Abstract
Currently, the identification of valuable candidate genes affecting milk fat synthesis in dairy cows is still limited, and the specific regulatory mechanism is still unknown. In this study, we used primary bovine mammary epithelial cells(BMECs)as a model and utilized overexpression and knockdown techniques for the PI4K2A gene to investigate the specific mechanisms by which it regulates lipid metabolism in BMECs. We studied whether PI4K2A regulates the inhibition of trans-10, cis-12 conjugated linoleic acid (t10,c12-CLA) on lipid synthesis in BMECs. The results revealed that PI4K2A plays a crucial regulatory role within the cytoplasm of BMECs, positively impacting fatty acid (FA) transport, triacylglycerol (TAG) synthesis, and lipid droplet secretion (p < 0.05 for all effects). Furthermore, PI4K2A promotes cell proliferation and inhibits apoptosis (p < 0.05). This study also confirmed that overexpressing the PI4K2A gene can effectively reduce the inhibitory effect of t10,c12-CLA on lipid synthesis in BMECs. Our findings have deepened the understanding of the regulatory mechanisms of milk fat synthesis and provide potential genetic targets for the improvement of lactation traits in dairy cows.
Similar content being viewed by others
Introduction
Milk fat is one of the most important milk components in dairy cattle1,2,3. The weight of milk fat in the performance index for dairy cows varies between 6% and 25% across different countries, which proves its importance in dairy breeding. Milk fat synthesis in dairy cows involves complex regulatory networks, including de novo synthesis of fatty acids (FA), uptake of long-chain fatty acids (LCFA) in blood, LCFA transport and desaturation, triglyceride (TAG) synthesis and lipid droplet secretion4. Thanks to the rapid advancement of next-generation sequencing technology and the subsequent reduction in sequencing cost, breeding researchers have successfully identified numerous effective genes and molecular markers, which play pivotal roles in milk fat metabolism. At present, the identification of valuable candidate genes of milk fat synthesis is still limited, and the regulatory mechanism of milk fat synthesis is still unclear5.
For this reason, our lab studied the expression of mRNA transcripts in primary bovine mammary epithelial cells (BMECs) at the peak of lactation in dairy cows with high and low milk fat content by using comparative transcriptome sequencing6. Weighted gene coexpression network analysis (WGCNA) was used to analyze the mRNA expression profile data, and identified 15 candidate differential genes (Table S1) that may regulate milk fat metabolism7. To further pinpoint and validate the most critical candidate genes, we analyzed the tissue expression profiles of 15 candidate genes by using quantitative real-time polymerase chain reaction (qRT-PCR). We found that the mRNA expression level of PI4K2A in mammary tissue (Table S2 for the full gene names) was significantly higher than that in small intestine, liver, kidney, heart and ovary. Furthermore, PI4K2A expression was higher and more stable in both mammary tissue and BMECs.
PI4KIIa (PI4K2A) is an integral membrane protein characterized by a proline-rich amino-terminal domain (1–93) and plays a pivotal role in cell signal transduction (Wnt) and transport processes8. PI4K2A is found to be highly expressed in prolactin-treated BMECs, indicating a potential involvement in lactation processes9. Palombo, et al.10 found that PI4K2A is an important candidate gene for milk fat metabolism in Italian Simmental and Holstein cows. Consequently, based on previous research conducted by our research group, we have identified PI4K2A as a key candidate gene that regulates lipid synthesis in BMECs. To understand the specific mechanism by which PI4K2A regulates milk fat synthesis in dairy cows, we used primary BMECs as a model to detect the expression of marker genes and transcription factors associated with milk fat synthesis. At the same time, we studied the effects on TAG synthesis, lipid droplet secretion, as well as the proliferation and apoptosis of BMECs. Trans-10, cis-12 conjugated linoleic acid (t10,c12-CLA) plays an important role in inhibiting fat deposition and lowering blood lipids11,12,13,14,15. The addition of t10,c12-CLA can increase the phosphatidylinositol (PI) content in milk while reducing the milk fat content16, and PI is the main substrate17 of PI4K2A. However, it still unclear whether PI4K2A will influence the inhibitory effect of t10,c12-CLA on milk fat. Therefore, another objective of this study was to investigate the relationship between PI4K2A and t10,c12-CLA in lipid synthesis. The results provide valuable genetic marker information for milk composition traits in dairy cows, and are crucial for genetic improvement.
Results
Identification of key candidate gene for milk fat metabolism in dairy cows
The qRT-PCR results demonstrated that the relative expression of the ENPP2 gene (2.17 ± 0.76) in the mammary gland was significantly higher than that in other tissues (p < 0.05). The expression level of the PI4K2A gene in the mammary gland (1.40 ± 0.26) was slightly higher than in the uterus (1.31 ± 0.14) and considerably higher than in other tissues (p < 0.05). CTSH gene was highly expressed in both the mammary gland and small intestine tissues, with expression levels of 1.18 ± 0.23 and 1.00 ± 0.03, respectively. The expression level of PTPRR was highest in the mammary gland (3.35 ± 0.67), significantly higher than that in small intestine, liver and kidney (p < 0.05). The expression of ID1 and PDGFD in the mammary gland were lower than those in the uterus but significantly higher than those in other tissues (p < 0.05). The expression level of the ATP8A2 gene in the mammary gland was lower than that in the ovary but significantly higher than that in other tissues (p < 0.05). The expression level of VEGFD, ZFYVE28, DKK1, APOL3, SLC16A1, CES4A, KCNMA1 and BCAT1 genes in mammary gland were moderate or low compared with those in other tissues (Fig. 1).
Expression of candidate differential genes for milk fat metabolism in different tissues of dairy cows, respectively. Distinct lowercase letters showed significant difference (p < 0.05), while the same lowercase letters showed no significant difference (p > 0.05). (mean ± SD, n = 3).
