Abstract
Dysregulated RNA processing is crucial in nasopharyngeal carcinoma (NPC) progression. Our research aimed to evaluate the prognostic values of RNA-processing genes (RPGs) in NPC through bioinformatic analysis of the GSE12452 and GSE102349 datasets, identifying differentially expressed RNA-processing genes (DE-RPGs). A prognostic model was developed using univariate and multivariate Cox analysis, with effectiveness assessed by ROC analysis. The correlation between risk scores, immune characteristics, and chemotherapy sensitivity was also analyzed. Model gene expression was validated by RT-qPCR, Western Blot, and immunohistochemistry, alongside functional assays. Bioinformatics indicated that RNA binding motif protein 20 (RBM20) and LSM5 are prognostic RPGs, with the ROC curve confirming their predictive ability for survival. Significant differences in drug sensitivity were noted between high- and low-risk groups. Experimental validation showed LSM5 is overexpressed in NPC tissues, correlating with poorer prognosis, and its down-regulation inhibits cell proliferation and migration. Thus, LSM5 is identified as a new adverse biomarker in NPC, with implications for targeted therapy and prognosis improvement.
Similar content being viewed by others
Introduction
Nasopharyngeal carcinoma (NPC) usually occurs in the posterior wall of the nasopharynx and pharyngeal recess with distinct epidemiological, histopathological, clinical, and therapeutic features. According to an epidemiological survey, the new global 2018 NPC medical records had approximately 129,000 cases1,2, with an unbalanced geographical distribution, as over 70% of the new medical records were concentrated within East and Southeast Asia3. Concurrent chemoradiotherapy (CCRT) or Radiation Therapy are recognized as the main treatment for nasopharyngeal carcinoma (NPC). However, there are still many patients with local recurrence or distant metastasis due to radiotherapy or chemotherapy resistance, which eventually leads to treatment failure4,5. Consequently, further investigation focusing on the molecular mechanisms underlying recurrence or metastasis processes is necessary to characterize potential targets for maintaining or reestablishing the sensitivity to therapy and improving the prognosis of the patient.
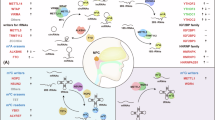
RNA processing is the process by which RNA transcripts are converted into mature RNA molecules6. Alterations in RNA processing, such as RNA splicing and polyadenylation, are major sources of variation in the cancer transcriptome and can play an important oncogenic role7. The abnormal expression of RNA processing factors can negatively affect mRNA transport and editing8. Furthermore, individual splicing changes and mutations in RNA splicing factors can be induced by cancer treatment, and such mutations can occur within single genes or in RNA processing factors themselves, which may affect the splicing of downstream target genes9. RNA processing factors also regulate intron removal and selective splicing of individual genes. Changes in alternative splicing are related to the initiation and progression of malignant tumors. Therefore, further research on RNA processing is necessary, as it may generate new opportunities for therapeutic intervention for cancer.
RNA binding proteins (RBPs) and RNA binding motifs (RBM) proteins are involved in the regulation of pre-mRNA splicing as they interact with pre-mRNA10. Studies have shown that RBPs, and CAPRIN1 in particular, are involved in the regulation of cell cycle, apoptosis, and drug sensitivity, suggesting that it may be a potential target for the treatment of NPC11. In essence, RBMS are key to understanding how RBPs regulate gene expression at the post-transcriptional level12. Xu et al.13 found that RBM47 largely regulates alternative splicing of pre-mRNAs in NPC cells, including those associated with cancer and thus, its upregulation in NPC specimens was associated with a poor prognosis in patients with NPC. In addition to RBM, the Like-Sm (LSM) protein may also be associated with NPCS. LSM proteins (such as Hfq) act as RNA chaperones and are involved in a variety of RNA metabolic reactions, including replication, stability, regulation of small RNA activity, and polyadenylation14. Studies have shown that LSM protein plays an important role in regulating mRNA function, maintaining RNA stability and post-transcriptional modification15. In addition, Ta et al.16 found that LSM4 was significantly correlated with tumorigenesis in breast cancer (BRCA) patients, suggesting that LSM4 may be a prospective prognostic indicator for BRCA. Therefore, it can be speculated that LSM protein may affect NPC cell cycle, stress response and other pathways by regulating RNA post-transcriptional modification and stability, thus promoting the occurrence and progression of tumors. RNA splicing-related circRNAs promote the proliferation and metastasis of NPC. circCAMSAP1 (highly expressed in NPC tissues) binds to SERPINH1 to enhance its expression, thus reducing the ubiquitination-based degradation of c-Myc. SRSF1 (RNA splicing factor) also promotes pre-mRNA transcription and back-splicing of CAMSAP1, forming a positive feedback induction of circPVT1 to induce NPC proliferation and metastasis17. circPVT1, which is highly expressed in NPC tissues, inhibits proteasomal degradation of c-Myc by binding to the E3 ubiquitin ligase β-TrCP and ultimately promotes invasion and metastasis of NPC cells by altering cytoskeleton remodeling and cell adhesion of NPC. Furthermore, c-Myc transcription upregulated the expression of SRSF1, which increased the generation of circPVT1 by coupling of transcription and splicing18. Luo et al.19 used transcriptome data to analyze alternative splicing events in 85 NPC samples, They found that GOLIM4-L and its splicing regulator RBFOX2 activate vesicles to mediate trafficking, which interacts with RAB26 to promote its tumorigenic potential in NPC cells. Combined with the effects of RNA processing factors that regulate alternative splicing events of single or multiple genes in NPC cells on the occurrence and progression of NPC, the abnormal expression of regulatory RPGs can drive important changes in the RNA expression profile observed in NPC.However, the role of RPGs in NPC is currently unexplored. Therefore, it is important to examine systematically the role of RPGs in NPC to create new opportunities for therapeutic intervention in NPC.
This study aims to analyze NPC expression profiles, identify prognostic markers related to RNA processing factors in NPC patients, and validate these findings. The goal is to develop more precise prognostic assessment tools for NPC patients and aid in identifying potential prognostic risks.
Results
Identification of differentially expressed RNA processing genes
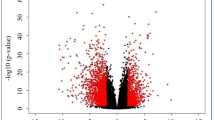
A total of 795 NPC-related DEGs were identified from the GSE12452 database (Fig. 1a,b, Supplementary Table S1). Among these, 293 genes were upregulated, and 502 genes were downregulated. Overlap analysis identified 10 common genes between NPC-related DEGs and RPGs: AHNAK2, ALYREF, FASTKD1, LSM5, PIH1D2, PNPT1, PUS7, RBM20, RBM24, and RRP15 (Fig. 1c). The expression levels of ALYREF, FASTKD1, LSM5, PNPT1, PUS7, and RRP15 were upregulated in the NPC group compared to healthy controls, while the expression levels of AHNAK2, PIH1D2, RBM20, and RBM24 decreased significantly (Fig. 1d). IHC results from the HPA database confirmed the expression of these genes (Fig. 1e).
Identification of differentially expressed genes (DEGs). (a) Volcano plot for DEGs identified between the tumor and normal sample groups. Orange, green, and black dots represent genes with upregulated, downregulated, and unaltered expression, respectively (Tumor/Normal). (b) Heat map for DEGs between tumor and normal sample groups. Green for normal samples; red for tumor samples. There was a positive correlation between gene expression and color change. The left dendrogram represents the cluster analysis of different genes from different samples. (c) Venn diagram for differentially expressed RNA-processing genes (DE-RPGs). (d) Heat map for expression patterns of 10 DE-RPGs. To interpret the legend, refer to Fig. 1b. (e) Expression of 10 differential RNA processing genes in normal nasopharyngeal tissues and nasopharyngeal carcinoma tissues (Immunohistochemical results from HPA database).
Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses elucidated the potential molecular mechanisms of NPC-related DE-RPGs in NPC. GO analysis identified 55 enriched biological process (BP) terms and 16 molecular function (MF) terms (Supplementary Table S2). The top five terms in the BP and MF categories are presented in Fig. 2a. In the BP category, these genes were predominantly associated with RNA processing activities, including splicing, catabolism, metabolism, processing, and translocation. Additionally, terms related to the cell cycle were significantly enriched. In the MF category, terms related to RNA binding were notably enriched. KEGG pathway analysis indicated that NPC-related DE-RPGs are involved in RNA degradation and spliceosome pathways (Fig. 2b, Supplementary Table S3).
Functional enrichment analysis of differentially expressed RNA-processing genes (DE-RPGs). (a) GO enrichment analysis of top five DE-RPGs. The proportion of the GO term is on the horizontal axis. The name of GO term is on ordinate. Black blots represent count, and color represents adjusted P-value. (b) KEGG enrichment analysis of DE-RPGs. Gene count is on the abscissa. The KEGG pathways are on the vertical axis. The color represents the adjusted P-value.
Construction and identification of the prognostic model
Univariate Cox regression analyses were conducted using the GSE102349 training set (n = 44) to assess the association of the identified DE-RPGs with disease-free survival (DFS) in patients with NPC. Table 1 shows that LSM5 (p = 0.022) and RBM20 (p = 0.035) were significantly correlated with DFS in patients with NPC. The regression coefficients of LSM5 and RBM20 were then calculated to construct the RNA-processing-related gene signature (Table 2).
\(1.6243\times expression of LSM5+(-0.9761)\times RBM20\) was used to perform risk scores for patients with NPC in the GSE102349-training set. The NPC samples in the GSE102349 training set were divided into high and low-risk groups based on the optimal cutoff value of 2.35 (Supplementary Fig. S1a). Figure 3a shows the risk curve and disease progression status (disease-free and progressive) of patients in the training set, indicating that patients with progressive disease had higher risk scores. The DFS of the low-risk group was significantly higher than that of the high-risk group (p = 0.00025) (Fig. 3b). Additionally, the area under the ROC curve (AUC) was 0.836 for one year and 0.652 for three and five years (Fig. 3c). Furthermore, in the high-risk group, LSM5 was expressed at a relatively high level, whereas the longer DFS rates of the low-risk group were associated with high expression of RBM20.
Construction and identification of the prognostic model. (a) Risk curves of patients in the high and low-risk groups in the training set. The abscissa of the upper and middle panels is the sample of patients sorted based on their risk scores. Risk scores increase from left to right; risk scores and survival times are in the appropriate order. The optimal risk score threshold and the corresponding number of patients are indicated by dashed lines. The lower panel represents the heatmap of the model gene expression. (b) Survival analysis for the training set. Ordinate represents survival rate, and abscissa represents disease-free survival. The red and blue curves indicate groups with high and low gene expression, respectively. (c) ROC curve for training set survival prediction, where ROC curve at three years overlaps with the five-year curve. (d) Risk curves of patients in the high and low-risk groups in the validation set. For legend interpretation, refer to Fig. 3a. (e) Survival analysis of the validation set. To interpret the legend, refer to Fig. 3b. (f) ROC curve for survival prediction of internal validation set, where the ROC curve at three years overlaps with the five-year curve.
The NPC samples in the GSE102349 testing set were divided into high-risk (n = 14) and low-risk (n = 30) groups based on the optimal cutoff value of 1.944 (Supplementary Fig. S1b). In the testing set, the results indicated a similar trend in survival (p = 0.023) (Fig. 3d,e), and the AUC for one, three, and five years was 0.740, 0.602, and 0.602, respectively (Fig. 3f).
Cox regression analysis based on the GSE102349 dataset indicated that the risk score could affect DFS in patients with NPC independently of stage characteristics (Supplementary Fig. S2). This implies that LSM5 and RBM20 may serve as independent prognostic factors for NPC.
The nomogram model constructed based on LSM5 and RBM20 had good predictive ability
Furthermore, the nomogram constructed on the basis of LSM5 and RBM20 showed good predictive ability (Supplementary Fig. S3a). The calibration curve for survival prediction closely aligned with the theoretical straight line (Supplementary Fig. S3b). In addition, the AUC value of the ROC curve exceeded 0.9, and the DCA curve showed that the net benefit level of the model was higher than the prediction of individual prognostic gene (Supplementary Fig. S3c-d). These results further confirmed the reliability of the prediction model.
Correlation of prognostic genes with immune cells
Next, we probed into the abundance of 24 types of immune cells in the high- and low-risk groups using ssGSEA (Fig. 4a). The Wilcoxon rank-sum test showed that the percentages of 21 immune cells were statistically different between the two risk groups (Fig. 4b). Furthermore, CD8 T cells, cytotoxic cells, Tem, Tcm, mast cells, eosinophils, T cells, pDC, and DC were considerably associated with the DFS rates in patients with NPC (Fig. 4c, Table 3). We also calculated Pearson’s correlations of these differential immune cells with RBM20 and LSM5 (Fig. 4d). In the low-risk group, DC had obvious and negative correlations with RBM20, CD8 + T cells, T cells, DC, eosinophils and Tem , whereas pDC negatively correlated with LSM5. However, no significant correlations between immune cells and prognostic genes were detected in the high-risk group.
Correlation of prognostic genes with immune cells. (a) Rank value of the 24 tumor-Infiltrating lymphocyte cells (TIICs) in risk groups. To interpret the legend, refer to Fig. 1b. (b) Expression of 24 TIICs in the risk group. The abscissa for 24 TIICs, the vertical coordinate for the fraction. Significant differences were marked with an asterisk (*p < 0.05, **p < 0.01, ***p < 0.001). (c) Univariate forest maps with significant differences in TIICS. (d) Correlation between DE-RPGs and immune cells.
Sensitivity analysis of chemotherapeutic agents
To further explore the guiding significance of the risk score for chemotherapy, we compared the half-maximal inhibitory concentration (IC50) of 138 chemotherapy drugs. The results suggested that, for 55 drugs, the high and low risk groups showed statistically significant differential sensitivity, as reflected in IC50 values (Supplementary Table S4). It can be inferred that patients in the low-risk group were more sensitive to these 55 drugs. The association of risk scores with 10 common chemotherapy drugs is demonstrated in Fig. 5. The differences between the two risk groups of the remaining 45 drugs are reviewed in Supplementary Fig. S4.
Sensitivity analysis of chemotherapeutic agents (partial display).
The prognostic genes showed consistent expression trends in GSE64634 and GSE12452
Subsequently, expression analysis of the prognostic genes was performed in the independent cohorts. The results found that the expression trends of the prognostic genes were consistent in the datasets GSE64634 and GSE12452: LSM5 was highly expressed in the disease group, while RBM20 was highly expressed in the control group. However, the expression difference of RBM20 in GSE64634 did not reach statistical significance, which may be related to the sample size (Supplementary Fig. S5a-b). The Protein–Protein Interaction (PPI) analysis showed that LSM5 interacted with 10 proteins, including LSM7, LSM3, and LSM6, while RBM20 interacted with 10 proteins, such as TNN and TCAP, to collaboratively exert its biological functions (Supplementary Fig. S5c).
Expression verification of RBM20 and LSM5
To verify the reliability of our predictions, we examined the expression of RBM20 and LSM5 in tissues and cell lines. Compared with normal nasopharyngeal tissues (n = 30), LSM5 expression was elevated and RBM20 expression was reduced in NPC tissues (n = 40) (Fig. 6a,b). Both protein and mRNA levels of LSM5 were elevated in nasopharyngeal carcinoma cells compared to NP69 cells (normal epithelial cells of the human nasopharynx), while RBM20 expression remained low (Fig. 6c–g).
