Abstract
Accurately extracting organs from medical images provides radiologist with more comprehensive evidences to clinical diagnose, which offers up a higher accuracy and efficiency. However, the key to achieving accurate segmentation lies in abundant clues for contour distinction, which has a high demand for the network architecture design and its practical training status. To this end, we design auxiliary and refined constraints to optimize the energy function by supplying additional guidance in training procedure, thus promoting model’s ability to capture information. Specifically, for the auxiliary constraint, a set of convolutional structures are involved into a conventional network to act as a discriminator, then adversarial network is established. Based on the obtained architecture, we further build adversarial mechanism by introducing a second discriminator into segmentor for refinement. The involvement of refined constraint contributes to ameliorate training situation, optimize model performance, and boost its ability of collecting information for segmentation. We evaluate the proposed framework on two public databases (NIH Pancreas-CT and MICCAI Sliver07). Experimental results show that the proposed network achieves comparable performance to current pancreas segmentation algorithms and outperforms most state-of-the-art liver segmentation methods. The obtained results on public datasets sufficiently demonstrate the effectiveness of the proposed model for organ segmentation.
Similar content being viewed by others
Introduction
Distinguishing organs of interest from medical images is critical to synthesis disease diagnosis or state analysis in the field of medical imaging analysis. However, multiple organs (i.e. target area and non-target area) are contained in a set of images, the boundaries between organ and background may exist indistinct phenomenon, and the tissues in morphology and outline tend to be variable. All these factors bring challenge to accurate organ segmentation. In recent years, several medical imaging analysis methods based on neural networks are proposed for organ segmentation1,2,3,4,5,6. For example, Hu et al. firstly trained a three-dimensional convolutional neural network to predict the probability maps of target regions. Then, time-implicit multi-phase level-set is used to optimize current abdominal multi-organ segmentation model1. Chlebus et al. proposed an automatic liver tumor segmentation method based on a two-dimensional convolutional neural network, following an object-based postprocessing to refine the segmentation results2. Liu et al. raised an automatic framework to achieve organ segmentation from CT scans, where simple linear iterative clustering and support vector machine are used to roughly classify pixels, and convolutional neural network is involved to achieve precise boundary3. Ren et al. presented an interleaved three-dimensional convolutional neural network to segment small tissues of head and neck from CT images. Multiscale patches are firstly collected for more image appearance information, then the individual CNNs are interleaved to refine segmented tissues4. Huo et al. designed an automated liver attenuation estimation method, i.e., liver attenuation ROI-based measurement (ALARM). A previous deep convolutional neural network SS-Net is firstly used to segment liver. Based on liver segmentation results and morphological operations, several regions of interests are obtained to achieve attenuation estimation5.
In this paper, in order to preserve more details for segmentation, we design two sets of novel constraints (i.e. auxiliary constraintand refined constraint) on the basis of convolutional neural network to extract organs of interest from medical images. To obtain auxiliary constraint, we involve convolutional structure into the conventional segmentation model, and these two parts are trained in a manner of minmax two-player game, then a novel adversarial mechanism is built. The conventional segmentation model acts as a generator while the convolutional structure acts as a discriminator. This contributes to enhance the similarity between the predicted segmentation maps and their corresponding ground truths. Specifically, in each training iteration, the output probability maps from generator (i.e. segmentation network) and the corresponding truths are sent into discriminator (i.e. convolutional structure). Then, the discriminator measures the nuance between the synthetic sample and the original image distributions, where auxiliary constraint is used as the performance measurement. In the meantime, the generator starts training according to the obtained feedback from discriminator, with the purpose of synthesizing so authentic samples that cannot be distinguished from original images by the discriminator. This helps to improve the quality of the output segmentation maps.
