Abstract
Vitiligo is a chronic autoimmune condition that leads to the loss of skin pigmentation in certain areas due to the destruction of melanocytes, which produce pigment. A topological index is a numerical value obtained from the structure of a chemical graph and is useful for studying the theoretical characteristics of organic molecules. It can also help determine the physico-chemical and biological aspects of various drugs. This article uses novel neighborhood degree-based topological indices to study vitiligo drugs and demonstrates a strong correlation with physico-chemical properties. Additionally, the results are compared with those obtained through degree-based topological indices.
Similar content being viewed by others
Introduction
Our interactions with the outside world are greatly influenced by our skin, and visible skin disorders may pose challenges to healthy psychosocial development. The skin disease that leads to the symptom of depigmentation of skin colour is called Vitiligo1. The skin develops smooth, white, or light patches, often known as macules. However, it can appear anywhere on your body, including your mucous membrane, eyes, and inner ears. Usually, it starts on your hands, forearms, feet, and face2. When the immune system destroys melanocytes in your body, this disease develops. Skin cells called melanocytes are responsible for producing melanin, the substance that gives pigmentation or colour to skin3. For each person with vitiligo, the extent of the damaged skin differs. While some people only have a few depigmented spots, others lose their skin’s pigmentation all over. Around 1% of people worldwide have vitiligo4. An important step in the management of vitiligo is to first acknowledge that it is not just a cosmetic disease but a multifactorial disease with an intricate interplay between genetic and non-genetic factors5. Numerous methods exist for treating vitiligo. Unfortunately, not every patient reacts well to the treatments available today. Therapy selection is based on the patient’s unique features, including age, disease kind and stage, and afflicted body site6. The etiology of vitiligo is still unclear and complicated. As such, it remains one of the most challenging dermatological problems.
Corticosteroids such as Fluticasone, Clobetasone, Beclomethasone Dipropionate, Desonide, Clobetasol propionate, Betamethasone Valerate, Hydrocortisone Valerate, and Fluticasone Propionate are used to inhibit the immune system and reduce inflammation7. For ailments like dermatitis, eczema, allergies, and asthma, they are frequently prescribed. On the other hand, Azathioprine is an immunosuppressive drug that is used to treat autoimmune conditions such as inflammatory bowel disease and rheumatoid arthritis, as well as to avoid organ transplant rejection8. To treat vitiligo, monobenzone is a depigmenting chemical that lightens the skin in unaffected areas to match the depigmented areas9. Psoralens are a class of artificial or naturally occurring substances that are used in ultraviolet (PUVA) therapy to treat vitiligo and psoriasis, among other skin conditions10.
Chemical graph theory is a subfield of mathematical chemistry. It focuses on the non-trivial application of graph theory to chemistry. This multidisciplinary field of study uses mathematics to tackle chemistry-related problems, potentially impacting both chemistry and mathematics11. A molecular graph can be used to perform a topological representation of a molecule. A molecular graph is made up of a set of lines that represent the covalent bonds between molecules and a collection of points that represent the atoms within them12. In graph theory terminology, these points are called vertices, and the lines are known as edges. A topological index is the graph invariant number calculated from a graph representing a molecule. The degree and distance-based topological indices are the two major classes that play a significant role in chemical graph theory. One advantage of topological indices is their direct application as fundamental numerical descriptors for comparing molecular properties, whether they are physical, chemical, or biological, in Quantitative Structure-Property Relationships (QSPR) and Quantitative Structure-Activity Relationships (QSAR). With the exponential growth of chemical data, the use of QSPR/QSAR models has become essential for managing the data and deriving valuable insights from its analysis13. There has been rapid development in the mathematical techniques used as regression tools in QSAR/QSPR analysis.
