Abstract
Leaf rust (LR) is one the most widely distributed and serious pathogen hampering the wheat production globally. To combat the continuous evolution of pathogens and breakdown of resistance, matching resistance genes must be discovered at the same pace from the diverse sources. In this study, we identified the US genome species Aegilops kotschyi acc pau 396, which is resistant to the Indian Puccinia triticina pathotypes 109R31-1 (77 − 5), 21R55 (104-2), and 121R60-1 (77 − 9). An LR resistance introgression line, ILkots was developed from this accession of Aegilops kotschyi in the background of Triticum aestivum cultivar PBW343. To examine the genetics of transferred resistance, a mapping population of 222 F3:4 plants was generated by crossing ILkots with the LR-susceptible cultivar WL711NN. Field experiments were conducted using a randomized complete block design (RCBD). Genetic analysis revealed that resistance was conferred by a single, dominant gene temporarily designated as Lrkots. Bulked segregant analysis (BSA) in combination with AFFYMETRIX 35 K Wheat Breeders’ AXIOM array mapped Lrkots on chromosome 3DL. A partial genetic linkage map of the genomic region carrying Lrkots included five Kompetitive allele-specific PCR (KASP) markers and two simple sequence repeat (SSR) markers spanning 8.2 cM. Lrkots was flanked by the KASP marker AX-94,443,154 (0.32 cM) on the proximal end and the SSR marker Xbarc71 (7.9 cM) on the distal end, defining a 3 Mb region in the RefSeq v1.0 genome assembly. This interval contains eight candidate genes encoding protein domains involved in disease resistance responses.
Similar content being viewed by others
Introduction
Cultivated wheat (Triticum aestivum L.) is predisposed to leaf rust (LR) (Puccinia triticina Eriks), a major fungal pathogen that can reduce grain yield by up to 50% under favourable conditions1. Given the projected 56% increase in global food consumption by 20502, there is a critical need to address key challenges associated with wheat production. India relies significantly on wheat production in its northern states, notably the North-West Plain Zone, which covers around 12.33 million hectares for wheat cultivation and accounts for nearly half of the country’s wheat production3. A decline in yield due to leaf rust could significantly impact national food security and agricultural economy.
Deploying rust resistance genes against leaf rust is an effective and sustainable strategy to mitigate yield losses4. Numerous efforts have been carried out to identify and screen germplasm for rust resistance5,6,7. However, artificial selection and domestication have led to genetic uniformity, reducing resistance diversity. Indian wheat cultivars carry several Lr genes (e.g., Lr1, Lr3, Lr9, Lr10, Lr13, Lr14a, Lr17, Lr19, Lr22, Lr23, Lr24, Lr26, Lr34), many of which have been rendered ineffective due to emerging virulent pathotypes6. For instance, the widespread use of Lr3 resulted in the emergence of pathotype 77, while the Lr23 + Lr26 combination was overcome by evolving pathotypes such as 12 − 1 and 77 − 18. The rapid evolution of race group 77 has led to the emergence of 13 distinct pathotypes in India9.
Rust resistance is categorized into seedling stage (all-stage resistance, ASR) and adult plant resistance (APR). ASR usually provides robust immune response but is race specific and susceptible to breakdown with the evolution of pathotypes. In contrast, APR provides broader but partial resistance against multiple strains and is characterized by more durability, thereby offering protection over an extended period10. To counter evolving rust pathotypes, a continuous introduction of both resistance types is necessary. In breeding efforts, wild sources of resistance have been rigorously exploited to address this problem. Around 50% of the rust resistance identified belongs to wild sources, with the genus Aegilops accounting for about 20% of the total11. A wealth of LR resistance genes has been incorporated from Aegilops, including Lr21/Lr40, Lr22a, Lr32, Lr39/Lr41, Lr42 from Aegilops tauschii; Lr28, Lr35, Lr36, Lr37, Lr47, Lr51, Lr66 from Aegilops speltoides, Lr9, Lr76 from Aegilops umbellulata, Lr56 from Aegilops sharonensis; Lr57 from Aegilops geniculata; Lr58, Lrtri from Aegilops triuncialis; Lr59, LrP, LrAp from Aegilops peregrina and Lr62 from Aegilops neglecta12,13,14,15.
Aegilops kotschyi (UUSS), a tetraploid wild relative of wheat, holds promise in enhancing wheat’s disease resistance and nutritional value. It has been identified as a source of LR and yellow rust (YR) resistance while also offering high levels of grain mineral content16,17,18,19. Till date, only a single Lr gene, Lr54 linked to Yr37 has been reported from Ae. kotschyi and transferred to hexaploid background by16 on long arm of chromosome 2D.
Understanding the genetic potential of Ae. kotschyi requires leveraging both traditional and modern gene identification techniques. Classical genetics, cytology, and morphological markers have historically played key roles in gene discovery. Controlled crosses and cytogenetic analyses have provided insights into trait inheritance and chromosome structure20. For instance, classical genetic studies have clarified the inheritance of disease resistance genes such as Lr3421, while cytological techniques, including chromosome staining and microscopy, have helped to identify structural variations such as deletions, translocations, and inversions22.
Advancements in molecular biology have revolutionized gene identification in wheat, enabling precise mapping through DNA markers. DNA markers offer advantages such as high abundance, reproducibility, and the ability to detect genetic variations at the molecular level. Techniques like Restriction Fragment Length Polymorphism (RFLP), Sequence Tagged Site (STS) and Simple Sequence Repeats (SSRs) have been extensively used for genetic mapping in wheat12,17. Moreover, Single Nucleotide Polymorphisms (SNPs) have emerged as valuable markers for wheat genomics due to their abundance and genome-wide distribution. These markers enable precise gene mapping and marker-assisted selection for breeding purposes21.
