Abstract
This study systematically investigated the molecular mechanisms underlying tetrahydrocannabinol (THC)-induced hepatotoxicity in humans through an integrated approach combining network toxicology, molecular docking, and experimental validation. Our analysis identified 22 core targets associated with THC-mediated hepatotoxicity. Protein-protein interaction (PPI) network analysis revealed significant functional associations among these 22 potential target proteins. KEGG pathway and GO term analyses demonstrated that THC potentially exerts hepatotoxic effects through multiple biological processes, including endocrine resistance, bile secretion, negative regulation of apoptosis, and cellular oxidant detoxification. Disease enrichment analysis further identified several pathological conditions closely associated with THC-induced hepatic damage. Molecular docking simulations demonstrated strong binding affinities between THC and functional domains of 17 target proteins that participated in the aforementioned enriched pathways. An in vitro model of THC-induced hepatocyte injury was successfully established and subsequently validated through RT-qPCR experiment. THC exposure significantly altered the expression patterns of 10 critical target genes: ERBB2, GPX1, MAPK14, NR1H4, SOD1, CXCR2, PPARG, EGFR, TYMS and KDR. The hepatotoxic effects of THC appear to arise from the synergistic interplay of multiple pathways and the coordinated dysfunction of various gene products. These findings elucidate key molecular pathways and therapeutic targets associated with THC-induced hepatotoxicity, providing a theoretical foundation for developing clinical interventions and hepatoprotective strategies against cannabis-related liver damage.
Similar content being viewed by others
Introduction
Cannabis has gained global popularity across diverse populations for medicinal, cosmetic, and recreational applications. Therapeutically, it demonstrates efficacy in alleviating pain, reducing chemotherapy-induced nausea, and managing neurological conditions such as epilepsy and multiple sclerosis1. Emerging evidence reveals multi-system health risks associated with cannabis use, particularly impacting respiratory, cardiovascular, neuropsychiatric, and gastrointestinal functions. Chronic inhalation may induce bronchial inflammation and elevate chronic obstructive pulmonary disease (COPD) susceptibility, while cardiovascular manifestations range from tachycardia to myocardial infarction. Neuropsychiatric complications encompass anxiety disorders, depressive episodes, and cannabis-induced psychosis2,3,4. Notably, some users develop cannabis use disorder, with detrimental effects on occupational performance and social functioning5. While tetrahydrocannabinol (THC), the principal psychoactive constituent, has been extensively investigated for its central nervous system effects, its gastrointestinal impacts remain understudied despite increasing clinical observations. Cannabinoids exhibit bidirectional modulatory effects on gastrointestinal physiology, influencing appetite regulation, emesis control, and intestinal inflammatory responses through endocannabinoid system interactions6,7,8,9. THC’s hyperphagic effects may promote hedonic eating patterns, potentially contributing to metabolic syndrome and obesity-related hepatobiliary complications in chronic users10,11. Paradoxically, despite transient symptom relief, cannabis use may exacerbate disease progression in inflammatory bowel disorders through immune modulation alterations12,13. The anticholinergic effects of smoked cannabis induce xerostomia, potentially impairing salivary enzyme function critical for initial digestive processes8. Hepatic metabolic interactions between cannabinoids and xenobiotics may potentiate drug-induced liver injury, particularly in polydrug users with alcohol co-consumption14,15. Preclinical studies suggest THC modulates lipid homeostasis through mediating mechanisms of some function protein, potentially accelerating hepatic steatosis progression16. Synthetic cannabinoid receptor agonists (SCRAs), exhibiting several-fold greater CB1 receptor affinity than THC, demonstrate heightened hepatotoxic potential alongside severe cardiotoxic and proconvulsant effects in clinical case reports17,18.
Hepatotoxicity refers to the toxicological injury induced by xenobiotics on hepatic tissue, manifesting as structural damage or functional impairment of the liver. Although research on THC-induced hepatotoxicity remains insufficient, emerging evidence indicates its hepatotoxic potentiation effects. Preclinical studies demonstrate that THC co-administration with chemotherapeutic agents such as irinotecan significantly exacerbates hematological toxicity and induces histopathological alterations in rodent hepatic tissue19,20. Notably, a documented case of an adult male developing fulminant hepatic failure attributed to cannabis exposure, culminating in fatal outcomes, has been reported in clinical literature18. Despite established recognition of THC’s hepatotoxic potential, critical knowledge gaps persist regarding its precise molecular targets, mechanisms of action, and pathophysiological pathways in human hepatotoxicity.
Network toxicology represents an emerging interdisciplinary field that synergizes bioinformatics, multi-omics integration, computational modeling, and systems biology approaches21,22. It focuses on toxic pathways of compounds and molecular mechanisms of diseases, using a network-based approach to understand how molecular interactions in biological systems lead to toxicity. This method establishes relationships between chemicals, biological targets, and adverse effects, offering insights into complex systems biology. Unlike traditional toxicology’s “one target, one drug” approach, network toxicology follows a “multiple targets, one drug” model, revealing the intricate interactions among genes, proteins, and metabolites related to diseases and drugs23,24. As a computational chemistry technique, molecular docking enables precise simulation of ligand-receptor interactions through three-dimensional conformation matching, allowing predictive analysis of pharmacodynamic profiles and off-target liabilities25. As a core tool in drug discovery, it has transformed the study of interactions between small molecules and biological targets26. In toxicology, molecular docking can predict interactions between toxins and biomolecules, revealing toxic mechanisms. Recent studies have employed this methodology to elucidate the molecular interactions of the environmental toxin acetyl tributyl citrate as well as the hepatotoxic mechanisms mediated by the mycotoxin aflatoxin B121,27.