Khan et al.14 discussed the computational and topological properties of neural networks by utilizing graph theoretic parameters. Hayat et al.15 examined structure-property modeling for the thermodynamic properties of benzenoid hydrocarbons using various temperature-based indices. Alsinai et al.16 studied the degree-based HDR indices and Mhr polynomial of drugs used to treat the COVID-19 pandemic. Julietraja et al.17 conducted a theoretical analysis of superphenalene using various VDB indices. In their research, Parveen et al.18 focused on degree-based topological indices of autoimmune disease vitiligo treatment and conducted QSPR modeling. Abubakar et al.19 carried out a QSPR analysis of the physical properties of antituberculosis drugs using neighborhood degree-based topological indices. Chamua et al.20 proposed some neighborhood degree sum-based topological indices that exhibit a strong correlation with certain physico-chemical parameters of 66 alkanes and octane isomers. They also demonstrated the superiority of neighborhood degree sum-based topological indices over traditional degree-based ones in terms of sensitivity. Ahmed et al.21 discussed molecular insights into anti-Alzheimer’s drugs through predictive modeling using linear regression and QSPR analysis. Mahboob et al.22 investigated QSPR analysis of anti-hepatitis drugs using a linear regression model. Vignesh et al.23 examined the QSPR analysis of some closed neighborhood degree-based topological indices for octane isomers and also performed sensitivity analysis for the proposed indices. Abirami et al.24 studied reverse neighborhood degree-based topological indices for transition metal phthalocyanine polymers and compared them with reverse degree-based topological indices. Shibsankar Das et al.25 executed a comparative analysis of neighborhood degree sum-based molecular indices on some silicon carbide networks. Mondal et al.26 introduced neighborhood degree-based topological indices named NDe indices and compared them to a few popular and frequently used indices for structure-property modeling and isomer discrimination. Vignesh et al.27 introduced open neighborhood indices, namely \(SK_{N}\), \(SK1_{N}\), \(SK2_{N}\), neighborhood degree versions of Modified Randic and Inverse Sum indices for graphene structure. Balasubramaniyan et al.28 performed QSPR analysis of asthma drugs using neighborhood degree-based topological indices. Kirmani et al.29 investigated the physico-chemical properties of anti-viral drugs used in COVID-19 treatment by establishing QSPR modeling using neighborhood degree-based topological indices. Balasubramaniyan et al.30 investigated the physico-chemical properties of prostate cancer drugs using degree and neighborhood degree-based topological indices.
Techniques and material
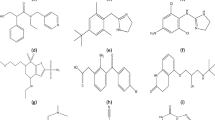
Molecular graphs are mathematical structures constructed by representing each atom as a vertex and connecting atom pairs with edges. Typically, in this representation, hydrogen atoms are not included. The molecular structure of the drugs used for vitiligo treatment is obtained from PubChem and is shown in Figure 1.
Chemical structure of Vitiligo treatment drugs.
Throughout our discussion, let \(G=(V, E)\) denote a molecular graph with the vertex set V and the edge set E. The following degree-based topological indices and their computed values (Table 1, Table 2) are chosen from18 for our comparative study.
Here, d(u) represents the degree of a vertex u in the molecular graph G.
We have selected ten neighborhood degree-based topological descriptors, which are listed in Table 3. We used the edge partitioning technique to analyze the molecular graphs of the drugs and obtained the calculated values for the set of indices, as shown in Table 4.
The neighborhood of a vertex u of a graph G is denoted by \(N_G(u)\), and defined as the set of all vertices that are adjacent to u in G. In symbolic form, it is expressed as \(N_G(u)=\{v\in V(G):u\) and v are adjacent in \(G\}\). The neighborhood degree of a vertex u in a molecular graph G is denoted by \(\delta _G(u)\), and it is defined as \(\delta _G(u)=\sum \limits _{v\in N_G(u)}d(v)\). That is the sum of all degrees of the vertices which are adjacent to u in G.
For instance, let us consider the drug \(\mathcal {P}\): Psoralen.

Let \(E_{ij}\) denote the set of edges with neighborhood degrees of end vertices i and j. We employ the edge partitioning technique to compute the \(ND_1\) index of \(\mathcal {P}\). The edge partition of this drug is as follows: \(E_{35}=1,\; E_{55}=2,\; E_{45}=2,\; E_{57}=3,\; E_{56}=1,\; E_{67}=5,\; E_{77}=2\). Now, by definition of \(ND_1\) index, we get
Similarly, all other neighborhood degree-based indices are computed for the chosen drugs and listed in Table 4.