The advancement and cost-effectiveness of high-throughput sequencing and genotyping technologies have revolutionized the process of candidate region identification, marker development, and the transfer of target traits into susceptible cultivars. The emergence of sequencing technologies has resulted in the development of various SNP array platforms for scanning genetic diversity in target germplasms. 35K SNP array23 is one of the important genotyping platforms known for its even genomic distribution of SNPs and suitability for genetic mapping in hexaploid germplasms5,24. Bulk Segregant Analysis (BSA) using 35K SNP chips offers a rapid approach to identifying genomic regions associated with leaf rust resistance.
Punjab Agricultural University has pioneered wheat-wide hybridization programs, facilitating the transfer of key resistance genes from wild relatives into hexaploid wheat. In this study, we investigated an introgression line, ILkots, developed by transferring LR resistance from the non-progenitor species Ae. kotschyi (UUSS). The objective was to characterize the genetic basis of LR resistance in ILkots and map the resistance gene using tightly linked, robust SNP markers.
Results
Inheritance of resistance
The non- progenitor species, Ae. kotschyi acc pau 396 and ILkots exhibited resistant infection type against the Indian Pt pathotypes 121R60-1 (77 − 9), whereas WL711NN showed susceptible reaction with infection type of 33+ (Fig. 1). In the F2 mapping population 171 plants were classified as resistant (R) and 51 plants as susceptible (S), at both seedling and adult plant stages, indicating a favorable fit to ratio of 3 R: 1 S (χ2 = 0.37, P = 0.54). These results were further affirmed in F3 and F4 families. Among the 222 F2:3 families, 76 were Homozygous Resistant (HR), 95 were Segregating (SEG), and 51 were Homozygous Suceptible (HS) (in ratio of 1HR: 2Seg: 1HS, χ2 = 2.59, P = 0.27) while in F4 there were 80HR, 56 Seg, and 86 HS (ratio of 1.5HR: 1Seg: 1.5HS,χ2 = 0.21, P = 0.90) at both the seedling and adult plant stages(Table 1). The segregation of HR, Seg and HS progenies in F2, F3 and F4 confirmed that a single dominant gene responsible for LR resistance was transferred from Ae. kotschyi acc pau 396 into ILkots and is temporarily designated as Lrkots, as used further in text.
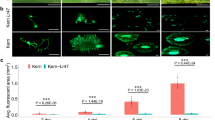
Phenotypic responses for leaf rust resistance. Infection type of (1) Ae. kotschyi acc pau 396, (2) ILkots, (3&4) WL711NN, (5–7) resistant and (8–10) susceptible progenies of ILkots/WL711NN against Pt pathotype 121R60-1 (77 − 9) at seedling stage.
Molecular mapping of Lr kots
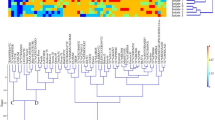
The genotypic data of 35,143 SNP markers were filtered in a sequentially by initially removing the SNPs without assigned location and chromosomal identification. From the remaining 25,996 SNPs, 4102 were found to be polymorphic between parental lines ILkots and WL711NN, distributed across all the 21 wheat chromosomes and out of these, 153 were polymorphic between Resistant Bulk (RB) and Susceptible Bulk (SB), present on all chromosomes except chromosomes 1B, 1D, 4 A, 5B, and 6D (Fig. 2). The majority of these 153 SNPs were distributed randomly across chromosomes, except chromosome 3DL where 28 SNPs were concentrated as a block in a 20 Mb region spanning from 588,341,372 bp to 613,631,868 bp. We tested 2–3 SNPs from each of chromosome by converting these into KASP assay but none of these showed polymorphism between parental lines. Twenty-one SNPs from chromosome 3DL were converted into KASP assays (Table 2) and amplified on the ILkots, WL711NN, RB and SB bulks to evaluate their discriminatory capability. Distinct cluster patterns were observed with six of these markers viz., AX-9443154, AX-94648333, AX-94752977, AX- 94874313, AX-94436339, and AX-95226287, which were subsequently amplified on 222 F4 progenies. Five KASP markers (AX-9443154, AX-94648333, AX-94752977, AX-94436339 and AX-95226287) exhibited distinct cluster patterns on F4 population with KASP marker AX-9443154, AX-94648333, AX-94752977, AX-94436339 and AX-9522,627 having 1, 6, 6, 11 and 27 recombinants, respectively with Lrkots.
Graphical representation of the distribution of polymorphic SNPs (153) differentiating between RB and SB across the chromosomes in F2:3 population derived from ILkots/WL711NN for leaf rust resistance. The chromosome and number of polymorphic SNPs are represented by X and Y axes respectively.
Five Simple Sequence Repeat (SSR) markers (Xbarc270, Xbarc323, Xbarc363, Xbarc71, and wms114) (Table 3), were also selected from 588 Mb to 609 Mb region on chromosome 3DL, where SNPs from 35 K chip showed probable linkage. These 5 SSRs were amplified on parents and bulks and the two SSRs, Xbarc363 and Xbarc71, were found to be polymorphic. SSR marker Xbarc71 amplified alleles of 115 bp in WL711NN and 100 bp in ILkots. The 100 bp allele was also amplified in the tetraploid wild accession, Ae. kotschyi pau acc 396 from which the introgression line ILkots was derived. SSR Xbarc363 behaved as a dominant marker which amplified 295 bp allele in susceptible parent WL711NN and a null allele in ILkots. When amplified on 222 F4 population, SSR Xbarc363 showed 7 recombinants and Xbarc71 showed 9 recombinants with Lrkots.