The expression patterns of most candidate genes in the mammary gland were consistent with those in BMECs (Fig. 2A,B). SLC16A1, PI4K2A, CTSH, APOL3, BCAT1 and DKK1 were highly expressed in both the mammary gland and BMECs, whereas ATP8A2, ID1 and ZFYVE28 showed low expression levels in both tissues. The expression of SLC16A1 in both the mammary gland and BMECs was higher than other genes. However, the expression of SLC16A1 gene in the mammary gland is not prevalent across various tissues. Therefore, it is no longer considered a key candidate gene for milk fat metabolism in cows. Except for small intestine, the expression of CTSH gene in the mammary gland was significantly higher than that in other tissues (p < 0.05). However, our previous study revealed that the CTSH gene was significantly down-regulated in the group with high milk fat content (p < 0.05), which has led us to decide against pursuing further functional verification. PI4K2A is an important candidate gene for milk fat metabolism in cows. It has the highest expression level in the mammary gland compared with other tissue, and the expression of PI4K2A is relatively high in both the mammary gland and BMECs (ranked third). Furthermore, our group has demonstrated that the PI4K2A gene is differentially expressed between groups with high and low milk fat content, specifically being upregulated in the group with high milk fat content (Table S1). Therefore, this study will focus on the functional mechanism of the PI4K2A gene in lipid metabolism of BMECs.
Subcellular localization of PI4K2A and differential expression of candidate genes in mammary gland and BMECs, respectively. (A) Differential expression of candidate genes involved in milk fat metabolism in mid-lactation mammary tissue, while the same lowercase letters indicate no significant difference; (B) Differential expression of candidate genes associated with milk fat metabolism variations in mid-lactation BMECs; (C) Results of total nuclear and plasma RNA extraction from BMECs; D: qRT-PCR analysis to detect the distribution of PI4K2A in BMECs. (Different lowercase letters indicate significant differences (p < 0.05). (mean ± SD, n = 3,*p < 0.05).
Subcellular localization of PI4K2A
The subcellular localization of the PI4K2A protein indicated that it was mainly located in the cytoplasm (86.0–87.1%, Table 1). Although the PSORT II software predicted a lower PI4K2A protein content in the cytoplasm (34.8%), the trend was consistent across the four software. To further confirm the authenticity of the prediction results, we extracted both nuclear and cytoplasmic RNA from BMECs using Cytoplasmic & Nuclear RNA Purification Kit (NORGREN, UK). The extraction of cytoplasmic and nuclear RNA was shown in Fig. 2C. The qRT-PCR analysis showed that PI4K2A was expressed in both the nucleus and cytoplasm, with approximately 80% in the cytoplasm, in agreement with the prediction (Fig. 2D).
The effect of PI4K2A on the expression of milk fat metabolism-related genes in BMECs
To investigate the effect of PI4K2A gene on milk fat metabolism, BMECs were infected with Ad-PI4K2A adenovirus for 48 h, and the mRNA expression levels of milk fat synthesis-related marker genes were detected using qRT-PCR. The results showed that overexpressing PI4K2A significantly upregulated the mRNA expression of the FA transport marker gene FABP3 (Fig. 3C, p < 0.01), but had no significant effect on the mRNA expression of genes involved in de novo FA synthesis, LCFA uptake from blood and desaturation (Fig. 3A,B,C). In addition, we found that overexpressing PI4K2A upregulated the expression of the ELOVL6 gene (Fig. 3D) and significantly increased the mRNA expression level of PPARG and ACOX1 (p < 0.01/p < 0.05, Fig. 3D). In contrast, PPARD showed an opposite trend (p < 0.05, Fig. 3D). Subsequently, we randomly selected three significantly different genes for Western Blot detection. The results showed PPARG, PPARD, and FABP3 protein levels matched their mRNA levels (Fig. 3E,F), suggesting that PI4K2A can promote FA transport and LCFA elongation. We hypothesize that PI4K2A can enhance lipid synthesis in BMECs because most lipogenic gene expression was upregulated after PI4K2A overexpression.
The effect of overexpression of PI4K2A gene on milk fat metabolism in dairy cows. (A) Marker genes related to de novo synthesis of FA; (B) Marker genes related to uptake of LCFA from blood; (C) Marker genes for FA transport and desaturation; D: Transcriptional regulators and other genes related to milk fat synthesis; (E, F) The detection of PPARG, FABP3 and PPAR protein expression by using Western Blot; (G) TAG synthesis-related marker genes; (H) TAG catabolism-related marker genes; (I, J) The detection of DGAT1 protein expression by using Western Blot; (K) The effect of overexpression of PI4K2A gene on triglyceride content of BMECs; (L) Effect of overexpression of PI4K2A gene on cholesterol content of BMECs; (M) The effect of overexpression of PI4K2A gene on marker genes related to lipid droplet secretion in BMECs; (N, O) The effect of overexpression of PI4K2A gene on lipid droplet secretion of BMECs; (P) Results of enzymatic assay of secreted lipid droplets. (mean ± SD, n = 3,*p < 0.05, **p < 0.01).
The qRT-PCR results showed that PI4K2A could significantly inhibit the marker genes of milk fat synthesis. Among these genes, the mRNA expression of key genes for de novo FA synthesis, LCFA uptake in blood and FA desaturation (SCD) was significantly reduced after PI4K2A interference (p < 0.01/p < 0.05, Fig. 4A,B,C). Additionally, although there was a decreasing trend in the mRNA expression of key genes related to FA synthesis and transport, the difference was not significant (p > 0.05). Furthermore, the mRNA expression of all genes and transcription factors involved in milk fat synthesis showed a decreasing trend (Fig. 4D), and the downregulation of ELOVL6, EGFR, PPARA, and PPARD genes reached a significant level (p < 0.01/p < 0.05). Then, we randomly selected four genes with decreased expression for Western Blot analysis, and the results showed that the protein expression of PPARG, PPARD, SCD, and FABP3 were consistent with their mRNA levels (Fig. 4E,F). These results suggest that interfering with the PI4K2A gene can inhibit de novo FA synthesis, LCFA uptake from blood, FA transport, and desaturation processes in BMECs.
Effect of interfering with PI4K2A gene on milk fat metabolism in dairy cows. (A) Marker genes related to de novo synthesis of FA; (B) Marker genes related to uptake of LCFA from blood; (C) Marker genes for FA transport and desaturation; (D) Other related genes involved in milk fat synthesis and their transcriptional regulators; (E, F) The expression of PPARG, FABP3, SCD and PPARD proteins by Western Blot; (G) TAG synthesis-related marker gene; (H) TAG catabolism-related marker gene; (I) Effect of interfering with PI4K2A gene on TAG content of BMECs; (J) The effect of interfering with PI4K2A gene on cholesterol content of BMECs; (K) The effect of interference with PI4K2A gene on marker genes related to lipid droplet secretion in BMECs; (L, M) The effect of interfering with PI4K2A gene on lipid droplet secretion of BMECs; (N) The results of enzymatic assay of secreted lipid droplets. (mean ± SD, n = 3,*p < 0.05, **p < 0.01).
