Abstract
Efficient segmentation of small hyperreflective dots, key biomarkers for diseases like macular edema, is critical for diagnosis and treatment monitoring.However, existing models, including Convolutional Neural Networks (CNNs) and Transformers, struggle with these minute structures due to information loss.To address this, we introduce EFCNet, which integrates the Cross-Stage Axial Attention (CSAA) module for enhanced feature fusion and the Multi-Precision Supervision (MPS) module for improved hierarchical guidance. We evaluated EFCNet on two datasets: S-HRD, comprising 313 retinal OCT scans from patients with macular edema, and S-Polyp, a 229-image subset of the publicly available CVC-ClinicDB colonoscopy dataset. EFCNet outperformed state-of-the-art models, achieving average Dice Similarity Coefficient (DSC) gains of 4.88% on S-HRD and 3.49% on S-Polyp, alongside Intersection over Union (IoU) improvements of 3.77% and 3.25%, respectively. Notably, smaller objects benefit most, highlighting EFCNet’s effectiveness where conventional models underperform. Unlike U-Net-Large, which offers marginal gains with increased scale, EFCNet’s superior performance is driven by its novel design. These findings demonstrate its effectiveness and potential utility in clinical practice.
Similar content being viewed by others
Introduction
Small medical objects, such as HyperReflective Dots (HRDs) observed on fundus Optical Coherence Tomography (OCT) are increasingly recognized by ophthalmologists as potential biomarkers for retinal diseases, including age-related macular degeneration (AMD), diabetic macular edema (DME), and macular edema caused by branch retinal vein occlusion (BRVO). The presence, number, and spatial distribution of HRDs provide key insights into the severity and progression of these disorders1,2,3,4. However, manual annotation of these small objects is both time-consuming and labor-intensive. Consequently, there is a growing need to develop automated segmentation methods for small medical objects using advanced computer vision algorithms. Figure 1A provides examples of such small medical objects.
Despite the success of numerous image segmentation models such as U-Net5, ResUNet6, DenseUNet7, ResUNet++8, TransFuse9, Swin-Unet10, and the recent SAM11, their effectiveness in segmenting small medical objects like HRDs is limited. Two primary limitations—significant information loss and inadequate feature utilization—suggest that these models might may not perform well for small object tasks. First, traditional methods5,12 restrict feature integration to the encoder-decoder stages, which limits the diversity of features used across decoding stages. Previous studies13–14, have shown that combining shallow and deep features improves segmentation accuracy. As a result, these models fail to use the full range of encoder information, leaving valuable data untapped. Second, many models use a single segmentation head for supervision at the final decoding stage6,12,15. Research indicates that early decoding stages, which process low-resolution features, are critical for accurately localizing small objects16,17,18. However, these stages often lose information during convolution and upsampling processes. Additionally, small medical objects inherently contain less data compared to larger ones, which worsens the impact of information loss. These limitations are evident in the SAM11 model, which performs poorly when segmenting small medical objects, as noted in prior research.
To address the challenge of information loss in segmenting small medical objects, we developed a novel encoder-decoder-based model that optimizes feature use at every stage. Figure 1B highlights the advantages of our method compared to conventional approaches. In the encoder, our model integrates the Cross-Stage Axial Attention Module (CSAA), which employs an attention mechanism to combine features from all stages. This enables the model to adjust to the informational needs of each decoding stage, minimizing the typical information loss. The CSAA ensures the decoder has access to relevant information throughout the process, improving segmentation accuracy.In the decoder, we introduce the Multi-Precision Supervision Module (MPS). This module applies segmentation heads with different precisions at various stages, effectively using low-resolution features. These features help capture broad contextual information while temporarily focusing less on fine local details. By using MPS, the model optimizes feature utilization at each stage, reducing information loss and enhancing segmentation precision.
To validate our model’s effectiveness, we conducted experiments on two datasets: S-HRD and S-Polyp, as shown in Fig. 1A. The S-HRD dataset was developed by our team, and the S-Polyp dataset is a subset of CVC-ClinicDB19. Our model’s performance on these datasets surpasses existing state-of-the-art models, as shown by higher scores in the Dice Similarity Coefficient (DSC) and Intersection over Union (IoU). These results demonstrate the potential of our approach to improve the segmentation of small medical objects, offering significant benefits for medical imaging diagnostics.
Related work
Medical image segmentation
Recently, researchers have developed several innovative methods for semantic segmentation in medical images20,21,22,23,24,25,26,27,28,29. Agarwal et al.25 proposed multi-scale dual-channel feature embedding decoder in the field of biomedical image segmentation. Zhang et al.9 proposed a unique hybrid architecture that integrates both CNN and Transformer technologies, which helps in preserving low-level details that are often lost. Chen et al.30 enhanced the U-Net encoder by incorporating a Transformer module, which improved the model’s capability for long-range dependency modeling. Wang et al.31 tackled the issue of overfitting through the use of a Transformer encoder and furthered this innovation by introducing a progressive locality decoder, aimed at optimizing local information processing within medical images. While these approaches have significantly advanced the field of medical image segmentation, they typically do not adequately address the impact of object size on segmentation outcomes.Lou et al.32 introduced the Context Axial Reverse Attention module (CaraNet) to address this challenge by focusing on small objects’ local details. However, CaraNet’s reliance on bilinear interpolation during the decode phase results in considerable information loss, negatively impacting small object segmentation. In contrast, our model mitigates such information loss through the integration of the CSAA and MPS, substantially enhancing segmentation precision for small objects.
