Abstract
Exophiala dermatitidis, a polyextremotolerant black yeast, thrives in diverse natural and human-made environments. Additionally, it is an opportunistic pathogen, capable of infecting immunocompromised individuals, particularly causing neurotropic infections. This study examined 41 E. dermatitidis strains from diverse environments, investigating their growth under different temperatures, NaCl concentrations, and pH levels. Optimal growth occurred at 28 °C, with large variations among strains at other temperatures, from 4 to 42 °C. Growth was enhanced at 5% NaCl, though strains also grew at 10% and 17% NaCl. Growth varied across different pH levels, from pH 2.5 to 12.5. Most strains showed the highest biofilm formation at 37 °C, α- and γ-hemolysis and resistance to antifungal agents. Better growth was detected on neurotransmitters than on (poly)aromatic compounds. High-throughput metabolic analyses revealed consistent oxidation patterns across 94 carbon sources in five selected strains. Genomic analysis revealed a diverse repertoire of carbohydrate-active enzymes and pathways for degrading polyaromatic hydrocarbons and neurotransmitters. Melanin biosynthesis inhibitor tricyclazole minimally affected E. dermatitidis growth under stress, but induced morphological changes in some cases. This study underscores E. dermatitidis’ urban extremophilic nature, with high resilience, metabolic adaptability, and potential for heightened pathogenicity in evolving global conditions.
Similar content being viewed by others
Introduction
The black yeast Exophiala dermatitidis (Ascomycota, Pezizomycotina, Eurotiomycetes, Chaetothyriales) is known for its remarkable ability to thrive in a wide range of harsh environmental conditions, demonstrating polyextremotolerance to various stressors1,2.This black yeast is an opportunistic human pathogen3, found in diverse settings such as drinking water systems, domestic environments like saunas, toilets, kitchens, steam rooms, drainpipes, and dishwashers2,4,5,6,7, as well as in extreme natural environments8,9 and human-impacted polycyclic aromatic hydrocarbon (PAH) contaminated sites10,11. In natural environments, it occurs with very low frequency, such as a few CFU –colony formation units– per several liters of water12. In contrast, in dishwashers, it can be found in extremely high numbers, reaching up to 106 cells per cm−1 of rubber seal4,6. As a polyextremotolerant microorganism, E. dermatitidis, can withstand temperatures ranging from 4 to 47 °C and thrives across a broad pH range from 2.0 to 12.0. It also shows resistance to oxidative stress, high osmolarity caused by various salts (NaCl, KCl, MgCl2, CaCl2, NaBr, MgSO4), and exposure to detergents4,9,12,13.
E. dermatitidis is a significant health risk as it can infect immunocompromised individuals, leading to a variety of infections including cutaneous, subcutaneous, systemic, gastrointestinal, pulmonary, and neurotropic conditions2,7,8,14,15,16,17,18. Its ability to grow at 37 °C and ability to synthesize toxic metabolites and produce melanized micro-vesicles enhances its pathogenic potential17,19,20. Additionally, its capacity to evade the host immune system contributes to its virulence, often resulting in severe central nervous system infections2,8,21.
E. dermatitidis is noted for its significant genomic diversity1 and unique metabolic capabilities, including the breakdown of aromatic hydrocarbons17, phenolic compounds22, and various neurotransmitters such as acetylcholine, gamma aminobutyric acid (GABA), glycine, glutamate, and dopamine, which are prevalent in the brain17. These neurotropic capabilities have contributed to its high mortality rates, particularly in healthy individuals in East Asia, where severe brain infections from strains of E. dermatitidis have been recorded2,8.
E. dermatitidis has been found in extremely nutrient-poor natural settings such as the Arctic and Antarctic regions, Alpine glaciers9, and hot springs8, as well as hydrocarbon polluted environments10,11. Notably, this black yeast thrives in oil-contaminated areas like creosote-treated railway sites and concrete near petroleum oil stains8,10. These environments often face harsh conditions such as high temperatures, acidic pH, oxygen depletion, elevated salinity, and intense ultraviolet exposure10,23. E. dermatitidis’ ability to metabolize PAHs as its sole carbon sources highlights its remarkable metabolic versatility and adaptability to challenging environments17,24.
The yeast’s polyextremotolerance is further demonstrated by its resistance to high radiation doses, extreme temperatures, osmotic stress, desiccation, and oxidative stress2,4,10,24,25,26. Although different metabolic and physiological adaptations enable E. dermatitidis’ resilience to extreme environments, they are relatively little investigated. These adaptations include melanin synthesis within its cell wall and secretion of extracellular polysaccharides for stress tolerance17,19,22,23,24,27. Additionally, E. dermatitidis exhibits polymorphism and high metabolic versatility. It is capable of utilizing diverse substrates like alcohols, sugars, neurotransmitters, and xenobiotics12,13,17,22.
Given global environmental changes, ranging from climatic shifts to increased urbanization and higher standards of indoor living—evidenced by the rising use of home appliances in currently less developed regions—E. dermatitidis represents a model organism for predicting potential evolutionary trends in microbial urban extremophiles. E. dermatitidis potentials might lead to its increased prevalence in extreme indoor habitats and polluted areas. Therefore, under the changing global conditions, particularly due to climate change and urbanization, novel pathogenic ecotypes of E. dermatitidis could potentially spread more widely through extreme environments and those impacted by human activities.
To deepen our understanding of E. dermatitidis’ ecology and adaptations to different extreme conditions, we investigated the cellular proliferation and stress responses of 41 strains of E. dermatitidis isolated globally from indoor, natural and clinical environments. We assessed their growth under various stress conditions, including changes in temperature, osmolarity, pH, presence of neurotransmitters, and exposure to (poly)aromatic compounds. High-throughput metabolic analyses using Biolog MicroPlate™ technology was employed to evaluate their metabolic adaptability. Additionally, genomic analysis of five selected strains identified their carbohydrate-active enzyme (CAZyme) profiles and metabolic pathways for PAH degradation. Our study also examined the role of melanin in enhancing E. dermatitidis’ tolerance to extreme conditions such as variations in temperature, pH, salinity, and exposure to neurotransmitters and polycyclic compounds. The aim was to define the plasticity of adaptations in E. dermatitidis strains and to highlight their evolutionary potential in an increasingly changing world.
Materials and methods
Exophiala dermatitidis strains, preservation, and culture conditions
E. dermatitidis strains used in this study were obtained from indoor environments (26 strains), natural habitats (10 strains), and clinical sources (5 strains) as detailed in Table 1 and Supplementary Fig. 1. The strains were cryopreserved at – 80 °C in 20% glycerol at the Ex Microbial Culture Collection of the Infrastructural Centre Mycosmo, Department of Biology, Biotechnical Faculty, University of Ljubljana, Slovenia. E. dermatitidis strains were routinely cultured and maintained on Malt Extract Agar (MEA) containing malt extract (20 g/L), casein peptone (1 g/L), and agar (20 g/L) at 37 °C for 7 days. For all experiments, five-day-old pre-inocula were prepared in Malt Extract Broth (MEB) at 37 °C with shaking at 150 rpm.
Tolerance to extreme temperatures, NaCl concentrations and pHs
The tolerance of E. dermatitidis strains to various temperatures (4 °C, 15 °C, 28 °C, 30 °C, 37 °C, and 42 °C), NaCl concentrations (5%, 10%, and 17%), and extreme pH values (2.5, 4, 10, and 12.5) was assessed using 96-well microplates. Each well was inoculated with 300 µL of MEB with an initial cell density of 0.1 (OD600 = 0.1). Microplates were then incubated for five days at 150 rpm. Yeast growth was monitored spectrophotometrically at 600 nm using an Epoch microplate spectrophotometer (BioTek Instruments Inc, Winooski, VT, USA). Abiotic controls were included in the experiments. The tolerance to NaCl and pH was examined at both 28 °C and 37 °C. The experiment was conducted twice with triplicate samples each time.
Biofilm formation
Biofilm formation by yeast strains was quantified using the crystal violet assay, with modifications as described by24. Briefly, cell suspensions in 0.1× MEB (OD600 = 0.2) were prepared from five-day-old pre-inoculum cultures. Nunc-TSP plates (Thermo Scientific, Waltham, MA, USA) were inoculated with 200 µL of the cell suspension and incubated statically for 48 h at both 28 °C and 37 °C. Subsequently, 100 µL of 1% (w/v) crystal violet solution was added to each well. After 30 min of incubation, the crystal violet solution was removed, and the wells were washed with deionized water. Later, 100 µL of 10% (w/v) sodium dodecyl sulfate (SDS) was added to solubilize the biofilm, and the absorbance was measured at 590 nm using an Epoch microplate spectrophotometer (BioTek Instruments Inc, Winooski, VT, USA). The experiment was performed twice with triplicate samples each time.
Hemolytic assay
The hemolytic activity of E. dermatitidis strains was assessed following the protocol outlined by28. Briefly, 5 µL of cell suspensions (OD600 = 0.1) were inoculated onto blood agar base plates (Dibico, Cuautitlán Izcalli, Mexico) supplemented with 5% defibrinated sterile bovine blood. After incubation for 7 days at both 28 °C and 37 °C, the hemolytic characteristics of the colonies were examined and classified according to29: α-hemolysis, indicated by partial hemolysis and a greenish-grey or brownish discoloration around the colonies; β-hemolysis, characterized by complete hemolysis and a clear halo around the colonies; and γ-hemolysis, where no hemolysis was observed and no change in the medium surrounding the colonies occurred. The experiment was conducted twice with triplicate samples each time.
Susceptibility to antifungal drugs
The susceptibility of yeast isolates to various antifungal agents, including terbinafine, ketoconazole, fluconazole, itraconazole, and nystatin (Merck, Darmstadt, Germany), was determined. Antifungal drugs were evaluated at different doses, encompassing both low and high concentrations (0.0313 and 16.0 µg/mL ketoconazole and itraconazole, 0.125 and 64.0 µg/mL fluconazole, 0.33 and 1.0 µg/mL terbinafine, 0.6024 and 32.0 µg/mL nystatin), to cover the spectrum of minimum inhibitory concentrations previously reported in30,31,32. In brief, Petri plates containing Müller-Hinton Agar (Biolife, Milan, Italy) supplemented with antifungal-loaded discs and sprayed with 5 µL of a cell suspension (OD600 = 0.1) were incubated for 72 h at both 28 °C and 37 °C. Based on inhibition of yeast growth, drug susceptibility was categorized as resistant, intermediate, or sensitive32. The experiment was conducted twice with triplicate samples each time.
Utilization of neurotransmitters and (poly)aromatic compounds as the sole carbon source
The metabolic capability of E. dermatitidis strains to utilize neurotransmitters such as serotonin, dopamine, norepinephrine, and epinephrine, and polycyclic aromatic compounds (phenol, 1,2,4-benzenetriol, phenanthrene, and naphthalene) as sole carbon sources was assessed using high-throughput experiments conducted in 96-well microplates. Each well contained 300 µL of mineral medium supplemented with 0.01 M of the respective tested compound, and the yeast density in each well was adjusted to 0.1 (OD600). The composition of the mineral medium was: 7.8 mg/L CuSO4.5H2O, 18 mg/L FeSO4.7H2O, 500 mg/L MgSO4.7H2O, 10 mg/L ZnSO4.7H2O, 50 mg/L KCl, 1 g/L K2HPO4, 2 g/L NH4NO3, 2 g/L KH2PO4, 100 mg/L CaCl2, 5 mg/L MnSO4, 0.1 mg/L H3BO3, 0.1 mg/L NaMoO4.2H2O and 1 mg/L CoCl2. To account for the potential influence of stored nutrients within the E. dermatitidis cells, we performed three successive washes of the yeast cells with 0.9% saline solution, followed by an 18-hour incubation in saline solution to deplete intracellular nutrients accumulated during the yeast growth in MEB medium. Then, yeast cells were recovered by centrifugation and inoculated in the growth specificity assays on aromatic substrates33,34. Microplates were incubated at 37 °C and 150 rpm. Yeast growth (OD600nm) was monitored over a five-day period using an Epoch microplate spectrophotometer (BioTek Instruments Inc, Winooski, VT, USA). Cultures grown in MEB served as controls. Yeasts were also inoculated in distilled water and mineral medium to confirm the absence of growth in the absence of substrates. Abiotic controls were included in the experiment. The experiment was conducted twice with triplicate samples each time.