The effect of PI4K2A on TAG synthesis and lipid droplet secretion in BMECs
To determine whether the PI4K2A gene affects TAG synthesis in BMECs, we examined the expression of TAG synthesis-related marker genes using qRT-PCR. Our results showed that overexpressing PI4K2A can significantly increase the mRNA and protein expression of DGAT1 (Fig. 3G,I,J, p < 0.05). Furthermore, we studied marker genes of TAG degradation and discovered that mRNA expression of the ATGL gene was significantly reduced after overexpressing PI4K2A (Fig. 3H, p < 0.01). So, the TAG content in BMECs was significantly increased (Fig. 3K, p < 0.05), while the cholesterol content remained unchanged (Fig. 3L). The expression of genes related to lipid droplet secretion in BMECs was significantly upregulated after overexpressing PI4K2A (Fig. 3M), and Oil Red O staining showed a significant increase in the lipid droplets secreted by BMECs (Fig. 3N,O). Furthermore, the results of absorbance values was consistent with that of Oil Red O staining (Fig. 3P). Therefore, the overexpression of PI4K2A in BMECs promotes lipid synthesis, fatty acid transport, TAG synthesis, and lipid droplet secretion, and significantly inhibits TAG degradation.
The results of marker genes TAG synthesis and decomposition showed that interfering PI4K2A could significantly reduce the mRNA expression of the AGPAT6 for TAG synthesis (Fig. 4G, p < 0.01), but had no effect on TAG decomposition (Fig. 4H). Compared with the NC group, interfering with PI4K2A significantly reduced TAG content in BMECs (Fig. 4I, p < 0.01). Additionally, the total cholesterol content exhibited a downward trend, although it did not reach statistical significance (Fig. 4J, P = 0.058). The mRNA expression of lipid droplet secretion-related marker genes (XDH, BTN1A1, and PLIN2) was significantly down-regulated after interfering with PI4K2A (Fig. 4K, p < 0.05). The lipid droplet secretion in the siRNA-PI4K2A group was significantly lower than that in the NC group (Fig. 4L,M,N, p < 0.05). In conclusion, interference with PI4K2A gene can inhibit the lipid synthesis. The results from overexpression and interference experiments indicated that PI4K2A gene positively regulated lipid synthesis of BMECs.
The effect of PI4K2A on proliferation and apoptosis in BMECs
After overexpressing PI4K2A gene for 48 h, the number of proliferating cells in Ad-PI4K2A group was significantly higher than that in Ad-EGFP group, indicating that overexpression of PI4K2A gene could significantly increase proliferation rate of BMECs (p < 0.05, Fig. 5A,B,C,D,E,F,G). The protein expression of CCND1 in Ad-PI4K2A group was also significantly higher than that in Ad-EGFP group (p < 0.05), aligning with the EdU detection results (Fig. 5H,I). The results of the PI4K2A gene interference experiments showed the opposite side, indicating that PI4K2A positively regulates the proliferation of BMECs (Fig. 5J,K,L,M,N,O,P,Q,R).
Effect of overexpression/interference of PI4K2A gene on BMECs proliferation. (A–C) Results of EdU treatment after Ad-EGFP adenovirus transfection (all taken from the same field of view, the same below). Where A is the cytoplasm of proliferating cells stained red; B is the nucleus of all cells stained blue; C is the combined image of A and B; (D–F) Results of EdU treatment after transfection with Ad-PI4K2A adenovirus; (G) EdU labeling index obtained by ImageJ software after counting proliferating cells and all cells in the field of view, respectively; (H) Expression of G1/S-specific CCND1 after overexpression of PI4K2A; (I) Relative protein expression calculated after grayscale scanning of protein bands by ImageJ software; (J–L) Results of EdU treatment after NC transfection; (M–O) Results of EdU treatment after transfection with siRNA-PI4K2A; (P) EdU labeling index obtained by ImageJ software after counting proliferating cells and all cells in the field of view, respectively; (Q) Expression of CCND1 after interference with PI4K2A; (R) Relative protein expression calculated after grayscale scanning of protein bands by ImageJ software. (mean ± SD, n = 3,*p < 0.05, **p < 0.01).
The apoptosis of BMECs was detected when BMECs were infected with Ad-EGFP and Ad-PI4K2A for 48 h, respectively. The results showed that overexpressing PI4K2A gene significantly reduced apoptosis of BMECs (p < 0.05, Fig. 6A,B,C,D,E,F,G,H,I). Furthermore, the protein expression of apoptotic marker genes (caspase-3) in the Ad-PI4K2A group was also significantly lower than that in the Ad-EGFP group (p < 0.05), which was consistent with the results of apoptosis kit (Fig. 6J,K). In addition, the results of interfering with PI4K2A gene showed the opposite side (Fig. 7), indicating that PI4K2A negatively regulates the apoptosis of BMECs.
Effect of overexpression of PI4K2A gene on apoptosis of BMECs. (A–D) Results of apoptosis assay after transfection with Ad-EGFP adenovirus (all taken in the same field of view, the same below). Where A is the BMECs under white light, B is the result of nuclear staining, the nuclei of apoptotic cells were stained with green fluorescence by SYTOX Green, C is the result of cell membrane staining, the cell membrane of apoptotic cells were stained with red fluorescence by Annexin V-mCherry, D is the combined image of B and C; (E–H) The results of apoptosis detection after transfection with Ad-PI4K2A adenovirus; (I) Apoptosis index calculated by ImageJ software after counting apoptotic cells and all cells, respectively; (J) Protein expression of caspase-3 after overexpression of PI4K2A; (K) Relative protein expression calculated by ImageJ software after grayscale scanning of protein bands. (mean ± SD, n = 3,*p < 0.05).
Effect of interfering with PI4K2A gene on apoptosis of BMECs. (A–D) Results of the apoptosis assay following transfection with NC; (E–H) Outcomes of apoptosis detection after transfection with siRNA-PI4K2A; (I) Apoptosis index determined by ImageJ software after counting apoptotic cells and total cells, respectively; (J) Protein expression of caspase-3 upon overexpression of PI4K2A; (K) Relative protein expression computed by ImageJ software after grayscale scanning of protein bands. (mean ± SD, n = 3,*p < 0.05, **p < 0.01).
