Abstract
Diarrheagenic Escherichia coli (DEC) are strains of Escherichia coli (E. coli) that can induce diarrhea symptoms in the host, as well as cause disease through contaminated food. To accurately and efficiently identify the five DEC types, this study retrieved the corresponding gene sequences from NCBI based on the virulence genes specified in Chinese national standards: invE, stx1, stx2, sth, stp, lt, aggR, astA, pic, bfpB, and escV. Probes and primers were designed for the conserved regions of these genes, and the amplification system, temperature, and other parameters were optimized. A TaqMan®single-plex real-time PCR assay was established for simultaneous detection of Enteropathogenic E. coli, Enteroinvasive E. coli, Enterotoxigenic E. coli, Enterohemorrhagic E. coli, and Enteroaggregative E. coli. The results demonstrated that the minimum detection limit for bacterial genomic DNA was 1.60 × 101 copies/μL (except for stx2 which was 1.60 × 102 copies/μL). The within-group variation rate ranged from 0.12 to 0.88%, while the between-group variation rate ranged from 0.67 to 1.62%. Moreover, R2 values for the standard curve generated by this method were between 0.999 and 1 with an amplification efficiency ranging from 98.4 to 100%. We evaluated a total of 122 clinical specimens using both conventional PCR and the qPCR method developed in this study. The findings indicated that the qPCR method exhibited high accuracy. Therefore, this study successfully developed a Taqman real-time fluorescence quantitative PCR assay with high specificity, sensitivity, repeatability, and amplification efficiency as well as significant regression effect. This assay can be utilized for clinical detection of DEC.
Similar content being viewed by others
Introduction
The DEC pathotypes differ regarding their virulence factors, pathogenesis, epidemiological characteristics, and are classified as EPEC, EIEC, ETEC, EHEC, and EAEC1,2,3. DEC is a common foodborne pathogen that poses a risk to public health. Timely identification of the source of DEC contamination is critical to effectively mitigate the impact of epidemics caused by this pathogen4. Therefore, it is essential to establish simple and efficient detection and typing techniques for diagnosing DEC infections accurately. These measures are necessary to ensure food safety and effectively control the spread of epidemics associated with DEC strains. The TaqMan method in real-time fluorescence quantitative PCR technology exhibits high specificity towards the target sequence and demonstrates excellent repeatability of results, thereby presenting evident application advantages in the rapid and large-scale screening of E. coli5,6.
Current conventional methods for the detection of E. coli typically require several days, particularly when E. coli concentrations are low. Enrichment is a commonly employed technique for bacterial isolation to enhance cell counts of target bacteria7. However, traditional culture methods also entail a slow and labor-intensive process that involves multiple steps and may necessitate definitive identification using ancillary techniques (such as biochemical, serological, nucleic acid-based methods), resulting in bacterial identification taking up to a week8. Culture-based approaches combined with other methodologies such as PCR9, immunoassay10, phage technology11, next-generation sequencing, biosensors, and matrix-assisted laser desorption ionization time-of-flight mass spectrometry (MALDI TOF MS) are increasingly utilized for the detection and identification of foodborne pathogens12.
Nucleic acid-based techniques like Real-Time PCR and viability PCR (vPCR) coupled with sequencing methods are more widely employed for pathogen detection compared to immunoassays and NGS-based approaches13. Quantitative Real-time PCR is a method employing fluorescent chemicals to measure the total amount of product after each polymerase chain reaction cycle in DNA amplification reactions,this enables quantitative analysis of specific DNA sequences14. The TaqMan method in real-time fluorescence quantitative PCR utilizes one or more fluorescent labeled oligonucleotide probes to detect the PCR products while employing fluorescence resonance energy transfer (FRET) to identify specific amplification products. The probe method exhibits high specificity towards the target sequence along with good reproducibility of results15. Fluorescence quantitative PCR technology is extensively utilized in the field of molecular biology and genetics to accurately quantify DNA, RNA, and other molecular substances in microorganisms, cells, viruses, parasites, and plants. In the detection of phytoplasma in plant phloem tissues, due to its low concentration, real-time PCR can be used to effectively detect and quantify phytoplasma16. For the detection of many viruses, real-time PCR also plays an excellent role in the detection of many viruses, such as polyomavirus, cytomegalovirus, Epstein-Barr virus, herpes simplex virus, etc.17,18. Moreover, the use of real-time PCR to detect DNA in the environment can also be effective in assessing parasitic infections in fish19.
