Abstract
Fluorescence recovery after photobleaching (FRAP) allows to study actin-turnover in dendritic spines by providing recovery trajectories over time within a nested data structure (i.e. spine/neuron/culture). Statistical approaches to FRAP usually consider one-phase association models to estimate recovery-curve-specific parameters and test statistical hypotheses on curve parameters either at the spine or neuron level, ignoring the nested data structure. However, this approach leads to pseudoreplication concerns. We propose a nonlinear mixed-effects model to integrate the one-phase association model estimate with the nested data structure of FRAP experiments; this also allows us to model heteroscedasticity and time dependence in the data. We used this approach to evaluate the effect of the downregulation of the actin-binding protein CAP2 on actin dynamics. Our model allows the additional modelling of the variance function across experimental conditions, which may represent a novel parameter of interest in FRAP experiments. Indeed, the detected differential effect of the experimental condition on the variance component captures the increased instability of time-specific observations around the spine-specific trajectory for the CAP2-downregulated spines compared to the control spines. We hypothesise that this parameter reflects the increased instability of the actin cytoskeleton in dendritic spines upon CAP2 downregulation. We developed an R-based Shiny application, termed FRApp, to fit the statistical models introduced without requiring programming expertise.
Similar content being viewed by others
Introduction
Fluorescence recovery after photobleaching (FRAP) is a live-cell functional imaging technique that allows the investigation of protein dynamic behaviour at the single-cell level by exploiting the properties of fluorescent proteins as observed by confocal microscopy systems1,2,3.
FRAP is considered a useful technique to investigate the dynamics of the actin cytoskeleton in dendritic spines4,5, which are small actin-rich protrusions from the dendritic shafts of neurons. Spines contain all the postsynaptic machinery required for the transmission of synaptic signalling, and their morphology, which is tightly coupled to their function, is finely regulated by the dynamics of the actin cytoskeleton6.
The actin cytoskeleton is composed of filamentous actin (F-actin), which is a polymer made of globular (G)-actin monomers. Actin filaments are polar structures with one end (barbed end) growing more rapidly than the other (pointed end). Constant removal of the ADP-G-actin subunits from pointed ends and addition of the ATP-actin to the barbed ends is called actin ‘treadmilling’, which is regulated by actin-binding proteins7. In dendritic spines, the addition of G-actin to the barbed ends pushes the plasma membrane and induces changes in spine shape.
To study actin dynamics in spines via FRAP, primary neuronal cultures are transfected with the actin-binding protein probe LifeAct-GFP to fluorescently label actin filaments. To conduct a FRAP experiment, a single spine is briefly illuminated with a strong laser to quench the GFP fluorescence. The mean fluorescence intensity of the spine area is measured from a time-lapse series of images acquired immediately after photobleaching.
After spine photobleaching, actin monomers normally diffuse into the spine and are incorporated into the barbed end of bleached filaments, predominantly in the juxtamembrane region of the spine head. Concomitantly, bleached actin molecules are removed from filament pointed ends (closer to the spine ‘core’) and exchanged out of the spine, which recovers its fluorescence8. Therefore, the rate of fluorescence recovery reflects the rate of actin treadmilling. FRAP experiments have demonstrated that spine F-actin can be divided into two pools based on their turnover rate: the dynamic mobile pool, which has a faster turnover rate, and the stable pool, which has a slower turnover rate9,10. Quantitative analysis of FRAP data via nonlinear (i.e. where parameters of interest occur nonlinearly in the expression of the expected response) regression models allows the estimation of parameters related to the shape of the recovery curves (i.e. curve parameters), including the mobile fraction of molecules (i.e. the fraction of molecules that are mobile and free to diffuse) and the half maximal recovery time of the mobile fraction (i.e. the time when the intensity equals one-half of the maximal intensity). Therefore, the data can provide insight into the mobility of the fluorescent actin and the percentage of dynamic and stable actin pools in the spine5,11.
FRAP experiments conducted on dendritic spines are associated with additional challenges, including large variation between measurements, rapid dendritic spine dynamics and changes in fluorescence intensity, and spine enlargement following photo-bleaching5. These features may increase the instability of spine-specific recovery curves.
With a few exceptions12,13, statistical analysis of FRAP data overlooks the nested hierarchical structure of FRAP experiments, where multiple spines belong to the same neuron, and different neurons are derived from one of the cultures. To evaluate differences between experimental groups, standard hypothesis testing is conducted either at the spine or at the neuron level after the one-phase modelling approach (pre-specified based on existing literature4,5) has been used to estimate curve parameters at the spine level. The presence of a nested structure likely leads to pseudoreplication, which occurs when observations are not statistically independent but are treated as if they are14.
Despite being conceptually straightforward and amenable to standard hypothesis testing, the use of summary statistics, such as cluster-based means or slopes, to account for the nested structure, implies a loss of information compared with using individual observations. As a generalisation of repeated-measures analysis of variance (ANOVA)15, mixed-effects or multilevel regression models correctly handle dependence in nested designs and perform the analysis in one step, while retaining information on the precision of the estimates. This results in increased power to detect the experimental effect of interest compared with the use of summary statistics16. Unfortunately, recent surveys16,17 have confirmed that only 50% of the studies presenting nested data that are published in prestigious neuroscience journals accounted for data dependence in any meaningful way.
In the current study, we explore the use of a nonlinear mixed-effects model to evaluate the effects of the downregulation of an actin-binding protein critical in spine morphology remodelling within the nested data structure of FRAP experiments. In this frame, we decided to focus on Cyclase-Associated Protein 2 (CAP2), which has been shown to be a regulator of actin treadmilling and a key player in spine shape remodelling18.
Results
We performed FRAP experiments to evaluate the effect of CAP2 downregulation on actin turnover in spines. Figure 1 shows the scheme of the FRAP experiment (Fig. 1a) and the corresponding raw data (neurons 1–4) grouped following their hierarchical structure (Fig. 1b): the spine level was nested into the neuron level, which was nested into the culture level. Possible sources of dependence were related to this data structure. Each primary neuronal culture was prepared from rat hippocampi dissected from littermates. Therefore, neurons from the same culture tended to be more similar to one another than neurons from different cultures, due to the shared genetics, common environment, and similar developmental conditions characteristic of littermates. Dendritic spines are subcellular compartments of neuronal cells. Therefore, spines on the same neuron tend to be more similar to each other than to spines on different neurons. Similarities between dendritic spines on the same neuron can be attributed to shared genetic instructions and localized biochemical signaling. In each culture, neurons were transfected with LifeAct-GFP and either CAP2-shRNA to downregulate CAP2 or a scrambled control sequence (SCR), as control condition. Hence, neurons and, consequently, their dendritic spines were assigned to one of two experimental conditions (Fig. 1a, right side). LifeAct is a peptide fused to GFP with a high affinity for actin filaments, although it also exhibits some binding to G-actin. While LifeAct may produce slightly different FRAP results compared to fluorescently labeled actin-commonly used in FRAP studies of dendritic spines, it remains a valuable tool. Its efficacy has been demonstrated in live neuron experiments, where it enables real-time visualization and analysis of F-actin dynamics19,20.
