate. The final concentration of the sample was 42.5 µg/mL. Donepezil was used as a positive control at the same concentration. The % inhibition was calculated.
Monoamine oxidase inhibitory assay
The effect of the extract on MAO (EC 1.4.3.4; purchased from Sigma-Aldrich Co., St. Louis, MO, USA) activity was measured according to a previously reported method34. In brief, the reaction mixture contained 0.025 M phosphate buffer (pH 7.0), 0.0125 M semicarbazide, 10 mM benzylamine, 0.67 mg of the enzyme, and 0–100 µL of extracts. After 30 min incubation, ace tic acid was added and boiled for 3 min in a boiling water bath, followed by centrifugation. The resulting supernatant (1 mL) was mixed with an equal volume of 2,4-dinitrophenylhydrazine, and 1.25 mL of benzene was added after 10 min incubation at room temperature. After separating the benzene layer, it was mixed with an equal volume of 0.1 N NaOH. The alkaline layer was decanted and heated at 80 °C for 10 min. The orange–yellow color developed was measured at 450 nm in a UV/visible spectrophotometer (Jenway 6305 model). The MAO activity was thereafter expressed as a percentage inhibition of the reference.
α-amylase inhibitory potential
This assay was conducted following the standard protocol to determine the α-amylase inhibitory potential of the extracts35. To begin, a fresh preparation of enzyme (porcine pancreatic α-amylase; purchased from Sigma-Aldrich Co., St. Louis, MO, USA) was prepared, comprising 5 units per mL, in pH 6.7 ice-cold PBS with a concentration of 20 mM and 6.7 mM NaCl. Then, 250 µL of the enzyme was combined with inhibitors (acarbose or test samples) at varying concentrations (excluding a blank sample), and the mixture was incubated at 37 °C for 20 min. Then, starch solution at a concentration of 0.5% (w/v) was added, and the mixture was incubated for an additional 15 min at 37 °C. Immediately after the DNS reagent was added, the mixture was mixed and placed in a water bath at 100 °C for 10 min. Finally, the absorbance was read at 540 nm.
α-glucosidase inhibitory potential
The effect of the extracts on intestinal α-glucosidase activity was evaluated using a technique described by Bouslamti et al.36 which quantified the glucose generated by sucrose breakdown. To perform the assay, 100 µL of sucrose (50 mM), 1000 µL of phosphate buffer (50 mM; pH = 7.5), and 100 µL of α-glycosidase enzyme (from Saccharomyces cerevisiae, purchased from Sigma-Aldrich Co., St. Louis, MO, USA) solution were prepared as the test solution (10 I.U.). Control (distilled water), positive control (acarbose), or test samples were all added to this mixture at varying concentrations. The absorbance was read at 500 nm.
In vivo assay
Dosing of experimental animals
Human therapeutic dosages of C. crepidiodes range from 1 to 9 g, according to the ethnobotanical survey that was carried out in Iwo, Nigeria, during the study (unpublished data). Rat dosage was determined based on body surface area and the conversion factor from human to albino rat (conversion factor = 0.162). To do this, it was divided by the 60 kg adult human weight and then multiplied by a factor to account for the animal’s body surface area37,38. About 100 mg/kg b.w. rat is the range of the calculated dose that was attained. Thus, in this investigation, a dose of 100 mg/kg was used. Gavage was used to dose albino rats. During the experiment, the dosage was once daily at a volume of 2 mL per kg body weight. Based on the animal’s most recent reported body weight, individual dose volumes were computed. Since oral delivery is the recommended method of human exposure, it was chosen.
MSG-induced uterine leiomyoma study
Healthy female albino rats (15) weighing 190 and 200 g each were procured from Bowen University in Iwo, Nigeria. All experimental rats used in this study were handled in accordance with the rules and regulations established for animal management in research, as outlined in NIH Publications No. 80-23 revised, 1996. The Institutional Animal Ethics Committee of the Department of Biochemistry at Bowen University, Iwo (BUI/BCH/2025/001) confirmed and approved that the experimental treatment of the rats is in accordance with ARRIVE guidelines. Female albino rats were grouped into three groups of five animals each. Uterine fibroid was induced by oral administration of 800 mg/kg body weight of MSG for 30 days. Group A was the no-treatment group (Control). MSG-treated and GR-treated groups received 800 mg/kg MSG, while the Control group received no MSG treatment. All groups were treated except the control group received 100 mg/kg of C. Crepidiodes extract for 30 days, after which they were euthanized by intraperitoneally administration of sodium pentobarbital (40 mg/kg) at the end of the experiment. Total plasma follicle-stimulating hormone (FSH), luteinizing hormone (LH), testosterone, and estradiol levels were determined. The cervix and uterus were harvested for histopathology.
Ovarian and cervical histology
The ovaries and cervixes were examined using a standard procedure. The sections were removed with clean, labeled slides from a water bath that Raymond Lamb had heated to 55 °C, dehydrated on a hotplate for an hour at 60 °C, and then inspected under a light microscope with x 40 objectives.
HPLC analysis
To identify the phytochemicals in the extracts, High-Performance Liquid Chromatography (HPLC) analysis was conducted using the protocol outlined by Olanrewaju et al.31 With a few modifications, the composition gradient was: 5% of methanol (B) for the first 2 min and then changed to obtain varying percentages of B at 10–60 min in 10-min intervals, respectively. The mobile phase was water containing 2% acetic acid (A) and methanol (B). The compounds identified were used for an in silico study.
In silico study
Protein structure preparation
The 3D-structures of backbone dynamics of the DNA-binding domain of FOXP1(PDBID: 2KIU) and GATA2 C-terminal zinc finger domain (GATA2) (PDBID: 5O9B) were retrieved from the Protein Data Bank (http://www.rcsb.org). The existing ligands and water molecules were removed from all the crystal structures, while missing hydrogen atoms were added using MGL-AutoDockTools (ADT, v1.5.6)39.
Ligand preparation
The retrieval of Structure Data Format (SDF) of HPLC-identified bioactive compounds was downloaded from the PubChem database (www.pubchem.ncbi.nlm.nih.gov). The compounds were further converted to the pdb chemical format by means of Open Babel40. Non-polar hydrogen molecules were merged with the carbons, while the polar hydrogen charges of the Gasteiger-type were assigned to atoms. Furthermore, ligand molecules were converted to dockable PDBQT format with the help of AutoDock Tools.
Molecular Docking of phytochemicals with the targeted active site
AutoDock Vina integrated with PyRx 0.8 was used to carry out an active site target molecular docking of the reference inhibitors and the HPLC-identified bioactive compounds to the binding site of the three target proteins41. Prior to the docking analysis, PyRx 0.8’s bioactive chemicals were imported using OpenBabel40. The bioactive compounds were further minimized using OpenBabel. The energy minimization parameter and conjugate gradient descent used were the Universal Force Field (UFF) and optimization algorithm, respectively. The binding site coordinates of the target proteins were identified by mapping the amino acid residues around the binding site of the native ligand. The dimension of the grid boxes formed was center x (-28.25), y (26.12), z (-22.72), and size x (33.95), y (36.43), z (54.20). The selected conformer from the docking analysis was further subjected to interactive analysis using Discovery Studio Visualizer version 16.