Foodborne E. coli has long been recognized as one of the primary causative agents of severe clinical diarrhea in humans, livestock, poultry, pets, and wildlife. The objective of this project is to develop a TaqMan real-time PCR method for the detection of five DEC pathotypes, thereby contributing to the prevention and control of DEC prevalence and transmission for wildlife.
Materials and methods
DEC standard strains and plasmid
ETEC (CICC10667), EAEC (CICC24186), EHEC (CICC24187), EPEC (CICC24189) and EIEC (CICC24188) are all standard strains, which were procured from the China Research Institute of Food and Fermentation Industry (CICC) and the China Industrial Microor-ganisms Preservation Management Center. Klebsiella pneumoniae (ATCC700603), Pasteurella multocida (ATCC43137), Staphylococcus aureus (ATCC25923), Proteus mirabilis (ATCC12453), Pseudomonas acruginosa, Enterococcus faccalis, Clostridium perfringens (ATCC13124), and Salmonella (H9812) were all maintained by the Sichuan Key Laboratory of Endangered Wildlife Conservation Biology. The virulence genes of five types of DEC were identified as invE, stx1, stx2, sth, stp, lt, aggR, astA, pic, bfpB and escV. The recombinant plasmid was synthesized by combining partial gene fragments from these virulence genes, which were synthesized by Sangon Biotech (Shanghai, China).
Clinical samples
In this study, 122 clinical test samples were selected, which were received by the Key Laboratory of Conservation Biology for Endangered Wildlife of Sichuan Province in 2023. The types of samples encompassed feces, kidney tissues, liver tissues, and other biological materials, all derived from naturally deceased animals. The specific sources were as follows: 7 samples were from wild monkeys, 10 from wild blue sheep, 92 from giant pandas, and 13 from red pandas. All the samples were collected from the Chengdu Research Base of Giant Panda Breeding.
Main reagents and instruments
During the bacterial culture process, brain heart infusion agar (BHI agar) (Hopebio, Qingdao, China)and brain heart infusion broth (BHI broth) (Hopebio, Qingdao, China) were utilized. The composition and proportions (g/L) of BHI agar are as follows: 4 g beef brain extract powder, 4 g beef heart extract powder, 5 g peptone, 16 g casein peptone, 5 g sodium chloride, 2 g glucose, 2.5 g disodium hydrogen phosphate, and 13.5 g agar. The composition and proportions (g/L) of BHI broth are as follows: 10 g peptone, 12.5 g dehydrated calf brain extract powder, 5 g dehydrated beef heart extract powder, 5 g sodium chloride, 2 g glucose, and 2.5 g disodium hydrogen phosphate.During the agarose gel electrophoresis procedure, we employed a nucleic acid staining reagent (Thermo Fisher, Massachusetts, USA), 6 × loading buffer (Takara, Beijing, China), TAE buffer (Takara, Beijing, China), and agarose (Takara, Beijing, China). Omega Bio-Tek (Atlanta, USA) provided the plasmid extraction kit and bacterial genomic DNA extraction kit, fluorescent quantitative PCR probe master mix were supplied by Yeason company (Shanghai, China); The mean instruments used in the experiment included PCR instrument (Thermo Fisher Scientific, Massachusetts, USA), fluorescence quantitative PCR instrument(Bio-Rad, Berkeley, USA), UV spectrophotometer (Thermo Fisher Scientific, Massachusetts, USA), shaking incubator (CRYSTAL, Texas, USA), electrothermal biochemical incubator (CRYSTAL, Texas, USA), nucleic acid electrophoresis instrument (Tanon, Shanghai, China), gel imaging system (Tanon, Shanghai, China).
Design and synthesis of probe and primer
The virulence genes (Table 1) for detecting five types of DEC were selected in accordance with the Chinese national food safety standard (E. coli, GB 4789.6 - 2016), and their corresponding gene sequences were retrieved from NCBI. The conserved regions of each virulence gene were determined through sequence homology analysis using Genbank and BLAST software. Primers were designed within these conserved regions using Oligo and DNAstar software, followed by screening for optimal primers. Subsequently, probes specific to the primers were designed using Oligo software, incorporating 5′ 6-FAM as a fluorophore and 3’ BHQ1 as a quenching group (Table 2).