General segmentation model
Kirillov et al. introduced the SAM11, a versatile segmentation model that has made significant strides in natural image segmentation. Despite its success, SAM is not well-suited for many medical image segmentation tasks, especially those involving small objects and complex boundaries, which often require manual intervention33,34. To address these limitations, Ma et al. developed MedSAM35, an adaptation of SAM tailored for medical images. MedSAM outperforms SAM in handling complex segmentation tasks but still struggles with small object segmentation.
Attention mechanism
Numerous attention-based methods have emerged in recent years36,37,38,39,40,41,42,43,44, and have been applied to diverse tasks in computer vision and natural language processing. Vaswani et al.41 introduced the Transformer, which marked a shift from traditional convolutional models to attention-based ones. Woo et al.42 enhanced this idea by adding Channel and Spatial Attention to CNNs, improving their ability to handle complex inputs. Zhang et al.43 proposed the Pyramid Squeeze Attention Module (PSA), which captures spatial information across multiple channels. Building on these developments, we introduce the CSAA, which promotes better feature integration across the model while minimizing information loss. Yadav et al.44 proposed a modified recurrent residual attention U-Net model, which has advanced the segmentation of brain tumors in MRI images, further influencing small object segmentation approaches in medical imaging.
Contribution
-
1.
Innovative segmentation approach: We introduce EFCNet, a novel method designed to address the challenges of segmenting small medical objects. This approach enhances the extraction of diverse information by systematically processing features at every stage, reducing information loss—a common issue in small object segmentation. Our approach directly improves on existing techniques by emphasizing comprehensive feature utilization.
-
2.
Development of key modules: We have designed two innovative modules: the CSAA and MPS. These modules specifically address information loss during the encoder and decoder phases of segmentation models. Their integration results in a significant improvement in segmentation accuracy, capturing fine-grained details more effectively.
-
3.
Benchmark construction and model validation: We established a robust benchmark for small medical object segmentation, developing specialized datasets, S-HRD and S-Polyp. Our model was rigorously tested against this benchmark and consistently outperformed existing state-of-the-art models, setting a new standard for performance in small object segmentation in medical imaging.
Methods
This section describes the development of a fundus OCT dataset, the architectural details of EFCNet, including its CSAA and MPS, and the formulation of the loss function used in our experiments.
Dataset establishment
For this study, we established two distinct datasets, with medical image samples shown in Fig. 1A:
S-HRD dataset
This dataset was compiled from 313 retinal OCT scans obtained from patients treated for DME or BRVO-induced macular edema at the EENT Hospital of Fudan University within the last six months. Informed consent was obtained from all participants involved in the study. The study protocol was approved by the Ethics Committee of the EENT Hospital, adhering to the principles of the Declaration of Helsinki. Patient confidentiality was strictly maintained through rigorous anonymization. Each OCT scan, performed using the Heidelberg Spectralis OCT + HRA (Heidelberg Engineering, Heidelberg, Germany), was centered on the fovea. HRDs were defined as discrete intraretinal reflectivity changes no larger than 30 μm, with reflectivity similar to that of the nerve fiber layer and no associated back shadowing. The images were annotated independently by two ophthalmologists who cross-referenced clinical records to ensure the accuracy of annotations. Discrepancies were resolved by a senior specialist. In this dataset, each lesion occupies less than 1% of the total image area.
S-Polyp dataset
A subset of 229 images was curated from the publicly available CVC-ClinicDB19, which includes 612 high-resolution images from 31 colonoscopy video sequences. This database is commonly used for polyp detection and includes ground truth masks that delineate the polyp regions. The subset was specifically selected to include images where each lesion occupies less than 5% of the total image area, emphasizing small-scale features. This selection challenges conventional segmentation methods and aims to improve the accuracy of small polyp detection.
Problem setup and notations
In our segmentation task, we define the medical image dataset as \(\:X=\left\{{x}_{1},\dots\:,{x}_{m}|{x}_{i}\in\:{\mathbb{R}}^{\left(C\times\:H\times\:W\right)},i=\text{1,2},\dots\:,m\right\}\), where each \(\:{x}_{i}\) represents an OCT image and \(\:i\) ranges over the dataset indices. Experienced ophthalmologists used ITK-SNAP, a versatile, open-source 3D medical image analysis software for user-guided segmentation, to segment HRD lesions on each OCT image. This meticulous process produced a corresponding ground truth dataset \(\:Y=\left\{{y}_{1},\dots\:,{y}_{m}|{y}_{i}\in\:{\text{0,1}}^{\left(1\times\:H\times\:W\right)},i=\text{1,2},\dots\:,m\right\}\), where each \(\:{y}_{i}\) is a binary mask of the segmented lesions in \(\:{x}_{i}\) The combined dataset \(\:D=\{X,\:Y\}\:\)is split into a training set \(\:{D}_{\text{train}}=\{{X}_{\text{train}},{Y}_{\text{train}}\}\) and a testing set \(\:{D}_{\text{test}}=\{{X}_{\text{test}},{Y}_{\text{test}}\}\),facilitating the model’s training and evaluation phases.