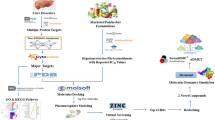
This study aims to elucidate the complex interactions and molecular mechanisms underlying THC-induced hepatotoxicity by combining the described methods with cellular functional experiments, thereby laying a scientific foundation for the development of relevant safety measures or safer alternatives. A key strength of this approach lies in its capacity to analyze extensive datasets and biological information to predict and identify potential toxic targets, thereby providing a comprehensive perspective on the risks associated with THC exposure. Moreover, understanding THC’s hepatotoxic potential not only aids in evaluating its safety for medical use but also provides a theoretical basis for the development of drugs targeting pathological changes. The study’s flowchart is shown in Fig. 1.
Flowchart of the network toxicology-based approach to investigate the toxic target of hepatotoxicity induced by THC.
v
Materials and methods
Toxicological analysis of THC
The toxicological assessment of THC was performed using two specialized databases: ADEMTlab2.0 (https://admetmesh.scbdd.com/service/evaluation/cal) and ADEMTsar (http://lmmd.ecust.edu.cn/admetsar2). Initially, the SMILES notation of THC was obtained from the comprehensive PubChem database (https://pubchem.ncbi.nlm.nih.gov/). Subsequently, the SMILES sequence was input into both aforementioned databases, resulting in the retrieval of predictive outcomes, which were then downloaded for this study.
Drug targets of THC
DrugBank is a comprehensive, freely accessible web-based bioinformatics and cheminformatics database that provides detailed information on drugs, including drug targets, drug actions, and drug interactions28. The primary advantage of DrugBank lies in its extensive and detailed data on drug targets, which are experimentally validated and regularly updated. Additionally, the SwissTargetPrediction database is used to predict the most likely protein targets of small molecules by analyzing similarities in both the two-dimensional and three-dimensional chemical structures of compounds29. The SwissTargetPrediction database provides the advantage of predicting small molecule drug targets based on structural similarity, thereby enhancing accuracy through prediction scoring. Drug targets for tetrahydrocannabinol (THC) were obtained from both DrugBank (accessed in September 2023) and the SwissTargetPrediction database (accessed in September 2023). To improve prediction accuracy, data from these sources were combined and duplicates were removed.
Therapeutic targets for hepatotoxicity diseases
DisGeNET provides one of the most comprehensive and publicly accessible catalogs of genes and genomic variants associated with diseases30. Its advantage lies in its ability to identify disease-related targets using gene-disease association (GDA) scores. GeneCards is a comprehensive and authoritative resource providing detailed annotations on genes31. The strength of the GeneCards database lies in its ability to identify gene-associated diseases by thoroughly calculating relevance scores between diseases and genes. The Therapeutic Target Database (TTD) is the world’s first online database providing free access to drug target information32. The therapeutic targets of agents in TTD are validated by clinical evidence, patents, or documented literature. The merged and de-duplicated targets from the DisGeNET (accessed in September 2023, GDA score > 0.06)30, GeneCards (accessed in September 2023, relevance score > 10)31, and TTD (accessed in September 2023) databases were selected as targets for hepatotoxic diseases.
The online tool Draw Venn Diagram (http://bioinformatics.psb.ugent.be/webtools/Venn/) was used to identify potential therapeutic targets of THC for hepatotoxic diseases. The functional classification of therapeutic targets was performed using the PANTHER database (accessed in September 2023)33.
Construction of the protein–protein interaction (PPI) network
Therapeutic targets of THC related to hepatotoxic diseases were uploaded to the STRING database online tool34 with a confidence score greater than 0.40. Subsequently, the PPI network was visualized using CytoHubba, a plugin of Cytoscape software (version 3.6.0), to calculate the degree of each protein node.
GO, KEGG and disease enrichment analysis
Functional enrichment analysis for Gene Ontology (GO), pathway analysis using the Kyoto Encyclopedia of Genes and Genomes (KEGG)35,36,37, and disease association analysis were performed using the DAVID database38. Therapeutic targets of THC in hepatotoxicity were uploaded to the DAVID database for analysis. Disease association analysis was performed using the DISGENET and GAD_DISEASE databases, focusing on diseases related to diabetes, kidney, and cardiovascular conditions, with a P-value less than 0.05 among the top 20 enrichment scores. Pathway enrichment analysis was performed using KEGG_PATHWAY, while GO functional enrichment analysis was conducted using GOTERM_BP_DIRECT, GOTERM_CC_DIRECT, and GOTERM_MF_DIRECT. For further analysis, the top 20 enriched KEGG pathways and GO terms with a P-value less than 0.05 were selected.
Molecular docking validation of the binding capacity between THC and targets
The 3-dimensional structure of THC was downloaded from PubChem database39 and transformed into Mol2 format using Open Babel software40. Hydrogen atoms were added using UCSF Chimera software41. The template structures of target proteins were obtained from the UniProt database42 and RCSB Protein Data Bank43. All the above databases were accessed in November 2023. Atom charges and hydrogen atoms were added to the protein, and solvents were removed using UCSF Chimera. Molecular docking of THC with target proteins was conducted using the SwissDock webserver (accessed December 2023)44 and visualized by UCSF Chimera.
Cell culture
The normal human liver cell lines (L- 02) were purchased from the Sunncell Biological Technology (Wuhan, China). L- 02 cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM, Cytiva, Marlborough, Massachusetts, United States) containing 10% fetal bovine serum (FBS, TRANS, China), 100 units/mL penicillin, and 100 µg/mL streptomycin at 37℃ in a 5% CO2 incubator (Panasonic MCO- 18 AC, Osaka, Japan). Cells were treated with Tetrahydrocannabinol (THC, 1 and 10 µM)45 dissolved in DMSO, and incubated for 48 h.