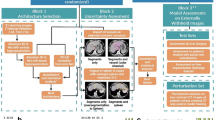
As for the refined constraint, we further apply a set of convolutional structure into the segmentation model with auxiliary constraint to build another group of adversarial mechanism, which contributes to preservation of detailed information for segmentation. Specifically, prior to training the adversarial network with auxiliary constraint, we firstly locate the deconvolutional layer which produces the predicted maps in current network. Then, we further involve an adversarial mechanism before the located deconvolutional layer. That is, a new convolutional structure is introduced as a discriminator to establish the second adversarial mechanism. At this moment, refined constraint is applied to measure the subtlety between the original and the synthetic distributions. The procedure described above is equivalent to supply two constraints on conventional segmentation model, contributing to preservation of details for segmentation. Figure 1 displays schematic illustration of the involved constraints in our proposed network.
Methods
Basic segmentation architecture
We employ U-Net7 to initially construct basic segmentation architecture in this work, which is an upgraded version of fully convolutional network (FCN). U-Net is primarily applied to medical image segmentation and most segmentation models apply it as baseline frame8,9,10,11. Compared to FCN, the main improvements of U-Net are the combined features from contracting path and symmetric expanding path, which contributes to yield more precise segmentation performance. The basic loss function is calculated as Eq. (1):
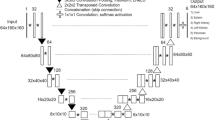
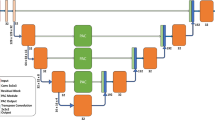
Ym denotes the synthetic samples predicted by generator while Yg denotes the corresponding images from original dataset. ω is the total number of volumes. The specific diagram of basic loss is displayed in Fig. 2. Specifically, the predicted probability maps from the fifth deconvolutional layer in basic segmentation model and their corresponding ground truths are measured by dice function, and then the Lbasic is obtained.
A schematic illustration of the proposed framework for extracting organs of interest from medical images, which includes different energy functions: Basic loss (LBasic), Auxiliary constraint (LAuxiliary), and Refined constraint (LRefined). Model 1 and Model 2 represent basic segmentation architectures with five and four deconvolutional layers, respectively. Structure 1 and Structure 2 represent the firstly and secondly involved convolutional layers which are used as discriminators, respectively.
Illustration of model architectures: deconvolutional layers (a) and (b) in segmentation frameworks Model 1 and Model 2, adversarial networks (c) and (b) which are consisted of convolutional layers. Samples from top to bottom: predicted maps from Model 1, corresponding ground truths, predicted maps from Model 2.
Auxiliary constraint for organ segmentation
The generative adversarial network proposed by Goodfellow et al.12 is capable of mastering the mapping rules between input and output samples, relying on its ability to capture spatial distributions knowledge of the training samples. Generative adversarial network consists of two competing parts (i.e. generator and discriminator) and it obeys a minmax two-player game to train the network. When inputting noise z into generator, a set of fake samples will be produced. And these samples and their corresponding ground truths are used as input images to train discriminator, which can be regarded as enlarging the training datasets for the discriminator. Then back propagation algorithm is involved to further update the parameters of generator. The above procedure is an iterative process. In this process, the ultimate goal of generator is to produce samples which are so authentic that the discriminator has difficulty distinguishing them from original images. The ultimate intention of discriminator is to accurately distinguish between synthetic samples and real images. The antagonistic and competitive relationship between generator and discriminator contributes to promote network performance of each other.
In order to preserve sufficient clues for organ segmentation, we introduce an auxiliary constraint into conventional U-Net7 to assist network training. Specifically, we involve three convolutional layers with kernels 7 × 7, 5 × 5 and 4 × 4 into conventional segmentation network, and these two structures are trained in a manner of minmax two-player game, then adversarial mechanism is obtained. The constructed adversarial learning provides an extra constraint for basic segmentation model, which ensures the segmentor training under the guidance of basic loss and the provided adversarial loss in each iteration process. This helps segmentor to obtain a further training and reach a steady state more quickly, thus achieving better segmentation performance. In this section, we defined the total energy function of segmentation model as auxiliary constraint. That is to say, the auxiliary constraint here performs a regression task, which consists of two items and lower values represent better performance, as shown in Eq. (2):
G and D represent generator and discriminator, and θG and θD respectively represent the parameters of G and D. In concrete practice process, the parameters θG are fixed when training D, while the parameters θD are fixed when training G. PG(seg) denotes the spatial distributions of the produced samples. DθD(GθG(Ym)) expresses the probability that the inputting images of D come from synthetic samples produced by G.