High-throughput metabolic fingerprinting of selected Exophiala dermatitidis strains using YT Biolog phenotypic microarrays
We employed Phenotype MicroArrays YT (YeasT) Biolog MicroPlate™ technology (Biolog, Hayward, CA, USA) to analyze the metabolic profiles of five E. dermatitidis strains: EXF-10123, EXF-8193, EXF-13310, EXF-5586, and EXF-9187 (Table 1). These strains were selected for high-throughput metabolic assays based on specific criteria: EXF-10123 is the type strain17, and the others have been previously characterized to varying extents in different studies, providing representation from diverse environments and lifestyles4 (our unpublished results).
The YT MicroPlate™ technology encompasses 94 simultaneous oxidation and assimilation assays to generate a phenotypic fingerprinting profile of yeast metabolism. In each well of a 96-well microplate, essential nutrients and biochemicals, including amino acids, organic acids, polyalcohols, and sugars (mono-, di-, and polysaccharides), are preloaded and desiccated. Specific wells are allocated for oxidation tests, where a highly sensitive colorimetric carbon source oxidation assay is conducted using color indicators and tetrazolium violet redox dye. Conversely, assimilation wells lack redox dye and are used for assessing substrate utilization. Yeast suspensions (OD600 nm = 0.1) obtained from five-day-old pre-inocula of E. dermatitidis strains were added to YT inoculating fluid following Biolog’s protocol. In each well of the YT MicroPlate, 200 µL of YT inoculating fluid containing yeasts were added. After inoculation, YT MicroPlates were incubated at 28 °C and 100 rpm for 20 days, with spectrophotometric monitoring at 590 nm using an Epoch microplate spectrophotometer (BioTek Instruments Inc, Winooski, VT, USA) to determine oxidation or assimilation of the different tested carbon sources35. Positive oxidation reactions were identified by a purple color change due to tetrazolium violet reduction, whereas increased turbidity indicated carbon source assimilation. Negative metabolic reactions exhibited no color change in oxidation tests and no increase in turbidity in assimilation wells36. Reactions with absorbance values ≥ 0.2 were considered positive37. Non-inoculated YT MicroPlates served as negative controls, with additional control wells containing Tween 80 and water. Experiments were performed in triplicate.
Genome sequencing and analysis of selected Exophiala dermatitidis strains
Whole genomes of five E. dermatitidis strains were sequenced and analysed: EXF-10123, EXF-8193, EXF-13310, EXF-5586, and EXF-9187. Genomic DNA was extracted from biomass grown in MEB with shaking at 180 rpm and 24 °C until mid-exponential growth phase. The isolation of DNA was performed as described previously38. Biomass was frozen in liquid nitrogen and kept at – 80 °C until DNA isolation using the UltraClean Microbial DNA isolation kit (MO BIO Laboratories, Carlsbad, CA, USA). The cells were homogenised with a pestle and mortar in liquid nitrogen and with 1 min of shaking in Retsch Mixer Mill 301 (ThermoFisher Scientific, Waltham, MA, USA) at 20 Hz. The isolated DNA was treated with RNAse A (ThermoFisher Scientific, Waltham, MA, USA). The final DNA was evaluated using agarose electrophoresis and by fluorometry (Qubit; ThermoFisher Scientific, Waltham, MA, USA).
The genome sequencing was performed at Novogene Europe on Illumina NovaSeq 6000, with 2 × 150-bp libraries. The quality of the reads was checked with FastQC. The ‘bbduk’ script was used to trim any remaining adapters and 3’ ends with Phred quality score below 20 (https://jgi.doe.gov/data-and-tools/bbtools/). Reads shorter than 50 bp after trimming were discarded. The genome-sequencing reads were assembled with SPAdes 3.13.039, using the option “--careful” and other parameters left at default values. Further scaffolding was attempted with Ragtag 2.1.040, using the E. dermatitidis genome (GenBank GCA_023621285.1) as a reference. Contigs and scaffolds shorter than 500 bp were removed from the final assemblies. Genomes were annotated with Augustus 3.441 after masking of the repeats with RepeatMasker 4.0.942. Annotation parameters were optimised with the scripts provided by Augustus authors using predicted transcripts and proteins of E. dermatitidis (GenBank GCA_023621285.1) as hints. These hints were also used for the final annotation. Predicted proteomes were benchmarked with the BUSCO 4.1.143 using the default parameter values and the data set of benchmarking universal single-copy orthologs (BUSCOs) for Chaetothyriales44.
The annotated genome sequences have been submitted to GenBank and are accessible under BioProject PRJNA1110056.
CAZyme motifs were identified in the predicted proteomes of E. dermatitidis by a standalone dbCAN2 server45 using DIAMOND46, HMMer47, and Hotpep48. Only identifications made by at least two of the three programs were used for downstream analyses.
Kyoto Encyclopedia of Genes and Genomes (KEGG) annotations were performed with a stand-alone version of KofamKOALA (‘kofam_scan’ 1.3.0)49. Homologues identified as KO components of KEGG modules linked to “Xenobiotics biodegradation” were extracted, counted and visualized, as well as genes related to neurotransmitter degradation.
Exophiala dermatitidis strains’ growth under melanin inhibition and exposure to varying pH, temperature, NaCl concentrations, neurotransmitters, and polycyclic compounds
Five-day-old pre-inocula of E. dermatitidis strains were prepared in 125 mL flasks. Subsequently, 20 mL of MEB with a yeast density of 0.5 OD600 nm were incubated in 125 mL flasks at 28 °C and 150 rpm for five days. Cultures were conducted with and without the melanin inhibitor tricyclazole (0.5 mM) for individual strains50.
The effect of melanin inhibition on selected E. dermatitidis strains’ tolerance to varying pHs (2.5, 10, 12.5), temperatures (4 °C, 28 °C, 37 °C), salinities (5%, 10%, 15% NaCl), and assimilation of neurotransmitters (0.01 M dopamine and serotonin) and polyaromatic compounds (0.01 M 1,2,4 benzenetriol and phenanthrene) was investigated in 96-well microplates containing 200 µL of culture per well. Microplates were incubated at 28 °C for 7 days. The effect of melanin inhibition on pH, temperature, and salinity tolerance was assessed using MEB, while assimilation of neurotransmitters and polyaromatic compounds was tested on minimal medium. The initial cell concentration in each well was 0.1 OD600 nm.
To maintain the inhibition of melanin synthesis by tricyclazole throughout the experiment, various dosing methodologies were standardized, including an initial addition of the inhibitor followed by subsequent additions of varying concentrations. An initial dose of 0.5 mM tricyclazole followed by a subsequent dose of 0.25 mM proved effective in suppressing melanin synthesis over the course of seven days, preventing the development of pigmented cultures during the study. Thus, two doses of the melanin inhibitor were administered to the microplate cultures.
Yeast growth was monitored spectrophotometrically at 600 nm using an Epoch microplate spectrophotometer (BioTek Instruments Inc, Winooski, VT, USA). Additionally, yeast cells from cultures supplemented with the melanin inhibitor were examined using a phase-contrast microscope (AmScope T720) with an Apo100×/1.25 N.A. objective (AmScope, Irvine, CA, USA). Data acquisition was performed using an AmScope MU1000 camera for all observations.
The experiment was conducted in triplicate for rigorous assessment and reproducibility of the results.
Statistical computations
We utilized Non-parametric Kruskal-Wallis tests to assess the influence of temperature, pH, and salinity on the growth of E. dermatitidis strains. Additionally, the Conover-Iman post hoc test was employed to examine different treatment combinations serving as explanatory variables, with the growth rate as the dependent variable (p ≥ 0.05).
To analyze the growth patterns of E. dermatitidis strains under the influence of neurotransmitters and (poly)aromatic compounds, we employed the Kruskal-Wallis test together with the Conover-Iman post hoc test (p ≥ 0.05). These analyses were conducted using the agricolae R package51. Furthermore, we performed a Pearson’s correlation test to examine temporal usage trends of polyaromatic compounds and neurotransmitters. For evaluating the profiles of carbon source assimilation, a principal component analysis (PCA) was conducted using the Vegan R package52, incorporating the origin of the strains as an explanatory variable.
In the Biolog YT MicroPlate experiment, absorbance data from oxidation and assimilation assays were adjusted by subtracting the absorbance value recorded at day zero from subsequent absorbance values for each well over time. To assess the temporal progression of substrate oxidation and assimilation, a Pearson’s correlation test was conducted among all strains of E. dermatitidis utilized in this study.
To analyze the effect of inhibiting melanin in selected E. dermatitidis strains on tolerance different stress conditions we employed Paired Student t Test (p ≥ 0.05; p ≥ 0.01; p ≥ 0.001).
All statistical computations and figure preparations were performed using R version 4.2.253 in the RStudio IDE (version 2022.07.1).
Results
Polyextremotolerance of Exophiala dermatitidis strains under varied temperatures, NaCl, and pH conditions
Figure 1A–F presents heat maps and box plot visualizations depicting the specific growth rates of 41 strains of E. dermatitidis at various temperatures, NaCl concentrations, and pH values. Most E. dermatitidis strains showed adaptability across the evaluated temperature range, with optimal growth observed at 28 °C. At this temperature, specific growth rates ranged from 0.002 to 0.017 h−1 (Fig. 1B). Growth rates decreased by half when the temperature was increased to 30 °C, except for strain EXF-10195, isolated from a cystic fibrosis patient, which demonstrated maximum growth (0.012 h−1) at 30 °C. Overall, growth rates were significantly reduced at 4 °C, 15 °C, 37 °C, and 42 °C compared to 28 °C and 30 °C (Fig. 1A and B). Notably, strain EXF-5586 from a dishwasher rubber, exhibited maximum growth (0.014 h−1) at 4 °C, while strains EXF-11756, EXF-6392, and EXF-9347 from domestic environments showed maximum growth at 37 °C. Strain EXF-13310, isolated from oily water in a car mechanic workshop, exhibited its highest growth rate at 42 °C (0.07 h−1). Additionally, environmental strain EXF-8193 and indoor strains EXF-8435, EXF-9187, and EXF-11966 showed remarkable growth at 42 °C. The lowest growth rates for all strains were observed at 15 °C (Fig. 1A and B).
Cell growth of 41 strains of Exophiala dermatitidis isolated from clinical, domestic, and environmental samples across a range of temperatures (4 °C, 15 °C, 28 °C, 30 °C, 37 °C, and 42 °C), NaCl concentrations (5%, 10%, and 17%), and pH levels (2.5, 4, 10, and 12.5). Heat maps (A, C, E) illustrate the growth rates (h−1) of E. dermatitidis strains under different temperature, NaCl concentration, and pH conditions, respectively. Statistical analysis was performed using the Kruskal-Wallis and Conover-Iman tests (p ≤ 0.05) (B, D, F).
At the temperatures studied (28 °C and 37 °C), the strains showed increased growth in the presence of 5% NaCl (Fig. 1C and D). Temperature did not have a statistically significant effect on strain growth in 5% NaCl. Generally, a decrease in growth rate was observed for E. dermatitidis strains at 10% and 17% NaCl, with significant inhibition of growth at 17% NaCl observed at both temperatures (Fig. 1C and D). The growth of strains in 17% NaCl was notably influenced by temperature, with higher growth observed at 28 °C compared to 37 °C. The heat map categorizes the E. dermatitidis isolates into three primary clusters (Fig. 1C). Notably, under conditions of 5% and 10% NaCl at 28 °C and 37 °C, the growth rates of certain strains, including clinical isolates EXF-10123 and EXF-10273, indoor strains EXF-8036 and EXF-9187, and strain EXF-6986 obtained from polluted ground near a gas station, significantly increased. Among all strains evaluated, clinical strain EXF-10123 exhibited exceptionally rapid growth in 17% NaCl at 28 °C (0.009 h−1). Conversely, strains EXF-16016, EXF-6942, and EXF-8192, showed the lowest growth rates across all conditions examined (0.001 h−1, 0.002 h−1, 0.003 h−1, respectively) (Fig. 1C). These findings provide evidence of halotolerance in E. dermatitidis strains.