The isolation of bacterial DNA and plasmids
The preserved standard strains of EPEC, EHEC, ETEC, EAEC, and EIEC were each inoculated by transferring 200 μL of the frozen glycerol stock into 5 mL of brain heart infusion (BHI) nutrient broth medium. After overnight incubation at 37 °C with 180 rpm/min in the shaker for 16 h, a volume of 5 mL bacterial suspension was collected, and DNA extraction was performed following the instructions provided by the DNA extraction kit. A 1 μL aliquot of DNA was added to the cuvette of the UV spectrophotometer to assess its quality. After inoculating and culturing synthetic recombinant plasmids of DEC overnight as described above, plasmid standards were extracted according to the instructions provided by the plasmid extraction kit. The concentration and mass were determined using a UV spectrophotometer.
Primer validation
The DNA of 11 gene groups was utilized as a template for conventional PCR amplification (Table 3). Subsequently, the obtained results were submitted to Sangon Biological Company (Shanghai, China) for sequencing and comparison in order to validate the primer efficiency for invE, stx1, stx2, sth, tp, lt, aggR, astA, pic, bfpB, escV genes. During nucleic acid electrophoresis analysis, 5 µl of PCR product were subjected to electrophoresis on a 1% agarose gel (prepared with 0.5 g agarose and 50 ml TAE buffer) at 140 V for 35 min. Following electrophoresis, the resulting bands were visualized and analyzed using a gel imaging system.
Optimization of the reaction system and assessment of specificity
The matrix method was employed to optimize the concentrations of primers and probes in the amplification system. The probe concentrations (pmol/μL) tested were 2, 2.5, 3, 3.5, and 4, while the primer concentrations (pmol/μL) ranged from 2 to 4. Subsequently, a range of annealing temperatures from 54 to 61 °C was explored to optimize the annealing temperature for fluorescence quantitative PCR amplification.The amplification system and reaction procedures can be found in (Table 4).
The DNA of the standard strains currently present in the laboratory, including Klebsiella pneumoniae (ATCC700603), Pasteurella multocida (ATCC43137), Staphylococcus aureus (ATCC25923), Proteus mirabilis (ATCC12453), Pseudomonas acruginosa, Enterococcus faccalis, Clostridium perfringens (ATCC13124) and Salmonella (H9812) were extracted for analysis. Negative controls were also included in the experiment to ensure accuracy. To verify the specificity of our method, the amplification technique established was utilized within this study.
The assessment of sensitivity and establishment of the standard curve
After detecting the concentration of the extracted plasmid using an ultraviolet spectrophotometer, the plasmid copy number was calculated using a formula (copies/ml = 6.02 × 1023 copies/mole × C/MWg/mol) and subjected to tenfold gradient dilution ranging from 1.6 × 101 copies/μL to 1.6 × 108 copies/μL. Each group had three parallel controls, and assay sensitivity was determined based on the lowest concentration’s copy number in 40 cycles of amplification curve.
After performing a tenfold gradient dilution of the plasmid standards, plasmids with concentrations ranging from 1.6 × 102 to 1.6 × 108 copies/μL were selected. Three parallel controls were included in each group, and the established RT-qPCR detection method was utilized to generate a standard curve.
The assessment of repeatability
The standard materials with plasmid concentrations of 1.6 × 108 ng/μL, 1.6 × 107 ng/μL and 1.6 × 106 ng/μL were selected as templates for inter-group and intra-group repeatability tests. Three replicates were performed for each plasmid content to obtain the within-group coefficient of variation (CV). The three concentration gradients of plasmid standards were treated as a group, and three independent experimental replicates were conducted using the optimized real-time fluorescence quantitative PCR method to determine the between-group coefficient of variation (CV). Intra-group and inter-group coefficients of variation were calculated using the formula: CV% = SD/X × 100%.
Evaluation of the qPCR assay analyzing of clinical samples
Clinical samples, including fecal specimens, were inoculated into 5 mL of BHI broth. The inoculated broth was subsequently incubated in a bacterial shaker at 180 revolutions per minute for 16 h. Following this, bacterial DNA was extracted from the cultured broth according to the manufacturer’s instructions provided with the DNA extraction kit. The clinical samples collected by the laboratory in 2023 from Sichuan Province were subjected to detection using the fluorescence quantitative PCR method established in this study, while simultaneously employing the PCR detection method specified in the national food safety standard (E. coli, GB 4789.6 - 2016). A comparison was made between the results obtained from both methods to determine the detection rate of DEC and assess the accuracy of test outcomes.