RNA isolation
After treatment, total RNA of L- 02 cells was extracted using TRIzol reagent according to the manufacturer’s instructions. The purity and concentration of the RNA were evaluated using Nanodrop micro-spectrophotometer (Thermo Fisher Scientific, USA) and agarose gel electrophoresis. RNA samples with a 260/280 and 260/230 absorbance ratio between 1.8 and 2.0, and undegraded electrophoretic band were considered to have acceptable quality. Subsequent analyses were performed.
Reverse transcription‑quantitative polymerase chain reaction (RT‑qPCR)
Reverse-transcription was performed using PrimeScript™ RT reagent Kit with gDNA Eraser (Perfect Real Time) (Takara, Dalian, China). In detail, the reverse transcription was performed at 37 ℃ for 15 min, 85 ℃ for 5 s, and 4 ℃ on an ABI VeritiPro Thermal Cycler (Thermo Fisher Scientific, USA), and a total of 1 µg RNA was taken to synthesize complementary DNA (cDNA). All primers for PCR amplification were designed using primer Express 3.0 software. The primer sequences are shown in Table 1. Then, qPCR was carried out in a QuantStudio® 5 Real-time PCR system (Thermofisher, USA) using TB Green® Advantage® qPCR Premix (Takara, Dalian, China). The cycling conditions were as follows: 95 °C for 30 s, followed by 40 cycles of 95 °C for 5 s and 60 °C for 31 s. Expression of ACTB gene was assayed simultaneously with samples as an internal control. Relative gene expression was determined by the 2−ΔΔCq method. All samples were performed three times in parallel. The specificity of each primer pair was verified both by in silico analysis (NCBI Primer-BLAST) and melting curve analysis after qPCR amplification.
Results
Toxicological analysis of THC
The co-toxicity profile of THC, as revealed by the ADMETlab2.0 and admetSAR databases (Table 2), prominently highlights its tendency toward human hepatotoxicity and liver injury. Additionally, these databases predict THC’s potential to cause acute oral and respiratory toxicity. Notably, there is an indication of THC’s potential carcinogenicity. Based on these database insights and supporting literature, the following section will explore the molecular mechanisms underlying THC-induced hepatotoxicity.
Identification of THC drug targets
There were 41 and 54 THC drug targets in Drugbank and SwissTargetPrediction databases respectively. We combined these drug targets and removed 4 duplicates. Finally, 91 drug targets were selected for further analysis.
Identification of disease targets
In the DisGeNET, Genecards, and TTD databases, 414, 356, and 6 targets were associated with hepatotoxicity, respectively. After removing redundant targets, 689 hepatotoxicity related targets were obtained.
Identification of therapeutic targets for THC caused hepatotoxicity
The above THC drug targets were intersected with hepatotoxicity targets to obtain potential therapeutic targets for THC caused hepatotoxicity. Finally, a total of 22 targets were identified (Fig. 2A). And these targets can be classified into 7 functional categories including metabolite interconversion enzyme (PTGS2, SMPD1, SOD1, TYMS, NOS2, GPX1, GSR, HMGCR, ACHE, CAT), gene-specific transcriptional regulator (ESR1, NR1H4, NR3C1, PPARG), transporter (CFTR), protein modifying enzyme (MAPK14), transmembrane signal receptor (EGFR, ERBB2, KDR, OPRM1), cell adhesion molecule (CXCR2), DNA metabolism protein (PARP1) (Fig. 2B).
Identification of toxic targets for THC induced hepatotoxicity. A Venn diagram of potential toxic targets. B Classification of the identified targets. 22 therapeutic targets for toxic can be obtained and classified into 7 functional categories.
PPI network construction
The identified 22 therapeutic targets for THC caused hepatotoxicity were used to construct the PPI network. A total of 20 nodes and 73 edges were included in the PPI network, with an average node degree of 6.95 (Fig. 3). The ranking of the therapeutic targets based on the weighted degree was MAPK14, PPARG, PTGS2, EGFR, ESR1, PARP1, KDR, NOS2, ERBB2, SOD1, GSR, NR3C1, TYMS, ACHE, HMGCR, NR1H4, OPRM1, CXCR2, CAT and CFTR.
Protein-protein interaction network of targets of THC induced hepatotoxicity.
GO, KEGG and disease enrichment analysis
GO, KEGG, and disease enrichment analyses of THC targets associated with hepatotoxicity were performed using the DAVID database. The top 20 enriched biological processes, cellular components, and molecular functions from the GO analysis are shown in Fig. 4A. Among the enriched biological process terms, response to xenobiotic stimulus (GO:0009410), negative regulation of apoptotic process (GO:0043066), and cellular oxidant detoxification (GO:0098869) may play important roles in hepatotoxicity. The cellular components are primarily associated with the mitochondrial matrix, nucleus, cytosol, and plasma membrane (Fig. 4A). The molecular functions are primarily associated with enzyme binding, ion binding, and protein binding (Fig. 4A). A total of 20 KEGG pathways were enriched. As shown in Fig. 4B, eight KEGG pathways are potentially involved in THC-induced hepatotoxicity, including the VEGF signaling pathway (hsa04370), bile secretion (hsa04976), AMPK signaling pathway (hsa04152), and FoxO signaling pathway (hsa04068). After removing duplicates, the top 20 enriched diseases from the DISGENET and GAD_DISEASE databases are shown in Table 3; Fig. 4C. Many genes are enriched in liver- and drug toxicity-related diseases, including intrahepatic cholangiocarcinoma (C0345905), extrahepatic cholangiocarcinoma (C3805278), drug toxicity (C0013221), and chronic persistent hepatitis (C0520463).