The impact of adding t10,C12-CLA on lipid synthesis in BMECs
The expression of marker genes for de novo FA synthesis (ACSS2, ACACA, FASN), LCFA uptake (ACSL1 and CD36), FA transport (FABP3), TAG synthesis (DGAT1, AGPAT6) and TAG degradation (ATGL) was significantly down-regulated in the t10,c12-CLA group by comparison with the control (Fig. 8A,B,C,D,E, p < 0.05). Notably, the expression of the TAG synthetic marker gene LPIN1 was significantly up-regulated following treatment with t10,c12-CLA (p < 0.05). Meanwhile, the mRNA expression of lipogenic metabolism transcription factors (PPARD, PPARG, ACOX1, CPT1A, ELOVL6, and EGFR) in the t10,c12-CLA group was significantly lower than that in the control group (Fig. 8F, p < 0.05). Therefore, t10,c12-CLA significantly inhibited FA de novo synthesis, LCFA uptake in blood, and FA transport process (p < 0.05). Additionally, it can regulate the lipid synthesis of BMECs by down-regulating transcription factors related to lipogenic metabolism.
Effect of adding t10,c12-CLA to BMECs after simultaneous overexpression or inhibition of PI4K2A on milk fat synthesis marker genes. (A) Marker genes related to fatty acid de novo synthesis; (B) Marker genes related to long-chain fatty acid uptake in blood; (C) Marker genes related to fatty acid transport and desaturation; (D) Marker genes related to TAG synthesis; (E) Marker genes related to TAG catabolism; (F) Other transcriptional regulators related to milk fat synthesis. Blank is the control group; t10,c12-CLA is the group with the addition of t10,c12-CLA group; t10,c12-CLA+(Ad-PI4K2A) is the group with the addition of t10,c12-CLA while overexpressing PI4K2A; t10,c12-CLA+(siRNA-PI4K2A) is the group with the addition of t10,c12-CLA while suppressing PI4K2A; Different lowercase letters in the same comparison group indicate significant differences (p < 0.05). (mean ± SD, n = 3).
The influence of the PI4K2A gene on the inhibition of lipid synthesis by t10, C12-CLA in BMECs
The expression of marker genes for FA de novo synthesis (ACSS2) and TAG degradation (LIPE) was significantly down-regulated in the t10,c12-CLA+(Ad-PI4K2A) group (p < 0.05, Fig. 8A,E) compared with the t10,c12-CLA group. However, the expression of marker genes for uptake of LCFA from the blood (ACSL1 and CD36), FA transport function (FABP3) and TAG synthesis process (DGAT1 and GPAM) was significantly up-regulated (p < 0.05, Fig. 8B,C,D). The mRNA expression of most milk fat metabolism regulators in t10,c12-CLA+(Ad-PI4K2A) group was higher than that in the t10,c12-CLA group. Therefore, adding t10,c12-CLA and Ad-PI4K2A in BMECs could attenuate the inhibitory effect of t10,c12-CLA on milk fat. When compared to the t10,c12-CLA group, the expression of most milk fat metabolism marker genes in the t10,c12-CLA + siRNA-PI4K2A group showed no significant difference (Fig. 8). However, when compared to the control group, these genes were significantly down-regulated (p < 0.05). This suggests that the inhibitory effect of t10,c12-CLA on milk fat marker genes is independent of PI4K2A.
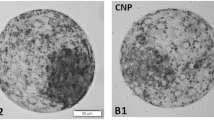
The inhibitory effect of t10,c12-CLA on milk fat in BMECs resulted in a significant decrease in TAG content (p < 0.05), whereas the total cholesterol content did not decrease significantly (p > 0.05). The TAG content in both the t10,c12-CLA+(Ad-PI4K2A) group and the t10,c12-CLA+(siRNA-PI4K2A) group was significantly higher than that in the t10,c12-CLA group (p < 0.05, Fig. 9). In addition, total cholesterol was less affected by t10,c12-CLA, and PI4K2A gene in BMECs. The Oil Red O staining showed that the secretion of milk fat formation in t10,c12-CLA group was significantly lower than that in control group. The secretion of lipid droplets in the t10,c12-CLA group was significantly lower compared to the t10,c12-CLA+(AD-PI4K2A) group, but it was not significantly different from the t10,c12-CLA+(siRNA-PI4K2A) group (Fig. 10A,B,C,D). Both the results of absorbance and the expression levels of lipid droplet secretion marker genes were consistent with those obtained from Oil Red O staining. (Fig. 10E,F, p < 0.05).
Effect of adding t10,c12-CLA with simultaneous overexpression or inhibition of PI4K2A on TAG and cholesterol levels. (A) The test results of TAG content; (B) The test results of cholesterol content.
Effect of adding t10,c12-CLA with simultaneous overexpression or inhibition of PI4K2A on lipid droplet secretion. (A–D) Blank, t10,c12-CLA, t10,c12-CLA+(Ad-PI4K2A), t10,c12-CLA+(siRNA-PI4K2A) groups, respectively; (E) OD values measured for lipid droplets dissolved in 100% isopropanol in different groups; (F) Expression of lipid droplet secretion-related marker genes.
Discussion
The effect of PI4K2A gene on lipid synthesis in BMECs
The synthesis of milk fat involves complex molecular regulatory networks, including FA de novo synthesis, uptake of LCFA from the blood, LCFA transport and desaturation, TAG synthesis, and lipid droplet secretion, with many key genes involved in each synthesis steps18. LCFA released into the blood is primarily activated by FABP3 and then undergo desaturation by SCD119. Our study revealed that PI4K2A had a significant positive regulatory effect on the FABP3 gene, suggesting that the PI4K2A gene could promote fatty acid transport during lipid synthesis in BMECs. A study of buffalo milk fat showed that PPARG positively regulates genes involved in FA de novo synthesis (FASN and ACSS2)20. In this study, we observed that overexpressing PI4K2A significantly up-regulated the expression of the PPARG gene, which promoted the FA de novo synthesis. ACOX1 is a crucial enzyme in fatty acid β-oxidation21. Overexpressing PI4K2A significantly up-regulated the expression of the ACOX1 gene, while interfering with PI4K2A had the opposite effect. This indicates that PI4K2A promotes the oxidation of fatty acid in milk fat metabolism. Although PI4K2A plays a role in promoting fatty acid oxidation, TAG synthesis and lipid droplet secretion by up-regulating the milk fat synthesis marker genes, we observed that PPARD expression was significantly down-regulated after overexpressing or interfering with PI4K2A gene. This finding appears to contradict the expectation that PI4K2A positive regulates lipid synthesis. Interestingly, when PPARD inhibited the expression of the FASN gene, it resulted in an increase in TAG content, suggesting that PPARD plays a crucial role in maintaining the dynamic balance of ruminant mammary gland cells22. It’s worth noting that knocking down PI4K2A results in numerous adipogenic marker genes being down-regulated, compared to overexpressing it23. These results suggest that inhibiting lipid synthesis by knocking down the PI4K2A gene may be more obvious than by overexpressing it.