Statement
This study and included experimental procedures were approved by the institutional animal care and use committee (IACUC) of Chengdu Research Base of Giant Panda Breeding (2023013). All animal housing and experiments were conducted in strict accordance with the procedures for care and use of laboratory animals. I confirm that all methods were performed in accordance with the relevant guidelines and regulations.
Results
Selection of primers and the fluorescent labeled probes
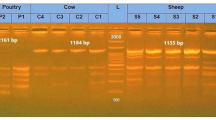
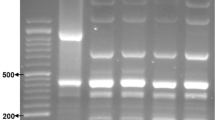
The forward and reverse primers, designed based on 11 genes, were utilized for PCR amplification. All PCR products were analyzed by electrophoresis, and the results shows that the sizes of all detection bands were consistent with the expected results (Fig. 1). Upon sequencing by Sangon Biological Company, the obtained result aligned with the target gene sequence, indicating successful amplification of the desired band.
PCR agarose gel electrophoresis results. 1 to 11 correspond to PCR bands for five DEC virulence genes (pic; sth; stx1; lt; escV; invE; bfpB; astA; stx2; stp; aggR). 12 serves as a negative control. M denotes a marker with a total length of 1000 base pairs (bp).
Optimization of RT-qPCR reaction conditions
After optimizing the primer concentration, probe concentration, and annealing temperature, the optimal working concentration for each primer was 3.5 pmol/μL, while the optimal working concentrations for each probe were as follows: invE (3 pmol/μL), stx1 (3.5 pmol/μL), stx2 (4 pmol/μL), sth (3.5 pmol/μL), stp (4 pmol/μL), It (3 pmol/μL), aggR (3.5 pmol/μL), astA (3.5 pmol/μL), pic (3.5 pmolμ/L), bfpB (3.5 pmol/μL), escV (3.5 pmol/μL). Additionally, the optimal annealing temperature was determined to be 57 °C.
Specificity of detection
Given the potential interference from certain intestinal bacteria on qPCR during clinical applications, we conducted specificity tests to evaluate the method’s robustness. For this purpose, we selected 11 bacterial strains for specificity analysis. Using the optimized primers and probes, we performed the experiments. The results indicated that the DNA of the 11 target genes served as positive templates for specificity testing, demonstrating high specificity with all non-specific amplification CT values exceeding 35 cycles. Notably, the bfpB and stx2 genes did not exhibit any non-specific amplification within 40 cycles, indicating superior specificity compared to the other nine genes (Fig. 2).
Specificity assay of reaction. (a–k) correspond to specific data for five DEC virulence genes (aggR; astA; bfpB; escV; invE; lt;pic; sth; stp; stx1; stx2). 1–2: Positive control; 3: Negative control; 4–19: Klebsiella pneumoniae, Pasteurella mul-tocida, Staphylococcus aureus, Proteus mirabilis, Pseudomonas aeruginosa, Enterococcus faecalis, Aeromonas veronii, Salmonella. The x-axis represents the Ct (Cycle Threshold) value, while the y-axis denotes RFU, which signifies the intensity of fluorescence signal.
Sensitivity of detection
In clinical applications, the sensitivity of a detection method is a critical evaluation metric. Therefore, this study aims to assess the sensitivity of the fluorescence quantitative detection method.By utilizing a positive standard plasmid with a concentration ranging from 1.60 × 108 to 1.60 × 101 copies/μL as the template, the optimized fluorescence quantitative PCR program was employed for amplification purposes. The final results of the amplification curve demonstrated that the method established in this experiment demonstrated a minimum detection limit of 1.60 × 101 copies, with the exception of gene stx2 which exhibited a limit of 1.60 × 102 copies (Fig. 3).
Sensitivity assay of reaction. (a–k) correspond to sensitivity data for five DEC virulence genes (aggR; astA; bfpB; escV; invE; lt;pic; sth; stp; stx1; stx2). The x-axis represents the Ct (Cycle Threshold) value, while the y-axis denotes RFU, which signifies the intensity of fluorescence signal.
