Abstract
Breast cancer remains a major global health challenge with high incidence and mortality rates among women. Recent studies have highlighted the critical role of the tumor microenvironment, particularly cancer-associated fibroblasts (CAFs), in tumor progression. However, current understanding of CAFs heterogeneity and its implications for breast cancer diagnosis and treatment remains limited. This study aimed to identify and validate refined marker genes for CAFs and to develop a diagnostic model to improve breast cancer diagnosis and therapeutic strategies. We employed various machine learning algorithms to identify feature genes associated with CAFs. Based on these genes, we constructed a high-precision diagnostic model for breast cancer. Furthermore, through single-cell analysis, we delved into the heterogeneity of CAFs and predicted the sensitivity of different CAF subsets to specific drugs. To validate the expression of these characteristic genes, immunohistochemical (IHC) experiments were also conducted. This study used machine learning to identify FXYD1, SULF1, and TNXB as refined biomarkers for CAFs in breast cancer. Among these evaluated algorithms, the Random Forest algorithm distinctly stood out as the best due to its robust classification accuracy and stability. Single-cell analysis provided insights into the heterogeneity of CAFs between Luminal and non-Luminal breast cancer, thereby enhancing our understanding of the tumor microenvironment. Drug sensitivity predictions indicated that distinct CAF subsets responded differently to specific drugs, laying a solid foundation for the development of personalized breast cancer treatment strategies. Through IHC, the expression patterns of these three biomarkers were verified: FXYD1 was expressed in myoepithelial and fibroblasts in normal breast tissue but was significantly absent in breast cancer; SULF1 was upregulated in fibroblasts of breast cancer; while the expression of TNXB did not exhibit notable variations between normal and cancerous tissues. These findings not only highlight the crucial roles played by FXYD1, SULF1, and TNXB in the development of breast cancer, but also uncover the heterogeneity CAFs. Consequently, our research provides a fresh perspective and a solid theoretical basis for advancing both early and precise diagnostic methods, as well as tailored therapeutic strategies.
Similar content being viewed by others
Introduction
Breast cancer, a prevalent malignancy among women globally, continues to exhibit high incidence and mortality rates, posing a significant public health challenge1,2. The tumor microenvironment (TME) serves as the fertile ground for tumor cell development and progression, supported by numerous studies3,4,5. Within this intricate ecosystem, cells can be broadly categorized into immune cells and stromal cells. Among stromal cells, a growing body of evidence underscores the pivotal role of specific subsets, particularly CAFs, in tumor progression6,7,8. CAFs have emerged as a central player, with multiple studies elucidating their essential functions in cancer proliferation, advancement, and invasion9. Existing research demonstrates that CAFs interact intimately with cancer cells and play a crucial role in mediating and facilitating the metastasis of breast cancer10,11,12.
The emerging evidence reveals that the paradigms of cancer-centric therapeutics have limited therapeutic options in the clinic13. Consequently, there is a pressing necessity for a deeper exploration of CAF heterogeneity. Current research endeavors aimed at CAF classification and marker identification, though ongoing, remain limited in scope, with minimal translation into clinical practice14. This study aimed to identify refined CAF-related markers by applying advanced machine learning algorithms to single-cell and bulk datasets. Consequently, we have identified three signature genes: FXYD1 (fxyd domain-containing transport regulator 1), SULF1, and TNXB, which have received limited attention in breast cancer research to date.
FXYD1, a crucial regulator of ion channel transport, encodes the phospholemman (PLM) protein, which plays a vital role in heart and brain tissue15. Given its importance in these critical systems, its potential involvement in breast cancer pathogenesis merits further examination. SULF1, a sulfatase enzyme, modulates tumor development by influencing the binding affinity of cell surface heparan sulfate proteoglycans16. Similarly, TNXB, an extracellular matrix protein, contributes to collagen network assembly and tissue integrity17. This study delves into the mechanisms of these genes in breast cancer initiation and progression through comprehensive analyses that encompass gene expression patterns, copy number alterations (CNAs), functional evaluations, and drug sensitivity predictions. Each of these components contributes to a holistic understanding of the genes’ roles in breast cancer development.
Immunohistochemical validation of these markers in both benign and malignant breast tissue samples provided a robust theoretical basis for advancing the diagnosis of breast cancer. However, to translate these groundbreaking discoveries into tangible clinical benefits, further rigorous clinical validation was imperative. Additionally, this study presented novel insights into CAF heterogeneity, uncovering promising avenues for the tailored development of therapeutic strategies.
We anticipate that our findings will establish a solid scientific foundation for earlier diagnosis, more accurate prognosis assessment, and the accelerated development of personalized therapeutic strategies. Ultimately, by elucidating the expression patterns of FXYD1, SULF1, and TNXB in breast cancer progression, we aim to enhance patient outcomes and quality of life, thereby paving the way for the development of targeted therapies that can more effectively address this devastating disease.
Results
Identification of feature genes of CAFs in breast cancer by machine learning
Three previous single-cell studies on breast cancer18,19,20 provided lists of differentially expressed genes for each cell subtype in their supplementary materials. We downloaded these files and extracted the CAFs subtype-specific differentially expressed genes. Next, we intersected these three lists to identify genes consistently reported across all studies. Subsequently, to further refine our candidate gene list, we intersected this consensus gene set with differentially expressed genes (DEGs) between breast cancer and normal breast tissues. Notably, these DEGs were downloaded from the GEPIA2 website (http://gepia2.cancer-pku.cn/)21. This dual intersection approach allowed us to identify 28 highly promising candidate genes (Fig. 1A, Supplementary Table 2).
Feature selection by machine learning. (A) Integration of CAFs-associated gene lists from 3 studies with TCGA breast cancer DEGs revealed 28 candidate CAFs markers. (B) Among six evaluated feature selection methods, random forest emerged as the optimal choice, achieving high accuracy and stability; it also determined the optimal number of variables to be 3. (C) The Sankey diagram presents the optimal variables selected by different algorithms during the feature selection process. (D-E) Boxplots display expressions of feature genes in GSE65194 and GSE233242, respectively. Both datasets underwent PCA and tSNE analyses, effectively differentiating cancer from normal breast tissues.
Next, we used the caret package to perform feature selection on the TCGA-Train dataset. After evaluating the performance of six built-in feature selection methods, the random forest algorithm emerged as the most suitable due to its superior classification accuracy and stability, determining the optimal number of variables to be 3 (Fig. 1B). To gain a deeper understanding of the results, we visually demonstrated them in the form of a Sankey diagram (Fig. 1C) and tables (Supplementary Tables 3–4), which clearly showed the process of feature selection.
In-depth analysis of the TCGA datasets revealed that FXYD1 and TNXB were significantly downregulated in breast cancer tissues, while SULF1 was significantly upregulated (Supplementary Fig. 1). This abnormal expression pattern strongly suggests the potential key roles of these three genes in the development of breast cancer.
To further validate the effectiveness of these feature genes in distinguishing cancer from normal breast tissues, we performed PCA and tSNE dimensionality reduction analyses. The results were compelling: Cancer tissues were effectively differentiated from normal breast tissues based on the expression profiles of these genes (Supplementary Fig. 1). Additionally, we conducted external validations using two independent datasets, GSE65194 and GSE233242. The results revealed consistent results with the TCGA dataset (Fig. 1D-E), further reinforcing our findings.
To clarify whether FXYD1, TNXB, and SULF1 are unique to breast cancer progression or exhibit conserved expression across other malignancies, we analyzed their pan-cancer expression patterns using GEPIA221. Consistent with their expression patterns in breast cancer, FXYD1 and TNXB were downregulated in most cancer types, while SULF1 was upregulated (Supplementary Fig. 2). These results indicate their abnormal expression is not restricted to breast cancer, supporting broader implications in tumorigenesis beyond breast-specific progression.
Subsequently, we conducted an extensive exploration of the clinical and prognostic implications of these three genes across diverse datasets utilizing the BEST portal22. It revealed significant correlations between the expression of FXYD1 and TNXB and breast cancer grade, with both genes exhibiting decreased mRNA levels as the grade increased. However, the expression levels of the three feature genes did not differ significantly among tumors with varying ER, PR, or HER2 statuses. Moreover, no significant associations were found between the three genes and patient outcomes, including overall survival (OS), disease-free survival (DFS), relapse-free survival (RFS), disease-specific survival (DSS), and progression-free survival (PFS). To avoid redundancy, detailed results are available at the BEST website.