The E. dermatitidis strains also displayed remarkable adaptability to pH variations, thriving in highly acidic and alkaline conditions (Fig. 1E and F). Overall, the strains showed a more pronounced growth rate at pH values of 4 and 10, regardless of the temperature (28 °C or 37 °C). As depicted in Fig. 1F, many isolates of E. dermatitidis exhibited their highest specific growth rates at 28 °C across pH values of 4, 10, and 12.5. In contrast, most isolates showed their highest specific growth rate at pH 10 at 37 °C (Fig. 1E). Notably, strain EXF-9187, obtained from a dishwasher, demonstrated the highest specific growth rate at pH 2.5 and 28 °C but not at 37 °C. Conversely, the indoor isolate EXF-5772 exhibited distinct growth rate patterns at 37 °C, with the highest growth rate observed at pH 2.5 (0.01 h−1) and 10 (0.005 h−1) (Fig. 1E). These findings highlight the significant impact of temperature on the growth of E. dermatitidis strains across different pH conditions (Fig. 1E and F).
Pathogenicity-related traits and metabolic capacity for utilizing neurotransmitters and polyaromatic compounds in Exophiala dermatitidis strains
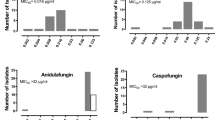
The biofilm formation, hemolytic activity, and antifungal resistance of E. dermatitidis strains were studied (Fig. 2). Most isolates showed the highest biofilm formation at 37 °C. However, strains EXF-8493 and EXF-8036 exhibited notable biofilm formation at 28 °C. The α-hemolytic phenotype was predominant among Exophiala strains, observed in 93% of strains, while a smaller proportion showed γ-hemolytic phenotypes at 28 °C and 37 °C (Fig. 2). β-hemolysis was not observed. Despite the high concentrations, the strains displayed considerable resistance to nystatin at 28 °C and 37 °C (Fig. 2). Additionally, a significant proportion of the strains showed resistance to fluconazole and itraconazole. However, yeast growth was substantially inhibited by ketoconazole and terbinafine at both temperatures.
Biofilm formation, hemolytic activity, and antifungal resistance of Exophiala dermatitidis strains grown at 28 °C and 37 °C. Antifungal drug susceptibility was tested at both low (LD*) and high (HD**) doses: (0.0313* and 16.0** µg/mL ketoconazole and itraconazole, 0.125* and 64.0** µg/mL fluconazole, 0.33* and 1.0** µg/mL terbinafine, 0.6024* and 32.0** µg/mL nystatin).
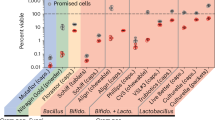
The growth response of E. dermatitidis strains was assessed using sole carbon sources, including neurotransmitters (serotonin, dopamine, norepinephrine, and epinephrine) and (poly)aromatic compounds (phenol, 1,2,4-benzenetriol, phenanthrene, and naphthalene) (Fig. 3A). Figure 3A and B indicate that E. dermatitidis strains exhibited significantly higher growth in the presence of neurotransmitters compared to (poly)aromatic compounds. Notably, most strains showed optimal growth when 1,2,4-benzenetriol was the sole carbon source, followed by serotonin (Fig. 3A and B). Strains EXF-5586 and EXF-5646 from indoor environments and strain EXF-16016 from an oil-contaminated site were independently clustered based on their growth responses to the hydrocarbons and neurotransmitters tested (Fig. 3A).
Growth of Exophiala dermatitidis strains using different neurotransmitters (serotonin, dopamine, norepinephrine, and epinephrine) and polyaromatic compounds (phenol, 1,2,4-benzenetriol, phenanthrene, and naphthalene) as sole carbon sources. Heatmap (A) illustrates the increase in absorbance (DO(λ=600 nm)) over time, indicating the growth of E. dermatitidis strains with the evaluated neurotransmitters and polyaromatic compounds. Pearson’s correlation coefficients (B, C) show significant correlations (p < 0.05) between the growth of Exophiala strainswith different neurotransmitters and aromatic compounds as sole carbon sources. Visualizing correlation with scatterplots (B) is shown. Principal component analysis (D) reveals patterns in the growth of E. dermatitidis strains under these conditions. Boxplots (E) display the distribution and mean growth values of all E. dermatitidis strains in the presence of neurotransmitters and aromatic compounds as sole carbon sources (p-value < 0.05). Density kernel distribution plots (F) visualize the principal component analysis results for E. dermatitidis strains grown with neurotransmitters and aromatic compounds as unique carbon sources.
A significant positive correlation (p < 0.01) was observed between the growth of E. dermatitidis strains with different neurotransmitters and (poly)aromatic compounds (Fig. 3C and D). Notably, the strongest positive correlations were observed with serotonin and 1,2,4-benzenetriol (r = 0.8), epinephrine and phenol (r = 0.8), epinephrine and naphthalene (r = 0.8), and norepinephrine and naphthalene (r = 0.8). Additionally, significant positive correlations (p < 0.01) were found between growth with polyaromatic compounds phenol and naphthalene (r = 0.8), as well as neurotransmitters dopamine and epinephrine (r = 0.9) used as carbon sources (Fig. 3C and D).
Principal component analysis (PCA) illustrated the correlation between E. dermatitidis growth in the presence of neurotransmitters and (poly)aromatic compounds (Fig. 3E). Three distinct clusters with comparable variances were identified. Phenol, dopamine, and epinephrine were primary contributors to PC1, while naphthalene, phenanthrene, and norepinephrine contributed significantly to PC2. The variance contributions of 1,2,4-benzenetriol and serotonin contrasted with those of the other carbon sources. A three-dimensional PCA analysis was also conducted to examine the relationship between yeast growth in various carbon sources (neurotransmitters and (poly)aromatic compounds) and the origin of E. dermatitidis strains (Fig. 3F). No discernible clustering based on isolation source was observed among E. dermatitidis isolates.
High-throughput metabolic characterization of selected Exophiala dermatitidis strains
The versatile metabolic capacity of E. dermatitidis strains EXF-5586, EXF-8193, EXF-9187, EXF-10123, and EXF-13310 was demonstrated using the Biolog MicroPlate assay (Fig. 4). These five strains were chosen for high-throughput metabolic assays based on specific criteria. EXF-10123 serves as the type strain17, while the other strains have been previously characterized to varying degrees in different studies, representing a range of environments and lifestyles4 (our unpublished results).
Metabolic characterization of selected Exophiala dermatitidis strains (EXF-5586, EXF-8193, EXF-9187, EXF-10123, and EXF-13310) using the Biolog YT phenotypic microplate methodology. Panels (A) and (E) show substrate oxidation and growth curves of the selected strains from day 0 to day 20 for the oxidation and assimilation tests, respectively. Panels (B) and (F) display correlation matrices of the oxidation and assimilation tests between the strains. Panels (C), (D), (G), and (H) present Z-scores of oxidation rate and maximum oxidation, as well as assimilation rate and maximum assimilation of substrates by the selected strains. Panels (I) and (J) depict correlations between oxidation and assimilation tests for the selected E. dermatitidis strains.
The oxidation kinetics of E. dermatitidis strains are illustrated in Fig. 4A, demonstrating consistent oxidation patterns across all evaluated substrates among the strains. This observation is further supported by statistical analysis revealing a highly significant positive correlation between the strains and their oxidation kinetics over 20 days (Fig. 4B). The Biolog results were used to develop a logistic regression model incorporating two parameters: the maximum rate of substrate oxidation (Fig. 4C) or assimilation (Fig. 4G), and the maximum observed oxidation (Fig. 4D) or assimilation (Fig. 4H). Among the tested strains, EXF-10123 (a clinical isolate) and EXF-13310 (isolated from a mechanic workshop) exhibited similar performance, displaying the highest oxidation rates across most evaluated carbon sources (Fig. 4C). Strain EXF-10123 also exhibited the highest maximum oxidation value for most evaluated carbon sources (Fig. 4D), except for monosaccharides, where strain EXF-8193 showed a higher maximum oxidation value. Notably, indoor environment-related strains EXF-5586 and EXF-9187 displayed the lowest oxidation rate values and the lowest maximum oxidation values compared to the other strains (Fig. 4C and D).
Figure 4E depicts the assimilation kinetics of various substrates by selected Exophiala strains. In contrast to the oxidation test, distinct assimilation profiles were observed for each strain, resulting in relatively low correlation coefficients (Fig. 4F). EXF-8193 and EXF-13310 exhibited similar growth rate profiles (Fig. 4G). Strain EXF-8193 demonstrated higher growth rates with polyalcohols, amino acids, and monosaccharides (Fig. 4G), aligning with its maximum growth observed with polyalcohols and amino acids (Fig. 4H). EXF-13310 exhibited higher growth rates in the presence of polysaccharides and disaccharides but slower growth with organic acids (Fig. 4G); nevertheless, it displayed high maximum growth in the presence of polysaccharides, disaccharides, and organic acids (Fig. 4H). Strain EXF-10123 exhibited low growth rates across most evaluated substrates, particularly with monosaccharides (Fig. 4G). However, strain EXF-10123 achieved high maximum growth levels with all substrates except disaccharides, indicating a slow growth rate but substantial growth potential compared to other strains (Fig. 4H).
Our findings revealed that strains EXF-8193 and EXF-13310 exhibited similar above-average oxidation and assimilation profiles across the evaluated substrates. Conversely, strains EXF-5586 and EXF-9187 demonstrated below-average similarity in their oxidation and growth rate profiles across the same substrates.
Furthermore, our results demonstrated that strain-specific ecotypes influenced the oxidation and assimilation of the tested substrates, and there was not always a direct correlation between oxidation and assimilation rates for substrates. Specifically, strain EXF-10123 exhibited the highest oxidation rates and maximum oxidation values across most evaluated substrates; however, it also displayed the lowest growth rate. Nonetheless, it showed a higher maximum assimilation value, suggesting metabolic versatility characterized by slow growth13.
We sought to explore the relationship between substrate oxidation and assimilation in selected strains of E. dermatitidis (Fig. 4I and J). Figure 4I illustrates the fitted curves and confidence intervals representing the correlation between oxidation and assimilation of various substrates for each strain. The E. dermatitidis strains were segregated into two distinct groups: one comprised of EXF-5586 and EXF-13310, and the other of EXF-10123, EXF-9187, and EXF-8193. Strains EXF-5586 and EXF-13310 exhibited lower oxidation values than the other strains, suggesting a different lifestyle.
The specific profiles of oxidation and assimilation are depicted in Fig. 4J, revealing that among 28 substrates, only a subset was utilized for oxidation and assimilation by the five E. dermatitidis strains. These included the saccharides L-sorbose, gentiobiose, D-galactose, turanose, D-melezitose, D-melibiose, and maltotriose, as well as the polyalcohols xylitol and D-sorbitol. Notably, these substrates exhibited significant correlations between oxidation and assimilation across all cases (p < 0.01, 0.6≤ r ≥ 1.0) (Fig. 4J). Strains EXF-10123 and EXF-13310 demonstrated the ability to oxidize and assimilate the highest number of substrates: 23 and 22 compounds, respectively (Fig. 4J). Interestingly, strain EXF-9187 displayed a distinct substrate utilization profile compared to the other four strains, utilizing the lowest number of substrates, likely indicative of a unique ecological niche related to nutrient availability at its isolation site (black dust from dry rubber seal in a dishwasher) (Fig. 4J).
Our results highlight the diversity among E. dermatitidis strains in substrate oxidation and assimilation profiles. While all strains oxidized the tested substrates with variations in rates and maximum oxidation, assimilation capabilities varied significantly among strains. Substrate oxidation did not always imply subsequent assimilation across all strains, and the indoor isolate strain EXF-9187 exhibited a markedly different substrate utilization profile.
Carbohydrate-active enzymes and xenobiotic and neurotransmitter metabolism in Exophiala dermatitidis genomes
Three of the sequenced genome sizes were close to 25 Mbp (EXF-8193: 26.43 Mbp, EXF-10123: 26.38 Mbp, and EXF-13310: 26.69 Mbp), while two genomes were nearly double that size (EXF-5586: 51.73 Mbp, EXF-9187: 51.74 Mbp), indicating that these two strains are diploid. The number of scaffolds was between 23 and 31 for haploid and almost 600 for diploid strains. All predicted proteomes contained over 97% of complete Chaetothyriales BUSCOs, with 88% representatives present in two copies in presumably diploid strains and over 97% present in a single copy in haploid strains.
The repertoire of CAZyme genes (Fig. 5) and genes involved in detoxification pathways of xenobiotics (Fig. 6) was studied for the five selected strains of E. dermatitidis (EXF-5586, EXF-8193, EXF-9187, EXF-10123, and EXF-13310).