The objective of this study is to develop an algorithm that enhances our model’s capability to effectively segment small medical objects from \(\:{D}_{\text{train}}\) and demonstrate its robustness on \(\:{D}_{\text{test}}\). The focus is on ensuring precise segmentation during training and consistent generalizability on unseen data during testing.
Overall architecture
We present EFCNet, a novel architecture tailored for precise segmentation of small medical objects. The model integrates two key modules: the CSAA and the MPS, both crucial for processing information throughout the encoding and decoding phases.
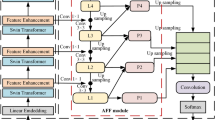
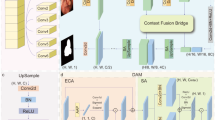
As shown in Fig. 1C, the process begins with an input image undergoing multi-stage encoding to produce detailed feature maps. The CSAA Module harmonizes these features across all encoder stages, refining the data for the decoding process. This ensures that each decoding stage leverages a complete dataset profile for accurate segmentation. In the decoding process, the MPS Module executes multi-precision predictions with each stage receiving specific supervision to fine-tune the segmentation results. During testing, the output from the final decoder stage serves as the definitive model result, optimized for high accuracy and reliability.
CSAA module
To address the challenge of critical information dispersion across encoder stages and reduce information loss during convolution and downsampling, we introduce the CSAA module. This module adaptively processes features from all encoder stages, refining their integration to efficiently inform the decoding process. As shown in Fig. 1D, the CSAA operates in four steps: resizing, W-dimensional axial attention, H-dimensional axial attention, and resizing back. This design optimizes feature fusion, ensuring better alignment between encoder and decoder stages, which ultimately enhances segmentation accuracy.
Resizing
To improve the integration of features across all encoder stages, each feature map within the encoder is resized to uniform dimensions \(\:\left({C}^{*},{H}^{*},{W}^{*}\right)\) using convolution operations. This adjustment aligns both spatial and channel dimensions to facilitate subsequent processing steps:
where \(\:{f}_{i}^{e}\) represents the feature map at stage \(\:i\) of the encoder, \(\:{\upsigma\:}\) signifies the ReLU activation function, and \(\:BN\:\)denotes batch normalization. \(\:k\) represents the total number of encoder and decoder stages. These resized feature maps, \(\:\{{f}_{i}^{*}{\}}_{i=1}^{k}\), are then prepared as inputs for the subsequent W-dimensional axial attention step.
W-dimensional CSAA
In the W-Dimensional CSAA module, we generate the Query \(\:\{{Q}_{i,w}{\}}_{i=1}^{k}\), Key \(\:\{{K}_{i,w}{\}}_{i=1}^{k}\), and Value \(\:\{{V}_{i,w}{\}}_{i=1}^{k}\) components based on the resized feature maps \(\:\{{\text{f}}_{\text{i}}^{\text{*}}{\}}_{\text{i}=1}^{\text{k}}\) in the width dimension. The process is defined by the following equations: \(\:{Q}_{i,w}={W}_{i}^{Q}\left({f}_{i,w}^{*}\right),\:\:{K}_{i,w}={W}_{i}^{K}\left({f}_{1,w}^{*},{f}_{2,w}^{*},\dots\:,{f}_{k,w}^{*}\right),\:{\:V}_{i,w}={W}_{i}^{V}\left({f}_{1,w}^{*},{f}_{2,w}^{*},\dots\:,{f}_{k,w}^{*}\right),\:\) where \(\:\{{\text{W}}_{\text{i}}^{\text{Q}}{\}}_{\text{i}=1}^{\text{k}}\), \(\:\{{\text{W}}_{\text{i}}^{\text{K}}{\}}_{\text{i}=1}^{\text{k}}\), and \(\:\{{\text{W}}_{\text{i}}^{\text{V}}{\}}_{\text{i}=1}^{\text{k}}\)are the weight matrices responsible for generating the corresponding Query, Key, and Value sets. Here, \(\:\text{k}\) denotes the number of stages in both the encoder and decoder, and \(\:{\text{f}}_{\text{i},\text{w}}^{\text{*}}\) represents the feature map \(\:{\text{f}}_{\text{i}}^{\text{*}}\) along the width dimension. These matrices integrate information from all encoder stages to generate a comprehensive feature representation for each stage.
Subsequently, the output \(\:\{{\text{f}}_{\text{i}}^{\text{w}}{\}}_{\text{i}=1}^{\text{k}}\) of the W-Dimensional Axial-Attention is computed as follows:
This equation merges information across all stages of the encoder along the width dimension, effectively utilizing axial attention to enhance the detail and specificity of feature maps prior to the decoding phase.