Error measurement (VOE, VD, ASD, RMSD, MSD and RMSE) distributions on NIH pancreas dataset from different models with basic loss (first column), auxiliary constraint (second column), and refined constraint (third column).
As displayed in Fig. 2, the predicted probability maps from the fifth deconvolutional layer in basic segmentation model and their corresponding ground truths are sent into the adversarial network, which are served as inputting images for discriminator. Meanwhile, the discriminator emits an extra constraint for segmentation network and an adversarial regression function for itself. The basic loss from segmentation model and the extra constraint from discriminator form the novel auxiliary constraint, as displayed in Eq. (2). And the adversarial regression function for discriminator is displayed in Eq. (3):
Similarity measurement (DSC, Precision, Recall and Jaccard) distributions on NIH pancreas dataset from different models with basic loss (first column), auxiliary constraint (second column), and refined constraint (third column).
PD(ground) denotes the spatial distributions of the original dataset. DθD(Yg) expresses the probability value that its inputting images are from original dataset.
Refined constraint for organ segmentation
A second set of convolutional layers are involved into the model with auxiliary constraint, to further refine network performance for more precise details. Specifically, based on the model with auxiliary constraint, another discriminator is introduced to construct a new adversarial mechanism, which is set behind the fourth deconvolutional layer in segmentor. In current segmentation model, the energy function from conventional network rules supreme compared with the involved adversarial constraints. This scheme gives the flexibility to use the specific competing relationship in GAN, which involves two constraints into segmentor to refine energy function, thus improve information preservation capacity to boost segmentation performance. The second set of convolutional layers in this part is identical to the discriminator in model with auxiliary constraint described in “Auxiliary constraint for organ segmentation”.
As shown in Fig. 2, on basis of the model with auxiliary constraint, the predicted probability maps from the fourth deconvolutional layer in basic segmentation model and their ground truths are measured by dice function, and then a second basic loss corresponding to current segmentor is obtained. Then, the predicted probability maps from the fourth deconvolutional layer and their corresponding ground truths are used as the inputting images for an adversarial network. Thus, an extra constraint for current segmentation network and a new adversarial regression function for this adversarial network are obtained. The refined constraint here performs a regression task, and it consists of four items, as shown in Eq. (4):
According to the primary and secondary relations of various factors affecting segmentation performance, the items of ω1, ω2, ω3, and ω4 in Eq. (4) are set to 1, 0.001, 0.1, and 0.0002, respectively.
The regression functions for discriminators are respectively displayed in Eqs. (5) and (6):
Results
Dataset and evaluation metrics
We involve two public datasets in this work to test model performance, including pancreas database collected by National Institutes of Health Clinical Center13,14 and Sliver07 provided by MICCAI liver segmentation competition15. The pancreas dataset consists of 82 contrast-enhanced CT volumes, and the size of each slice is 512 × 512. According to the training mode in16,17,18,19,20, we employ 4-fold cross validation to spit the dataset in this work. The Sliver07 contains 20 contrast-enhanced volumes with standard truths, and the size of each slice is 512 × 512.
Examples of pancreas segmentation results from models with different constraints. From left to right: original volumes from NIH dataset, the corresponding ground truths, predicted maps from model with basic loss, predicted maps from model with auxiliary constraint, and predicted maps from model with refined constraint.
Error measurement (VOE, VD, ASD, RMSD and RMSE) boxplots on Sliver07 dataset from model with basic loss (a), model with auxiliary constraint (b), and model with refined constraint (c).
There are two groups of metrics are applied for segmentation evaluation in this work, including error measurement and similarity measurement. The error measurement consists of six indexes: Volumetric Overlap Error (VOE), Relative Volume Difference (VD), Average Symmetric Surface Distance (ASD), Root Mean Square Symmetric Surface Distance (RMSD), Maximum Symmetric Surface Distance (MSD) and Root Mean Squared Error (RMSE). The similarity measurement consists of four indexes: Dice similarity coefficient (DSC), Precision, Recall and Jaccard. All experiments are carried on PyTorch environment and run on an NVIDIA GeForce GTX 1080Ti GPU of 11 GB memory.
