Abstract
MtbClpC1 is a promising drug target against tuberculosis. Recent studies have shown that several natural product antibiotics targeting the unfoldase N-terminal domain can impair MtbClpC1 function resulting in cell death. While the pharmacological properties of these natural product antibiotics prevent their use in the clinic, similar molecules binding to the same binding pockets can result in new drugs against Mtb. Here we demonstrate that we successfully used in silico screening to identify new ClpC1 N-terminal domain binders with micromolar affinity from a small compound library. In addition, we experimentally demonstrate that the new compounds bind to the same pockets used by the natural product antibiotics and inhibit ClpC1 function.
Similar content being viewed by others
Introduction
The unfoldase ClpC1 is one of the most promising drug targets against tuberculosis1,2,3,4. This AAA + unfoldase works in cooperation with the ClpP1P2 protease and is the target of at least four natural product antibiotics (NPAs): cyclomarin5,6,7,8, ecumicin9,10, lassomycin11,12 and rufomycin13,14,15. Indeed, ecumicin, cyclomarin, rufomycin and lassomycin, all targeting ClpC1, are among the most powerful anti-TB molecules to have emerged recently. For example, ecumicin displays potent selective anti-tuberculosis activity with an MIC value 50 times lower than that of rifampicin or isoniazid, the first line drugs for the treatment of tuberculosis10. ClpC1 is a member of the class II AAA + family of proteins, which contains a N-terminal domain (NTD) and two distinct ATP-binding modules, D1 and D2. We have recently determined the full-length structure of ClpC1 in its apo and 2 different antibiotic bound states2. Despite representing only a small portion of the full protein, all the NPAs have been shown to bind to the ClpC1-NTD domain and high-resolution X-ray structures of the binding sites are available for cyclomarin, ecumicin and rufomycin (Fig. 1A–C)5,9,12,13. Recently the X-ray structure of the ClpC1-NTD bound to lassomycin has also been published12. While this allows a proper mapping of the NPAs binding pockets, it is still not clear how binding to the NTD can translate into functional impairment of the remaining protein.
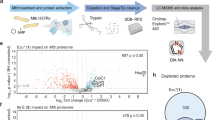
In silico screen structures of the antibiotic bound ClpC1 NTD. (A) Cyclomarin (PDBid: 3WDC), (B) Ecumicin (PDBid: 6PBS) and (C) Rufomycin (PDBid: 6CN8). Antibiotics are shown in dark blue stick representation. Coloured ClpC1 NTD binding sites are: A) blue, residues: M1, F2, E3, Q17, F80, K85, E89. (B) pink: M1, F2, E3, R4, V14, Q17, H77, K85, L92, L96. (C) green, residues: M1, F2, V13, H77, F80, E89. (D) The 9 first structures selected through an Autodock Vina energy score ranking based on the three binding sites described in (A–C). A black reference line is marking the cut off at 9 compounds.
Although, these NPAs are promising starting points, due to their complex multiring structures, they are challenging for medicinal chemistry and display poor pharmacological properties. Nevertheless, considering the chemical diversity, the sterilizing properties and the variety of mechanisms of action displayed by ClpC1 inhibitors, we believe that small molecules acting on the same binding sites will be equally able to block ClpC1P1P2 activity and represent valid drug candidates.
In recent years, in silico screening has emerged as a reliable approach for rapid compound screening with multiple advantages over classical screening protocols. Most importantly a larger number of compounds can be tested at lower costs and in less time, enabling the screen of libraries comprising millions of compounds, and thus, far beyond the scale accessible to even the most ambitious conventional HTS. In addition, recent developments have far improved the percentage of in silico hits that display biological activity. Indeed, although in silico screening is often regarded with some scepticism it is worth mentioning a report from Bayer HealthCare16 stating that half of the 20 new chemical entities currently being tested in phase I clinical trials significantly benefited from computer-aided design methods.
Here we show that in silico screening can be used to successfully find ClpC1 inhibitors with micromolar affinity.
Results
In silico screen
Our goal was to use structural information from all three published ClpC1-NTD antibiotic complexes to design an in-silico screen that would take into account all the possible interactions within the binding site. Thus, increasing our chances of finding small-molecule ClpC1 binders, ideally with the same diverse biochemical features as the NPAs. The structural data of lassomycin was published only during the preparation of this work and was not used for our in-silico screening approach. Nevertheless, lassomycin shares the same binding site with the remaining NPAs12. (see Supplementary Fig. 1). A description of the pharmacophore units displayed by the different natural product antibiotics is shown in Supplementary Fig. 6.
The employed strategy of our in-silico approach is presented in Fig. 1 together with a flow chart of the approach in Supplementary Fig. 2. The library used for our in-silico screen was the ICSN chemical library, which contains 5481compounds, made up of 33% natural products, 33% analogues of natural products and 33% synthetic products. This represents a very high level of molecular diversity. It includes steroids, terpenes, sugars, nucleosides, taxoids, macrolides, pyrole and flavonoids and numerous synthetic compounds with different scaffolds. Of note the chemical library contains no commercial compounds. After a virtual screen of the ICSN library with the three ClpC1-NTD NPA structures (PDBID: 3WDC, 6PBS, 6CN8), a ranked list for each of the structures was created. To select compounds that target the common binding site in ClpC1-NTD a list of consensus hits was created adjusting the cutoff of the AutoDock VINA ranking score. This way we reduced the number of hits to be experimentally tested by employing the criterion of consensus docking to find a hit for all three conformations. We decided for an AutoDock VINA energy score cutoff of 46–56, including 9 compounds to be tested (see Fig. 1D).