Molecular dynamics
For a 100 ns molecular dynamics simulation, the complexes of the top two docked bioactive chemicals with the GATA2 (5O9B) were further chosen. The study was conducted using GROMACS 2019.2 and the GROMOS96 43a1 force field42,43,44. The proteins and ligands topology files were generated using Charmm GUI45,46. The simulation used a solvation system, periodic boundary conditions, physiological conditions, system minimisation, equilibration in a constant number of atoms, constant pressure, and constant temperature (NPT), all of which were similar to those in our previous study47,48,49. The velocity rescales and Parrinello-Rahman barostat were used to maintain the temperature and pressure at 310 K and 1 atm, respectively. A 2-femtosecond time step was used with a leap-frog integrator. Each system underwent a 100 ns simulation, with snapshots being taken every 0.1 nanosecond and totaling 1000 frames for each system. From the MDs trajectories, the RMSD and RMSF, ROG, SASA, and H-bonds.
Binding free energy calculation using MM-GBSA
The Molecular Mechanics Generalised Born Surface Area (MM-GBSA) method and decomposition analysis using the gmx MMPBSA package were used to obtain the binding energies of amino acids within 0.5 nm of the ligand in order to determine the binding free energy of the two top docked phytochemicals from the initial docking analysis50,51. The methods used were the same as those published in our previous manuscripts47,48.
Statistical analysis
Values were displayed as Mean ± Standard deviation (SD). Data were analyzed using one-way analysis of variance (ANOVA), and group means were compared using Dunnett’s Multiple Comparison and Bonferroni tests using GraphPad Prism version 5.01 for Windows, GraphPad Software, San Diego, California, USA. P values that were less than 0.05 were deemed significant.
Results
Antioxidant and Inhibition of α-amylase, α-glucosidase, cholinesterase, and monoamine oxidase activities
The antioxidant activity of the aqueous extract of C. crepidiodes leaves is displayed in Table 1. When compared to quercetin, the standard utilized, the extract was found to have a lesser capacity to scavenge DPPH radicals. It is demonstrated that the NO radical ability was equivalent to quercetin (Fig. 1), the criterion that was employed. The extract’s capacity to chelate iron (Fe2+) was less than that of the employed control, quercetin. The α-amylase activity of the extract was marginally lower than that of the control, as seen in Table 2. As extract concentrations rise, Fig. 2A illustrates an increase in α-amylase activity. In Fig. 2B, the extract’s capacity to inhibit α-glucosidase activity was lower than that of the control. The effects of C. crepidioides (CC) leaf extract on acetylcholinesterase (AChE) and monoamine oxidase (MAO) activities are presented in Figs. 3A,B. The extract exhibited dose-dependent inhibitory activity against both enzymes. Similarly, CC extract significantly inhibited MAO activity in a dose-dependent manner, with an IC50 value of 0.179 ± 0.003 µg/mL, which was slightly higher than that of donepezil (0.155 ± 0.005 µg/mL).
(A) % DPPH scavenging ability, (B) Ferric Reducing antioxidant property, (C) Hydroxyl radical scavenging ability, (D) % Iron chelating activity, (E) Nitric oxide radical scavenging ability, and MDA inhibition in (F) the brain and (G) pancreas, of Crassocephalum crepidioides. Control for all is Quercetin, and DPPH radical assay, where ascorbic acid was used. Control for MDA inhibition assay is Donepezil.
Percentage Inhibitory effect of Crassocephalum crepidioides on (A) α-amylase, (B) α-glucosidase. Control (Gallic acid).
Percentage Inhibitory effect of Crassocephalum crepidioides on (A) acetylcholinesterase, and (B) monoamine oxidase. Control (Donepezil).
In vivo study
While the levels of luteinizing hormone (LH) in the MSG-treated group and the C. crepidioides-treated group were not similar, Fig. 4 demonstrates that the MSG-induced uterine leiomyoma rats had a significantly higher level of testosterone. Crassocephalum crepidioides successfully corrected the hormonal anomalies in the albino rats, as evidenced by the treatment group’s reduced levels of testosterone and oestradiol in comparison to the control group. Figure 5 presented the photomicrograph of the ovaries and uterus of the rats. Photomicrograph of a uterine section showed a normal endometrium epithelial layer, normal endometrial gland, and mild to moderate infiltration of the endometrial stroma by inflammatory cells. The cervix showed developing follicles, including secondary follicles, Graafian follicles, as well as atretic follicles within the ovarian cortex. The ovarian stroma shows normal connective tissues and luteinized cells.
Effect of Crassocephalum crepidiodes leaf extract on (A) FSH, (B) LH, (C) Testosterone, and (D) Estradiol levels in control and treated rats. GR = Amaranthus hybridis leaves aqueous extract. * Means there is a significant difference compared to the positive control group.
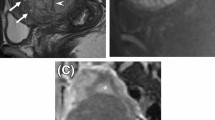
Photomicrograph of ovaries and uterine sections of the control (Ai and Aii), MSG-treated (Bi and Bii), and Crassocephalum crepidiodes-treated (Ci and Cii) groups, respectively, stained by hematoxylin and eosin (Mag. X 40). For the ovary, the Graafian follicle (red arrow), and the Atresia follicle (black arrow) within the ovarian cortex, an antral follicle with an oocyte (green arrow). For the cervix, endometrium epithelial layer (white arrow), normal endometrial gland (blue arrow).
In silico study
The binding scores of the HPLC-detected bioactive compounds (Fig. 6) against the GATA2 and FOXP1 targets are represented in Table 3.
HPLC Chromatogram of Crassocephalum crepidioides aqueous extract.
Interaction of the two top-ranked bioactive compounds from the docking analysis with the protein targets
Ellagic acid was stabilized in the binding to GATA2 by a covalent bond Asn41 and Asn39 and several van der Waals to Asn41, Tyr34, Tyr35, Arg42, Asn39, His38, while chrologenic acid interaction with GATA2 was stabilized by hydrogen bonds with Thr15, Thr14, Thr13, Glu49, Gln11, Lys48, pi-alkyl contact with Ala30 and van der Waals with Asn29, Thr45, Cys28 and Thr12. Kaempferol, on the other hand, was stabilized in the binding to FOXP1 by a hydrogen bond with Asn44, amide-pi stacking to Tyr9, pi-pi stacking to Thr47, pi-alkyl contact with Ala51, and van der Waals to Ile13, Asn55, Val52, Thr8, Phe7, Pro6, Phe41, Tyr41, Tyr40, and Trp48. Chlorogenic acid was stabilized by hydrogen bonds with Asn44, Tyr40, and Tyr9, pi-pi T shaped contact with Phe38, pi-alkyl contact with Arg4 and van der Waals interaction with Val52, Ala51, Pro6, Val3,, Thr47, Phe41, Pro5, Leu12 and Peh38 (Fig. 7).
Interactive plots of op ranked compounds from the docking analysis with amino acids in the binding site of (a) GATA2 (i) chrologenic acid (ii) kaempferol and (b) FOXP1 (i) Ellagic acid (ii) chrologenic acid. The ligands are displayed as sticks.