In addition, FRAP experiments provided 65 measures of normalized fluorescence intensity equally spaced in time over a period of 100 seconds, thereby revealing time dependence in the data structure (Fig. 1b, Movies S1–S2). Spine-specific normalized fluorescence intensity trajectories generally showed nondecreasing patterns21 over time across plots in Fig. 1b. Most of them were compatible with the representation of the pseudo-first order association kinetics of the interaction between a ligand and its receptor, or a substrate and an enzyme provided by the one-phase association curves5,22. The one-phase association curves are known as asymptotic exponential curves in the statistical literature on growth curve models, as they describe a limited growth, with Y approaching a horizontal asymptote as X tends to infinity23. In detail, the asymptotic exponential curves are characterised by three parameters describing the starting point, the steepness, and the asymptote of the curve. The intercept (R0) is the normalized fluorescence intensity immediately after photobleaching (i.e. at time 0), the log rate constant (lrc) measures how fast the normalized fluorescence intensity recovers, and the horizontal asymptote (Asym) is the plateau of the recovered normalized fluorescence intensity over time. The lrc is expressed in the reciprocal of the x-axis time units and is related to the recovery half-time as \(t_{1/2} =\frac{\ln (2)}{rc}\), where \(rc = \exp (lrc)\) is the rate constant and ln indicates the natural logarithm (i.e. base e). Such parameters provide insights into actin turnover in spines. From a fitted curve, the mobile fraction of actin in spines is obtained as the difference between the Asym and R0 values, whereas lrc describes the kinetics of actin turnover5,24 (Fig. S1).
The raw data highlighted that differences were present among spines of the same neuron (e.g. culture 4/experimental condition SCR/neuron 1), among neurons of the same culture (e.g. culture 4/experimental condition SCR), and among cultures (e.g. cultures 5 and 6/experimental condition SCR), indicating the presence of potential variability to be modelled at each hierarchical level by mixed-effects models (Fig. 1 b). Mixed-effects models combine fixed effects describing the global population mean and random effects accounting for variation among units at a specific level23. Hierarchical data structures are easily handled, as random effects parameters can be specified for each level of the hierarchy25. As an example, a random effect on a given parameter at the culture level estimates the variability of that parameter among cultures; similarly, a random effect at the neuron level estimates the variability among neurons within the same culture. Therefore, adding the fixed-effects estimates to the random-effects ones up to a specific level allows us to obtain a fitted curve for that level.
The nonlinear mixed-effects model allowed the integration of the one-phase association model estimates for spine-specific curves within the typical nested data structure of FRAP experiments. While an asymptotic exponential curve is assumed to model normalized fluorescence intensity over time accounting for the hierarchical structure and the experimental condition, the within-group errors are additionally allowed to be correlated and/or have unequal variances. Model selection was therefore performed following the approach suggested by Pinheiro and Bates23, where the final model is constructed through the following key steps: 1) defining the random-effects structure, 2) specifying the variance function to account for potential heteroscedasticity, and 3) establishing the temporal dependence structure. At each step, the significance of each addition/deletion or refinement was evaluated using the Akaike Information Criterion (AIC), the Bayesian Information Criterion (BIC), and the likelihood ratio test (LRT) (when appropriate), along with visual inspection of diagnostic plots. Details are provided in the Supplementary Information (SI).
The selected model included all three available random effects for R0 at the spine, neuron, and culture levels, as well as two random effects for Asym and lrc, both at the spine and culture levels (Table 1). Random effects for R0 at the three hierarchical levels suggested that variability was present in the starting point across spines, neurons, and cultures, whereas the random effects on the curve plateau, Asym, and steepness, lrc, indicated variability between different spines and cultures. No statistically significant interactions were detected between the three fixed-effects parameters and the experimental conditions (CAP2-shRNA vs. SCR); similarly, interactions between random effects (at neuron and spine levels) and the experimental conditions were excluded in the absence of model improvement according to information criteria and LRTs (Table S1). Overall, the estimated fixed- and random-effect structure suggests that we did not identify any significant effects of CAP2 downregulation on curve parameters, especially the recovery rate of actin (i.e. lrc) and the actin mobile fraction in spines (\(Asym - R0\)).
An exponential variance function handled residual heteroscedasticity; indeed, residuals (i.e. differences between single observations and the corresponding fitted values obtained from the statistical model) exhibited a non-proportional increasing variance at increasing normalized fluorescence intensities (Fig. S3a). Compared with the SCR spines, CAP2-shRNA spines exhibited higher instability in normalized fluorescence intensity recovery. This led us to extend the model by adding an extra parameter to the variance function to account for a different within-group residual variance in the experimental conditions (CAP2-shRNA and SCR). Model comparison (Table S1) and the analysis of residuals (Fig. S3b) confirmed this experimental observation. The corresponding estimates were: \(\hat{\delta }_{g=\text {SH}} = 1.159\) vs. \(\hat{\delta }_{g=\text {SCR}}=1.065\) (Table 1), thus suggesting that the residual variance was higher when CAP2 downregulation occurred.
An autoregressive model of order one was then added to account for the temporal dependence of the sampled values, which is due to the recording of normalized fluorescence intensity at successive time points (Table S1). This final step led to the nonlinear mixed-effects model for which parameter estimates and 95% approximate confidence intervals are reported in Table 1 and the analysis of residuals is shown in Fig. S4.