The final linkage map of Lrkotswas generated using 5 KASP markers and 2 SSR markers, covering a genetic distance of 28.5 cM on the long arm of chromosome 3D (Fig. 3). Lrkots was flanked by KASP marker AX-94443154 (0.32 cM) (Fig. 4) on the proximal end and SSR marker Xbarc71 (7.9 cM) on the distal end, spanning 8.2 cM on the map. Other markers were closely placed on genetic linkage map in order of AX-94443154, AX-94648333, AX-94752977, Xbarc363, AX-94436339 and AX-95226287 at a distance of 0.32, 1.31, 1.32, 2.63, 5.0, 8.45, and 20.6 cM respectively, from leaf rust resistance gene Lrkots. The KlusterCaller output view of segregation of KASP marker AX-9443154 is provided in Fig. 4.
(a) Genetic linkage map showing the location of resistance gene Lrkots on the long arm of chromosome 3D. (Map unit - cM) (b) Physical location of KASP & SSR markers on chromosome 3D (Map unit - Mbp) (c) Annotated disease resistance genes in the 3Mbp interval (600731378 bp to 603734145 bp) containing Lrkots. Gene IDs in orange colour are predicted to encode TIR-NBS-LRR disease resistance protein.
KlusterCaller output view of segregation of KASP marker AX-9443154 in F4 lines of mapping population derived from the ILkots/WL711NN cross. Co-dominant KASP marker segregating for FAM tailed WL711NN (blue colour) and HEX tailed ILkots alleles (red colour) on X and Y axes of the plot, respectively and heterozygous individuals for that particular SNP on mid-axis (green colour) in 384-well format. Position of each dot represents genotype of the allele and colour represents phenotype of the respective allele (colour figure online).
The genomic region containing Lrkots, between the flanking markers AX-94443154 and Xbarc71, was physically mapped onto the Chinese Spring reference genome (Fig. 3b) to identify putative candidate genes within a comparable wheat genomic region. However, since Lrkots originates from Ae. kotschyi, which lacks a reference genome, this mapping does not confirm whether the resistance gene is located on chromosome 3S or 3U in Ae. kotschyi. Despite this limitation, the flanking markers AX-94443154 and Xbarc71 were positioned at 600.73 and 603.73 Mb, respectively, in Chinese Spring. Xbarc363 and AX-94436339, which were putatively linked with Lrkots, were also positioned within this 3 Mb region at 602.11 and 603.71 Mb, respectively. Eight high-confidence putative genes associated with disease resistance were identified within this interval. Among these, four genes (TraeCS3D02G513400, TraeCS3D02G515400, TraeCS3D02G515500, and TraeCS3D02G516300) code for nucleotide-binding and leucine-rich repeats (NLR) and could be putative candidate genes for Lrkots. The details of these genes are provided in Table 4”.
Lr kots on 3D homeologue
The KASP markers AX-94443154, AX-9464833, AX-94752977, AX-94436339, AX- 95226287 and SSR markers Xbarc71, Xbarc363 found to be putatively linked with Lrkots were amplified on the nulli-tetrasomic lines CS N3A–T3B, CS N3B–T3D and CS N3D–T3A to confirm their homeologue position. The KASP markers AX-94443154, AX-9464833, AX-94752977 and AX-94436339 and SSR marker Xbarc363 amplified the alleles in nulli-tetrasomic lines CS N3A–T3B and CS N3B–T3D but not CS N3D–T3A (Fig. 5, Supplementary Figs. 1 & 2) indicating these markers were specific to homeologue D of chromosome 3. However, amplification of Xbarc71 and AX- 95226287 was observed in all the nulli-tetrasomic lines which are mapped at 7.9 cM and 20.65 cM away from the gene Lrkots indicating that these markers are not chromosome 3 homeologue specific.
Amplification of SSR (simple sequence repeats) markers (a) Xbarc363 (Supplementary Fig. 1) & (b) Xbarc71 (Supplementary Fig. 2) linked with Lrkots. Lanes: (L) 100 bp ladder, (1) CS N3A–T3B (2) CS N3D–T3A (4) CS N3B–T3D.
Discussion
Stability of disease resistance relies on the identification, mapping and deployment of new disease resistance loci and is crucial, particularly in diseases which are widespread due to the rapid evolution of pathogens. Rust resistance is a kind of threat to wheat, which poses a serious challenge to wheat growers all over the world and the scarcity of genetic variations in cultivated wheat germplasm necessitates exploration wild relatives12,25.
The genus Aegilops is the nearest relative of wheat, which includes 11 diploid, 10 tetraploid, and 2 hexaploid species, with majority of the species concentrated in the Fertile Crescent region in the Middle East and around the Mediterranean Sea26. The genomes C, D, M, N, S, T, and U found in these species have all evolved from a common ancestral source. The goldmine of genetic diversity hosted by this species is an answer to the diversity problem faced by cultivated wheat27,28,29,30. The utilization of different Aegilops species has been well-documented, showing their effectiveness in improving different traits like biotic stress of rust resistance and powdery mildew resistance26. Several LR resistance genes have been identified from Aegilops species viz., Lr9, Lr76 from Ae. umbellulata; Lr28, Lr35, Lr36, Lr37, Lr47, Lr51, Lr66 from Ae. speltoides; Lr54 from Ae. kotschyi; Lr56 from Ae. sharonensis; Lr57 from Ae. geniculata; Lr58 from Ae. triuncialis; Lr59, LrP & YrP from Ae. peregrina; Lr62 from Ae. neglecta31,32,33.