In vitro hit validation
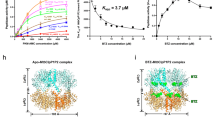
The top ranked consensus hits (Fig. 1D) were selected for validation and biochemical testing. Although, we have previously developed enzymatic based methods for the identification of ClpC1P1P2 inhibitors, we opted here to use isothermal titration calorimetry (ITC) as a first approach to confirm ClpC1-NTD binding1. This option derives mainly from 2 considerations. First, some compounds found to be true binders of ClpC1-NTD and active against Mtb do not display dramatic effects on the enzymatic activity of ClpC1. An example of this is cyclomarin, active in vivo against Mtb and binding ClpC1-NTD with nM affinity, but not a strong inhibitor of ClpC1P1P2 protein degradation5,7. Second, activity-based assays, as we and others have described, are prone to false positives, for example derived from pan-assay interference1,17, particularly when the concentrations of enzyme used in the assays are high. In this concern, ITC, although a low throughput technique, is considered a gold-standard for assessing biological interactions18. ITC can be reasonably used for testing a small number of hits and has the great advantage of helping to avoid nonspecific binders and, in addition, determining compound binding affinity irrespective of a biological effect (inhibition or activation of an enzyme reaction). Indeed, using ITC, from the 9 hits tested, only one compound, (2R,4aR,5S,8aR)-2-(2-((4-(benzylamino)benzoyl)oxy)ethyl)-5-((Z)-2-(4-carboxyphenyl)-2-chlorovinyl)-7-methyl-1,5,8,8a-tetrahydronaphthalene-4a(2H)-carboxylic acid (called here F5) was confirmed as a binder with a Kd of 1.6 µM (see Fig. 2). The ITC results for the remaining compounds are displayed in Supplementary Fig. 3.
Isothermal titration calorimetry curves of compound F5. Isothermal titration calorimetry of compound F5 into ClpC1 NTD WT (A) and mutants (B: V13A, C: F80A, D: K85A). For the WT protein a D Kd of 1.6 μM was determined assuming a binding stoichiometry of 1:1, for the V13A mutant the binding affinity was 15 × lower (24 μM) and for the F80A and K85A mutants no binding could be observed.
As F5 represented a new ClpC1-NTD binder scaffold we decided to further test 17 F5 analogues by ITC (Supplementary Fig. 4 and Table 1). Although with varying affinities, all analogues were shown to bind to the ClpC1-NTD domain. Both F5 and analogues are derivatives of meiogynin A, a natural sesquiterpenoid dimer isolated from a Malaysian tree bark19. They all present a cis-decalin carbon skeleton with a carboxylic acid at the ring junction to which a propenyl benzoic acid or a chloro-propenyl benzoic acid moiety is attached. This basic structure is functionalized by a side chain of variable nature and length which gives, each of the C1-C18 compounds, its originality.
Hit binding interface
While ITC confirms binding of the compound to the ClpC1-NTD domain, it does not provide insights about the actual binding site. As our compounds have been designed to bind to the same pocket as the antibiotics ecumicin, cyclomarin and rufomycin, we needed to confirm that this is actually the site of interaction with F5. Our first approach was to use X-ray crystallography since many high-resolution structures of the ClpC1-NTD in apo and antibiotic bound form have been published. Indeed, we were able to obtain many conditions with high resolution apo ClpC1-NTD structures (1–1.3 Å), but were unable to introduce our compound either through soaking or co-crystallization. This might be due to a preferred crystallization of the very stable apo ClpC1-NTD. Instead of embarking in a more time-consuming extended crystallization screen, we opted for solution NMR spectroscopy as the resonance assignment of the ClpC1-NTD has been published and HSQC type NMR spectra can be recorded within minutes, allowing rapid screening of many compounds8. This provides a much faster way to immediately identify residues interacting with the compound and thus the binding site. To this end, we recorded 1H–15N HSQC spectra of the apo ClpC1-NTD and ClpC1-NTD bound to one and two equivalents of compound F5. The resulting spectra are shown in Fig. 3A. Residues that are most affected by the binding are shown in black boxes, among them are E3, R4, F5, Q17, I78, F80, K85, L90 and L92. This is in perfect agreement with residues reported to interact with cyclomarin (M1, F2, E3, Q17, F80, K85, E89)5,8, ecumicin (M1, F2, E3, R4, V14, Q17, H77, K85, L92, L96)9 and rufomycin (M1, F2, V13, H77, F80, E89)13. Figure 3B shows the binding site marked in orange on the ClpC1-NTD structure. Due to the nature of the NMR chemical shift, we observe a much larger interaction interface as the ones reported for X-ray structures and validated by mutational experiments (Fig. 1A–C), but, as can be seen in Supplementary Fig. 5 the interaction pattern mapped by NMR of ecumicin and cyclomarin matches very well to the one of compound F5 and is located in the binding pocket formed by helices H1 and H5 and the adjacent loop regions. Due to the smaller size of F5 the binding profile resembles more the one created by cyclomarin, as for example the region around L96, one of the key interacting residues with ecumicin, might not be reached anymore by our compound.