The model provided fixed and random effects at the distinct levels of the hierarchical structure. From the estimates, we built the estimated level-specific curves by adding level-specific random effects, if present, to the fixed effects. In the absence of significant interactions between fixed effects and experimental condition (CAP2-shRNA and SCR), estimates of fixed effects-which are shared across all spines-allowed to draw the global population curve (Fig. 2a, dashed line). This was characterised by the following parameter estimates: \(\widehat{R0} = 0.252\) [Standard Error (SE) \(= 0.041\)], \(\widehat{lrc} = -2.713\) (SE \(= 0.135\)), and \(\widehat{Asym} = 0.956\) (SE \(= 0.036\)) (all p-values \(< 0.001\), from t-tests assessing the null hypothesis that the parameter is equal to 0). It follows that the estimated recovery half-time is \(t_{1/2} = ln(2)/\widehat{rc}=10.449\) seconds, and the estimated mobile fraction is equal to \(\widehat{Asym}-\widehat{R0}=0.705\). The lack of significance for the interaction terms between any of the fixed parameters and the experimental condition implied that the fitted global population curve was the same for the two experimental conditions. Indeed, each characteristic of the curve (starting point, steepness, and plateau) did not depend on the specific experimental condition applied. The six culture-specific curves (Fig. 2a, solid curves) were obtained by summing the fixed effects with the corresponding culture-level random effects shown in Fig. 2b. In the impossibility of properly defining an interaction term between random effects at culture level and experimental condition, the lack of significance of the interaction terms between fixed effects and the experimental condition for any of the three parameters led to draw the same culture-specific curve for both experimental conditions. In detail, random effects at the culture level on the lrc and R0 parameters revealed a common pattern for cultures 1, 2, 4, and 5, which started at a higher point and grew slower than the global population curve (Fig. 2b, \(\rho _c\) and \(\lambda _c\)). A different pattern was observed for the Asym parameter, where cultures 1, 5, and 6 reached a lower plateau than cultures 2, 3, and 4 (Fig. 2b, \(\alpha _c\)). These behaviors were depicted in more detail in Fig. 3, where spine-specific normalized fluorescence intensities over time were grouped by culture and experimental condition and compared with the only global population curve and the six culture-specific curves. Due to the final model selected and the adopted experimental design, the fitted curves were drawn regardless of the experimental condition.
Random effects at culture and neuron levels, in addition to fixed effects, allowed to draw the neuron-specific curves (Fig. 4; Fig. S5). Similarly, fixed and random effects at the three levels allowed us to plot spine-specific curves represented in Fig. 4 (Fig. S6). Selected panels in Fig. 4 (e.g. culture 2/experimental condition SH/neuron 2; culture 4/experimental condition SCR/neuron 1) allow us to highlight the major strengths of mixed-effects models, including the management of nested data structures and the advantages of shrinkage effects25. Indeed, in panel culture 4/experimental condition SCR/neuron 1, we observed that: 1) spine-specific curves adapted to the different increasing patterns present in the data (i.e. dots in the panel) and 2) accounting for the hierarchical structure allowed us to shrink the spine-specific curves towards the neuron-specific curve, which lay in the middle of the spine-specific curves. The latter advantage of mixed-effects models was further highlighted in panel culture 2/experimental condition SH/neuron 2, where the observed patterns exhibited a strong increasing trend; however, spine-specific fitted curves, although increasing, were moderate towards the global trend due to the shrinkage effect. In addition, neuron-specific curves generally reflected the culture-specific curves, except for the starting point, due to the lack of any random effect for the lrc and Asym parameters at the neuron level (Fig. 4).
Figure 5 shows 100 simulated samples (of the original 132 spines) generated from the final selected model, which accounted for the random effects and within-group error structure. Spines were grouped by experimental condition, CAP2-shRNA (right panel) and SCR (left panel). The differential effect of the experimental condition on the variance components reflected the increased instability of time-specific observations around the spine-specific trajectory for the CAP2-shRNA spines, compared to the SCR spines. This greater instability at each observation level also resulted in higher variability of the simulated spines under the CAP2-shRNA condition. This variability is particularly evident at the plateau, where the \(5{\rm th}\) and \(95{\rm th}\) quantiles were equal to 0.605 and 1.211 for the SCR condition, while they were 0.510 and 1.501 for the CAP2-shRNA condition. Additionally, the mean curve at plateau was higher under the CAP2-shRNA condition compared to the SCR condition, with a difference of 0.081. This difference was largely explained by the random effects for Asym at the spine level, as the difference between the conditioned means of these effects was given by: \(E({\hat{\alpha }}_{cgns}^{\left( 3\right) }| g=\text {SH}) - E({\hat{\alpha }}_{cgns}^{\left( 3\right) }| g=\text {SCR})= 0.073\).
To validate the selected model, Fig. 6 compares observed and simulated spines from a sample of 50 randomly selected simulated datasets, based on the final model chosen. The simulated spines generally aligned well with the observed data, supporting the validity of the selected model. This alignment is attributed to the inclusion of random effects at the spine level, which provides flexibility in the model. However, some observed spines did not fully adhere to the exponential functional form selected (leading to the heavy tails in the residual plots (Fig. S4)). In instances where such deviations occurred, the corresponding simulated spines tended to be regularized by the shrinkage effect inherent in the mixed-effects model, which tends to pull extreme trajectories toward the global population curve. In addition, when excluding ten randomly selected spines at a time and refitting the final model on the restricted dataset, the point estimates and corresponding approximate 95% confidence intervals were not materially different across the three scenarios under evaluation (Table S2). Moreover, these results were comparable with those from the main analysis (Table 1). Finally, when we predicted the excluded spines based on the restricted dataset, most predicted curves aligned with the excluded observed spines (Fig. S9). These findings globally supported the validity of the final model selected.
Figure 7 summarizes how our R-based Shiny application, FRApp, handles uploading and analysis of the data described in this paper according to the final statistical model selected. The application accommodates the specification of models including up to three hierarchical levels, each with corresponding level-specific random effects. Moreover, it offers the opportunity to modify: (1) the variance function, to account for the heteroscedasticity and (2) the autocorrelation order of the model, to account for the temporal dependence. In detail, after a dataset is uploaded, the FRApp application allows to fit several statistical models on it, by: (1) adding the response variable (i.e. y), (2) adding the explanatory variable (i.e. time), (3) adding or deleting (level-specific) random effects, (4) adding or deleting possible interaction terms involving the experimental condition and fixed effects, random effects at any hierarchical level, or the variance coefficient, and (5) adding or changing the variance function and the autocorrelation order. Models are saved to perform model comparison, and their results can be exported to carry out additional analyses. A brief report is also available for each model that provides summary results and residual plots. To showcase the functionality and usability of FRApp, we used the dataset presented in this paper. A vignette integrated in FRApp provides details on how to install and launch FRApp, prepare and import data, specify and fit suitable statistical models, and compare them to choose the best one.