In the current study, we explore the novel leaf rust resistance derived from tetraploid non-progenitor of wheat, Ae. kotschyi acc pau 396 with US genome. Ae. kotschyi, was reported to evolve by hybridization between diploid Ae. longissima (genome SlSl) and diploid Ae. umbellulata (genome UU)34. This species, native to the Irano-Turanian region, is well-suited to survive in desert climates, particularly in the Saharo-Arabian region. It is widely distributed across Transcaucasia, the Republics of Central Asia, Afghanistan, Pakistan, Turkey, Kuwait, Lower Egypt, Tunisia, and the Mediterranean region and can thrive in different soil types34,35. The US genome of Ae kotschyi, a tetraploid wild relative of wheat, holds promise in enhancing wheat’s disease resistance and nutritional value. It has been recognized as a source of resistance against Lr and Yr, as well as having high levels of grain mineral content19,36,37. The partial amphiploids of wheat - Ae kotschyi have elevated levels of Zn and Fe in the grain, than the parental wheat line38,39. It was reported that the Zn grain content was three times higher in wheat- Ae. kotschyi addition/substitution lines than the wheat parent40. Further, High-Molecular-Weight Glutenin Subunits (HMWGS) of 1U from a wheat- Aegilops substitution line were transferred with 1U chromosome to wheat41.
To effectively utilise resistance from Ae. kotschyi in the present study, it needs to be characterized by transferring into hexaploid background. Since US genome of Ae. kotschyi differs substantially from ABD genome of wheat, it’s transfer into susceptible hexaploid background PBW343 was done by inducing homoeologous pairing between these two genomes using Ph1b suppressor stock of Chinese Spring (CS Ph1b)42. The introgression line ILkots was selected for their leaf rust resistance, chromosome number 2n = 42, and high similarity to the recurrent parent PBW343, as an indicative of alien introgression13,27. Novelty of leaf rust resistance in ILkots was evident from its resistance against multiple leaf rust races, including one of the most prevalent pathotype 121R60-1 (77 − 9)6,43 both at seedling and adult plant stages for multiple years (Fig. 6). Pathotype 121R60-1 (77 − 9) is a variant of race 77, which became prevalent from the year 20161,6. It is very similar to pathotype 121R63-1 (77 − 5), which has remained as one of the most prevailing and virulent pathotype for the past 20 years43.
Flow chart illustrating (a) the development Introgression line and (b) the development of mapping population. (a) Leaf rust (LR) resistant Introgression line Kotschyi 396 (ILkots) was derived from wild tetraploid non-progenitor Ae. kotschyi (UUSS) accession pau 396, through first crossing with Ph1b mutant version of CS(S) and then backcrossing with LR susceptible wheat cultivar PBW343. (b) The leaf rust resistant ILkots was crossed with leaf rust susceptible cultivar WL711NN and selfed for few generations to develop mapping population.
A total of 83 leaf rust resistance genes (Lr1 to Lr83) have been identified on the 21 wheat chromosomes44,45,46,47. Still, efforts are going on to discover novel resistance genes from diverse sources because of wheat rust’s rapid ability to counter the host’s defence mechanisms. Through mutations or recombination, the pathogen undergoes a continuous evolution that gives rise to more aggressive races with improved patterns of virulence. In addition, new races migrate from other geographic regions, introducing one or more new avirulence genes that may not be present in the current host cultivars’ corresponding R genes48. As a result, the host resistance breaks down and is transient. Hence, it is crucial to discover novel R genes that can confer resistance to highly virulent avirulence genes.
The genetic analysis on F2 and F3 mapping population indicated transfer of a single dominant all stage leaf rust resistance gene from Ae. kotschyi supporting the fact that minimal US genome introgression happened in ILkots led to the transfer of one LR resistance gene only. The seedlings screened against Pt pathotypes were transplanted in the same order to the field such that the same plant is evaluated against the Pt pathotypes at seedling and adult plant stages to confirm all stage resistance. Bulked segregant analysis (BSA) on F2 mapping population combined with 35K SNP chip analysis comprehensively identified SNPs linked with leaf rust resistant loci on chromosome 3DL. BSA can rapidly identify the genetic locus underlying a phenotypic trait, by avoiding the need to genotype the entire population, making it a major improvement over traditional linkage mapping. The wheat genome is highly repetitive, owing to its polyploid nature which poses a challenge in the mapping of desirable genes. To reduce the impact of genome complexity, a blend of BSA with latest sequencing technologies49,50,51 has proved to be game changing for efficient mapping of important genes25,52. The development of high throughput and cost-effective genotyping platforms such as genotype by sequencing53 and high-density SNP arrays21 has revolutionized the genetic mapping of economically important traits. Various high density SNP genotyping arrays like 35K have been used for mapping LR resistance gene from Triticum spelta54, 90 K has been employed to map Yr80 from a common wheat landrace Aus2728455 and Lr82 from wheat landrace Aus2735256. A stripe rust resistance QTL on the long arm of chromosome 7B, was mapped using a combination of bulk segregant analysis and 600 K genotyping array57.
The linked SNPs identified in silico from the 35K SNP chip were validated through Kompetitive aAllele specific (KASP) assays. SNP markers have emerged as the most informative and efficient molecular markers for creating high-density maps30,53,58. The KASP technology has revolutionized the validation of SNPs and has been extensively implemented for tagging of rust resistance genes in wheat59 such as Lr76 - Yr7030, Yr80 tagged with KASPs at 3 cM55, Lr80 tagged with KASPs at 0.2 cM60, Lr82 KASPs mapped at 0.8 cM56.