Molecular dynamics
At the conclusion of the simulation, there was noticeable convergence in the unbound (5O9B) and 5O9B_Chlorogenic acid continuous evolution (Fig. 8). The plot of the RMSF system shows high fluctuations at the beginning of the simulation, which correspond to motion around terminals and around amino acid residues 31 to 40. The mean RMSF value of 6.66 ± 1.99 Å represents the baseline of the unbound system (5O9B). 5O9B_Chlorogenic acid system with RMSF value of 5.89 Å slightly reduced flexibility compared to the unbound form, suggesting some stabilization. 5O9B_Ellagic acid system with 5.46 Å) shows the lowest flexibility, indicating stronger stabilization of residues (Table 4).
The plots of thermodynamic parameters computed from the analysis of the MDs trajectories of unbound and bound 5O9B complex systems (a) The Backbone-Root Mean Square Deviation (RMSD) (b) Per residue Root Mean Square Fluctuations (RMSF) (c) radius of gyration (d) Surface Accessible Surface Area (SASA) (e) number of hydrogen atoms.
Molecular mechanics generalized born surface area (MMGBSA) analysis
Figure 9 shows the per-residue contribution to the total binding free energy. For the 5O9B_Ellagic acid system, the ligand contributed − 10.37 kcal/mol to the total binding free energy. The 5O9B_Ellagic acid system (-20.71 ± 4.94 kcal/mol) presented higher binding free energy when compared to 5O9B_Chlorogenic acid (− 10.95 ± 8.72), as shown in Table 5. The top contributing residues include LEU36 (-2.48), ARG20 (-1.74), PHE16 (-1.49), TRP37 (-1.03 kcal/mol), GLY18 (-0.74 kcal/mol), ARG21 (-0.67 kcal/mol), GLN17 (-0.50 kcal/mol) (Fig. 9a). The complex’s structural integrity is maintained by moderately supportive residues such as ARG20, TRP37, and GLY18, which do not dominate interactions. Overall contributions from residues like LEU36, PHE16, VAL14, and HID8 are lower, indicating that they promote or play a supporting function in localised binding dynamics. Chlorogenic contributed about − 3.82 kcal/mol to the overall binding to GATA2. This interaction was majorly stabilized by ΔVDWAALS (-6.80 kcal/mol). Its role in solvation and non-covalent interactions makes it pivotal for the system. By means of electrostatics and solvation, residues such as THR35 (-1.20 kcal/mol), THR34 (-1.01 kcal/mol), and ARG20 (-0.42 kcal/mol) have mild stabilising effects (Fig. 9b).
Molecular Mechanics Generalized Born Surface Area (MM-GBSA) plot of the contributing amino acid residues of (a) 5O9B_Ellagic acid (b) 5O9B_Chlorogenic acid systems.
Discussion
Antioxidant and Inhibition of α-amylase, α-glucosidase, cholinesterase, and monoamine oxidase activities
The antioxidant evaluation of C. crepidioides leaf extract revealed substantial radical scavenging and metal-chelating properties across multiple assays, supporting its traditional use and potential antifibrotic efficacy. The DPPH assay, which assesses the ability of the extract to neutralize free radicals, demonstrated a concentration-dependent scavenging effect comparable to that of ascorbic acid, the standard antioxidant used. This suggests that C. crepidioides contains potent hydrogen-donating phytochemicals, possibly flavonoids and phenolic compounds, known for their strong antioxidant activities52. As free radicals are major contributors to oxidative stress, which in turn fuels the pathogenesis of fibrosis53, the potent DPPH scavenging effect positions C. crepidioides as a promising source of natural antioxidants for antifibrotic interventions. Similarly, the ferric reducing antioxidant power (FRAP) assay showed that the extract effectively reduced Fe3+ to Fe2+, indicating its electron-donating capacity. This reducing power is essential in mitigating oxidative stress, as it helps neutralize reactive oxygen species (ROS) before they can initiate cellular damage. Given the critical role of oxidative stress in the pathogenesis of fibrosis54, the strong ferric-reducing activity of C. crepidioides highlights its potential therapeutic value. The FRAP results align with findings by Aryal et al.55, who emphasized that plants rich in phenolic and flavonoid content often exhibit strong reducing power, supporting the likelihood that bioactive polyphenols in C. crepidioides contribute to its antioxidative effects.
The hydroxyl radical scavenging activity was notably high, suggesting that the extract can effectively counter the most reactive oxygen species in biological systems. Hydroxyl radicals are known to cause extensive lipid peroxidation, DNA damage56, and protein modification, all of which contribute to fibrotic tissue remodeling57. The robust hydroxyl radical scavenging activity observed implies that C. crepidioides could mitigate oxidative tissue injury and subsequent fibrosis by suppressing lipid peroxidation pathways. In addition to neutralizing free radicals, the extract exhibited significant iron-chelating ability. Since free iron can catalyze the Fenton reaction, leading to hydroxyl radical generation, iron chelation represents a critical antioxidant defense mechanism58, generating harmful hydroxyl radicals. The ability to chelate iron reduces the risk of oxidative damage at the molecular level59. This metal ion sequestration further underlines the multifaceted antioxidant defense offered by the plant.
The nitric oxide (NO) scavenging assay revealed that C. crepidioides could substantially inhibit NO production. While NO is a vital physiological signaling molecule, its overproduction, especially under inflammatory conditions, could facilitate its reaction with superoxide, forming peroxynitrite that may cause nitrosative stress, thereby exacerbating tissue damage and promoting fibrosis53,60. Thus, the NO scavenging capacity of the extract suggests an additional anti-inflammatory dimension to its antifibrotic mechanism. Collectively, these antioxidant results strongly support the hypothesis that C. crepidioides exerts its antifibrotic effects, at least in part, by mitigating oxidative and nitrosative stress. Notably, recent research emphasizes that plant-derived antioxidants not only neutralize free radicals but also regulate fibrogenic signaling pathways such as TGF-β/Smad and NF-κB61. This is consistent with emerging research emphasizing the role of natural antioxidants in modulating fibrosis through redox balance restoration62. Furthermore, the use of in vitro antioxidant assays aligns with contemporary strategies that combine biochemical evaluations with in silico modeling to elucidate pharmacological potentials of medicinal plants63.
The IC50 value for AChE inhibition by the extract was 0.113 ± 0.010 µg/mL, which is approximately 1.8-fold higher than that of the standard inhibitor, donepezil (0.061 ± 0.009 µg/mL), indicating considerable AChE inhibition. Similarly, C. crepidioides extract significantly inhibited MAO activity in a dose-dependent manner. These findings suggest that C. crepidioides possesses intrinsic inhibitory activity against AChE and MAO, two enzymes responsible for the metabolism of acetylcholine and monoamines, respectively. While the inhibitory potency of the extract was lower than that of donepezil, the observed activity indicates a potential modulatory role of C. crepidioides in cholinergic and monoaminergic signalling pathways. Acetylcholine, beyond its role in neurotransmission, is known to mediate the cholinergic anti-inflammatory pathway64. This pathway plays a key role in limiting inflammatory responses associated with fibrogenesis. In models of pulmonary fibrosis, acetylcholine has been shown to inhibit myofibroblast differentiation via activation of α7 nicotinic acetylcholine receptors (α7nAChRs), leading to suppression of TGF-β signalling and promotion of myofibroblast dedifferentiation mechanisms that collectively reduce fibrotic tissue remodelling65. Similarly, monoamine oxidases (particularly MAO-A) catalyse the oxidative deamination of monoamines such as serotonin, norepinephrine, and dopamine, producing reactive oxygen species (ROS) such as hydrogen peroxide (H2O2) as metabolic by-products66. Elevated ROS levels contribute to fibrotic progression by promoting extracellular matrix (ECM) deposition and activating fibroblasts67,68. Therefore, MAO inhibition may attenuate ROS-mediated tissue injury and fibrosis.