We used the following MATLAB algorithm to validate our computations of neighbourhood degree-based indices given in Table 4.
Computation of neighbourhood degree-based indices
The physico-chemical properties chosen for our analysis are Boiling Point (BP), Enthalpy of Vapourization (EV), Molar Refraction (MR), Polar Surface Area (PSA), Refractivity (REF), Polarity (P), Surface Tension (ST), Molar Volume (MV), Molecular Weight (MW), Exact Mass (EM), Heavy Atom Count (HAC) and Complexity (C). These property values are derived from PubChem and ChemSpider which are given in Table 5.
Investigating the functional relationship between variables can be done theoretically with regression analysis. It has extensive application in numerous subject areas. The relationship between the response and predictor variables is represented as an equation or model. We represent the response variable by and the set of predictor variables by \(\mathcal {X}_1,\mathcal {X}_2,\cdots,\mathcal {X}_n,\) where n depicts the number of predictor variables, \(\epsilon\) is random error. The relationship by regression model is
Results and discussions
In this section, we will conduct a QSPR analysis using selected neighborhood degree-based indices to identify the best predictors for the twelve physico-chemical properties of Vitiligo drugs. Additionally, we will compare our results with those obtained from QSPR analysis using degree-based indices. To accomplish this, we will expand on the analysis conducted in18 by including the physico-chemical properties of MR, PSA, ST, MW, EM, and HAC. Statistical parameters such as the coefficient of determination (\(R^2\)), standard error (SE), F-statistic, and the significance p-value are essential for validating the regression models.
QSPR analysis using neighborhood degree-based topological indices
This subsection deals with the QSPR analysis of the chosen physico-chemical properties of drugs using neighborhood degree-based indices.
Linear regression model
Table 6 depicts the \(R^2\) value acquired among the indices and the physico-chemical properties in the linear regression model (LREM). The maximum \(R^2\) concerning the minimum SE is highlighted in bold. . The model equations are given below.
The index \(ISI_N\) performed as the best predictor for the properties MR, P and REF. The index \(ND_1\) serves as the best predictor for MV.
The properties MW, EM and C are best predicted by the index \(SK_N\). However, the properties BP, EV, PSA, ST and HAC are not predicted in LREM model.
Table 7 represents the best reliable fit for the properties obtained in LREM.
Quadratic regression model
In the quadratic regression model (QREM), Table 8 shows the \(R^2\) values obtained for the neighborhood degree-based indices and the properties. The highest \(R^2\) and lowest standard error (SE) are highlighted in bold. The model equations can be found below.
The index \(ND_2\) acts as the best predictor for the physico-chemical properties of EV, MR, P, MV, MW, and EM. The index \(ND_3\) is the most reliable predictor for the property C.
The index \(ND_5\) acts as the best predictor for the property REF.
The properties of BP, PSA, ST, and HAC are not predicted by the QREM. Table 9 depicts the best predictive fits obtained in QREM.
Cubic regression model
Table 10 shows the \(R^2\) value obtained through the cubic regression model (CREM) between indices and the physico-chemical properties. The maximum \(R^2\) and minimum SE values are marked in bold. The model equations are given below.
The index \(ND_5\) acts as the best predictor for the properties of EV, PSA, ST, and MV. The property BP is best predicted by \(SK_N\).
The properties of MR, P, MW, and EM are best predicted by the index \(ND_4\) and the index \(ND_3\) for C. The index \(mR_N\) is used to predict the property REF. The property HAC is not predicted by CREM. Table 11 depicts the best predictive fit obtained in CREM.
Multiple linear regression model
The following are the multiple linear regression model (MLREM) equations obtained between the response variable and the two or more predictor variables. The best predictive fits are shown in Table 12. It is observed that the properties PSA, ST, and HAC are not predicted by MLREM.
