Abstract
The aim of this study was to develop and validate a machine learning-based mortality risk prediction model for patients with severe community-acquired pneumonia (SCAP) in the intensive care unit (ICU). We collected data from two centers as the development and external validation cohorts. Variables were screened using the Recursive Feature Elimination method. Five machine learning algorithms were used to build predictive models. Models were evaluated through nested cross-validation to select the best one. The model was interpreted using Shapley Additive Explanations. We selected the optimal model to generate the web calculator. A total of 23 predictive features were selected. The Light Gradient Boosting Machine (LightGBM) model had an area under the receiver operating characteristic curve (AUC) of 0.842 (95% CI: 0.757–0.927), with an external 5-fold cross-validation average AUC of 0.842 ± 0.038, which was superior to the other models. External validation results also demonstrated good performance by the LightGBM model with an AUC of 0.856 (95% CI: 0.792–0.921). Based on this, we generated a web calculator by combining five high importance predictive factors. The LightGBM model was confirmed to be efficient and stable in predicting the mortality risk of patients with SCAP admitted to the ICU. The web calculator based on the LightGBM model can provide clinicians with a prognostic evaluation tool.
Similar content being viewed by others
Introduction
Community-acquired pneumonia (CAP) in adults, as distinct from hospital-acquired pneumonia (HAP), is an acute pulmonary infection that occurs outside the hospital or within the average incubation period after hospital admission1. Patients with CAP are admitted to the intensive care unit (ICU) primarily because they require a higher level of life support or close monitoring of vital signs. This subset of CAP cases is typically classified as severe community-acquired pneumonia (SCAP). Jain et al. (2015) demonstrated that approximately 21% of patients with CAP were admitted to the ICU2. To effectively manage SCAP, clinicians must carefully monitor the patient’s condition and promptly adjust treatment due to its rapidly variable nature3. Despite advancements in SCAP management, the condition continues to have a high mortality rate, posing a significant global public health challenge. A secondary analysis of a population-based prospective cohort study revealed a 30-day mortality rate of 27% and a 1-year mortality rate of 47% for ICU patients with SCAP4. Dupuis et al. (2021) reported a 28-day in-hospital mortality rate of 22.8% and a 1-year mortality rate of 32.3% for patients with SCAP in ICUs5. These outcomes contribute to substantial costs not only for the treatment of SCAP but also for associated comorbidities, such as chronic heart failure and chronic respiratory diseases, placing a heavy burden on healthcare systems6. Accurate mortality prediction in patients with SCAP facilitates the strategic allocation of limited ICU resources to those at greatest risk, enabling clinicians to develop personalized treatment plans and improve treatment efficacy. Such measures have the potential to reduce mortality rates. Proactive management strategies are essential to prevent SCAP-related deaths. Early implementation of advanced life support measures, such as mechanical ventilation or hemodynamic stabilization, is critical for mitigating disease progression. Furthermore, personalized treatment plans, including the use of more potent antibiotics or additional therapeutic interventions, are crucial. By identifying patients at heightened risk of mortality early, these interventions can be targeted more effectively, empowering ICU physicians to make decisive clinical decisions that save lives and alleviate the financial burden on patients and healthcare systems7.
Several studies have reported independent risk factors for mortality in patients with SCAP. However, traditional statistical methods have limited predictive power8. Although some studies have developed models to predict mortality risk, these single-center studies lacked external validation, limiting their generalizability9,10. A common observation in previous research is the multitude of variables associated with SCAP mortality, including age, comorbidities, severity scores, laboratory abnormalities, and the timeliness of treatment. Traditional models are unable to fully capture the complex interactions among these variables in predicting mortality. Machine learning, a subset of artificial intelligence, offers significant potential for handling large-scale data and improving predictive accuracy. It enables computers to learn patterns and relationships from data without explicit programming and encompasses supervised, unsupervised, and reinforcement learning techniques, commonly used for classification, regression, and prediction tasks. Machine learning has gained increasing traction in the medical field due to its efficiency and accuracy11,12, particularly in the ICU, where it helps clinicians manage information overload more effectively13,14,15. In recent years, several studies have applied machine learning algorithms to predict the risk of mortality in patients with severe pneumonia. However, some studies employed only a single model for training, resulting in less accurate predictions16. Others compared multiple machine learning models but failed to develop practical clinical application tools17. Additionally, these studies often relied on a single random grouping method for model training, which compromised the stability of their models.
This study aims to train and compare five different machine learning models to predict the risk of mortality in patients with SCAP admitted to the ICU. The strengths of this study include in the comparison of multiple machine learning models, external validation, and the development of a practical web calculator. By using the web calculator, clinicians can assess the risk of mortality for patients with SCAP and implement measures to reduce mortality rates. Furthermore, the interpretive tools of the model can help clinicians intuitively understand key predictive factors, enabling the formulation of more targeted treatment plans.
Methods
Study design
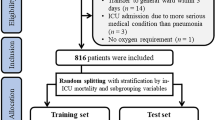
This study utilized datasets from two research centers. The development cohort included patients with SCAP (n = 455) admitted to the Department of Critical Care Medicine at the First Affiliated Hospital of Anhui Medical University between June 2019 and October 2022. Data from patients with SCAP (n = 120) admitted to the Respiratory Intensive Care Unit at Anhui Chest Hospital between July 2021 and January 2023 were used as the external validation cohort. This study was approved by the Ethics Committee of the First Affiliated Hospital of Anhui Medical University (No. PJ2022-01-09) and Anhui Chest Hospital (No. KJ2023-40). It strictly adhered to the principles of the Declaration of Helsinki. As this was a retrospective study, the requirement for informed consent was waived with the approval of the Ethics Committees of both hospitals.