In vivo study
We measured serum levels of follicle-stimulating hormone (FSH), luteinizing hormone (LH), testosterone, and estradiol in control, MSG-treated, and Crassocephalum crepidiodes treated rats to determine the endocrine-modulatory effects of C. crepidioides leaf extract. Compared to the control, the MSG-treated group showed a notable rise in FSH levels. Treatment with C. crepidioides extract, on the other hand, caused a slight, non-significant drop in FSH levels as compared to the control. MSG may cause neurotoxic effects on the hypothalamus by overactivating NMDA glutamate receptors, hence generating oxidative stress and inflammatory responses69. This changes pituitary FSH production by gonadotropin-releasing hormone (GnRH) pulsatile release. Additionally, increased FSH production is elevated by higher GnRH levels via the GnRH receptor–Gq/11–PLC–IP3/Ca²⁺–PKC signaling pathway70. The slight reduction in FSH following C. crepidoides treatment could be ascribed to its antioxidant-rich phytochemical profile, especially flavonoids and terpenoids, which can reduce glutamate-induced excitotoxicity and restore normal hypothalamus function. This antioxidant effect normalizes GnRH pulsatility and FSH output via inhibition of NF-κB signaling, hence regulating neuroinflammation71.
For LH, the MSG-treated group showed a slight increase compared to the control; the C. crepidioides-treated group exhibited a modest reduction in LH level. Though not statistically significant, this drop suggests a possible modifying influence of the extract. GnRH pulse frequency controls LH secretion as well. MSG-induced changes in hypothalamic function increase LH production by the cAMP/PKA pathway in pituitary gonadotrophs. Oxidative damage and inflammation also disturb the hypothalamus’ regulation of gonadotropin release72. By reducing glutamate toxicity and re-establishing appropriate GnRH signaling frequency, C. crepidioides extract probably restored LH balance. Furthermore, C. crepidioides might modulate kisspeptin-GPR54 signaling, a key upstream control of GnRH neurons, and reduce hypothalamus inflammation by blocking pro-inflammatory cytokines (IL-1β, TNF-α) and MAPK/ERK pathways73,74,75. The MSG-treated group had significantly higher serum testosterone levels than the control one. The C. crepidioides-treated group had testosterone levels slightly lower than the MSG-treated group but higher than the control, suggesting partial reversal of MSG-induced elevation. In MSG-treated rats, higher LH levels activate Leydig cell steroidogenesis via the LH receptor-mediated cAMP–PKA–StAR–CYP11A1 pathway, hence promoting testosterone production. MSG might potentially cause lipid peroxidation in Leydig cells, enhancing steroidogenic activity as a compensatory response76. By downregulating steroidogenic enzymes as StAR, 3β-HSD, and CYP17A1, the C. crepidioides extract partially restore testosterone synthesis77. Moreover, C. crepidioides’s antioxidant properties could also improve testicular redox homeostasis by means of Nrf2-mediated antioxidant defense mechanisms, hence safeguarding Leydig cells from oxidative stress and controlling testosterone production78.
The elevated estradiol in MSG-treated rats may stem from enhanced aromatization of testosterone via aromatase, which is upregulated by pro-inflammatory cytokines and oxidative stress through the COX-2–PGE2–cAMP/PKA and STAT3/NF-κB pathways79,80. Increased aromatase expression in adipose and gonadal tissues underlies the hyperestrogenic state observed. Remarkably, the MSG-treated group had higher estradiol levels than the control (p < 0.05). On the other hand, C. crepidioides extract administration lowered estradiol levels significantly compared to both the control and MSG-treated groups (p < 0.05). The inhibition of NF-κB and STAT3 signaling and COX-2 expression suppression may cause downregulation of aromatase expression. Moreover, C. crepidioides might increase AMPK–SIRT1 axis activity or directly suppress aromatase enzymatic activity, hence lowering estrogen production. These results imply that C. crepidioides protects the hypothalamic-pituitary-gonadal axis and may help to correct MSG-induced disturbance of reproductive hormones, through modulation of oxidative stress, inflammation, and neuroendocrine signaling.
In silico study
GATA2 inhibition reduces smooth muscle cell growth, which could result in fewer and smaller fibroids. It may prevent the production of genes involved in the survival and proliferation of fibroid cells by inhibiting the transcriptional activity of GATA28,9,10. The binding scores were between − 4.1 to -6.9 kcal/mol and − 4.9 to -7.5 kcal/mol for GATA2 and FOXP1 targets, respectively. Chrologenic acid (-7.4 for FOXP1) has the most negative binding score for FOXP1, which suggests that it has a strong binding affinity for this protein. Kaempferol (-7.5 for FOXP1) also has a very low score for FOXP1, indicating a strong binding to this target. Rutin (-6.4 for FOXP1) and quercetin (-7.1 for FOXP1) have good binding affinities as well, while ellagic acid and chrologenic acid show strong binding to GATA2 (-6.9 and − 6.4, respectively), indicating that these compounds are likely to interact significantly with GATA2. Although the difference is not very great, Ellagic acid (-6.9 for GATA2 and − 6.5 for FOXP1) indicates a greater binding score for GATA2 than for FOXP1. For FOXP1, chlorogenic acid exhibits a higher binding score (-7.4) than GATA2 (-6.4). Compared to GATA2 (-5.6), kaempferol binds to FOXP1 more strongly (-7.5). For both targets, compounds such as Rutin (-5.7 for GATA2, -6.4 for FOXP1), quercetin (-5.5 for GATA2, -7.1 for FOXP1), and aromadendrene (-5.9 for GATA2, -5.6 for FOXP1) exhibit intermediate binding scores. Although not as effective as the top-scoring compounds, these compounds may have a reasonable ability to target both proteins. With scores closer to -5, some compounds, such as benzofuran (-4.6 for GATA2, -5.9 for FOXP1) and eugenol (-4.9 for GATA2, -6.2 for FOXP1), have poorer binding, indicating that they would not be as effective in interacting with either target. For target-specific binding, chrologenic acid and kaempferol have much stronger binding to FOXP1, while chrologenic acid and ellagic acid show a stronger interaction with GATA2. This diversity in structure might contribute to a wide range of potential interactions with the target.