The SNPs are the most popular and abundant markers in the current time but the SSRs still have not lose their charm due to their easy to detect and co-dominance qualities. Two SSR markers Xbarc71 and Xbarc363, were also used in present study to saturate the linkage map on chromosome 3DL containing the linked SNPs with Lrkots. Due to the unavailability of the Aegilops kotschyi genome, the linkage map was constructed using the reference genome of Chinese spring, which may not perfectly align with the Aegilops kotschyi genome61. This disparity may lead to discrepancies between the physical and genetic maps. Furthermore, the nulli-tetrasomic lines; CS N3A–T3B, CS N3B–T3D and CS N3D–T3A of chromosome 3 confirmed the presence of Lrkots on D homeologue62,63. The non-amplification of AX-94443154, AX-9464833, AX-94752977, AX-94436339 and Xbarc363 markers in nulli-tetrasomic line without homeologue 3D (Fig. 5, Supplementary Fig. 1&2) proved their location on chromosome 3D alongwith gene Lrkots. Comparison of genetic map with physical positions of all the markers showed that five markers (AX-94752977, AX-94648333, AX-94443154, Xbarc363, AX-94436339 and Xbarc71) are located within 3 Mbp region (600.73 Mbp to 603.73 Mbp) (Fig. 3a and b).
So far, only the associated leaf rust and stripe rust resistance genes Lr54-Yr37 have been documented from Ae. kotschyi and is located on chromosome 2DL of hexaploid wheat16,19. In the present work, the Lrkots resistance gene identified from Ae. kotschyi is a single gene, possibly derived from chromosome 3U or 3S, and is different from the Lr54-Yr37 cluster. Wheat Chromosome 3DL has been reported to carry other leaf rust resistance genes such as Lr5R, Lr24, and Lr69. Of these, Lr24 is very effective against the dominant leaf rust races in India64. However, it was originally introgressed from the Agropyron elongatum (= Thinopyrum elongatum or Lophopyrum elongatum; 2n = 2x = 14, EE)65,66. The presence of Lr24 in ILkots was excluded since the Lr24-associated STS marker (J09)65,67 did not amplify in ILkots (Fig. 7, Supplementary Fig. 3). In the same manner, Lr5R and Lr6968, which are indigenous to bread wheat, were ruled out as candidate genes for the expressed resistance. The results of this research indicate that Lrkots is a new leaf rust resistance gene, providing high promise for wheat rust resistance breeding programs.
Amplification of STS (sequence-tagged sites) marker linked for Lr24 gene Lanes: (L) 1000 bp ladder, (1&2) WL711NN (3) Ae. kotschyi acc pau 396, (4) ILkots (Supplementary Fig. 3).
Although our research emphasizes the potential utility of Lrkots in wheat breeding, we do recognize some limitations that need to be addressed. Further research is required to fine-map the Lrkots locus to gene identity to enable functional characterization. Inconsistencies exist between physical and genetic maps generated in this study, due to the lack of a reference genome for Aegilops kotschyi. We intend to address these issues in further research to provide a comprehensive evaluation of Lrkots in practical breeding applications. In addition, the potential effects of gene-environment interactions on its effectiveness need to be analysed. The necessity for pyramiding with other resistance genes to guarantee stability, and the possible difficulties in transferring the gene to various genetic backgrounds are the major issues associated with characterization and transfer of this locus.
Overall, we have identified and mapped a new gene from Aegilops kotschyi, designated Lrkots, that contributes resistance to the predominant races of leaf rust in India both at seedling and adult plant stage. The linked markers will help marker assisted transfer of this gene into cultivated wheat varieties with enhanced resistance to leaf rust reducing yield losses, ultimately contributing to food security and sustainable agriculture. Deployment of genetic resistance always reduce the reliance on chemical fungicides, promoting environmentally friendly and economically viable disease management strategies.
Materials and methods
Plant materials
The plant material used in this study includes LR resistant Introgression line ILkots, LR susceptible cultivar WL711NN, and the F3:4 mapping population derived from the cross of ILkots/WL711NN (Fig. 6). The wild accession of Aegilops kotschyi acc pau 396 has been collected from National Bureau of Plant Genetic Resources (NBPGR), New Delhi, a national repository with number EC573277. The Introgression line ILkots, was formally identified and developed by Dr. Satinder Kaur at Punjab Agricultural University, using established morphological traits and taxonomic keys (Fig. 6). The Indian Agricultural Research Institute, New Delhi, India developed WL711 as one of the wheat derivatives in 1965 under the All India Coordinated Wheat Improvement Project (AICWIP)14. It became a popular wheat variety in Punjab state and the northwestern plain zone. WL711NN is a near isogenic line (NIL) that was developed from T. aestivum WL711 and is characterized by the presence of kr (crossability) alleles69. LR resistant ILkots was derived from wild tetraploid non-progenitor Ae. kotschyi (UUSS) accession pau 396, through first crossing with Ph1b mutant version of CS(S)42 and then backcrossing with LR susceptible wheat cultivar PBW343. In each backcross generation, the LR resistant plants, phenotypically like PBW343 were carried forward, in BC1F6 one LR resistant plant with chromosome number 2n = 42 was finally selected and designated as ILkots. To understand the genetics of LR resistance and map resistance associated genomic regions, the F2, F3 and F4 mapping populations derived from the ILkots/ WL711NN cross were used in the current study. The studies were conducted at the School of Agricultural Biotechnology (SOAB), Punjab Agricultural University (PAU), located in Ludhiana, Punjab has a latitude of approximately 30°54’03.5″N and longitude of approximately 75°48’41.5″E.