The inclusion criteria for this study were: (1) age ≥ 18 years, (2) admission to the ICU with a primary diagnosis of SCAP, and (3) diagnosis of SCAP in accordance with the Infectious Diseases Society of America/American Thoracic Society (IDSA/ATS) consensus guidelines18. Patients were excluded if: (1) the diagnosis was HAP, or (2) the final outcome was unclear. Patients with SCAP were identified using International Classification of Diseases- 10 codes. Only the initial ICU hospitalization data for each patient was included, eliminating potential bias from duplicate records. ICU hospital mortality in patients with SCAP was used as the study endpoint.
Candidate predictors
This study used 23 clinically accessible predictors as baseline data, selected based on clinical relevance and factors previously demonstrated in the literature to be associated with mortality risk in patients with SCAP. Basic demographic information included sex and age. Comorbidities included diabetes, hypertension, and tumors. Disease severity was assessed using the Acute Physiology and Chronic Health Evaluation (APACHE) II score and the Sequential Organ Failure Assessment (SOFA) score. Laboratory indicators included C-reactive protein (CRP), procalcitonin (PCT), lymphocytes, neutrophils, fibrinogen, D-dimer, urea nitrogen, total bilirubin, albumin, blood glucose, lactate, arterial carbon dioxide partial pressure (PaCO2), and arterial oxygen partial pressure (PaO2) /inhaled oxygen concentration (FiO2). Clinical characteristics included the duration of mechanical ventilation (MV), shock, and the use of glucocorticoids. All data were extracted from the Hospital Information System (HIS). To facilitate timely clinical interventions, the model emphasized early data. Therefore, the worst values within 48 h of ICU admission were used for severity scores and laboratory indicators.
Preprocessing of data
The data cleaning strategy included removing duplicate records and correcting obvious errors or inconsistencies. For missing data, multiple imputation methods were used to estimate missing values. Aggregation steps involved consolidating multiple data points, such as repeated blood pressure readings or scores, into a single representative value. The worst value recorded within 48 h of ICU admission was used, as it was considered the most accurate reflection of the patient’s disease severity. Outliers were identified using box plots and handled based on clinical significance and data distribution. Depending on the context, outliers were either corrected or removed. Continuous variables were normalized using the Z-score method, while categorical variables were encoded using the One-Hot Encoding method.
Selection of variables
The Recursive Feature Elimination (RFE) method was used to select characteristic variables and improve model stability. RFE is a feature selection technique that iteratively trains and removes the least important features until a predetermined number of features or performance criteria are met19. The process began with all candidate predictors and progressively reduced the feature set through RFE until the optimal subset of features for model training and testing was identified. This method effectively highlights the key features that contribute the most to the predictive power of the model.
Statistical analysis
Differences in baseline variables between the development and the validation cohort were compared. Additionally, differences in clinical information between the non-survivor and survivor groups within the development group were analyzed. The normal distribution of continuous variables was assessed using the Kolmogorov-Smirnov test. Continuous variables were compared using either Student’s t-test or the Mann-Whitney U-test, as appropriate. Categorical variables were analyzed using the Pearson chi-square test. Statistical significance was defined as p < 0.05 (two-sided). All analyses were conducted using Python (Version 3.10.10), Scikit-learn (Version 1.2.2), Shiny (Version 0.5.1), and R (Version 3.6.3).
Machine learning algorithms
The machine learning algorithms used in this study include Logistic Regression (LR), Random Forest (RF), Extreme Gradient Boosting (XGBoost), Light Gradient Boosting Machine (LightGBM), and Support Vector Machine (SVM). All five algorithms fall under the category of supervised learning.
LR is a statistical model commonly used for binary classification20. It estimates the probability that a given input belongs to a particular class using the logistic function. The algorithm models the relationship between input features and the log-odds of the target class probability. Specifically, LR finds the best-fitting linear equation for the log-odds of the outcome variable, rather than the outcome itself.
RF is an ensemble learning method based on decision trees21. It generates multiple decision trees during training and outputs the class that represents the majority vote across all trees. Each tree is trained on a random subset of the data, and at each node, a random subset of features is selected for splitting. This approach reduces overfitting and improves the model’s generalization ability.
XGBoost is an optimized implementation of the gradient boosting algorithm22. It builds an ensemble of decision trees by iteratively training new trees to correct the errors made by previous trees. The algorithm incorporates techniques like regularization and parallelization to enhance computational efficiency and reduce overfitting, making it particularly effective for large datasets.
LightGBM is an optimized gradient boosting framework that builds trees using a leaf-wise approach, enabling faster training. Two key innovations in LightGBM are Gradient-Based One-Side Sampling, which removes data with small gradients to focus on more informative data points, and Exclusive Feature Bundling, which combines mutually exclusive features to reduce dimensionality and enhance efficiency.
SVM is a classification algorithm that identifies a hyperplane to best separate data points of different classes23. The algorithm maximizes the margin between classes, ensuring the largest separation. For non-linear separations, SVM uses kernel functions to map data into higher-dimensional spaces where a linear separation becomes possible.