The two top-ranked compounds that were selected based on the binding energy and interaction with the target proteins were further subjected to interactive analysis. Ellagic acid was stabilized in the binding to GATA2 by a covalent bond Asn41 and Asn39 and several van der Waals to Asn41, Tyr34, Tyr35, Arg42, Asn39, His38, while chrologenic acid interaction with GATA2 was stabilized by hydrogen bonds with Thr15, Thr14, Thr13, Glu49, Gln11, Lys48, pi-alkyl contact with Ala30 and van der Waals with Asn29, Thr45, Cys28 and Thr12. Kaempferol, on the other hand, was stabilized in the binding to FOXP1 by a hydrogen bond with Asn44, amide-pi stacking to Tyr9, pi-pi stacking to Thr47, pi-alkyl contact with Ala51, and van der Waals to Ile13, Asn55, Val52, Thr8, Phe7, Pro6, Phe41, Tyr41, Tyr40, and Trp48. Chlorogenic acid was stabilized by hydrogen bonds with Asn44, Tyr40, and Tyr9, pi-pi T shaped contact with Phe38, pi-alkyl contact with Arg4 and van der Waals interaction with Val52, Ala51, Pro6, Val3,, Thr47, Phe41, Pro5, Leu12 and Peh38.
A vital computational method for understanding ligand-protein interactions, molecular dynamics (MD) simulations shed light on atomic-level binding processes, stability, and dynamic behaviour81. The RMSD calculates the simulation’s overall structural stability. A well-formed and stable complex is indicated by a decreased or steady RMSD81. Before 20 nanoseconds, the RSMD systems were in equilibrium. At the conclusion of the simulation, there was noticeable convergence in the unbound (5O9B) and 5O9B_Chlorogenic acid continuous evolution. The baseline mean RMSD value for the 5KC5 (Unbound) was 9.88 ± 2.70 Å, indicating relatively high structural variation during the simulation. The 5O9B_Chlorogenic acid (9.98 Å) system presented close structural deviation to the unbound form, suggesting comparable stability. The 5O9B_Ellagic acid system (7.29 Å) demonstrated lower RMSD compared to the others, reflecting greater structural stability with ellagic acid81.
The flexibility of each protein residue is measured by RMSF. Reduced flexibility and a more rigid structure are indicated by lower RMSF values82. The plot of the RMSF system shows high fluctuations at the beginning of the simulation, which correspond to motion around terminals and around amino acid residues 31 to 40. The mean RMSF value of 6.66 ± 1.99 Å represents the baseline of the unbound system (5O9B). 5O9B_Chlorogenic acid system with RMSF value of 5.89 Å slightly reduced flexibility compared to the unbound form, suggesting some stabilization. 5O9B_Ellagic acid system with 5.46 Å) shows the lowest flexibility, indicating stronger stabilization of residues. Ellagic acid’s variance in RMSF values (with a greater standard deviation) points to a potential area with notable interaction or flexibility alterations83. RoG indicates protein compactness during the simulation. A higher RoG may suggest unfolding or structural expansion, while a lower RoG suggests a more stable, compact structure (Hong and Gierasch 2010). The mean RoG value of 13.06 ± 0.64 Å represents the baseline of the unbound system (5O9B). The 5O9B_Chlorogenic acid system (14.27 ± 0.92 Å) with increased RoG implies the structure is slightly less compact when bound to chlorogenic acid. The 5O9B_Ellagic acid system (13.73 ± 0.91 Å) with a slight increase in RoG, but not as much as chlorogenic acid, suggesting moderate compactness. The 5O9B_Ellagic acid complex has a relatively large RoG compared to 5O9B, but it is still more compact than the Chlorogenic acid complex84.
The amount of protein surface exposed to the solvent is measured by SASA. More exposure is indicated by a higher SASA, whereas a more compact or buried building is indicated by a lower SASA85. The mean SASA value of 5084.4 ± 266.5 Ų represents the baseline solvent accessibility of the unbound system (5O9B). The 5O9B system has the smallest SASA, indicating that it is more tightly packed or shielded from the solvent, which might be an indicator of more stable binding. The 5O9B_Chlorogenic acid system with SASA value of 5227.8 Ų presents increased SASA, suggesting more surface exposure to solvent. The 5O9B_Ellagic acid system (5537.2 Ų) presents the highest SASA, likely due to conformational changes that expose more of the structure to solvent85. Hydrogen bonds refer to the number of hydrogen bonds formed between the protein and the ligand. H-bonds are crucial for ligand-protein stability; a higher mean H-bonds number indicates stronger interactions. The mean number of H-bonds, 21.16 ± 3.68 represents the baseline of the unbound system (5O9B). As observed in the graph, there is no major difference in the progression between the bound and the unbound system. The 5O9B_Chlorogenic acid (21.87) with a slight increase in H-bonds, indicating a modest stabilizing effect. The 5O9B_Ellagic acid (17.76) presents fewer H-bonds formed compared to chlorogenic acid, potentially indicating weaker interactions. Despite having a larger SASA and fewer hydrogen bonds, ellagic acid exhibits superior structural stability with lower RMSD and reduced flexibility (RMSF). Stabilisation via non-H-bond interactions may be the cause of this. Although chlorogenic acid forms slightly more H-bonds than ellagic acid, indicating moderately strong interactions, it also shows greater SASA and RoG, indicating a more extended conformation86.
The energy difference between the bound and unbound components of a complex (ligand and receptor) is measured by the binding free energy; the bigger the negative value, the more strongly the ligand binds to the protein87. Binding free energy estimates provide comprehensive information on the binding mechanisms of the top docked compounds throughout the early stages of drug discovery and development (Kollman et al. 2000). The binding free energy of the two top-docked compounds against GATA2 (5O9B) was calculated using the MMGBSA technique. The 5O9B_Ellagic acid system (-20.71 ± 4.94 kcal/mol) presented higher binding free energy when compared to 5O9B_Chlorogenic acid (− 10.95 ± 8.72), suggesting that it has a more stable and favorable binding interaction with the protein compared to chlorogenic acid, despite the less favorable solvation and electrostatic solvation energies. Ellagic Acid exhibited stronger van der Waals (-30.67 ± 4.07 kcal/mol) and gas-phase interactions (-47.10 ± 10.42 kcal/mol), leading to more robust binding. Despite higher solvation energy costs (ΔGSOLV) (26.39 ± 6.68 kcal/mol), its efficient packing (lower ΔESURF) ((-3.69 ± 0.45 kcal/mol) and stronger gas-phase binding outweigh these penalties. Ellagic acid emerges as the better lead for stabilizing the 5O9B protein. Chlorogenic acid, on the other hand, exhibited strong electrostatic interactions (-109.87 ± 44.4 kcal/mol) but suffers from high surface area energy (ΔESURF) (114.72 ± 44.69 kcal/mol), which decreases its efficiency. Its binding appears highly dependent on solvation effects, which may reduce its stability in less aqueous environments. To understand the contribution of amino acids to the total binding free energy, especially for the 5kc5_Kaemferol system, the total binding free energy was decomposed. Ellagic acid was a major stabilizer, contributing the strongest van der Waals interactions, favorable solvation energy, and overall binding energy. This probably has a major impact on preserving the complex’s stability. THR35 and THR34 display ΔVDWAALS and ΔEEL values that are favourable despite being rather tiny, indicating moderate contributions to binding. Both top docked compounds demonstrated a high binding profile and stability with the target, suggesting high binding tendencies. These results demonstrate the unique functions that particular residues and their electrostatic and non-electrostatic interactions play in maintaining protein-ligand complex stability.
