Abstract
Relaxations Along a Fictitious Field in the rotating frame of rank 2 (RAFF2) was used to characterize fibrosis-related myocardial diseases with MRI. RAFF2 can be sensitized to molecular dynamics processes by changing the durations of the RAFF2 pulses. Here, we studied the effect of altering the RAFF2 pulse durations for characterization of myocardial infarction (MI). 10 mice underwent myocardial infarct surgery, and the hearts were collected 7 days after the operation. 3 mice hearts were served as healthy control. We tested the influence of stretching factor (TL) on relaxations, where TL was between 0.6 and 2.0, and TL = 1 corresponds to the duration of the RAFF2 pulses TL = 2.25 ms with α2 = 45°. Here, α2 = 45° indicates location of the fictitious field in the second rotating frame. The contrast between MI and remote areas were measured as Relative Relaxation Time Difference (RRTD). Significant increase in relaxation time constants between MI and remote areas with relatively high RRTD were found in RAFF2 TL0.6, TL0.8, TL1.8 and TL2.0. With protein analysis, fibronectin was upregulated in MI area and histology showed area of fibrosis. By altering durations RAFF2 pulse, MI areas can be captured from the myocardium, which provides a novel way to evaluate molecular dynamics in MI area. The findings were validated by the results of other MRI methods, protein analyses and histological sections. Enhancement of MRI contrast between MI, remote and healthy myocardium without using any contrast agents can be achieved by changing the duration RAFF2 pulse.
Similar content being viewed by others
Introduction
Occlusion of coronary artery leads to oxygen deprivation, myocyte loss and subsequently to scar formation1. The development of myocardial infarction (MI) after the occlusion of left anterior descending artery (LAD) affects biological processes, such as protein up- and downregulation and furthermore in the protein content (Fig. 1). Previous studies have shown that fibronectin (Fn1) plays a critical role in the development of MI since it is a component of the extracellular matrix (ECM)2. Fn1 is involved in cellular adhesion, growth, angiogenesis, the formation of the fibronectin matrix and collagen fibrils within the damaged myocardium since polymerization of Fn1 is required for collagen deposition2. Thus, it contributes to the remodeling, and to fibrosis of the myocardium after the MI (Fig. 1)2. There are also several proteins which are involved in the development of MI, such as tenascin-C, osteopontin, periostin (postn), and thrombospondins3.
A schematic diagram of the ischemia-reperfusion injury, which is causing fibronectin (Fn1) to upregulate. This is causing the inhibition of liver-kinase B1 AMP-activated protein kinase (LKB1-AMPK) signaling, which is further promoting nuclear factor kappa-light-chain-enhancer of activated B cells (NF-KB) and transforming growth factor beta 1 (TGF-B1) signaling pathways leading to cardiomyocyte apoptosis, fibrosis and inflammation. The figure is modified from2.
Conventionally, in vivo MI is determined with late gadolinium enhancement (LGE) magnetic resonance imaging (MRI), which is based on the differences in wash-out rates of gadolinium from MI and remote tissues4,5. However, LGE has major drawbacks, since the hyperintensity areas are generated only by the expansion of the extracellular volume (ECV). Gadolinium wash-out similarly from areas with scar, fibrosis and mild collagen formation2, making LGE incapable of distinguishing these biological processes behind different cardiovascular diseases2. Other drawbacks are related to invasiveness, time consumption, and toxicity for using LGE6.
Conventional T1 and T2 relaxation time mapping methods can be used to distinguish the MI area from the remote and healthy myocardium4,7. T1 relaxation time constant is sensitive to MI and fibrotic areas, which is seen as enhanced T1 relaxation time constant in that area, whereas T2 relaxation time constant is sensitive to edema caused by ischemic and inflammation reactions in the MI area, which is seen as enhanced T2 relaxation time constant4,5. Despite the success of T1 and T2 mapping methods for determining MI areas, these methods have some limitations, such as limited robustness for cardiac fibrosis determination by T1 and lack of specificity in the relaxation dispersion of the T2 measurements8. Diffusion tensor imaging is also one promising MRI method to delineate MI area from remote area9.
One MRI contrast, which has been tested in humans is a longitudinal rotating frame relaxation, is characterized by T1ρ relaxation time constant. T1ρ occurs when the magnetization is aligned along the applied radiofrequency (RF) pulse effective magnetic field. T1ρ measurements can be performed with either a continuous wave (CW) spin-lock (SL) technique with the SL pulses at the constant amplitude10,11, or with a train of adiabatic pulses placed prior to the readout portion of the sequence12,13,14,15. CW T1ρ is increased in MI compared to remote areas, which have been demonstrated earlier in mice16,17,18, in swine10 and in humans19. Additionally, a good correlation between CW T1ρ relaxation time constant maps and LGE based determination of MI area was found in mouse16 and swine in vivo models10. Adiabatic rotating frame measurements can be modulated with the choice of frequency- and amplitude-modulation functions of adiabatic pulses, which allows them to generate tissue contrast non-invasively and tamper the contrast in the myocardium after MI16. Adiabatic T1ρ relaxation measurements have been used to image MI area in mice in vivo and ex vivo hearts where adiabatic T1ρ relaxation time constants were elevated in myocardial injury regions compared to remote areas20,21.
In addition to T1ρ, magnetization can be aligned perpendicular to the effective field. Then, magnetization decays with transversal rotating frame relaxation time constant T2ρ7,22. Adiabatic T2ρ has been used successfully to determine MI area from the rest of the myocardium in acute and subchronic phases of MI in both ex vivo and in vivo mice21 indicating greater potential of adiabatic T1ρ and T2ρ measurements to assess the specific pathological processes, such as fibrosis, during the development of the MI area as compared to conventional free-precession T2 method.