Binding site confirmation by NMR. Evaluation of on target action of the in silico designed compounds. (A) 1H-15N HSQC spectrum of 15N labelled ClpC1 NTD in the apo (black) and compound bound state (1:1 orange, 1:2 red ClpC1 NTD:compound). Residues with large chemical shift differences and intensity changes are highlighted. (B) Interacting residues with high chemical shift changes are marked in orange on the structure of an MD simulation of ClpC1 NTD together with compound F5. (C) Combined chemical shift change between apo and compound bound ClpC1 NTD. The position of the 8 alpha helices in ClpC1 NTD are marked in grey and named H1-H8.
To further investigate the interaction between the ClpC1-NTD and F5 we performed 0.5 µs Molecular Dynamics (MD) simulations of the complex obtained from AutoDock. MD simulations were performed in explicit solvent to let the protein–ligand complex explore the conformational space in an NPT ensemble at 300 K and 1 bar. As can be seen in Fig. 4A, F5 never leaves the binding pocket but adopts two slightly different conformations in the same binding site as ecumicin, lassomycin and rufomycin (Fig. 4B). The presence of these two populations can be also seen in the RMSD histogram of the MD simulation in Fig. 4A. End-point free energy calculations, using MM-PB(SA) and MM-GB(SA), on these two different sections of the MD (0–170 ns) and (170–500 ns) demonstrate a stabilization of the second conformation by − 3.9 kcal/mol and − 3.0 kcal/mol respectively, that can be mainly explained by a gain in electrostatic energy (for calculation parameters see Table 2).
Molecular Dynamics simulation. 0.5 μs MD simulation of the ClpC1 NTD bound to the identified ligand F5 in explicit solvent at 300 K and 1 bar. (A) RMSD vs. time of the MD simulation and the corresponding histogram. (B) Representation of the 2 conformations adopted during the MD simulation. The F5 ligand coloured in blue, shows a differential interaction pattern in the 2 conformations. ClpC1 NTD residues interacting in conformation 1 are: F5, R10, V13, V14, Q17, I28, H77, F80, K85, E89, L92 and in conformation 2: M1, F2, E3, F5, R10, V13, H77, F80, K85, E89, L92.
To corroborate our findings of the NMR and MD experiments, we introduced single-point mutations in the binding site (V13A, F80A, K85A). These residues, were identified as the most effected by F5 binding in our NMR analysis, they are key residues for the interaction with cyclomarin, rufomycin and ecumicin in our pharmacophore analysis (see Supplementary Fig. 6B) and they have been reported in the literature to confer resistance to the NPAs cyclomarin, rufomycin and ecumicin. Consistent with our previous results, these mutations (V13A, F80A and K85A) resulted in dramatic changes in F5 binding (Fig. 2B-D). While the mutation V13A lead to a 15 × increase in the Kd, no binding was observed for the F80 and K85 mutants.
To conclude, F5 shares as predicted the same binding site with cyclomarin, ecumicin and rufomycin.
Enzymology and biological activity
ClpC1 function alone and in complex with ClpP1P2 is frequently tested using either ClpC1 intrinsic ATPase activity or following protein substrate degradation in association with ClpP1P21. Although the NPAs targeting ClpC1-NTD share the same binding surface, they display considerable differences in the effects they mediate on ClpC1P1P2 activities. For example, while ecumicin is a strong activator of ATPase activity and an inhibitor of FITC-casein degradation10, cyclomarin has very modest effects on those activities7. Taking this into consideration it was important to test what was the effect of F5 and its analogues on ClpC1 mediated activities, namely ATPase activity and degradation of a folded, GFP, and unfolded, FITC-casein, substrate in association with ClpP1P2. In the ATPase assay we measure the ClpC1 use of ATP that, coupled with pyruvate kinase and phosphoenolpyruvate dehydrogenase, leads to the consumption of NADH and formation of NAD +. The FITC-casein and GFPssra assays, measure the degradation of an unfolded and folded substrate respectively taking advantage of the FITC fluorescein reporter and the intrinsic fluorescence of GFP1.
In Fig. 5 we attempt to group compounds according to their biochemical profile. Compounds with an inhibitory effect on ATPase activity and GFPssra degradation are colored green, while compounds with an additional inhibitory effect on FITC-casein degradation are marked yellow. 4 compounds, although binders of the ClpC1-NTD, do not seem to have any biochemical effect and are marked grey. The mother compound F5, is marked in blue, but shares the same profile as the compounds marked in green. The first striking feature is an overall lack of correlation between the compound affinity (KdD) determined by ITC (Table 1) and the impact on enzymatic activities (see Figs. 5 and 6). What appears nevertheless clear is that a more significant inhibition of ClpC1 is obtained with bulkier molecules that likely better occupy the NTD substrate binding surface. Indeed, plotting compound solvent accessible surface area (SASA) versus enzymatic activity (Fig. 6B and Supplementary Fig. 8A,B), it is clear that smaller compounds, even when displaying lower Kd values (Fig. 6A) are less powerful inhibitors. Curiously, contrary to our expectation that larger molecules, resembling the larger cyclic NPA structures, would be better binders was not confirmed. C1 and C13, have a simplified structure and lower molecular weight compared to compound F5, but show a similar Kd to several larger analogs in our list.