Conclusion
Based on the data obtained, along with its demonstrated antioxidant, α-amylase, and α-glucosidase inhibitory properties, Crassocephalum crepidiodes was effective in improving the fibrotic conditions in female albino rats. HPLC identified key bioactive molecules—clorogenic acid and ellagic acid in the plant extracts. These bioactives exhibited strong binding affinity and stability with protein targets GATA2 and FOXP1, supporting the plant’s observed antifibrotic activity in vivo. Together, the findings suggest that C. crepidiodes may have multifaceted effects on the molecular pathways involved in uterine fibroids, potentially contributing to the restoration of normal ovarian morphology, hormonal balance, and insulin sensitivity.
Data availability
The datasets used and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Pinto, A. Uterine smooth muscle tumors: an overview. Adv. Anat. Pathol. 31(6), 397–410 (2024).
Borah, B. J., Nicholson, W. K., Bradley, L. & Stewart, E. A. The impact of uterine leiomyomas: a National survey of affected women. Am. J. Obstet. Gynecol. 209(4), 319–e1 (2013).
Aninye, I. O., Chew, S. & Goulmamine, S. 2025 SWHR Women’s Health Research Agenda: Prioritizing Uterine Fibroids, Lupus, and Metabolism (Journal of Women’s Health, 2025).
Sung, J. H., Kim, K. S., Han, K. & Park, C. Y. Association of uterine leiomyoma with type 2 diabetes mellitus in young women: a population-based cohort study. Diabetes Metabolism J. 48(6), 1105–1113 (2024).
Anaziah, G. The prevalence and post-operative complications of uterine leiomyomas in South-Western Nigeria and Northern Nigeria: A comparartive review ([Accessed on 16/04/2025] (2023).
Maekawa, R. & Sugino, N. Epigenetics and uterine fibroids. Uterine Fibroids Adenomyosis 69–85 (2018).
Yang, X. et al. Lichong Decoction improves inflammatory microenvironment and alleviates fibrosis in uterine leiomyoma via targeting CXCL8. J. Ethnopharmacol. 340, 119276 (2025).
Glembocki, A. I. & Somers, G. R. Prognostic and predictive biomarkers in paediatric solid tumours. Pathology 56(2), 283–296 (2024).
Pavlovic, Z. J. et al. Dysregulated expression of GATA2 and GATA6 transcription factors in adenomyosis: implications for impaired endometrial receptivity. F&S Sci. 5(1), 92–103 (2024).
MacLean, J. A. & Hayashi, K. Progesterone actions and resistance in gynecological disorders. Cells 11(4), 647 (2022).
Ajayi, R. O., Adeyemi-Benson, O. S., Adeyemi-Benson, O. A. & Ogunjobi, T. T. Chronic disease management in families: A public health and biomedicine perspective. Medinformatics (2025).
Ogunlakin, A. D. et al. Prospects, mechanisms, and limitations of botanicals’ application in the management of uterine leiomyomas. Vegetos 1–8 (2025).
Rahm, C. General obstetrics and gynecology (OB-GYN) and global traditional and natural treatment options. Clin. Med. Eng. Live. 3(1), 1–38 (2025).
Panay, N. et al. Menopause and MHT in 2024: addressing the key controversies–an international menopause society white paper. South. Afr. Gen. Practitioner. 5(3), 119–134 (2024).
Bahar, E. et al. β-cell protection and antidiabetic activities of Crassocephalum crepidioides (Asteraceae) Benth. S. Moore extract against alloxan-induced oxidative stress via regulation of apoptosis and Reactive Oxygen Species (ROS). BMC Complement Altern Med. 17(1), 179 (2017).
Omotayo, M. A., Avungbeto, O., Sokefun, O. O. & Eleyowo, O. O. Antibacterial activity of Crassocephalum Crepidioides (fireweed) and Chromolaena odorata (siam weed) hot aqueous leaf extract. Int. J. Pharm. Biol. Sci. 5(2), 114–122 (2015).
Ajibesin, K. K. Ethnobotanical survey of plants used for skin diseases and related ailments in Akwa Ibom State, Nigeria. Ethnobot Res. Appl. 10, 463–522 (2012).
Dhanabal, M. V. N. L. C., Rajendran, S. P. & Rajan, I. Pharmacodynamic and ethnomedicinal uses of weed species in Nilgiris, Tamil Nadu State, india: a review. Afr. J. Agric. Res. 8(27), 3505–3527. https://doi.org/10.5897/AJAR2013.7042 (2013).
Sakpere, A. M. A., Adedeji, O. & Folashade, A. T. Flowering, postpollination development and propagation of Ebolo (Crassocephalum Crepidioides (Benth.) S. Moore) in Ile-Ife, Nigeria. Japan Sci. Technol. (Corporation). 33(2), 37–49 (2013).
Tomimori, K. et al. Antitumor activity and macrophage nitric oxide producing action of medicinal herb, Crassocephalum Crepidioides. BMC Complement. Altern. Med. 12(78), 78. https://doi.org/10.1186/1472-6882-12-78 (2012).
Ayodele, O. O., Onajobi, F. D. & Osoniyi, O. In vitro anticoagulant effect of Crassocephalum Crepidioides leaf methanol extract and fractions on human blood. J. Exp. Pharmacol. 11, 99–107. https://doi.org/10.2147/JEP.S218261 (2019).
Ayodele, O. O., Onajobi, F. D. & Osoniyi, O. Modulation of blood coagulation and hematological parameters by Crassocephalum crepidioides leaf methanol extract and fractions in STZ-induced diabetes in the rat. Sci. World J. https://doi.org/10.1155/2020/1036364 (2020).
Adjatin, A. et al. Ethnobotanical investigation and diversity of Gbolo (Crassocephalum rubens (Juss. ex Jacq.) S. Moore and Crassocephalum crepidioides (Benth.) S. Moore), a traditional leafy vegetable under domestication in Benin. Genet Resour Crop Evol. 59(8), 1867–1881. https://doi.org/10.1007/s10722-012-9901-z (2012).
Bogning, C. et al. In vitro anthelmintic activity of aqueous extract of Crassocephalum Crepidioides (Benth.) S. Moore on Haemonchus contortus. J. Exp. Integr. Med. 6(1), 31–37. 10.5455/je im.061215.or.144 (2016).
Owokotomo, I. A., Ekundayo, O., Oladosu, I. A. & Aboaba, S. A. Analysis of the essential oils of leaves and stems of Crassocephalum Crepidioides growing in South Western Nigeria. Int. J. Chem. 4(2), 34–37. https://doi.org/10.5539/ijc.v4n2p34 (2012).
Joshi, R. K. Study on essential oil composition of the roots of Crassocephalum Crepidioides (Benth.) S. Moore. J. Chil. Chem. Soc. 59(1), 2363–2365. 10.4067/ (2014).
Chung, C. et al. A galactolipid possesses novel cancer chemopreventive effects by suppressing inflammatory mediators and mouse B16 melanoma. Cancer Res. 67, 6907–6915 (2007).
Wijaya, S., Nee, T. K., Jin, K. T., Din, W. M. & Wiart, C. Antioxidant, anti-inflammatory, cytotoxicity and cytoprotection activities of Crassocephalum Crepidioides (Benth.) S. Moore. Extracts and its phytochemical composition. Eur. J. Sci. Res. 67(1), 157–165 (2011).