At high magnetic field, adiabatic T1ρ and T2ρ relaxation mapping methods are limited by specific absorption rate (SAR)16. One alternative method is entitled Relaxation Along a Fictitious Field in the rotating frame of rank n (RAFFn). Magnetization is swept under sub-adiabatic conditions during RAFFn which significantly reduces power deposition without compromising final effective field amplitude. The delivered specific absorption rate is significantly lower compared to T1ρ and T2ρ16,23,24. Image contrast enhancement based on periodic RF irradiation has been under research interest for a long time25,26,27,28,29,30,31. By varying the period of refocusing of the rotating frame rotary echoes generated during RAFFn, the sensitivity of RAFFn to exchange-induced relaxations can be tuned with periodic RF irradiation to the differences in chemical shifts between the exchanging sites21. Notably, RAFFn (n is rank of the rotating frame) was designed to operate in different rotating frames (e.g., n = 1–5), and the RAFF2 (rank = 2) was used in cardiac MRI applications16,17,24,25,32. For example, it was demonstrated in several prior in vivo studies that RAFF2 can successfully characterize fibrosis-related myocardial diseases16,17,25, with a good correlation to LGE16, it owes potential to be further tuned to increase contrast between MI and remote myocardium by altering the refocusing period of the RAFF2 pulses.
In this work, we measured and localized MI area in mouse ex vivo hearts using RAFFn to study the effect of the refocusing period on RAFF2 relaxation time constants. In addition, single protein up- or downregulation effects on RAFFn relaxations were studied.
Results
Increased relaxation times in remote areas compared to healthy hearts
Relaxation time constants RAFF1 (p < 0.05), RAFF2 TL1.0 (p < 0.01), RAFF4 (p < 0.05) and RAFF5 (p < 0.05) were significantly higher in remote areas compared to healthy hearts (Fig. 2). Other methods were unable to detect the difference (Fig. 2). Significant differences between MI and healthy control hearts were found in RAFF1-5 (p < 0.05) relaxation time constants (Fig. 2).
Averaged rotating frame relaxation times of 10 infarcted hearts and 3 healthy control hearts used in ex vivo measurements. Statistical significances were tested with Two-way ANOVA with Bonferroni post hoc correction test (*P < 0.05, **P < 0.005 ***P < 0.001).
Increased relaxation times in MI compared to remote areas
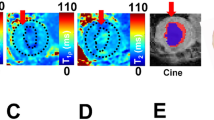
In all calculated relaxation maps, higher relaxation time constants are associated with MI as compared to the remote myocardium (Figs. 2 and 3). Calculated steady state (SS)-maps from different RAFFn methods are presented in Fig. 4. The effect of periodicity of irradiation by increasing the TLs of P-packets from 0.6 to 2.0, where TL = 1 corresponds to the duration 2.25 ms can be seen as monotonic decrease in the RAFF2 in MI, remote area and healthy control hearts by being the highest in the TL0.6 and the lowest in TL2.0 (Fig. 2). Furthermore, significantly increased relaxation time constants were found in RAFF2 TL0.6 (p < 0.01), RAFF2 TL0.8 (p < 0.01) and RAFF2 TL1.0 (p < 0.01) maps in the MI area as compared to remote myocardium (Fig. 2). As a comparison to RAFF2 results, significant increase in relaxation time constants were found with RAFF1,4,5 (p < 0.05), T1ρ (p < 0.05) and T2ρ (p < 0.05) in the MI area as compared to remote myocardium (Fig. 2).
Example of rotating frame and T2 relaxation time constant maps in one infarcted heart. Black ROI outline indicates the area of MI, and the remote area was defined in the septum, which was selected in RAFF2 TL1.0 relaxation map and subsequently this MI area was used for the analysis in all other relaxation maps.
Example of SS-maps from the same heart as in Fig. 3. There are SS-maps from RAFF2 TL 0.6 (A), RAFF2 TL0.8 (B), RAFF2 TL1.0 (C), RAFF2 TL1.2 (D), RAFF2 TL1.4 (E), RAFF2 TL1.6 (F), RAFF2 TL1.8 (G), RAFF2 TL2.0 (H), RAFF1 (I), RAFF3 (J), RAFF4 (K) and RAFF5 (L).
RAFF2 TL2.0, RAFF2 TL1.8 and RAFF1 exhibited the highest contrast between MI and remote areas as described by RRTD values
The contrast difference between MI and remote areas was studied visually and with relative relaxation time difference (RRTD)-values (Figs. 3, 4 and 5). The highest RRTD values were seen with RAFF2 TL2.0, RAFF2 TL1.8 and RAFF1 methods (Fig. 5). By shortening the P-packets as they had significant difference between MI and remote areas, the RRTD values were in RAFF2 TL0.6 (18.2 ± 0.8%), in RAFF2 TL0.8 (17.0 ± 0.9%), and in RAFF2 TL1.0 (12.3 ± 0.8%). Significant differences were also found between RAFFn and different RAFF2 TL RRTD values (Fig. 5). However, no significant differences in RRTD values between longer P-packets (TL1.4-TL2.0) were found, but there were contrast differences between the MI and remote areas as demonstrated in Fig. 3.
RRTD values calculated from relaxation times between MI and remote areas. The statistical comparisons were calculated between methods with Students’ T-Test and One-way ANOVA Dunn’s multiple comparison test (* p < 0.05, *** p < 0.001).
For comparison to RAFFn measurements, the RRTD values from all T1ρ methods were lower than RRTD values of RAFF2 TL2.0 and RAFF2 TL1.8 methods and higher than RAFF1 RRTD values (Fig. 5). Additionally, significant differences in RRTD values between T1ρ (hyperbolic secant) HS1- T1ρ HS4 (p < 0.05), and CW T1ρ - T1ρ HS1 (p < 0.05) were found (Fig. 5). For the other comparison to RAFFn and all T1ρ to T2ρ measurements, significant differences in RRTD values were found between RAFF3,4 - T2ρ (p < 0.05), RAFF5 - T2ρ (p < 0.01), all T1ρ - T2ρ methods (p < 0.001) (Fig. 4). Comparing the rotating frame methods to T2 method, there were no significant differences found (Fig. 5). B1 measurements resulted in 620 ± 11 Hz.