Establishment of a standard curve
The specific amplification curve was obtained through real-time fluorescence quantitative PCR amplification of the gradient diluted standard (Fig. 4). The probe and matching primers exhibited a robust amplification relationship within the range of 1.6 × 109–1.6 × 103 copies/μL. Notably, aggR demonstrated an impressive amplification efficiency of 98.1%, with R2 = 1 and y-int = 40.345. The same pattern was seen for the amplification efficiency of astA (E = 99.2%, R2 = 1, y-int = 39.781), bfpB (E = 99.5%, R2 = 0.999, y-int = 41.621), escV (E = 99.6%, R2 = 1, y-int = 39.484), invE (E = 100%, R2 = 0.999, y-int = 40.755), lt (E = 99.8%, R2 = 1, y-int = 40.927), pic (E = 99.8%, R2 = 1, y-int = 39.682), sth (E = 99.5%, R2 = 1, y-int = 41.295), stp (E = 98.4%, R2 = 1, y-int = 40.798), stx1 (E = 98.8%, R2 = 0.999, y-int = 39.622), stx2 (E = 98.8%, R2 = 1, y-int = 40.875). The standard curve established in the experiment exhibited excellent linearity, significant regression impact, and high amplification efficiency.
Standard curve as was calculated by the BIO-RAD CFX Maestro qPCR software. (a–k) correspond to standard curve for five DEC virulence genes (aggR; astA; bfpB; escV; invE; lt;pic; sth; stp; stx1; stx2). The x-axis represents the logarithm of varying concentrations of gradient plasmids, while the y-axis denotes the Cq value, also referred to as the threshold cycle number.
Repeatability of the real-time PCR assay
To verify the stability of this detection method, the experiments on intra-group and inter-group repeatability were conducted using plasmid standards as templates, with concentrations of 1.6 × 108 copies/μL, 1.6 × 107 copies/μL and 1.6 × 106 copies/μL, respectively. The results revealed that the variability within the intra-group ranged from 0.07 to 0.88%, while the variability between inter-group ranged from 0.67 to 1.62% (Table 5). The CV is a measure of data dispersion. Generally, a CV below 10% indicates low variability in the data. The experimental results demonstrate that the CV for the method developed in this study was consistently below 1.63%. These findings indicate that the detection method established in this study exhibits excellent repeatability and stability.
Detection of clinical samples
The clinical samples were analyzed using both the TaqMan real-time fluorescent quantitative PCR method established in this study and the traditional PCR method specified by national standards for detecting five types of DEC. A total of 122 clinical samples were tested, yielding positive detection rates of 36 and 33 for the two methods, respectively, with three types of DEC virulence genes identified (Table 6). The detection frequencies of the astA and escV genes were largely consistent, whereas the detection frequency of the pic gene was marginally higher compared t o that obtained using traditional PCR. The results indicate that the detection method developed in this study demonstrates a high level of consistency with the national standard detection method while exhibiting greater sensitivity (Fig. 5).
The radar charts presented in (a, b) illustrate the clinical detection results of Taqman qPCR and conventional PCR, respectively. The various colors in the diagram represent different detection results of the virulence genes identified. Based on the graphical representation, it is evident that both detection methods exhibit a high level of consistency in their results.
Discussion
E. coli is one of the most important and widely studied etiologic agents of diarrhea worldwide20,21. However, not all E. coli strains isolated from diarrheal patients or animals are pathogenic because commensal E. coli also extensively colonize the gut22,23. Therefore, distinguishing pathogenic strains from nonpathogenic ones is critical for controlling DEC infections24. Unfortunately, molecular diagnostics do not work well for all DEC pathotypes, in particular due to issues defining molecular targets for EAEC25. In the selection of virulence genes, we conducted a comprehensive review of pertinent research reports26,27,28, and compared our findings with the testing standards outlined in Chinese national standard. A high degree of concordance was observed between the selected virulence genes in both sources. Given that this methodology is primarily intended for application within China, we opted to adopt the virulence genes specified in the Chinese national standards as the target genes for this study. In addition, high rates of asymptomatic carriage in epidemiological studies indicate inconsistent relationships between marker gene presence and diarrhea symptoms29. qPCR and ddPCR are extremely sensitive, allowing for the detection of very small amounts of pathogen, which may or may not represent biologically or clinically relevant infections30. Nonetheless, both established and emerging PCR-based methods are critical for detecting and diagnosing DEC pathotypes in both clinical and research settings.