Model construction and comparison of diagnostic performance
Using FYXD1, SULF1, and TNXB as feature genes, we establish diagnostic models utilizing various algorithms to distinguish between breast cancer and normal tissue. The performance of the different models across the datasets is summarized in Supplementary Table 5. Across the internal datasets, all models exhibit robust performance, achieving AUC and accuracy scores exceeding 0.9. Notably, the RF model stands out, demonstrating particularly significant performance (Supplementary Fig. 3A-B and Supplementary Table 5). Specifically, on the TCGA-test and TCGA-all datasets, the RF model achieves AUC values of 0.9941 and 0.9944, respectively, and accuracy values of 0.9655 and 0.9319. The high true positive and true negative rates in the confusion matrices further validate its excellent diagnostic capability (Supplementary Fig. 3 A). The SVM and XGB models also performed well on the testing datsets. However, the GLM and NB models slightly lag behind on the internal validation datsets (Supplementary Fig. 3A-B and Supplementary Table 5). Importantly, even in the imbalanced TCGA-all dataset, where cancer samples significantly outnumber normal ones, all models demonstrate exceptional ability in identifying minority class samples (i.e., normal tissue), as evidenced by the high PR-AUC values (Supplementary Fig. 3 C). This finding underscores the robustness of our models in handling imbalanced data.
When applying these models to the external validation datsets GSE233242 and GSE65194, results varied. On GSE233242, the AUC and accuracy of the RF model decreased slightly but remained within an acceptable range (AUC = 0.8732, accuracy = 0.6744). In contrast, the SVM and KNN models saw significant decline in performance, almost losing their predictive power (Fig. 2A-B, Supplementary Table 5). On GSE65194, the RF model maintained its superior performance (AUC = 0.904, accuracy = 0.9085), while the GLM and KNN models showed notable improvements. However, the SVM model struggled to maintain its initial performance. Notably, despite achieving a high true positive rate on GSE65194, the NB model had an extremely limited ability to recognize normal samples (correctly identifying only one case) (Fig. 2A, Supplementary Table 5). Based on the PR-AUC values from both external datasets, the RF model remained the top performer (Fig. 2C).
Diagnostic models’ performance across six machine learning algorithms on external validation sets. (A) Confusion matrices for six machine learning algorithms on GSE65194 and GSE233242 datasets. ROC Curves (B) and PR-RPC Curves (C) of Different Models. Mean absolute SHAP values and detailed SHAP values of feature genes in GSE65194 (D) and GSE233242 (E).
Upon comprehensive analysis of these results, the RF model not only excelled on internal datasets but also demonstrated robust generalization ability on external validation sets, further emphasizing the pivotal role of FYXD1, SULF1, and TNXB as feature genes in breast cancer diagnosis. Moving beyond the assessment of the model’s performance, we conducted an in-depth analysis of Shapley Additive exPlanations (SHAP) values, revealing variations in the importance of these feature genes across the diverse datasets. In contrast to TNXB’s consistent prominence in both the testing and internal validation sets, FYXD1 showed a distinct lead in the rankings of the two external validation sets (Fig. 2D-E; Supplementary Fig. 3D-E). We primarily attributed this discrepancy to the inherent diversity in data distributions across different datasets. Having explored the diagnostic potential of these three feature genes to some extent, we intend to further investigate their potential in subsequent sections of this paper.
CNA analysis
In the TCGA database, a comprehensive analysis was conducted on the variations of the three feature genes: FXYD1, SULF1, and TNXB. Specifically, FXYD1 alterations were observed in 44 samples (approximately 5% of the cohort), primarily manifesting as amplification (2.5%, n = 24) and mRNA high (1.67%, n = 16) (Supplementary Fig. 4A-B). Likewise, SULF1 exhibited a broader spectrum of variations across 114 samples (approximately 12%), with amplification being the most prevalent (9.27%, n = 89), reinforcing its potential significance in tumorigenesis and progression (Supplementary Fig. 4A-B). The variation landscape of TNXB was comparatively intricate, with alterations detected in 45 samples (approximately 5%), encompassing mutation (0.94%, n = 9), amplification (1.04%, n = 10), mRNA high (1.15%, n = 12), mRNA low (1.15%, n = 11), and multiple alterations (0.31%, n = 3) (Supplementary Fig. 4A-B). These diverse variation patterns may mirror the multifaceted roles played by TNXB in tumor biology.
Shifting attention to the METABRIC dataset, we observed similar yet distinct trends. Variations in FXYD1 were detected in 104 samples (approximately 6%), with amplification (1.93%, n = 36) and mRNA high (3.59%, n = 67) remaining the predominant forms (Supplementary Fig. 5A-B). Notably, the frequency of SULF1 variations significantly increased, observed in 384 samples (approximately 21%), with amplification accounting for the vast majority (15.86%, n = 296), further corroborating the high prevalence of SULF1 variations in breast cancer (Supplementary Fig. 5A-B). The variation pattern of TNXB in the METABRIC dataset mirrored that in TCGA, but with distinct numerical specifics—specifically, amplification was observed in 0.86% of samples (n = 16), mRNA high in 2.84% (n = 53), and mRNA low in 1.23% (n = 23) (Supplementary Fig. 5A-B).
In terms of survival analysis, no significant associations were observed between genetic alterations in all feature genes and either OS or RFS in the TCGA dataset (Supplementary Fig. 4C-D). However, interestingly, in the METABRIC dataset, genetic alterations in FXYD1 were significantly associated with improved OS; specifically, the altered group exhibited significantly better prognosis compared to the non-altered group, suggesting a potentially favorable prognostic effect of FXYD1 variations (Supplementary Fig. 5C-D). In contrast, SULF1 was negatively correlated with DFS, with the non-altered group faring better, which may be attributed to the promoting role of SULF1 in tumor progression (Supplementary Fig. 5C-D).
The expression patterns of the feature genes at the single-cell resolution
In this section, we executed a series of systematic strategies for cell classification. Initially, based on the specific expression patterns of EPCAM and PTPRC, cells were categorized into three distinct groups: epithelial cells, immune cells, and a non-specific stroma cell population (Supplementary Fig. 6). UMAP plots show the expression patterns of three feature genes within the stroma cells: Specifically, FXYD1 expression was significantly downregulated in cancerous tissues compared to adjacent normal tissues; conversely, SULF1 expression was markedly upregulated; whereas TNXB did not display any notable difference in expression levels between cancerous and normal tissues, thereby providing essential insights for our subsequent investigations (Supplementary Fig. 6).
To comprehensively unravel the heterogeneity of stroma cells, we conducted an extensive secondary clustering analysis, refining them into three major subpopulations: EPCAM1 + endothelial cells, RGS5 + pericytes, and PDGFRA + fibroblasts (Fig. 3A). Notably, although endothelial cells from both normal and cancerous tissues exhibited overlap in their expression profiles, posing a challenge for clear distinction, pericytes and fibroblasts could be distinctly categorized based on tissue type (Fig. 3B). Within the stromal cell subpopulations, cancer cells from different types of breast cancer were intermixed, lacking distinct subpopulation differentiation or heterogeneity (Fig. 3C). Through further refined clustering analysis, we segmented the stroma cells into five specific subpopulations: endothelial cells; normal pericytes; cancer pericytes; normal fibroblasts; and CAFs (Fig. 3D). Specifically, FXYD1 was predominantly expressed in normal fibroblasts and, to a lesser degree, in pericytes; SULF1 was enriched primarily in CAFs; and TNXB was expressed in both normal fibroblasts and CAFs (Fig. 3E-F).
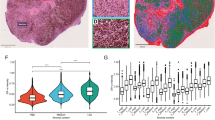
Stroma cell heterogeneity. (A) Stroma cells can be categorized into three major subpopulations: endothelial cells, pericytes, and fibroblasts. (B) Distributions of normal and cancerous samples across stroma subpopulations. (C) Distributions of different types of breast cancer across stroma subpopulations. (D) The stroma cell subpopulations were further refined based on the distributions of benign and malignant samples. UMAP plots (E) and violin plots (F) show the expression levels of selected genes. (G) Fibroblasts were extracted and subjected to dimensional reduction and clustering, yielding normal fibroblasts, iCAFs, and mCAFs, UMAP plots and violin plots show the expression levels of selected genes. (H) Trajectory reconstruction illuminates the differentiation process of fibroblasts in breast cancer. (I) Feature genes show varied expression patterns during the trajectory reconstruction. (G-K) CAFs exhibit heterogeneity in breast cancer. (L) Dot plots show the expression levels of selected genes across different CAFs subpopulations.