Simulated RAFF2 TL 0.6–2.0.6.0 and RAFFn relaxation times were not affected by single protein up or down regulation
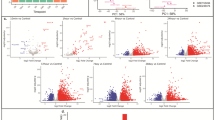
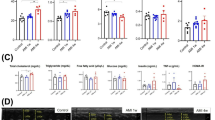
The top 40 proteins were found when the protein analysis of MI and healthy myocardium were done (Fig. 6). There were multiple proteins, which were downregulated in healthy myocardium and upregulated in MI area, like fibronectin (Fn1), Reticulocalbin-3 (Rcn3), Hemopexin (Hpx), Isoform 2 of Periostin (Postn), Inter-alpha-trypsin inhibitor heavy chain H3 (Itih3) and Protein S100-A4 (S100a4) proteins (Fig. 6). On the other hand, for example, Myozenin-2 (Myoz2), Troponin T cardiac muscle (Tnnt2), Troponin I cardiac muscle (Tnni3), Myosin light chain 1/3 skeletal muscle isoforms (Myl; Myl3) and Actin, gamma-enteric smooth (Actc1/Actg2) proteins were upregulated in healthy myocardium but downregulated in MI tissue (Fig. 6). The most important finding was that Fn1 protein (approximately 8.9-fold) and Postn protein (approximately 9.0-fold) were significantly upregulated in the MI area compared to healthy myocardium supporting knowledge of the development of the MI (Fig. 6).
(A) Top 40 proteins which are statistically significant (Adjusted p value < 0.05, logFC > 2) and differentially expressed in MI mouse model (MI15, MI16, MI18) compared to healthy wildtype littermates (H19, H20, H21). Red colour is showing the upregulation and the blue the downregulation. (B) Fn1 and in (C) Postn intensity in the MI and healthy myocardia. (D) RAFF2 relaxation time map 7 days after the MI. (E) Sirius Red histology section from one heart, where red area indicates the area of fibrosis and MI. Red arrows are pointing the area of MI and blue arrows are pointing the remote myocardium.
Results of simulations showed that the concentrations of Fn1 and Postn proteins are too low to affect the relaxation time constants and SS-values of different RAFFn and RAFF2 methods after the MI (Table 2). The SS-values in MI area were higher for different RAFF2 TL’s and similar with the original RAFF1-4 in simulations as compared to SS-values from the measurements (Table 2; Figs. 2 and 4). Additionally, the results of simulations are in line with measured relaxation time constants since RAFF2 TL0.6 and RAFF2 TL0.8 have smaller R-values as compared to bigger R-values in RAFF2 TL1.0–TL2.0 methods (Table 2; Figs. 2 and 4).
Histology revealed the MI area with elevated fibrotic collagen content
Picro Sirius Red-stained histology revealed the MI area with elevated collagen content and fibrosis in the myocardium as compared to the rest of the myocardium (Fig. 6). Small, delicate stained areas were also seen in the remote areas and the original RAFF2 relaxation time map showed the increase of RAFF2 relaxation time constant in the same remote areas as in the Picro Sirius Red stained section (Fig. 6). MRI imaging slices were acquired in the same location as the histology slices were cut, which supports our MRI findings. In addition, as the protein analysis was done by cutting the hearts in half, protein analysis in the MI area included also some parts of remote areas of the myocardium, which can be seen in the histology (Fig. 6). These observations are supporting also the protein findings.
Discussion
In this study, the determination of MI, remote and healthy areas were done by either compressing or stretching the P-packets in RAFF2 pulse train. As a comparison, other rotating frames and T2 relaxation time constants were determined in the same areas as with the RAFFn method and the change of P-packets in RAFF2 pulse. Additionally, MS proteomics analysis was done for the determination of MI and Picro Sirius Red-stained histology validated the MI area.
Increased relaxation times in remote areas as compared to healthy hearts
We found that relaxation time constants in RAFF1, RAFF2 TL1.0, RAFF4 and RAFF5 maps differed significantly between remote areas and healthy hearts. This is interesting, since it is known that remote areas are affected by stress, edema and inflammation reactions. This is supported by the earlier study, where it was found that CW T1ρ relaxation time constant increased in the remote myocardium as compared to healthy myocardium; however, the difference between those regions were not significant18. Histology revealed tissue changes detected by a delicate Picro Sirius Red staining in the remote areas as compared to healthy myocardium (Fig. 6). In addition, original RAFF2 relaxation time map showed the increase of RAFF2 relaxation time constant in the same remote areas as in the Picro Sirius Red stained section (Fig. 6).
Increased RAFF2 and other rotating frame relaxation time constants in MI compared to remote areas
MI area had increased rotating frame relaxation time constants in RAFF1, RAFF2 TL0.6, RAFF2 TL0.8, RAFF2 TL1.0, RAFF4, RAFF5, CW T1ρ, adiabatic T1ρ, and adiabatic T2ρ relaxation maps as compared to relaxation times in the remote area (Figs. 3 and 4), which agrees with previous studies10,16,20,21 and supports the idea of RAFF2 TL0.6-TL2-0 together with other rotating frame methods being a contrast agent free method to separate the MI area from the rest of the myocardium. Relaxation time constants measured with RAFF2 TL0.6, RAFF2 TL0.8 and RAFF2 TL1.0 relaxation times increased significantly in MI area as compared with remote areas, which indicates that reducing the durations of P-packets in RAFF2 pulse train affect the relaxation time constants in MI area while keeping the contrast difference between MI and remote areas (Figs. 3 and 4). The durations of P-packets affect the time to repeat the P-packets in the preparations to acquire different weighting images, which could have potential to add the sensitivity to reveal the MI area from the rest of the myocardium. Furthermore, elevated relaxation time constants were found in the same MI areas as determined with RAFFn, T1ρ and T2ρ, indicating the benefits of the shortening the P-packets in RAFF2 pulse.