QSPR analysis using degree-based topological indices
This subsection deals with the QSPR analysis of the twelve physico-chemical properties of Vitiligo drugs using a set of nine degree-based topological indices. The best predictors are identified in each of the following models.
Linear regression models
Table 13 depicts the \(R^2\) value acquired among the indices and the physico-chemical properties of Vitiligo drugs in the LREM. The maximum \(R^2\) and minimum SE values are marked in bold. The model equations are given below.
We infer that the properties \(MR, P, MW, EM \text { and } C\) are best predicted by the index F, and the property REF is best predicted by the index \(M_{1}\), whereas the properties \(BP, EV, PSA, ST \text { and } HAC\) are not predicted in the LREM. Table 14 represents the most reliable fit for the properties obtained in LREM.
Quadratic regression models
Table 15 presents the \(R^2\) obtained among each degree-based index and the properties, whereas the indices exhibiting higher \(R^2\) values are identified and marked in bold.
We infer that the property EV is best predicted by the index H and the properties \(MR, P, \text { and } REF\) are best predicted by the index RA. The index ABC acts as the best predictor for the properties \(MV, MW \text { and } EM\) and F for the property C, whereas the properties \(BP, PSA, ST, \text { and } HAC\) are not predicted in the QREM. Table 16 shows the best predictors and the statistical values obtained in the QREM.
Cubic regression models
Table 17 presents \(R^2\) acquired between the degree-based index and the properties. The indices exhibiting higher \(R^2\) values are identified as the optimal predictors in the CREM. The best predictive models are as follows:
We infer that the property BP is best predicted by the index ABC, the property EV is best predicted by the index GA, and the properties \(MR, P, REF, MW\text { and } EM\) are best predicted by the index RA. The index \(M_1\) is used to forecast the property ST, and the properties MV and C are best predicted by the index F, whereas the properties \(PSA \text { and } HAC\) are not predicted in the CREM. Table 18 shows the best predictors and the statistical parameters obtained through the CREM.
Multiple linear regression model
In this subsection, we perform the MLREM regression analysis on degree-based topological indices for the twelve physico-chemical properties. The MLREM models are as follows:
The corresponding \(R^{2}\), Adj-\(R^{2}\), SE values, F-Stat, and p-value are given in Table 19, whereas the properties BP, EV, PSA, ST, MV, and HAC are not predicted by this MLREM using the set of chosen degree-based topological indices.
Result and analysis
In this section, we present the results of the single variate regression models using neighborhood and degree-based topological indices. We compared the most accurate models to obtain effective predictive models for each physico-chemical property, with the exception of the property HAC, which is displayed in Table 20.
The graphical visualization of the best-fit models is shown in Figure 2- Figure 4
ABC against BP, \(mR_N\) index against REF in CREM and \(ND_3\) index against C in QREM.
\(ND_5\) index against EV, PSA, ST and MV in CREM.
\(ND_4\) index against MR, P, MW and EM in CREM.
The table (Table 21) presents a comparison of multiple linear regression (MLRE) models that utilize degree-based and neighborhood degree-based indices. It is noted that among the twelve properties selected, the properties HAC, PSA, and ST are not predicted by any of these MLRE models. Furthermore, upon analyzing the results in Tables 20 and 21, we observe that the single variate regression models perform better than the MLRE models.
The comparative analysis has been made with the best models obtained from the earlier sections: Single-variate regression models along with the existing experimental values. Figures 5 - 10 give the graphical representation of the comparison between the actual and predicted values.
Graphical Depiction of Actual Vs Predicted values for BP and EV.
Graphical Depiction of Actual Vs Predicted values for MR and PSA.
Graphical Depiction of Actual Vs Predicted values for REF and P.
Graphical Depiction of Actual Vs Predicted values for ST and MV.
Graphical Depiction of Actual Vs Predicted values for MW and EM.
Graphical Depiction of Actual Vs Predicted values for C.