To gain a deeper understanding of the molecular mechanisms underlying the transformation of fibroblasts into CAFs, we further subdivided the fibroblast population, distinguishing three key subpopulations: normal fibroblasts; myofibroblastic CAFs (mCAFs), marked by ACTA2 expression; and inflammatory CAFs (iCAFs), characterized by CXCL14 expression. Notably, we found that FXYD1 and TNXB were more prominently expressed in iCAFs, suggesting a potential link to their inflammatory regulatory roles within the tumor microenvironment. Conversely, SULF1 was preferentially enriched in mCAFs, indicating its pivotal role in the development of myofibroblastic CAFs (Fig. 3G).
To dynamically simulate the transition from normal fibroblasts to CAFs, we employed advanced pseudotime analysis techniques. Our findings indicate that mCAFs occupy the terminal stage of development. Notably, during this transition, FXYD1 expression gradually diminishes, which may correlate with the loss of certain functions as fibroblasts transform into CAFs. Conversely, TNXB expression exhibits an initial surge followed by a decline, mirroring the dynamic shifts in extracellular matrix remodeling that accompany the transition. Furthermore, SULF1 expression consistently intensifies, emphasizing its central role in CAF development and functional preservation (Fig. 3H-I). Upon deeper exploration of CAF subdivision, we observed marked heterogeneity between Luminal and non-Luminal breast cancer CAF populations (Fig. 3J), allowing for their classification into four distinct subgroups: Luminal iCAFs, Luminal mCAFs, non-Luminal iCAFs, and non-Luminal mCAFs (Fig. 3K). Finally, a bubble plot visually represents the expression profiles of ACTA2, CXCL14, and CAF-specific genes across these diverse subpopulations, revealing their unique expression signatures within the CAF subgroups (Fig. 3L).
Functional analysis
Through GO and KEGG analyses, we have unraveled the unique biological functions and pathways associated with different types of fibroblasts in breast cancer. For the functional analysis, we present only the top 10 results (Fig. 4), with marker genes for each fibroblast type listed in Supplementary Tables 6 and 7. Our analysis underscores the complexity of CAFs’ roles within the tumor microenvironment. Specifically, mCAFs play a pivotal role in extracellular matrix remodeling and nutritional support, whereas normal fibroblasts are intimately linked to immune responses and inflammatory processes, potentially maintaining immune homeostasis via signaling pathways such as IL-17 and TNF. iCAFs play a pivotal role in regulating inflammation, immune responses, and cellular signaling, crucial for both physiological homeostasis and pathological conditions. Upon further examination, distinct functional characteristics between Luminal mCAFs and non-Luminal mCAFs have been discerned regarding protein synthesis and immune modulation. Notably, Luminal mCAFs exhibit significant enrichment in pathways related to ribosomal function, emphasizing their crucial role in protein synthesis. Conversely, non-Luminal mCAFs demonstrate greater enrichment in pathways associated with autoimmune diseases and pathogen infections, suggesting unique functions in immune regulation and resistance to infections. Regarding iCAFs, Luminal iCAFs are prominently associated with inflammation- and tumor-related signaling pathways, indicating their pro-inflammatory and pro-tumorigenic effects within the tumor microenvironment. Meanwhile, non-Luminal iCAFs are enriched in pathways linked to complement and coagulation cascades, as well as cytokine-receptor interactions, highlighting their significant roles in regulating inflammatory responses and blood coagulation. These results underscore not only the functional diversity of CAFs in the cancer microenvironment but also pave the way for novel research avenues and potential therapeutic interventions.
Functional analysis. (A) GO and KEGG analyses were performed using marker genes of normal fibroblasts, iCAFs, and mCAFs, respectively showing the top 10 enriched outcomes for each subpopulation. (B) GO and KEGG analyses were performed using marker genes of different CAFs, respectively showing the top 10 enriched outcomes for each subpopulation.
Drug sensitivity prediction
Based on BCs, we successfully subdivided Luminal CAFs into four subgroups (TC0 to TC3) and non-Luminal CAFs into three subgroups (TC0 to TC2). However, due to the scarcity of TC2 subgroup cells in non-Luminal CAFs, we excluded the analysis results for this subgroup. To provide an intuitive illustration, we employed UMAP dimensionality reduction plots to showcase the distribution of these distinct CAF types in the reduced space (Fig. 5A and G). Additionally, regarding drug sensitivity prediction, we conducted detailed calculations for TCs and CAFs classifications. The detailed information of TOP Differential High Sensitivity Drugs across all classifications is listed in Supplementary Tables 8–11.
Drug Sensitivity prediction for luminal and non-luminal CAFs. luminal and non-luminal CAFs were subjected to dimensional reduction and clustering respectively, based on Beyondcell scores of individual cells. (A) UMAP plots present two views: the left depicting TC clusters specific to Luminal CAFs, the right illustrating the distribution patterns of Luminal iCAFs and Luminal mCAFs. (B) Top differential sensitivity drugs of Luminal iCAFs and mCAFs. (C) Intersection of high sensitivity drugs for Luminal mCAFs and low sensitivity drugs for Luminal iCAFs. (D) UMAP plots show top 5 high sensitivity drugs for Luminal mCAFs. (E) Intersection of high sensitivity drugs for Luminal iCAFs and low sensitivity drugs for Luminal mCAFs. (F) UMAP plots show top 5 high sensitivity drugs for Luminal iCAFs. (G) UMAP plots present two views: the left depicting TC clusters specific to non-Luminal CAFs, the right illustrating the distribution patterns of non-Luminal iCAFs and non-Luminal mCAFs. (H) Top differential sensitivity drugs of Luminal iCAFs and mCAFs. (I) Intersection of high sensitivity drugs for non-Luminal mCAFs and low sensitivity drugs for non-Luminal iCAFs. (J) UMAP plots show top 5 high sensitivity drugs for non-Luminal mCAFs. (K) Intersection of high sensitivity drugs for non-Luminal iCAFs and low sensitivity drugs for non-Luminal mCAFs. (L) UMAP plots show top 5 high sensitivity drugs for non-Luminal iCAFs.
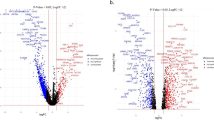
Firstly, concerning the drug sensitivity prediction results for TC classifications, we present the top 5 differential high sensitivity drugs in each TC cluster through volcano plots (Supplementary Fig. 7 A and Supplementary Fig. 7 C). Furthermore, the UMAP plots show the distribution of cells sensitive to the respective top differential high sensitivity drug for each TC cluster (Supplementary Fig. 7B and Supplementary Fig. 7D). Specifically, in Luminal CAFs, TC0 was most sensitive to GSK-J4, TC1 to SCH-900,776, TC2 to TENIPOSIDE, and TC3 to GSK525762A. For non-Luminal CAFs, TC0 favored AZD8055, while TC1 preferred SORAFENIB. Notably, the distribution patterns of these drug-sensitive cells were highly consistent with the TC classifications.
Next, our findings provide important insights into CAF heterogeneity, providing robust theoretical support for the development of targeted therapeutic strategies aimed at specific CAF subgroups, but also significantly deepening our understanding of this complex phenomenon. Subsequently, we focused our efforts on predicting drug sensitivity within various CAF classifications. The volcano plots revealed the top five drugs with differential high sensitivity for each CAF classification (Fig. 5B and H). Intriguingly, we discovered that drugs that are sensitive to mCAFs tend to be insensitive to iCAFs, and conversely, drugs that are sensitive to iCAFs are often insensitive to mCAFs (Fig. 5C, E and I, and 5K), this finding offers a novel perspective on CAF heterogeneity. In Luminal CAFs, mCAFs exhibited sensitivity to drugs such as DASATINIB and SKI-II (Figs. 5D), whereas iCAFs responded more favorably to ENTINOSTAT and MUBRITINIB (Figs. 5F). For non-Luminal CAFs, a similar pattern of distinct drug sensitivity between mCAFs and iCAFs was observed (Figs. 5J). Notably, DASATINIB and SKI-II played pivotal roles in both Luminal and non-Luminal mCAFs (Fig. 5D and J), whereas MUBRITINIB demonstrated high sensitivity specifically to iCAFs (Fig. 5F and L). Additionally, it is noteworthy that a previous study has validated the efficacy of DASATINIB in inhibiting CAFs23, thereby further enhancing the credibility of our drug sensitivity prediction results.