TAG synthesis and lipid droplet secretion represent the final two stages of milk fat synthesis, involving many genes23. DGAT1 is the primary enzyme responsible for attaching the third long-chain fatty acid (LCFA) to 1-diacylglycerol (DAG) to produce TAG24. In our study, the overexpression of PI4K2A significantly increased the mRNA expression and protein of DGAT1, and increased TAG content and lipid droplet secretion in BMECs. Conversely, knocking down PI4K2A had the opposite result. Moreover, overexpressing PI4K2A also significantly reduced the expression of the ATGL25, ultimately leading to an increase in lipid synthesis. In summary, PI4K2A positively regulates lipid synthesis in BMECs and can also indirectly regulate this process via two transcription factors (PPARG and PPARD), which are known to be involved in regulating lipid metabolism.
The effects of PI4K2A gene on proliferation and apoptosis of BMECs
The mammary glands of cows undergo many changes in structure and function from birth to adulthood. Mammary cells proliferate, differentiate, apoptosis at different stages in response to different stimuli, ultimately remodeling glandular tissue structure26. Our study found that PI4K2A plays a role in regulating the balance between proliferation and apoptosis in fibroblasts27, aligning with previous research. CCND1 is a key regulator of cell cycle. It binds to cyclin-dependent kinase (CDK) and phosphorylates the tumor suppressor protein Rb, facilitating the transition of the cell cycle from the G1 phase to the S phase and promoting cell proliferation28. PI4K2A positively regulates the expression of CCND1 protein in BMECs, indicating that PI4K2A-induced proliferation of BMECs may occur during the transition from the G1 to the S phase of the cell cycle, thus ensuring proper DNA replication during S phase. The role of PI4K2A gene in promoting the proliferation of BMECs provides a theoretical basis for processes such as mammary wound healing, tissue regeneration and pathological tissue repair. A prominent characteristic of apoptosis is the activation of cysteine aspartic proteases (caspase). The activation of caspases leads to the degradation of cytoskeletal proteins during apoptosis. Therefore, phosphatidylserine is exposed on the cell surface and act as a marker of apoptotic cells. Macrophages recognize and phagocytose these apoptotic bodies through exposed phosphatidylserine and complete the clearance of apoptotic cells29. PI4K2A negatively regulates the expression of caspase-3 protein in BMECs, and then inhibits the apoptosis of BMECs. The disorder of apoptosis may be directly or indirectly related to the occurrence of many diseases. The inhibitory effect of PI4K2A on BMECs apoptosis has certain significance for the treatment of breast-related diseases, such as breast cancer30.
The effect of PI4K2A gene on the inhibition of lipid synthesis by t10, c12-CLA
Many studies have shown that t10,c12-CLA can reduce TAG content and promote fatty acid decomposition and oxidation31 in many species, and our study further validates these results. Notably, DGAT1 was significantly up-regulated and XDH was significantly down-regulated after adding t10,c12-CLA to GMECs, resulting in the accumulation of neutral lipids in GMECs. However, t10,c12-CLA did not reduce the lipid content of GMECs13, which contradicts the results of this study. The controversy may arise from species differences, the concentration of t10,c12-CLA in cells, and the upregulation of DGAT1 is more likely than the downregulation of XDH. Moreover, t10,c12-CLA can also up-regulate the expression of genes related to milk fat synthesis and promote lipid accumulation32 in human hepatocellular carcinoma HepG2 cells. The above studies reflect that the regulatory mechanism of t10,c12-CLA on lipid metabolism may vary across different species or tissues. In lipid synthesis, when LCFA are released from the blood into the extracellular environment, they are actively taken up by BMECs via CD36 and SLC27A, and subsequently activated by ACSL33. This study found that t10,c12-CLA could significantly reduce the expression of CD36 and ACSL1 in BMECs, which is consistent with the results of Zhang, et al.13 and Peterson, et al.34.
Furthermore, studies have found that t10,c12-CLA can increase the content of PI while reducing milk fat content16, and PI is a major substrate of PI4K2A17. In this study, we initially investigated the effect of PI4K2A on the inhibition of lipid synthesis in BMECs by t10,c12-CLA. The results showed that overexpressing PI4K2A could attenuate the inhibitory effect of t10,c12-CLA on lipid synthesis, while knocking down PI4K2A had no significant effect, suggesting that the inhibitory effect of t10,c12-CLA on lipid synthesis in BMECs may not depend on the PI4K2A gene35,36. Our research is still in its preliminary stage, and further investigation is needed to study the specific pathways or mechanisms.
Conclusion
In this study, we found that the PI4K2A gene positively regulates multiple crucial stages of milk fat metabolism in dairy cows, including de novo synthesis of FA, FA transport and desaturation, FA oxidation, TAG synthesis and lipid droplet secretion. The PI4K2A gene can promote the proliferation and inhibit apoptosis of BMECs, and has potential value in the treatment of breast-related diseases. This study further confirmed the inhibitory effect of t10,c12-CLA on lipid synthesis and lipid droplet secretion in primary BMECs. Additionally, our results indicate that overexpressing the PI4K2A gene can effectively reduce the inhibition of t10,c12-CLA on lipid synthesis. Next, we plan to conduct additional in vivo experiments to further investigate the precise mechanisms through which the PI4K2A gene regulates milk fat synthesis.
Materials and methods
Ethics statement
Animal experiments were conducted in accordance with the Regulations for the Administration of Affairs Concerning Experimental Animals (Ministry of Science and Technology, China, 2004). Cows in the experiment were electrocuted before collecting samples. All animal protocols have been approved by the Animal Ethics Committee of Ningxia University (permit number NXUC20200620). Our study is reported in full compliance with the ARRIVE guidelines (https://arriveguidelines.org/).