Expression verification of RBM20 and LSM5. (a, b) expression of RBM20 and LSM5 in normal nasopharyngeal tissue and nasopharyngeal carcinoma tissue. (c–e) protein expression of RBM20 and LSM5 in normal nasopharyngeal epithelial cells(NP69) and nasopharyngeal carcinoma cells. The original blots of Western blot, the samples derive from the same experiment and that blots were processed in parallel. Original blots are presented in Supplementary Fig. 6. (f, g) RBM20 and LSM5 mRNA expression in normal nasopharyngeal epithelial cells and nasopharyngeal carcinoma cells. Significant differences were marked with an asterisk (*p < 0.05, **p < 0.01), ns indicates that there is no statistical difference.
Functional verification of LSM5 in nasopharyngeal carcinoma cells
The follow-up rate for the 40 NPC patients was 100%. The median survival time for the 16 patients with high LSM5 expression was 7.75 months, while the median survival time for the 14 patients with low LSM5 expression was 21 months (Fig. 7a,b). The chi-square value of the Log-Rank test was 15.93, and the survival curve is shown in Fig. 7c.
Functional verification of LSM5 in nasopharyngeal carcinoma cells. (a, b) expression of LSM5 in nasopharyngeal carcinoma. (c) Survival curve of nasopharyngeal carcinoma patients with different expression of LSM5. (d, e) Validation of protein expression after RNA interference with LSM5. The original blots of Western blot, the samples derive from the same experiment and that blots were processed in parallel. Original blots are presented in Supplementary Fig. 7. (f) Validation of mRNA expression after RNA interference with LSM5. (g-h) Colony formation assay analysis of cell survival and cloning ability after RNA interference with LSM5. (i, j) EdU assay analysis of cell proliferation after RNA interference with LSM5. (k, l) Migration assay analysis of cell migration capacity after RNA interference with LSM5. (m, n) Wound healing assay analysis of cell migration capacity after RNA interference with LSM5. Significant differences were marked with an asterisk (*p < 0.05; **p < 0.01), ns indicates that there is no statistical difference.
To investigate the role of LSM5 in the development of nasopharyngeal carcinoma, We efficiently transfected three small interfering Rnas targeting LSM5 into 5-8F and CNE2 cells, and verified the transfection effect with WB (Fig. 7d) and RT-qPCR. (Fig. 7e). Clonogenesis and EdU assays showed that LSM5 knockdown inhibited the viability and proliferation of 5-8F and CNE2 cells (Fig. 7f–j). Wound healing and invasion assays demonstrated that downregulation of LSM5 reduced the migration and invasion abilities of nasopharyngeal carcinoma cells (Fig. 7k–n).
Discussion
Dysregulated gene expression plays a key role in cancer, and RNA processing is the crucial step in altering gene expression. Abnormal expression of RNA processing factors in tumors can promote tumor occurrence, invasion and drug resistance20,21. Therefore, it is urgently needed to identify RNA processing factors associated with the prognosis of NPC to guide treatment and improve the prognosis of the patient.
In this study, we obtained 10 DE-RPGs from the GEO and AmiGO databases, including PNPT1, AHNAK2, ALYREF, PIH1D2, FASTKD1, RBM24, RBM20, PUS7, RRP15, and LSM5. Except for AHNAK2, PIH1D2, RBM20, and RBM24, the expression levels of the six other genes were considerably upregulated in the NPC group compared to healthy controls. PNPT1 is a highly evolved conserved gene encoding a protein localized in the mitochondrial membrane space, regulating mitochondrial homeostasis, mtRNA import, and aging-related inflammation22. Studies have shown that consumption of PNPT1 increases double-stranded RNA (dsRNA) accumulation, suggesting that PNPT1 helps regulate dsRNA levels. PNPT1 interacts with AGO1x, which may help buffer endogenous DsrNA-induced interferon response and prevent overactivation of interferon response and apoptosis of breast cancer cells, thereby maintaining cell balance23. At present, there have been no reports on the correlation between PNPT1 and NPC, and our findings provide a new perspective for the study of this disease. AHNAK2 serves as a novel prognostic biomarker in various cancers24,25,26,27. Studies have shown that AHNAK2 may promote the metastasis and proliferation of thyroid cancer through the PI3K/AKT signaling pathway28. In addition, AHNAK2 is highly expressed in lung adenocarcinoma tissues and plays a role through TGF-β/Smad3 signaling pathway, which may provide a new idea for the treatment of lung adenocarcinoma29. At present, no studies on AHNAK2 and NPC have been reported, and our findings provide a new target for the treatment of this disease. ALYREF is a nuclear export adaptor protein that couples histone mRNA processing and nuclear export30. Yanan Jin et al. suggested that ALYREF, as an m5C recognition factor, played an important role in NPC transfer31. In addition, ALYREF plays an important carcinogenic role in the progression of hepatocellular carcinoma (LIHC) by stabilizing EGFR mRNA and activating STAT3 signaling pathways32. We identified ALYREF as a gene associated with NPCS, which is consistent with previous findings. Further investigation of the mechanism of ALYREF in these diseases will provide an important theoretical basis for the development of new treatment strategies. PIH1D1, an unstable protein, enhances ribosomal RNA transcription in breast cancer by interacting with mTOR133. It was found that PIH1D1 regulates the stability of the R2TP complex by binding to RPAP3 isoform 1 to maintain its protein stability, thereby promoting cell survival34. At present, there is no literature report on the correlation between PIH1D1 and NPC, and its function in NPC can be further studied in the future to provide valuable reference for the treatment of this disease. FASTKD1 has a high mutation rate in various cancers and is an RNA biomarker for endometrial cancer35, also important for post-transcriptional mRNA stability and mitochondrial gene expression36. In addition, FASTKD1 may be involved in the interaction of cancer signaling pathways37, which provides the basis for future studies to use it as a diagnostic and therapeutic target. However, no studies have explored the relationship between FASTKD1 and NPCS, and our study provides new insights into this area. RBM24 is a splicing regulator downregulated in NPC, suppressing its progression by regulating miR-2538,39. In another study, miR-383 interfered with the activation of the NF-κB signaling pathway by inhibiting the expression of RBM24, thereby affecting chemotherapy resistance in lung adenocarcinoma cells40. Further study of the molecular mechanism of RBM24 is expected to provide a new direction for the treatment of NPC. RBM20 is crucial for RNA spliceosome formation and alternative splicing41, with low expression linked to poor liver cancer prognosis42. Our research shows low levels of RBM20 in NPC, suggesting it may be a protective factor. In addition, the MAPK/ELK1 signaling pathway influences the splicing of precursor mRNA by regulating the expression of RBM20, which may have an important impact on the function of genes associated with heart disease43. Our study identified RBM20 as a gene associated with NPC for the first time, providing a new theoretical basis for the study of this disease. PUS7 is an RNA modification enzyme influencing RNA metabolism44, with its pseudouridylation affecting codon translation in glioblastoma stem cell regulators45. Studies have shown that PUS7 interacts with Sirtuin 1 (SIRT1), stabilizes SIRT1, and then activates the Wnt/β-catenin signaling pathway, playing a carcinogenic function46. Our study shows for the first time that PUS7 is related to NPC, providing a new research target for this field. RRP15, located in the nucleolus, processes ribosomal RNA and regulates the cell cycle; its knockdown can induce apoptosis in tumor cell lines47. RRP15 has been found to promote hepatocellular carcinoma (HCC) migration by activating the LAMC2/ITGB4/FAK pathway, thus providing a new target for HCC therapy48. For the first time, we have found that RRP15 is associated with NPC, and further study of its role in NPC treatment will be of great significance. LSM5 is a component of the U4/U6-U5 tri-snRNP complex and pre-spliceosome B complex, involved in spliceosome assembly and splicing catalysis49.