Biochemical evaluation of ClpC1 NTD binders. (A) Biochemical characteristics of the 18 analogs selected based on the identified hit (F5 coloured in blue). Kd in μM measured by ITC, ATPase activity, FITC-casein (as example of unfolded protein degradation) and GFPssra (as example of folded protein degradation) degradation activity. Values are given as % enzymatic activity vs DMSO control. Compounds are coloured according to their biochemical behaviour: blue (original compound), grey (no inhibition), green (ATPase and GFPssra degradation inhibition), yellow (ATPase, GFPssra and FITC-casein degradation inhibition). As a cutoff 30% inhibition are used. (B) Structures of the analogs grouped and coloured based on table A).
Correlation between molecule size and activity. Correlation between different biochemical and biophysical parameters. Shown are: (A) Solvent-accessible surface area (SASA) vs Kd and (B) SASA vs ATPase activity. While the affinity is not correlated with the size of the molecule, the size of a molecule (expressed as SASA) seems to strongly influence if a compound is biochemically active. A dashed reference line is drawn at a SASA of 700 Å2 indicating a possible cutoff for active vs inactive molecules. A grey background shows molecules with a lower Kd (less than 10 μM) in panel (A), as well as molecules with a strong inhibitory effect on ATPase activity (less than 25%) in panel (B).
While none of the compounds identified in this screen can be compared with the described NPAs, either in terms of Kd or inhibition potency, they are superior to compounds discovered previously using a 115,000 compound library in an expensive and time consuming HTS based on a ClpC1P1P2 functional assay1. For example, the IC50 for FITC-casein ClpC1P1P2 degradation of compound C9 was determined as 6 µM (Supplementary Fig. 7), while the best inhibitor retrieved from the previous HTS effort displayed an IC50 of 50 µM using a 20 × bigger library1.
In addition and although we are aware that the small size of the pilot library screened, only 5481 compounds, was unlikely to result immediately in biologically active molecules, we nevertheless tested F5 and analogues for activity against M. smegmatis and M. tuberculosis using protocols previously established in our laboratory4. As expected, considering the presence of two ionizable carboxy groups which prevent lipophilic membrane crossing (21), no activity was observed when the compounds were tested at a concentration of 50 µM.
Discussion
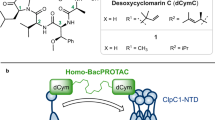
ClpC1 has been recognized as an important drug target against Mtb. This is supported not only by strong data obtained in different laboratories, but also by the discovery of 4 natural product antibiotics that target specifically this protein. In addition, the interest in ClpC1 has also been fueled by the recent rise of PROTACs as an alternative approach to drug development and the ClpC1 system has been the subject of several approaches to use targeted protein degradation to kill Mtb20,21,22,23. Yet and despite these advances, evolution did not take into account the pharmacological properties of the natural product antibiotics it created and all the NPAs found so far do not display good drug profiles, even when used as bacPROTAC molecules. This leads us to the question that motivated the present study: Can we find molecules that mimic the NPAs as a first step towards drug development?
So far, all efforts have struggled to identify molecules that can do that. As ClpC1-NTD does not have intrinsic enzymatic activity, drug screens can only be performed either using full-length ClpC1 or in association with ClpP1P2, in case protein degradation is measured1,24. This can result in the identification of a larger number of false positives or compounds that are hard to validate and develop due to their low affinity or specificity24. False positives can derive from multiple factors such as aggregation, covalent modification of the target, autofluorescence or interactions with the reporter and not the target. We propose that better results could be obtained if we reduce the screening space to just the ClpC1-NTD and use instead a structural screening approach based on the starting NPAs structures instead of a functional screening approach. The overall strategy employed here, is shown in Fig. 7.
Novel approach for finding ClpC1 NTD binders. Flow chart of the proposed innovative approach to identify small molecule binders using a structure based in silico screen and the known binding sites of three natural product antibiotics, including biochemical and biophysical methods for hit validation.
Reinforcing the importance of a proper hit validation using biophysical methods, only one of the 9 top ranked hits was found to be indeed a ClpC1-NTD binder according to ITC. This discrepancy can derive from limitations in our in-silico approach, nevertheless, and despite the small library screened, we were able to validate a bona fide binder of the ClpC1-NTD and demonstrate that it binds to the same binding interface as the NPAs inhibiting ClpC1 function using solution NMR. In addition, we observed inhibition of ATPase activity and the degradation of FITC-casein and GFPssra in association with ClpP1P2. Curiously, analogue search showed that despite several of the analogues displaying affinity for the ClpC1-NTD, they do not result in ClpC1 inhibition. From our data, it appears that in order to obtain enzymatic inhibition a significant occupancy of the ClpC1-NTD binding surface is required as bigger molecules lead to stronger enzymatic inhibition. This perhaps explains the low number of hits we identified in a recent activity screen on a small compound library (MW < 500 Da)1 as smaller binders were likely discarded due to the weak inhibition displayed.