Experimental materials
The experimental subjects were dairy cows, and the experiment was based on Ningxia University’s “Guide to Laboratory Animals.” The Holstein cows in this experiment were from Ningxia Agriculture Reclamation Dairy Co.,Ltd, and all cows were physiologically healthy and without any udder disease. Three cows were selected in the middle to late stages of lactation (150–220 days) of second parity. The tissue samples of small intestine, liver, kidney, heart, ovary, uterus, and mammary gland were collected after the animals were electrocuted, and then stored in liquid nitrogen. In addition, Our laboratory has already completed the isolation, culture, and identification of BMECs in the early stage6.
Primer design and synthesis
PCR primers were designed using Primer Premier 5.0 software and gene sequences were obtained from NCBI (species is bovine). The designed primers were synthesized from Shanghai Bioengineering Technology Service Co.,Ltd and the primer sequences can be found in Table S3.
RNA extraction and cDNA synthesis
Total RNA was extracted from mammary gland and BMECs using RNA Extraction Kit (Tiangen Biochemical Technology Co., Ltd.) and RNAiso Plus (TaKaRa), respectively. The total RNA was analyzed using 1% agarose gel electrophoresis, and all three RNA bands (28 S, 18 S, and 5 S) were clearly visible (Figure S1). The spectrophotometric results showed that the OD 260/OD 280 ratio ranged from 2.0 to 2.1, indicating a purity suitable for further experiments, and the total RNA concentration was approximately 800 ng/µL (Table S4). After quality control, 1 µg of total RNA was used as a template synthesize the first strand of cDNA in a 20 µL reverse transcription reaction system, prepared according to the HiScript III 1st Strand cDNA Synthesis Kit (+ Gdna wiper).
The qRT-PCR and amplification procedure
The qRT-PCR solution system (20 µL): SYBR Green Pro Taq HS qPCR kit:10 µL, each of upstream and downstream primers: 0.4 µL, cDNA: 1 µL, ddH2O: 8.2 µL. The qRT-PCR amplification program: 95 °C for 30 s, 95 °C for 5 s, annealing (Table S3) for 30 s, 40 cycles from the second step, 60 ℃ for 5 s and 95 ℃ for 5 s.
Western blot
After washing the BMECs twice with pre-chilled PBS, they were digested with 0.25% trypsin (containing EDTA) and collected in 1.5 mL sterile centrifuge tubes. Then, 80 µL of lysis solution (1 mL RIPA: 25 µL of 100 mM PMSF) was added to each well to extract the total protein. A quarter volume of 5x protein loading buffer was added to the total protein and the mixture was denatured in a metal bath at 100°C for 5 minutes. Once the denatured mixture is removed, let it cool on ice before storing it in a -80°C freezer. Protein gel separation (20 µg loading volume, 5 µL protein marker loading volume) was performed using 4–20% precast gels with 11 spotting wells. After treatment with 160 V for 35 min, membrane transfer (25 V, 1.0 A, 15 minutes) was performed using Bio-Rad’ semi-dry transfer system. Then the membranes were blocked and incubated with antibodies. The PVDF membranes were exposed to ECL for 3 min, imaged in a gel imaging system, and finally protein abundance was calculated by Image J software37.
Lipid analysis
The TAG and total cholesterol were extracted according to the enzymatic assay kits (Beijing Pulley Gene Technology Co., Ltd.). Lipid droplet content was determined by Oil Red O staining kit (Beijing Solaibao Technology Co., Ltd). The main steps are as follows: (1) add 1.5 mL Oil Red O fixative to 6-well plate for 25 min, discard the fixative and wash twice with sterile water; (2) Add 1.5 mL 60% isopropanol to 6-well plate for 5 min, and add freshly prepared Oil Red O staining solution for 10 min. (3) Discard the staining solution and wash with sterile water until no excess dye remains. (4) Use hematoxylin to restain the cell nuclei for 1–2 min. (5) Add Oil Red O buffer for 1 min, then cover the cells with sterile water and observe under the microscope. (6) Discard the sterile water and let it dry for a while, then add appropriate amount of 100% isopropanol to dissolve the lipid droplets bound with Oil Red O. (7) Take 100 µL mixture of lipid droplets and 100% isopropanol from each well, and measured absorbance at 510 nm.
Packaging and vector construction of viruses
Hanheng biotechnology Co.,Ltd performed the packaging of overexpressed adenovirus and interfering sequence lentivirus of PI4K2A gene. The upstream primer sequence of PI4K2A gene is Adeasy-c-PI4K2A-K/X-F: CTGTGACCGGCGCCTACTCTGGTACCGCCACCATGGACG AGACGAG, and the downstream primer sequence is Adeasy-c-PI4K2A-K/X-R: TCATCGTCATCCTTGTAGTCCTCGAGCCACCAGGAGAAGAAGG GCT. The infection efficiency of overexpression group (Ad-PI4K2A) and control group (Ad-EGFP) were above 70% (Figure S2). Compared with Ad-EGFP group, the mRNA expression level of PI4K2A gene in Ad-PI4K2A group was significantly upregulated (p < 0.01). The expression level of the tagged protein 3*flag was also significantly higher in the Ad-PI4K2A group (p < 0.01). These findings indicated that the recombinant adenovirus Ad-PI4K2A was successfully constructed.
The sense strand (5’-3’) in the interfering RNA sequence of PI4K2A gene: GCUACAAAGAUGCAG ACUATT; Antisense strand (5’-3’): UAGUCUGCAUCUUUGUAGCTT; Target sequence: GCUACAAAG AUGCAGACUA. The infection efficiency for both the control group (NC) and interference group (siRNA-PI4K2A) were above 85% after BMECs were infected for 48 h (Figure S3). Compared with NC, the mRNA expression of PI4K2A gene was significantly downregulated (p < 0.01). The recombinant lentivirus siRNA-PI4K2A was successfully constructed when the interference efficiency reached 59.75%,
The optimal concentration of t10,c12-CLA to inhibit TAG synthesis in BMECs
The t10,c12-CLA was purchased from Matreya LLC in the United States. The t10,c12-CLA (25 mg) were dissolved in 100 µL ethanol and then add 900 µL DMEM-F12. When the cell density reaches about 70% of 6-well plate, different concentrations of t10,c12-CLA (85 µM, 170 µM, 225 µM, and 340 µM) were added to different wells 48 h (with 3 biological replicates for each concentration, and t10,c12-CLA was not added to the control). Then, the TAG content was detected (Figure S4). The TAG content decreased at first and then increased in BMECs with the increase of t10,c12-CLA concentration. The TAG content in BMECs was significantly lower than that in other treatment groups when the concentration of t10,c12-CLA was 170 µM (p < 0.05), and the TAG content suddenly increases (p < 0.05) when the concentration of t10,c12-CLA was 225 µM. Therefore, the optimal concentration of t10,c12-CLA was 170 µM, which inhibited the lipid synthesis of BMECs.