Development and validation of the models
The dataset from the development cohort was randomly divided into a 70% training set and a 30% testing set. Using the 70% training set, the models were trained multiple times with internal 5-fold cross-validation to optimize the hyperparameters, fit the parameters, and develop five machine learning models. The models were then tested on the 30% testing set to generate receiver operating characteristic (ROC) curves. Considering the APACHE II score as a premier predictive indicator for mortality among ICU patients, we conducted a performance comparison between the optimal machine learning model and the APACHE II score, as well as a model that excludes the APACHE II components. The area under the receiver operating characteristic curve (AUC) with 95% confidence intervals (CIs) was used to compare the performance of the models. To ensure the completeness of the evaluation, additional metrics such as the F1 score, accuracy, recall, precision, sensitivity, and specificity were also reported.
External 5-fold cross-validation was conducted using the entire development cohort to calculate and compare the average AUC for each model’s predictive performance. This step aimed to validate performance across different data divisions and reduce selection bias.
Finally, the dataset from the validation cohort served as an external validation center to assess the generalization ability of the models24. The AUC was recalculated to evaluate the predictive performance of each model.
Variable importance and web calculator
Using the model with the best average performance, Shapley Additive Explanations (SHAP) were applied to visualize the relative importance of variables in predicting mortality risk for patients with SCAP. Variables were ranked, and each was iteratively added to the model to calculate cumulative AUC values. When the cumulative AUC values stabilized, it indicated that adding additional variables had minimal impact on model performance. A web calculator was developed based on the most concise combination of variables. By entering the values of these key variables, the web calculator predicts the probability of in-hospital mortality for patients with SCAP, providing clinicians with a practical prognostic tool.
Results
Baseline characteristics and feature selection
A total of 455 patients with SCAP were included in the development cohort, and 120 patients with SCAP were included in the external validation cohort. The baseline characteristics of the two cohorts are summarized in Table 1. Patients in the development cohort were significantly younger than those in the external validation cohort (61.93 ± 17.77 vs. 69.63 ± 14.13 years, P < 0.001). The proportion of patients with tumors was lower in the development cohort compared to the external validation cohort (3.52% vs. 33.33%, P < 0.001). In contrast, the development cohort had significantly higher SOFA scores than the external validation cohort (8.37 ± 3.99 vs. 6.28 ± 2.77, P < 0.001). Laboratory indicators that showed significant differences between the development and external validation cohorts included PCT (10.41 ± 24.61 vs. 7.27 ± 22.34 ng/mL, P < 0.001), lymphocytes (1.07 ± 0.92 vs. 0.71 ± 0.90 × 10⁹/L, P < 0.001), D-dimer (8.07 ± 9.56 vs. 1.73 ± 3.22 µg/mL, P < 0.001), urea nitrogen (15.77 ± 11.37 vs. 12.00 ± 8.82 mmol/L, P < 0.001), albumin (33.56 ± 6.18 vs. 29.27 ± 4.87 g/L, P < 0.001), blood glucose (11.63 ± 5.24 vs. 9.65 ± 4.39 mmol/L, P < 0.001), and PaO₂/FiO₂ (247.17 ± 96.46 vs. 139.93 ± 64.37, P < 0.001). The proportion of patients with shock was higher in the development cohort compared to the external validation cohort (60.88% vs. 28.33%, P < 0.001). However, there was no statistically significant difference in mortality between the two cohorts (21.32% vs. 23.33%, P = 0.634).
Table 2 compares the characteristics of the non-survivor group (n = 97) and the survivor group (n = 358) in the development cohort. Variables with significant differences included APACHE II score (29.11 ± 8.67 vs. 19.86 ± 7.10, P < 0.001), SOFA score (10.61 ± 4.17 vs. 7.76 ± 3.72, P < 0.001), lymphocytes (1.57 ± 1.49 vs. 0.93 ± 0.62 × 10⁹/L, P < 0.001), D-dimer (11.91 ± 15.76 vs. 7.03 ± 6.66 µg/mL, P < 0.001), blood glucose (13.82 ± 8.22 vs. 11.04 ± 3.89 mmol/L, P < 0.001), lactate (6.92 ± 6.61 vs. 4.29 ± 3.42 mmol/L, P < 0.001), PaCO₂ (51.85 ± 22.12 vs. 43.70 ± 12.84 mmHg, P < 0.001), PaO₂ (218.54 ± 103.84 vs. 254.93 ± 93.01 mmHg, P < 0.001), and shock (90.72% vs. 52.79%, P < 0.001).
RFE method identified 23 characteristic predictors for model training and testing: APACHE II score, lymphocytes, PaCO₂, blood glucose, shock, PaO₂/FiO₂, D-dimer, PCT, lactate, CRP, total bilirubin, fibrinogen, urea nitrogen, albumin, SOFA score, duration of MV, age, neutrophils, use of glucocorticoids, diabetes, sex, hypertension, and tumors.
Comparison and selection of models
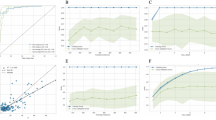
Five machine learning models were constructed and evaluated using internal 5-fold cross-validation. Figure 1 presents the ROC curves and AUC values for each model. The AUC for each model was as follows: LR 0.825 (95% CI: 0.732–0.919), RF 0.819 (95% CI: 0.732–0.905), XGBoost model 0.832 (95% CI: 0.743–0.921), LightGBM model 0.842 (95% CI: 0.757–0.927), and SVM 0.817 (95% CI: 0.735–0.900). Among these, the LightGBM model demonstrated the highest level of discrimination and accuracy. The results depicted in Fig. 2 demonstrate that the LightGBM model, which includes all features, outperforms both the APACHE II score (AUC = 0.763, 95% CI 0.625–0.902) and the LightGBM model without the APACHE II score (AUC = 0.822, 95% CI 0.740–0.903). Table 3 provides a detailed performance comparison of the five machine learning models. The LightGBM model achieved an F1 Score of 0.588, an accuracy of 0.745, a recall (sensitivity) of 0.893, a specificity of 0.706, and a precision of 0.439.