Screening against leaf rust
Puccinia triticina (Pt) pathotype 109R31-1 (77 − 5), 21R55 (104-2), and 121R60-1 (77 − 9) were used to screen the mapping population, ILkots, and WL711NN. The Pt pathotype 109R31-1 (77 − 5) is avirulent on Lr9, Lr19, Lr24, Lr25, Lr29, Lr28, Lr39, Lr32, Lr43, Lr42, Lr45 & Lr47, 104-2 is avirulent on Lr9, Lr10, Lr13, Lr15, Lr19, Lr20, Lr24, Lr25, Lr28, Lr29, Lr32, Lr36, Lr40, Lr41, Lr42, Lr43 & Lr45) and 121R60-1 (77 − 9) is avirulent on Lr2a, Lr2b, Lr2c, Lr9, Lr19, Lr24, Lr28, Lr25, Lr32, Lr39, Lr42, Lr45 & Lr4743.
At the seedling stage, screening was conducted using the Pt pathotype 121R60-1 (77 − 9). The seeds were sown in bread boxes in a glasshouse. Ten days old seedlings were inoculated with rust spores well mixed with the talcum powder, at a rate of 1 V of fresh urediniospores to 20 V of talcum powder70. The inoculated seedlings were incubated in a humid chamber (100% relative humidity) for 24 h and transferred to the benches of a glasshouse maintained at a temperature of 20 ºC ± 2ºC. Infection types were recorded 14 days post-inoculation according to Ref.71.
After recording data on seedlings, the same seedlings were transplanted in open field and grown into adult plants, to score LR data of seedling stage and adult plant stage on same plant. The LR inoculum load in the field was maintained by planting and inoculating rows of LR susceptible cultivar all around the experimental plot. Suspensions of mixture from urediniospores of Pt pathotypes 109R31-1 (77 − 5), 21R55 (104-2), and 121R60-1 (77 − 9) were sprayed in the experimental area between January to February. The LR responses were recorded thrice from February to March, as percentage of leaf area covered using a modified Cobb’s scale, taking into consideration the type of spore and the percentage of leaf area covered72.
Genomic DNA extraction and bulk prepration
Genomic DNA was extracted from 35 to 40 days old leaf tissue though modified CTAB (Cetyl Trimethyl Ammonium Bromide) method73. After assessment of DNA quality the DNA concentration was normalized. Twenty resistant and 20 susceptible F2 plants were selected and equal amounts of DNA from 10 Resistant and 10 Susceptible F2 plants were pooled to form the two Resistant Bulks (RB) and two Susceptible Bulks (SB). For bulk preparation only those F2 plants were selected whose F3 progenies were homozygous resistant/homozygous susceptible.
Markers analysis
Genotyping of ILkots, WL711NN, RB and SB were carried out using the AFFYMETRIX 35 K wheat breeder’s axiom array23. SNPs lacking physical descriptions were eliminated. SNPs showing polymorphism between resistant and susceptible parents and RB & SB were filtered out. Selected SNPs were converted to KASP (Kompetitive allele Specific PCR) markers and were PCR amplified as described by55. The fluorescence signals were read by Tecan Safire infinite F200 PRO plate reader. These signals were imported and visualized using the Kluster caller software (version 2.22.0.5, LGC genomics) and were analysed.
In addition to SNPs, SSR markers were also designed from selected chromosomal region and amplified in 10 µl reaction mixtures containing 50–100 ng DNA, 0.375 µM of each forward and reverse primer and PCR master mix (Emerald Amp GT 2X) at profile: 94 °C for 4 min, 94 °C for 30 s, 60 °C for 30 s, 72 °C for 20 s, 72 °C for 10 min (30 cycles). The PCR amplified products were separated on a 3.5% Agarose gel and visualized on UV-trans illuminator.
Confirmation of Lr kots on 3D homeologue
To confirm the position of markers linked with LR resistance, the markers were amplified on the nullisomic- tetrasomic lines in wheat cultivar Chinese Spring background CS N3A–T3B, CS N3B–T3D and CS N3D–T3A (obtained from Wheat Genetics Resource Center, Kansas State University, Manhattan).
Statistical analysis and genetic map construction
The F2 plants derived from ILkots/WL711NN, were categorized into resistant (R) and susceptible (S) while the F3 and F4 generation were classified into homozygous resistant (HR), segregating (Seg), and homozygous susceptible (HS) families. Chi-squared (χ2) tests were used to determine goodness of fit of the observed rust response and marker segregation with the expected genetic segregation ratios. The linkage map was generated using QTL-Ici mapping software at a LOD score of 3.0, and genetic distances in centiMorgans (cM) based on recombination fractions were calculated using the Kosambi mapping function.
Data availability
Supporting data is provided within the manuscript. The datasets generated and analysed during this study are based on SNP array data. These datasets and the plant material used in this study are available from the corresponding author upon reasonable request to ensure research reproducibility and transparency.
References
Bhardwaj, S. C. et al. Physiologic specialization and shift in Puccinia triticina pathotypes on wheat in Indian Subcontinent during 2013–2016. Indian Phytopathol. 72, 23–34 (2019a).
Van Dijk, M., Morley, T., Rau, M. L. & Saghai, Y. A meta-analysis of projected global food demand and population at risk of hunger for the period 2010–2050. Nat. Food. 2, 494–501 (2021).
AICRP on Wheat & Barley. http://www.aicrpwheatbarleyicar.in (2025).
Basnet, B., Juliana, P., Bhattarai, K. & Upreti, U. A review on major rust resistance gene and amino acid changes on wheat (Triticum aestivum L). SSRN Electron. J. https://doi.org/10.2139/ssrn.4258549 (2022).
Kumar, S. et al. Mining of Indian wheat germplasm collection for adult plant resistance to leaf rust. PLoS One. 14, e0213468 (2019a).
Raghunandan, K. et al. Identification of novel broad-spectrum leaf rust resistance sources from Khapli wheat landraces. Plants 11, 1965 (2022).