Verification of the expression patterns of the feature genes by IHC
We have thoroughly analyzed the IHC results and made several significant discoveries. In normal breast tissue, FXYD1 protein is predominantly located in myoepithelial and stromal cells (Supplementary Fig. 8A-B). However, within carcinoma in situ, a marked decline in FXYD1 expression is observed within these cells, with occasional expression noted in the peritumoral stromal area. (Supplementary Fig. 8A-B). Notably, in contrast to these findings, the expression of FXYD1 is completely absent in the invasive carcinoma cases (Supplementary Fig. 8A-B). In fibroadenomas, the proliferation of fibrous tissue specifically leads to an enhanced expression of FXYD1 in fibroblasts, which is clearly detectable by IHC (Supplementary Fig. 9 A). By analyzing a pathological slide encompassing normal breast tissue, carcinoma in situ, and invasive carcinoma (Supplementary Fig. 10 A), we directly observe variations in FXYD1 expression across these distinct pathological stages. The gradual loss of FXYD1 expression pattern suggests a possible suppressive role in breast tumor progression. Upon investigation of SULF1 protein, we have observed a markedly elevated expression level in stromal fibroblasts within cancer tissue, in comparison to normal breast tissue and fibroadenomas (Supplementary Fig. 8 C and Supplementary Fig. 9B). Our analysis, however, did not uncover any significant correlation between SULF1 expression and various clinicopathological features (Supplementary Figs. 8D). Similarly, Supplementary Fig. 10B presents another pathological slide showcasing normal breast tissue, carcinoma in situ, and invasive carcinoma. Upon examination, this slide reveals variations in the level of SULF1 expression across these distinct pathological stages. Notably, the distinct expression pattern of SULF1 hints at a potentially significant contribution to the formation and modulation of the breast tumor microenvironment. However, no significant difference in TNXB expression was observed in benign and malignant breast tissues (Supplementary Fig. 11). This finding highlights the need for further investigation into the potential mechanisms underlying the regulation of TNXB expression in the context of breast cancer development. We have selected two consecutive histological sections of breast cancer tissue to demonstrate the distinct expression patterns of FXYD1 and SULF1. Specifically, the expression levels of FXYD1 protein are notably higher in normal breast tissue compared to those in cancerous tissue (Fig. 6A), whereas SULF1 expression levels are more pronounced in cancerous tissue (Fig. 6B). Notably, FXYD1 is expressed not only within myoepithelial cells but also prominently on vascular walls. Based on our previous single-cell analysis, which provided insights into the cellular distribution of FXYD1, we hypothesize that FXYD1 may also be expressed in perivascular cells (Figs. 3E and 6A). To attain a deeper comprehension of the expression patterns exhibited by FXYD1 and SULF1, we compared the expression patterns of FXYD1 and SULF1 with some important protein markers. The use of α-SMA (Fig. 7A), P63 (Fig. 7B), and Calponin (Fig. 7C) as markers for identifying myoepithelial cells is consistent and reliable in the daily practice of clinical pathological diagnosis. On the other hand, α-SMA and Vimentin are known as markers of CAFs. Upon close comparison, it is evident that in normal breast tissue, similar to Vimentin (Fig. 7D), FXYD1 is expressed in both myoepithelial cells and stromal fibroblasts (Fig. 7E). Notably, however, SULF1 expression is scarcely detectable within the stromal compartment of normal breast tissue (Fig. 7F). As the tissue transitions towards malignancy, resulting in carcinoma in situ, markers like P63, Calponin, α-SMA, and Vimentin continue to be expressed in myoepithelial cells; additionally, Vimentin expression is markedly elevated in fibroblasts (Fig. 8A). Conversely, FXYD1 expression is drastically reduced, becoming absent in both myoepithelial cells and fibroblasts. This inverse trend is observed with SULF1, whose expression increases in the stromal compartment, suggesting a potential tumor-suppressive function for FXYD1 and a tumor-promoting role for SULF1 during early tumorigenesis. In the context of invasive carcinoma, the persistent absence of FXYD1 expression underscores its vital role in inhibiting tumor progression (Fig. 8B). Furthermore, the notable upregulation of SULF1 expression in fibroblasts, echoing the pattern seen with α-SMA and Vimentin, underscores its potential contribution to tumor progression (Fig. 8B).
Two consecutive pathological slides intuitively demonstrate the distinct expression Patterns of FXYD1 and SULF1. (A) The expression of FXYD1 in myoepithelial cells. (B) The expression of FXYD1 in fibroblasts. (C) The expression of SULF1 in fibroblasts. (D) The relationship between SULF1 and different clinicopathological characteristics, including tissue type, age, T stage, N stage, ER status, PR status, and Her2 status.
A Comparison of the IHC results for FXYD1 and SULF1 with other markers in normal breast tissues. α-SMA (A), P63 (B), and Calponin (C) are markers of myoepithelial cells. α-SMA (A), and Vimentin (D) are markers of CAFs. The expression of FXYD1 (E) is observed in myoepithelial cells and fibroblasts. The expression of SULF1 (F) is nearly undetectable in normal mammary tissue.
A Comparison of the IHC results for FXYD1 and SULF1 with other markers in malignant breast tissues. In carcinoma in situ (A), FXYD1 expression is notably absent in myoepithelial cells and markedly downregulated in fibroblasts, whereas SULF1 expression undergoes upregulation in fibroblasts. As the disease progresses to invasive carcinoma (B), the absence of FXYD1 expression remains consistent, and the upward trend in SULF1 expression persists. Compared to other markers, these findings provide a more intuitive understanding of the expression patterns.
Discussion
This study applied multiple approaches to Recursive Feature Elimination (RFE), a statistical technique that enhances model performance by iteratively discarding the least significant features. This method effectively identified the CAF-associated genes FXYD1, SULF1, and TNXB, which are closely associated with breast cancer. This finding underscores the vital role of cancer-associated fibroblasts in breast cancer pathology and suggests these genes as promising new therapeutic targets. Among the machine learning algorithms assessed, the RF model particularly excelled, greatly improving diagnostic accuracy and paving the way for new personalized treatment options. Data analysis from the BEST database revealed noteworthy correlations between the expression of FXYD1 and TNXB and tumor grade, providing essential insights. We do not claim these genes to be entirely new, but emphasize three incremental advances: (1) a refined three-gene signature that robustly discriminates breast cancer from normal tissue across five independent cohorts (TCGA, METABRIC, GSE65194, GSE233242, GSE161529); (2) Single-cell transcriptomic analysis associated mCAFs and iCAFs—further classified as Luminal and non-Luminal subtypes—with distinct drug-sensitivity profiles that await experimental validation; and (3) IHC evidence of gradual expression changes across normal, DCIS and invasive carcinoma in an untreated clinical cohort.
FXYD family consists of seven members (FXYD1 to FXYD7), which serve as tissue-specific regulators of Na+/K+-ATPase activity in cellular membranes, influencing its function based on tissue type24. Given the well-documented roles of FXYD3, FXYD5, and FXYD6 across various cancer types25,26,27,28,29,30,31,32, we aimed to investigate the expression patterns of FXYD1 in benign and malignant breast tissues, an area that has not been thoroughly explored. FXYD1 shows a distinct expression pattern in normal tissues, with significantly higher levels in the heart, kidneys, placenta, skeletal muscle, gastrointestinal tract, and colon, while moderate levels are found in breast samples33. Research involving quantitative real-time PCR of clinical samples has indicated a notable downregulation of FXYD1 in ovarian cancer tissues, associating its overexpression with enhanced migratory and invasive characteristics of ovarian cancer cells, unrelated to proliferation34. Our current study specifically demonstrated that FXYD1 had higher immunohistochemical expression in normal breast tissue but was significantly reduced in breast cancer tissues, particularly in myoepithelial cells and CAFs. Although FXYD1 is well established as a crucial regulator of neural and cardiovascular development and exerts protective effects against pulmonary hypertension and arterial remodelling35, its role in breast cancer progression remains unknown. As FXYD1 fine-tunes Na⁺/K⁺-ATPase activity, its depletion could weaken myofibroblast contractility and thereby facilitate CAF activation and collagen re-organization. Therefore, we speculate that the down-regulation of FXYD1 may be closely related to CAF activation and the modulation of the breast-tumor microenvironment. Functional assays are required to validate this hypothesis.