The receiver operating characteristic (ROC) curves for five machine learning models evaluated through internal 5-fold cross validation. The area under the curve (AUC) values are as follows: the LR model 0.825 (95% CI 0.732–0.919), the RF model 0.819 (95% CI 0.732–0.905), the XGBoost model 0.832 (95% CI 0.743–0.921), the LightGBM model 0.842 (95% CI 0.757–0.927), and the SVM model 0.817 (95% CI 0.735-0.900).
The ROC Curves for Mortality Prediction Models in Patients with Severe Community-Acquired Pneumonia (SCAP). The ROC curves illustrate the performance of three different models: (1) the APACHE II score (AUC = 0.763, 95% CI 0.625–0.902), (2) the LightGBM model excluding the APACHE II score components (AUC = 0.822, 95% CI 0.740–0.903), and (3) the comprehensive LightGBM model including all features (AUC = 0.842, 95% CI 0.757–0.927).
The results of the external 5-fold cross-validation are shown in Fig. 3. The solid lines represent the average ROC curves for each machine learning algorithm, while the shaded areas indicate their fluctuation ranges. The average AUC values for external 5-fold cross-validation were as follows: LR 0.813 ± 0.063, RF 0.819 ± 0.051, XGBoost 0.822 ± 0.032, LightGBM 0.842 ± 0.038, and SVM 0.806 ± 0.078. These results further confirmed that the LightGBM model outperformed the other models.
The average performance of five machine learning models evaluated through external 5-fold cross-validation. The solid line represents the average ROC curve, and the shaded area indicates the 95% confidence interval. The AUC values are: LR 0.813 ± 0.063, RF 0.819 ± 0.051, XGBoost 0.822 ± 0.032, LightGBM 0.842 ± 0.038, SVM 0.806 ± 0.078.
External validation results showed that the LightGBM model was reliable, with consistent predictive ability (AUC = 0.856, 95% CI: 0.792–0.921; Fig. 4). Based on these findings, the LightGBM model was selected as the final mortality risk prediction model.
External validation of the LightGBM model. The AUC value is 0.856 (95% CI 0.792–0.921).
Variable importance and explanation of the model
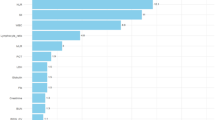
SHAP values were plotted based on the LightGBM model and are shown in Fig. 5. These visualizations illustrate the impact of each variable on the predicted outcome. Positive SHAP values, represented in red, indicate that higher feature values increase the probability of mortality, whereas negative SHAP values, represented in blue, indicate that lower feature values are associated with survival. The analysis revealed that APACHE II score, lymphocytes, PaCO₂, blood glucose, and shock were the most influential variables in predicting mortality. Higher values for these variables were strongly associated with increased mortality risk. Figure 6 displays a bar chart ranking features in decreasing order of importance, alongside a line chart of cumulative AUC values. The cumulative AUC values plateaued after including the top 10 variables, indicating that the addition of further variables had minimal impact on model performance. This finding suggests that using the top 10 predictors achieved the best balance between predictive performance and model complexity.
SHapley Additive Explanations (SHAP) are used to quantify the contribution of each variable in the LightGBM model. High feature values (red) and low feature values (blue) show their influence on model predictions. Positive contributions (right side) increase the prediction, while negative contributions (left side) decrease it.
LightGBM model’s feature importance and cumulative AUC fold plots. The bar chart illustrates the descending order of importance of the features. The line graph shows how the predictive parameters of the model change as the number of included features increases.
Implementation of the web calculator
To facilitate real-world application, we developed a web-based mortality calculator accessible at https://surge-ustc.shinyapps.io/acquired_pneumonia/. The calculator is based on five key predictors: APACHE II score, lymphocytes, PaCO₂, blood glucose, and shock. Users can input a patient’s clinical information into the provided fields to obtain a predicted risk of mortality (Fig. 7).
The web calculator for estimating the risk of in-hospital mortality in patients with SCAP admitted to the ICU based on the LightGBM model.
Discussion
The increasing availability of large datasets has enhanced the utility of machine learning algorithms, which often outperform traditional statistical methods in terms of accuracy and robustness25. A growing number of medical studies are adopting machine learning techniques to predict disease outcomes, including mortality risk26,27,28. This study is a retrospective study based on machine learning algorithms to develop and validate predictive models for mortality risk in patients with SCAP in ICUs at two hospitals in China. The study focused on creating clinically practical and accurate prediction models, using ICU hospital mortality as the study endpoint. This approach minimized bias that could arise from missing follow-up data or changes in treatment regimens after patient discharge. Results showed that all five models performed well, with AUC values above 0.8 during internal 5-fold cross-validation. Among them, the LightGBM model demonstrated the best performance (AUC = 0.842). External 5-fold cross-validation confirmed the LightGBM model’s superiority, with an average AUC of 0.842 ± 0.038, indicating stable predictive performance. Additionally, external validation at a separate center yielded an AUC of 0.856, further demonstrating the model’s strong generalizability.