Discussion
The in-depth analysis of dynamic data in living cells is a crucial aspect of modern biology. Here, we challenged the use of nonlinear mixed-effects models to the analysis of FRAP data in dendritic spines. Nonlinear mixed-effects models leads to the following advantages: (1) use of random effects to model variability at the different levels of the hierarchical structure, typical of the experimental design of FRAP in dendritic spines, (2) shrinkage of spine-specific curves towards the global population mean, which allows the analysis of curves whose trajectories suffer from spine enlargement phenomena, typical of FRAP experiments for dendritic spines, (3) assessment of the presence of potential treatment-specific effects on fixed effects, random effects, and on the variance function that are meant to account for residual heteroscedasticity, and (4) modelling of the potential dependence of observations over time with autoregressive structures.
FRAP was originally developed over 45 years ago26, and it has been extensively used to study the kinetics of labelled molecules in cells. In neuroscience research, FRAP sparked interest in the investigation of actin dynamics in a specific subcellular compartment, that is, dendritic spines. However, FRAP in spines can have several drawbacks, such as the large variation between measurements, rapid dendritic spine dynamics and changes in fluorescence intensity, and spine enlargement following photo-bleaching5. Furthermore, the analysis of FRAP recovery curves remains an active area of research that strives to take advantage of more comprehensive and reliable analysis strategies. Currently, statistical analyses of FRAP data are performed by fitting spine-specific recovery curves with a single exponential equation and by deriving biologically relevant parameters to study the kinetics of actin recovery (i.e. recovery half-times) and the composition of the actin cytoskeleton in spines (i.e. plateau). Hypothesis testing is used to compare curve parameters across experimental conditions by considering as single experimental units either: (1) spines (e.g. Mikhaylova et al.27 and George et al.28), or (2) neurons, after averaging over (curve) parameters of spines from the same neuron (e.g. Koskinen et al.4). However, both approaches lead to pseudoreplication because the standard assumption that experimental units are independent from one another is no longer valid. Spines are biological entities that belong to neurons, and neurons are plated in different culture preparations. When spines are considered as experimental units, their belonging to the same neuron-leading to dependence-is ignored; when neurons are considered as experimental units, their belonging to the same culture-which still leads to dependence-is also ignored. If standard statistical tests are applied with an invalid independence assumption, the well-known consequences are underestimated standard errors, deflated p-values, and inflated type I error rates (i.e. number of false positives)17.
In recent years, concerns about the low standard of statistical analysis in preclinical studies have arisen and have promoted discussion about the use of more sophisticated statistical methods15,29. To manage such a multilevel data structure, we used a mixed-effects model, which consists of both fixed and random effects, to account for different sources of variability in the data. Compared with other cellular systems, FRAP in spines is vulnerable to technical and biological issues that increase variability at different levels5. The bleaching laser pulse may increase spine size, leading to overshooting of fluorescence from pre-bleaching levels and high fluctuations in the post-bleaching fluorescence. These well-known issues have traditionally led to discarding curves exceeding one at the final recording stages because negative values in the stable component of actin cytoskeleton are biologically difficult to interpret and cannot be safely included in hypothesis testing procedures comparing different experimental conditions. To include all curves derived from well-conducted experiments in the analysis, a more conservative, geometric approach (i.e. using a linear approximation of the stable component) was developed to downgrade highly variable curves5. By taking into account the different hierarchical levels of the experiment, mixed-effects models provide a statistical approach to compensate for the expected additional variability of spine-level curves with the naturally occurring shrinkage of any spine-specific curve towards the global mean. The combined role of random effects and shrinkage helps mitigate biases that could arise from the subjective exclusion of curves exceeding one at the plateau. In our application, even though spines almost reaching three at their final recording stages were included, the final global mean estimate of \(\widehat{Asym}\) was equal to 0.956, with the \(95\%\) approximate confidence interval given by \(0.885-1.027\).
Analysis of the random effects revealed that their variability is different at the specific levels taken into consideration. The culture level is characterised by random effects for all the three parameters of the recovery curves (i.e. R0, lrc, and Asym), suggesting that cell cultures prepared from different animals or in different experimental settings and time periods produce recovery curves with markedly different shapes. Therefore, considering neurons/spines from distinct cultures as independent units can remarkably affect the statistical analysis. Furthermore, a random effect was selected at the neuron level for the starting point (i.e. R0) of the curve. We can hypothesise that such variability among neuronal cells depends on the expression levels of LifeAct-GFP in different neurons (excluding the variability related to CAP2 treatment). At the spine level, all the random effects were selected for the R0, lrc, and Asym parameters. Even though spines belong to the same neuron, they can produce different recovery curve shapes, likely due to the different abundances of actin-binding proteins. To summarise, in the current application, mixed-effects models were useful to capture variability from both technical and biological sources. When not accounted for by mixed-effects models, this variability is likely to lead to biased results when comparing experimental conditions using standard statistical approaches for FRAP data.
Independently from the hierarchical structure and from the treatment effect, it is reasonable to expect that FRAP experiments in spines exhibit fluctuations of fluorescence recovery observations that are not constant, but rather increase over time5. To model this phenomenon, the regression models are adapted to include a variance function that models the effect of time using a pre-specified structure (e.g. linear, power, or exponential function); model selection via information criteria is also amenable to identify the best structure. In addition, a differential effect of treatment over the variance function can be assessed and incorporated into the final model by adding an interaction term between the variance function and the experimental condition. When present, this different effect on spine-specific-curve variability over time might be related to the intrinsic instability of actin dynamics. Similarly, the acquisition of information on normalized fluorescence intensity at successive time points justifies the inclusion of an autoregressive structure into the final model, to take into account the temporal dependence of the sampled values.
To our knowledge, a Bayesian approach to nonlinear mixed-effects models has been previously used once in the analysis of FRAP experiments obtained from multiple cells, to study the cell cycle-dependent binding behaviour30. While the Bayesian approach is advantageous in that it naturally turns the attention towards prediction31, in the current application we preferred to adopt the more standard frequentist approach. Although a Bayesian model similar to the selected one could be easily estimated using the R package brms, our model is not available in its current formulation in this package; the corresponding code has to be implemented by hand. Given the large number of data considered, the impact of vague prior distributions on final results is likely negligible. In addition to an easier implementation, the frequentist approach asks for shorter computation times, while still providing reliable results. This might also be crucial when using our FRApp application. The derived parameter estimates may anyways be used for prior specification in a next application of Bayesian mixed-effects models to the current data.