Here, we evaluated the correlation of 10 DE-RPGs with patient survival and identified LSM5 and RBM20 as prognostic genes. LSM5 overexpression was considered a risk factor, whereas RBM20 was considered a protective factor for patients with NPC. We constructed a risk model with the LSM5 and RBM20 genes. To the best of our knowledge, our study is the first to develop a prognostic model for NPC based on RNA-processing factors. To assess the independence and applicability of LSM5 and RBM20, patients were regrouped according to different clinicopathological characteristics before performing a survival analysis. LSM5 and RBM20 were both confirmed to be independent prognostic factors for NPC (Supplementary Fig. S2). However, these two genes have not been previously reported in NPC. LSM5 plays a key role in U6 function, pre-mRNA splicing, and mRNA de-capping, which modulates various aspects of RNA metabolism50. The study found that LSM5 may promote colon cancer progression by down-regulating the expression of p53, CDKN1A, and TNFRSF10B51. In addition, RBM20, as a tissue-specific RNA binding protein, is involved in regulating post-transcriptional splicing of the FHOD3 gene52. In the study of Julia Kornienko et al., the pathogenic variant of RBM20 affected its normal splice regulation function by disrupting its interaction with TNPO3, causing it to mislocate to the cytoplasm53. Therefore, it can be speculated that LSM5 and RBM20 may play an important role in the biological processes such as proliferation, apoptosis and migration of NPC cells by regulating specific mRNA or non-coding RNA. Our findings are consistent with those of previous studies showing that DE-RPGs affect the occurrence and development of human diseases.
Immune response plays an important role in tumor growth, invasion and metastasis, so TIICs can be a target for radiotherapy and chemotherapy. Our research indicated that twenty-two immune cells were significantly differentially present among the risk groups. Among these, CD8 + T cells, cytotoxic cells, Tem, Tcm, mast cells, eosinophils, T cells, pDC, and DC were significantly correlated with DFS and were potential protective factors. The main roles of DC in the antitumor immune response include phagocytosis of dead tumor cells, capture, and presentation of tumor-associated antigens, and activation of various T cells, thus collectively stimulating a series of immune responses to kill tumor cells. In tumors, a decrease in DC count leads to a weaker antigen processing, which alters the proliferation and differentiation of T cells, negatively influences their tumor killing effect, and induces immune tolerance. In our study, RBM20 and LSM5 were strongly and negatively correlated with DC within the low-risk group. Therefore, DC cells are most likely to play a role as drug targets in nasopharyngeal carcinoma immunotherapy. In addition, the study found that RBM20 and its related gene variants may be involved in immune regulation in colorectal cancer by influencing the diversity and number of T cell responses54. In the study of Marek Lenart et al., downregulation of LSM5 may regulate the activation and function of CD8 T lymphocytes by affecting the stability and degradation of RNA, thus affecting the immune response and disease process of breast cancer patients55. This suggests that RBM20 and LSM5 are closely related to the immune microenvironment, and exploring the relationship between them and immune cells will provide more valuable insights for future disease research.
Furthermore, we performed RT-qPCR and western blot analysis using hunan NPC cell lines to verify mRNA and protein levels of prognostic genes. As expected, compared with normal nasopharyngeal tissues and cells, the expression of RBM20 and LSM5 in NPC tissues and cells is significantly different, corroborating the expression patterns obtained in the bioinformatic results.
LSM5 expression was upregulated in hepatocellular carcinoma tissues and identified as a new unfavorable prognostic biomarker56, consistent with our findings. However, functional studies of LSM5 have primarily focused on clock regulation in plants and humans57,58 and have not reported its role in cancer, particularly in NPC. Therefore, we investigated the function of LSM5 in NPC cells using gene interference techniques.
In our study, NPC patients with high LSM5 expression had a worse prognosis than those with low expression, with a median survival of only 7.75 months. Inhibiting LSM5 expression not only effectively reduced the viability and growth of nasopharyngeal carcinoma cells but also inhibited their migration and invasion. Our data fully validated the potential of LSM5 as a prognostic marker for NPC.
We investigated the potential guiding significance of the risk score for chemotherapy. Low-risk patients were sensitive to 55 chemotherapy agents. The IC50 values of AZD6244, Bicalutamide, BMS.536924, DMOG, FTI.277, JNK.9L, PD.0332991, RDEA119, and CI.1040 were lower in the low-risk group, suggesting that low-risk patients may benefit more from a chemotherapeutic regimen that includes these agents. Furthermore, we performed RT-qPCR and western blot analysis using seven cell lines (NP69, 5-8F, 6-10B, CNE-2Z, CNE1, S18, and S26) to verify mRNA and protein levels of prognostic genes. As expected, the expressions of RBM20 and LSM5 were significantly different within different types of NPC cells and nasopharyngeal epithelial cells, corroborating the expression patterns obtained in the bioinformatic results. In addition, the two prognostic genes exhibited a weak correlation in mRNA expression and were common to 27 KEGG signaling pathways related to NPC, such as cell cycle, DNA replication59, and spliceosome60, reflecting the potential functional similarity between the two.
Constructing a prognostic model of NPC using RPGs, our study displays two main strengths. Our model had high accuracy and good efficacy when tested using public datasets. Secondly, the expression levels of prognostic genes were verified at the horizontal tissue and cell levels. It is important to emphasize that we performed functional validation of LSM5 in nasopharyngeal carcinoma cells. However, we also recognize several limitations in our study. For example, due to the small sample size of NPC cases in the online database, the sample size was limited, and there was a lack of external cohorts with NPC-related survival data to verify the predictive performance of the prognostic risk models. Furthermore, while we identified 10 representative drugs to which low-risk patients in the Genomics of Drug Sensitivity in Cancer (GDSC) database may be sensitive, we acknowledge that IC50 data for these drugs in the seven cell lines used in our study are unavailable. Additionally, the drug sensitivity analysis needs to be further validated in both cell lines and larger clinical cohorts from different risk groups. Regarding the validation of diagnostic biomarkers, we also recognize the differences in protein or gene expression levels between tissues, cell lines, and serum or blood samples due to volume and local heterogeneity. This is an important consideration, and future studies should aim to validate our findings in human serum or blood samples. Finally, the significant functional correlation between RBM20 and LSM5 merits further investigation.
In conclusion, this paper is the first to reveal the influence of RNA processing genes on nasopharyngeal carcinoma. The RNA processing factors LSM5 and RBM20 could be promising prognostic marker genes for NPC. The functionality of LSM5 has been proven. These results show promise for targeted therapy and improved prognosis for patients with NPC.
Materials and methods
Data source
NPC-related transcriptome data were downloaded from the Gene Expression Omnibus (GEO) database. The GSE12452 dataset (31 NPC and 10 normal nasopharyngeal epithelial tissues)61,62,63 was used for the analysis of differentially expressed genes (DEGs), with the platform GPL570 (https://ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE12452). The GSE102349 dataset (88 patients with NPC)64 was used to construct and evaluate prognostic signatures with the platform GPL11154 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE102349). The GSE64634 dataset (4 nasopharyngeal carcinoma and 12 normal nasopharyngeal tissues)65 was used for prognostic genes expression analysis, with the platform GPL570. The RPGs were obtained from the AmiGO database (http://amigo.geneontology.org)66. Briefly, 6734 genes were obtained by searching the AmiGO database with the keyword “RNA processing” and subsequently defined as RPGs for this study. Typically, chip data undergo quality control (QC) after being generated, such as generating box plots and PCA plots to remove outlier samples. Subsequently, standardization is performed using the Robust Multi-array Average (RMA) algorithm to eliminate technical errors. The normalized data were modeled using the “limma” package (v 3.50.1)67 to identify DEGs. To control the false positive rate (FDR), the corrected p-values were directly output using the adjust.method = “BH” option in the topTable() function of limma.
Identification differentially expressed RNA processing genes
Differential expression analysis identified DEGs in NPC and normal nasopharyngeal epithelial tissues. We applied the R package “limma” based on the GSE12452 dataset to identify differential expression genes using the threshold p < 0.05 and |log2 fold change (FC)|> 1. Overlapping genes are genes common between two or more groups of elements. We used Jvenn (http://jvenn.toulouse.inra.fr/app/example.html) to identify common genes within the lists of DEGs and RPGs, which were considered NPC-related differentially expressed RNA processing genes (DE-RPGs) in this study. We used cluster Profiler68 to perform GO and KEGG69,70,71 functional enrichment analysis, which was aimed at finding the common functions and pathways of DE-RPG related to NPCs. The significance threshold was established at p < 0.05.