The introduction of machine learning techniques significantly enhanced model performance. Compared to previous studies, the LightGBM model exhibited greater accuracy and generalizability, primarily due to the efficient processing of large clinical datasets and the algorithm’s ability to capture complex relationships among variables. This capability allows the model to make reliable predictions in diverse clinical scenarios. This innovation not only improves the scientific rigor of predictive modeling but also provides clinicians with a practical and powerful tool for clinical applications.
GBDT is a well-established model in machine learning, widely recognized for its effectiveness in handling structured data. LightGBM is a powerful framework that implements the GBDT algorithm while offering efficient parallel training capabilities. LightGBM provides several key advantages over traditional GBDT implementations, including faster training speeds, lower memory consumption, improved accuracy, and distributed support, enabling it to handle large datasets efficiently29,30. Our study selected a predictive model based on the LightGBM algorithm to assess mortality risk in patients with SCAP. The model incorporated 23 clinical predictors that are readily accessible in clinical practice, enhancing its applicability. While traditional feature importance methods provide only the relative importance of each feature variable, SHAP offers additional unique advantages. SHAP not only reflects the contribution of each feature to the model’ s predictions but also indicates the direction (positive or negative) of its impact. This improves both the interpretability and stability of the predictive model31,32,33,34. We applied SHAP to the LightGBM model to analyze the key predictors of mortality in patients with SCAP in the ICU. The results demonstrated that mortality risk increased with higher values of APACHE II score, lymphocytes, PaCO₂, blood glucose, and shock. Based on these five primary predictors, we developed a web calculator to predict the probability of SCAP-related mortality, providing a practical tool for clinical decision-making.
The classical APACHE II scoring system, one of the most commonly used indicators of disease severity in critically ill patients, played a vital role in our prediction model. High APACHE II scores at or shortly after ICU admission indicate poor organ function and are associated with worse outcomes. In a multicenter thromboprophylaxis study database, Guowei Li et al. identified higher APACHE II scores on admission as an independent risk factor for in-hospital mortality in ICU patients with pneumonia35. This was also verified in our study, where SHAP analysis showed the importance of the APACHE II score over other characteristics. Moreover, the integration of APACHE II with other variables improved the performance of the model. This result suggests that our model may be more specialized for the prognostic assessment of patients with SCAP.
The second significant component of the model was the blood lymphocyte count, which was collected within 48 h of ICU admission. Innate lymphocytes play a crucial role in the early inflammatory and immune response in patients with severe pneumonia, as evidenced by their increased numbers36,37,38,39. Furthermore, organ failure caused by infection can be classified as sepsis40. Sepsis initially manifests as a cytokine storm, whereas later stages are characterized by immune cell depletion, often signaling a poor prognosis41. In this study, some patients with severe pneumonia met the diagnostic criteria for sepsis, placing them at higher risk of mortality.
Elevated PaCO₂, or hypercapnia, was also identified as a risk factor for mortality in patients with SCAP. This may be attributed to the use of small tidal volume lung-protective ventilation strategies for patients with Acute Respiratory Distress Syndrome (ARDS) resulting from SCAP. Additional contributors could include undiagnosed Chronic Obstructive Pulmonary Disease (COPD) or acute airway obstruction due to sputum accumulation. Hypercapnia on admission is associated with the severity of CAP42, and some studies have confirmed that hypercapnia worsens the prognosis of patients with ARDS43. Specifically, hypercapnic acidosis within the first 24 h of ICU admission is strongly associated with increased in-hospital mortality44.
Acute illnesses, including severe infections, can disrupt glucocorticoid and cytokine metabolism, leading to increased hepatic glucose output and the occurrence of stress hyperglycemia45. Hyperglycemia is an independent determinant of in-hospital mortality in critically ill patients and has been shown to have high predictive value for poor clinical outcomes in patients with severe pneumonia46,47. The findings of this study further supported that elevated blood glucose is a significant predictor of mortality risk in patients with SCAP. Effective management of glucose metabolism disorders is crucial for reducing mortality in these patients. Shock was also identified as a critical factor in the prediction model. In this study, all cases of shock were classified as septic shock. As one of the primary diagnostic criteria for severe pneumonia18, the presence of shock predicts adverse clinical outcomes, with mortality rates as high as 51%48. Previous research has also demonstrated that septic shock is a major risk factor for death in patients with severe pneumonia6,49.
The strengths of this study are as follows. First, we trained and compared five different machine learning models and selected the best-performing model for predicting mortality risk in patients with SCAP. Second, external 5-fold cross-validation was employed for each model, minimizing bias associated with a single data partition. Third, the robust performance of the LightGBM model during external validation further confirmed its stability and reliability. Finally, this study offers clinical translational significance by providing a practical online prediction tool for ICU clinicians. The web-based calculator enables clinicians to identify high-risk patients in real time using their clinical and laboratory data, facilitating timely interventions. Interventions such as early adjustments to ventilation strategies and rapid treatment of infections are critical for improving survival outcomes in patients with SCAP.
Although we implemented various methods to improve the accuracy of our findings, several limitations of this study must be acknowledged. First, the study was conducted in two centers using retrospective data, which limits its geographic and population generalizability. Second, while the dataset size was sufficient for model development and validation, there remains a risk of model overfitting when handling high-dimensional features. Third, the data were extracted from the HIS of the two hospitals, which may have contained incomplete or inconsistent records, such as missing laboratory values or delays in scoring. Additionally, SHAP, while useful for interpretability, is computationally expensive for complex models. The analysis of high-dimensional data may also lead to information overload, potentially making the results less intuitive. Finally, manual data entry in the web calculator introduces operational complexity, which could hinder its adoption in clinical practice. Future prospective studies with larger sample sizes are needed to further validate the model’s predictive performance. Moreover, integrating the web calculator with electronic health record (EHR) systems and exploring alternative tools to improve the interpretability of machine learning models are key areas for future research50.