In addition to the improvement in the statistical analysis of FRAP curves, our study may provide relevant information about the biological role of CAP2 in the regulation of spine actin cytoskeleton. The downregulation of CAP2 does not significantly affect the recovery rate of the global population curve or the mobile fraction of actin in the spines, as our model did not reveal significant differences in the fixed effects for Asym, lrc, or R0 between CAP2 shRNA and SCR control conditions. The use of nonlinear mixed-effects models enabled, however, the identification of a novel parameter that may offer additional insights into actin cytoskeleton dynamics, complementing the widely recognized parameters derived from one-phase association curves. Indeed, the additional modelling of the variance function across treatments allowed us to identify a greater variability of the time-specific observations around the spine-specific trajectory in the CAP2 shRNA experimental condition. This pointwise instability is supported by approximate confidence intervals for the treatment-specific variance parameter estimates, which do not overlap. Even though we cannot exclude a contribution to this effect of the variability in the efficiency of the CAP2 downregulation, we hypothesise that the higher variance identified for CAP2 shRNA spines at the observation level can mirror a higher instability of actin cytoskeleton in CAP2-downregulated spines. Indeed, data from FRAP analysis indicating cytoskeletal instability may have biological implications, potentially contributing to the alterations in spine morphology observed in CAP2 shRNA-transfected neurons18. This hypothesis is consistent with previous findings indicating that CAP2 plays a crucial role in the control of spine morphology, ultimately affecting synaptic function18. Indeed, CAP2 downregulation has been shown to disrupt a cofilin-mediated mechanism essential for spine shape remodeling and the potentiation of synaptic transmission18. Furthermore, a decrease in CAP2 protein level has been reported in neurodegenerative disorders, such as Alzheimer’s disease18, and we can hypothesise that the instability of the actin cytoskeleton under such pathological conditions contributes to the defects in synaptic plasticity phenomena described in Alzheimer’s disease32,33. Further studies are required to better understand how CAP2 downregulation influences the mechanisms that govern actin cytoskeleton stability.
Materials and methods
Ethical approvals
Sprague-Dawley pregnant rats were purchased from Charles River Laboratories (Calco, Italy). Embryos were used at embryonic day 19 to prepare hippocampal neuronal cultures. All procedures performed in studies involving animals were in accordance with the European Guidelines for Animal Experiments (Directive 2010/63/EU), the Italian Animal Welfare Law and the ARRIVE guidelines34. The study protocol was approved by the Institutional Animal Care and Use Committee of University of Milan and by the Italian Ministry of Health (Permission number #5247B.N.YCK/2018). Animals were maintained on a 12-h light/dark cycle in a temperature-controlled room (22°C) in cages with free access to food and water. Housing in the animal facility is performed under the control of veterinarians with the assistance of trained personnel. The study employed anesthesia (i.e. isoflurane 5%) and euthanasia methods consistent with the commonly accepted norms of veterinary best practice.
Neuronal culture preparation and transfection
Primary hippocampal neuronal cultures were prepared from embryonic day 19 rat hippocampi as previously described in Piccoli et al.35. Neurons were transfected using calcium-phosphate method at day in vitro (DIV) 10 with LifeAct-GFP plus plasmids expressing both the fluorescent mCherry marker (under the control of the neuron-specific Ca2+/calmodulin-dependent protein kinase II promoter) and either CAP2 small harpin RNA (CAP2-shRNA, sequence CCAAGTCGTCCGAGATGAATGTCCTGGTC) or a control scramble sequence (SCR, sequence GCACTACCAGAGCTAACTCAGATAGTACT)18. LifeAct-GFP was a gift from Dr. Margarita Dinamarca (University of Basel, Switzerland).
FRAP experiment
At DIV12, FRAP assays were performed at 37°C and with \(5\%\) CO2 in culture medium [Neurobasal medium, supplemented with B27, without phenol red (Thermofisher)]. Inverted Zeiss LSM confocal microscopes, equipped with a temperature-controlled chamber and CO2 supply, and Plan-Apochromat 63x/1.40 Oil were used (Zeiss LSM 510: cultures 1 and 2; Zeiss LSM 900: cultures 3, 4, 5, and 6) were used for the acquisition with settings of 512 x 512 pixel resolution. The assay was performed in neurons that were co-expressing both LifeAct-GFP and either CAP2-shRNA or SCR (visualized by the fluorescent mCherry marker). Dendritic spines with a large bulbous spine-head, corresponding to mushroom mature spines, were selected. FRAP was performed only on the GFP channel. Prebleach fluorescent signal of GFP was acquired using a 488 nm line at \(2.2\%\) laser power. The frame including the ROI (region of interest containing the whole spine) was imaged three times before bleaching. Photobleaching of the ROI was achieved by scanning with the 488 nm argon laser line at 100% laser power28. Postbleach fluorescent signal was acquired using a 488 nm line at \(2.2\%\) laser power (prebleach: 3 frames; postbleach acquisition: 65 frames).
Data processing
The ImageJ software was used to measure the fluorescence (integrated density) recovery in the dendritic spines. After image acquisition, the dendritic spines lacking fluorescence recovery because of focus drift were excluded from the quantitative analysis. We carried out a double normalization to analyze the spine fluorescence recovery. The intensity of the bleached spine is normalized to a neighboring unbleached dendritic area to diminish error caused by normal photobleaching during the monitoring period4. The fluorescence values of the three prebleach sequences were then averaged and all the fluorescence values of the postbleach frames were normalized by dividing each value by the average of all prebleach measurements. This normalization of the prebleaching fluorescence to 1.0 allows to compare different curves from different spines to each other. The first post-bleach measurement was set to 0 s. To prevent potential experimental bias due to insufficient bleaching efficiency11, we excluded spines with a normalized fluorescence intensity level at time 0 greater or equal to 0.6. From the original 136 dendritic spines, 4 spines (\(3\%\) of the total; 3 SCR, 1 CAP2-shRNA) were removed from the analysis, leaving a total of 132 spines (61 SCR, 71 CAP2-shRNA).
Data structure
The 132 selected dendritic spines provided 65 measures equally spaced in time over a period of 100 s, for a total of 8580 observations. Spines belonged to 54 neurons, grouped in 6 cultures. The number of spines within neurons, and of neurons within cultures was not constant. CAP2-shRNA neurons were 28 (versus 26 SCR controls) and the resulting CAP2-shRNA spines were 71 (versus 61 SCR controls) (Figs. 1 and S2).