The sulfatase family, comprising sulfatase 1 (SULF1) and sulfatase 2 (SULF2), is important for controlling the sulfation of heparan sulfate proteoglycans (HSPGs). This modification greatly affects various physiological and pathological functions, including cell signaling, proliferation, migration, and differentiation36,37. The importance of SULF1 in a range of cancers, including prostate, ovarian, esophageal, hepatocellular, gastric, and colon cancers, has been widely recognized38,39,40,41,42,43. The expression level of SULF1 is nearly absent in normal breast tissue and low in benign and hyperplastic lesions. In contrast, SULF1 expression significantly rises in triple-positive and triple-negative breast cancers, particularly during the later stages of tumors, where its short splice variants are the most prevalent44. Notably, A previous study45 indicated that Sulf-1 exerted three distinct tumor-suppressive effects in breast cancer: it inhibited the proliferation of Sulf-1-deficient MDA-MB-468 cells after stable re-expression of Sulf-1; it lowered tumor-derived heparan-sulfate 6-O-sulfation, thereby blunting VEGFR signaling and reducing micro-vessel density by more than 60% in nude-mouse xenografts; and it markedly induced apoptosis and necrosis of malignant cells. Mechanistically, another study demonstrated that Sulf-1 halted cell-cycle progression by down-regulating cyclin D1 via non-canonical AKT/STAT3 and ERK1/2–STAT3 pathways46. Although SULF1 is important, research on its complex relationship with CAFs has been limited until recently. A groundbreaking study has shed light on this area, showing that SULF1, a signaling molecule secreted by CAFs, promotes metastasis and cisplatin (CDDP) resistance in gastric cancer cells by binding to TGFBR3 on their surfaces, thereby activating the TGF-β signaling pathway38. Our study examined the previously overlooked relationship between SULF1 and breast cancer, revealing significant changes in SULF1 expression in breast fibroblasts throughout cancer progression. In normal breast tissues, SULF1 levels were low; however, they increased significantly in breast cancer tissues, especially in aggressive tumors. This change suggested that SULF1 may activate and enhance the functions of CAFs. As tumors transitioned from in situ to invasive stages, SULF1 expression rose in fibroblasts, potentially aligning with the increased pro-tumorigenic activities of CAFs in the tumor microenvironment. These findings underscored the importance of SULF1 as a marker of functional changes in CAFs and introduced potential molecular targets for developing therapies aimed at CAFs in breast cancer.
The tenascin family comprises four members—tenascin-C (TNC), tenascin-R (TNR), tenascin-X (TNXB), and tenascin-W (TNW)—each playing a pivotal role in diverse biological processes, including tissue regeneration, inflammatory diseases, tumorigenesis, and wound healing47. Under physiological conditions, TNXB functions as a crucial regulator of collagen deposition, fibril spacing, mechanical properties, and fibrillogenesis in various physiological contexts48,49,50. As early as 2002, researchers uncovered the intricate relationship between TNXB and fibroblasts, notably observing that B16 melanoma cells demonstrated reduced adhesion and spreading capabilities, coupled with increased detachment, when cultured on TNXB-null fibroblasts51. Recently, a pan-cancer analysis of TNXB significantly highlighted its reduced expression in breast cancer tissues compared to normal tissues, as determined by IHC52. Our bioinformatics analysis, using multiple datasets, clearly showed the downregulation of TNXB mRNA in breast cancer and identified TNXB as a marker for CAFs through single-cell analysis. However, the IHC findings did not lead to conclusive results. Therefore, the precise role of TNXB in breast cancer progression remains unclear. It is still unknown whether TNXB functions as a tumor suppressor or promoter, and how its expression in CAFs might influence the tumor microenvironment and cancer cell behavior. Our ongoing research aims to address this issue and provide a deeper insight into the interactions between TNXB, CAFs, and breast cancer.
Analysis of changes in the FXYD1, SULF1, and TNXB genes from the TCGA and METABRIC datasets provided important insights into their roles in breast cancer. Notably, the significant amplification of SULF1 in many samples highlighted its critical role in tumor development, warranting further investigation into the mechanisms behind this amplification. Interestingly, alterations in FXYD1 were associated with improved overall survival in the METABRIC dataset, suggesting a protective effect, while the TCGA dataset did not show similar positive results. Conversely, changes in SULF1 were associated with worse outcomes and shorter disease-free survival. This contrast emphasized the complex interactions between these genes in tumor progression and opened up opportunities for targeted therapies. Furthermore, the diverse patterns of TNXB alterations indicated its various contributions to the tumor microenvironment, reinforcing the need for a deeper understanding of its biological significance. Given the discrepancies across different datasets, future research should have focused on validating these findings in various populations and cancer subtypes, enhancing our knowledge of breast cancer biology and guiding the development of more effective treatments.
In this study, we conducted a comparative analysis of normal mammary fibroblasts with iCAFs and mCAFs, revealing differences in their functionalities and signaling pathways. A previous study suggested that immune modulators, including myeloperoxidase (MPO) and inflammatory cytokines such as tumor necrosis factor alpha (TNF-α), may contribute to the development of high breast density by modulating gene expression patterns and collagen production in fibroblasts, ultimately influencing the risk of breast cancer53. Researchers have previously summarized that mCAFs are primarily responsible for the generation and remodeling of the extracellular matrix, providing support and migration pathways for tumor cells; on the other hand, iCAFs influence the tumor immune microenvironment by secreting inflammatory cytokines and immunoregulatory molecules, thereby facilitating tumor immune evasion and further progression54. Our study found that normal fibroblasts serve as the “baseline,” primarily involved in regulating immune responses and inflammatory processes, while mCAFs are mainly engaged in the remodeling of the extracellular matrix, and iCAFs play a role in inflammation, immune reactions, and cellular signaling. Notably, for the first time, we introduce the distinction between Luminal and non-Luminal CAFs, highlighting the diversity and complexity of CAFs in breast cancer. Luminal CAFs exhibit a pronounced ability to promote protein synthesis and inflammatory responses, potentially accelerating tumor growth and progression. Conversely, non-Luminal CAFs contribute uniquely to immune regulation, anti-infection, and blood coagulation, which may modulate the tumor microenvironment dynamics. These discoveries not only offer novel insights and potential therapeutic targets for precision breast cancer therapy but also pave the way for future research endeavors and therapeutic interventions. By attaining a deeper comprehension of the intricate functions and functional heterogeneity of CAFs within the breast cancer microenvironment, we can devise more targeted treatment strategies, with the ultimate goal of effectively suppressing tumor growth.
In our efforts to find new therapies for breast cancer, we have identified CAFs as significant therapeutic targets. Therapeutic strategies that focus on CAFs can involve targeting their surface markers, secreted factors, metabolic pathways, epigenetic modifications, immunoregulatory roles, and mechanical characteristics, along with specific interventions for different subgroups54. However, the heterogeneity of CAFs—evident in their varied functions, phenotypic traits, and drug sensitivities across subgroups—poses challenges for any single treatment method. Therefore, it is vital to have a clear understanding of CAF classification when designing effective therapeutic strategies. By predicting sensitive drugs for iCAFs and mCAFs in Luminal and non-Luminal breast cancer subtypes, respectively, we have provided a crucial foundation for the development of targeted therapies directed at specific CAF subgroups. Critically, we have also unveiled an intriguing phenomenon of mutual exclusivity in drug sensitivity among CAF subgroups, whereby certain drugs effective against iCAFs may be ineffective for mCAFs, and vice versa. This revelation not only offers a fresh perspective on CAF heterogeneity but also lays the groundwork for formulating combinatorial therapeutic strategies. By combining drugs sensitive to different CAF subgroups, we can more effectively inhibit their tumor-promoting effects while mitigating adverse effects and minimizing the risk of drug resistance. Previous research has demonstrated that Dasatinib can substantially inhibit the growth of CAFs in lung cancer, potentially augmenting the effectiveness of anticancer therapies23. Our analysis highlights the sensitivity of Luminal mCAFs to Dasatinib. This revelation not only reinforces the credibility of our results but also emphasizes Dasatinib’s promising potential as a therapeutic target for CAFs.
This study has several limitations. First, the model’s generalizability to external cohorts and the full spectrum of CAF heterogeneity remain to be evaluated. While additional CAF subsets (e.g., apCAFs) have been reported, they could not be robustly separated in the dataset; thus, the present work concentrates on mCAFs and iCAFs. Second, functional validation of the three identified genes is lacking, so their role in breast-cancer progression remains unclear. Third, drug-sensitivity predictions were not experimentally verified, and in vitro assays of canonical CAF markers (α-SMA, P63, Calponin, Vimentin) and the new signatures were not performed. Finally, immunohistochemistry was performed in a limited cohort; larger, multi-centre samples and multi-omic integration are required to substantiate these findings and thus advance clinical translation.