GO, KEGG and disease enrichment analysis for hepatotoxicity related genes associated with THC. A: The top 20 terms of GO enrichment (green: biological process; orange: cellular component; blue: molecular function); B: The top 20 pathways of KEGG enrichment; C: Chord diagram of top 20 results for disease enrichment.
Validation of binding capacity between THC and therapeutic targets by molecular docking
The binding capacities of THC and the target proteins were predicted using the SwissDock web server. The docking models with the lowest binding energies (Delta G) were recorded. Docking sites of THC with ESR1, HMGCR, NOS2, NR3C1, and PARP1 were not obtained from the SwissDock database. The remaining molecular docking results of THC with the target proteins are shown in Table 4. The results showed that THC exhibited the strongest and most stable binding affinity toward CFTR, followed by CXCR2, TYMS, SMPD1, PPARG, NR1H4, ACHE, EGFR, GSR, CAT, PTGS2, MAPK14, KDR, OPRM1, SOD1, ERBB2, and GPX1. Among these, hydrogen bonds were observed between THC and the five target proteins (ERBB2, GPX1, MAPK14, NR1H4, and SOD1). The results indicate that THC does not bind directly to the other target proteins but interacts through a pocket located between the proteins. The specific docking sites of THC with the target proteins are shown in Fig. 5.
The 3-dimensional map of the binding sites between THC and target proteins. A ERBB2, B GPX1, C MAPK14, D NR1H4, E SOD1, F ACHE, G CAT, H CFTR, I CXCR2, J EGFR, K GSR, L KDR, M OPRM1, N PPARG, O PTGS2, P SMPD1, Q TYMS. THC is shown in baby blue. Target proteins are displayed as brown. The places where THC and the target proteins are connected represent specific docking sites between THC and target proteins.
Efect of THC on hepatotoxicity model
Normal human liver cell (L- 02) lines were treated with NC, THC(1µM, 10µM) for 48 h. mRNA expression levels of the candidate target genes were analyzed using RT-qPCR. The relative mRNA levels of nine genes (ERBB2, GPX1, MAPK14, NR1H4, SOD1, CXCR2, PPARG, EGFR, and TYMS) were significantly downregulated, while the mRNA level of one gene (KDR) was significantly upregulated in the THC group compared with the NC group (Fig. 6). No statistically significant differences were observed in the expression of the other genes identified in the molecular docking results between the THC and NC groups.
Efect of THC on target proteins. The mRNA expression levels of different genes. 0µM represented the control group without THC treatment, and 1µM and 10µM represented the THC treatment group with different concentrations, respectively. *P < 0.05, **P < 0.01, ***P < 0.001.
Discussion
Marijuana legalization has gained public acceptance over the past few years; however, research on the potential negative health effects of dabbing is still ongoing46. As the predominant constituent of cannabis, THC is primarily responsible for its classification as a narcotic. Prolonged use can lead to dependence and cause substantial harm to the human nervous system. Research has also revealed that THC poses significant health risks to the liver, potentially inducing hepatotoxicity. However, the molecular mechanisms underlying THC-induced hepatotoxicity in humans remain unclear. This study aims to elucidate the potential targets involved in THC-induced hepatotoxicity through network toxicology, molecular docking, and functional assays. Our findings suggest that these identified targets may contribute to liver toxicity through coordinated action across multiple pathways.
Using Venn diagrams, a total of 22 potential targets associated with THC-induced hepatotoxicity were identified from various databases. Subsequently, the general interactions among the 22 targets were identified using the PPI network. Gene Ontology (GO), KEGG, and disease enrichment analyses of these potential targets revealed associations between liver disease and drug toxicity. In addition, molecular docking of these core targets demonstrated that THC’s binding affinity to 17 targets may result in adverse effects. Finally, RT-qPCR experiments showed that ten targets were significantly differentially expressed.