Heatmap depicting the abundance of CAZy (Carbohydrate-Active Enzyme) families in selected strains of Exophiala dermatitidis (EXF-5586, EXF-9187, EXF-13310, EXF-8193, and EXF-10123). CAZy families include GH (Glycoside Hydrolases), AA (Auxiliary Activities), GT (Glycosyl Transferases), CE (Carbohydrate Esterases), PL (Polysaccharide Lyases), and CBM (Carbohydrate-Binding Modules).
(A) Genomic capabilities associated with xenobiotic metabolism, encompassing hydrocarbons and aromatic compounds, and neurotransmitter transformation within selected Exophiala dermatitidis strains. (B) Representation of the key pathways involved in the degradation of aromatic compounds such as phenol, phenanthrene, 1,2,4-benzenetriol, and naphthalene, as well as neurotransmitters including dopamine, epinephrine, and norepinephrine, with corresponding KO (KEGG Orthology) annotations in the genomes of the selected strains. (C) Cell growth kinetics for the selected strains cultivated in the presence of neurotransmitters (dopamine, epinephrine, norepinephrine, and serotonin) and aromatic compounds (phenol, phenanthrene, naphthalene, and 1,2,4-benzenetriol). Panel (A) displays the KO annotations associated with the catabolism of: Aminobenzoate (K14338: cytochrome P450; K10215: monooxygenase; K07824: benzoate 4-monooxygenase; K03380: phenol 2-monooxygenase (NADPH); K01692: enoyl-CoA hydratase; K01512: acylphosphatase; K01501: nitrilase; K01426: amidase; K01101: 4-nitrophenyl phosphatase), Atrazine (K23359: biuret amidohydrolase; K06035: cyanamide hydratase; K01941: urea carboxylase; K01429: urease subunit beta; K01428: urease subunit alpha; K01427: urease), Benzoate (K14334: carboxy-cis, cis-muconate cyclase; K07824: benzoate 4-monooxygenase; K04098: hydroxyquinol 1,2-dioxygenase; K03381: catechol 1,2-dioxygenase; K01857: 3-carboxy-cis, cis-muconate cycloisomerase; K01692: enoyl-CoA hydratase; K01607: 4-carboxymuconolactone decarboxylase; K00626: acetyl-CoA C-acetyltransferase; K00252: glutaryl-CoA dehydrogenase; K00217: maleylacetate reductase; K00074: 3-hydroxybutyryl-CoA dehydrogenase), Caprolactam (K03379: cyclohexanone monooxygenase; K01692: enoyl-CoA hydratase; K01053: gluconolactonase), Chloroalkane and chloroalkene (K13953: alcohol dehydrogenase; K08726: soluble epoxide hydrolase; K01560: 2-haloacid dehalogenase; K00148: glutathione-independent formaldehyde dehydrogenase; K00128: aldehyde dehydrogenase (NAD+); K00121: S-(hydroxymethyl)glutathione dehydrogenase), Chlorocyclohexae and chlorobenzene (K10676: 2,4-dichlorophenol 6-monooxygenase; K06912: alpha-ketoglutarate-dependent 2,4-dichlorophenoxyacetate dioxygenase; K04098: hydroxyquinol 1,2-dioxygenase; K03381: catechol 1,2-dioxygenase; K01560: 2-haloacid dehalogenase; K01061: carboxymethylenebutenolidase; K00217: maleylacetate reductase), Dioxin (K00480: salicylate hydroxylase), Drug metabolism (K13953: alcohol dehydrogenase; K13299: glutathione S-transferase kappa 1; K00799: glutathione S-transferase; K00485: dimethylaniline monooxygenase (N-oxide forming); K00274: monoamine oxidase; K00121: S-(hydroxymethyl)glutathione dehydrogenase), Drug metabolism-other enzymes (K13800: UMP-CMP kinase; K10808: ribonucleoside-diphosphate reductase subunit M2; K10807: ribonucleoside-diphosphate reductase subunit M; K03782: catalase-peroxidase; K01951: GMP synthase (glutamine-hydrolysing); K01520: dUTP diphosphatase; K01519: XTP/dITP diphosphohydrolase; K01489: cytidine deaminase; K00940: nucleoside-diphosphate kinase; K00876: uridine kinase; K00799: glutathione S-transferase; K00106: xanthine dehydrogenase; K00088: IMP dehydrogenase), Fluorobenzoate (K03381: catechol 1,2-dioxygenase; K01061: carboxymethylenebutenolidase; K00217: maleylacetate reductase), Metabolism of xenobiotics (K15303: aflatoxin B1 aldehyde reductase; K13953: alcohol dehydrogenase; K13299: glutathione S-transferase kappa 1; K00799: glutathione S-transferase; K00121: S-(hydroxymethyl)glutathione dehydrogenase; K00079: carbonyl reductase 1; K00078: dihydrodiol dehydrogenase), Naphthalene (K13953: alcohol dehydrogenase; K00480: salicylate hydroxylase; K00121: S-(hydroxymethyl)glutathione dehydrogenase), PAHs (K00480: salicylate hydroxylase), Steroid (K03333: cholesterol oxidase), Styrene (K10438: 3-hydroxyphenylacetate 6-hydroxylase; K10437: phenylacetate 2-hydroxylase; K01800: maleylacetoacetate isomerase; K01555: fumarylacetoacetase; K01501: nitrilase; K01426: amidase; K00451: homogentisate 1,2-dioxygenase), Toluene (K05797: 4-cresol dehydrogenase (hydroxylating) flavoprotein subunit; K03381: catechol 1,2-dioxygenase; K03380: phenol 2-monooxygenase (NADPH); K01061: carboxymethylenebutenolidase; K00217: maleylacetate reductase; K00055: aryl-alcohol dehydrogenase), Xylene (K00055: aryl-alcohol dehydrogenase), Tyrosine (K00545: catechol O-methyltransferase; K00274: monoamine oxidase). Neurotransmitter degradation (K14338: cytochrome P450; K00669: arylalkylamine N-acetyltransferase; K00545: catechol O-methyltransferase; K00463: indoleamine 2,3-dioxygenase; K00274: monoamine oxidase).
The abundance of CAZyme genes in the E. dermatitidis strains corresponded to the genome size. The number of genes in EXF-5586 and EXF-9187 was nearly double that of the other strains. To compensate for this presumable difference in ploidy (as discussed below), the gene numbers used in downstream analyses were divided by two for strains EXF-5586 and EXF-9187 (Fig. 5). As depicted in Fig. 5, across all strains, the most abundant CAZyme genes were those encoding some families of glycosyl hydrolases (GH), glycosyl transferases (GT), and auxiliary activity enzymes (AA). Genes encoding carbohydrate esterases (CE), carbohydrate-binding modules (CBM), and polysaccharide lyases (PL) were less represented. The glycosyl transferase family 2 (GT2) was found to be the most prevalent among all genomes of E. dermatitidis, followed by glycosyl transferase family 2 (GT2), glycoside hydrolase family 16 (GH16), glycoside hydrolase family 13 (GH13), auxiliary activity family 1 (AA1), glycosyltransferase family 4 (GT4), auxiliary activity family 3 (AA3), glycoside hydrolase family 5 (GH5), and glycosyltransferase family 22 (GT22). The distribution of these enzymes in the CAZyme profiles of E. dermatitidis strains was generally similar in all genomes. Nevertheless, based on their CAZyme gene content, the strains grouped into two distinct clusters: one comprising EXF-5586 and EXF-9187, both isolated from household dishwashers, and another group consisting of EXF-10123, EXF-13310, and EXF-8193.
E. dermatitidis strains exhibited a broad diversity of genes associated with 17 biochemical pathways of xenobiotic degradation, including catabolism for PAH depletion (Fig. 6A). Although the selected E. dermatitidis strains showed similar copy numbers for most genes related to PAH degradation, indicating a high degree of conservation in these pathways, two groups of genes contained a high number of representatives in all strains: glutathione S-transferases (K00799) and K01426 amidases were notably enriched. These enzymes play crucial roles in several of the studied detoxification pathways. Genes for carbonyl reductase 1 (K00079) were found only in two of the five strains.
Given our interest in investigating the metabolism of neurotransmitters and hydrocarbons by Exophiala strains, we identified potential metabolic pathways for the catabolism of these compounds by annotating genes involved in these metabolic pathways within the studied genomes (Fig. 6B). We identified key genes responsible for the breakdown of aromatic compounds such as phenol, phenanthrene, 1,2,4-benzenetriol, and naphthalene, along with genes involved in the biodegradative pathways of neurotransmitters like dopamine, epinephrine, and norepinephrine. For instance, the genomes of E. dermatitidis contain a complete repertory of cytochrome P450 genes encoding various oxidoreductases that act on donors with CH-OH and CH-CH groups, as well as transferases. These include carbonyl reductase, aldehyde reductase, glutathione dehydrogenase, glutathione S-transferase, dihydrodiol dehydrogenase, alcohol dehydrogenase, hydroxy-1,2-dihydronaphthalene glutathione-lyase, 4,5-dihydro-4-hydroxy-5-S-glutathionyl-benzo[a]pyrene glutathione-lyase, and 7,8-dihydro-7-hydroxy-8-S-glutathionyl-benzo[a]pyrene hydrolase. Additionally, other genes encoding enzymes such as hydroxyquinol 1,2-dioxygenase (K04098) and salicylate hydroxylase (K00480) that play direct roles in the biotransformation of aromatic hydrocarbons were also identified54,55. These enzymes are involved in the synthesis and cleavage of catechol, catechol derivatives, and various diols and dihydroxy-compounds, which are critical intermediates in the biotransformation of PAHs56,57. In addition, genes encoding K00545 catechol O-methyltransferase, which facilitates the initial biochemical step in neurotransmitter biotransformation by transferring a methyl group, and K00274 monoamine oxidase, also known as epinephrine oxidase or serotonin deaminase, responsible for the oxidative deamination of neurotransmitters and biogenic amines, are recognized as key genes in neurotransmitter catabolism58,59. These genes were identified within the studied E. dermatitidis genomes (Fig. 6B).
To corroborate the predicted genomic potential for these strains, we studied the cell growth kinetics of the five selected strains when cultivated in neurotransmitters and (poly)aromatic compounds (Fig. 6C). Despite sharing the same genetic background, notable differences in cell growth on these diverse aromatic substrates were observed among the different strains of E. dermatitidis. For instance, the strains exhibited distinct growth patterns in response to various neurotransmitters. While serotonin supported discrete growth, other neurotransmitters were more conducive to the growth of black yeast strains. The clinically derived strain EXF-10123 demonstrated its best growth in epinephrine but not in norepinephrine, despite their high structural similarity. Strains EXF-5586 and EXF-8193 displayed exceptional growth in the presence of dopamine, with EXF-8193 also exhibiting notable growth in norepinephrine.
Regarding aromatic compounds, phenol and naphthalene were the least supportive substrates for strain growth, while 1,2,4-benzenetriol was identified as the most favorable substrate for yeast growth. Interestingly, although structurally similar, with only one hydroxyl group difference, EXF-5586 showed robust growth in dopamine and minimal growth in benzenetriol. Similarly, EXF-8193 achieved comparable growth levels on these substrates, albeit with different kinetics. These results demonstrate the remarkable phenotypic plasticity of E. dermatitidis strains.
Melanin inhibition and growth responses of Exophiala dermatitidis under varying pHs, temperatures, NaCl levels, neurotransmitters, and polycyclic compounds
The impact of tricyclazole-mediated chemical inhibition of melanin biosynthesis on the cell growth of E. dermatitidis strains (EXF-5586, EXF-8193, EXF-9187, EXF-10123, and EXF-13310) under various stresses was comprehensively evaluated. Additionally, the cell growth of these strains was also assessed in the presence of tricyclazole using neurotransmitters (dopamine and serotonin) and polyaromatic hydrocarbons (1,2,4-benzenetriol and phenanthrene) as the sole carbon source (Fig. 7). Overall, melanin inhibition did not significantly affect the growth of E. dermatitidis strains under most tested conditions, including high salinity (15% NaCl), cold temperature (4 °C), extreme acidic pH (2.5), and highly alkaline pHs (10.0 and 12.5) (Fig. 7A). However, the melanin inhibitor negatively impacted the cell growth of strain EXF-13310 at 37 °C. Tricyclazole also slightly inhibited the psychrotolerant growth of EXF-10123 at 4 °C. In general, no statistically significant differences were observed in yeast growth under conditions of melanin inhibition. However, some differences were observed; they were minor in magnitude.