Methods
Data acquisition
In our study, we utilized the TCGAbiolinks package to access TCGA-BRCA TPM (Transcripts Per Kilobase Million) data along with corresponding patient clinical profiles. We applied the following data exclusion criteria: (1) genes with low expression, defined as those having an expression level of zero in more than 10% of the samples; (2) cases with incomplete clinical information; (3) male cases. We downloaded gene expression profile data (GSE65194 and GSE233242) and corresponding clinical information from the public Gene Expression Omnibus database (GEO, http://www.ncbi.nlm.nih.gov/geo/). Additionally, we obtained a single-cell dataset (GSE161529) from the GEO database, selecting 13 normal breast tissue samples, 6 Her2-positive breast cancer tissue samples, 17 ER-positive breast cancer tissue samples, and 8 triple-negative breast cancer tissue samples.
Identification of feature genes of CAFs in breast cancer by machine learning
We employed the caret package55 to implement our machine learning pipeline, with the code being available at https://topepo.github.io/caret/index.html. Initially, we divided the paired samples from the TCGA-BRCA dataset into a training set (TCGA-train, comprising 68 pairs of cancer and normal samples) and a test set (TCGA-test, containing 29 pairs), following a 7:3 ratio. Furthermore, the entire TCGA-BRCA dataset (TCGA-all), encompassing 1076 cancer samples and 99 normal samples, served as the internal validation set to ensure robustness.
To broaden the model’s applicability and assess its generalizability, we incorporated two external validation sets: GSE65194, a microarray dataset consisting of 153 cancer samples and 11 normal samples, and GSE233242, a high-throughput sequencing dataset composed of 43 pairs of cancer and normal samples.
Prior to model training, data preprocessing was crucial. To improve model performance and reduce redundant information, we eliminated collinear variables using a threshold of r > = 0.756. Continuous variables underwent z-scoring normalization using the ‘center’ and ‘scale’ methods provided by the caret preProcess function. This standardization step was essential to ensure that all variables contributed equally to the model, regardless of their original scales.
For feature selection, we opted for the Recursive Feature Elimination (RFE) method with 10-fold cross-validation. Six algorithms—Random Forest (RF), Generalized Additive Models (GAM), Linear Discriminant Analysis (LDA), Logistic Regression (LR), Naive Bayes (NB), and Bagged Trees (BT)—built in the caret package were utilized to guide this process. Through this rigorous selection, the optimal subset of features was identified.
Survival analysis of the feature genes in breast cancer
Survival analyses were performed using the BEST database (https://rookieutopia.hiplot.com.cn/app_direct/BEST)22. Patients were stratified into high- and low-expression groups based on the median mRNA expression level of each gene (FXYD1, SULF1, and TNXB). Kaplan–Meier survival curves were generated, and differences between groups were evaluated using the log-rank test. A p-value < 0.05 was considered statistically significant.
Model construction and comparison of diagnostic performance
We employed six machine learning methods to develop diagnostic models using the TCGA-train dataset with 10-fold cross-validation. These methods include RF, NB, K-Nearest Neighbors (KNN), Generalized Linear Model (GLM), XGBoost (XGB), and Support Vector Machine (SVM). To comprehensively evaluate the performance of the models, we utilized confusion matrices (generated using the caret package), Receiver Operating Characteristic (ROC) curves (calculated with the pROC package), and Precision-Recall (PR) curves (computed with the PRROC package) on the test set, as well as the internal and external validation sets. For global interpretation of the models, we employed the fastshap package, with results visualized using the shapviz package. The code for interfacing fastshap with caret models is available at https://harpomaxx.github.io/post/shap-values/.
Copy number alteration (CNA) analysis
We conducted a comprehensive analysis of genomic data utilizing the cBioPortal platform (https://www.cbioportal.org)57,58. Our investigation encompassed two datasets: the TCGA-BRCA (Breast Invasive Carcinoma, Firehose Legacy), and the METABRIC dataset. The ‘OncoPrint’ module was used to visualize variants. The ‘Cancer Types Summary’ module was used to gain an overview of genomic alterations. To further advance our analysis, we employed the ‘Survival’ module to assess the potential correlation between alterations in feature genes and patient survival time. This enabled us to gain deeper insights into the prognostic implications of specific genomic alterations in breast cancer.
Single-cell analysis
In this section, we used Seurat v459 while referencing the data quality control protocols outlined in the ‘scCancer’60 package (code is available at https://github.com/wguo-research/scCancer). To ensure data integrity, we employed the “DoubletFinder"61 package to meticulously remove potential doublets from our dataset. Furthermore, to enhance the comprehensiveness of our analysis, we leveraged the “Harmony"62 package to seamlessly integrate data from multiple samples. Using previously validated markers from prior studies63,64, we achieved a precise delineation of distinct cell subgroups, thereby solidifying the foundation for our subsequent investigations. By using the “Findmarkers” function in Seurat, unique markers for each subgroup were identified, applying parameters as follows: min.pct = 0.1, logfc.threshold = 0.25, and p_adjust < 0.05. Subsequently, we conducted trajectory inference using the monocle3 package (code is available at https://github.com/cole-trapnell-lab/monocle3), which provided us with insights into the developmental pathways of the cells.
Moving on to enrichment analysis, we chose the ‘ClusterGVis’ package (code is available at https://github.com/junjunlab/ClusterGVis), specifically utilizing the ‘prepareDataFromscRNA’ function to transform the single-cell data. Notably, the ‘diffData’ parameter was configured to encompass the top 20 marker genes for each identified cell subgroup. Following this, we applied the ‘enrichCluster’ function to perform extensive gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analyses.
Furthermore, we used the Drug sensitivity signature collection (SSC) from the ‘beyondcell’65 package for drug sensitivity prediction (code is available https://github.com/cnio-bu/beyondcell). SSC identified transcriptional state changes occurring before and after drug treatment, by collecting and analyzing data from extensive drug sensitivity databases. Through differential expression analysis, SSC screened for gene expression patterns indicative of drug sensitivity and constructed gene expression signatures. These signatures were subsequently utilized to calculate Beyondcell scores for individual cells, accurately quantifying their sensitivity to specific drugs. Lastly, we performed cluster analysis to form treatment clusters (TC), further refining our prioritization of potential drug candidates.
Clinical sample collection
In 2023, we collected surgical samples from the Breast and Thyroid Surgery Department of Zibo Maternity and Child Health Hospital, including breast fibroadenoma, breast carcinoma in situ, and invasive breast cancer. Each cancer sample was accompanied by adjacent normal tissue taken at least 5 cm away from the respective tumor margin. The clinical and histopathologic characteristics of the patient cohort are shown in supplementary Table 1. After collecting the samples, to ensure the quality of our dataset, we applied rigorous criteria in selecting the samples for inclusion in the study. Specifically, we chose patients who had not received any prior treatment and had undergone a modified radical mastectomy for breast cancer. The study was approved by the Ethics Review Committee of Zibo Maternal and Child Health Hospital, all methods were performed in accordance with the relevant guidelines and regulations, and all participants provided written informed consent.
Immunohistochemistry (IHC)
Immunohistochemical experiments were conducted adhering to a standardized protocol. Initially, paraffin-embedded tissue sections underwent deparaffinization with xylene, followed by a series of ethanol solutions for rehydration. After antigen retrieval, using an EDTA buffer at a pH of 9.0 in a DAKO PT Link device heated to 97 °C for 20 min, the samples were cooled to 65 °C and rinsed with Tris-buffered saline. To inactivate endogenous peroxidase activity, 3% hydrogen peroxide was applied. The primary antibodies employed in this study included rabbit polyclonal antibodies specific to FXYD1, SULF1, and TNXB, sourced exclusively from Abcam, and respectively diluted at ratios of 1:200, 1:400, and 1:100. Additionally, as part of routine pathology practice, ready-to-use primary antibodies for P63, vimentin, α-SMA, and calponin, obtained from Fuzhou Maisen Biotechnology Co., Ltd., were also utilized. These latter antibodies, which are routinely employed in the pathology department for diagnostic purposes, were each applied to the sections for 20 min at room temperature. Subsequently, the sections were incubated with a secondary antibody, anti-mouse IgG from Dako, for an additional 20 min at room temperature. The final stage involved color development utilizing a DAB chromogen for a duration of one minute. The intensity and distribution of the staining signal were independently assessed by three qualified pathologists. IHC staining percentage was graded as follows: 1 = 0–25%; 2 = 26–50%; 3 = 51–75%; 4 = 76–100%. The IHC intensity was scored as follows: 0 = none (-); 1 = weak (1+); 2 = moderate (2+); 3 = strong (3+). The IHC score was calculated by multiplying the intensity and percentage scores.