H-dimensional CSAA
Continuing from the W-dimensional axial attention, we extend our methodology to the height (\(\:H\)) dimension by generating Query \(\:\{{Q}_{i,h}{\}}_{i=1}^{k}\), Key \(\:\{{K}_{i,h}{\}}_{i=1}^{k}\), and Value \(\:\{{V}_{i,h}{\}}_{i=1}^{k}\) sets based on the feature maps \(\:\{{f}_{i}^{w}{\}}_{i=1}^{k}\) processed in the previous step:\(\:{Q}_{i,h}={W}_{i}^{Q}\left({f}_{i,h}^{w}\right),\) \(\:{K}_{i,h}={W}_{i}^{K}\left({f}_{1,h}^{w},{f}_{2,h}^{w},\dots\:,{f}_{k,h}^{w}\right),\) \(\:{V}_{i,h}={W}_{i}^{V}\left({f}_{1,h}^{w},{f}_{2,h}^{w},\dots\:,{f}_{k,h}^{w}\right),i=\text{1,2},.,k,\) where \(\:\{{W}_{i}^{Q}{\}}_{i=1}^{k}\), \(\:\{{W}_{i}^{K}{\}}_{i=1}^{k}\), and \(\:\{{W}_{i}^{V}{\}}_{i=1}^{k}\) are the weight matrices that facilitate the generation of Query, Key, and Value sets respectively. These matrices serve to synthesize information across the encoder stages, now focusing on the height dimension.
The outcome of this axial attention process is computed as follows:
This equation efficiently combines information from all stages of the encoder, both in width and height dimensions, to produce feature maps that are rich in detail and spatial context. This enhanced feature integration ensures that each stage of the decoder is informed by a comprehensive, multi-dimensional understanding of the input data, which is crucial for accurate segmentation of small medical objects.
Resizing back
To facilitate the decoding process, the output features\(\:{\:f}_{i}^{h}\) from the axial attention stages are resized to align with the dimensions of the corresponding decoder stage feature maps. This resizing involves adjusting both spatial and channel dimensions through convolution operations:
where σ represents the ReLU activation function, \(\:BN\) indicates batch normalization, and k denotes the number of stages in both the encoder and decoder. The resized feature maps, denoted as \(\:{f}_{i}^{attn}\), are then concatenated to the feature map of the corresponding decoder stage along the channel dimension.
This approach addresses the computational challenges often associated with traditional two-dimensional attention mechanisms45, which require substantial resources. By employing a two-stage one-dimensional attention structure within the CSAA, our model processes feature maps sequentially across both width and height dimensions, optimizing computational efficiency without compromising performance.
The CSAA is instrumental in enhancing the model’s ability to extract and utilize detailed information about small medical objects throughout the encoding and decoding processes. This method ensures that each decoder stage is equipped with comprehensive and relevant information from all encoder stages, thereby improving segmentation accuracy and efficiency. Through this integration, the CSAA strengthens the connection between the encoder and decoder, reinforcing the model’s overall performance in segmenting small medical objects.
MPS module
The MPS is designed to optimize the utilization of low-resolution features within the decoder, which are pivotal for their robust global perception capabilities. Previous models such as those detailed in Ronneberger et al.5, Chen et al.25, and Lou et al.27 often failed to fully capitalize on this globally perceptual information, resulting in significant data loss during convolution and upsampling stages. Our MPS addresses this deficiency by preserving and enhancing the information extracted from these low-resolution features throughout the decoding process.
Segmentation process
In the segmentation step of MPS, feature maps from each decoder stage are processed through dedicated segmentation heads tailored to their respective resolutions. This segmentation is conducted as follows:
where \(\:{f}_{i}^{d}\:\)is the feature map at decoder stage i, σ represents the ReLU function, \(\:BN\) denotes batch normalization, and S is the sigmoid function. These operations yield segmentation results at varying resolutions, which are crucial for detailed object analysis.
Upsampling and integration
Following segmentation, these results are upsampled using a neighbor interpolation method to align with the ground truth dimensions, thereby facilitating multi-precision segmentation:
This step ensures that each segmentation output matches the scale of the ground truth, enhancing the accuracy of the model’s output against actual data.
Supervision strategy
The MPS adopts a varied precision supervision strategy, recognizing that while low-resolution features offer substantial global insight, they lack finer local details. By applying a lower precision of supervision to these features, the module maintains an emphasis on global attributes without overfitting to local noise. This approach not only preserves the benefits of traditional single-segmentation heads but also enhances them by integrating broad-scale perceptual strengths, thereby significantly improving the model’s efficacy in segmenting small medical objects.
Loss function
In the context of small medical object segmentation, where there is a significant imbalance between positive and negative pixels, we employ a hybrid loss function that combines DiceLoss46 and Binary Cross-Entropy (BCE) Loss. This approach is designed to optimize our model’s performance by addressing both the spatial and class imbalance challenges inherent in such tasks. The loss function for the segmentation outputs at each decoder stage is defined as follows:
where i = 1,2,…,k indexes the decoder stages, \(\:{M}_{i}\:\)represents the predicted segmentation map at stage i, and Y is the ground truth. The hyperparameters \(\:{{\uplambda\:}}_{1}\) and \(\:{{\uplambda\:}}_{2}\) are used to balance the contributions of Dice Loss and BCE Loss, respectively.