In the enrichment analysis of candidate targets, we found that certain KEGG pathways may play an important role in THC-induced liver toxicity. For example, the endocrine resistance pathway has been associated with liver cirrhosis and is involved in insulin secretion47,48. This pathway contains candidate target genes including ERBB2, MAPK14, ESR1, and EGFR, among which ERBB2, MAPK14, and EGFR showed decreased expression at various THC concentrations according to RT-qPCR results at the cellular level (Fig. 6). The downregulated expression of ERBB2, MAPK14 and EGFR within the endocrine resistance pathway may indirectly contribute to the pathogenesis of liver cirrhosis through mechanisms involving impaired hepatic repair, exacerbated fibrotic progression, and dysregulation of metabolic homeostasis. Long-term use of hepatotoxic drugs can cause liver damage, leading to liver fibrosis that affects cell proliferation and the cell cycle. Pathway enrichment related to liver fibrosis includes the VEGF signaling pathway, MAPK signaling pathway, relaxin signaling pathway, and FoxO signaling pathway49,50,51,52,53,54. These pathways include ERBB2, KDR, MAPK14, EGFR, NOS2, and CAT. Among these genes, ERBB2, MAPK14, and EGFR showed significantly decreased expression, whereas KDR exhibited a significant increase (Fig. 6). The differential gene expression variations within signaling pathways can exert substantial functional impacts on the corresponding pathways. For instance, MAPK14 in the MAPK pathway, which is primarily involved in inflammatory responses and stress signaling, may demonstrate compromised regulation of cellular apoptosis or fibrogenesis upon its downregulation. Manifestations of liver toxicity are accompanied by abnormal bile secretion, and KEGG enrichment results show involvement of the bile secretion pathway, which includes the target gene NR1H4 that is significantly downregulated (Fig. 6). Liver toxicity can also damage liver cells, impairing their normal function and leading to fat accumulation, which results in fatty liver formation. Notably, the KEGG enrichment results include the HIF- 1 signaling pathway and the AMPK signaling pathway, both of which have been significantly associated with fatty liver in previous studies55,56,57,58. Both pathways involve ERBB2, EGFR and PPARG, all of which exhibit significant differences in expression levels (Fig. 6). In the biological process category of the GO enrichment results, several terms potentially important in liver toxicity—such as response to xenobiotic stimulus (GO:0009410), negative regulation of apoptotic process (GO:0043066), and cellular oxidant detoxification (GO:0098869)—are associated with 12 genes (GPX1, SMPD1, CAT, PPARG, TYMS, PTGS2, SOD1, ERBB2, NR1H4, KDR, EGFR, and GSR). Among these genes, GPX1, PPARG, TYMS, SOD1, ERBB2, NR1H4, KDR and EGFR show significant differences in expression levels (Fig. 6). Disease enrichment analysis revealed a significant number of conditions related to THC-induced liver toxicity, including intrahepatic cholangiocarcinoma, extrahepatic cholangiocarcinoma, inflammation, cholangiocarcinoma, marijuana abuse, drug toxicity, adverse drug reactions, cholestasis, chronic hepatitis, chronic persistent hepatitis, and cryptogenic chronic hepatitis. Among the aforementioned disease enrichment results, the expression levels of eight potential target genes (EGFR, ERBB2, GPX1, MAPK14, NR1H4, PPARG, SOD1 and TYMS) exhibited significant differences. These genes are also represented in the KEGG and GO enrichment results above, suggesting that THC-induced hepatotoxicity may be mediated through multiple pathways.
Of the 17 candidate target genes that performed well in molecular docking, 10 exhibited significant differential expression in the RT-qPCR results. Nine of these genes were significantly enriched in pathways, terms, or disease types associated with THC induced hepatotoxicity. The previous study revealed that ERBB2 promotes ACC1 activity through TAp63 upregulation, thereby activating hepatic stellate cells and driving hepatic fibrosis59. THC may attenuate liver fibrosis by indirectly suppressing HSC activation via modulation of TAp63 expression or interference with the ERBB2 signaling pathway. In parallel, SOD1 exerts hepatoprotective effects by scavenging superoxide radicals60. However, THC-induced oxidative stress may inhibit SOD1 activity, leading to lipid peroxidation and mitochondrial dysfunction, thereby exacerbating the progression of non-alcoholic fatty liver disease. GPX1 is a critical antioxidant gene, and its downregulation exacerbates oxidative stress61. THC may suppress GPX1 expression, leading to the accumulation of reactive oxygen species (ROS) in hepatocytes, thereby triggering mitochondrial dysfunction and apoptosis. This mechanism could represent a key contributor to THC-induced hepatotoxicity. MAPK14, a downstream effector of the EGF signaling pathway, is highly expressed in hepatocellular carcinoma (HCC) and correlates with poor prognosis62. THC may reduce MAPK14 activity by inhibiting EGF signaling or disrupting the TPRG1-AS1/CLTC complex formation, consequently suppressing HCC cell proliferation and invasion. The FXR protein encoded by NR1H4 regulates bile acid metabolism and anti-inflammatory responses63. THC might activate FXR to enhance bile acid efflux and inhibit pro-inflammatory cytokine release, partially alleviating liver injury; however, chronic overactivation could disrupt bile acid homeostasis, resulting in hepatocyte damage. KDR encodes VEGFR2, which promotes angiogenesis in HCC64. THC may upregulate KDR expression, activating the VEGF signaling pathway to facilitate tumor neovascularization and metastasis, a potential mechanism underlying THC-driven HCC progression. TYMS, a key enzyme in one-carbon metabolism, is regulated by MTHFR to maintain DNA stability65. THC may indirectly impair TYMS function by interfering with folate metabolism, leading to nucleotide synthesis defects and hepatocyte DNA damage, ultimately promoting carcinogenesis. CXCR2 aggravates tacrolimus-induced hepatotoxicity via activation of the JAK3/STAT3 pathway66. THC might upregulate CXCR2 expression, enhancing STAT3 phosphorylation to amplify inflammatory responses and hepatocyte apoptosis, thereby exacerbating liver injury. PPARG is upregulated in liver fibrosis and modulates inflammation and apoptosis67. THC could activate PPARG to promote lipid accumulation and fibrogenic gene expression, accelerating hepatic fibrogenesis. EGFR is upregulated in liver fibrosis and drives hepatic stellate cell (HSC) activation68. THC may enhance EGFR expression to stimulate HSC proliferation and collagen deposition, while methyl donor therapy reverses this effect, suggesting THC might interfere with DNA methylation-dependent regulation of EGFR. Notably, under THC exposure, the downregulation of CXCR2, PPARG, and EGFR in hepatocytes may initially reflect compensatory protective responses (e.g., suppressing excessive inflammation). However, prolonged THC exposure likely overwhelms these adaptive mechanisms, ultimately accelerating hepatic injury progression. This finding further suggests that THC-induced hepatotoxicity is not mediated by a single pathway but rather constitutes a multidimensional injury network through disrupting redox balance (GPX1/SOD1), metabolic homeostasis (NR1H4/TYMS), fibrogenic pathways (ERBB2/PPARG/EGFR), and inflammation-to-cancer transition (MAPK14/CXCR2/KDR). Notably, THC exhibits bidirectional or damage severity-dependent regulatory effects on specific genes (e.g., EGFR and PPARG).