Conclusion
In this study, we calculated ten neighborhood degree-based topological indices for vitiligo treatment drugs using the edge partitioning technique. We conducted QSPR modeling to analyze the relationship between these indices and their physico-chemical properties. We compared the regression models obtained from the neighborhood degree and degree-based indices and identified the best predictive fits. With reference to Tables 20 and 21, we observe the following:
-
Neighborhood degree-based indices \(ND_3,\;ND_4,\;ND_5\), and \(mR_N\) outperformed the degree-based indices in predicting physico-chemical properties \(EV,\;MR,\;PSA,\;REF,\;P,\;ST,\;MV,\;MW,\;EM,\) and C of drugs used for vitiligo treatment.
-
The degree-based index ABC turned out to be the best predictor for the property BP.
-
Despite the analysis, none of the regression models were able to predict the property HAC.
The pharmaceutical sector could benefit from this information to develop better medications for preventing vitiligo. These findings offer valuable insights for drug science experts and provide a method for forecasting the physico-chemical characteristics of drugs for various other illnesses.
Data availability
The chemical structures of the drugs are obtained from the National Library of Medicine whose URL is: www.pubchem.ncbi.nlm.nih.gov.
References
Wankowicz-Kalinska, A., van den Wijngaard, R. M., Nickoloff, B. J., and Das, P. K. (2004, January). Autoimmune aspects of depigmentation in vitiligo. In Journal of Investigative Dermatology Symposium Proceedings (Vol. 9, No. 1, pp. 68-72). Elsevier.
Bergqvist, C. & Ezzedine, K. Vitiligo: a review. Dermatology 236(6), 571–592 (2020).
Kutlubay, Z., Karakus, O., Engin, B. & Serdaroglu, S. Vitiligo as an autoimmune disease. J Turk Acad Dermatol 6(2), 1262 (2012).
Le Poole, I. C. & Luiten, R. M. Autoimmune etiology of generalized vitiligo. Dermatologic Immunity 10, 227–243 (2008).
Frisoli, M. L., Essien, K. & Harris, J. E. Vitiligo: mechanisms of pathogenesis and treatment. Annual review of immunology 38, 621–648 (2020).
Kostoviæ, K., Nola, I., Buèan, Ž., and Šitum, M. (2003). Treatment of vitiligo: current methods and new approaches. LOGICA, 163.
Senyigit, T. & Ozer, O. Corticosteroids for skin delivery: challenges and new formulation opportunities (In Glucocorticoids-New Recognition of Our Familiar Friend, IntechOpen, 2012).
Hoffmann, M., Rychlewski, J., Chrzanowska, M., and Hermann, T. (2001). Mechanism of activation of an immunosuppressive drug: azathioprine. Quantum chemical study on the reaction of azathioprine with cysteine. Journal of the American Chemical Society, 123(26), 6404-6409.
Arowojolu, O. A., Orlow, S. J., Elbuluk, N. & Manga, P. The nuclear factor (erythroid-derived 2)-like 2 (NRF 2) antioxidant response promotes melanocyte viability and reduces the toxicity of the vitiligo-inducing phenol monobenzone. Experimental dermatology 26(7), 637–644 (2017).
Ortonne, J. P. Psoralen therapy in vitiligo. Clinics in dermatology 7(2), 120–135 (1989).
Bernard, N. M. & Prijith, G. S. Graph theory in other subjects 175 (Sustainable Development for Society, Industrial, 2022).
Babujee, J. B. & Ramakrishnan, S. Topological indices and new graph structures. Applied Mathematical Sciences 6(108), 5383–5401 (2012).
Liu, P. & Long, W. Current mathematical methods used in QSAR/QSPR studies. International Journal of Molecular Sciences 10(5), 1978–1998 (2009).
Khan, A. et al. Computational and topological properties of neural networks by means of graph-theoretic parameters. Alexandria Engineering Journal 66, 957–977 (2023).
Hayat, S., Khan, A., Ali, K. & Liu, J. B. Structure-property modeling for thermodynamic properties of benzenoid hydrocarbons by temperature-based topological indices. Ain Shams Engineering Journal 15(3), 102586 (2024).
Alsinai, A., Ahmed, H., Alwardi, A. & Soner, N. D. HDR degree-based indices and Mhr-polynomial for the treatment of COVID-19. Biointerface Research in Applied Chemistry 12(6), 7214–7225 (2021).