RAFF2 TL2.0, RAFF2 TL1.8 and RAFF1 exhibited the highest contrast between MI and remote areas as described by RRTD values
The highest contrast differences between MI and remote areas described by RRTD values were RAFF2 TL2.0, RAFF2 TL1.8 and RAFF1 methods, while mean RRTD values in RAFF2 TL0.6, in RAFF2 TL0.8, and in RAFF2 TL1.0 were higher than 10% indicating the contrast differences between MI and remote areas. The highest RRTD among the methods was RAFF2 TL2.0. With shortening the P-packets in RAFF2 pulse, the relaxation time constants had a significant difference between MI and remote areas; however, those methods did not have as high RRTD-values as stretching the P-packets but still RAFF2 TL0.6, in RAFF2 TL0.8, and in RAFF2 TL1.0 had high RRTD-values. These findings can be interpreted by the effect of chemical exchange since exchange-induced relaxation rate constants causing regional change of relaxation time constants, especially in MI area, where macromolecular tissues like collagen, fibrosis and scar are present. As compared to the original RAFF2, which had the smallest RRTD-value, by changing the length of P-packets in RAFF2 pulse can increase the RRTD between MI and remote areas.
Interestingly, T1ρ with HS1 had the largest RRTD value between the MI and remote areas when compared to CW T1ρ and T1ρ with HS4 methods. The substantial differences observed in T1ρ with HS1 pulse in comparison with other T1ρ techniques may be related to the specific motional regime selected by the adiabatic T1ρ approach utilizing HS1 pulse modulation, which was also seen in previous study20,21. RRTD values of T1ρ significantly differed from T2ρ, which is attributed to the differences in relaxation channels which govern T1ρ and T2ρ relaxations. We also found significant difference in RRTD values between RAFFn and T2ρ, which supports the findings of the rotating frame relaxation time constant differences between MI and remote areas. This is interesting since it has been shown before that TRAFFn comprises both T1ρand T2ρ relaxation pathways23 but we did not see any significant differences between RAFFn and T1ρ, which might indicate that the increase of relaxation time constant in MI area is due to similarity of the relaxation mechanisms and the biological tissues, such as collagen and fibrosis20.
Comparing the rotating frame methods to T2, there were no statistically significant differences in RRTD values between these methods even though the relaxation pathways are different between these methods. It should be noted that we have done ex vivo measurements, where the relaxation mechanisms are affected due to formalin fixation, since protein and water molecule interactions in ex vivo and in vivo MRI could substantially differ in their spin dynamics, which leads to almost uniform fixation of the ECM and causes heart dehydration leading to alterations in proton diffusion patterns as compared to in vivo situation33. Additionally, in ex vivo MRI, the maximum temperature of the heart is room temperature, which is different than during in vivo measurements and needs to be taken account34.
RAFF2 TL 0.6–2.0.6.0 and RAFFn were not affected by single protein up or down regulation
Proteomics analysis revealed that cell adhesion proteins like Postn and Fn1 are related to outside of myocyte in ECM, fibronectin matrix, fibril formation and cardiac repairing (by Postn protein) and these proteins were highly upregulated in MI area in our results and previous findings2,35. Additionally, other upregulated proteins in MI area like Itih3 and S100a4 are related to other ECM proteins, calcium-binding, angiogenesis, and various cellular processes, indicating that there have been tissue changes in the ECM as compared to healthy myocardium by determining the changes in protein content in MI area. Fn1 protein is interesting because it is related to ECM, fibronectin matrix and fibril formation since polymerization of Fn1 is required for collagen deposition and thus, it plays a critical part in fibrosis in MI area2.
The gold standard in MI visualization with MRI is LGE, which is related to Gd’s wash-in-wash-out-rate from MI area ECM compared to remote area ECM. In addition, relaxation time constants are increasing due to the increase of free water in MI area, which is increasing the ECV and ECM. Thus, protein findings in this study support the idea of rotating frame relaxation time methods to efficiently determine the MI area from the remote areas, which is successfully shown in previous MI studies10,16,19,20,21. Furthermore, Fn1 has been previously analyzed with nuclear magnetic resonance36, indicating that Fn1 has enough MR properties to be able to get detectable MR-signal from Fn1; however, further studies of how Fn1 itself influences the detectable MRI-signal. For the detectability of Fn1, the actual concentration of Fn1 in MI area, how the pH is changing during the MI development and how the free water molecules can access the exchange site of Fn1 need to be figured out to consider Fn1 role in the effect to the relaxation. In addition to protein findings, there have been studies about cardiac ECM target imaging for the fibrotic collagen content with MRI37,38 and with collagen targeted contrast agent39, which indicate that more specific MRI methods can be optimized to sensitively capture the ECM related proteins in the MI area.
Our simulations of the relaxation time constants and SS formation for Fn1 and Postn protein upregulations in MI area showed that the concentrations of these proteins are too low to affect the RAFF2 TL and RAFFn relaxation times or steady states, even though Fn1 can produce the detectable NMR-signal36. Straightforward connection between Fn1 and the increase of rotating frame relaxation times cannot be made based on our simulations; however, the sum of Fn1 and other upregulated ECM proteins have a minor effect on detectable MR-signal. Based on our histology results, the major source of detectable MR-signal is from increased fibrosis and water accumulation, which might overshadow the MR-signal from the upregulated ECM proteins in heart imaging. In addition to that, there were also differences in remote and healthy myocardium detected with the RAFFn method. Here, one possible factor in that result is related to the role of Fn1 and Postn proteins in whole MI differences compared to healthy myocardium.
Limitations
There were some limitations in this 2-D ex vivo study. The sample size was small, particularly healthy hearts, which affected the robustness of the statistical analysis both MRI and MS proteomics. Exact location of 2-D short-axis MRI slice might not always be optimal as the more serious MI area might have been lower in the myocardium. Furthermore, the relaxation times measured with RAFF2 with increased TL were more prone to were B0 artifacts than RAFF2 TL1.0 (Fig. 4). As original RAFF1-5 pulses have been applied for human brain imaging and RAFF2 for human heart imaging, due to similarities of different lengths of RAFF2 pulses to original RAFF1-5 pulses, this method owes potential to be transferred to in vivo preclinical and clinical devices. However, the results of this study may not be transferrable as is, due to differences between in vivo and ex vivo tissue conditions. To the best of our knowledge, this study is the first demonstration of a novel contrast in cardiac MRI which is generated during periodic irradiation.
As a conclusion, RAFF2 rotating frame relaxation contrast between infarct and remote myocardium depends on and can be optimized by altering the duration of refocusing time. RAFFn were able to distinguish MI, remote and healthy myocardia without a need of contrast agent.