The strengths of this study lie in its comprehensive identification of therapeutic targets for THC-induced hepatotoxicity in humans, accomplished through an innovative network toxicology analysis integrating multiple databases. The identified targets were further validated using molecular docking simulations and in vitro experiments. To the best of our knowledge, no previous studies have performed such extensive screening and validation of therapeutic targets for THC-induced hepatotoxicity in humans. The results of this study reveal that the molecular mechanism of THC-induced hepatotoxicity may arise from the synergistic actions of multiple pathways and gene functions. This investigation highlights the intricate biochemical mechanisms underlying THC-induced hepatotoxicity, providing potential targets for drug development and offering insights into therapeutic strategies to mitigate the adverse effects associated with this cannabinoid.
The primary objective of this study was to preliminarily identify potential targets of THC-induced hepatotoxicity through network toxicology and molecular docking, and to validate their expression alterations at the mRNA level via RT-qPCR. This integrated strategy has been extensively employed in the initial phase of mechanistic exploration27,69. However, this study has several limitations. For example, the identified targets were validated only through computer simulations and in vitro experiments for mRNA, without protein experiment and any in vivo validation. Additionally, this study did not investigate the variations in the degree of hepatotoxicity caused by different doses of THC. Therefore, further western blot for protein experiment and in vivo animal model experiments are needed in subsequent stages to examine the effects of varying doses on the liver and provide a theoretical basis for clinical medication.
Conclusions
Beyond the aforementioned limitations, we conclude that therapeutic targets for THC-induced hepatotoxicity were identified through network toxicology, bioinformatics analysis, molecular docking simulations, and in vitro cellular evaluations. THC treatment resulted in abnormal expression of 10 target genes (ERBB2, GPX1, MAPK14, NR1H4, SOD1, CXCR2, PPARG, EGFR, TYMS, and KDR), involving various pathways and biological processes. These findings highlight potential pathways and targets affected by THC-induced hepatotoxicity in humans, providing a theoretical foundation for clinical treatment strategies and drug development aimed at addressing this toxicity.
Data availability
All data generated or analysed during this study are included in this published article.
References
Grotenhermen, F. & Muller-Vahl, K. The therapeutic potential of cannabis and cannabinoids. Dtsch. Arztebl Int. 109, 495–501 (2012).
Tashkin, D. P. Effects of marijuana smoking on the lung. Ann. Am. Thorac. Soc. 10, 239–247 (2013).
Ravi, D., Ghasemiesfe, M., Korenstein, D., Cascino, T. & Keyhani, S. Associations between marijuana use and cardiovascular risk factors and outcomes: A systematic review. Ann. Intern. Med. 168, 187–194 (2018).
Volkow, N. D. et al. Effects of cannabis use on human behavior, including cognition, motivation, and psychosis: A review. JAMA Psychiatry. 73, 292–297 (2016).
Lopez-Quintero, C. et al. Probability and predictors of transition from first use to dependence on nicotine, alcohol, cannabis, and cocaine: results of the National epidemiologic survey on alcohol and related conditions (NESARC). Drug Alcohol Depend. 115, 120–130 (2011).
Parker, L. A., Rock, E. M. & Limebeer, C. L. Regulation of nausea and vomiting by cannabinoids. Br. J. Pharmacol. 163, 1411–1422 (2011).
Hasenoehrl, C., Taschler, U., Storr, M. & Schicho, R. The Gastrointestinal tract - a central organ of cannabinoid signaling in health and disease. Neurogastroenterol Motil. 28, 1765–1780 (2016).
Huestis, M. A. Human cannabinoid pharmacokinetics. Chem. Biodivers. 4, 1770–1804 (2007).
Goyal, H., Singla, U., Gupta, U. & May, E. Role of cannabis in digestive disorders. Eur. J. Gastroenterol. Hepatol. 29, 135–143 (2017).
Fearby, N., Penman, S. & Thanos, P. Effects of Delta9-Tetrahydrocannibinol (THC) on obesity at different stages of life: A literature review. Int. J. Environ. Res. Public. Health 19, 3174 (2022).
Eitan, A., Gover, O., Sulimani, L., Meiri, D. & Schwartz, B. The effect of orally administered Delta9-Tetrahydrocannabinol (THC) and Cannabidiol (CBD) on obesity parameters in mice. Int. J. Mol. Sci. 24,13797 (2023).
Perisetti, A., Rimu, A. H., Khan, S. A., Bansal, P. & Goyal, H. Role of cannabis in inflammatory bowel diseases. Ann. Gastroenterol. 33, 134–144 (2020).
Carvalho, A. C. A. et al. Cannabis and canabidinoids on the inflammatory bowel diseases: going beyond misuse. Int. J. Mol. Sci. 21, 2940 (2020).
Cohen, L. & Neuman, M. G. Cannabis and the Gastrointestinal tract. J. Pharm. Pharm. Sci. 23, 301–313 (2020).
Gotfried, J., Naftali, T. & Schey, R. Role of cannabis and its derivatives in Gastrointestinal and hepatic disease. Gastroenterology 159, 62–80 (2020).
Mallat, A., Teixeira-Clerc, F. & Lotersztajn, S. Cannabinoid signaling and liver therapeutics. J. Hepatol. 59, 891–896 (2013).
McCain, K. R. et al. Impaired driving associated with the synthetic cannabinoid 5f-Adb. J. Forensic Sci. Criminol. 6,1-4 (2018).