Construction and identification of the prognostic model
Gene signatures involved in RNA processing were screened and evaluated from the GSE102349 dataset. After matching the expression profiles of NPC-related DE-RPGs, we randomly divided 88 NPC patients into the GSE102349-training set (n = 44) and GSE102349-test set (n = 44) in a 1:1 ratio. Univariate Cox analysis for DFS was performed to identify NPC-related DE-RPGs with prognostic values. The significance threshold was established at p < 0.05.
In this study, the R package “survminer” was used to perform univariate and multivariate COX analysis of NPC-related DE-RPGs, and the threshold P value was < 0.05 to screen the genes related to the risk model of NPC. We obtained risk values for each patient by simulating gene expression and risk coefficients (coef) derived from multivariate Cox regression. Subsequently, patients were divided into high- and low-risk groups according to the optimal threshold of the risk score. Samples above the optimal threshold were the high-risk group and samples below the optimal threshold were the low-risk group. The optimal threshold was determined using histogram and dot plot. The histogram displayed the density distribution of high and low immune score groups based on the threshold, while the dot plot showed the standardized log-rank statistic for each threshold. We selected the threshold with the highest log-rank statistic as the optimal threshold (indicated by the dashed line). The formula used for the risk score was as follows.
Kaplan–Meier survival analysis was used to visualize the results of the prognostic survival analysis between groups. The validity of the risk model was evaluated using the ROC curve drawn with the “pROC” R package. Finally, the independent prognosis of the risk characteristics was verified through a univariate and multivariate Cox analysis based on risk scores and clinicopathological factors.
Construction and evaluation of nomogram
In order to assess the overall ability of prognostic genes to predict disease, a nomogram was constructed based on prognostic genes in all samples of the GSE102349 using the rms R package (version 6.5-0)72. Each prognostic gene was assigned a score, and the total score, which represented disease risk, was the sum of the individual scores, with higher scores indicating a higher risk of disease. Subsequently, the predictive performance of the model was further evaluated by ROC, DCA and calibration curves.
Characterization of immune cell profiles in risk groups
First, using the single-sample GSEA method, we obtained 24 immune-related gene sets for each sample in the GSE102349 dataset. Next, we classified samples into high- and low-risk groups based on the richness of immune-related information. To compare the differences in tumor-infiltrating immune cells between these risk groups, we used the Wilcoxon rank sum test. The significance threshold was set at p < 0.05.
In addition, to correlate the differential immune cell profile with the DFS rates in patients with NPC, we performed a univariate Cox regression analysis. Subsequently, we used Pearson’s correlation analysis to reveal immune cells correlated with prognostic genes. |Correlation coefficients (cor)|> 0.3 and p < 0.05 were established as significance thresholds.
Exploration of sensitivity to chemotherapeutic agents
The IC50 value can reflect tumor sensitivity to anticancer drugs. In order to understand the difference in response to drug therapy between high and low risk groups, predicting the IC50 value helps to identify which drugs patients may be more sensitive or resistant to, thus helping to optimize the treatment regimen. In this study, we used the “pRRophetic package”73 in R to predict the IC50 of 138 common chemotherapeutic agents in the risk groups using the Wilcoxon test. Results with p < 0.05 demonstrated statistical significance. Lower IC50 values implied greater sensitivity to chemotherapeutic agents.
Expression and interaction analysis of prognostic genes
To verify the difference in expression levels of prognostic genes between the disease and control groups, expression matrices of datasets such as GSE12452 and GSE64634 were obtained from the GEO database and analyzed. In addition, to explore the interactions of prognostic genes with other proteins, a protein interaction network was constructed based on the String website (https://cn.string-db.org/), and the screening threshold was set at 0.4.
Cell culture and transfection
Human NPC cells 5-8F, CNE2, S18, 6-10B, CNE1, S26 along with human nasopharyngeal epithelial cell NP69, were purchased from the BeNa Culture Collection, Hebei, China. The culture medium for human NPC cells and NP69 cell was RPMI-1640 supplemented with 10% fetal bovine serum and K-SFM containing 10% fetal bovine serum, respectively.
CNE2 and 5-8F cells were cultured under standard conditions to ensure that the cells were in good condition. The day before transfection, the cells were inoculated into 6-well plates with appropriate density, and 2 mL of complete medium (RPMIS-1640 supplemented with 10% fetal bovine serum) was added to each well, so that the cell density reached 70–80% at transfection time. The sirnas used in the experiment (siRNA1-LSM5, siRNA2-LSM5, siRNA3-LSM5 and siRNA-NC) were provided by German Biotechnology (Shanghai) Co., LTD., and their sequences are listed in supplementary Table S5. Lipofectamine 2000 was selected as the transfection reagent. 5 µL siRNA was mixed with 50 µL serum-free medium (RPMI-1640). At the same time, 5 µl Lipofectamine 2000 was mixed with 50µL serum-free medium (RPMI-1640) and left for 5 min at room temperature. Subsequently, the diluted siRNA was mixed with Lipofectamine 2000, gently blown and mixed, and left at room temperature for 15–20 min to form the siRNA-transfection reagent complex.
Next, the old medium in the 6-well plate was removed, the cells were gently washed with PBS once, and 1.8 mL of fresh complete medium was added to each well. The SirNA-transfection reagent complex (100 µL) was added drop by drop into the cell culture hole and the culture plate was gently shaken to ensure even mixing. The experimental groups were transfected with siRNA1-LSM5, siRNA2-LSM5, and siRNA3-LSM5, respectively, while a negative control group (siRNA-NC) and a blank control group (no addition of siRRNA) were set up.
After transfection, the 6-well plates were placed in cell incubators for further culture. The culture medium was replaced with fresh and complete medium 4–6 h after transfection. Cells were collected within 24 h of transfection, and transfection efficiency and gene expression levels were measured by Western Blot (WB) and RT-qPCR (RT-qPCR). Western Blot assay was used to extract total cell protein. LSM5 specific antibody was used to detect protein expression levels. The internal reference protein (beta-actin) is used for standardization. RT-qPCR assay: Total cell RNA was extracted and reverse-transcribed into cDNA. qPCR was performed using LSM5 specific primers. Internal reference genes, such as beta-actin, are used for standardization.
Tumor tissue collection and immunohistochemical (IHC) analysis in nasopharyngeal carcinoma patients
From January 2019 to December 2020, We retrospectively collected paraffin embedded specimens of 40 cases of nasopharyngeal carcinoma (NPC) and 30 cases of normal NPC tissues from Xiangya Hospital of Central South University. We conducted outpatient or telephone follow-up visits every 3–6 months for the 40 NPC patients to collect data on survival and death times. The follow-up cut-off points were death, loss to follow-up, or the end of the follow-up period (December 2023).
In this study, 70 paraffin-embedded tissues (40 nasopharyngeal carcinoma tissues and 30 normal nasopharyngeal tissues) were stained by immunohistochemical SP method. First, paraffin embedded tissue blocks were cut into 4 µm thick slices, attached to polylysine-treated slides, and baked in an oven at 60 °C for 1 h. Subsequently, xylene I and xylene II were dewaxed in turn, followed by gradient ethanol (100%, 95%, 80%, 70%) hydration, and finally rinsed with distilled water. For antigen repair, the slices were immersed in sodium citrate buffer (pH 6.0), heated in a pressure cooker to 95 °C for 15 min, cooled to room temperature and rinsed with PBS (pH 7.4) 3 times for 3 min each time. Next, the primary antibody was diluted according to the antibody instructions, and the drops were added to the slices, incubated overnight at 4 °C, rinsed with PBS, and the secondary antibody in PV-9000 universal two-step test kit (purchased from Zhongshan Jinqiao, Beijing, China) was incubated at room temperature for 30 min, and rinsed with PBS for 3 times. DAB color developing solution (purchased from Zhongshan Jinqiao, Beijing, China) was added to the section by drops, and the color development was observed under a microscope. When the positive signal turned brown and yellow, the reaction was immediately terminated with distilled water. Finally, the nucleus was re-stained with hematoxylin for 1–2 min, rinsed with tap water to return to blue, dehydrated with gradient ethanol, xylene transparent, neutral gum seal. The staining results were collected by microscope, and the average optical density (MOD) value was calculated by Image J image analysis software for quantitative analysis. Positive and negative controls were set up for each batch of experiments to ensure the reliability of experimental results.