Conclusion
This study demonstrated that the LightGBM model is effective and stable in predicting mortality risk for patients with SCAP admitted to the ICU. Based on this model, we developed a web calculator incorporating five commonly used clinical characteristics. This tool has significant potential for clinical application and is worth promoting to aid in the early prediction of mortality risk in patients with SCAP.
Data availability
The data supporting this study’s findings are available from the corresponding author upon reasonable request.
Abbreviations
- CAP:
-
Community-acquired pneumonia
- HAP:
-
Hospital-acquired pneumonia
- ICU:
-
Intensive care unit
- SCAP:
-
Severe community-acquired pneumonia
- APACHE:
-
Acute Physiology and Chronic Health Evaluation
- SOFA:
-
Sequential Organ Failure Assessment
- CRP:
-
C-reactive protein
- PCT:
-
Procalcitonin
- PaCO2:
-
Arterial carbon dioxide partial pressure
- PaO2 :
-
Arterial oxygen partial pressure
- FiO2 :
-
Inhaled oxygen concentration
- MV:
-
Mechanical ventilation
- RFE:
-
Recursive feature elimination
- LR:
-
Logistic regression
- RF:
-
Random forest
- XGBoost:
-
Extreme gradient boosting
- LightGBM:
-
Light gradient boosting machine
- SVM:
-
Support vector machine
- GBDT:
-
Gradient boosting decision tree
- ROC:
-
Receiver operating characteristic
- AUC:
-
Area under the receiver operating characteristic curve
- CIs:
-
Confidence intervals
- SHAP:
-
Shapley addictive explanations
- ARDS:
-
Acute respiratory distress syndrome
- COPD:
-
Chronic obstructive pulmonary disease
- EHR:
-
Electronic health record
- HIS:
-
Hospital information system
References
Aliberti, S., Dela Cruz, C. S., Amati, F., Sotgiu, G. & Restrepo, M. I. Community-acquired pneumonia. Lancet 398, 906–919. https://doi.org/10.1016/S0140-6736(21)00630-9 (2021).
Jain, S. et al. Community-acquired pneumonia requiring hospitalization among U.S. adults. N. Engl. J. Med. 373, 415–427. https://doi.org/10.1056/NEJMoa1500245 (2015).
Martin-Loeches, I. et al. ERS/ESICM/ESCMID/ALAT guidelines for the management of severe community-acquired pneumonia. Eur. Respir J. 61 https://doi.org/10.1183/13993003.00735-2022 (2023).
Cavallazzi, R. et al. The burden of community-acquired pneumonia requiring admission to ICU in the United States. Chest 158, 1008–1016. https://doi.org/10.1016/j.chest.2020.03.051 (2020).
Dupuis, C. et al. Burden of pneumococcal pneumonia requiring ICU admission in France: 1-year prognosis, resources use, and costs. Crit. Care. 25 https://doi.org/10.1186/s13054-020-03442-z (2021).
Espinoza, R. et al. Factors associated with mortality in severe community-acquired pneumonia: A multicenter cohort study. J. Crit. Care. 50, 82–86. https://doi.org/10.1016/j.jcrc.2018.11.024 (2019).
Phua, J. et al. Severe community-acquired pneumonia: timely management measures in the first 24 hours. Crit. Care. 20, 237. https://doi.org/10.1186/s13054-016-1414-2 (2016).
Ullah, A. R., Masood, A., Amin, S. & Ali, I. Predictive factors and outcomes of severe community acquired pneumonia in patients with respiratory failure. Pak J. Med. Sci. 38, 1031–1037. https://doi.org/10.12669/pjms.38.4.5312 (2022).
Wang, X. et al. A new method to predict hospital mortality in severe community acquired pneumonia. Eur. J. Intern. Med. 40, 56–63. https://doi.org/10.1016/j.ejim.2017.02.013 (2017).
Song, Y. et al. Development and validation of a nomogram for predicting 28-day mortality on admission in elderly patients with severe community-acquired pneumonia. J. Inflamm. Res. 15, 4149–4158. https://doi.org/10.2147/JIR.S369319 (2022).
Handelman, G. S. et al. eDoctor: Machine learning and the future of medicine. J. Intern. Med. 284, 603–619. https://doi.org/10.1111/joim.12822 (2018).
Gutierrez, G. Artificial intelligence in the intensive care unit. Crit. Care. 24, 101. https://doi.org/10.1186/s13054-020-2785-y (2020).
Houthooft, R. et al. Predictive modelling of survival and length of stay in critically ill patients using sequential organ failure scores. Artif. Intell. Med. 63, 191–207. https://doi.org/10.1016/j.artmed.2014.12.009 (2015).
Giannini, H. M. et al. A machine learning algorithm to predict severe sepsis and septic shock: Development, implementation, and impact on clinical practice. Crit. Care Med. 47, 1485–1492. https://doi.org/10.1097/CCM.0000000000003891 (2019).