Methods
Animals
The hearts of C57BL mice (n = 13) were obtained from the Animal Center of the University of Eastern Finland. The hearts from 10 mice were collected after 7 days of Left Anterior Descending LAD occlusion and 3 hearts were from healthy controls and no sham operations were done to healthy controls. Animal experiments were performed according to the national and international guidelines for laboratory animal use, and the experiments were approved by the National Animal Experiment Board of Finland under license ESAVI-270-04.10.10.07–2017 and carried out following the guidelines of the Finnish Act on Animal Experimentation and Directive 2010/63/EU of the European Parliament. All experiments were following the Animal Research Reporting In Vivo Experiments (ARRIVE) guidelines.
LAD was ligated in a surgical operation40, where mice were the first anesthetized with inhalation anesthesia (4% induction, 2% maintenance with 30/70 oxygen/nitrogen ratio). Mouse chest was opened at the 4th intercostal space of the sternum. After that, the heart was pulled out, pericardial sac was removed and then LAD occlusion was done. The heart was then returned to its initial place and the chest was closed. After the operation, analgesia was given subcutaneously to mice on the day of the operation and the following three days. The analgesia included buprenorphine (0.3 mg/ml), Temgesic (0.05–0.1 mg/kg), Carprofen (50 mg/ml) and Rimadyl (5 mg/kg). After 7 days, mice were sacrificed with CO2 and their hearts perfused with phosphate buffer saline through the left ventricle. Then the hearts were collected and fixed with 4% paraformaldehyde in 7.5% sucrose for 4 h and after that, the hearts were stored in 15% sucrose.
MRI
MRI was performed with a vertical 9.4 T magnet using volume RF transceiver coil with diameter of 15 mm (Rapid Biomedical GmbH, Rimpar, Germany) and Vnmrj3.1 Varian/Agilent DirectDrive console. All imaging was conducted ex vivo. The hearts were placed in 8-mm NMR glass tube, and the tube were filled with perfluoropolyether (Galden HS 240, Solvay Solexis, Italy), since Galden is a fluorinated heat transfer fluid, which is not giving any MR signal.
RAFFn pulse wave forms were generated as before23,26 with α1 = 45° and refocusing durations were calculated with the equation of 4π/(\(\:\sqrt{2{{\upomega\:}}_{1}^{max}}\)), where \(\:{{\upomega\:}}_{1}^{max}\:\)is maximum frequency of applied B1 as described in23,26,41. Imaging protocol contained experiments with RAFFn (n = 1–5), and RAFF2 with stretching factors (TL)s of 0.6–2.0.6.0, where TL0.6 is shortened by factor 0.6 from the original TL1.0 RAFF2 P-packet with the duration 2250 µs at the nominal peak power 625 Hz with α2 = 45°, where, α2 = 45° indicates location of the fictitious field in the second rotating frame, and TL2.0 is stretched by factor 2.0 (Fig. 7; Table 1) similarly as in23,26,41. The parameters of the pulses, i.e., the nominal pulse peak powers and the durations of P-packets with refocusing durations for all used RAFF2 pulses are presented in Table 1. The maximum number of pulses in the pulse train for the RAFF2 TL0.6-TL2.0 varied from 0 to 32 and the durations of the pulse trains were between 0 and 72.32 ms, respectively. The same pulse train lengths were applied for RAFFn as RAFF2 TL1.0 and the peak nominal pulse power ranged between 625 and 245 Hz, respectively23. Relaxation time constant maps for RAFFn measurements were calculated pixel-by-pixel manner with monoexponential decay function with SS-formation using Aedes Software (http://aedes.uef.fi) in MATLAB R2023b (Mathworks Inc., Natick, CA, USA). Signal intensity decay curves (SI(t)) were measured using two consecutive acquisitions, first starting with magnetization oriented along + z, and second when magnetization is initially along -z. Two SI curves were simultaneously fitted using non-linear fitting with a least square cost function using exponential decay and recovery to the SS41:
Effect of the stretching factor on the amplitude of P-packets and to the phases in RAFF2 pulse. Here, the RAFF2 TL0.6 (A), RAFF2 TL1.0 (B), and RAFF2 TL2.0 (C) methods are shown.
Here S0 is initial SI, R is the relaxation rate constant which describes the decay of the signal, SSS is the steady-state SI for \({\text{t}} \to \infty\)SS is a fraction \(SS={S_{SS}}/{\text{S}}{{\text{I}}_0}\)
For comparison, CW T1ρ, adiabatic T1ρ with both HS1 and HS4 pulses with R = 2014, adiabatic T2ρ with both HS1 and HS4 pulses and R = 20, T2 and B1 were measured. For CW T1ρ measurements, the flip of M to xy-plane was done with adiabatic half passage (AHP) pulse, which duration was 4.0 ms and the RF power was 2500 Hz42. After the flip, CW pulses with durations of 0, 18, 36 and 54 ms were used with pulse power of 1250 Hz to produce the CW SL weighting. After CW pulse, AHP reverse pulse was used to return M back to its initial orientation along the z axis. For both adiabatic T1ρ and T2ρ measurements, number of HS1 and HS4 pulses were 0, 8, 16, and 32 with duration of 4.5 ms and RF peak power of 2500 Hz in the pulse train15,20,21. For T2ρ measurements, similar AHP and reverse pulses as in CW T1ρ measurement was used before and after the adiabatic full passage (AFP) pulse train15,21. For T2 measurements, number 0, 4, 8 and 16 pulses with duration of 4.0 ms, delays of 0.55 ms and power of 2500 Hz were used to make the echo times of 0, 18.2, 36.4 and 72.8 ms. Relaxation time constant maps were calculated pixel-by-pixel manner with monoexponential decay function. B1 measurements for B1 mapping included a hard pulse with power of 625 Hz with increasing durations from 0 to 1.0 ms, Δ = 0.125 ms43.
2D fast spin echo (FSE) sequence was used as a readout in all measurements after the preparation pulse. Parameters of FSE were TR = 3000 ms, effective TE = 8.44 ms, averages = 5 with data matrix = 192 × 192 and 1 mm slice thickness. The axial imaging plane was placed in the lower mid level of the heart and imaged as a short-axis view (Table 2).