Segmentation results on NIH dataset
Figure 3 shows error measurement distributions on NIH pancreas dataset from models with basic loss, auxiliary constraint and refined constraint, including metrics VOE, VD, ASD, RMSD, MSD and RMSE. Whereas Fig. 4 shows similarity measurement distributions of DSC, Precision, Recall and Jaccard from these three models. Figures 3 and 4 intuitively display segmentation result distributions from different models on NIH pancreas dataset, and the specific values of these ten sets of metrics are correspondingly shown in Table 1. Furthermore, Fig. 5 displays five groups of predicted maps from these three models to observe segmentation performance in an intuitive mode.
Similarity measurement (DSC, Precision, Recall and Jaccard) boxplots on Sliver07 dataset from (a) model with basic loss, (b) model with auxiliary constraint, and (c) model with refined constraint.
Segmentation results on Sliver07 dataset
Figure 6 presents boxplots of error measurement on Sliver07 dataset from segmentation models with basic loss, auxiliary constraint and refined constraint, including VOE, VD, ASD, RMSD and RMSE. And the similarity measurement distributions of DSC, Precision, Recall and Jaccard for these three models are displayed in Fig. 7. Figures 6 and 7 intuitively display boxplots from different models on Sliver07 dataset, and the specific values of these nine sets of metrics are correspondingly shown in Table 2. In order to intuitively exhibit the liver segmentation results, we display the output probability maps in Fig. 8.
Discussion
Segmentation results on NIH dataset
For error measurement, lower values equate to better segmentation performance in figures showing quantitative comparisons. It is obvious from Fig. 3 that the obtained error measurement from model with refined constraint represent strongest convergence in each indicator, compared with models with basic loss and auxiliary constraint. This indicates the involvement of refined constraint contributes to a more stable pancreas segmentation state. It also can be seen from Fig. 3 that the average level of each index corresponding to model with refined constraint show optimal state. This illustrates the application of refined constraint improves the model performance, which also effectively demonstrates its validity for pancreas segmentation.
For similarity measurement, higher values equate to better segmentation performance. As displayed in Fig. 4, the obtained similarity measurement from model with refined constraint show strongest convergence than another two models. This indicates the introduction of refined constraint improves the liver segmentation stability. It also can be observed from Fig. 4 that the mean values of each measurement corresponding to model with refined constraint show optimal level. This indicates refined constraint effectively improves liver segmentation performance, confirming the effectiveness of refined constraint for organ segmentation.
As displayed in Table 1, mean and standard deviation (std) of VOE for models with basic loss, auxiliary constraint and refined constraint are 36.3 ± 9.5, 32.5 ± 8.6 and 29.0 ± 7.4, while mean and std of ASD for these three models are 1.0 ± 0.8, 0.7 ± 0.5 and 0.5 ± 0.4. From these error measurement on pancreas dataset, we can conclude that the involvement of refined constraint achieves an optimized VOE of 7.3% and ASD of 0.5 mm, which strongly suggests its profit for boosting segmentation quality. Furthermore, the depressed std values illustrates the model performance gradually stabilized. Besides, the obtained results of DSC and Precision are 77.4 ± 7.6, 80.3 ± 6.4, 82.8 ± 5.3, 77.6 ± 9.7, 79.9 ± 9.6 and 83.1 ± 7.9, consecutively. This further demonstrates the improved maps quality and stabilized model performance. It can be observed from Fig. 5 that the predicted probability maps from basic model tend to represent chunks of deviations, and the involvement of auxiliary constraint ameliorates the situation. After this, refined constraint further improves segmentation performance even in tiny positions which are difficult to segment.
Examples of liver segmentation results from models with different constraints. From left to right: original volumes from Sliver07 dataset, the corresponding ground truths, predicted maps from model with basic loss, predicted maps from model with auxiliary constraint, and predicted maps from model with refined constraint.