Statistical analysis
In our study, all statistical analyses and graphical presentations were executed using the R software, version 4.2.2, aided by a selection of tailored R packages that met our analytical requirements. For scenarios involving nonparametric data, we employed the Wilcoxon rank-sum test for pairwise comparisons and the Kruskal-Wallis H test when multiple groups were analyzed. This rigorous methodology allowed us to accurately gauge statistical significance within our dataset, ensuring comprehensive evaluation of our results.
Data availability
The dataset of TCGA-BRCA is available at the TCGA database (https://cancergenome.nih.gov/). We obtained the dataset by using the TCGAbiolinks package in R.The datasets analyzed during the current study are available at GEO: GSE65194, GSE233242, and GSE161529.The raw data generated and/or analyzed during the current study are not publicly available for the IHC results due to ethical and privacy concerns regarding patient samples. However, key findings and analyses based on these data are reported in the manuscript, and any additional information necessary to reproduce the results may be obtained upon reasonable request to the corresponding author.
Abbreviations
- α-SMA:
-
Alpha Smooth Muscle Actin
- BC:
-
Breast Cancer
- CAF:
-
Cancer-Associated Fibroblasts
- CNA:
-
Copy Number Alteration
- DAB:
-
3,3’-Diaminobenzidine
- DEGs:
-
Differentially Expressed Genes
- EDTA:
-
Ethylenediaminetetraacetic Acid
- SSC:
-
sensitivity signature collection
- ER:
-
Estrogen Receptor
- FXYD1:
-
fxyd domain-containing transport regulator 1
- GAM:
-
Generalized Additive Models
- GEO:
-
Gene Expression Omnibus
- GLM:
-
Generalized Linear Model
- GO:
-
Gene Ontology
- HER2:
-
Human Epidermal Growth Factor Receptor 2
- HSPG:
-
Heparan Sulfate Proteoglycan
- IHC:
-
Immunohistochemistry
- iCAFs:
-
Inflammatory CAFs
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- KNN:
-
K-Nearest Neighbors
- LDA:
-
Linear Discriminant Analysis
- LR:
-
Logistic Regression
- mCAFs:
-
Myofibroblastic CAFs
- METABRIC:
-
Molecular Taxonomy of Breast Cancer International Consortium
- NB:
-
Naive Bayes
- OS:
-
Overall Survival
- PCA:
-
Principal Component Analysis
- PR:
-
Precision-Recall
- ROC:
-
Receiver Operating Characteristic
- RFE:
-
Recursive Feature Elimination
- RF:
-
Random Forest
- RFS:
-
Relapse-Free Survival
- SULF1:
-
Sulfatase 1
- SULF2:
-
Sulfatase 2
- SVM:
-
Support Vector Machine
- TCGA:
-
The Cancer Genome Atlas
- TC:
-
Treatment Cluster
- TME:
-
Tumor Microenvironment
- TNC:
-
Tenascin-C
- TNR:
-
Tenascin-R
- TNXB:
-
Tenascin-X
- TNW:
-
Tenascin-W
- tSNE:
-
t-Distributed Stochastic Neighbor Embedding
- TPKM:
-
Transcripts Per Kilobase Million
- XGB:
-
xgboost
References
Han, L. et al. LncRNA HOTTIP facilitates the stemness of breast cancer via regulation of miR-148a-3p/WNT1 pathway. J. Cell. Mol. Med. 24, 6242–6252 (2020).
Zuo, S., Yu, J., Pan, H. & Lu, L. Novel insights on targeting ferroptosis in cancer therapy. Biomark. Res. 8, 1–11 (2020).
Granucci, F. The family of LPS signal transducers increases: the arrival of chanzymes. Immunity 48, 4–6 (2018).
Zheng, S. et al. Development and validation of a stromal immune phenotype classifier for predicting immune activity and prognosis in triple-negative breast cancer. Int. J. Cancer. 147, 542–553 (2020).
Tamborero, D. et al. A pan-cancer landscape of interactions between solid tumors and infiltrating immune cell populations. Clin. Cancer Res. 24, 3717–3728 (2018).
Lambrechts, D. et al. Phenotype molding of stromal cells in the lung tumor microenvironment. Nat. Med. 24, 1277–1289 (2018).
Auciello, F. R. et al. A stromal lysolipid-autotaxin signaling axis promotes pancreatic tumor progression. Cancer Discov. 9, 617–627 (2019).
Bertero, T. et al. William. Tumor-stroma mechanics coordinate amino acid availability to sustain tumor growth and malignancy Thomas. Cell Metab 29, 124–140 (2019).
Sahai, E. et al. A framework for advancing our Understanding of cancer-associated fibroblasts. Nat. Rev. Cancer. 20, 174–186 (2020).
Gagliano, T. et al. PIK3Cδ expression by fibroblasts promotes triplenegative breast cancer progression. J. Clin. Invest. 130, 3188–3204 (2020).
Alcaraz, L. B. et al. A 9-kDa matricellular SPARC fragment released by cathepsin D exhibits pro-tumor activity in the triple-negative breast cancer microenvironment. Theranostics 11, 6173–6192 (2021).
Al-Ansari, M. M., Hendrayani, S. F., Shehata, A. I. & Aboussekhra, A. P16 INK4A represses the paracrine tumor-promoting effects of breast stromal fibroblasts. Oncogene 32, 2356–2364 (2013).
Yang, P. et al. CAF-derived exosomal WEE2-AS1 facilitates colorectal cancer progression via promoting degradation of MOB1A to inhibit the Hippo pathway. Cell Death Disease 13(9), 796 (2022).
Chen, X. & Song, E. Turning foes to friends: targeting cancer-associated fibroblasts. Nat. Rev. Drug Discovery. 18, 99–115 (2019).
Cuomo, M. et al. Epigenetic remodelling of Fxyd1 promoters in developing heart and brain tissues. Sci. Rep. 12, 1–11 (2022).
Zhu, W. et al. SULF1 regulates malignant progression of colorectal cancer by modulating ARSH via FAK/PI3K/AKT/mTOR signaling. Cancer Cell Int. 24, 1–19 (2024).
Matsumoto, K. I., Higuchi, T., Umeki, M., Ono, M. & Sakamoto, S. Tenascin-X is increased with decreased expression of miR-378a-5p and miR-486-5p in mice fed a methionine-choline-deficient diet that induces hepatic fibrosis. Biomedical Res. (Japan). 45, 67–76 (2024).
Wu, S. Z. et al. A single-cell and spatially resolved atlas of human breast cancers. Nat. Genet. 53, 1334–1347 (2021).
Wu, S. Z. et al. Stromal cell diversity associated with immune evasion in human triple-negative breast cancer. EMBO J. 39, 1–20 (2020).
Kieffer, Y. et al. Single-cell analysis reveals fibroblast clusters linked to immunotherapy resistance in cancer. Cancer Discov. 10, 1330–1351 (2020).
Tang, Z., Kang, B., Li, C., Chen, T. & Zhang, Z. GEPIA2: an enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Res. 47, W556–W560 (2019).
Liu, Z. et al. BEST: a web application for comprehensive biomarker exploration on large-scale data in solid tumors. J. Big Data 10 Suppl 1(Suppl 1), 12 (2023).
Haubeiss, S. et al. Dasatinib reverses Cancer-associated fibroblasts (CAFs) from primary lung carcinomas to a phenotype comparable to that of normal fibroblasts. Mol. Cancer. 9, 1–8 (2010).
Geering, K. et al. FXYD proteins: new tissue- and isoform-specific regulators of Na,K-ATPase. Ann. N. Y. Acad. Sci. 986, 388–394 (2003).
Gao, Q. et al. FXYD6: A novel therapeutic target toward hepatocellular carcinoma. Protein Cell. 5, 532–543 (2014).
Zhu, Z. L. et al. Overexpression of FXYD-3 is involved in the tumorigenesis and development of esophageal squamous cell carcinoma. Dis. Markers. 35, 195–202 (2013).
Liu, J., Zhou, N. & Zhang, X. A monoclonal antibody against human FXYD6. Hybridoma 30, 487–490 (2011).
Kayed, H. et al. FXYD3 is overexpressed in pancreatic ductal adenocarcinoma and influences pancreatic cancer cell growth. Int. J. Cancer. 118, 43–54 (2006).