The detection of cell proliferation and apoptosis
The BMECs were inoculated in 6-well plates and transfected with overexpression or interfering viruses when the cells reached 50–70% of each plate. After 48 h, BMECs were washed twice with pre-chilled PBS. The proliferation and apoptosis of BMECs were detected by EdU Cell Proliferation Kit and Apoptosis Kit (Beyoncé Biotechnology), respectively. Images were captured and stored using a fluorescent inverted microscope (DMI4000B, Weztlar, Germany).
Statistical analysis
The treatment was replicated at least 3 times and the results were expressed as means ± standard error of the means (SEM). Data from qRT-PCR were analyzed using the 2^(-ΔΔCt) method and normalized to the respective control (Ad-PI4K2A or siRNA-PI4K2A)38. The proportion of genes in nuclear RNA (%) =\(\:{2}^{-\text{nuclear}\:\text{CT}\:\text{value}}\)/ (\(\:{2}^{-\text{cytoplasmic}\:\text{CT}\:\text{value}}\) +\(\:{\:2}^{-\text{nuclear}\:\text{CT}\:\text{value}}\)), and the proportion of genes in cytoplasmic RNA (%) =1 - proportion of genes in nuclear RNA (%). The one-way ANOVA function of the SAS (v.9.2) software was used to test the statistical difference.
Data availability
Previous studies on transcriptome sequencing and WGCNA analysis have been published, and the raw data and analysis results can be accessed through the published articles with the following DOIs respectively: 10.3389/fgene.2022.772115 and 10.1038/s41598-022-10435-1. All data generated or analyzed during this study are included in this published article and its supplementary information files.
References
Sun, C. Q., O’Connor, C. J. & Roberton, A. M. The antimicrobial properties of milk fat after partial hydrolysis by calf pregastric lipase. Chem. Biol. Interact 140, 185–198. https://doi.org/10.1016/s0009-2797(02)00016-9 (2002).
Zhou, C. et al. Comparative transcriptomic and proteomic analyses identify key genes associated with milk fat traits in Chinese holstein cows. Front. Genet. 10, 672. https://doi.org/10.3389/fgene.2019.00672 (2019).
Belury, M. A. Dietary conjugated linoleic acid in health: physiological effects and mechanisms of action. Annu. Rev. Nutr. 22, 505–531. https://doi.org/10.1146/annurev.nutr.22.021302.121842 (2002).
Bauman, D. E., Mather, I. H., Wall, R. J. & Lock, A. L. Major advances associated with the biosynthesis of milk. J. Dairy Sci. 89, 1235–1243. https://doi.org/10.3168/jds.S0022-0302(06)72192-0 (2006).
Xu, B. et al. Multiple roles for the non-coding RNA SRA in regulation of adipogenesis and insulin sensitivity. PLoS ONE 5, e14199. https://doi.org/10.1371/journal.pone.0014199 (2010).
Mu, T. et al. Screening and conjoint analysis of key lncRNAs for milk fat metabolism in dairy cows. Front. Genet. 13, 772115. https://doi.org/10.3389/fgene.2022.772115 (2022).
Mu, T. et al. Identifying key genes in milk fat metabolism by weighted gene co-expression network analysis. Sci. Rep. 12, 6836. https://doi.org/10.1038/s41598-022-10435-1 (2022).
Waugh, M. G. Phosphatidylinositol 4-kinases, phosphatidylinositol 4-phosphate and cancer. Cancer Lett. 325, 125–131. https://doi.org/10.1016/j.canlet.2012.06.009 (2012).
Kučka, M. et al. Calcium-prolactin secretion coupling in rat pituitary lactotrophs is controlled by PI4-kinase alpha. Front. Endocrinol. 12, 790441. https://doi.org/10.3389/fendo.2021.790441 (2021).
Palombo, V. et al. Genome-wide association study of milk fatty acid composition in Italian simmental and Italian holstein cows using single nucleotide polymorphism arrays. J. Dairy Sci. 101, 11004–11019. https://doi.org/10.3168/jds.2018-14413 (2018).
Wang, H. F., Liu, H. Y., Yang, W. R., Liu, J. X. & Yang, Z. B. Effects of foreign trans-10, cis-12 conjugated linoleic acid on SREBP-1 gene expression and protein synthesis in cultured bovine mammary epithelial cells. Scientia Agric. Sin. 44, 4892–4901. https://doi.org/10.3864/j.issn.0578-1752.2011.23.016 (2011).
Yin, F. Q. et al. Effects of different vegetable oils on fatty acids in ruminal fluid and blood plasma and milk CLA of lactating cows. Chin. J. Anim. Nutr. 20, 63–68. https://doi.org/10.3969/j.issn.1006-267X.2008.01.011 (2008).
Zhang, T. et al. Trans10, cis12 conjugated linoleic acid increases triacylglycerol accumulation in goat mammary epithelial cells in vitro. Anim. Sci. J. 89, 432–440. https://doi.org/10.1111/asj.12935 (2018).
Wang, H. et al. High-level exogenous trans10, cis12 conjugated linoleic acid plays an anti-lipogenesis role in bovine mammary epithelial cells. Anim. Sci. J. 85, 744–750. https://doi.org/10.1111/asj.12204 (2014).
Shi, H. et al. Trans-10,cis-12-conjugated linoleic acid affects expression of lipogenic genes in mammary glands of lactating dairy goats. J. Agric. Food Chem. 65, 9460–9467. https://doi.org/10.1021/acs.jafc.7b02377 (2017).
Yang, J. N. et al. Analysis of milk fat globule membrane glycerolphospholipids with low-fat depression induced by conjugated linoleic acid in dairy cows. Acta Vet. Zootech. Sin. 53, 791–801 (2022).
Berridge, M. J. & Irvine, R. F. Inositol trisphosphate, a novel second messenger in cellular signal transduction. Nature 312, 315–321. https://doi.org/10.1038/312315a0 (1984).
Osorio, J. S., Lohakare, J. & Bionaz, M. Biosynthesis of milk fat, protein, and lactose: roles of transcriptional and posttranscriptional regulation. Physiol. Genomics 48 231–56. https://doi.org/10.1152/physiolgenomics.00016.2015 (2016).