ClpC1-NTD has recently been proposed to function as a receptor for unfolded substrates and therefore bulkier compounds that better occupy the surface of the domain, are likely better preventing substrate binding, translocation and posterior degradation by the ClpP1P2 protease21. This type of protein receptors creates a problem for drug development as they are designed to bind multiple sites with low affinity while the protein target is recognized and translocated into the unfoldase pore. Good binding and inhibition as observed with peptide derived natural product antibiotics is achieved by large surface molecules that are able to cover the ClpC1-NTD hydrophobic pocket. These peptide derived antibiotics have however some inherent drawbacks such as short half-life owing to the susceptibility to protease degradation and cytotoxicity to host cells. This appears to be the case o of the NPA´s targeting ClpC1-NTD that display non optimal pharmacological properties.
In this concern our results are particularly interesting as they show that non peptide derived molecules can also be used to target these surfaces, opening perspectives for other molecules, with distinct pharmacological properties, to still be identified. Indeed, we believe better binders will likely be retrieved once larger libraries are used as has been the standard in recent in silico screening efforts25.
It is important to note, that our approach represents only the first step towards drug development as all the compounds identified in our screen do not show biological activity versus either M. smegmatis or M. tuberculosis in our assays. This is a frequent problem in drug development and can derive from multiple factors, including poor membrane permeability, considerable efflux transport, poor target specificity or in cell drug degradation/metabolism. Indeed, one common feature of the binders found here is the presence of two ionizable carboxy groups that appear to be relevant in our models for the interaction with the NTD residues Q17 and K85, also key residues ruling cyclomarin binding. These charged residues can be responsible for the poor membrane permeability of the compounds as we have previously shown that carboxy acetylation improves biological effect26,27. These in vivo results stress a current limitation of in silico screening, which despite the impressive advances in recent years, is still far from being able to predict if a binder, regardless of its high affinity, will be useful in vivo as multiple other variables are involved.
Overall, we have demonstrated here that in silico screening is a valid approach in order to identify new binders of the ClpC1-NTD. Recent state of the art in silico screening using artificial intelligence led to an average confirmation rate by biological assays of 5.5% of the hits found, and it is worth mentioning that in the referred study, validation was obtained mainly via functional assays, that as we stress above are intrinsically inferior to ITC28. Taking the previous, a logic conclusion is that our approach is not inferior and should be applied to much larger libraries in order to expand the number of chemical scaffolds found and increase the probability of finding drug-like molecules. In addition, the fact that we were able to identify small ligands with micromolar affinities and low ClpC1P1P2 inhibition is particularly encouraging for the use with the ClpC1P1P2 system for targeted protein degradation. Indeed, so far all the molecules obtained derive from the natural product cyclomarin, taking advantage of its intrinsic high affinity for the ClpC1-NTD, but thus having the same pharmacological problems as the parent compound. Smaller molecules with sufficient affinity for ClpC1-NTD would certainly represent a major advantage towards this goal.
Methods
Structural based virtual screening
Virtual screening calculations were conducted using the ICSN chemical library, comprising 5481 compounds, both synthetic and natural. This library consists of substances sourced from various laboratories within the ICSN institute. Notably, 33% of the compounds are natural products, 33% are natural product analogs, and the remaining 33% are synthetic compounds (the collection is composed of steroids, alkaloids, sugars, nucleosides, taxoides, macrolides, pyroles, flavonoids, etc. and excludes any commercial products or compounds sourced from commercial vendors).
The conversion from SDF format to 3D structures for the ICSN chemical library was made in automatic pipelines implemented by home-made python routines using openbabel29. The hydrogens were automatically added by assuming a protonation state of the molecules at pH 7.4. 3D coordinates were generated by gen3d with the openbabel default option. Then conformer search was allowed to explore different ligand conformation and generate a lowest-energy set of conformers.
The employed target protein was the ClpC1-NTD in three different antibiotic-bound states, (PDBid 3wdc, 6cn8 and 6pbs bound to cyclomarin, rufomycin and ecumicin respectively).
A minimization step with obminimize was performed on each molecule with all the default parameters in order to create a pdb for further pdbqt conversion.
For the conversion of the pdb to pdbqt AutoDockTools30 was utilized with default values by the means of pythonsh routines and the prepare_ligand4.py script.
SBVS calculations were performed on the 3 different ClpC1 structures employing AutoDock VINA. A grid spacing of 1.0 Å was selected for AutoDock VINA (default value) centered on the ClpC1 ligands. The size of the grid was defined in order to embed all the Cyclomarin, Ecumicin and Rufomycin interactions with the protein in every ClpC1 NTD conformation.
An ensemble protein receptor consensus docking strategy was followed to score the compound leads for further experimental validation as follow:
First, we did virtual screening with the ICSN library, on each of the 3 ClpC1 X-ray receptors (3wdc, 6pbs and 6cn8) and as criterium, a rank with the best 100 scored compounds (lowest energy) was created for every receptor.
In the next step, a second criterium for the selection relies on finding a consensus of solutions between the 3 receptors, and only consider those hits that are in the intersection of the three list. In that way, decreasing the AD vina ranking, from 1 to 100, only the consensus number of hits was kept (see Fig. 1D and Supplementary Fig. 2) allowing to reduce the number of hits to be experimental validated (e.g. from 50 based on the AD ranking score, to 9 according to the consensus pool).
This pool contains ligands potentially suitable for binding to any of the different conformations present in the ClpC1 NTD containing the most relevant interacting elements, independent of conformational changes of the receptor. Only hits that are part of the first pool were then selected for experimental validation.