To integrate the contributions from each decoder stage, the total model loss is computed as:
where the set of hyperparameters \(\:\{{{\upalpha\:}}_{\text{i}}{\}}_{\text{i}=1}^{\text{k}}\) weights the losses from different stages, acknowledging the varying precision of segmentation results across the decoder stages.
This formulation of the loss function ensures that the model is finely tuned to maximize accuracy across all stages of decoding, thereby improving the overall segmentation performance and addressing the unique challenges posed by small medical object segmentation. This balanced and stage-wise approach to loss computation helps to mitigate the potential for overfitting to less informative regions and emphasizes learning from critical features relevant to medical diagnostics.
(A) Samples of small medical objects from two datasets (S-HRD and S-Polyp). This panel displays examples from our datasets showing small medical objects, with emphasis on detailed views and annotated ground truths, to underscore the challenges in segmenting such minute features. (B) Methodological comparison. Panels (a) and (b), traditional methods5,6 merge encoder and decoder features by concatenation or addition, ending with a single segmentation head. Panel (c), advanced methods32,47 integrate attention mechanisms at single encoder stages but still use only one segmentation head. Panel (d), Our EFCNet employs the Cross-Stage Axial Attention Module (CSAA) for comprehensive feature integration across all encoder stages and the Multi-Precision Supervision Module (MPS) to refine outputs across decoder stages. (C) EFCNet architecture overview: Provides a comprehensive depiction of EFCNet, highlighting the integration of CSAA and MPS to mitigate information loss and enhance segmentation fidelity across encoder and decoder stages. (D) Cross-stage axial attention module (CSAA) details: Focuses on the mechanics of CSAA, showcasing how multi-stage encoder features are dynamically integrated and directed through the decoding process to enhance the detail and accuracy of the segmentation. (E) Visualization of EFCNet and Previous Methods on S-HRD and S-Polyp. Our EFCNet compared to Attn-UNet37 on S-HRD and SSFormer31 on S-Polyp. Green circles highlight where EFCNet successfully captures extremely small medical objects that previous methods did not detect. Yellow circles show EFCNet’s enhanced accuracy in segmenting the boundaries of small medical objects compared to previous methods. Red circles indicate areas where previous methods incorrectly segmented small medical objects, whereas EFCNet provides correct segmentation.
Experiment
Evaluation metrics
To assess the performance of our model relative to existing state-of-the-art methods, we utilize two widely recognized metrics. DSC, which quantifies the similarity between the predicted labels and the ground truth. It is mathematically defined as:
where P denotes the area covered by the predicted labels, and G represents the area covered by the ground truth labels. IoU, which measures the overlap between the predicted labels and the ground truth relative to their union. It is defined as:
where \(\:{S}_{i}\) is the intersection area between the predicted labels and the ground truth, and \(\:{S}_{u}\:\)is their union area. These metrics are critical for validating the accuracy and reliability of segmentation methods in medical imaging, providing a comprehensive evaluation of model performance.
Implementation details
Our experiments were conducted using an NVIDIA RTX A6000 GPU with 48GB of memory, utilizing the Stochastic Gradient Descent (SGD) optimizer with an initial learning rate of 0.01. The training procedure spanned 200 epochs with a batch size of 4. All input images were uniformly resized to 352 × 352 pixels. The architecture of our model includes four stages each in the encoder and decoder.
To fine-tune the loss function, we adjusted the weights of Dice Loss and BCE Loss using the hyperparameters \(\:{\lambda\:}_{1}\) = 0.7 and \(\:{\lambda\:}_{2}\) = 0.3. Additionally, to effectively manage the contributions from multi-precision segmentation outputs, we set the balancing coefficients \(\:{{\upalpha\:}}_{1}\) = 1.0, \(\:{\alpha\:}_{2}\) = 0.9, \(\:{\alpha\:}_{3}\) = 0.8, and \(\:{\alpha\:}_{4}\) = 0.7 for the respective stages of the decoder.
This experimental setup ensures rigorous testing and validation of our model under controlled and replicable conditions, allowing for reliable comparison of its performance against established benchmarks in medical image segmentation.
Comparative models
To benchmark our approach in small medical object segmentation, we compare against a diverse set of state-of-the-art models: 1.CNN-Based Methods: Includes foundational U-Net5 and its advanced variants Attention-UNet37, MSU-Net48, and CaraNet32. 2. Transformer-Based Methods: Incorporates TransFuse9, TransUNet30, SSFormer31, and Swin-UNet10, which utilize self-attention to enhance detail processing.3. SAM11 and the related works: SAM without any prompt, SAM with point, SAM with box and MedSAM35. Additionally, we assess the impact of model scale on segmentation accuracy by testing an expanded version of U-Net (U-Net-Large) with 12 layers in both encoder and decoder, exploring the relationship between model size and performance efficacy.
Quantitative results analysis
As illustrated in Table 1, EFCNet consistently surpasses previous state-of-the-art models in both DSC and IoU across all folds for the S-HRD and S-Polyp datasets.