MRI data analysis
MI, remote and healthy areas were assessed using regions of interest (ROIs) analysis. MI and remote ROIs were the first drawn in the original RAFF2 (TL1.0) relaxation map39 and used that ROI in all other maps in all MI hearts. For the healthy myocardium, the whole myocardium was selected as one ROI and it was used in all healthy control hearts. The ROIs were then used to collect the mean values in specific regions of the data in relaxation maps. The contrast between remote and sub-chronic MI areas was measured as RRTD. The RRTD was calculated as (T(infarct)-T(remote))/T(remote)·100%, where T represents the relaxation time constant in that specific ROI. Results are given in the form of the mean ± standard deviation. Student’s t-test, One-way ANOVA Dunn’s multiple comparison test and Two-way ANOVA with Bonferroni’s post hoc tests were performed to analyze statistically the results, and p < 0.05 was considered statistically significant.
Proteomics
After the MRI, the hearts were frozen and cut in half to separate ventricles and atriums. Infarcted and healthy ventricular and atrial samples (n = 3 per group) were homogenized in Abcam protein lysis buffer (ab193970) using the efficient bead-beating protein extraction method44. The solubilized heart proteome was extracted by centrifugation at 18,000 × g for 20 min. The extracted proteins were digested and processed using the filter-aided sample preparation (FASP) protocol as previously described45,46. The resulting tryptic peptides were analyzed in full-scan and data-independent acquisition (DIA) modes using high-resolution mass spectrometry (Orbitrap Q Exactive Classic) coupled with an ultra-high-performance liquid chromatography system (Vanquish Flex UHPLC, Thermo Scientific, Bremen, Germany) as previously described46. The MS/MS spectral library and peptide retention times were predicted using the UniProt reference proteome database for mouse (UP000000589, updated in April 2022, containing 21,957 protein entries) along with an additional UniProt mouse database comprising 41,543 protein isoform entries. Cysteine residues were set as static modifications, while oxidation of methionine and N-terminal acetylation were designated as variable modifications, with a maximum of two variable modifications per peptide. The predicted MS library was used to search the raw data, applying a 1% false discovery rate (FDR) threshold for both precursor and protein groups and requiring at least one proteotypic peptide per protein (7–30 amino acids in length)47. The normalized MaxLFQ quantities were subsequently subjected to downstream comparative statistical analysis using Perseus software48,49. Categorical groups were analyzed using the LIMMA test48, with multiple testing corrections applied via the false discovery rate (FDR), and statistical significance set at q < 0.05.
Simulations
Two-pool Bloch-McConnell equations were used to model the effect of Fn1 and Postn proteins on RAFF1-5 and RAFF2 TL0.6-TL2.026,29. The identical pools of the T1 and T2 were calculated by using the model of dipolar interactions between isolated spins using rotational correlation time τc = 10·10− 12 s, similarly to26. Other parameters for the simulations were 9.4 T external magnetic field, gyromagnetic ratio was 42.576 MHz/T for 1H in 9.4 T, Planck’s constant was 1.054571628·10− 34 Js, hydrodynamic radius was 158·10− 12 m50, permeability of vacuum was 4π·10− 7, τex was 0.001 Hz for -OH-group, off-resonance frequency for both pools were set to be ΔA = 2·π ·PB·500 rad/s and be ΔB = 2·π ·PA·500 rad/s, respectively, where PA=(c(Fn1)/c(water)) or PA=(c(Postn)/c(water)) and PB=1-PA. PA included healthy concentration values of 1.2·10− 6 mol/l for Fn1 (FN1 protein expression summary - The Human Protein Atlas), of 1.1·10− 9 mol/l for Postn51 and of 80 mol/l for the water concentration. For the upregulated concentration values, we used 1.2·10− 5 mol/l for Fn1 and 1.1 ·10− 8 for Postn. The α1 was 45° and ω1max = 2·π·625 rad/s, which was decreasing as the n in the RAFFn was increasing. The number of points in PPπ−1PπP− 123 waveform was 128 for original RAFFn and were changed as the TL factor changed. This affects the refocusing pulse duration which is calculated with as follows: Tp=(4·π)/(\(\:\sqrt{2}\) ·ω1max). A total of 64 pulses were used in the pulse train to form up to 144 ms long RF irradiation, where time points 0 and 32 points were evenly distributed. Also, magnetizations from both + Z and -Z were used to calculate relaxation rates in the simulations and then the magnetizations were summed up, and SS formation23,41 of exponential decay function was taken account to fit the data points by using non-linear least square fitting. Waveforms and refocusing schemes were formed with equations previously derived23,52. Simulations were performed in Matlab R2023b by numerically solving Bloch-McConnell equations by ordinary differential equation solver (ode45) time point-by time point fashion.
Histology
For the two hearts, standard tissue processing and paraffin embedding methods were used. 4 μm thick cross-sections of the hearts were cut and stained with Picro Sirius Red staining. The slides were mounted with Permount (ThermoFisher Scientific), and the sections were photographed using a Nikon Eclipse microscope with a Ds-Ri2 camera (Nikon Instruments Europe BV).
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Falk, E., Shah, P. K. & Fuster, V. Coronary plaque disruption. Circulation 92 (3), 657–671. https://doi.org/10.1161/01.CIR.92.3.657 (1995).
Blockage of Fibronectin 1 Ameliorates Myocardial Ischemia/Reperfusion Injury in Association with Activation of AMP-LKB1-AMPK Signaling Pathway Yun-Long Zhang, Li, P. B., Han, X. & Zhang, B. Hui-Hua Li 12 May 2022 https://doi.org/10.1155/2022/6196173
Frangogiannis, N. G. The extracellular matrix in myocardial injury, repair, and remodeling. J. Clin. Invest. 127 (5), 1600–1612. https://doi.org/10.1172/JCI87491 (2017).
Raijah, P. S. et al. Cardiac MRI: state of the Art. Radiology 307 (3), e223008 (2023).
[5] Ylä-Herttuala, E. et al. Curr. Cardiovasc. Imaging Rep. 12:11. https://doi.org/10.1007/s12410-019-9487-3 (2019).