Bionaz, M. & Loor, J. J. Gene networks driving bovine milk fat synthesis during the lactation cycle. BMC Genom. 9, 366. https://doi.org/10.1186/1471-2164-9-366 (2008).
Zhou, F., Ouyang, Y. & Miao, Y. Peroxisome proliferator-activated receptor gamma regulates genes involved in milk fat synthesis in mammary epithelial cells of water buffalo. Anim. Sci. J. 92, e13537. https://doi.org/10.1111/asj.13537 (2021).
Wang, Z. J., Ye, P., Zhang, X. J. & Wang, L. H. Effects of aging on the expression of Acy-l CoA oxidase in rat liver. Chin. J. Arterioscler. 13, 10–12. https://doi.org/10.3969/j.issn.1007-3949.2005.01.003 (2005).
Shi, H. B., Zhang, C. H., Zhao, W., Luo, J. & Loor, J. J. Peroxisome proliferator-activated receptor delta facilitates lipid secretion and catabolism of fatty acids in dairy goat mammary epithelial cells. J. Dairy Sci. 100, 797–806. https://doi.org/10.3168/jds.2016-11647 (2017).
Li, J. et al. Adipose triglyceride lipase regulates lipid metabolism in dairy goat mammary epithelial cells. Gene 554, 125–130. https://doi.org/10.1016/j.gene.2014.10.020 (2015).
Mu, T. et al. Regulation of key genes for milk fat synthesis in ruminants. Front. Nutr. 8, 765147. https://doi.org/10.3389/fnut.2021.765147 (2021).
Deiuliis, J. et al. Bovine adipose triglyceride lipase is not altered and adipocyte fatty acid-binding protein is increased by dietary flaxseed. Lipids 45, 963–973. https://doi.org/10.1007/s11745-010-3476-2 (2010).
Inman, J. L., Robertson, C., Mott, J. D. & Bissell, M. J. Mammary gland development: cell fate specification, stem cells and the microenvironment. Development 142, 1028–1042. https://doi.org/10.1242/dev.087643 (2015).
Mohamed, M. et al. Novel defect in phosphatidylinositol 4-kinase type 2-alpha (PI4K2A) at the membrane-enzyme interface is associated with metabolic cutis laxa. J. Inherit. Metab. Dis. 43, 1382–1391. https://doi.org/10.1002/jimd.12255 (2020).
Long, J., Ou, C., Xia, H., Zhu, Y. & Liu, D. MiR-503 inhibited cell proliferation of human breast cancer cells by suppressing CCND1 expression. Tumour Biol. 36 8697 – 702. https://doi.org/10.1007/s13277-015-3623-8 (2015).
Schlegel, R. A. & Williamson, P. Phosphatidylserine, a death knell. Cell. Death Differ. 8, 551–563. https://doi.org/10.1038/sj.cdd.4400817 (2001).
Li, Q. et al. Donkey milk inhibits triple-negative breast tumor progression and is associated with increased cleaved-caspase-3 expression. Food Funct. 11, 3053–3065. https://doi.org/10.1039/c9fo02934f (2020).
Baldin, M. et al. CLA induced milk fat depression reduced dry matter intake and improved energy balance in dairy goats. Small Rumin. Res. 116, 44–50. https://doi.org/10.1016/j.smallrumres.2013.10.001 (2014).
Go, G. W. et al. T10, c12 conjugated linoleic acid upregulates hepatic de novo lipogenesis and triglyceride synthesis via mTOR pathway activation. J. Microbiol. Biotechnol. 23, 1569–1576. https://doi.org/10.4014/jmb.1308.08008 (2013).
Doege, H. & Stahl, A. Protein-mediated fatty acid uptake: novel insights from in vivo models. Physiology 21, 259–268. https://doi.org/10.1152/physiol.00014.2006 (2006).
Peterson, D. G., Matitashvili, E. A. & Bauman, D. E. The inhibitory effect of trans-10, cis-12 CLA on lipid synthesis in bovine mammary epithelial cells involves reduced proteolytic activation of the transcription factor SREBP-1. J. Nutr. 134, 2523–2527. https://doi.org/10.1093/jn/134.10.2523 (2004).
Jacometo, C. B. et al. Linoleic and α-linolenic fatty acid consumption over three generations exert cumulative regulation of hepatic expression of genes related to lipid metabolism. Genes Nutr. 9, 405. https://doi.org/10.1007/s12263-014-0405-7 (2014).
Khan, M. J. & Sun, P. Changes of metabolism and gene network expression in subcutaneous adipose tissue of perinatal dairy cows and their genetic quality differences. China Anim. Husb. Vet. Med. 40, 1 (2013).
Rueden, C. T. et al. ImageJ2: ImageJ for the next generation of scientific image data. BMC Bioinform. 18, 529. https://doi.org/10.1186/s12859-017-1934-z (2017).
Zhang, W. et al. The LXRB-SREBP1 network regulates lipogenic homeostasis by controlling the synthesis of polyunsaturated fatty acids in goat mammary epithelial cells. J. Anim. Sci. Biotechnol. 13, 120. https://doi.org/10.1186/s40104-022-00774-4 (2022).
Acknowledgements
We are grateful to Ningxia Agricultural Reclamation Helan Mountain Dairy Industry Corporation for facilitating the data collection and providing access to production records.
Funding
This study was supported by the special breeding project of high-quality and high yield dairy cows in the Ningxia Autonomous region (Grant No: 2019NYYZ05) and the doctoral research start-up project of Yan’an University (YAU202407517, YAU202411766).
Author information
Authors and Affiliations
Contributions
T. M. and H. H: Data processing and article writing. X. F.: Article modification and visualization. J. Z. and C. W.: Sample collection and verified by qRT-PCR. Z. S.: Article grammar modification. Y. G.: Conceptual analysis, writing-review and editing. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Institutional review board statement
The Animal Ethics Committees of Ningxia University approved the experimental design and animal sample collection for the present study (permit number NXUC20200620). And animal experiments were conducted strictly followed the guidelines of the Regulations for the Administration of Affairs Concerning Experimental Animals (Ministry of Science and Technology, China, 2004).
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Mu, T., Hu, H., Feng, X. et al. The PI4K2A gene positively regulates lipid synthesis in bovine mammary epithelial cells and attenuates the inhibitory effect of t10,c12-CLA on lipid synthesis. Sci Rep 15, 3456 (2025). https://doi.org/10.1038/s41598-025-87089-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-87089-2