For the S-HRD dataset, EFCNet shows a significant improvement with an average increase of 4.88% in DSC and 3.77% in IoU compared to earlier methods. In the S-Polyp dataset, the model achieves an enhancement of 3.49% in DSC and 3.25% in IoU. These results highlight EFCNet’s robust performance, particularly in handling datasets with smaller medical objects where other models underperform due to limited available information.
The analysis reveals that the smaller the medical objects in the datasets, the greater the relative improvement of EFCNet over previous state-of-the-art methods, underscoring its effectiveness in small medical object segmentation. Notably, scaling up the architecture of U-Net (U-Net-Large) results in only marginal gains, suggesting. that EFCNet’s superior performance is primarily driven by its innovative design rather than merely increased model size.
The integration of CSAA and MPS modules within EFCNet, although increasing computational costs, is justified by the significant performance gains in critical segmentation tasks. A detailed comparison of model costs and performance benefits is provided in the supplementary materials.
Visualization
Figure 1E presents visual comparisons of segmentation results using EFCNet against previous state-of-the-art methods on the S-HRD and S-Polyp datasets. As indicated by our quantitative analysis in Table 1, the prior state-of-the-art models include Attn-UNet37 for S-HRD and SSFormer26 for S-Polyp. The visual results distinctly highlight several advantages of our EFCNet:1. Precision in Detecting Small Objects: EFCNet demonstrates superior ability to capture extremely small medical objects, which are clearly marked within the green circled areas in Fig. 1E.2. Boundary Accuracy: Our method exhibits enhanced accuracy in delineating the boundaries of small medical objects, as shown in the yellow circled areas.3. Reduced False Positives: EFCNet is markedly less prone to incorrectly segmenting the background as part of the medical objects, as indicated by the red circled areas.
These visual outcomes underscore the effectiveness of the CSAA and MPS modules. CSAA effectively utilizes local information from low-level features in the encoder to refine the segmentation details, while MPS leverages global perceptual insights from low-resolution features early in the decoder, significantly improving the detection and segmentation of small medical objects.
Ablation studies
We conducted a series of ablation experiments on the S-HRD and S-Polyp datasets to confirm the individual and synergistic effects of the CSAA and MPS modules on segmentation performance.
Initially, CSAA and MPS were integrated into the U-Net backbone separately to assess their standalone contributions. The results, presented in Table 2, reveal that each module significantly enhances segmentation accuracy. Moreover, the concurrent application of both modules leads to even greater improvements, demonstrating their combined efficacy. To explore the optimal configuration for CSAA, we manipulated the number of encoder stages involved in feature aggregation: The ‘Concat-One’ configuration merges feature from each encoder stage directly to the corresponding decoder stage without additional modifications; The ‘AA-One’ setup applies axial attention to features from a single encoder stage; The ‘AA-All’ arrangement, which represents our final model configuration, aggregates features across all encoder stages. As indicated in Table 3, ‘AA-All’ outperforms the other configurations, underscoring the importance of extensive feature integration for enhancing segmentation performance.
Different levels of supervision within the MPS were evaluated by varying the number of connected segmentation heads: The ‘MPS-4’ configuration, which connects segmentation heads to all decoder stages, was tested against setups with fewer connections (‘MPS-3’, ‘MPS-2’, and ‘MPS-1’). As detailed in Table 4, increased supervision levels correlate positively with segmentation accuracy, confirming the benefits of comprehensive supervision in complex segmentation tasks.
The results from these ablation studies validate the effectiveness of CSAA and MPS in our model, particularly highlighting their roles in significantly improving the precision and reliability of medical object segmentation. The configurations tested affirm the model’s robustness and adaptability, making it particularly suited for challenging segmentation environments.
Conclusion
In this study, we introduced EFCNet, a novel framework specifically designed to enhance the segmentation of small medical objects in medical images. EFCNet addresses a common challenge in these tasks, significant information loss, by ensuring that features at every stage of the model are fully utilized. This is achieved through the integration of two key modules: the CSAA and MPS. These modules are strategically developed to minimize information loss during both the encoding and decoding phases, resulting in a substantial improvement in segmentation performance. Additionally, EFCNet establishes a new benchmark in the field of small medical object segmentation, offering a solid reference point for future research and development.
Our extensive experimental evaluation, conducted across two distinct datasets, demonstrates that the incorporation of CSAA and MPS enhances segmentation accuracy, allowing EFCNet to consistently outperform previous state-of-the-art methods. These results highlight the potential of EFCNet as a powerful tool in medical imaging, particularly in applications where accurate segmentation of small objects is essential.
However, it is important to acknowledge the limitations of our study. First, our method requires significant computational resources, which may pose challenges for its practical application in resource-constrained environments. Reducing the resource consumption of the model will be a key area of future work. Second, our approach primarily focuses on a single modality (medical images) and does not yet incorporate other modalities, such as natural language, to guide the segmentation process. Expanding the model to integrate multi-modal data will be another important direction for future research.
Despite these limitations, the insights gained from the development and evaluation of EFCNet provide valuable contributions to the field of medical image analysis and lay the groundwork for further advancements in small object segmentation.