Sayed, M., Riano, D. & Villar, J. Novel criteria to classify ARDS severity using a machine learning approach. Crit. Care. 25, 150. https://doi.org/10.1186/s13054-021-03566-w (2021).
Odeyemi, Y. E. et al. Early machine learning prediction of hospitalized patients at low risk of respiratory deterioration or mortality in community-acquired pneumonia: derivation and validation of a multivariable model. Biomol. Biomed. https://doi.org/10.17305/bb.2023.9754 (2023).
Wang, B. et al. Novel pneumonia score based on a machine learning model for predicting mortality in pneumonia patients on admission to the intensive care unit. Respir. Med. 217, 107363. https://doi.org/10.1016/j.rmed.2023.107363 (2023).
Mandell, L. A. et al. Infectious Diseases Society of America/American Thoracic Society consensus guidelines on the management of community-acquired pneumonia in adults. Clin. Infect. Dis. 44 (Suppl 2), 27–72. https://doi.org/10.1086/511159 (2007).
Escanilla, N. S. et al. Recursive feature elimination by sensitivity testing. Proc. Int. Conf. Mach. Learn. Appl. 40–47 (2018). https://doi.org/10.1109/ICMLA.2018.00014 (2018).
Anley, D. T. et al. Prognostication of treatment non-compliance among patients with multidrug-resistant tuberculosis in the course of their follow-up: A logistic regression-based machine learning algorithm. Front. Digit. Health. 5, 1165222. https://doi.org/10.3389/fdgth.2023.1165222 (2023).
Zhou, C. M. et al. Machine learning predicts lymph node metastasis of poorly differentiated-type intramucosal gastric cancer. Sci. Rep. 11, 1300. https://doi.org/10.1038/s41598-020-80582-w (2021).
Hou, N. et al. Predicting 30-days mortality for MIMIC-III patients with sepsis-3: A machine learning approach using XGboost. J. Transl Med. 18, 462. https://doi.org/10.1186/s12967-020-02620-5 (2020).
Ozer, M. E., Sarica, P. O. & Arga, K. Y. New machine learning applications to accelerate personalized medicine in breast cancer: Rise of the support vector machines. Omics 24, 241–246 (2020). https://doi.org/10.1089/omi.2020.0001
Ramspek, C. L., Jager, K. J., Dekker, F. W., Zoccali, C. & van Diepen, M. External validation of prognostic models: What, why, how, when and where? Clin. Kidney J. 14, 49–58. https://doi.org/10.1093/ckj/sfaa188 (2021).
Segar, M. W. et al. Machine learning-based models incorporating social determinants of health vs traditional models for predicting in-hospital mortality in patients with heart failure. JAMA Cardiol. 7, 844–854. https://doi.org/10.1001/jamacardio.2022.1900 (2022).
Li, L. et al. Development and validation of machine learning-based models to predict in-hospital mortality in life-threatening ventricular arrhythmias: Retrospective cohort study. J. Med. Internet Res. 25, e47664. https://doi.org/10.2196/47664 (2023).
Ke, J. et al. Machine learning-based in-hospital mortality prediction models for patients with acute coronary syndrome. Am. J. Emerg. Med. 53, 127–134. https://doi.org/10.1016/j.ajem.2021.12.070 (2022).
Moll, M. et al. Machine learning and prediction of all-cause mortality in COPD. Chest 158, 952–964. https://doi.org/10.1016/j.chest.2020.02.079 (2020).
Rufo, D. D., Debelee, T. G., Ibenthal, A. & Negera, W. G. Diagnosis of diabetes mellitus using gradient boosting machine (LightGBM). Diagnostics (Basel). 11 https://doi.org/10.3390/diagnostics11091714 (2021).
Fu, X. Y. et al. Development and validation of LightGBM algorithm for optimizing of Helicobacter pylori antibody during the minimum living guarantee crowd based gastric cancer screening program in Taizhou, China. Prev. Med. 174, 107605. https://doi.org/10.1016/j.ypmed.2023.107605 (2023).
Zou, Y. et al. Extreme gradient boosting model to assess risk of central cervical lymph node metastasis in patients with papillary thyroid carcinoma: Individual prediction using SHapley Additive exPlanations. Comput. Methods Programs Biomed. 225, 107038. https://doi.org/10.1016/j.cmpb.2022.107038 (2022).
Valiente Fernandez, M., Gonzalez de Aledo, L., Delgado Moya, A. & Martin Badia, I. SHAP model explainability in ECMO-PAL mortality prediction: A critical analysis. Intensive Care Med. 49, 1559. https://doi.org/10.1007/s00134-023-07252-z (2023).
Wang, K. et al. Interpretable prediction of 3-year all-cause mortality in patients with heart failure caused by coronary heart disease based on machine learning and SHAP. Comput. Biol. Med. 137, 104813. https://doi.org/10.1016/j.compbiomed.2021.104813 (2021).
You, J. et al. Development of a novel dementia risk prediction model in the general population: A large, longitudinal, population-based machine-learning study. EClinicalMedicine 53, 101665. https://doi.org/10.1016/j.eclinm.2022.101665 (2022).
Li, G. et al. Risk factors for mortality in patients admitted to intensive care units with pneumonia. Respir. Res. 17, 80. https://doi.org/10.1186/s12931-016-0397-5 (2016).
Cheng, H. et al. Guards at the gate: Physiological and pathological roles of tissue-resident innate lymphoid cells in the lung. Protein Cell. 8, 878–895. https://doi.org/10.1007/s13238-017-0379-5 (2017).