Statistical analysis
We referred to a nonlinear mixed-effects model where an asymptotic exponential curve describes the normalized fluorescence intensity recovery over time and random-effects parameters allow for the potential variability of the dynamic of normalized fluorescence intensity recovery to be captured across spines, neurons, experimental conditions, and cultures.
Model selection was performed by following the approach outlined in Pinheiro and Bates23, where the final model is constructed through a series of incremental additions/deletions and refinements. This process involved three key steps: (1) defining the random-effects structure, (2) specifying the variance function to account for potential heteroscedasticity, and (3) establishing the temporal dependence structure. At each step the significance of each addition/deletion was evaluated by using the AIC, the BIC, and the LRT (when appropriate), along with visual inspection of diagnostic plots. A detailed description of the selected model, including the performance of model selection criteria (Table S1 and Fig. S3) and diagnostic checks (Figs. S4, S7, and S8) is available in the SI.
Briefly, in the selected model \(y_{cgnsi}\), the normalized fluorescence intensity measured at time \(t_i\) (\(i=1,\ldots ,{65}\)) for the spine s (\(s=1,\ldots ,S_{cgn}\)) in neuron n (\(n=1,\ldots ,N_{cg}\)) of the experimental condition g (\(g=(\text {SCR},\text {SH})\)) in culture c (\(c=1,\ldots ,C\)), is modeled as
where R0, the response y at time 0, is the normalized fluorescence intensity after photobleaching, the horizontal asymptote Asym is the normalized fluorescence intensity recovered in time, and the log rate constant lrc measures how fast the normalized fluorescence intensity recovers. Thus, the mobile fraction, or dynamic actin pool, can be estimated as the difference between the asymptote Asym and the normalized fluorescence recovery at time 0, R0 (Fig. S1).
In the selected model, the parameters vary according to
where \(\alpha ^{(0)}\), \(\lambda ^{(0)}\), and \(\rho ^{(0)}\) are fixed-effects parameters, whereas \(\alpha ^{(j)}\), \(\lambda ^{(j)}\), and \(\rho ^{(j)}\), \(j>0\), are random-effects parameters at culture (\(j=1\)), neuron (\(j=2\)), and spine (\(j=3\)) levels. They are assumed independent within each level and normally distributed. At the culture level, \(\alpha ^{(1)}_{c} \sim N(0, \sigma ^{2}_{1})\), \(\lambda ^{(1)}_{c} \sim N(0, \tau ^2_{1})\), and \(\rho ^{(1)}_{c} \sim N(0, \nu ^2_{1})\); at the neuron level, \(\rho ^{(2)}_{cgn} \sim N(0, \nu ^2_{2})\); at the spine level, \(\alpha ^{(3)}_{cgns} \sim N(0, \sigma ^{2}_{3})\), \(\lambda ^{(3)}_{cgns} \sim N(0, \tau ^2_{3})\), and \(\rho ^{(3)}_{cgns} \sim N(0, \nu ^2_{3})\).
Lastly, \(\varepsilon _{cgnsi}\) are heteroscedastic and correlated within-group errors assumed independent of the random effects. In particular:
where \(\delta _{g}\), for \(g=(\text {SCR},\text {SH})\), are the variance parameters for the two experimental conditions and \(v_{cnsi}\) are the fitted values, \(k=0,1,\ldots\) is the time index, \(|\phi |<1\) is the lag-1 correlation parameter of the autoregressive model, and \(i \ne i'\).
We simulated 100 datasets based on the model’s parameter estimates to: (1) underscoring the differential effect of the experimental condition at the variance level and (2) validating the selected model. Specifically, the simulated data were generated by sampling from the parameter vector, assuming normality, and incorporating two key factors: (1) an exponential variance structure interacting with the experimental condition and (2) a first-order autocorrelation structure. To underscore the differential effect of the experimental condition at the variance level, we provided a visual representation of the simulated spine-specific trajectories, as well as their mean and selected quantiles. To validate the selected model, we provided a visual representation of the observed and simulated spine-specific trajectories, grouped by neuron as well as culture and experimental condition combined. Finally, as a second assessment of model validity, we conducted a robustness analysis excluding ten randomly selected spines at a time and refitting the final model on the restricted dataset. We repeated the same procedure for three times by using a different seed. We then compared: (1) the point estimates and the corresponding confidence intervals across the scenarios under investigation and with the results from the main analysis and (2) the excluded spines with the predicted curves obtained from the selected model, as estimated over the restricted dataset.
FRApp Shiny application
In order to popularize the proposed statistical methodology in the neuroscience community, we developed an R-based package using the Shiny user interface, termed FRApp, based on the originally used R package \(\texttt {nlme}\)36. This interactive application serves at least two purposes: (1) non-R users can easily accommodate fitting of nonlinear mixed-effects models with asymptotic exponential growth curve to their data using the R software, which otherwise would require programming skills37,38 and (2) the reproducibility of our analysis can be easily assessed by separating out the different steps of the performed analysis and playing around with the options proposed in FRApp. In addition, sensitivity analyses can be easily carried out to assess the robustness of the statistical model when specific subsets of the data are considered.
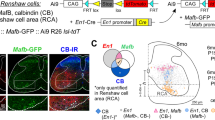
Scheme of the FRAP experiment and corresponding raw data grouped according to their hierarchical structure. (a, left panel) Schematic representation of the experimental plan highlighting the hierarchical structure of the data. The dendritic spines analyzed belonged to hippocampal neurons, grouped in different hippocampal neuronal cultures. In each culture, neurons were transfected to express LifeAct-GFP and downregulate the actin-binding protein CAP2 (SH). As control condition, neurons were transfected with a scramble sequence (SCR). (a, right panel) Representative images of baseline GFP signal (pre-bleach) in neurons transfected with either CAP2-shRNA (SH) or SCR. The visualized neurons were presented in false-color according to the schematic representation in (a). A Region of Interest (ROI, white dashed circle) was photobleached and the dendrite was imaged immediately thereafter (t=0) and at the indicated times post-bleach (t=50s and t=100s). (b) Spine-specific normalized fluorescence intensities over time, as grouped by neuron (rows), and culture and experimental condition [CAP2-shRNA (SH) or SCR] combined (columns). Normalized fluorescence intensity measures over time were depicted as connected points. SCR and SH spines are identified with red and blue color, respectively; excluded spines were represented in grey color. The complete dataset was presented in Fig. S2.