Effect of the melanin inhibitor tricyclazole on the growth of selected Exophiala dermatitidis strains (EXF-5586, EXF-8193, EXF-9187, EXF-10123, and EXF-13310) was investigated under various culture conditions. (A) Boxplots depict the growth of E. dermatitidis strains at different temperatures (4 °C, 28 °C, and 37 °C), NaCl concentrations (5%, 10%, and 15%), extreme pH levels (2.5, 10, and 12.5), and in the presence of 0.01 M of 1,2,4-benzenetriol (1,2,4-BT), dopamine, serotonin, and phenanthrene. Asterisks denote statically significant differences as determined by Paired Student t Test (p ≥ 0.05; p ≥ 0.01; p ≥ 0.001). (B) Microscopic observations illustrate E. dermatitidis growth on 1,2,4-benzenetriol, dopamine, serotonin, and phenanthrene in the presence of the melanin inhibitor tricyclazole.
Regarding salinity, statistically significant growth differences were noted when melanin biosynthesis was inhibited at 5%, 10%, and 15% NaCl (Fig. 7A). Specifically, at 5% NaCl, the cell growth of strains EXF-13310 was significantly reduced in the presence of tricyclazole. Similarly, at 10% NaCl, the cell growth of EXF-13310 and EXF-5586 was decreased considerably. At 15% NaCl, only the growth of the glacier-derived strain EXF-8193 was impacted when melanin biosynthesis was inhibited. Additionally, the melanin inhibitor significantly decreased the growth of certain yeast strains at both acid and alkaline pHs (Fig. 7A). For example, the growth of strains EXF-8193 and EXF-13310 at pH 2.5 and 12.5 was markedly inhibited by tricyclazole. Strain EXF-13310 exhibited reduced growth across all tested pH values. The extremotolerant growth of E. dermatitidis strain EXF-13310 was reduced when melanin synthesis was inhibited (Fig. 7A), suggesting that melanin plays a crucial role in conferring resistance to various types of stress in this strain.
Melanin inhibition did not affect yeast growth in the presence of dopamine, serotonin, or phenanthrene (Fig. 7A). However, tricyclazole significantly increased the growth of EXF-8193 and EXF-9187 when grown on 1,2,4-benzenetriol. No significant differences were observed in the biofilm formation of the five selected E. dermatitidis strains when grown with and without the melanin biosynthesis inhibitor (Fig. 7A).
Finally, selected E. dermatitidis strains were examined under a microscope to assess their growth in the presence and absence of tricyclazole. Figure 7B illustrates significant changes in their micromorphology when the melanin inhibitor was added. Only three strains (EXF-8193, EXF-10123, and EXF-13310) revealed distinct micromorphological differences in their growth with and without tricyclazole (Fig. 7B). Strains EXF-8193 exhibited pseudohyphae formation when grown with 0.01 M of 1,2,4-benzenetriol and phenanthrene without tricyclazole. Similarly, pseudohyphae formation was observed for EXF-10123 in the presence of 1,2,4-benzenetriol without tricyclazole.
In contrast, cultures of these strains with tricyclazole showed mainly isolated cells grouped in clusters. Also, strains EXF-10123 and EXF-13310 formed pseudohyphae when grown in 0.01 M of dopamine and serotonin without tricyclazole, whereas cultures with tricyclazole showed isolated cells (Fig. 7B). These results suggest that melanin inhibition induces a transition from pseudohyphal to unicellular growth.
Discussion
E. dermatitidis is known to inhabit various extreme environments, often of human origins, such as dishwashers, saunas, and bathrooms7,60. While the ecology of E. dermatitidis has been partially characterized for some isolates, there remains a limited understanding of the ecological traits across a diverse collection of this species, particularly considering strains isolated from varied environments.
In this study, we aimed to enhance our understanding of E. dermatitidis ecology and adaptation through comprehensive investigations into cellular proliferation and stress responses of 41 E. dermatitidis strains isolated from indoor and natural environments and clinical samples. We systematically evaluated their growth under diverse stress conditions, including temperature, osmolarity, and pH variations, and in the presence of neurotransmitters and (poly)aromatic compounds. High-throughput metabolic analyses using Biolog YT MicroPlate™ technology were employed to assess their metabolic adaptability. Additionally, genomic analysis of five selected strains was conducted to identify carbohydrate-active enzyme (CAZyme) repertoires and metabolic pathways involved in the degradation of PAHs and neurotransmitters. Furthermore, our study explored into the role of melanin in E. dermatitis tolerance to extreme conditions such as temperature, pH, and salinity, along with exposure to neurotransmitters and polycyclic aromatic compounds.
Exophiala dermatitidis: a polyextremotolerant black yeast with versatile physiology and metabolism
E. dermatitidis exhibits a broad thermotolerance range from 4 to 47 °C and even at 60 °C in mixed bacterial fungal biofilms6,7,60, enabling it to survive in diverse conditions, including the human body. Studies have primarily focused on its growth at 37 °C due to its clinical significance as an opportunistic pathogen17,61. Optimal growth for most strains occurred at 28 °C, although they adapted well across a wide temperature range (4–42 °C). This adaptability is crucial for survival in habitats with temperature fluctuations, from dishwashers to glaciers7. Gene expression adjustments in response to temperature shifts –such as changes in lipid production and cytoskeleton arrangement at lower temperatures and increased Golgi apparatus activity at higher temperatures– contribute to E. dermatitis thermotolerance60.
This study also confirmed E. dermatitidis’ halotolerance, with most strains capable of growing in NaCl concentrations from 5 to 10%. Growth, however, was significantly inhibited at 37 °C compared to 28 °C when exposed to higher (10% and 17%) NaCl concentrations. E. dermatitidis can tolerate high concentrations of various salts, growing in conditions of up to 3 M NaCl, 2.5 M KCl, 0.75 M MgCl2, 0.5 M CaCl2, 0.75 M NaBr, or 3 M MgSO4, which is particularly beneficial for strains isolated from environments like bath facilities and dishwashers that periodically encounter high salt levels4,12.
E. dermatitidis also exhibited a high tolerance for a broad range of pH levels (2.5–12.5), with a preference for alkaline conditions; pH 10 was optimal for the growth of most strains. However, certain strains (EXF-9187 and EXF-5772) have shown peak growth at unusual values for this species, such as pH 2.5 or 4. The ability to thrive in alkaline environments is advantageous for strains residing in indoor environments4, where exposure to alkaline detergents is common. On the other hand, for most E. dermatitidis strains grown at 28 °C, the most favorable growth was within pH 4 to 12.5. At 37 °C, the optimal pH decreased to pH 10. Wang and Szaniszlo62 also observed temperature-dependent variations in pH tolerance affecting the growth of E. dermatitidis, noting that the fungus grew better at pH 3.5, 4, and 8 at 37 °C compared to 25 °C.
Our findings suggest that E. dermatitidis could serve as a versatile model for studying pH tolerance in eukaryotic organisms. Little is known about the regulation of the Pal/Rim pathway in this black yeast, which is a key adaptive mechanism involved in alkalotolerance in Aspergillus nidulans and Saccharomyces cerevisiae – fungi where gene regulation by pH has been extensively investigated63,64. It has been suggested that alkalotolerance adaptation in E. dermatitidis is regulated by PacC at transcriptional level62. Chen et al.65 observed that the primary cellular responses to acidity in E. dermatitidis included strengthening the cell wall, slowing down metabolism, and pumping out cations. Further studies are needed to fully understand the expression profile and regulation of alkaline-expressed genes to comprehend the high adaptation of E. dermatitidis to pH variations.
To explore the phenotypic versatility of E. dermatitidis, we investigated various characteristics related to its pathogenicity, given its role as an opportunistic pathogen responsible for phaeohyphomycosis, central nervous system infections, and respiratory tract infections in individuals with cystic fibrosis2. Biofilm formation by E. dermatitidis, characterized by exopolysaccharide production, hinders infection elimination and leads to chronic recurring infections. This trait has also been linked to thermotolerance, drug resistance, and immune evasion2,8,66. In this study, all evaluated E. dermatitidis strains demonstrated biofilm production. Strains isolated from wet surfaces (EXF-8493 and EXF-8036), classified as belonging to genotype A –the pathogen phenotypic cluster2 – showed strong biofilm formation. Contrary to previous studies suggesting a correlation between biofilm formation and specific environmental conditions67, our findings suggest that biofilm formation is influenced by temperature, with most strains showing increased production at 37 °C. This underscores a potential relationship between biofilm formation and E. dermatitidis pathogenicity, even in strains isolated from outdoor environments.
Hemolysin production is another mechanism involved in pathogenicity, where certain fungi secrete proteins capable of lysing red blood cells68. Research on the hemolytic capabilities of E. dermatitidis is limited. In a prior study, Seneviratne et al.69 reported hemolysis of sheep blood agar by a clinical isolate of E. dermatitidis. However, Sav et al.67 observed that hemolytic activity was absent in environmental E. dermatitidis strains. In the current study, the hemolytic activity of E. dermatitidis strains predominantly exhibited α-hemolysis at both tested temperatures (28 °C and 37 °C). Environmental and indoor strains exhibited similar hemolytic potential to those isolated from clinical samples.
Limited data are available regarding the susceptibility patterns of E. dermatitidis to antifungal agents despite its clinical significance2,70. Kirchhoff et al.2 previously observed the sensitivity of E. dermatitidis strains to amphotericin B, itraconazole, voriconazole, flucytosine, and terbinafine. In our study, we assessed the susceptibility of E. dermatitidis strains to four ergosterol biosynthesis inhibitors (terbinafine, ketoconazole, itraconazole, and fluconazole), as well as the pore-forming antifungal nystatin. Our findings indicated that E. dermatitidis strains were susceptible to terbinafine and ketoconazole at 37 °C but showed reduced sensitivity to fluconazole and itraconazole. Notably, most strains exhibited resistance to nystatin, possibly due to differences in the fungal membrane sterol composition that affect nystatin’s efficacy71,72,73.
In summary, the opportunistic pathogen E. dermatitidis exhibits a range of traits contributing to its ecological versatility and pathogenicity. This black yeast is particularly notable for its ability to thrive at 37 °C, a critical factor for colonization and infection in warm-blooded hosts, including humans. Additionally, E. dermatitidis demonstrates hemolytic activity, associated with tissue invasion and damage, and biofilm production, enhancing its persistence and resistance to host defenses and antifungal treatments. The species also shows resistance to various antifungal agents, aiding in its evasion of therapeutic interventions. These phenotypic characteristics underscore the pathogenic potential of E. dermatitidis and highlight its adaptability to diverse environmental conditions, positioning it as an emerging pathogen of concern in the context of global environmental changes.
Understanding the ecological and pathogenic phenotypes of E. dermatitidis is crucial for predicting its spread from extreme environments to new habitats and elucidating its pathogenic mechanisms, particularly in the context of increasingly aggressive mycoses. This knowledge is essential for developing effective strategies to manage and prevent fungal diseases caused by this adaptable pathogen.
The phenotypic plasticity of strains of E. dermatitidis is highlighted by their ability to utilize both (poly)aromatic compounds and neurotransmitters as sole carbon sources, with neurotransmitter metabolism playing a significant role in their pathogenicity17. The transformation of specific hydrocarbons has been linked to fungal pathogenicity. For example, Prenafeta-Boldú et al.23 suggested that the ability to metabolize alkylbenzenes and phenols could enhance fungal pathogenicity within the family Herpotrichiellaceae, which includes the genus Exophiala. This study proposed a connection between neurotropism and hydrocarbon degradation among pathogenic fungi. Certain neurotrophic fungi, such as Aspergillus fumigatus, Candida albicans, Cladophialophora spp., and Pseudallescheria boydii, have shown the ability to assimilate hydrocarbons or phenols as sole carbon sources23.