At present, PCR-based molecular methods are widely employed for the detection and research of DEC, given their sensitivity, specificity, relative swiftness, and ease of application. In contrast to conventional PCR, which observes PCR products via gel electrophoresis, qPCR amplification is measured by fluorescent reporter molecules and quantifies the target in relation to a standard curve. Currently, some laboratories have developed real-time PCR detection methods for E. coli detection. The minimum detection concentration of avian E. coli can be detected by real-time PCR methods ranging from 1 × 102 to 4.69 × 102 copies/μL31,32. For porcine E. coli’s five virulence factors’ single fluorescent quantitative PCR assay, the minimum detection limit ranged from 3.05 × 101 to 1.20 × 102 copies/μL33. The sensitivity of our study’s method was determined as 1.60 × 102 copies while other genes had a higher sensitivity at 1.60 × 101 copies. Newer tools have also been utilized for DEC pathotype detection. Among them are Luminex and BioFire panels aimed at detecting and diagnosing a series of gastrointestinal pathogens34,35. Both the Luminex and BioFire platforms leverage known DEC pathotype targets to furnish rapid diagnostic outcomes in clinical contexts36. Other emerging approaches, such as digital droplet PCR (ddPCR) and Taqman array card technology37, are conducive to enhancing the amplification, detection, multiplexing, and automation capabilities for DEC pathotypes.
Furthermore, the method established herein also presents certain limitations. For example, within the reaction systems of different genes, there exist considerable disparities in fluorescence intensity: the minimum fluorescence intensity is merely 3000 RFU, while the maximum can reach 12,000 RFU. Additionally, during the experimentation, we observed that the stability of the experimental results in systems with lower fluorescence signals is marginally inferior to that in systems with higher fluorescence signals. Secondly, in the specificity experiment, we identified the possibility of false positives emerging after more than 35 cycles. Hence, although the qPCR method demonstrates superior sensitivity, in practical clinical applications, we ought to impose constraints on the interpretation criteria of the detection results to more effectively avoid false positive outcomes.
In the future, this method is slated to be incorporated into commercial detection kits in the form of prefabricated plates, not only further escalating the detection rate but also significantly simplifying the operational process. Even non-professional researchers can promptly commence the testing by merely adding the sample DNA into the corresponding wells, offering substantial facilitation for clinical applications.
Conclusion
In conclusion, this study successfully developed a qPCR method capable of accurately detecting five types of DEC. This novel approach demonstrates high sensitivity and specificity, providing a reliable tool for the detection and monitoring of DEC in clinical and research settings. The RT-qPCR detection approach established in this research has been successfully utilized for the routine surveillance of clinical samples from both giant pandas and red pandas at the Chengdu Research Base of Giant Panda Breeding. Previously, this base adopted the traditional PCR method as stipulated in the national standard (GB4789.6 - 2016) for detection. By contrast, the RT-qPCR method developed in this study exhibited remarkable superiority in terms of sensitivity and efficiency. Furthermore, the outcomes derived from the new method were highly congruent with those from the national standard detection procedures.
Data availability
The datasets used and analysed during the current study available from the first author Bingyu Xue.
References
Allocati, N., Masulli, M., Alexeyev, M. F. & Di Ilio, C. Escherichia coli in Europe: An overview. Int. J. Environ. Res. Public Health 10(12), 6235–6254. https://doi.org/10.3390/ijerph10126235 (2013).
Gomes, T. A. et al. Diarrheagenic Escherichia coli.. Braz. J. Microbiol. 47(Suppl 1), 3–30. https://doi.org/10.1016/j.bjm.2016.10.015 (2016).
Vila, J. et al. Escherichia coli: An old friend with new tidings. FEMS Microbiol. Rev. 40(4), 437–463. https://doi.org/10.1093/femsre/fuw005 (2016).
Manzanas, C. et al. Molecular testing devices for on-site detection of E. coli in water samples. Sci. Rep. 13(1), 4245. https://doi.org/10.1038/s41598-023-31208-4 (2023).
Molina, F. et al. Improved detection of Escherichia coli and coliform bacteria by multiplex PCR. BMC Biotechnol. 15, 1–9. https://doi.org/10.1186/s12896-015-0168-2 (2015).
Xue, Y., He, S., Li, M. & Qiu, Y. Development and application of four foodborne pathogens by TaqMan multiplex real-time PCR. Foodborne Pathog. Dis. https://doi.org/10.1089/fpd.2023.0134 (2024).
Nurliyana, M. et al. The detection method of Escherichia coli in water resources: A review. J. Phys. Conf. 995, 012065. https://doi.org/10.1088/1742-6596/995/1/012065 (2018).