Fadogba, O. A., Ogunlakin, A. D., Ajayi, A. M. & Sonibare, M. A. Antioxidant and anti-arthritic activity of Bombax buonopozense P. Beauv. Leaves. Ann. Pharm. Fr. 82, 673–684 (2024).
Ogunlakin, A. D. et al. 3-(4-methoxyphenyl) acrylic acid halts redox imbalance and modulate purinergic enzyme activity in iron-induced testicular injury. Pure Appl. Chem. 96(5), 757–765 (2024).
Olanrewaju, A. A. et al. Nutritional, phytochemistry, antioxidant, and antidiabetic potentials of Hippocratea velutina (Afzel.) leaves: in vitro, ex vivo and in Silico studies. Phytomedicine Plus. 4(4), 100638 (2024).
Oboh, G., Ademiluyi, A. O. & Akinyemi, A. J. Inhibition of acetylcholinesterase activities and some pro-oxidant induced lipid peroxidation in rat brain by two varieties of ginger (Zingiber officinale). Exp. Toxicol. Pathol. 64(4), 315–319 (2012).
Elufioye, T. O., Obuotor, E. M., Sennuga, A. T., Agbedahunsi, J. M. & Adesanya, S. A. Acetylcholinesterase and butyrylcholinesterase inhibitory activity of some selected Nigerian medicinal plants. Revista Brasileira De Farmacognosia. 20, 472–477 (2010).
Oboh, G. et al. Cabbage and cucumber extracts exhibited anticholinesterase, antimonoamine oxidase and antioxidant properties. J. Food Biochem. 41(3), e12358 (2017).
Nasir, A. et al. Evaluation of alpha-amylase inhibitory, antioxidant, and antimicrobial potential and phytochemical contents of polygonum hydropiper L. Plants 9(7), 852 (2020).
Bouslamti, M. et al. Phenolic profile, inhibition of α-amylase and α-glucosidase enzymes, and antioxidant properties of solanum elaeagnifolium cav.(Solanaceae): in vitro and in Silico investigations. Processes 11(5), 1384 (2023).
Nair, A. B. & Jacob, S. A simple practice guide for dose conversion between animals and human. J. basic. Clin. Pharm. 7(2), 27 (2016).
Ogunlakin, A. D. et al. Therapeutic effects of Artocarpus communis JR Forst. & G. Forst. Seeds on letrozole-induced polycystic ovary syndrome Wistar rats. Phytomedicine Plus. 4(3), 100583 (2024).
Morris, G. M. et al. AutoDock4 and AutoDockTools4: automated Docking with selective receptor flexibility. J. Comput. Chem. 30(16), 2785–2791 (2009).
O’Boyle, N. M. et al. Open babel: an open chemical toolbox. J. Cheminform. 3, 1–4 (2011).
Trott, O. & Olson, A. J. AutoDock vina: improving the speed and accuracy of Docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31(2), 455–461 (2010).
Bekker, H. et al. World Scientific Publishing,. Gromacs-a parallel computer for molecular-dynamics simulations. In 4th international conference on computational physics (PC 92) 252–256 (1993).
Oostenbrink, C., Villa, A., Mark, A. E. & Van Gunsteren, W. F. A biomolecular force field based on the free enthalpy of hydration and solvation: the GROMOS force-field parameter sets 53A5 and 53A6. J. Comput. Chem. 25(13), 1656–1676 (2004).
Abraham, M. J. et al. GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1, 19–25 (2015).
Lee, J. et al. CHARMM-GUI input generator for NAMD, GROMACS, AMBER, OpenMM, and CHARMM/OpenMM simulations using the CHARMM36 additive force field. Biophys. J. 110(3), 641 (2016).
Lee, J. et al. CHARMM-GUI supports the amber force fields. J. Chem. Phys. 153(3) (2020).
Gyebi, G. A., Ogunyemi, O. M., Ibrahim, I. M., Afolabi, S. O. & Adebayo, J. O. Dual targeting of cytokine storm and viral replication in COVID-19 by plant-derived steroidal pregnanes: an in Silico perspective. Comput. Biol. Med. 134, 104406 (2021).
Ogunyemi, O. M. et al. Dietary stigmastane-type saponins as promising dual-target directed inhibitors of SARS-CoV-2 proteases: A structure-based screening. RSC Adv. 11(53), 33380–33398 (2021).
Ogunyemi, O. M. et al. Identification of promising multi-targeting inhibitors of obesity from Vernonia amygdalina through computational analysis. Mol. Diversity. 27(1), 1–25 (2023).
Miller, I. I. I. B. R. et al. MMPBSA. Py: an efficient program for end-state free energy calculations. J. Chem. Theory Comput. 8(9), 3314–3321 (2012).
Valdés-Tresanco, M. S., Valdés-Tresanco, M. E., Valiente, P. A. & Moreno, E. gmx_MMPBSA: a new tool to perform end-state free energy calculations with GROMACS. J. Chem. Theory Comput. 17(10), 6281–6291 (2021).
Shehna, S., Sreelekshmi, S., Remani, P. R., Padmaja, G. & Lakshmi, S. Anti-cancer, anti-bacterial and anti- oxidant properties of an active fraction isolated from curcuma Zedoaria rhizomes. Phytomed Plus 2, 100195 (2022).
Li, X. et al. Zingiberis rhizoma recens: A review of its traditional uses, phytochemistry, pharmacology, and toxicology. Evid. Based Complement. Alternat Med. 6668990. https://doi.org/10.1155/2021/6668990 (2021).
Muruthi, C. W., Ngugi, M. P., Runo, S. M. & Peter Githaiga M. In vitro antioxidant activities of Carissa edulis ((Forssk) Vahl) and Pappea capensis (Eckyl. & Zeyh) extracts. Heliyon (2023). https://doi.org/10.1016/j.heliyon.2023.e12965
Aryal, S. et al. Total phenolic content, flavonoid content and antioxidant potential of wild vegetables from Western Nepal. Plants 8(4), 96 (2019).
Saleem, A., Saleem, M. & Akthar, M. F. Antioxidant, anti-inflammatory and antiarthritic potential of Moringa Oleifera lam: an ethnomedicinal plant of Moringaceae family. South Afr. J. Bot. 128, 246–256. https://doi.org/10.016/j.sajb.20119.11.023 (2020).
Al-Owaisi, A. et al. In vitro detoxification of aflatoxin B1 by aqueous extracts of medicinal herbs. Life 15, 314–324. https://doi.org/10.1080/26895293.2022.2049900 (2022).
Yusuf, M., Sadiya, Ahmed, B. & Gulfishan, M. Modern perspectives of Curcumin and its derivatives as promising bioactive and pharmaceutical agents. Biointerface Res. Appl. Chem. 12(6), 7177–7204 (2022).
Sidor, A. & Gramza-Michałowska, A. Black Chokeberry Aronia melanocarpa L.: A qualitative composition, phenolic profile and antioxidant potential. Molecules 24, 3710 (2019).
Ajiboye, T. O. et al. Involvement of oxidative stress in Protocatechuic acid-mediated bacterial lethality. Microbiol. Open. 6(4), e00472 (2017).