All the SBVS and their furthers analysis, was performed by multithread python-homemade pipelines.
Chemistry
Synthesis of compounds C1-C18 has been described elsewhere26,27,31.
Biochemistry and in vivo assays
Protein expression and purification (ClpC1-NTD, ClpC1, ClpP1, ClpP2 and GPFssra) were executed as previously described1,2,8. The ATPase, GFP and FITC-casein assays were performed as described in1,2. In vivo compound activity versus M. smegmatis and M. tuberculosis (H37rv) was tested as previously described4.
Isothermal titration calorimetry
The interaction between the selected compounds (F1–F9 and C1–C18) and ClpC1 NTD was evaluated using a high-sensitivity VP-ITC microcalorimeter and a high-sensitivity automated Auto-iTC200 microcalorimeter (MicroCal – Malvern Instruments, Malvern, UK). The protein solution in the calorimetric cell (100 mM Hepes pH 7.4, 100 mM NaCl, 2% DMSO, at 10 or 15 μM, for VP-ITC and Auto-iTC200, respectively) was titrated with compound solution (same buffer as protein, at 150 or 200 μM, for VP-ITC and Auto-iTC200, respectively). Solutions were degassed at 15 °C for 2 min before each assay. A sequence of 10 or 2 μL injections (for VP-ITC and Auto-iTC200, respectively) of titrant solution every 300 or 150 s (for VP-ITC and Auto-iTC200, respectively) was programmed and the stirring speed was set to 459 or 750 rpm (for VP-ITC and Auto-iTC200, respectively). The cell volume was 1.4534 mL or 0.2007 mL (for VP-ITC and Auto-iTC200, respectively). The association constant, Ka, and the enthalpy of binding, ΔH, were estimated through non-linear regression of the experimental data employing a single ligand binding site model (1:1 protein:compound stoichiometry) implemented in Origin (OriginLab, Northampton, MA). The background heat effect per injection (the so-called “heat of dilution”) was accounted for by introducing an adjustable parameter in the fitting routine.
NMR
Samples for NMR spectroscopy are composed of 50 mM HEPES at pH 6.5 buffer, supplemented by 50 mM NaCl in 95%/5% H2O/D2O. NMR experiments were acquired on a 0.2 mM U-[1H–15N] ClpC1-NTD sample at 298 K, on a Bruker Avance III HD spectrometer operating at a magnetic field strength corresponding to the 1H Larmor frequencies of 800 MHz equipped with a 5 mm z-gradient TCI (H/C/N) cryoprobe.
In order to determine the interaction interface on ClpC1-NTD with the F5 compound, HSQC 2D 1H–15N correlation via double inept transfer using sensitivity improvement, phase sensitive Echo/Antiecho-TPPI gradient selection with decoupling during acquisition using trim pulses in inept transfer NMR experiments were recorded in the absence and in the presence of 0.2 mM (1 eq) and 0.4 mM (2 eq) of F5. F5 was dissolved in 100% deuterated DMSO at a final concentration of 40 mM in the stock solution.
Molecular dynamics
Protein coordinates were retrieved from the Protein Data Bank with PDB 3wdc12 and F5 ligand positions were imposed from the first pose proposed by the docking calculation performed with Autodock vina through the computational virtual screening calculations.
F5 parameters and files for the molecular dynamics were prepared by the use of antechamber and parmchk2 to generate the atomic charges added by employing the AM1-BCC (assuming a spin multiplicity equal to 1 and a F5 charge to -1) and the parameter files that comprises bonded terms and VDW, using GAFF force field respectively32.
Standard protonation states were assigned to titratable residues (Asp and Glu are negatively charged; Lys and Arg are positively charged). Histidine protonation was assigned favouring formation of hydrogen bond in the initial structure. 4 Na+ were added to the system to electro neutralize it, by posing them around the protein using a coulombic potential on a grid. Protein was then solvated by a truncated octahedral box of TIP3P waters, ensuring that the distance between the biomolecule surface and the box limit was at least 10 Å.
The NTD-ClpC1-F5 complex coordinates were first optimized using a conjugate gradient algorithm for 23,000 steps, followed by 1 ns. constant volume MD equilibration during which the temperature of the system was slowly raised from 0 to 300 K. The heating was followed by a 1 ns. constant temperature and constant volume MD simulation and then by a subsequent 1 ns constant temperature and constant pressure to equilibrate the system density.
All simulations were performed with the amber package33,34,35 of programs using the ff19SB force field for all aminoacidic residues36.
Temperature was kept constant using the Langevin thermostat and pressure using the defaults Berendsen barostat with amber default coupling parameters.
All simulations were performed with periodic boundary conditions using the particle mesh Ewald summation method for long-range electrostatic interactions. The SHAKE algorithm was applied to all hydrogen-containing bonds, allowing the use of a 2 fs. time step. MDs trajectory post-processing and analysis was performed by combining cpptraj and VMD.
In particular, calculations with end point binding free energy calculation were performed by Molecular mechanics Poisson–Boltzmann and Generalized born surface area (MM-PB(SA) and MM-GB(SA) respectively), were computed with the MMPBSA.py37 included in the Amber suite package. Snapshots were extracted from the MDs trajectories every 1 ns.