Segmentation results on Sliver07 dataset
As shown in Fig. 6, the mean level of each metric box uniformly progressively decreased from (a) model with basic loss, (b) model with auxiliary constraint to (c) model with refined constraint. This declares the contribution of auxiliary constraint and refined constraint on promoting liver segmentation capability. Meanwhile, the mean level of each box in Fig. 7 progressively increased from (a) basic loss, (b) auxiliary constraint to (c) refined constraint, which further verifies the effectiveness of the involved auxiliary and refined l constraints. Table 2 displays mean and std of metrics for liver segmentation models with basic loss, auxiliary constraint and refined constraint. Their corresponding VOE values are 19.6 ± 3.2, 12.3 ± 3.1 and 7.2 ± 2.2 while ASD values are 1.8 ± 0.9, 0.7 ± 0.4 and 0.3 ± 0.2. Besides, the obtained liver segmentation results of DSC and Jaccard are 89.1 ± 2.0, 93.4 ± 1.7, 96.3 ± 1.2, 80.4 ± 3.2, 87.7 ± 3.1 and 92.8 ± 2.2, consecutively. The predicted segmentation maps are displayed in Fig. 8, it is obvious that the output maps from model with auxiliary constraint can deal better with main frame areas than basic loss. Furthermore, the employment of refined constraint helps to polish minor positions which are challengeable to distinguish from background in other two models.
Comparison to state-of-the-art
Our proposed model is compared with the-state-of-art pancreas segmentation methods to test its performance6,17,18,19,20,21,22,23,24. As can be shown in Table 3, the model with refined constraint achieves improvements with DSC of 0.47% (82.8% vs. 82.33%), Precision of 0.98% (83.1% vs. 82.12%) and Jaccard of 4.45% (74.8% vs. 70.35%) compared to Li et al.21. The mean DSC increases 1.53% (82.8% vs. 81.27%) and 0.4% (82.8% vs. 82.4%) compared to Roth et al.19 and Cai et al.22 and their Jaccard respectively increases 5.93% (74.8% vs. 68.87%) and 4.2% (74.8% vs. 70.6%). The DSC score of methods Liu et al.23 and Cai et al.24 are respectively 1.30% (82.8% vs. 84.10%) and 0.90% (82.8% vs. 83.70%) higher than the proposed model with refined constraint. While its obtained Jaccard optimizes 1.94% and 2.50% compared to Liu et al.23 (74.8% vs. 72.86%) and Cai et al.24 (74.8% vs. 72.30%). It can be observed that our proposed model exceeds most current technologies from different evaluation metrics.
The comparison in Table 4 shows our proposed model exceeds most liver segmentation methods3,25,26,27,28,29,30. The proposed model with refined constraint achieves the best liver segmentation performance with a mean DSC of 96.3%. The obtained RMSD optimizes 0.2 mm and 0.3 mm respectively compared to Wu et al.26 (2.3 mm vs. 2.5 mm) and Gauriau et al.27 (2.3 mm vs. 2.6 mm). The obtained ASD optimizes 0.93 mm, 0.99 mm and 1.00 mm compared to Wang et al.28 (0.3 mm vs. 1.23 mm), Wu et al.26 (0.3 mm vs. 1.29 mm) and Gauriau et al.27 (0.3 mm vs. 1.3 mm). Meanwhile, the obtained VOE optimizes 0.37% and 0.67% compared to Wang et al.28 (7.2% vs. 7.57%) and Wu et al.26 (7.2% vs. 7.87%). The DSC score of methods Liu et al.3 and Liao et al.29 are respectively 1.13% (96.3% vs. 97.43%) and 0.9% (96.3% vs. 97.2%) higher than our proposed model. While its obtained ASD optimizes 1.18 mm and 0.8 mm compared to Liu et al.3 (0.3 mm vs. 1.48 mm) and Liao et al.29 (0.3 mm vs. 1.1 mm).