Royer, C. et al. Comparison of metagenomic and traditional methods for diagnosis of E. coli enteric infections. MBio 15(4), e03422-e3423. https://doi.org/10.1128/mbio.03422-23 (2024).
Zimoń, B. et al. Novel multiplex-PCR test for Escherichia coli detection. Microbiol Spectr. 12(6), e0377323. https://doi.org/10.1128/spectrum.03773-23 (2024).
Zhuang, L. et al. Progress in methods for the detection of viable Escherichia coli. Analyst 149(4), 1022–1049. https://doi.org/10.1039/d3an01750h (2024).
Hong, B., Li, Y., Wang, W., Ma, Y. & Wang, J. Separation and colorimetric detection of Escherichia coli by phage tail fiber protein combined with nano-magnetic beads. Microchim. Acta 190(6), 202. https://doi.org/10.1007/s00604-023-05784-1 (2023).
Kabiraz, M. P., Majumdar, P. R., Mahmud, M. C., Bhowmik, S. & Ali, A. Conventional and advanced detection techniques of foodborne pathogens: A comprehensive review. Heliyon https://doi.org/10.1016/j.heliyon.2023.e15482 (2023).
Aladhadh, M. A review of modern methods for the detection of foodborne pathogens. Microorganisms 11(5), 1111. https://doi.org/10.3390/microorganisms11051111 (2023).
Hawkins, S. F. C. & Guest, P. C. Multiplex analyses using real-time quantitative PCR. Methods Mol. Biol. 1546, 125–133. https://doi.org/10.1007/978-1-4939-6730-8_8 (2017).
Dymond, J. S. Explanatory chapter: quantitative PCR. Methods Enzymol. 529, 279–289. https://doi.org/10.1016/B978-0-12-418687-3.00023-9 (2013).
Yusuf, A.-J., Aknadibossian, V., Jawhari, M., Tawidian, P. & Abrahamian, P. Real-time PCR protocol for phytoplasma detection and quantification. Methods Mol Biol. 1875, 117–130. https://doi.org/10.1007/978-1-4939-8837-2_9 (2019).
Xu, Y. et al. A novel multiplex real-time PCR assay for the detection of cytomegalovirus, Epstein-Barr virus, herpes simplex virus 1/2 and strategies for application to blood screening. Diagn. Microbiol. Infect. Dis. 109(1), 116234. https://doi.org/10.1016/j.diagmicrobio.2024.116234 (2024).
Arvia, R. et al. Droplet digital PCR (ddPCR) vs quantitative real-time PCR (qPCR) approach for detection and quantification of Merkel cell polyomavirus (MCPyV) DNA in formalin fixed paraffin embedded (FFPE) cutaneous biopsies. Virol. Methods 246, 15–20. https://doi.org/10.1016/j.jviromet.2017.04.003 (2017).
Berger, C. S. & Aubin-Horth, N. An eDNA-qPCR assay to detect the presence of the parasite Schistocephalus solidus inside its threespine stickleback host. J. Exp. Biol. 221(9), jeb178137. https://doi.org/10.1242/jeb.178137 (2018).
Pearson, J. S., Giogha, C., Wong Fok Lung, T. & Hartland, E. L. The genetics of enteropathogenic Escherichia coli virulence. Annu. Rev. Genet. 50, 493–513. https://doi.org/10.1146/annurev-genet-120215-035138 (2016).
GBD Diarrhoeal Diseases Collaborators. Estimates of global, regional, and national morbidity, mortality, and aetiologies of diarrhoeal diseases: A systematic analysis for the Global Burden of Disease Study 2015. Lancet Infect. Dis. 17(9), 909–948. https://doi.org/10.1016/S1473-3099(17)30276-1 (2017).
Riley, L. W. Distinguishing pathovars from nonpathovars: Escherichia coli.. Microbiol. Spectr. https://doi.org/10.1128/microbiolspec.ame-0014-2020 (2020).
Foster-Nyarko, E. & Pallen, M. J. The microbial ecology of . in the vertebrate gut. FEMS Microbiol. Rev. 46(3), fuac008. https://doi.org/10.1093/femsre/fuac008 (2022).
Bian, S. et al. Molecular serotyping of diarrheagenic Escherichia coli. with a MeltArray assay reveals distinct correlation between serotype and pathotype. Gut Microbes 16(1), 2401944. https://doi.org/10.1080/19490976.2024.2401944 (2024).