Our findings illustrate the significant capacity of the black yeast E. dermatitidis to assimilate (poly)aromatic compounds, including neurotransmitter-like catecholamines such as dopamine, norepinephrine, epinephrine, and serotonin10,17,23. This suggests that various strains of E. dermatitidis, including those from indoor environments, have the potential to induce systemic mycoses through neurotransmitter degradation. This could lead to the emergence of virulent ecotypes capable of causing new neuropathies when transitioning from natural and indoor environments to human hosts17,23. Zajc et al.24. and Lavrin et al.17 observed that diverse strains of E. dermatitidis, including clinical, environmental, and indoor isolates, were proficient in utilizing toluene, n-hexadecane, and mineral oil as sole carbon sources. Our study consistently found that most E. dermatitidis strains thrived on phenol, naphthalene, and phenanthrene. Notably, 1,2,4-benzenetriol, a byproduct of benzene metabolism74 and a close structural analogue of dopamine75, was exceptionally suitable as a substrate for a broad range of E. dermatitidis strains. The assimilation of these carbon sources supports the hypothesis that neurotransmitter assimilation may be linked with PAH metabolism. Indeed, as shown in Fig. 3B and C, a strong correlation was observed between 1,2,4-benzenetriol and serotonin assimilation.
The dual biodegradation capabilities of E. dermatitidis and related black yeasts, which enable them to metabolize both PAHs and neurotransmitters like serotonin and dopamine, represent a significant study area with broad implications. This ability highlights their adaptive strategies, allowing them to survive and thrive in diverse habitats – from indoor environments to potentially within hosts during infections. The metabolic versatility of E. dermatitidis underscores its potential pathogenicity and capability to persist in various ecological niches. Elucidating these metabolic pathways can deepen our knowledge of fungal physiology, ecology, and practical applications, such as developing bioremediation strategies for environmental cleanup and targeting these pathways for novel antifungal therapies. Therefore, exploring the biodegradative processes of Exophiala spp. is crucial for advancing both fundamental fungal biology and applied sciences in environmental and medical fields.
E. dermatitidis strains demonstrated broad metabolic versatility, capable of metabolizing a diverse array of carbohydrates, including D-glucose, D-galactose, D-xylose, L-arabinose, sucrose, maltose, α-trehalose, cellobiose, salicin, raffinose, and melezitose. This reflects their ability to adapt their metabolism under challenging environmental conditions23. Additionally, these strains showed growth on alcohols such as glycerol, mesoerythritol, xylitol, D-glucitol, and ethanol, as well as ethylamine, lysine, and cadaverine, similar to clinical Exophiala isolates12. This metabolic plasticity confirms their ability to utilize a broad range of saccharides, primarily monosaccharides, organic acids, alcohols, and amino acids, essential for “invasive microorganisms” like E. dermatitidis to locate and utilize nutrient sources within their natural habitats76.
Significant differences in the assimilation of various carbon sources among the five strains highlighted their considerable metabolic diversity. Strains EXF-5586 and EXF-9187, isolated from indoor environments, exhibited the lowest consumption rates and cell growth across most assessed carbon sources (Fig. 4). Conversely, strains EXF-13310 and EXF-10123 displayed the highest consumption rates and cell growth with most of the carbon sources tested (Fig. 4).
The observed metabolic diversity across different strains of the black yeast E. dermatitidis has significant implications for microbial evolution, phenotypic plasticity, and adaptive fitness, particularly in global environmental changes. The ability of E. dermatitidis to assimilate and grow using a wide array of carbon sources, including various sugars, alcohols, and amino acids, underscores its metabolic versatility and capacity to adapt to diverse environmental conditions12,23. This metabolic flexibility is crucial for the survival and proliferation of E. dermatitidis in changing habitats, enabling it to exploit available nutrients and occupy new ecological niches. Such metabolic adaptations not only indicate phenotypic plasticity but also contribute to the pathogenic potential of E. dermatitidis, as it thrives in various environments and can potentially cause infections in diverse host organisms. The spread of E. dermatitidis to new habitats and its success as an opportunistic pathogen in the era of global change underscores the importance of understanding its metabolic diversity and adaptive strategies. This knowledge can inform environmental monitoring, disease prevention, and the development of targeted therapeutic interventions. By elucidating the metabolic underpinnings of E. dermatitidis and its strain-specific variations, we gain insights into the evolutionary dynamics and ecological fitness of this yeast species amidst ongoing environmental shifts.
The genomes of Exophiala dermatitidis showcase a broad repertoire of genes encoding enzymes for carbohydrate degradation, (poly)aromatic compound metabolism, and neurotransmitter processing
Based on the presence of single-copy orthologues in Chaetothyriales, all five genomes were annotated to a high level of completeness. The two presumably diploid strains EXF-5586 and EXF-9187 were isolated from dishwashers several years apart (Table 1). Although typically haploid, some species of melanized yeasts have previously been reported to harbor diploids13. Nevertheless, with only two presumable diploids known in E. dermatitidis and a relatively small number of its genomes sequenced, the sample size is, for now, too small to estimate the variability of ploidy within the species. Similarly, it is too early to speculate on potential links between different ploidies and specific habitats, such as the dishwasher interior, which seem particularly favorable for E. dermatitidis4.
The genomes displayed an extensive array of genes for CAZymes and for degrading (poly)aromatic compounds and neurotransmitters (Figs. 5 and 6A and B), illustrating the fungus’s capacity to interact with and utilize various complex carbon sources. This genetic diversity points to E. dermatitidis’ adaptability to diverse ecological niches and its potential pathogenicity in environmental and clinical contexts. Regarding metabolic adaptability, the diversity of CAZyme families is crucial, supporting growth in diverse ecological niches72,77,78. The identified CAZymes in these E. dermatitidis strains corroborate our findings from Biolog phenotyping (Fig. 4), where glycoside hydrolases (GH), glycosyl transferases (GT), and auxiliary activities (AA) were the most prevalent (Fig. 5).
These strains demonstrated a diverse array of CAZymes and metabolic enzymes that facilitate the degradation of complex compounds, such as (poly)aromatic compounds and neurotransmitters. Notably, the enzyme families like glycoside hydrolases (GH13), transglycosylases (GH16), flavoproteins with FAD-binding domains (AA3), and oxidoreductases (AA1), including laccases, provide a metabolic toolkit for the biotransformation and assimilation of complex carbon sources, such as phenolic compounds79,80,81.
The families of CAZyme encoding genes and the numbers of their representatives were broadly similar between the strains. This is not surprising, given the close relatedness between the strains and the increasingly common observation that the CAZyme content is mainly shaped by phylogenetic history rather than habitat requirements82. Nevertheless, based on the CAZyme gene content, two distinct clusters formed: one comprising the diploid strains EXF-5586 and EXF-9187 isolated from household dishwashers and another consisting of EXF-10123, EXF-13310, and EXF-8193. This clustering might suggest specific enzymatic capabilities selected by the dishwasher habitat or indicate evolutionary differences between diploid and haploid strains. The difference might also be an artifact of the genome ploidy since diploid strains present a much greater challenge for genome assembly. This is illustrated by the fragmented nature of the genome assemblies of diploid strains, which contain approximately 20-times as many scaffolds than haploid strains. The identification of genes associated with pathways for the degradation of monoaromatic hydrocarbons and other volatile pollutants further underscores the metabolic versatility of E. dermatitidis strains. These genes are involved in diverse metabolic processes, including the degradation of benzoate, PAHs, and other xenobiotics, which are critical for fungal adaptation and survival in different ecological niches83.
Specifically, genomic analysis of five E. dermatitidis strains revealed genes encoding enzymes such as catechol o-methyltransferase, monoamine oxidase, catechol 1,2-dioxygenase, salicylate hydroxylase, and hydroxyquinol 1,2-dioxygenase. These enzymes are involved in degrading polyaromatic compounds and neurotransmitters, which can be utilized as carbon sources for cell growth (Fig. 6B). The consistent presence of these genes across all strains suggests a shared capability to utilize PAHs and neurotransmitters for growth. Differences in cell growth observed among strains (Fig. 6C) may be attributed to other cellular responses, such as transcriptional regulation. These findings highlight the metabolic versatility and adaptive strategies of E. dermatitidis strains, contributing to their pathogenesis and ecological success in diverse environments.
Melanin-mediated responses to polyextremotolerant growth and utilization of polyaromatic compounds and neurotransmitters
Melanin, a pigment widely distributed among fungi84, has emerged as a crucial player in facilitating polyextremotolerance – a trait allowing fungi to thrive in extreme environments characterized by high levels of stressors such as UV radiation, desiccation, and oxidative stress85,86. The protective role of melanin is versatile, contributing significantly to fungal adaptation and survival under adverse conditions. Melanin acts as a potent antioxidant, scavenging free radicals generated by UV exposure and other stressors, thereby shielding cellular components from oxidative damage86. Additionally, melanin absorbs and dissipates harmful UV radiation, preventing DNA damage and preserving cellular integrity87. Furthermore, melanin confers resistance against desiccation by enhancing cell wall rigidity and reducing water loss, thus enabling fungi to endure dry conditions prevalent in arid environments50. The adaptive significance of melanin in polyextremotolerance is underlined by its correlation with the ecological distribution of melanized fungi in extreme habitats such as hot springs, deep-sea vents, salterns, and polar regions8,86,88,89. Understanding the mechanistic reinforcements of melanin-mediated polyextremotolerance is essential for elucidating fungal ecological dynamics and has implications for biotechnological applications and fungal pathogenicity research.
Three distinct pathways are involved in melanin production: the DHN-melanin pathway, the DOPA-melanin pathway, and the L-tyrosine degradation pathway65. In E. dermatitidis, although genes associated with all three biosynthetic pathways are present, melanin is predominantly synthesized through the DHN-melanin pathway65. Within the DHN-melanin pathway, melanin synthesis occurs from 1,8-dihydroxynaphthalene (1,8-DHN) catalyzed by a polyketide synthase enzyme. Notably, this pathway can be inhibited by tricyclazole (5-methyl-1,2,4-triazolo[3,4-b] benzothiazole), a fungicide targeting melanin biosynthesis2,65.
To assess the potential role of melanin in polyextremotolerance among the five selected E. dermatitidis strains, we evaluated their response to various stresses following the addition of tricyclazole. The growth of E. dermatitidis strains was not notably affected by diverse stressors, including temperature (4 °C, 28 °C, and 37 °C), NaCl concentrations (5%, 10%, and 15%), pH extremes (2.5, 10.0, and 12.5), and the presence of 1,2,4-benzenotriol, dopamine, serotonin, and phenanthrene when melanin was inhibited (Fig. 7). The varying impact of tricyclazole on yeast growth under different stress conditions among strains highlights their intraspecies diversity in responding to environmental perturbations. Specifically, strain EXF-13310 exhibited decreased cell growth at 37 °C in the presence of the inhibitor. Additionally, the growth of EXF-13310 was adversely affected by tricyclazole under different NaCl concentrations, such as 5% and 10%, and also in the presence of extreme pH values (2.5, 10.0, and 12.5). Also, the growth of strains EXF-5586 and EXF-8193 was reduced under hypersaline conditions (10% and 15%, respectively) when melanin synthesis was inhibited. Overall, melanin did not play a significant role at 15% NaCl, as only one strain was affected by tricyclazole under this condition. These findings suggest that melanin plays a protective role in enabling survival under specific abiotic stresses, consistent with previous studies2. Notably, strain EXF-13310 displayed sensitivity to tricyclazole across multiple stress parameters (temperature, NaCl concentrations, and pH), implying a potential link between melanin biosynthesis and other stress response mechanisms, such as the involvement of the PacC transcription factor in adaptation to alkaline pH and high NaCl concentrations62. In contrast, strains EXF-10123 and EXF-9187 showed no significant impact on cell growth in the presence of the inhibitor under the evaluated stress conditions (Fig. 7).
In cultures supplemented with neurotransmitters and polyaromatic hydrocarbons, specifically with 1,2,4-benzenotriol as the sole carbon source, the addition of tricyclazole resulted in increased cell growth (measured by optical density) for strains EXF-8193 and EXF-9187. This suggests a potential synergistic effect between tricyclazole and 1,2,4-benzenotriol, enhancing fungal growth. The synergistic effect could stem from several possible interactions. Tricyclazole, as a melanin biosynthesis inhibitor, may disrupt fungal stress responses or alter metabolic pathways, potentially making the fungal cells more adept at assimilating 1,2,4-benzenotriol as a carbon source. Another plausible explanation is that 1,2,4-benzenotriol could mitigate the stress or toxicity induced by tricyclazole, thus facilitating more robust growth. Additionally, tricyclazole-induced melanin inhibition might reduce competition within metabolic pathways or alter fungal membrane and cell wall dynamics, improving the uptake or utilization of 1,2,4-benzenotriol. These interactions highlight the potential interplay between stress modulation and carbon source metabolism. Further investigation, particularly into the metabolic and physiological changes induced by these compounds, is required to elucidate the mechanisms underpinning this apparent synergistic effect.