Fitted curves and random effects at culture level. (a) Estimates of the global population curve (dashed line) and the culture-specific curves (solid lines). Horizontal red dotted lines represent estimates of \(\rho ^{(0)}\) and \(\alpha ^{(0)}\). (b) Random-effects estimates at the culture level. The parameters of the culture-specific curves are obtained by adding the random-effect estimates from the (b) panel to the fixed- effect estimates. Notably, in the absence of significant interaction terms between any of the three fixed parameters and the experimental condition in the final selected model, the fitted global population curve was the same for both experimental conditions. Similarly, the six fitted culture-specific curves were the same for both experimental conditions. Indeed, the fixed effects at culture level were the same for any of the three parameters. In addition, the experimental condition was applied at the neuron-and not at the culture level-in this experimental design, thus preventing from properly defining the interaction term between random effects at culture level and experimental condition.
Spine-specific normalized fluorescence intensities over time (grey points), grouped by culture (rows) and experimental condition (columns). Notably, across the twelve plots, the dark grey dashed line represents the global population curve, which holds for both the two experimental conditions and the six cultures. In each row, the dark grey solid line represents the fitted culture-specific curve, which holds for both experimental conditions and the specific culture under examination.
Spine-specific normalized fluorescence intensities over time (grey dots), grouped by neuron (rows), and culture and experimental condition [CAP2-shRNA (SH) or SCR] combined (columns). Grey dashed lines represent fitted curves at the neuron level. Red and blue solid lines represent fitted curves at the spine level for SCR and CAP2-shRNA (SH) conditions, respectively.
Monte Carlo simulation samples of normalized fluorescence intensity \(y_{cgns}\) according to the final model selected for the two experimental conditions. Grey lines represent 100 simulated datasets. The solid line represents the mean, while dotted lines represent quantiles of order 0.05, 0.25, 0.5, 0.75, and 0.95, from the bottom to the top. Notably, the sampled trajectories, the mean, and the quantiles simultaneously accounted for both the fixed and the random effects included in the final model, as well as the variance components and the autoregressive structure. Therefore, all trajectories could accommodate for the random effects, particularly those for Asym at the spine level. The two red lines highlight that a difference could be appreciated between the mean curves at plateau and this was due to the random effects for Asym at the spine level (see Fig. S6). The simulated trajectories differed from the fitted global population curve presented in Fig. 2a, which only accounted for the fixed effects.
Model validation: comparison of observed data with 50 randomly selected Monte Carlo simulation samples of normalized fluorescence intensity \(y_{cgns}\) according to the selected model (grey lines) at the spine-specific level. Observed spines were grouped by neuron (rows), and culture and experimental condition [blue for CAP2-shRNA (SH) or red for SCR] combined (columns). These observed spines were the same reported in Fig. 1b for neurons 1-4 and in Fig. S2. Notably, the simulated spines generally aligned well with the observed data, due to the inclusion of random effects at the spine level. When deviations occurred, the corresponding simulated spines were regularized by the shrinkage effect inherent in the mixed-effects model.
Using FRApp to upload and analyze the data described in this paper with the final statistical model selected. (a) Data loading: data upload options are presented including the Load example data button that automatically loads the dataset and selects the specification of the final model presented in the paper. (b) Model specification: dependent and explanatory variables, as well as the hierarchical structure subsumed by the random effects, the variance function, the autocorrelation order, and possible interaction terms involving the experimental condition can be specified. (c) Model results, model summary: summary information on the model including estimates of the fixed and random effects, the correlation structure, and the variance function are displayed. (d) Model results, plots: scatterplot of residuals vs. fitted values (upper) and quantile-quantile plot of the residuals (lower) are displayed. (e) Model results, confidence intervals: approximate estimates and 95% confidence intervals of parameters under investigation are displayed.
Data availibility
Data and corresponding code for data cleaning and statistical analysis, together with the R package source file, are provided in the SI. The R package is available on the first author’s GitHub public repository (https://doi.org/10.5281/zenodo.14849826) and published on the Comprehensive R Archive Network (CRAN).
References
Mueller, F., Mazza, D., Stasevich, T. J. & McNally, J. G. FRAP and kinetic modeling in the analysis of nuclear protein dynamics: What do we really know?. Curr. Opin. Cell Biol. 22, 403–411 (2010).
Giakoumakis, N. N., Rapsomaniki, M. A. & Lygerou, Z. Analysis of protein kinetics using fluorescence recovery after photobleaching (FRAP). In Markaki, Y. & Harz, H. (eds.) Light Microscopy: Methods and Protocols, 243–267, https://doi.org/10.1007/978-1-4939-6810-7_16 (Springer New York) (2017).
Reits, E. A. & Neefjes, J. J. From fixed to FRAP: Measuring protein mobility and activity in living cells. Nat. Cell Biol. 3, E145-7 (2001).
Koskinen, M., Bertling, E. & Hotulainen, P. Methods to measure actin treadmilling rate in dendritic spines. Methods Enzymol. 505, 47–58 (2012).
Koskinen, M. & Hotulainen, P. Measuring F-actin properties in dendritic spines. Front. Neuroanat. 8, 74 (2014).
Sala, C. & Segal, M. Dendritic spines: The locus of structural and functional plasticity. Physiol. Rev. 94, 141–188 (2014).
Pollard, T. D. The cytoskeleton, cellular motility and the reductionist agenda. Nature 422, 741–745 (2003).
Hotulainen, P. et al. Defining mechanisms of actin polymerization and depolymerization during dendritic spine morphogenesis. J. Cell Biol. 185, 323–339 (2009).
Star, E. N., Kwiatkowski, D. J. & Murthy, V. N. Rapid turnover of actin in dendritic spines and its regulation by activity. Nat. Neurosci. 5, 239–246 (2002).
Honkura, N., Matsuzaki, M., Noguchi, J., Ellis-Davies, G. C. & Kasai, H. The subspine organization of actin fibers regulates the structure and plasticity of dendritic spines. Neuron 57, 719–729 (2008).
Rapsomaniki, M. A. et al. easyFRAP: An interactive, easy-to-use tool for qualitative and quantitative analysis of FRAP data. Bioinformatics 28, 1800–1801 (2012).
Wilson, M. D., Sethi, S., Lein, P. J. & Keil, K. P. Valid statistical approaches for analyzing sholl data: Mixed effects versus simple linear models. J. Neurosci. Methods 279, 33–43 (2017).