Julietraja, K., Alsinai, A. & Alameri, A. Theoretical analysis of superphenalene using different kinds of VDB indices. Journal of Chemistry 2022(1), 5683644 (2022).
Parveen, S., Awan, N. U. H., Farooq, F. B., Fanja, R. & Anjum, Q. U. A. Topological indices of novel drugs used in autoimmune disease vitiligo treatment and its QSPR modeling. BioMed Research International 2022(1), 6045066 (2022).
Abubakar, M. S., Aremu, K. O., Aphane, M., and Amusa, L. B. (2024). A QSPR Analysis of Physical Properties of Antituberculosis Drugs Using Neighborhood Degree-Based Topological Indices and Support Vector Regression. Heliyon.
Chamua, M., Buragohain, J. & Bharali, A. Some novel neighborhood degree sum-based versus degree-based topological indices in QSPR analysis of alkanes from n-butane to nonanes. International Journal of Quantum Chemistry 124(1), e27292 (2024).
Ahmed, W., Ali, K., Zaman, S., and Raza, A. (2024). Molecular insights into anti-Alzheimer’s drugs through predictive modeling using linear regression and QSPR analysis. Modern Physics Letters B, 2450260.
Mahboob, A., Rasheed, M. W., Dhiaa, A. M., Hanif, I., and Amin, L. (2024). On quantitative structure-property relationship (QSPR) analysis of physicochemical properties and anti-hepatitis prescription drugs using a linear regression model. Heliyon.
Ravi, V., and Desikan, K. (2024). Quantitative Structure-Property Relationship (QSPR) Analysis of Some Closed Neighborhood Degree Based Topological Indices for Octane Isomers. Authorea Preprints.
Abirami, S. J., Raj, S. A. K. & Siddiqui, M. K. Computation of reverse neighborhood degree-based topological indices for the transition metal phthalocyanine polymers (poly-TMPc). Physica Scripta 99(2), 025025 (2024).
Das, S., and Kumar, V. (2024). Neighborhood Degree Sum-based Molecular Indices and Their Comparative Analysis of Some Silicon Carbide Networks. Physica Scripta.
Mondal, S., Dey, A., De, N. & Pal, A. QSPR analysis of some novel neighborhood degree-based topological descriptors. Complex and intelligent systems 7, 977–996 (2021).
Ravi, V. & Desikan, K. Neighborhood degree-based topological indices of graphene structure. Biointerface Research in Applied Chemistry 11(5), 13681–13694 (2021).
Balasubramaniyan, D. & Chidambaram, N. On some neighborhood degree-based topological indices with QSPR analysis of asthma drugs. The European Physical Journal Plus 138(9), 823 (2023).
Kirmani, S. A. K., Ali, P. & Azam, F. Topological indices and QSPR/QSAR analysis of some antiviral drugs being investigated for the treatment of COVID-19 patients. International Journal of Quantum Chemistry 121(9), e26594 (2021).
Balasubramaniyan, D., Chidambaram, N., and Ravi, V. (2024). Estimating Physico-chemical Properties of Drugs for Prostate Cancer using Degree-Based and Neighborhood Degree-Based Topological Descriptors. Physica Scripta.
Acknowledgements
The author sincerely thank the editor and anonymous referees for their careful reading of the manuscript and suggestions to improve the article in its present form.
Funding
There is no funding to support this article.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, X., Balasubramaniyan, D., Chidambaram, N. et al. Predictive analysis of vitiligo treatment drugs using degree and neighborhood degree-based topological descriptors. Sci Rep 15, 5218 (2025). https://doi.org/10.1038/s41598-025-89603-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-89603-y
Keywords
This article is cited by
-
Degree-based graph energies as molecular descriptors in QSPR analysis
Chemical Papers (2026)
-
Predicting flavonoid physicochemical properties using topological indices and regression modeling
Scientific Reports (2025)
-
On analysis of two-dimensional nickel-based organometallic network via statistical method
Chemical Papers (2025)