Though the proposed model achieves desirable results on both pancreas and liver datasets, the network improvement of accuracy and application is still expected. Future research will fine-tune current architecture to cope better with pancreas and liver segmentation. Also, future work will apply the proposed model on multiple datasets to explore the latent potential to solve different organ segmentation tasks. In addition, we intend to conduct the existing model on medical imaging materials collected from clinical application, to explore its potential ability of addressing the clinical scenario.
Conclusion
This paper designs a convolutional neural network with auxiliary and refined constraints to segment organs from medical image. We introduce convolutional structure into basic segmentor, which is used as a discriminator, to construct adversarial mechanism for a higher segmentation quality. Auxiliary constraint at this moment is used to represent network regression condition. Furthermore, a new convolutional structure is involved into current framework to construct twofold adversarial mechanism. Aiming at collecting more sufficient information for segmentation through imposing dual constraints on segmentor. Refined constraint at this moment is used to represent network regression condition. Experiments are conducted on two public datasets and the obtained results demonstrate its superior performance over several state-of-the-art algorithms on pancreas and liver segmentation tasks. This adequately indicates the proposed method is a powerful and promising tool for organ segmentation.
Data availability
The Pancreas-CT and Sliver07 datasets are publicly available, and can be downloaded using the following links: Pancreas-CT: https://wiki.cancerimagingarchive.net/display/Public/Pancreas-CTSliver07: https://sliver07.grand-challenge.org/.
References
Hu, P. et al. Automatic abdominal multi-organ segmentation using deep convolutional neural network and time-implicit level sets. Int. J. Comput. Assist. Radiol. Surg. 12 (3), 399–411 (2017).
Chlebus, G. et al. Automatic liver tumor segmentation in CT with fully convolutional neural networks and object-based postprocessing. Sci. Rep. 8, 15497 (2018).
Liu, X., Guo, S., Yang, B., Ma, S. & Yu, F. Automatic organ segmentation for CT scans based on Super-pixel and Convolutional neural networks. J. Digit. Imaging. 31 (6), 748–760 (2018).
Ren, X. et al. Interleaved 3D-CNNs for joint segmentation of small-volume structures in Head and Neck CT images. Med. Phys. 45 (5), 2063–2075 (2018).
Huo, Y. et al. Fully automatic liver attenuation estimation combing CNN segmentation and morphological operations. Med. Phys. 46 (8), 3508–3519 (2019).
Fang, C., Li, G., Pan, C., Li, Y. & Yu, Y. Globally guided progressive fusion network for 3D pancreas segmentation. In Paper presented at: International Conference on Medical Image Computing and Computer-Assisted Intervention (2019).
Ronneberger, O., Fischer, P., Brox, T. & U-Net convolutional networks for biomedical image segmentation. In Paper presented at: International Conference on Medical Image Computing and Computer-Assisted Intervention (2015).
Sundaresan, V., Zamboni, G., Rothwell, P. M., Jenkinson, M. & Griffanti, L. Triplanar ensemble U-Net model for white matter hyperintensities segmentation on MR images. Med. Image. Anal. 73, 102184 (2021).
Zeng, G., Xin, Y., Jing, L., Yu, L. & Zheng, G. 3D U-net with multi-level deep supervision: fully automatic segmentation of proximal femur in 3D MR images. In Paper presented at: Machine Learning in Medical Imaging (2017).
Hao, D., Yang, G., Liu, F., Mo, Y. & Guo, Y. Automatic brain tumor detection and segmentation using U-Net based fully convolutional networks. In Annual Conference on Medical Image Understanding and Analysis (2017).
Balagopal, A., Kazemifar, S., Dan, N., Lin, M. H. & Jiang, S. B. Fully automated organ segmentation in male pelvic CT images. Phys. Med. Biol. 63 (24), 245015 (2018).
Goodfellow, I. et al. Generative adversarial nets. In Paper presented at: Advances in Neural Information Processing Systems (2014).
Clark, K. et al. The Cancer Imaging Archive (TCIA): maintaining and operating a public information repository. J. Digit. Imaging. 26 (6), 1045–1057 (2013).
Roth, H. R. et al. Data from Pancreas-CT. (2016). The cancer imaging archive.