Mehdi, R. R. et al. Non-Invasive diagnosis of chronic myocardial infarction via composite In-Silico-Human data learning. Adv. Sci. 12, e06933. https://doi.org/10.1002/advs.202406933 (2025).
Wu et al. Alternating Look-Locker for quantitative T1, T1 and B1 3D MRI mapping. Magn. Reson. Med. 91, 149–161 (2024).
Pavel Filip, A. et al. Rotating frame MRI relaxations as markers of diffuse white matter abnormalities in multiple sclerosis, . NeuroImage: Clinical https://doi.org/10.1016/j.nicl.2020.102234 (2020).
Mendiola, E. A. et al. Progressive microstructural remodeling in the infarcted left ventricle analyzed by diffusion tensor MRI. In Functional Imaging and Modeling of the Heart. FIMH 2025 Vol. 15673 (eds Chabiniok, R. et al. et al.) (Springer, 2025). https://doi.org/10.1007/978-3-031-94562-5_3.
Witschey, W. R. et al. Rotating frame spin lattice relaxation in a swine model of Chronic, left ventricular myocardial infarction. Magn. Reson. Med. 64 (5), 1453–1460. https://doi.org/10.1002/mrm.22543 (2010).
Berisha, S., Han, J., Shahid, M., Han, Y. & Witschey, W. R. T. Measurement of myocardial T1ρ with a motion Corrected, parametric mapping sequence in humans. PLoS ONE. 11 (3), e0151144. https://doi.org/10.1371/journal.pone.0151144 (2016).
Kettunen, M. I., Gröhn, O. H., Penttonen, M. & Kauppinen, R. A. Cerebral T1ρ relaxation time increases immediately upon global ischemia in the rat independently of blood glucose and anoxic depolarization. Magn. Reson. Med. 46, 565–572. https://doi.org/10.1002/mrm.1228 (2001).
Michaeli, S., Sorce, D. J., Springer, C. S. Jr, Ugurbil, K. & Garwood, M. T1rho MRI contrast in the human brain: modulation of the longitudinal rotating frame relaxation shutter-speed during an adiabatic RF pulse. J. Magn. Reson. 181 (1), 135–147. https://doi.org/10.1016/j.jmr.2006.04.002 (2006).
Garwood, M. & DelaBarre, L. The return of the frequency sweep: designing adiabatic pulses for contemporary NMR. J. Magn. Reason. 153 (2), 155–177 (2001).
Michaeli, S. et al. Exchange-influenced T2rho contrast in human brain images measured with adiabatic radio frequency pulses. Magn. Reson. Med. 53 (4), 823–829. https://doi.org/10.1002/mrm.20428 (2005).
Yla-Herttuala, E. et al. Quantification of myocardial infarct area based on TRAFFn relaxation time maps - comparison with cardiovascular magnetic resonance late gadolinium enhancement, T1ρ and T2 in vivo. J. Cardiovasc. Magn. Reson. 20, 34. https://doi.org/10.1186/s12968-018-0463-x (2018).
Ylä-Herttuala, E. et al. Lymphatic insufficiency leads to distinct myocardial infarct content assessed by magnetic resonance TRAFFn, T1ρ and T2 relaxation times. Sci. Rep. 13, 1579. https://doi.org/10.1038/s41598-023-28219-6 (2023).
Musthafa, H. S. et al. Longitudinal rotating frame relaxation time measurements in infarcted mouse myocardium in vivo. Magn. Reson. Med. 69 (5), 1389–1395. https://doi.org/10.1002/mrm.24382 (2013).
van Oorschot, J. W. et al. Endogenous assessment of diffuse myocardial fibrosis in patients with T1r- mapping. J. Magn. Reson. Imaging. 45 (1), 132–138 (2017).
Iida Räty, A., Aarnio, M. J., Nissi, S. & Kettunen, A. K. Ex vivo imaging of subacute myocardial infarction with ultra-short echo time 3D quantitative T1- and T1ρ-mapping magnetic resonance imaging in mice. Eur. Heart J. - Imaging Methods Pract. 3(Issue 1), qyae131. https://doi.org/10.1093/ehjimp/qyae131 (2025).
Ylä-Herttuala, E. et al. Capturing acute and subchronic myocardial infarct by MRI rotating frame relaxation times in mice in and ex vivo. NMR Biomed. 38, e70061. https://doi.org/10.1002/nbm.70061 (2025).
Michaeli, S., Source, D. J., Idiyatullin, D., Ugurbil, K. & Garwood, M. Transverse relaxation in the rotating frame induced by chemical exchange. J. Magn. Reason. 169 (2), 293–299 (2004).
Liimatainen, T. et al. MRI contrasts in high rank rotating frames. Magn. Reson. Med. 73, 254–262 (2015).
Mirmojarabian, S. A. et al. Relaxation along fictitious field (RAFF) provide an appropriate alternative method for imaging chronic myocardial infarct without exogenous contrast media. Interv Cardiol. 13 (5), 381–388 (2021).
Khan, M. A. et al. The follow-up of progressive hypertrophic cardiomyopathy using magnetic resonance rotating frame relaxation times. NMR Biomed. 31, e3871. https://doi.org/10.1002/nbm.3871 (2018).
Timo Liimatainen, H., Laakso, D., Idiyatullin, S. & Mangia, S. Capturing exchange using periodic radiofrequency irradiation. J. Magn. Reson. https://doi.org/10.1016/j.jmr.2018.09.001 (2018).
Idiyatullin, D., Corum, C. A. & Garwood, M. Multi-band-SWIFT. J. Magn. Reson. 251, 19–25 (2015).
Brunner, D. O. et al. Sideband excitation for concurrent rf transmission and reception. Proc. Int. Soc. Magn. Reson. Med. 19, 625 (2011).
Portnoy, S. & Stanisz, G. J. Modeling pulsed magnetization transfer. Magn. Reson. Med. 58, 144–155 (2007).
Zhang, S. & Gorenstein, D. G. Adiabatic decoupling sidebands. J. Magn. Reson. 144, 316–321 (2000).