Silver, J. S. et al. Inflammatory triggers associated with exacerbations of COPD orchestrate plasticity of group 2 innate lymphoid cells in the lungs. Nat. Immunol. 17, 626–635. https://doi.org/10.1038/ni.3443 (2016).
Huang, Y. et al. S1P-dependent interorgan trafficking of group 2 innate lymphoid cells supports host defense. Science 359, 114–119. https://doi.org/10.1126/science.aam5809 (2018).
Yang, D., Guo, X., Huang, T. & Liu, C. The role of group 3 innate lymphoid cells in lung infection and immunity. Front. Cell. Infect. Microbiol. 11, 586471. https://doi.org/10.3389/fcimb.2021.586471 (2021).
Angus, D. C. & van der Poll, T. Severe sepsis and septic shock. N. Engl. J. Med. 369, 840–851. https://doi.org/10.1056/NEJMra1208623 (2013).
Chousterman, B. G., Swirski, F. K. & Weber, G. F. Cytokine storm and sepsis disease pathogenesis. Semin. Immunopathol. 39, 517–528. https://doi.org/10.1007/s00281-017-0639-8 (2017).
Iqbal, N., Irfan, M., Zubairi, A. B. S., Awan, S. & Khan, J. A. Association of hypercapnia on admission with increased length of hospital stay and severity in patients admitted with community-acquired pneumonia: A prospective observational study from Pakistan. BMJ Open. 7, e013924. https://doi.org/10.1136/bmjopen-2016-013924 (2017).
Gendreau, S., Geri, G., Pham, T., Vieillard-Baron, A. & Mekontso Dessap, A. The role of acute hypercapnia on mortality and short-term physiology in patients mechanically ventilated for ARDS: A systematic review and meta-analysis. Intensive Care Med. 48, 517–534. https://doi.org/10.1007/s00134-022-06640-1 (2022).
Tiruvoipati, R., Pilcher, D., Buscher, H., Botha, J. & Bailey, M. Effects of hypercapnia and hypercapnic acidosis on hospital mortality in mechanically ventilated patients. Crit. Care Med. 45, e649–e656. https://doi.org/10.1097/CCM.0000000000002332 (2017).
Dungan, K. M., Braithwaite, S. S. & Preiser, J. C. Stress hyperglycaemia. Lancet 373, 1798–1807. https://doi.org/10.1016/S0140-6736(09)60553-5 (2009).
Lee, T. F. et al. Relative hyperglycemia is an independent determinant of in-hospital mortality in patients with critical illness. Crit. Care Med. 48, E115–E122. https://doi.org/10.1097/Ccm.0000000000004133 (2020).
Liu, B., Chen, Y., Yu, L. P. & Zhou, M. Stress hyperglycemia ratio is associated with systemic inflammation and clinical outcomes in diabetic inpatients with pneumonia on admission. J. Diabetes. https://doi.org/10.1111/1753-0407.13398 (2023).
Speiser, J. L. et al. Predicting in-hospital mortality in pneumonia-associated septic shock patients using a classification and regression tree: A nested cohort study. J. Intensive Care. 6, 66. https://doi.org/10.1186/s40560-018-0335-3 (2018).
Ferrer, M. et al. Severe community-acquired pneumonia: Characteristics and prognostic factors in ventilated and non-ventilated patients. PLoS One. 13, e0191721. https://doi.org/10.1371/journal.pone.0191721 (2018).
Rajkomar, A. et al. Scalable and accurate deep learning with electronic health records. NPJ Digit. Med. 1 https://doi.org/10.1038/s41746-018-0029-1 (2018).
Acknowledgements
Thanks for the data support provided by the First Affiliated Hospital of Anhui Medical University and Anhui Chest Hospital. We would like to thank Editage (www.editage.cn) for English language editing.
Funding
This work was supported by the Postgraduate Innovation Research and Practice Program of Anhui Medical University (No. YJS20230127 to Z.G.) as well as the Key Research and Development Program of Anhui Province (No. 2022e07020039).
Author information
Authors and Affiliations
Contributions
JP, TG, MS, and ZG conceived and designed the research. JP and TG drafted the manuscript. JP, HK and WB collected the data. JP and TG analyzed the data. JP and TG prepared the figures and tables. MS and ZG edited and revised the manuscript. MS and ZG approved the final version of manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
Informed consent of patients was waived by the First Affiliated Hospital Ethics Committee of Anhui Medical University and Anhui Chest Hospital Ethics Committee because this study was retrospective. This study was approved by the Ethics Committee of First Affiliated Hospital of Anhui Medical University (No. PJ2022-01-09) and Anhui Chest Hospital(No. KJ2023-40). We conducted the study in strict compliance with the Declaration of Helsinki.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Pan, J., Guo, T., Kong, H. et al. Prediction of mortality risk in patients with severe community-acquired pneumonia in the intensive care unit using machine learning. Sci Rep 15, 1566 (2025). https://doi.org/10.1038/s41598-025-85951-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-85951-x
Keywords
This article is cited by
-
Revolutionizing sepsis diagnosis using machine learning and deep learning models: a systematic literature review
BMC Infectious Diseases (2025)
-
Development and multi-database validation of interpretable machine learning models for predicting In-Hospital mortality in pneumonia patients: A comprehensive analysis across four healthcare systems
Respiratory Research (2025)
-
Artificial intelligence can extract important features for diagnosing axillary lymph node metastasis in early breast cancer using contrast-enhanced ultrasonography
Scientific Reports (2025)