Jesser, K. J. & Levy, K. Updates on defining and detecting diarrheagenic Escherichia coli. pathotypes. Curr. Opin. Infect. Dis. 33(5), 372–380. https://doi.org/10.1097/QCO.0000000000000665 (2020).
Tsobeng, O. D. et al. Detection of blaTEM, blaOXA, blaCTX-M, and blaSHV genes of antibiotic resistance in diarrheagenic E coli causing enteric infection in hypertensive patients at Laquintinie Hospital, Littoral Region of Cameroon. J. Infect. Public Health 18(1), 102617. https://doi.org/10.1016/j.jiph.2024.102617 (2025).
Jesser, K. J. & Levy, K. Updates on defining and detecting diarrheagenic Escherichia coli pathotypes. Curr. Opin. Infect. Dis. 33(5), 372–380. https://doi.org/10.1097/QCO.0000000000000665 (2020).
Du, C. et al. Molecular serotyping of diarrheagenic Escherichia coli with a MeltArray assay reveals distinct correlation between serotype and pathotype. Gut microbes 16(1), 2401944. https://doi.org/10.1080/19490976.2024.2401944 (2024).
Liu, J. et al. Use of quantitative molecular diagnostic methods to identify causes of diarrhea in children: A reanalysis of the GEMS case-control study. Lancet (London, England) 388(10051), 1291–1301. https://doi.org/10.1016/S0140-6736(16)31529-X (2016).
Levine, M. M. & Robins-Browne, R. M. Factors that explain excretion of enteric pathogens by persons without diarrhea. Clin. Infect. Dis. 55(Suppl 4), S303–S311. https://doi.org/10.1093/cid/cis789 (2012).
Gao, Y. et al. Establishment and application of real-time fluorescence quantitative PCR assay for chicken Escherichia Coli. Hunan J. Anim. Sci. Vet. Med. 1, 31–35 (2024) (in Chinese).
Jinnan, C. et al. Establishment and application on a duplex FQ-PCR for detection of infectious bursal disease virus and pathogenic Escherichia coli.. China Poultry 45(09), 31–37. https://doi.org/10.16372/j.issn.1004-6364.2023.09.006 (2023).
Xia, C. Establishment and Preliminary Application of Single and Multiplex qPCR Methods for Detection of Five Virulence Factors of Porcine E. coli. Dissertation (Huazhong Agricultural University, 2022). (in Chinese)
Punia, M. et al. Development of a multiplexed Luminex assay for simultaneous detection of enteric viruses in cattle. Vet. Res. Forum 15(1), 13–19. https://doi.org/10.30466/vrf.2023.2005728.3925 (2024).
Suleiman, M., Iqbal, M., Tang, P. & Pérez-López, A. Comparison of QIAstat-Dx and BioFire FilmArray gastrointestinal panels in a pediatric population. Microorganisms 12(11), 2282. https://doi.org/10.3390/microorganisms12112282 (2024).
Huang, R. S. et al. Performance of the Verigene® enteric pathogens test, Biofire FilmArray™ gastrointestinal panel and Luminex xTAG® gastrointestinal pathogen panel for detection of common enteric pathogens. Diagn. Microbiol. Infect. Dis. 86(4), 336–339. https://doi.org/10.1016/j.diagmicrobio.2016.09.013 (2016).
Tere-Peña, C. P., Calderon-Ozuna, M. N. & Leguizamón Guerrero, J. E. Digital PCR validation for characterization of quantitative reference material of Escherichia coli. O157:H7 genomic DNA. Methods Protoc. 7(6), 94. https://doi.org/10.3390/mps7060094 (2024).
Funding
Funder Chengdu Research Base of Giant Panda Breeding; Grant Number: 2024CPB-B11.
Author information
Authors and Affiliations
Contributions
BX conceived the experiments, completed the experiment and subsequently composed the initial version of the manuscript. HZ and LL supervised the work/project, provided guidance for the experiments and conducted revisions on the paper. XY and XS; project administration. YZ, JX, SL and YZ; data processing. All authors have read and agreed to the published version of the manuscript.All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xue, B., Zhang, H., Yan, X. et al. A TaqMan qPCR for precise detection and quantification of diarrheagenic Escherichia coli. Sci Rep 15, 16728 (2025). https://doi.org/10.1038/s41598-025-96960-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-96960-1