Bai, Y. et al. A FXYD5/TGF-β/SMAD positive feedback loop drives epithelial-to-mesenchymal transition and promotes tumor growth and metastasis in ovarian cancer. Int. J. Oncol. 56, 301–314 (2020).
Loftås, P. et al. Expression of FXYD-3 is an independent prognostic factor in rectal cancer patients with preoperative radiotherapy. Int. J. Radiat. Oncol. Biol. Phys. 75, 137–142 (2009).
Liu, J. et al. Extracellular vesicles-encapsulated let-7i shed from bone mesenchymal stem cells suppress lung cancer via KDM3A/DCLK1/FXYD3 axis. J. Cell. Mol. Med. 25, 1911–1926 (2021).
Wang, L. J. et al. Prognostic significance of sodium-potassium ATPaseregulator, FXYD3, in human hepatocellular carcinoma. Oncol. Lett. 15, 3024–3030 (2018).
Floyd, R. V., Wray, S., Martín-Vasallo, P. & Mobasheri, A. Differential cellular expression of FXYD1 (phospholemman) and FXYD2 (gamma subunit of Na, K-ATPase) in normal human tissues: A study using high density human tissue microarrays. Annals Anat. 192, 7–16 (2010).
Zhao, E. et al. The roles of FXYD family members in ovarian cancer: an integrated analysis by mining TCGA and GEO databases and functional validations. J. Cancer Res. Clin. Oncol. 149, 17269–17284 (2023).
Hansen, T. S. et al. The FXYD1 protein plays a protective role against pulmonary hypertension and arterial remodeling via redox and inflammatory mechanisms. Am. J. Physiol. - Heart Circ. Physiol. 326, H623–H635 (2024).
Ai, X. et al. SULF1 and SULF2 regulate Heparan sulfate-mediated GDNF signaling for esophageal innervation. Development 134, 3327–3338 (2007).
Morimoto-Tomita, M. et al. Sulf-2, a proangiogenic Heparan sulfate endosulfatase, is upregulated in breast cancer. Neoplasia 7, 1001–1010 (2005).
Fang, X. et al. Cancer associated fibroblasts-derived SULF1 promotes gastric cancer metastasis and CDDP resistance through the TGFBR3-mediated TGF-β signaling pathway. Cell. Death Discovery. 10, 1–12 (2024).
Liu, C. T. et al. SULF1 inhibits proliferation and invasion of esophageal squamous cell carcinoma cells by decreasing heparin-binding growth factor signaling. Dig. Dis. Sci. 58, 1256–1263 (2013).
Hur, K. et al. Up-regulated expression of sulfatases (SULF1 and SULF2) as prognostic and metastasis predictive markers in human gastric cancer. J. Pathol. 228, 88–98 (2012).
Lai, J. P. et al. SULF1 inhibits tumor growth and potentiates the effects of histone deacetylase inhibitors in hepatocellular carcinoma. Gastroenterology 130, 2130–2144 (2006).
Ouyang, Q. et al. Loss of ZNF587B and SULF1 contributed to cisplatin resistance in ovarian cancer cell lines based on Genome-scale CRISPR/Cas9 screening. Am. J. Cancer Res. 9, 988–998 (2019).
Brasil da Costa, F. H., Lewis, M. S., Truong, A., Carson, D. D. & Farach-Carson, M. C. SULF1 suppresses Wnt3A-driven growth of bone metastatic prostate cancer in perlecan-modified 3D cancer-stroma-macrophage triculture models. PLoS ONE. 15, 1–25 (2020).
Gill, R. M., Mehra, V., Milford, E. & Dhoot, G. K. Short SULF1/SULF2 splice variants predominate in mammary tumours with a potential to facilitate receptor tyrosine kinase-mediated cell signalling. Histochem. Cell Biol. 146, 431–444 (2016).
Narita, K. et al. HSulf-1 inhibits angiogenesis and tumorigenesis in vivo. Cancer Res. 66, 6025–6032 (2006).
Chen, F., Zhang, Z., Yu, Y., Liu, Q. & Pu, F. HSulf-1 and Palbociclib exert synergistic antitumor effects on RB-positive triple-negative breast cancer. Int. J. Oncol. 57, 223–236 (2020).
Tucker, R. P. et al. Phylogenetic analysis of the Tenascin gene family: evidence of origin early in the chordate lineage. BMC Evol. Biol. 6, 1–17 (2006).
Okuda-Ashitaka, E. & Matsumoto, K. I. Tenascin-X as a causal gene for classical-like Ehlers-Danlos syndrome. Front. Genet. 14, 1–7 (2023).
Valcourt, U., Alcaraz, L. B., Exposito, J. Y., Lethias, C. & Bartholin, L. Tenascin-X: beyond the architectural function. Cell. Adhes. Migration. 9, 154–165 (2015).
Matsumoto, K. I. & Aoki, H. The roles of tenascins in Cardiovascular, Inflammatory, and heritable connective tissue diseases. Front. Immunol. 11, 1–10 (2020).
Minamitani, T., Ariga, H. & Matsumoto, K. I. Adhesive defect in extracellular matrix tenascin-X-null fibroblasts: A possible mechanism of tumor invasion. Biol. Pharm. Bull. 25, 1472–1475 (2002).
Liot, S. et al. Loss of Tenascin-X expression during tumor progression: A new pan-cancer marker. Matrix Biology Plus. 6–7, 6–7 (2020).
Archer, M. et al. Immune regulation of mammary fibroblasts and the impact of mammographic density. J. Clin. Med. 11(3), 799 (2022).
Caligiuri, G. & Tuveson, D. A. Activated fibroblasts in cancer: perspectives and challenges. Cancer Cell. 41, 434–449 (2023).
Kuhn, M. Building predictive models in R using the caret package. J. Stat. Softw. 28, 1–26 (2008).
López-Delgado, J. & Meirmans, P. G. History or demography? Determining the drivers of genetic variation in North American plants. Mol. Ecol. 31, 1951–1962 (2022).
Cerami et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 32, 736–740 (2017).
Gao, J. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 6, pl1 (2013).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587e29 (2021).
Guo, W. et al. ScCancer: A package for automated processing of single-cell RNA-seq data in cancer. Brief. Bioinform. 22, 10–11 (2021).
McGinnis, C. S., Murrow, L. M., Gartner, Z. J. & DoubletFinder Doublet detection in Single-Cell RNA sequencing data using artificial nearest neighbors. Cell. Syst. 8, 329–337e4 (2019).
Korsunsky, I. et al. Fast, sensitive, and accurate integration of single cell data with harmony. Nat. Methods. 16, 1289–1296 (2019).
Wu, S. Z. et al. A single-cell and spatially resolved atlas of human breast cancers. Nat. Genet. 53, 1334–1347 (2022).
Pal, B. et al. A single-cell RNA expression atlas of normal, preneoplastic and tumorigenic States in the human breast. EMBO J. 40, 1–23 (2021).
Fustero-Torre, C. et al. Beyondcell: targeting cancer therapeutic heterogeneity in single-cell RNA-seq data. Genome Med. 13, 1–15 (2021).
Acknowledgements
We express our gratitude to GEO, and the TCGA database, along with all contributors who have shared their codes online.
Funding
This work received support from the Zibo City Medical and Health Science Research Projects (No. 2023030926), and Zibo Maternal and Child Health Hospital.
Author information
Authors and Affiliations
Contributions
XZ: Writing – review & editing, Writing – original draft, Validation, Supervision, Project administration, Investigation, Data curation, Conceptualization. Na Wang: Writing – original draft, Resources, Investigation, Formal analysis. Ling Shi: Validation. Xindong Wei: Writing – original draft, Methodology, Investigation, Data curation. Xiaoqin Sun: Resources. Mingxiu Shao: Investigation. Xiaolong Guo: Investigation. Liang Tian: Data curation. Fangyuan Zhang: Data curation. Hui Lyu: Writing – review & editing, Writing – original draft, Validation, Supervision, Project administration, Investigation, Data curation, Conceptualization.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
This study was reviewed and approved by the Ethics Review Committee of Zibo Maternal and Child Health Hospital (approval no. 202106073, data: 2022-06-23). Patient informed consent was obtained as part of surgical consent at the time of surgery for scientific research. The patient’s information was kept confidential.
Competing interests
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhou, X., Wang, N., Shi, L. et al. Identification and validation of a refined CAF-Associated diagnostic signature in breast cancer. Sci Rep 16, 4664 (2026). https://doi.org/10.1038/s41598-025-34923-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-34923-2