Author contributions
Katharina Weinhäupl: Investigation, Writing, Visualisation; Louis Meuret: Investigation; Sandy Desrat: Investigation; Fanny Roussi: Investigation; Nelly Morellet: Investigation; Sandra Beaupierre: Investigation; Catherine Guillou: Investigation , Carine van Heijenoort Investigation; Olga Abian: Investigation; Sonia Vega: Investigation; Ian Wolf: Investigation; Tatos Akopian: Investigation; Olga Krandor: Investigation; Eric Rubin: Supervision; Adrian Velazquez-Campoy: Investigation, Supervision; Diego Gauto: Project Administration, Conceptualization, Funding acquisition, Supervision, Investigation, Methodology, Formal analysis and Hugo Fraga: Project Administration, Conceptualization, Funding acquisition, Supervision, Investigation, Methodology, Formal analysis.
Data availability
Data is provided within the manuscript or supplementary information files.
Change history
10 March 2025
A Correction to this paper has been published: https://doi.org/10.1038/s41598-025-92778-z
References
Fraga, H. et al. Development of high throughput screening methods for inhibitors of ClpC1P1P2 from Mycobacteria tuberculosis. Anal. Biochem. 567, 30–37 (2019).
Weinhäupl, K. et al. Structure of the drug target ClpC1 unfoldase in action provides insights on antibiotic mechanism of action. J. Biol. Chem. 298, 102503 (2022).
Raju, R. M. et al. Mycobacterium tuberculosis ClpP1 and ClpP2 function together in protein degradation and are required for viability in vitro and during infection. PLoS Pathog. 8, 1002511 (2012).
Akopian, T. et al. Cleavage specificity of Mycobacterium tuberculosis ClpP1P2 protease and identification of novel peptide substrates and boronate inhibitors with anti-bacterial activity. J. Biol. Chem. 290, 11008–11020 (2015).
Vasudevan, D., Rao, S. P. S. & Noble, C. G. Structural basis of mycobacterial inhibition by Cyclomarin A. J. Biol. Chem. 288, 30883–30891 (2013).
Renner, M. K. et al. Cyclomarins A-C, new antiinflammatory cyclic peptides produced by a marine bacterium (Streptomyces sp.). J. Am. Chem. Soc. 121, 11273–11276 (1999).
Schmitt, E. K. et al. The natural product cyclomarin kills mycobacterium tuberculosis by targeting the ClpC1 subunit of the caseinolytic protease. Angew. Chem. Int. Ed. 50, 5889–5891 (2011).
Weinhäupl, K. et al. The antibiotic cyclomarin blocks arginine-phosphate–induced millisecond dynamics in the N-terminal domain of ClpC1 from Mycobacterium tuberculosis. J. Biol. Chem. 293, 8379–8393 (2018).
Wolf, N. M. et al. Structure of the N-terminal domain of ClpC1 in complex with the antituberculosis natural product ecumicin reveals unique binding interactions. Acta Crystallogr. D Struct. Biol. 76, 458–471 (2020).
Gao, W. et al. The cyclic peptide ecumicin targeting CLpC1 is active against Mycobacterium tuberculosis in vivo. Antimicrob. Agents Chemother. 59, 880–889 (2015).
Gavrish, E. et al. Lassomycin, a ribosomally synthesized cyclic peptide, kills mycobacterium tuberculosis by targeting the ATP-dependent protease ClpC1P1P2. Chem. Biol. 21, 509–518 (2014).
Jagdev, M. K. et al. Crystal structure of the N-terminal domain of MtClpC1 in complex with the anti-mycobacterial natural peptide Lassomycin. Int. J. Biol. Macromol. 253, 126771 (2023).
Wolf, N. M. et al. High-resolution structure of ClpC1-rufomycin and ligand binding studies provide a framework to design and optimize anti-tuberculosis leads. ACS Infect. Dis. 5, 829–840 (2019).
Choules, M. P. et al. Residual complexity does impact organic chemistry and drug discovery: The case of rufomyazine and rufomycin. J. Organ. Chem. 83, 6664–6672 (2018).
Choules, M. P. et al. Rufomycin targets clpc1 proteolysis in mycobacterium tuberculosis and M. Abscessus. Antimicrob. Agents Chemother. 63, 18 (2019).
Hillisch, A., Heinrich, N. & Wild, H. Computational chemistry in the pharmaceutical industry: From childhood to adolescence. ChemMedChem 10, 1958–1962 (2015).
Erlanson, D. A. Learning from PAINful lessons. J. Med. Chem. 58, 2088–2090 (2015).
Bastos, M. et al. Isothermal titration calorimetry. Nat. Rev. Methods Primers 3, 17 (2023).
Litaudon, M. et al. A dimeric sesquiterpenoid from a Malaysian Meiogyne as a new inhibitor of Bcl-xL/BakBH3 domain peptide interaction. J. Nat. Prod. 72, 480–483 (2009).
Morreale, F. E. et al. BacPROTACs mediate targeted protein degradation in bacteria. Cell 185, 2338-2353.e18 (2022).
Hoi, D. M. et al. Clp-targeting BacPROTACs impair mycobacterial proteostasis and survival. Cell 186, 2176-2192.e22 (2023).