Data availability
The datasets used and analysed during the current study available from Mr. Lingjie Kong on reasonable request.
References
Chung, Y. R. et al. Role of inflammation in classification of diabetic macular edema by optical coherence tomography. J. Diabetes Res. 2019, 1582 (2019).
Arthi, M., Sindal, M. D. & Rashmita, R. Hyperreflective foci as biomarkers for inflammation in diabetic macular edema: retrospective analysis of treatment Naïve eyes from South India. Indian J. Ophthalmol. 69 (5), 1197 (2021).
Huang, H. et al. Algorithm for detection and quantification of hyperreflective Dots on optical coherence tomography in diabetic macular edema. Front. Med. 8, 688986 (2021).
Qin, S. et al. Hyperreflective foci and subretinal fluid are potential imaging biomarkers to evaluate anti-vegf effect in diabetic macular edema. Front. Physiol. 2337, 1661 (2021).
Ronneberger, O., Fischer, P. & Brox, T. U-net: convolutional networks for biomedical image segmentation. In Medical Image Computing and Computer-Assisted Intervention—MICCAI 2015 234–241 (Springer, 2015).
Xiao, X., Lian, S., Luo, Z. & Li, S. Weighted res-unet for high-quality retina vessel segmentation. In 2018 9th International Conference on Information Technology in Medicine and Education (ITME), 327–331 (2018).
Guan, S., Khan, A. A., Sikdar, S. & Chitnis, P. V. Fully dense Unet for 2-d sparse photoacoustic tomography artifact removal. IEEE J. Biomed. Health Inf. 24 (2), 568–576 (2020).
Jha, D. et al. Resunet++: An advanced architecture for medical image segmentation. (2019).
Zhang, Y., Liu, H., Hu, Q. Transfuse Fusing Transformers and CNNs for medical image segmentation. In Medical Image Computing and Computer-Assisted Intervention—MICCAI 2021 14–24 (Springer, 2021).
Cao, H. et al. Swin-unet: Unet-like pure transformer for medical image segmentation. In European Conference on Computer Vision. 205–218 (Springer, 2022).
Kirillov, A. et al. Segment anything. arXiv preprint arXiv:2304.02643 (2023).
Jha, D., Riegler, M. A., Johansen, D., Halvorsen, P. & Johansen, H. D. Doubleu-net: A deep convolutional neural network for medical image segmentation. In 2020 IEEE 33rd International Symposium on Computer-Based Medical Systems (CBMS), 558–564 (IEEE, 2020).
Cheng, H. K., Chung, J., Tai, Y. W. & Tang, C. K. Cascadepsp: Toward class-agnostic and very high-resolution segmentation via global and local refinement. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 8890–8899 (2020).
Poudel, R. P. K., Bonde, U., Liwicki, S., Zach, C. & Contextnet Exploring context and detail for semantic segmentation in real-time. arXiv preprint arXiv:1805.04554 (2018).
Huang, H. et al. Unet 3+: A full-scale connected unet for medical image segmentation. In ICASSP 2020–2020 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP), 1055–1059 (IEEE, 2020).
Chen, L. C., Papandreou, G., Kokkinos, I., Murphy, K. & Yuille, A. L. Semantic image segmentation with deep convolutional nets and fully connected crfs. arXiv preprint arXiv:1412.7062 (2014).
Chen, L. C. et al. Semantic image segmentation with deep convolutional nets, atrous convolution, and fully connected Crfs. IEEE Trans. Pattern Anal. Mach. Intell. 40 (4), 834–848 (2017).
Zhang, Z. et al. Enhancing feature fusion for semantic segmentation. In Proceedings of the European Conference on Computer Vision (ECCV), 269–284 (2018).
Bernal, J. et al. Wm-dova maps for accurate polyp highlighting in colonoscopy: validation vs. saliency maps from physicians. Comput. Med. Imaging Graph.. 43, 99–107 (2013).
Chen, L. et al. DRINet for medical image segmentation. IEEE Trans. Med. Imaging. 37 (11), 2453–2462 (2018).
Gu, Z. et al. CE-Net: context encoder network for 2D medical image segmentation. IEEE Trans. Med. Imaging. 38 (10), 2281–2292 (2019).
Hatamizadeh, A. et al. UNETR: Transformers for 3D medical image segmentation. In Proceedings of the IEEE/CVF Winter Conference on Applications of Computer Vision, 574–584 (2022).
Milletari, F., Navab, N. & Ahmadi, S. A. V-net: Fully convolutional neural networks for volumetric medical image segmentation. In 2016 Fourth International Conference on 3D Vision (3DV), 565–571 (IEEE, 2016).
Valanarasu, J. M. J. & Patel, V. M. UNeXt: MLP-based rapid medical image segmentation network. In Medical Image Computing and Computer Assisted Intervention—MICCAI 2022: 25th International Conference, Singapore 23–33 (Springer, 2022).
Agarwal, R. et al. Deep quasi-recurrent self-attention with dual encoder-decoder in biomedical CT image segmentation. IEEE J. Biomed. Health Inform. (2024).