Närväinen, J., Hubbard, P. L., Kauppinen, R. A. & Morris, G. A. Z-spectroscopy with alternating-phase irradiation. J. Magn. Reson. 207, 242–250 (2010).
Tourais, J. et al. Feasibility of relaxation along a fictitious field in the 2nd rotating frame (TRAFF2) mapping in the human myocardium at 3 T.. Front. Cardiovasc. Med. 11, 1373240 (2024).
Thavarajah, R., Mudimbaimannar, V. K., Elizabeth, J., Rao, U. K. & Ranganathan, K. Chemical and physical basics of routine formaldehyde fixation. J. Oral Maxillofac. Pathol. 16, 400–405. https://doi.org/10.4103/0973-029X.102496 (2012).
Holz, M., Heil, S. R. & Sacco, A. Temperature-dependent self-diffusion coefficients of water and six selected molecular liquids for calibration in accurate 1H NMR PFG measurements. Phys. Chem. Chem. Phys. 2, 4740–4742. https://doi.org/10.1039/B005319H (2000).
J Clin Invest ;127(5):1600–1612. (2017). https://doi.org/10.1172/JCI87491
Copié, V. et al. Solution structure and dynamics of linked cell attachment modules of mouse fibronectin containing the RGD and synergy regions: comparison with the human fibronectin crystal structure. J. Mol. Biol. 277(3), 663–682 (1998).
Pinkert, M. A. et al. Adv. Exp. Med. Biol. :1098:21–44. doi: https://doi.org/10.1007/978-3-319-97421-7_2. (2018).
Won, S. et al. Curr. Opin. Cardiol. ;28(3):282–289. doi: https://doi.org/10.1097/HCO.0b013e32835f5a2b. (2013).
Helm, P. A. et al. Postinfarction myocardial scarring in mice: molecular MR imaging with use of a collagen-targeting contrast agent. Radiology 247(3), 788–796 (2008).
Gao, E. et al. A novel and efficient model of coronary artery ligation and myocardial infarction in the mouse. Circ. Res. [Internet]. 107 (12), 1445–1453 (2010).
Liimatainen, T., Sorce, D. J., O’Connell, R., Garwood, M. & Michaeli, S. MRI contrast from relaxation along a fictitious field (RAFF). Magn. Reson. Med. 64, 983–994. https://doi.org/10.1002/mrm.22372 (2010).
Gröhn, O. H. J. et al. Early detection of irreversible cerebral ischemia in the rat using dispersion of the magnetic resonance imaging relaxation time, T1rho. J. Cereb. Blood Flow. Metab. 20 (10), 1457–1466. https://doi.org/10.1097/00004647-200010000-00007 (2000).
Vaughan, J. T. et al. Detunable transverse electromagneetic (TEM) volume coil for high-field NMR. Magn. Reason Med. 47, 990–1000 (2002).
Neumann, D. et al. ; (2014). https://us.omni-inc.com/amfile/file/download/file/4/
Wiśniewski, J. R. & Anal Chim. Acta ;1090:23–30. doi:https://doi.org/10.1016/j.aca.2019.08.032. (2019).
Retinal Proteome Profiling of Inherited Retinal Degeneration Across Three Different Mouse. Models Suggests Common Drug Targets in Retinitis Pigmentosa Montaser, Ahmed B. et al. Molecular & Cellular Proteomics, Volume 23, Issue 11, 100855
Demichev, V. et al. Nat. Methods 17, 41–44 https://doi.org/10.1038/s41592-019-0638-x. (2020).
Cox, J. et al. Mol. Cell. Proteomics, 13(9), 2513–2526. (2014).
Tyanova, S. et al. Nat. Methods 13, 731–740 https://doi.org/10.1038/nmeth.3901 (2016).
Mangia, S., Liimatainen, T., Garwood, M. & Michaeli, S. Rotating frame relaxation during adiabatic pulses vs. conventional spin lock: simulations and experimental results at 4 T. Magn. Reson. Imaging. 27, 1074–1087 (2009).
Xia, Y. et al. The comprehensive study on the role of POSTN in fetal congenital heart disease and clinical applications. J. Transl Med. 21, 901. https://doi.org/10.1186/s12967-023-04529-1 (2023).
Liimatainen, T. et al. Relaxation dispersion in MRI induced by fictitious magnetic fields. J. Magn. Reson. 209, 269–276 (2011).
Acknowledgements
Authors also want to thank Kuopio biomedical imaging unit (BIU) for using their infrastructure and GeneCellNano flagship for supporting this project.
Funding
Authors thank, Finnish Cultural Foundation, Finnish Foundation for Cardiovascular Research, Valtion tutkimusrahoitus (VTR) also known as state funding for research, Mauri and Sirkka Wiljasalo Foundation, Paavo Nurmi Foundation, Emil Aaltonen Foundation, Matti and Vappu Maukonen Foundation, Maud Kuistila Memorial Foundation, the Orion Research Foundation, the Urho Känkänen Foundation, the Vilho, Yrjö, and Kalle Väisälä Foundation, Research Council of Finland #340761, Flagship of Advanced Mathematics for Sensing Imaging and Modelling grant #359186 and NIH grant P41 EB027061 for supporting this research project.
Author information
Authors and Affiliations
Contributions
Each author contributed significantly to the submitted work: EYH was involved in the design of the study, performed the MRI measurements, performed data analysis, did the simulations, interpreted the data and drafted the manuscript. IR did some MRI measurements, did the histology, performed the histological data analyses and revised the manuscript. AM did the proteomic measurements, proteomic analysis and drafted the manuscript. SL ordered the animals, performed the LAD operations, did the ex vivo heart operations, handled the hearts after the euthanasia and revised the manuscript. SM and TL participated in the design of the study, data interpretation and modelling, writing and editing the manuscript and drawing the conclusions. All authors read and approved of the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ylä-Herttuala, E., Räty, I., Montaser, A. et al. Enhancing MRI contrast between sub-chronic myocardial infarct, remote and healthy myocardium using periodic irradiation in the rotating frames. Sci Rep 16, 3888 (2026). https://doi.org/10.1038/s41598-025-34949-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-34949-6