Solimini, R. et al. Hepatotoxicity associated to synthetic cannabinoids use. Eur. Rev. Med. Pharmacol. Sci. 21, 1–6 (2017).
Prester, L. et al. Effects of Delta(9)-tetrahydrocannabinol on irinotecan-induced clinical effects in rats. Chem. Biol. Interact. 294, 128–134 (2018).
Lucic Vrdoljak, A. et al., Irinotecan and Delta(8)-Tetrahydrocannabinol Interactions in Rat Liver: A Preliminary Evaluation Using Biochemical and Genotoxicity Markers. Molecules 23, (2018).
Huang, S. Efficient analysis of toxicity and mechanisms of environmental pollutants with network toxicology and molecular Docking strategy: acetyl tributyl citrate as an example. Sci. Total Environ. 905, 167904 (2023).
Tao, W. et al. Network pharmacology-based prediction of the active ingredients and potential targets of Chinese herbal radix curcumae formula for application to cardiovascular disease. J. Ethnopharmacol. 145, 1–10 (2013).
Zhang, Y. et al. A network pharmacology-based strategy Deciphers the underlying molecular mechanisms of Qixuehe capsule in the treatment of menstrual disorders. Chin. Med. 12, 23 (2017).
Zhang, S. et al. Systematic analysis of the multiple bioactivities of green tea through a network Pharmacology approach. Evid. Based Complement. Alternat Med. 2014, 512081 (2014).
Stanzione, F., Giangreco, I. & Cole, J. C. Use of molecular Docking computational tools in drug discovery. Prog Med. Chem. 60, 273–343 (2021).
Pinzi, L. & Rastelli, G. Molecular docking: shifting paradigms in drug discovery. Int. J. Mol. Sci. 20, 4331 (2019).
Wishart, D. S. et al. DrugBank 5.0: a major update to the drugbank database for 2018. Nucleic Acids Res. 46, D1074–D1082 (2018).
Daina, A., Michielin, O. & Zoete, V. SwissTargetPrediction: updated data and new features for efficient prediction of protein targets of small molecules. Nucleic Acids Res. 47, W357–W364 (2019).
Pinero, J. et al. The disgenet knowledge platform for disease genomics: 2019 update. Nucleic Acids Res. 48, D845–D855 (2020).
Safran, M. et al. GeneCards Version 3: the human gene integrator. Database (Oxford) baq020 (2010). (2010).
Chen, X., Ji, Z. L. & Chen, Y. Z. TTD: therapeutic target database. Nucleic Acids Res. 30, 412–415 (2002).
Mi, H. et al. PANTHER version 16: a revised family classification, tree-based classification tool, enhancer regions and extensive API. Nucleic Acids Res. 49, D394–D403 (2021).
Szklarczyk, D. et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 47, D607–D613 (2019).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Kanehisa, M. Toward Understanding the origin and evolution of cellular organisms. Protein Sci. 28, 1947–1951 (2019).
Kanehisa, M., Furumichi, M., Sato, Y., Kawashima, M. & Ishiguro-Watanabe, M. KEGG for taxonomy-based analysis of pathways and genomes. Nucleic Acids Res. 51, D587–D592 (2023).
Dennis, G. Jr Database for annotation, visualization, and integrated discovery. Genome Biol. 4, P3 (2003).
Kim, S. et al. PubChem substance and compound databases. Nucleic Acids Res. 44, D1202–1213 (2016).
O’Boyle, N. M. et al. Open babel: an open chemical toolbox. J. Cheminform. 3, 33 (2011).
Huang, C. C., Meng, E. C., Morris, J. H., Pettersen, E. F. & Ferrin, T. E. Enhancing UCSF chimera through web services. Nucleic Acids Res. 42, W478–484 (2014).
UniProt C. UniProt: the universal protein knowledgebase in 2023. Nucleic Acids Res. 51, D523–D531 (2023).
Berman, H. M. et al. The protein data bank. Nucleic Acids Res. 28, 235–242 (2000).
Grosdidier, A., Zoete, V. & Michielin, O. SwissDock, a protein-small molecule Docking web service based on EADock DSS. Nucleic Acids Res. 39, W270–277 (2011).
Nasrin, S., Watson, C. J. W., Perez-Paramo, Y. X. & Lazarus, P. Cannabinoid metabolites as inhibitors of major hepatic CYP450 enzymes, with implications for Cannabis-Drug interactions. Drug Metab. Dispos. 49, 1070–1080 (2021).
Zhang, Z., Zheng, X., Zeng, D. D. & Leischow, S. J. Tracking dabbing using search query surveillance: A case study in the united States. J. Med. Internet Res. 18, e252 (2016).
Bai, X. P., Fan, Y. M., Zhang, L., Yang, G. H. & Li, X. Influence of liver cirrhosis on blood glucose, insulin sensitivity and islet function in mice. Am. J. Med. Sci. 362, 403–417 (2021).
Zhang, W. et al. Di-(2-ethylhexyl) phthalate could disrupt the insulin signaling pathway in liver of SD rats and L02 cells via PPARgamma. Toxicol. Appl. Pharmacol. 316, 17–26 (2017).
Jiang, H. et al. Hepatoprotective Effect of Medicine Food Homology Flower Saffron against CCl(4)-Induced Liver Fibrosis in Mice via the Akt/HIF-1alpha/VEGF Signaling Pathway. Molecules 28, (2023).
Chen, F. et al. VEGF-FGF signaling activates quiescent CD63(+) liver stem cells to proliferate and differentiate. Adv. Sci. (Weinh). 11, e2308711 (2024).
Lan, T. et al. Tianhuang formula ameliorates liver fibrosis by inhibiting CCL2-CCR2 axis and MAPK/NF-kappaB signaling pathway. J. Ethnopharmacol. 321, 117516 (2024).