RT-qPCR assay
First, total RNA is extracted from the cells. The control group consisted of normal nasopharyngeal epithelial cells (NP69), while the experimental group included different nasopharyngeal carcinoma cell lines such as CNE1, S18, S26, 5-8F, 6-10B, and CNE2. After cell growth to the logarithmic phase, RNA was extracted using TRIzol reagent (A33250, Thermo Fisher Technologies, Waltham, MA, USA). Next, mRNA reverse transcription kit (CW2569, Beijing Kangwei Century, China) was used to reverse transcribe the extracted RNA into cDNA, and the amount of RNA in the reaction system was 1 μg. The reverse transcription reaction was set up according to the kit instructions. After reverse transcription, quantitative PCR was performed using primers to measure the expression levels of LSM5, RBM20 and internal reference gene GAPDH. The relative mRNA expression of each gene was calculated by 2−ΔΔCT. The PCR primers for LSM5 and RBM20 were purchased from Tsingke Biotechnology Co., Ltd. Beijing, China. Human specific primers for LSM5, Forward primer: 5′-TGTCCGATTCCTTCACCTCC-3′, reverse primer 5′-CATTTGTCCACAAGCTCTCACC-3′. Human specific primers for RBM20, Forward primer: 5′-CCTCCCTTGAGCTCTCTCGC-3′, reverse primer 5′-AGCTTGGCGGCATTTTGGAT-3′. GAPDH, Forward primer: 5′-GAAAGCCTGCCGGTGACTAA-3′, reverse primer 5′-GCCCAATACGACCAAATCAGAG-3′.
Western Blot (WB) assay
Normal nasopharyngeal epithelial cells (NP69) were used as the control group, and the experimental group included different nasopharyngeal carcinoma cell lines (CNE1, S18, S26, 5-8F, 6-10b, and CNE2). All cells were cultured in medium at 37 °C, 5% CO₂. Cells in the logarithmic growth phase were collected, and protein concentrations were determined using the BCA kit (Lianke, Hangzhou, China). A total of 80 μg of protein per group was loaded onto 8% polyacrylamide gels and separated by SDS-PAGE. The proteins were then transferred onto PVDF membranes. Primary antibodies were incubated overnight at 4 °C (RBM20, 1:500, bs-9606R, Bioss, China; LSM5, 1:500, ab184568, Abcam, UK). Signals were visualized using a dual-color infrared laser imaging system (Odyssey CLx, LI-COR, New Jersey, USA). Endogenous Beta Actin (1:1000, 66009-1-Ig, Proteintech, China) and GAPDH (1:20,000, 10,494-1-AP, Proteintech, China) were used for normalization of protein expression.
Colony formation assay and 5-ethynyl-20-deoxyuridine (EdU) assay
After transfection of CNE2 and 5-8F cells, clonogenesis experiment was performed to evaluate the effect of LSM5 gene expression on cell clonogenesis. 48 h after transfection, the cells were inoculated on 6-well plates. Each group was inoculated with about 300 cells per well. Experimental Settings included blank control group (no siRNA was added), negative control group (siRNA-NC group), and experimental group (siRNA1-LSM5, siRNA2-LSM5, siRNA3-LSM5). The cells continue to be cultured at 37 °C, 5% CO₂, with medium changes every 2–3 days, usually 10 days. After the cells formed obvious clones (> 50 cells), the clones were stained and counted, and the effect of LSM5 gene inhibition on cell proliferation was analyzed according to the number of clones.
To evaluate the effect of LSM5 gene on the proliferation of CNE2 and 5-8F cells, cells transfected 48 h later were inoculated into 6-well plates with 1 × 104 cells per well and cultured for 24 h. Subsequently, 50 μM EdU solution was added and the cells were co-cultured for 2 h. After culture, the cells were fixed with 4% paraformaldehyde for 20 min. Then, each well was added with 500 μl of click-reaction solution and incubated at room temperature for 30 min to perform the EdU labeled reaction. To stain the nucleus, the cells are stained using DAPI solution. Finally, cell images were taken using a fluorescence microscope (Axio Observer 3, Carl Zeiss, Oberkochen, Germany) to record the number of EdU positive cells to assess cell proliferation. All reagents were tested using EdU cell proliferation imaging assay kit, which was purchased from Wuhan Elabscience Biotechnology Co., LTD.
Migration assay and wound healing assay
In order to evaluate the effect of LSM5 gene on the migration and healing ability of CNE2 and 5-8F cells, we conducted migration and scratch experiments, respectively. The experiment set up a blank control group (without adding any siRNA), SIRNA-NC group (negative control group) and experimental group (siRNA1-LSM5, siRNA2-LSM5, siRNA3-LSM5).
In the migration experiment, the cells transfected 48 h later were inoculated in the upper chamber of the Transwell chamber, with about 1 × 104 cells per well, and cultured for 24 h until the cells were stable and attached. Subsequently, a medium containing 15% fetal bovine serum was added to the lower compartment to induce cell migration. After 24 h of culture, the non-migrating cells in the upper chamber were gently removed with a cotton swab, and then the cells that migrated to the lower chamber were fixed with 4% paraformaldehyde. Migrating cells were stained with crystal violet solution for 15 min and images were taken through a microscope to count the number of cells migrating to the lower chamber, thereby assessing cell migration ability.
In the scratch experiment, 48 h after transfection, the cells were inoculated into 6-well plates with about 1 × 106 cells per well, and cultured to 70% fusion. Use a 200 μL sterile suction head to make even scratches on the cells. The cells were placed back in an incubator at 37 °C and 5% CO₂ for further culture. At the beginning of the scratch experiment (i.e. 0 h after scratch formation), an initial image of the scratched area is taken using a microscope. Images were then taken 24 h a day to see how the cells were healing. By measuring the width of the scratched area, the healing rate was calculated, the cell healing ability was assessed, and the differences between different groups were compared.
Ethics declarations
This retrospective study was carried out using the opt-out method for the case series of our hospital. The study was approved by the Medical Ethics Committee of Xiangya Hospital, Central South University (approval no. 202405111) and was conducted in accordance with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. Informed consent was waived by our Institutional Review Board (Medical Ethics Committee of Xiangya Hospital, Central South University) because of the retrospective nature of our study.
Statistical analysis
All data in this study were analyzed using R software (version 3.6.1). The significance thresholds are described in the corresponding sections. Differences between groups were analyzed using t-tests, while one-way analysis of variance (ANOVA) was used to compare multiple groups. Data are expressed as mean ± SD. A p-value of < 0.05 was considered statistically significant. Each experiment was repeated three times (n = 3).
Data availability
The datasets analyzed during the current study are available in the [Gene Expression Omnibus] repository [https://www.ncbi.nlm.nih.gov/geo, GSE12452 and GSE102349] and [AmiGO] repository [http://amigo.geneontology.org, “RNA processing”].