Heimann, T., Ginneken, B. V., Styner, M. A., Arzhaeva, Y. & Wolf, I. Comparison and evaluation of methods for liver segmentation from CT datasets. IEEE Trans. Med. Imaging. 28 (8), 1251–1265 (2009).
Roth, H. R. et al. Deeporgan: Multi-level deep convolutional networks for automated pancreas segmentation. In Paper presented at: International Conference on Medical Image cComputing and Computer-Assisted Intervention (2015).
Yu, Q. et al. Recurrent saliency transformation network: Incorporating multi-stage visual cues for small organ segmentation. In Paper presented at: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (2018).
Zhou, Y. et al. A fixed-point model for pancreas segmentation in abdominal CT scans. In Paper presented at: International Conference on Medical Image Computing and Computer-Assisted Intervention (2017).
Roth, H. R. et al. Spatial aggregation of holistically-nested convolutional neural networks for automated pancreas localization and segmentation. Med. Image. Anal. 45, 94–107 (2018).
Ma, J., Lin, F., Wesarg, S. & Erdt, M. A novel bayesian model incorporating deep neural network and statistical shape model for pancreas segmentation. In Paper presented at: International Conference on Medical Image Computing and Computer-Assisted Intervention (2018).
Li, M., Lian, F. & Guo, S. Pancreas segmentation based on an adversarial model under two-tier constraints. Phys. Med. Biol. 65 (22), 225021 (2020).
Cai, J., Lu, L., Xie, Y., Xing, F. & Yang, L. Improving Deep Pancreas Segmentation in CT and MRI Images Via Recurrent Neural Contextual Learning and Direct Loss Function (Medical Image Computing and Computer-Assisted Intervention (MICCAI), 2017).
Liu, S. et al. Automatic pancreas segmentation via coarse location and ensemble learning. IEEE Access. 8, 2906–2914 (2020).
Cai, J., Lu, L., Xing, F. & Yang, L. Pancreas segmentation in CT and MRI images via domain specific network designing and recurrent neural contextual learning. arXiv:180311303. (2018).
Dou, Q. et al. 3D deeply supervised network for automatic liver segmentation from CT volumes. In International Conference on Medical Image Computing and Computer-Assisted Intervention (2016).
Wu, W., Zhou, Z., Wu, S. & Zhang, Y. Automatic liver segmentation on volumetric CT images using supervoxel-based graph cuts. Comput. Math. Methods Med. 2016, 9093721 (2016).
Gauriau, R., Cuingnet, R., Prevost, R., Mory, B. & Bloch, I. A generic, robust and fully-automatic workflow for 3D CT liver segmentation. Comput. Clin. Appl. ABD-MICCAI. (2013).
Wang, J., Cheng, Y., Guo, C., Wang, Y. & Tamura, S. Shape-intensity prior level set combining probabilistic atlas and probability map constrains for automatic liver segmentation from abdominal CT images. Int. J. Comput. Assist. Radiol. Surg. 11 (5), 817–826 (2016).
Liao, M. et al. Efficient liver segmentation in CT images based on graph cuts and bottleneck detection. Phys. Med. 32 (11), 1383–1396 (2016).
Li, G. et al. Automatic Liver Segmentation based on shape constraints and deformable graph cut in CT images. IEEE Trans. Image Process. 24 (12), 5315–5329 (2015).
Author information
Authors and Affiliations
Contributions
L.F.H.: Conceptualization, methodology, formal analysis, writing-original draft. S.Y.J.: Software, validation. L.M.Y.: Conceptualization, formal analysis, supervision, funding acquisition, project administration, writing-review and editing. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
In this paper, all methods were carried out in accordance with relevant guidelines and regulations. All cases were used with the approval of the National Institutes of Health Clinical Center (NIH) and MICCAI liver segmentation competition, and all participants provided informed consent.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lian, F., Sun, Y. & Li, M. Extracting organs of interest from medical images based on convolutional neural network with auxiliary and refined constraints. Sci Rep 15, 2036 (2025). https://doi.org/10.1038/s41598-025-86087-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-86087-8