Cai, S. et al. Carvacrol alleviates liver fibrosis by inhibiting TRPM7 and modulating the MAPK signaling pathway. Eur. J. Pharmacol. 898, 173982 (2021).
Ezhilarasan, D. Relaxin in hepatic fibrosis: What is known and where to head? Biochimie 187, 144–151 (2021).
Feng, X. et al. Yinchen Gongying Decoction mitigates CCl(4)-induced chronic liver injury and fibrosis in mice implicated in Inhibition of the FoxO1/TGF-beta1/ Smad2/3 and YAP signaling pathways. J. Ethnopharmacol. 327, 117975 (2024).
He, Y. et al. Silencing HIF-1alpha aggravates non-alcoholic fatty liver disease in vitro through inhibiting PPAR-alpha/ANGPTL4 singling pathway. Gastroenterol. Hepatol. 44, 355–365 (2021).
Yuan, S. et al. Sorafenib attenuates liver fibrosis by triggering hepatic stellate cell ferroptosis via HIF-1alpha/SLC7A11 pathway. Cell. Prolif. 55, e13158 (2022).
Marcondes-de-Castro, I. A., Reis-Barbosa, P. H., Marinho, T. S., Aguila, M. B. & Mandarim-de-Lacerda, C. A. AMPK/mTOR pathway significance in healthy liver and non-alcoholic fatty liver disease and its progression. J. Gastroenterol. Hepatol. 38, 1868–1876 (2023).
Guo, T. et al. Liraglutide attenuates type 2 diabetes mellitus-associated non-alcoholic fatty liver disease by activating AMPK/ACC signaling and inhibiting ferroptosis. Mol. Med. 29, 132 (2023).
Fondevila, M. F. et al. p63 controls metabolic activation of hepatic stellate cells and fibrosis via an HER2-ACC1 pathway. Cell. Rep. Med. 5, 101401 (2024).
Yang, C. et al. Effects of dietary amylose/amylopectin ratio on antioxidant ability and amino metabolism in the liver of weaned piglets undergoing feed transition and challenged with lipopolysaccharide. Front. Nutr. 11, 1435051 (2024).
Huang, T. Y. et al. Furanocoumarin Notopterol: Inhibition of Hepatocellular Carcinogenesis through Suppression of Cancer Stemness Signaling and Induction of Oxidative Stress-Associated Cell Death. Nutrients 15, (2023).
Moon, S. U., Shah, M., Thao, T. T. & Woo, H. G. Long Non-coding RNA TPRG1-AS1 interacts with CLTC in liver cancer cells. Anticancer Res. 44, 4813–4824 (2024).
Mohammed, T. A. & Zalzala, M. H. Hepatoprotective effects of cilnidipine in cholestatic liver disease: role of FXR and NRF2 signalling. J. Exp. Pharmacol. 17, 93–105 (2025).
Abusaif, M. S. et al. Exploring a novel thiazole derivatives hybrid with fluorinated-indenoquinoxaline as dual inhibitors targeting VEGFR2/AKT and apoptosis inducers against hepatocellular carcinoma with Docking simulation. Bioorg. Chem. 154, 108023 (2025).
Wu, M. T. et al. MTHFR knockdown assists cell defense against folate depletion induced chromosome segregation and uracil misincorporation in DNA. Int. J. Mol. Sci. 22, 9392 (2021).
Chen, X., Hu, K., Zhang, Y., He, S. M. & Wang, D. D. CXCR2 activated JAK3/STAT3 signaling pathway exacerbating hepatotoxicity associated with tacrolimus. Drug Des. Devel Ther. 18, 6331–6344 (2024).
Yu, L., Yu, P., Cao, Y. & Cao, W. Mechanism of radix bupleuri and hedysarum multijugum Maxim drug pairs on liver fibrosis based on network pharmacology, bioinformatics and molecular dynamics simulation. PLoS One. 20, e0318336 (2025).
Bishnolia, M. et al. Methyl donor ameliorates CCl(4)-induced liver fibrosis by inhibiting inflammation, and fibrosis through the downregulation of EGFR and DNMT-1 expression. Food Chem. Toxicol. 196, 115230 (2025).
Ge, B. et al. Integrated network toxicology, molecular docking, and in vivo experiments to elucidate molecular mechanism of aflatoxin B1 hepatotoxicity. Ecotoxicol. Environ. Saf. 275, 116278 (2024).
Chen, Q. et al. Study on the mechanism of Mesaconitine-Induced hepatotoxicity in rats based on metabonomics and toxicology network. Toxins (Basel) 14, 486 (2022).
Acknowledgements
The authors would like to thank the researchers and staff of the above software and databases.
Funding
This work was supported by the NHC Key Lab of Drug Addiction Medicine (Kunming Medical University) Open Projects Fund (KN202411 and KN202420), and Kunming Medical University Talent Introduction Research Project (K132310511 and K132310529).
Author information
Authors and Affiliations
Contributions
S.J.H., L.L.L. and T.T.Z. designed the research; T.T.Z. and M.Q.W. analyzed data; L.P.H., G.M.Y., X.Y.C. and T.T.C. carried out cell experiments. T.T.Z. and L.L.L. wrote the paper; S.J.H, S.J.N. and R.L.Z. revised the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zou, T., Zhang, R., Hu, L. et al. Uncovering mechanism of hepatotoxicity diseases caused by tetrahydrocannabinol based on novel network toxicology and experimental verification. Sci Rep 15, 13712 (2025). https://doi.org/10.1038/s41598-025-97523-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-97523-0