In cultures of all five E. dermatitidis strains grown with neurotransmitters and PAHs in the presence of tricyclazole, a distinct differentiation was observed, with fewer budded yeast and more pseudo-hyphae cells compared to cultures without tricyclazole. This observation underscores the significance of DHN-melanin production20 and the transition from hyphal to yeast forms during skin infection2, suggesting a potential linkage between these responses activated by polyaromatic compounds.
In the context of black yeasts, which prominently feature melanin in their cell walls, comprehending the influence of melanin on adaptability and pathogenicity is crucial90. Black yeast, including species like E. dermatitidis, exhibit exceptional resilience in diverse environments, often characterized by extreme conditions. Melanin production in these fungi has been associated with enhanced resistance to antifungal agents, phagocytosis by immune cells, and particularly environmental stresses such as high temperatures and ultraviolet radiation, as informed in Exophiala viscosa91,92,93,94. Moreover, melanin’s role in modulating immune responses and host-pathogen interactions underscores its significance in fungal pathogenicity. A deeper understanding of how melanin contributes to the adaptability and virulence of black yeasts is critical for developing targeted strategies to control fungal infections and mitigate their impact on human health and agriculture. Leveraging insights into melanin-associated mechanisms of polyextremotolerance and pathogenicity could pave the way for innovative approaches in antifungal drug development, bioremediation, and environmental management, highlighting the importance of fully comprehending the multifaceted roles of melanin in the adaptive strategies and virulence of black yeast fungi.
Final remarks: Introducing E. dermatitis as an “urban extremophile”
Exophiala dermatitidis is an urban extremophile predominantly found in PAH-polluted areas and indoor environments near humans. Adapted to niches like dishwashers, characterized by alkaline pH, high temperatures, and oxidative stress, it demonstrates the potential for global spread through urbanization and increased living standards. Known as an opportunistic pathogen, it causes infections ranging from cutaneous to neurotropic, with entry via aerosols, aspiration, or trauma.
This study links E. dermatitidis’ neurotropism to its ability to degrade hydrocarbons. It showcases its capacity to utilize (poly)aromatic compounds and neurotransmitters as carbon sources, supporting a connection between neurotransmitter assimilation and PAH metabolism. Its extensive CAZyme repertoire and metabolic versatility enable colonizing diverse habitats, from man-made materials to neural systems. Traits such as thermotolerance, biofilm formation, hemolysin production, and antifungal resistance highlight its adaptability and pathogenic potential, particularly under global environmental changes. The study also explored the role of melanin in E. dermatitidis’ tolerance to extreme conditions. It is generally acknowledged that melanin plays a significant role in the polyextremotolerance and pathogenicity of black yeasts, including E. dermatitidis. One of the important contributions of this study is the nuanced understanding of melanin’s role in polyextremotolerance. While melanin inhibition by tricyclazole did not significantly affect growth under most tested conditions, it induced changes in some strains, mainly related to salinity and pH, causing, in some cases, changes in morphology, suggesting that the importance of melanin may be context- and strain-dependent.
The study underscores E. dermatitidis’ resilience, adaptability, and potential for further spread, emphasizing the need for deeper exploration of its genetic and metabolic pathways to mitigate its impact on human health and ecosystems.
Data availability
Data obtained from this research is available. The annotated genome sequences have been submitted to GenBank and are accessible under BioProject PRJNA1110056.The sample and raw data are now publicly available, so the bioproject accession number should work for everyone: https://www.ncbi.nlm.nih.gov/bioproject/?term=PRJNA1110056This manuscript does not contain gels or blots image(s).
Abbreviations
- PAH:
-
Polycyclic aromatic hydrocarbon
- CFU:
-
Colony formation units
- GABA:
-
Gamma aminobutyric acid
- CAZyme:
-
Carbohydrate-active enzyme
- MEA:
-
Malt Extract Agar
- MEB:
-
Malt Extract Broth
- OD600 :
-
Cell density at 600 nm
- SDS:
-
Sodium dodecyl sulfate
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- PCA:
-
Principal component analysis
- GH:
-
Glycosyl hydrolases
- GT:
-
Glycosyl transferases
- AA:
-
Auxiliary activity enzymes
- CBM:
-
Carbohydrate-binding modules
- PL:
-
Polysaccharide lyases
References
Teixeira, M. M. et al. Exploring the genomic diversity of black yeasts and relatives (Chaetothyriales, Ascomycota). Stud. Mycol. 86, 1–28 (2017).
Kirchhoff, L., Olsowski, M., Rath, P. M. & Steinmann, J. Exophiala dermatitidis: key issues of an opportunistic fungal pathogen. Virulence 10, 984–998 (2019).
Ma, N. et al. Concurrent infection of Exophiala dermatitidis and Angiostrongylus cantonensis in central nervous system of a child with inherited CARD9 deficiency: a case report and literature review. J. Med. Mycol. 34, 101455 (2024).
Zalar, P., Novak, M., de Hoog, G. S. & Gunde-Cimerman, N. Dishwashers—a man-made ecological niche accommodating human opportunistic fungal pathogens. Fungal Biol. 115, 997–1007 (2011).
Gümral, R. et al. Dishwashers provide a selective extreme environment for human-opportunistic yeast-like fungi. Fungal Divers. 76, 1–9 (2016).
Raghupathi, P. K. et al. Microbial diversity and putative opportunistic pathogens in dishwasher biofilm communities. Appl. Environ. Microbiol. 84, 258 (2018).
Kulesza, K. et al. Dishwashers as an extreme environment of potentially pathogenic yeast species. Pathogens 10, 446 (2021).
Sudhadham, M. et al. The neurotropic black yeast Exophiala dermatitidis has a possible origin in the tropical rain forest. Stud. Mycol. 61, 145–155 (2008).
Branda, E. et al. Yeast and yeast-like diversity in the southernmost glacier of Europe (Calderone Glacier, Apennines, Italy). FEMS Microbiol. Ecol. 72, 354–369 (2010).
Döğen, A., Kaplan, E., Ilkit, M. & de Hoog, G. S. Massive contamination of Exophiala dermatitidis and E. phaeomuriformis in railway stations in subtropical Turkey. Mycopathologia 175, 381–386 (2013).
Baron, N. C. et al. Black fungi and hydrocarbons: an environmental survey for alkylbenzene assimilation. Microorganisms 9, 1008 (2021).
Babič, M. N., Zupančič, J., Gunde-Cimerman, N., de Hoog, S. & Zalar, P. Ecology of the human opportunistic black yeast Exophiala dermatitidis indicates preference for human-made habitats. Mycopathologia 183, 201–212 (2018).
Gostinčar, C., Zalar, P. & Gunde-Cimerman, N. No need for speed: slow development of fungi in extreme environments. Fungal Biol. Rev. 39, 1–14 (2022).
Zeng, J. S. et al. Spectrum of clinically relevant Exophiala species in the United States. J. Clin. Microbiol. 45, 3713–3720 (2007).
Kondori, N. et al. High rate of Exophiala dermatitidis recovery in the airways of patients with cystic fibrosis is associated with pancreatic insufficiency. J. Clin. Microbiol. 49, 1004–1009 (2011).
Pinheiro, R. L. et al. Peritonitis by Exophiala dermatitidis in a pediatric patient. Med. Mycol. Case Rep. 24, 18–22 (2019).
Lavrin, T. et al. The neurotropic black yeast Exophiala dermatitidis induces neurocytotoxicity in neuroblastoma cells and progressive cell death. Cells 9, 963 (2020).
Usuda, D. et al. Exophiala dermatitidis. World J. Clin. Cases. 9, 7963–7972 (2021).
Tesei, D. et al. Proteome of tolerance fine-tuning in the human pathogen black yeast Exophiala dermatitidis. J. Proteom. 128, 39–57 (2015).
Poyntner, C., Mirastschijski, U., Sterflinger, K. & Tafer, H. Transcriptome study of an Exophiala dermatitidis PKS1 mutant on an ex vivo skin model: is melanin important for infection? Front. Microbiol. 9, 89 (2018).
Chowdhary, A. et al. ESCMID and ECMM joint clinical guidelines for the diagnosis and management of systemic phaeohyphomycosis: diseases caused by black fungi. Clin. Microbiol. Infect. 20, 47–75 (2014).
De Hoog, G. S. & Haase, G. Nutritional physiology and selective isolation of Exophiala dermatitidis. Antonie Van Leeuwenhoek 64 (1993).
Prenafeta-Boldú, F. X., Summerbell, R. & de Sybren, G. Fungi growing on aromatic hydrocarbons: biotechnology’s unexpected encounter with biohazard? FEMS Microbiol. Rev. 30, 109–130 (2006).
Zajc, J., Gostinčar, C., Černoša, A. & Gunde-Cimerman, N. Stress-tolerant yeasts: opportunistic pathogenicity versus biocontrol potential. Genes (Basel) 10, 42 (2019).
Malo, M. E. et al. Transcriptomic and genomic changes associated with radioadaptation in Exophiala dermatitidis. Comput. Struct. Biotechnol. J. 19, 196–205 (2021).
Schultzhaus, Z., Chen, A., Shuryak, I. & Wang, Z. The transcriptomic and phenotypic response of the melanized yeast Exophiala dermatitidis to ionizing particle exposure. Front. Microbiol. 11, 25 (2021).
Song, Y. et al. Comparative ecology of capsular Exophiala species causing disseminated infection in humans. Front. Microbiol. 8, 36 (2017).
Černoša, A. et al. Virulence traits and population genomics of the black yeast Aureobasidium melanogenum. J. Fungi. 7, 665 (2021).
Buxton, R. Blood agar plates and hemolysis protocols. Am. Soc. Microbiol. 15, 1–9 (2005).
Vitale, R. G., De Hoog, G. S. & Verweij, P. E. Vitro vitro activity of amphotericin B, itraconazole, terbinafine and 5-fluocytosine against Exophiala spinifera and evaluation of post-antifungal effects. Med. Mycol. 41, 301–307 (2003).
Arikan, S. Vitro activity of nystatin comparedthose tholiposomalonystatinstamphotericinericin B, and fluconazole against clinical Candida isolates. J. Clin. Microbiol. 40, 1406–1412 (2002).
Barbara D. Alexander. National Committee for Clinical Laboratory Standards, Reference Method for Broth Dilution Antifungal Susceptibility Testing of Yeasts, 4th Edition (2017).
Martínez-Ávila, L. et al. Tracking gene expression, metabolic profiles, and biochemical analysis in the halotolerant basidiomycetous yeast Rhodotorula mucilaginosa EXF-1630 during benzo[a]pyrene and phenanthrene biodegradation under hypersaline conditions. Environ. Pollut. 271, 475 (2021).
Peidro-Guzmán, H. et al. Transcriptomic analysis of polyaromatic hydrocarbon degradation by the halophilic fungus Aspergillus sydowii at hypersaline conditions. Environ. Microbiol. 23, 3435–3459 (2021).
Troncoso, E. et al. Identification and characterization of yeasts isolated from the South Shetland Islands and the Antarctic Peninsula. Polar Biol. 40, 649–658 (2017).
Kumari, A. et al. Description of lipase producing novel yeast species Debaryomyces apis f.a., sp. nov. and a modified pH indicator dye-based method for the screening of lipase producing microorganisms. Sci. Rep. 13, 11819 (2023).
Rico, A. & Preston, G. M. Pseudomonas syringae pv. tomato DC3000 uses constitutive and apoplast-induced nutrient assimilation pathways to catabolize nutrients that are abundant in the tomato apoplast. Mol. Plant-Microbe Interactions 21, 269–282 (2008).
Gostinčar, C., Stajich, J. E., Zupančič, J., Zalar, P. & Gunde-Cimerman, N. Genomic evidence for intraspecific hybridization in a clonal and extremely halotolerant yeast. BMC Genom. 19, 364 (2018).
Prjibelski, A., Antipov, D., Meleshko, D., Lapidus, A. & Korobeynikov, A. Using SPAdes De Novo Assembler. Curr. Protoc. Bioinf. 70, 69 (2020).