Change history
30 September 2025
This article has been retracted. Please see the Retraction Notice for more detail: https://doi.org/10.1038/s41598-025-20783-3
Abbreviations
- NPC:
-
Nasopharyngeal carcinoma
- RPG:
-
RNA processing gene
- DE-RPG:
-
Differentially expressed RNA processing gene
- GEO:
-
Gene expression comprehensive database
- ROC:
-
Receiver operator characteristic
- GSEA:
-
Gene cluster enrichment analysis
- IC50:
-
Half-maximal inhibitory concentration
- FBS:
-
Foetal bovine serum
- GO:
-
Gene ontology
- MF:
-
Molecular functions
- CC:
-
Cellular components
- BP:
-
Biological processes
- KEGG:
-
Kyoto encyclopedia of genes and genomes
- DFS:
-
Disease-free survival
References
Zhou, X., Liu, P. & Wang, X. Temporal lobe necrosis following radiotherapy in nasopharyngeal carcinoma: New insight into the management. Front. Oncol. 10, 593487 (2020).
Campion, N. J., Ally, M., Jank, B. J., Ahmed, J. & Alusi, G. The molecular march of primary and recurrent nasopharyngeal carcinoma. Oncogene 40, 1757–1774 (2021).
Kang, Y. et al. Advances in targeted therapy mainly based on signal pathways for nasopharyngeal carcinoma. Signal Transduct. Target. Ther. 5, 245 (2020).
Guan, S., Wei, J., Huang, L. & Wu, L. Chemotherapy and chemo-resistance in nasopharyngeal carcinoma. Eur. J. Med. Chem. 207, 112758 (2020).
Xu, C. et al. Chemoradiotherapy versus radiotherapy alone in Stage II nasopharyngeal carcinoma: A systemic review and meta-analysis of 2138 patients. J. Cancer 8, 287–297 (2017).
Manning, K. S. & Cooper, T. A. The roles of RNA processing in translating genotype to phenotype. Nat. Rev. Mol. Cell Biol. 18, 102–114 (2017).
Tollervey, D. & Caceres, J. F. RNA processing marches on. Cell. 103, 703–709 (2000).
Obeng, E. A., Stewart, C. & Abdel-Wahab, O. Altered RNA processing in cancer pathogenesis and therapy. Cancer Discov. 9, 1493–1510 (2019).
Bradley, R. K. & Anczuków, O. RNA splicing dysregulation and the hallmarks of cancer. Nat. Rev. Cancer 23, 135–155 (2023).
He, S., Valkov, E., Cheloufi, S. & Murn, J. The nexus between RNA-binding proteins and their effectors. Nat. Rev. Genet. 24, 276–294 (2023).
Yang, T., Huang, L., Qin, H. & Mai, S. STRESS granule-associated RNA-binding protein CAPRIN1 drives cancer progression and regulates treatment response in nasopharyngeal carcinoma. Med. Oncol. 40, 47 (2022).
Ray, D. et al. Nature. 499, 172–177 (2013).
Xu, X. C. et al. RNA-binding motif protein RBM47 promotes tumorigenesis in nasopharyngeal carcinoma through multiple pathways. J. Genet. Genomics 20, 595–605 (2021).
Le Derout, J., Boni, I. V., Régnier, P. & Hajnsdorf, E. Hfq affects mRNA levels independently of degradation. BMC Mol. Biol. 11, 17 (2010).
Tharun, S. Roles of eukaryotic Lsm proteins in the regulation of mRNA function. Int. Rev. Cell Mol. Biol. 272, 149–189 (2009).
Ta, H. D. K. et al. Potential therapeutic and prognostic values of LSM family genes in breast cancer. Cancers 13(19), 4902 (2021).
Wang, Y. et al. Splicing factor derived circular RNA circCAMSAP1 accelerates nasopharyngeal carcinoma tumorigenesis via a SERPINH1/c-Myc positive feedback loop. Mol. Cancer 21, 62 (2022).
Mo, Y. et al. Circular RNA circPVT1 promotes nasopharyngeal carcinoma metastasis via the β-TrCP/c-Myc/SRSF1 positive feedback loop. Mol. Cancer 21, 192 (2022).
Luo, C. L. et al. RBFOX2/GOLIM4 splicing axis activates vesicular transport pathway to promote nasopharyngeal carcinogenesis. Adv. Sci. 8, e2004852 (2021).
Barbieri, I. & Kouzarides, T. Role of RNA modifications in cancer. Nat. Rev. Cancer 20, 303–322 (2020).
Conn, V. M., Chinnaiyan, A. M. & Conn, S. J. RNA in cancer. Nat. Rev. Cancer 21, 22–36 (2021).
Sokhi, U. K. et al. Identification of genes potentially regulated by human polynucleotide phosphorylase (hPNPase old-35) using melanoma as a model. PloS One. 8, e76284 (2013).
Ghosh, S. et al. Prevention of dsRNA-induced interferon signaling by AGO1x is linked to breast cancer cell proliferation. EMBO J. 39, e103922 (2020).
Zhang, S. et al. AHNAK2 is associated with poor prognosis and cell migration in lung adenocarcinoma. BioMed Res. Int. 2020, 8571932 (2020).
Zheng, L., Li, S., Zheng, X., Guo, R. & Qu, W. AHNAK2 is a novel prognostic marker and correlates with immune infiltration in papillary thyroid cancer. Int. Immunopharmacol. 90, 107185 (2021).
Dumitru, C. A. et al. AHNAK and inflammatory markers predict poor survival in laryngeal carcinoma. PloS One. 8, e56420 (2013).
Wang, M. et al. AHNAK2 is a novel prognostic marker and oncogenic protein for clear cell renal cell carcinoma. Theranostics 7, 1100–1113 (2017).
Xu, M. et al. AHNAK2 promotes the progression of differentiated thyroid cancer through PI3K/AKT signaling pathway. Curr. Cancer Drug Targets 24, 220–229 (2024).
Liu, G., Guo, Z., Zhang, Q., Liu, Z. & Zhu, D. AHNAK2 Promotes migration, invasion, and epithelial-mesenchymal transition in lung adenocarcinoma cells via the TGF-β/Smad3 pathway. OncoTargets Ther. 13, 12893–12903 (2020).
Fan, J. et al. ALYREF links 3’-end processing to nuclear export of non-polyadenylated mRNAs. EMBO J. 38, e99910 (2019).
Jin, Y. et al. ALYREF promotes the metastasis of nasopharyngeal carcinoma by increasing the stability of NOTCH1 mRNA. Cell Death Disease 15, 578 (2024).
Nulali, J. et al. ALYREF-mediated RNA 5-methylcytosine modification promotes hepatocellular carcinoma progression via stabilizing EGFR mRNA and pSTAT3 activation. Int. J. Biol. Sci. 20, 331–346 (2024).
Kamano, Y. et al. PIH1D1 interacts with mTOR complex 1 and enhances ribosome RNA transcription. FEBS Lett. 587, 3303–3308 (2013).
Yoshida, M. et al. RPAP3 splicing variant isoform 1 interacts with PIH1D1 to compose R2TP complex for cell survival. Biochem. Biophys. Res. Commun. 430, 320–324 (2013).
Colas, E. et al. Molecular markers of endometrial carcinoma detected in uterine aspirates. Int. J. Cancer 129, 2435–2444 (2011).
Magraner-Pardo, L., Gobelli, D., de la Fuente, M. A., Pons, T. & Simarro, M. Systematic analysis of FASTK gene family alterations in cancer. Int. J. Mol. Sci. 22, 1133 (2021).
Magraner-Pardo, L., Gobelli, D., de la Fuente, M. A., Pons, T. & Simarro, M. Systematic analysis of FASTK gene family alterations in cancer. Int. J. Mol. Sci. 22, 11337 (2021).
Xia, R. M., Liu, T., Li, W. G. & Xu, X. Q. RNA-binding protein RBM24 represses colorectal tumourigenesis by stabilising PTEN mRNA. Clin. Transl. Med. 11, e383 (2021).
Hua, W. F. et al. RBM24 suppresses cancer progression by upregulating miR-25 to target MALAT1 in nasopharyngeal carcinoma. Cell Death & Disease 7, e2352 (2016).