Junk, L. et al. Homo-BacPROTAC-induced degradation of ClpC1 as a strategy against drug-resistant mycobacteria. Nat. Commun. 15, 46218 (2024).
Won, H. I. et al. Chemically-induced targeted protein degradation in mycobacteria uncovers antibacterial effects and potentiates antibiotic efficacy. Biorxiv 2023, 528552 (2023).
Bhanot, A. et al. Discovery of small molecule inhibitors of Mycobacterium tuberculosis ClpC1: SAR studies and antimycobacterial evaluation. Results Chem. 5, 100904 (2023).
Lyu, J. et al. Ultra-large library docking for discovering new chemotypes. Nature 566, 224–229 (2019).
Dardenne, J., Desrat, S., Guéritte, F. & Roussi, F. Asymmetric synthesis of two analogues of meiogynin A. Eur. J. Org. Chem. 2013, 2116–2122 (2013).
Abou Samra, A. et al. Dual inhibitors of the pro-survival proteins Bcl-2 and Mcl-1 derived from natural compound meiogynin A. Eur. J. Med. Chem. 148, 26–38 (2018).
AI is a viable alternative to high throughput screening: A 318-target study. Sci. Rep. 14, 7526 (2024).
O’Boyle, N. M. et al. Open babel: An open chemical toolbox. J. Cheminform. 3 (2011).
Trott, O. & Olson, A. J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31, 455–461 (2010).
Desrat, S. et al. From meiogynin A to the synthesis of dual inhibitors of Bcl-xL and Mcl-1 anti-apoptotic proteins. Chem. Commun. 50, 8593 (2014).
Wang, J., Wang, W., Kollman, P. A. & Case, D. A. Automatic atom type and bond type perception in molecular mechanical calculations. J. Mol. Graph Model 25, 247–260 (2006).
Salomon-Ferrer, R., Götz, A. W., Poole, D., Le Grand, S. & Walker, R. C. Routine microsecond molecular dynamics simulations with AMBER on GPUs. 2. Explicit solvent particle mesh ewald. J. Chem. Theory Comput. 9, 3878–3888 (2013).
Götz, A. W. et al. Routine microsecond molecular dynamics simulations with AMBER on GPUs. 1. Generalized born. J. Chem. Theory Comput. 8, 1542–1555 (2012).
Le Grand, S., Götz, A. W. & Walker, R. C. SPFP: Speed without compromise—A mixed precision model for GPU accelerated molecular dynamics simulations. Comput. Phys. Commun. 184, 374–380 (2013).
Tian, C. et al. ff19SB: Amino-acid-specific protein backbone parameters trained against quantum mechanics energy surfaces in solution. J. Chem. Theory Comput. 16, 528–552 (2020).
Miller, B. R. et al. MMPBSA.py: An efficient program for end-state free energy calculations. J. Chem. Theory Comput. 8, 3314–3321 (2012).
Acknowledgements
This work was supported by MOSBRI, a project receiving funding from the European Union’s Horizon 2020 Research and Innovation Program under grant agreement No. 101004806 (MOSBRI-2022-94 and MOSBRI-23-162); Ministerio de Ciencia e Innovación MCIN/AEI/10.13039/501100011033/ and ‘ERDF A way of Making Europe’ [PID2021-127296OB-I00 to A.V.C.]; Fondo de Investigaciones Sanitarias from Instituto de Salud Carlos III and European Union (ERDF/ESF, ‘Investing in your future’) [PI21/00394 to O.A.]. Louis Meuret’s internship was funded as part of the France 2030 programme ANR-11-IDEX-0003, awarded by the interdisciplinary program OI LivingMachines@Work from Université Paris-Saclay. Financial support from the Research Infrastructure INFRANALYTICS (CNRS FR2054) for conducting the NMR research is gratefully acknowledged. MD simulation was performed using HPC resources from GENCI–IDRIS (Grant 2023-A0110712950)
Author information
Authors and Affiliations
Contributions
Katharina Weinhäupl: Investigation, Writing, Visualisation; Louis Meuret: Investigation; Sandy Desrat: Investigation; Fanny Roussi: Investigation; Nelly Morellet: Investigation; Sandra Beaupierre: Investigation; Catherine Guillou: Investigation , Carine van Heijenoort Investigation; Olga Abian: Investigation; Sonia Vega: Investigation; Ian Wolf: Investigation; Tatos Akopian: Investigation; Olga Krandor: Investigation; Eric Rubin: Supervision; Adrian Velazquez-Campoy: Investigation, Supervision; Diego Gauto: Project Administration, Conceptualization, Funding acquisition, Supervision, Investigation, Methodology, Formal analysis and Hugo Fraga: Project Administration, Conceptualization, Funding acquisition, Supervision, Investigation, Methodology, Formal analysis.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this Article was revised: The original version of this Article contained an error in the spelling of the author Fanny Roussi which was incorrectly given as Fanny Roussy.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Weinhäupl, K., Meuret, L., Desrat, S. et al. Identification of new ClpC1-NTD binders for Mycobacterium tuberculosis drug development. Sci Rep 15, 4146 (2025). https://doi.org/10.1038/s41598-025-87535-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-87535-1
Keywords
This article is cited by
-
Natural pigments from the endophyte Aspergillus westerdijkiae and evaluation of their bioactivities
Microbial Cell Factories (2025)