Agarwal, R. et al. Multi-scale dual-channel feature embedding decoder for biomedical image segmentation. Comput. Methods Progr. Biomed. 257108464 (2024).
Mandal, B. Optimization of quadratic curve fitting from data points using real coded genetic algorithm. In Emerging Technologies in Data Mining and Information Security: Proceedings of IEMIS 2020, vol. 1. 419–428 (Springer, 2021).
Agarwal, R. et al. Spiking neural network in computer vision: techniques, tools and trends. In International Conference on Advanced Computational and Communication Paradigms 201–209 (Springer Nature Singapore, 2023).
Chowdhury, A. et al. U-Net based optic cup and disk segmentation from retinal fundus images via entropy sampling. In Advanced Computational Paradigms and Hybrid Intelligent Computing: Proceedings of ICACCP 2021. 479–489 (Springer, 2022).
Chen, J. et al. TransUNet: Transformers make strong encoders for medical image segmentation. arXiv preprint arXiv:2102.04306. (2021).
Wang, J. et al. Stepwise feature fusion: Local guides global. In Medical Image Computing and Computer Assisted Intervention—MICCAI 2022: 25th International Conference, Singapore. 110–120 (Springer, 2022).
Lou, A., Guan, S., Ko, H., Loew, M. H. & CaraNet Context axial reverse attention network for segmentation of small medical objects. In Medical Imaging 2022: Image Processing, 81–92 (SPIE, 2022).
He, S., Bao, R., Li, J., Grant, P. E. & Ou, Y. Accuracy of segment-anything model (SAM) in medical image segmentation tasks. arXiv preprint arXiv:2304.09324. (2023).
Huang, Y. et al. Segment anything model for medical images. arXiv preprint arXiv:2304.14660. (2023).
Ma, J. & Wang, B. Segment anything in medical images. arXiv preprint arXiv:2304.12306. (2023).
Huang, L., Wang, W., Chen, J. & Wei, X. Y. Attention on attention for image captioning. In Proceedings of the IEEE/CVF International Conference on Computer Vision, 4634–4643 (2019).
Oktay, O. et al. Attention U-Net: learning where to look for the pancreas. arXiv preprint arXiv:1804.03999 (2018).
Shen, T. et al. Reinforced self-attention network: A hybrid of hard and soft attention for sequence modeling. ArXiv Preprint arXiv :180110296. (2018).
Sinha, A. & Dolz, J. Multi-scale self-guided attention for medical image segmentation. IEEE J. Biomed. Health Inf. 25 (1), 121–130 (2020).
Tao, A., Sapra, K. & Catanzaro, B. Hierarchical multi-scale attention for semantic segmentation. arXiv preprint arXiv:2005.10821. (2020).
Vaswani, A. et al. Attention is all you need. Adv. Neural Inf. Process. Syst. 30. (2017).
Woo, S., Park, J., Lee, J. Y. & Kweon, I. S. CBAM: Convolutional block attention module. In Proceedings of the European Conference on Computer Vision (ECCV) 3–19 (2018).
Zhang, H., Zu, K., Lu, J., Zou, Y. & Meng, D. EPSANet: An efficient pyramid squeeze attention block on convolutional neural network. In Proceedings of the Asian Conference on Computer Vision 1161–1177 (2022).
Yadav, A. C., Kolekar, M. H. & Zope, M. K. Modified recurrent residual attention U-Net model for MRI-based brain tumor segmentation. Biomed. Signal Process. Control. 102, 107220 (2025).
Wang, H. et al. Axial-DeepLab: Stand alone axial-attention for panoptic segmentation. In Computer Vision—ECCV 2020: 16th European Conference, Glasgow, UK. 108–126 (Springer, 2020).
Sudre, C. H., Li, W., Vercauteren, T., Ourselin, S. & Cardoso, M. J. Generalised dice overlap as a deep learning loss function for highly unbalanced segmentations. In Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support: Third International Workshop, DLMIA 2017, and 7th International Workshop, ML-CDS 2017, Held in Conjunction with MICCAI 2017, Québec City, QC, Canada, 240–248 (Springer, 2017).
Zhang, M. et al. Shape matters for infrared small target detection. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 877–886 (2022).
Su, R., Zhang, D., Liu, J. & Cheng, C. MSU-Net: Multi-scale U-Net for 2D medical image segmentation. Front. Genet. 12, 639930 (2021).
Author information
Authors and Affiliations
Contributions
Q.W., H.C., X.Y., W.L., and M.W. spearheaded the project, offering clinical expertise and guidance on the study design. L.K., C.X., and Y.F. were instrumental in designing the network architecture and developing the data/modeling infrastructure. They also managed the training and testing setups and conducted the statistical analysis. Q.W. and H.C. were responsible for collecting the datasets and segmenting the HRDs in OCT data. L.K. and Q.W. made equal contributions to this article.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kong, L., Wei, Q., Xu, C. et al. EFCNet enhances the efficiency of segmenting clinically significant small medical objects. Sci Rep 15, 12813 (2025). https://doi.org/10.1038/s41598-025-93171-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-93171-6