Kaur, S. et al. Identification of leaf rust resistance loci in a geographically diverse panel of wheat using genome-wide association analysis. Front. Plant Sci. 14 (2023).
Tomar, S. M. S., Singh, S. K., Sivasamy, M. & Vinod Wheat rusts in India: resistance breeding and gene deployment -A review. Indian J. Genet. Plant. Breed. (The). 74, 129 (2014).
Pal, D., Bhardwaj, S. C., Khan, H. & Patial, M. ’HS628’-A potential genetic stock for resistance to new virulent pathotypes of black, brown and yellow rusts of wheat (Triticum aestivum L). Wheat Barley Res. 10, (2018).
Koláriková, L. et al. Leaf rust resistance genes in Aegilops genus: occurrence and efficiency. Eur. J. Plant Pathol. 167, 335–348 (2023).
Kishii, M. An update of recent use of Aegilops species in wheat breeding. Front. Plant Sci. 10, (2019).
Bansal, M. et al. Mapping of Aegilops umbellulata-derived leaf rust and Stripe rust resistance loci in wheat. Plant. Pathol. 66, 38–44 (2016).
Narang, D. et al. Discovery and characterisation of a new leaf rust resistance gene introgressed in wheat from wild wheat Aegilops Peregrina. Sci. Rep. 10 (2020).
Arora, S. et al. Introgression and genetic mapping of leaf rust and Stripe rust resistance in Aegilops triuncialis. J. Genet. 100, (2021).
Prasad, P., Savadi, S., Bhardwaj, S. C. & Gupta, P. K. The progress of leaf rust research in wheat. Fungal Biol. 124, 537–550 (2020).
Marais, G. F., McCallum, B., Snyman, J. E., Pretorius, Z. A. & Marais, A. S. Leaf rust and Stripe rust resistance genes Lr54 and Yr37 transferred to wheat from Aegilops kotschyi. Plant. Breed. 124, 538–541 (2005).
Chhuneja, P., Dhaliwal, H. S., Bains, N. S. & Singh, K. Aegilops kotschyi and Aegilops tauschii as sources for higher levels of grain Iron and zinc. Plant. Breed. 125, 529–531 (2006).
Verma, S. K. et al. Transfer of useful variability of high grain iron and zinc from Aegilops kotschyi into wheat through seed irradiation approach. Int. J. Radiat. Biol. 92, 132–139 (2016).
Kwiatek, M. T. et al. Molecular identification of triticale introgression lines carrying leaf rust resistance genes transferred from Aegilops kotschyi Boiss. and Ae. tauschii Coss. J. Appl. Genet. https://doi.org/10.1007/s13353-021-00635-2 (2021).
Luo, M. C. et al. Genome sequence of the progenitor of the wheat D genome Aegilops tauschii. Nature 551 (7681), 498–502 (2017).
Wang, S. et al. Characterization of polyploid wheat genomic diversity using a high-density 90 000 single nucleotide polymorphism array. Plant Biotechnol. J. 12, 787–796 (2014b).
Dubcovsky, J. & Dvorak, J. Genome plasticity a key factor in the success of polyploid wheat under domestication. Science 316 (5833), 1862–1866 (2007).
Allen, A. M. et al. Characterization of a wheat breeders’ array suitable for high-throughput SNP genotyping of global accessions of hexaploid bread wheat (Triticum aestivum). Plant Biotechnol. J. 15, 390–401 (2016).
You, Q. et al. Development and applications of a high throughput genotyping tool for polyploid crops: single nucleotide polymorphism (SNP) array. Front. Plant Sci. 9, (2018).
Kaur, S., Jindal, S. & Kaur, M. Utilization of wild species for wheat improvement using genomic approaches. Biotechnologies Crop Improv. 3, 105–150 (2018).
Kumar, A., Kapoor, P., Chunduri, V., Sharma, S. & Garg, M. Potential of Aegilops Sp. for improvement of grain processing and nutritional quality in wheat (Triticum aestivum). Front. Plant Sci. 10 (2019).
Kuraparthy, V. et al. Characterization and mapping of cryptic alien introgression from Aegilops geniculata with new leaf rust and Stripe rust resistance genes Lr57 and Yr40 in wheat. Theor. Appl. Genet. 114, 1379–1389 (2007).
Kuraparthy, V., Sood, S., Guedira, G. B. & Gill, B. S. Development of a PCR assay and marker-assisted transfer of leaf rust resistance gene Lr58 into adapted winter wheats. Euphytica 180, 227–234 (2011).
Narang, D., Kaur, S., Saini, J. & Chhuneja, P. Development and molecular characterization of Wheat-Aegilops Peregrine introgression lines with resistance to leaf rust and Stripe rust. J. Crop Improv. 32, 59–70 (2017).
Bansal, M. et al. Aegilops umbellulata introgression carrying leaf rust and Stripe rust resistance genes Lr76 and Yr70 located to 9.47-Mb region on 5DS telomeric end through a combination of chromosome sorting and sequencing. Theor. Appl. Genet. 133, 903–915 (2020).
Helguera, M. et al. PCR markers for Triticum Speltoides leaf rust resistance gene Lr51 and their use to develop isogenic hard red spring wheat lines. Crop Sci. 45, 728–734 (2005).
Marais, G. F., Badenhorst, P. E., Eksteen, A. & Pretorius, Z. A. Reduction of Aegilops sharonensis chromatin associated with resistance genes Lr56 and Yr38 in wheat. Euphytica 171, 15–22 (2009).