Syeda, A. M. & Riazunnisa, K. Data on GC-MS analysis, in vitro antioxidant and anti-microbial activity of the catharanthus roseus and Moringa Oleifera leaf extracts. Data Brief. 29, 105258 (2020).
Zhou, J. et al. Antioxidant activities of Clerodendrum cyrtophyllum Turcz leaf extracts and their major components. PLoS ONE 15(6), e0234435 (2020).
Kalemba, M. R. K., Makhuvele, R. & Njobeh, P. B. Phytochemical screening, antioxidant activity of selected methanolic plant extracts and their detoxification capabilities against AFB1 toxicity. Heliyon 10, e24435. https://doi.org/10.1016/j.heliyon.2024.e24435 (2024).
Pavlov, V. A. & Tracey, K. J. Neural circuitry and immunity. Immunol. Res. 63, 38–57 (2015).
Sun, P. et al. Deficiency of α7 nicotinic acetylcholine receptor attenuates Bleomycin-Induced lung fibrosis in mice. Mol. Med. 23, 34–49. https://doi.org/10.2119/molmed.2016.00083 (2017).
Maggiorani, D. et al. Monoamine Oxidases, oxidative Stress, and altered mitochondrial dynamics in cardiac ageing. Oxid. Med. Cell. Longev. 2017, 3017947. https://doi.org/10.1155/2017/3017947 (2017).
Lijnen, P. J., van Pelt, J. F. & Fagard, R. H. Stimulation of reactive oxygen species and collagen synthesis by angiotensin II in cardiac fibroblasts. Cardiovasc. Ther. 30(1), e1–e8. https://doi.org/10.1111/j.1755-5922.2010.00205.x (2012).
Nabeebaccus, A., Zhang, M. & Shah, A. M. NADPH oxidases and cardiac remodelling. Heart Fail. Rev. 16(1), 5–12. https://doi.org/10.1007/s10741-010-9186-2 (2011).
Ingledine, R. et al. Excessive Glutamate receptor activation and neurological disorders. In Basic Neurochemistry: Molecular, Cellular and Medical Aspects (eds. Siegel, G. J., Agranoff, B. W. & Albers, R. W.) 6th edition ( Lippincott-Raven, 1999). https://www.ncbi.nlm.nih.gov/books/NBK28148/
Durán-Pastén, M. L. & Fiordelisio, T. GnRH-Induced Ca(2+) signaling patterns and gonadotropin secretion in pituitary Gonadotrophs. Functional adaptations to both ordinary and extraordinary physiological demands. Front. Endocrinol. 4, 127. https://doi.org/10.3389/fendo.2013.00127 (2013).
Pekdemir, B. et al. Mechanisms and potential benefits of neuroprotective agents in neurological health. Nutrients 16(24), 4368. https://doi.org/10.3390/nu16244368 (2024).
Marques, P. et al. Physiology of GnRH and Gonadotrophin Secretion. [Updated 2024 Oct 15]. In Endotext (eds. Feingold, K. R., Ahmed, S. F. & Anawalt, B.) (MDText.com, Inc., 2000). https://www.ncbi.nlm.nih.gov/books/NBK279070/
Kirilov, M. et al. Dependence of fertility on kisspeptin-Gpr54 signaling at the GnRH neuron. Nat. Commun. 4, 2492. https://doi.org/10.1038/ncomms3492 (2013).
Wang, D. et al. KP-10/Gpr54 attenuates rheumatic arthritis through inactivating NF-κB and MAPK signaling in macrophages. Pharmacol. Res. 171, 105496. https://doi.org/10.1016/j.phrs.2021.105496 (2021).
Magata, F., Tsukamura, H. & Matsuda, F. The impact of inflammatory stress on hypothalamic Kisspeptin neurons: mechanisms underlying inflammation-associated infertility in humans and domestic animals. Peptides 162, 170958. https://doi.org/10.1016/j.peptides.2023.170958 (2023).
Wang, Y., Chen, F., Ye, L., Zirkin, B. & Chen, H. Steroidogenesis in Leydig cells: effects of aging and environmental factors. Reprod. (Cambridge England). 154(4), R111–R122. https://doi.org/10.1530/REP-17-0064 (2017).
Ye, L., Zhao, B., Hu, G., Chu, Y. & Ge, R. S. Inhibition of human and rat testicular steroidogenic enzyme activities by bisphenol A. Toxicol. Lett. 207(2), 137–142. https://doi.org/10.1016/j.toxlet.2011.09.001 (2011).
Guo, J., Ma, F., Li, Y., Ge, R. & Wu, A. Curcumin ameliorates Hydrocortisone-Induced Leydig cell injury and testicular dysfunction by upregulating the Nrf2 pathway. Andrologia 2025(1), 5596226 (2025).
Haverfield, J. T., Ham, S., Brown, K. A., Simpson, E. R. & Meachem, S. J. Teasing out the role of aromatase in the healthy and diseased testis. Spermatogenesis 1(3), 240–249. https://doi.org/10.4161/spmg.1.3.18037 (2011).
Cannarella, R. et al. Molecular insights into Sertoli cell function: how do metabolic disorders in childhood and adolescence affect spermatogonial fate? Nat. Commun. 15(1), 5582. https://doi.org/10.1038/s41467-024-49765-1 (2024).
Elegheert, J. et al. Structural basis for integration of GluD receptors within synaptic organizer complexes. Science 353(6296), 295–299 (2016).
Bornot, A., Etchebest, C. & De Brevern, A. G. Predicting protein flexibility through the prediction of local structures. Proteins Struct. Funct. Bioinform. 79(3), 839–852 (2011).
Narwani, T. J. et al. In Silico prediction of protein flexibility with local structure approach. Biochimie 165, 150–155 (2019).
Goldenberg, D. P. Computational simulation of the statistical properties of unfolded proteins. J. Mol. Biol. 326(5), 1615–1633 (2003).
Durham, E., Dorr, B., Woetzel, N., Staritzbichler, R. & Meiler, J. Solvent accessible surface area approximations for rapid and accurate protein structure prediction. J. Mol. Model. 15, 1093–1108 (2009).
Mlodawska, O. W. et al. Epigenomic and enhancer dysregulation in uterine leiomyomas. Hum. Reprod. Update. 28(4), 518–547 (2022).
Du, X. et al. Insights into protein–ligand interactions: mechanisms, models, and methods. Int. J. Mol. Sci. 17(2), 144 (2016).
Acknowledgements
None.
Author information
Authors and Affiliations
Contributions
Conceptualization: ADO; Formal Analysis: ADO, OA, DSA, OVO, HB, POA, ADA, GAG,; Software: ADO, GAG, POA, and OAO; Writing (original draft): all authors; Writing (review & editing): ADO, GAG, and OSA; Supervision: ADO.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
Ethical approval was obtained from the Institutional Animal Ethics Committee of the Department of Biochemistry at Bowen University, Iwo (BUI/BCH/2025/001) before this study.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ogunlakin, A.D., Asaleye, O., Anejukwo, D.S. et al. Understanding the mechanism of Crassocephalum crepidiodes (Benth.) S. Moore leaf antifibrotic activity using in vivo and in silico methods. Sci Rep 16, 4365 (2026). https://doi.org/10.1038/s41598-025-34489-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-34489-z