Alonge, M. et al. Automated assembly scaffolding using RagTag elevates a new tomato system for high-throughput genome editing. Genome Biol. 23, 258 (2022).
Stanke, M. & Morgenstern, B. AUGUSTUS: a web server for gene prediction in eukaryotes that allows user-defined constraints. Nucleic Acids Res. 33, W465–W467 (2005).
Smit, A., Hubley, R. & Grenn, P. RepeatMasker Open-4.0. Preprint at (2015).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Kriventseva, E. V. et al. OrthoDB v8: update of the hierarchical catalog of orthologs and the underlying free software. Nucleic Acids Res. 43, D250–D256 (2015).
Zhang, H. et al. dbCAN2: a meta server for automated carbohydrate-active enzyme annotation. Nucleic Acids Res. 46, W95–W101 (2018).
Buchfink, B., Reuter, K. & Drost, H. G. Sensitive protein alignments at tree-of-life scale using DIAMOND. Nat. Methods. 18, 366–368 (2021).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comput. Biol. 7, e1002195 (2011).
Busk, P. K., Pilgaard, B., Lezyk, M. J., Meyer, A. S. & Lange, L. Homology to peptide pattern for annotation of carbohydrate-active enzymes and prediction of function. BMC Bioinform. 18, 214 (2017).
Aramaki, T. et al. KofamKOALA: KEGG Ortholog assignment based on profile HMM and adaptive score threshold. Bioinformatics 36, 2251–2252 (2020).
Kejžar, A., Gobec, S., Plemenitaš, A. & Lenassi, M. Melanin is crucial for growth of the black yeast Hortaea werneckii in its natural hypersaline environment. Fungal Biol. 117, 368–379 (2013).
De Mendiburu, F. & Simon, R. Agricolae—ten years of an open source statistical tool forexperiments in breeding, agriculture and biology. PeerJ Prepr 3, e1404v1 (2015).
Jari et al. Vegan: Community ecology package (2022).
R Core Team. R Version 4.2. 2. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2022).
Ferraroni, M. et al. Crystal structure of the hydroxyquinol 1,2-dioxygenase from nocardioides simplex 3E, a key enzyme involved in polychlorinated aromatics biodegradation. J. Biol. Chem. 280, 21144–21154 (2005).
Costa, D. M. A. et al. Catalytic mechanism for the conversion of salicylate into catechol by the flavin-dependent monooxygenase salicylate hydroxylase. Int. J. Biol. Macromol. 129, 588–600 (2019).
Semana, P. & Powlowski, J. Four aromatic intradiol ring cleavage dioxygenases from Aspergillus niger. Appl. Environ. Microbiol. 85, 74 (2019).
Lubbers, R. J. M. et al. Discovery and functional analysis of a salicylic acid hydroxylase from Aspergillus niger. Appl. Environ. Microbiol. 87, 789 (2021).
Axelrod, J. & Tomchick, R. Enzymatic O-methylation of epinephrine and other catechols. J. Biol. Chem. 233, 702–705 (1958).
Edmondson, D. E., Mattevi, A., Binda, C., Li, M. & Hubalek, F. Structure and mechanism of monoamine oxidase. Curr. Med. Chem. 11, 1983–1993 (2004).
Blasi, B., Tafer, H., Tesei, D. & Sterflinger, K. From Glacier to Sauna: RNA-Seq of the human pathogen black fungus Exophiala dermatitidis under varying temperature conditions exhibits common and novel fungal response. PLoS One 10, e0127103 (2015).
de Hoog, G. S., Matos, T., Sudhadham, M., Luijsterburg, K. F. & Haase, G. Intestinal prevalence of the neurotropic black yeast Exophiala (Wangiella) dermatitidis in healthy and impaired individuals. Mycoses 48, 142–145 (2005).
Wang, Q. & Szaniszlo, P. J. Roles of the pH signaling transcription factor PacC in Wangiella (Exophiala) dermatitidis. Fungal Genet. Biol. 46, 657–666 (2009).
Peñalva, M. A. & Arst, H. N. Jr. Recent advances in the characterization of ambient pH regulation of gene expression in filamentous fungi and yeasts. Annu. Rev. Microbiol. 58, 425–451 (2004).
Fernández-López, M. G., Batista-García, R. A. & Aréchiga-Carvajal, E. T. Alkaliphilic/alkali-tolerant fungi: molecular, biochemical, and biotechnological aspects. J. Fungi 9, 652 (2023).
Chen, Z. et al. Comparative genomic and transcriptomic analysis of Wangiella dermatitidis, a major cause of phaeohyphomycosis and a model black yeast human pathogen. G3 Genes|Genomes|Genetics. 4, 561–578 (2014).
Boral, H., Metin, B., Döğen, A., Seyedmousavi, S. & Ilkit, M. Overview of selected virulence attributes in Aspergillus fumigatus, Candida albicans, Cryptococcus neoformans, Trichophyton rubrum, and Exophiala dermatitidis. Fung. Genet. Biol. 111, 92–107. https://doi.org/10.1016/j.fgb.2017.10.008 (2018).
Sav, H. et al. Virulence markers of opportunistic black yeast in Exophiala. Mycoses 59, 343–350 (2016).
Nayak, A. P., Green, B. J. & Beezhold, D. H. Fungal hemolysins. Med. Mycol. 51, 1–16. https://doi.org/10.3109/13693786.2012.698025 (2013).
Seneviratne, C. J., Fong, P. H., Wong, S. S. & Lee, V. H. Antifungal susceptibility and phenotypic characterization of oral isolates of a black fungus from a nasopharyngeal carcinoma patient under radiotherapy. BMC Oral Health. 15, 39 (2015).
Badali, H., De Hoog, G. S., Sudhadham, M. & Meis, J. F. Microdilution in vitro antifungal susceptibility of Exophiala dermatitidis, a systemic opportunist. Med. Mycol. 49, 819–824 (2011).
Lazardi, K., Urbina, J. A. & de Souza, W. Ultrastructural alterations induced by two ergosterol biosynthesis inhibitors, ketoconazole and terbinafine, on epimastigotes and amastigotes of Trypanosoma (Schizotrypanum) cruzi. Antimicrob. Agents Chemother. 34, 2097–2105 (1990).
Pasko, M. T., Stephen, C., Piscitelli A. & Van Slooten, D. Fluconazole: a new triazole antifungal agent. DICP 24, 860–867 (1990).
dos Santos, A. G. et al. The molecular mechanism of Nystatin action is dependent on the membrane biophysical properties and lipid composition. Phys. Chem. Chem. Phys. 19, 30078–30088 (2017).
Snyder, R. Xenobiotic metabolism and the mechanism(s) of benzene toxicity. Drug Metab. Rev. 36, 531–547 (2004).
Dreyer, D. R., Miller, D. J., Freeman, B. D., Paul, D. R. & Bielawski, C. W. Perspectives on poly(dopamine). Chem. Sci. 4, 3796 (2013).
North, M. et al. Genome-wide functional profiling reveals genes required for tolerance to benzene metabolites in yeast. PLoS One 6, 23 (2011).
Benoit, I. et al. Closely related fungi employ diverse enzymatic strategies to degrade plant biomass. Biotechnol. Biofuels 8, 107 (2015).
Moreno, L. F. et al. Genomic analysis of ant domatia-associated melanized fungi (Chaetothyriales, Ascomycota). Mycol. Prog. 18, 541–552 (2019).
Rafiei, V., Vélëz, H. & Tzelepis, G. The role of glycoside hydrolases in phytopathogenic fungi and oomycetes virulence. Int. J. Mol. Sci. 22, 9359 (2021).
Reyre, J. L. et al. The maize pathogen Ustilago maydis secretes glycoside hydrolases and carbohydrate oxidases directed toward components of the fungal cell wall. Appl. Environ. Microbiol. 88, 89 (2022).
Matsumoto, R. et al. Genomic and secretomic analyses of the newly isolated fungus perenniporia fraxinea SS3 identified CAZymes potentially related to a serious pathogenesis of hardwood trees. Appl. Environ. Microbiol. 89, 145 (2023).
López-Mondéjar, R., Tláskal, V., da Rocha, U. N. & Baldrian, P. Global distribution of carbohydrate utilization potential in the prokaryotic tree of life. mSystems 7, 47 (2022).
Song, Y. et al. Comparative genomic analysis of capsule-producing black yeasts Exophiala dermatitidis and Exophiala spinifera, potential agents of disseminated mycoses. Front. Microbiol. 11, 25 (2020).
Toledo, A. V. et al. Melanins in fungi: types, localization and putative biological roles. Physiol. Mol. Plant. Pathol. 99, 2–6 (2017).
Grube, M., Muggia, L. & Gostinčar, C. Niches and Adaptations of Polyextremotolerant Black Fungi. In Polyextremophiles: Life under multiple forms of stress, pp. 551–566. Dordrecht: Springer Netherlands (2013).
Duarte, A. W. F. et al. Antarctic fungi as producers of pigments. In Fungi of Antarctica 305–318 (Springer International Publishing, 2019). https://doi.org/10.1007/978-3-030-18367-7_14.
Mohania, D. et al. Ultraviolet Radiations: Skin Defense-Damage Mechanism 71–87 (2017).
Butinar, L., Sonjak, S., Zalar, P. & Plemenitaš, A. & Gunde-Cimerman, N. Melanized halophilic fungi are eukaryotic members of microbial communities in hypersaline waters of solar salterns. Bot. Mar. 48, 896 (2005).
Van Dover, C. L. et al. A fungal epizootic in mussels at a deep-sea hydrothermal vent. Mar. Ecol. 28, 54–62 (2007).
Schnitzler, N. et al. Effect of melanin and carotenoids of Exophiala (Wangiella) dermatitidis on phagocytosis, oxidative burst, and killing by human neutrophils. Infect. Immun. 67, 94–101 (1999).
Paolo, W. F. et al. Effects of disrupting the polyketide synthase gene WdPKS1 in Wangiella [Exophiala] dermatitidis on melanin production and resistance to killing by antifungal compounds, enzymatic degradation, and extremes in temperature. BMC Microbiol. 6, 55 (2006).
Smith, D. F. Q. & Casadevall, A. The Role of Melanin in Fungal Pathogenesis for Animal Hosts. Fungal Physiol. Immunopath. 1–30 (2019).
Romsdahl, J. et al. Phenotypic characterization and comparative genomics of the melanin-producing yeast Exophiala lecanii-corni reveals a distinct stress tolerance profile and reduced ribosomal genetic content. J. Fungi 7, 1078 (2021).
Carr, E. C. et al. Characterization of anovel polyextremotolerant fungus, Exophiala viscosa, with insights into its melanin regulation and ecological niche. G3:Genes, Genomes, Genetics, 13(8), jkad110 (2023).
Acknowledgements
Acknowledgments: LR-L received a PhD fellowship from the National Council of Humanities, Sciences and Technologies (CONAHCyT), Government of Mexico. TM-P and TC-M received a post-doc fellowship from CONAHCyT, Government of Mexico. RAB-G received a Sabbatical fellowship (CVU: 389616) from CONAHCyT, Government of Mexico. The authors thank Luis Pacheco for his technical support.
Funding
This work was supported by RYC2022-037554-I project funded by MCIN/AEI/10.13039/501100011033 and FSE+ (RAB-G). RAB-G received a Sabbatical fellowship (CVU: 389616) from the National Council of Humanities, Sciences and Technologies (CONAHCyT), Government of Mexico. This work was also supported by CONAHCyT-Project 315114 (RAB-G) and Slovenian Research Agency to Infrastructural Centre Mycosmo (MRIC UL, I0-0022), programs P4-0432 and P1-0198.
Author information
Authors and Affiliations
Contributions
Authors´ contributions: RAB-G and NG-C conceived and designed the study. All authors contributed to the investigation. The first draft of the manuscript was written by LR-L, TC-M, and RAB-G. Figures were prepared by TM-P, LR-L, CG, and RAB-G. All authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Consent for publication
All authors consent to the publication of this work.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
de León, L.R., Moreno-Perlín, T., Castillo-Marenco, T. et al. Polyextremotolerant, opportunistic, and melanin-driven resilient black yeast Exophiala dermatitidis in environmental and clinical contexts. Sci Rep 15, 6472 (2025). https://doi.org/10.1038/s41598-025-88595-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-88595-z