Paternoster, V. et al. The importance of data structure in statistical analysis of dendritic spine morphology. J. Neurosci. Methods 296, 93–98 (2018).
Lazic, S. E., Clarke-Williams, C. J. & Munafò, M. R. What exactly is ‘N’ in cell culture and animal experiments?. PLoS Biol. 16, e2005282 (2018).
Lazic, S. E. The problem of pseudoreplication in neuroscientific studies: Is it affecting your analysis?. BMC Neurosci. 11, 5 (2010).
Aarts, E., Verhage, M., Veenvliet, J. V., Dolan, C. V. & Van Der Sluis, S. A solution to dependency: Using multilevel analysis to accommodate nested data. Nat. Neurosci. 17, 491–496 (2014).
Yu, Z. et al. Beyond t test and ANOVA: Applications of mixed-effects models for more rigorous statistical analysis in neuroscience research. Neuron 110, 21–35 (2021).
Pelucchi, S. et al. Cyclase-associated protein 2 dimerization regulates cofilin in synaptic plasticity and Alzheimer’s disease.. Brain Communications 2, fcaa086 (2020).
Belin, B. J., Goins, L. M. & Mullins, R. D. Comparative analysis of tools for live cell imaging of actin network architecture. BioArchitecture 4, 189–202 (2014).
Wegner, W. et al. In vivo mouse and live cell STED microscopy of neuronal actin plasticity using far-red emitting fluorescent proteins. Sci. Rep. 7, 11781 (2017).
Konietzny, A. et al. Myosin V regulates synaptopodin clustering and localization in the dendrites of hippocampal neurons. J. Cell Sci. 132, jcs230177 (2019).
Lachowski, D. et al. G protein-coupled estrogen receptor regulates actin cytoskeleton dynamics to impair cell polarization. Front. Cell Dev. Biol. 8 (2020).
Pinheiro, J. & Bates, D. Mixed-Effects Models in S and S-PLUS (Springer, Berlin, 2000).
Carnell, M., Macmillan, A. & Whan, R. Fluorescence recovery after photobleaching (FRAP): Acquisition, analysis, and applications. Methods Membrane Lipids 1232, 255–271 (2015).
Gelman, A. & Hill, J. Data Analysis Using Regression and Multilevel/Hierarchical Models (Cambridge University Press, Cambridge, 2006).
Axelrod, D., Koppel, D., Schlessinger, J., Elson, E. & Webb, W. W. Mobility measurement by analysis of fluorescence photobleaching recovery kinetics. Biophys. J . 16, 1055–1069 (1976).
Mikhaylova, M. et al. Caldendrin directly couples postsynaptic calcium signals to actin remodeling in dendritic spines. Neuron 97, 1110–1125 (2018).
George, J., Soares, C., Montersino, A., Beique, J.-C. & Thomas, G. M. Palmitoylation of LIM Kinase-1 ensures spine-specific actin polymerization and morphological plasticity. Elife 4, e06327 (2015).
Freedman, L. P., Cockburn, I. M. & Simcoe, T. S. The economics of reproducibility in preclinical research. PLoS Biol. 13, e1002165 (2015).
Feilke, M., Schneider, K. & Schmid, V. J. Bayesian mixed-effects model for the analysis of a series of FRAP images. Stat. Appl. Genetics Molecular Biol. 14, 35–51 (2015).
Lazic, S. E., Mellor, J. R., Ashby, M. C. & Munafò, M. R. A Bayesian predictive approach for dealing with pseudoreplication. Sci. Rep. 10, 2366 (2020).
Pelucchi, S., Stringhi, R. & Marcello, E. Dendritic spines in Alzheimer’s disease: How the actin cytoskeleton contributes to synaptic failure. Int. J. Mol. Sci. 21, 908 (2020).
Pelucchi, S., Gardoni, F., Di Luca, M. & Marcello, E. Synaptic dysfunction in early phases of Alzheimer’s disease. Handb. Clin. Neurol. 184, 417–438 (2022).
McGrath, J. C., Drummond, G., McLachlan, E., Kilkenny, C. & Wainwright, C. Guidelines for reporting experiments involving animals: The arrive guidelines. Br. J. Pharmacol. 160, 1573–1576 (2010).
Piccoli, G. et al. Proteomic analysis of activity-dependent synaptic plasticity in hippocampal neurons. J. Proteome Res. 6, 3203–3215 (2007).
Pinheiro, J., Bates, D., DebRoy, S., Sarkar, D. & R Core Team. nlme: Linear and Nonlinear Mixed Effects Models (2023). R package version 3.1-162.
Chang, W. et al. Shiny: Web Application Framework for R (2023). R package version 1.8.0.
R Core Team. R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna (2023).
Acknowledgements
We thank Elisa Zianni for technical assistance in laboratory, Filippo La Greca for excellent practical work, and Dariush Khaleghi Hashemian for technical assistance in testing the Shiny app.
Funding
This work was supported by the Giovanni Armenise Harvard Foundation and AIRALZH ONLUS (2023 Armenise Harvard-AIRALZH Mid-Career Award in Neurodegenerative Diseases - AHA MCA- to EM), by the Italian Ministry of Enterprises and Made in Italy (PNRR - Next generation EU funding, “SEED for Innovation Patent 2.0” program, project TT_MIN23_SEED4IP2.0_07 to EM), the Fondazione Cariplo (grant number 2018-0511 to EM), the Italian Ministry of University and Research (grant numbers PRIN 202039WMFP and PRIN 2022 PNRR P2022R2E8N to EM, PRIN 20227YCB5P to VE (PI), FP (co-PI), and GDC (project member), Università degli Studi di Milano (grant number PSR2022 DIP_022_AZIONE_A_SPELU to SP), and MUR Progetto Eccellenza to DiSFeB, and MUR Progetto Eccellenza to DISCCO). The authors acknowledge support from the University of Milan through the APC initiative.
Author information
Authors and Affiliations
Contributions
E.M., V.E. designed research; S.P. and R.S. performed research; G.D.C. analyzed data and developed the interactive application; E.M., S.P. and V.E. obtained funding; F.P. critically revised the paper; G.D.C., S.P., E.M., V.E. wrote the paper. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Supplementary Video S1.
Supplementary Video S2.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Di Credico, G., Pelucchi, S., Pauli, F. et al. Nonlinear mixed-effects models to analyze actin dynamics in dendritic spines. Sci Rep 15, 5790 (2025). https://doi.org/10.1038/s41598-025-87154-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-87154-w