He, B. et al. miR-383 increases the cisplatin sensitivity of lung adenocarcinoma cells through inhibition of the RBM24-mediated NF-κB signaling pathway. Int. J. Oncol. 59, 87 (2021).
Upadhyay, S. K. & Mackereth, C. D. Structural basis of UCUU RNA motif recognition by splicing factor RBM20. Nucleic Acids Res. 48, 4538–4550 (2020).
Yao, J. et al. Application value of whole exome sequencing in screening and identifying novel mutations of hypopharyngeal cancer. Sci. Rep. 13, 107 (2023).
Cai, H. et al. Angiotensin II influences pre-mRNA splicing regulation by enhancing RBM20 transcription through activation of the MAPK/ELK1 signaling pathway. Int. J. Mol. Sci. 20, 5059 (2019).
Guegueniat, J. et al. The human pseudouridine synthase PUS7 recognizes RNA with an extended multi-domain binding surface. Nucleic Acids Res. 49, 11810–11822 (2021).
Cui, Q. et al. Targeting PUS7 suppresses tRNA pseudouridylation and glioblastoma tumorigenesis. Nat. Cancer 2, 932–949 (2021).
Zhang, Q. et al. PUS7 promotes the proliferation of colorectal cancer cells by directly stabilizing SIRT1 to activate the Wnt/β-catenin pathway. Mol. Carcinog. 62, 160–173 (2023).
Dong, Z., Zhu, C., Zhan, Q. & Jiang, W. The roles of RRP15 in nucleolar formation, ribosome biogenesis and checkpoint control in human cells. Oncotarget. 8, 13240–13252 (2017).
Pan, T. et al. Knockdown of ribosome RNA processing protein 15 suppresses migration of hepatocellular carcinoma through inhibiting PATZ1-associated LAMC2/FAK pathway. BMC Cancer 24, 334 (2024).
Bertram, K. et al. Cryo-EM structure of a pre-catalytic human spliceosome primed for activation. Cell. 170, 701-713.e11 (2017).
He, W. & Parker, R. Functions of Lsm proteins in mRNA degradation and splicing. Curr. Opin. Cell Biol. 12, 346–350 (2000).
Mo, C. J. et al. Sm-like 5 knockdown inhibits proliferation and promotes apoptosis of colon cancer cells by upregulating p53, CDKN1A and TNFRSF10B. World J. Gastrointest. Oncol. 16, 2716–2726 (2024).
Lorenzi, P. et al. RNA-binding proteins RBM20 and PTBP1 regulate the alternative splicing of FHOD3. Int. J. Biochem. Cell Biol. 106, 74–83 (2019).
Kornienko, J. et al. Mislocalization of pathogenic RBM20 variants in dilated cardiomyopathy is caused by loss-of-interaction with Transportin-3. Nat. Commun. 14, 4312 (2023).
Schmit, S. L. et al. Germline genetic regulation of the colorectal tumor immune microenvironment. BMC Genomics 25, 409 (2024).
Lenárt, M. et al. Peripheral blood CD8+ T-lymphocyte immune response in benign and subpopulations of breast cancer patients. Int. J. Mol. Sci. 25, 6423 (2024).
Zhuang, H. et al. Identification of LSM family members as novel unfavorable biomarkers in hepatocellular carcinoma. Front. Oncol. 12, 871771 (2022).
Perez-Santángelo, S. et al. Role for LSM genes in the regulation of circadian rhythms. Proc. Natl. Acad. Sci. U.S.A. 111, 15166–15171 (2014).
Okamoto, M. et al. Sm-like protein-mediated RNA metabolism is required for heat stress tolerance in Arabidopsis. Front. Plant Sci. 7, 1079 (2016).
Zou, Z., Gan, S., Liu, S., Li, R. & Huang, J. Investigation of differentially expressed genes in nasopharyngeal carcinoma by integrated bioinformatics analysis. Oncol. Lett. 18(1), 916–926 (2019).
Yang, H., Beutler, B. & Zhang, D. Emerging roles of spliceosome in cancer and immunity. Protein Cell. 13, 559–579 (2022).
Dodd, L. E. et al. Genes involved in DNA repair and nitrosamine metabolism and those located on chromosome 14q32 are dysregulated in nasopharyngeal carcinoma. Cancer Epidemiol. Biomark. Prev. 15, 2216–2225 (2006).
Sengupta, S. et al. Genome-wide expression profiling reveals EBV-associated inhibition of MHC class I expression in nasopharyngeal carcinoma. Cancer Res. 66, 7999–8006 (2006).
Hsu, W. L. et al. Evaluation of human leukocyte antigen-A (HLA-A), other non-HLA markers on chromosome 6p21 and risk of nasopharyngeal carcinoma. PloS One. 7, e42767 (2012).
Zhang, L. et al. Genomic analysis of nasopharyngeal carcinoma reveals TME-based subtypes. Mol. Cancer Res. 15, 1722–1732 (2017).
Bo, H. et al. Upregulated long non-coding RNA AFAP1-AS1 expression is associated with progression and poor prognosis of nasopharyngeal carcinoma. Oncotarget. 6, 20404–20418 (2015).
Carbon, S. et al. AmiGO: Online access to ontology and annotation data. Bioinformatics. 25(2), 288–289 (2009).
Law, C. W. et al. RNA-seq analysis is easy as 1–2–3 with limma, Glimma and edgeR. F1000Research. 5, 1408 (2016).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: An R package for comparing biological themes among gene clusters. Omics J. Integr. Biol. 16, 284–287 (2012).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 28(11), 1947–1951 (2019).
Kanehisa, M., Furumichi, M., Sato, Y., Matsuura, Y. & Ishiguro-Watanabe, M. KEGG: Biological systems database as a model of the real world. Nucleic Acids Res. 53(D1), D672–D677 (2025).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28(1), 27–30 (2000).
Li, M. et al. Recognition of refractory Mycoplasma pneumoniae pneumonia among Myocoplasma pneumoniae pneumonia in hospitalized children: Development and validation of a predictive nomogram model. BMC Pulm Med. 23, 383 (2023).
Geeleher, P., Cox, N. & Huang, R. S. pRRophetic: An R package for prediction of clinical chemotherapeutic response from tumor gene expression levels. PloS One 9, e107468 (2014).
Acknowledgements
The authors would like to thank Editage (www.editage.com) for English language editing.
Funding
We gratefully acknowledge the patients who allowed their clinical data and tumor specimens to be used for research purposes. This work was supported by the National Natural Science Foundation of China (grant number 82405496), the Project of the Hunan Provincial Department of Education (grant number 23B0355, 23A0298), the Hunan Provincial Natural Science Foundation Project (grant numbers 2025JJ90022), the Domestic First-Class Discipline Construction Project of Chinese Medicine of the Hunan University of Chinese Medicine (grant numbers 2022ZYX03, 2022ZYX22), and Work Station Guidance Project of Academician Liu Liang, Hunan University of Chinese Medicine (grant numbers 22YS001).
Author information
Authors and Affiliations
Contributions
T.L. and GH.G. proposed the concept and design of the study. T.L., Y.G.T., and F.L.Z. performed the data collection. T.L., L.Y.D, Y.C. and S.Y.L. conducted all the experiments. T.L. drafted the manuscript. T.L. and G.H.G. performed data analysis and interpretation. Y.C.H. proofread and revised the manuscript. All authors have read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article has been retracted. Please see the retraction notice for more detail:https://doi.org/10.1038/s41598-025-20783-3
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lin, T., Deng, L., Liu, S. et al. RETRACTED ARTICLE: Identification of RNA processing factor LSM5 as a new adverse biomarker in nasopharyngeal carcinoma. Sci Rep 15, 9901 (2025). https://doi.org/10.1038/s41598-025-94968-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-94968-1
Keywords
This article is cited by
-
Unveiling the role of protein palmitoylation in gastric cancer diagnosis via machine learning
Discover Oncology (2025)