Marais, F., Marais, A., McCallum, B. & Pretorius, Z. Transfer of leaf rust and Stripe rust resistance genes Lr62 and Yr42 from Aegilops neglecta req. Ex Bertol. To common wheat. Crop Sci. 49, 871–879 (2009).
Weissmann, S., Feldman, M. & Gressel, J. Sequence evidence for sporadic intergeneric DNA introgression from wheat into a wild Aegilops species. Mol. Biol. Evol. 22, 2055–2062 (2005).
Kimber, G. & M. Feldman. Wild Wheat, an Introduction. Special Report 353, College of Agriculture, 1–142 (University of Missouri, 1987).
Chhuneja, P., Dhaliwal, H. S., Bains, N. S. & Singh, K. Aegilops kotschyi and Aegilops tauschii as sources for higher levels of grain Iron and zinc. Plant. Breed. 125, 529–531 (2006).
Verma, S. K. et al. Transfer of useful variability of high grain iron and zinc from Aegilops kotschyi into wheat through seed irradiation approach. Int. J. Radiat. Biol. 92, 132–139 (2016).
Tiwari, V. K. et al. Development of Triticum turgidum subsp. durum–Aegilops longissima amphiploids with high iron and zinc content through unreduced gamete formation in F1 hybrids. Genome 51, 757–766 (2008).
Tiwari, V. K. et al. Random chromosome elimination in synthetic Triticum–Aegilops amphiploids leads to development of a stable partial amphiploid with high grain micro- and macronutrient content and powdery mildew resistance. Genome 53, 1053–1065 (2010).
Rawat, N. et al. Development and molecular characterization of wheat –Aegilops kotschyi addition and substitution lines with high grain protein, iron, and zinc. Genome 54, 943–953 (2011).
Singh, J. et al. Transfer of HMW glutenin subunits from Aegilops kotschyi to wheat through radiation hybridization. J. Food Sci. Technol. 53, 3543–3549 (2016).
Chen, P. D., Tsujimoto, H. & Gill, B. S. Transfer of PhI genes promoting homoeologous pairing from Triticum Speltoides to common wheat. Theor. Appl. Genet. 88, 97–101 (1994).
Kaur, H. et al. Postulation of leaf rust resistance genes in Indian and exotic wheat germplasm using near-isogenic lines (NILs) and molecular markers. Crop Prot. 174, 106431 (2023b).
McIntosh, R. A., Dubcovsky, J., Rogers, W. J., Xia, X. C. & Raupp, W. J. Catalogue of gene symbols for wheat. Annual Wheat Newsl. 66 (2020).
Kumar, K. et al. An update on resistance genes and their use in the development of leaf rust resistant cultivars in wheat. Front. Genet. 13, (2022).
Bokore, F. E. et al. Genetic mapping of leaf rust (Puccinia triticina Eriks) resistance genes in six Canadian spring wheat cultivars. Front. Plant. Sci. 14, 575. https://doi.org/10.3389/fpls.2023.1130768 (2023).
Kolmer, J. A., Bajgain, P., Rouse, M. N., Li, J. & Zhang, P. Mapping and characterization of the recessive leaf rust resistance gene Lr83 on wheat chromosome arm 1DS. Theor. Appl. Genet. 136, (2023).
Bolton, M. D., Kolmer, J. A. & Garvin, D. F. Wheat leaf rust caused by Puccinia triticina. Mol. Plant Pathol. 9, 563–575 (2008).
Trick, M. et al. Combining SNP discovery from next-generation sequencing data with bulked Segregant analysis (BSA) to fine-map genes in polyploid wheat. BMC Plant Biol. 12, (2012).
Dong, C. et al. Combining a new exome capture panel with an effective VarBScore algorithm accelerates BSA-based gene cloning in wheat. Front. Plant Sci. 11, (2020).
Klymiuk, V. et al. Discovery of Stripe rust resistance with incomplete dominance in wild emmer wheat using bulked Segregant analysis sequencing. Commun. Biology 5, (2022).
Chhuneja, P., Kaur, S. & Dhaliwal, H. S. Introgression and exploitation of biotic stress tolerance from related wild species in wheat cultivars. Mol. Breed. Sustainable Crop Improv. 2, 269–324 (2016).
Poland, J. A. & Rife, T. W. Genotyping-by‐sequencing for plant breeding and genetics. Plant. Genome 5, (2012).
Dinkar, V. et al. Molecular mapping of a new recessive wheat leaf rust resistance gene originating from Triticum spelta. Sci. Rep. 10 (2020).
Nsabiyera, V. et al. Characterisation and mapping of adult plant Stripe rust resistance in wheat accession Aus27284. Theor. Appl. Genet. 131, 1459–1467 (2018).
Bariana, H. S., Babu, P., Forrest, K. L., Park, R. F. & Bansal, U. K. Discovery of the new leaf rust resistance gene Lr82 in wheat: molecular mapping and marker development. Genes 13, 964 (2022).
Mu, J. et al. Genetic architecture of wheat Stripe rust resistance revealed by combining QTL mapping using SNP-based genetic maps and bulked Segregant analysis. Theor. Appl. Genet. 132, 443–455 (2018).
Hussain, W. et al. Genotyping-by-sequencing derived high-density linkage map and its application to QTL mapping of flag leaf traits in bread wheat. Sci. Rep. 7 (2017).
Ramirez-Gonzalez, R. H., Uauy, C. & Caccamo, M. PolyMarker: A fast polyploid primer design pipeline. Bioinformatics 31, 2038–2039 (2015).
Kumar, S. et al. Lr80: A new and widely effective source of leaf rust resistance of wheat for enhancing diversity of resistance among modern cultivars. Theor. Appl. Genet. 134, 849–858 (2021).
Brenchley, R. et al. Analysis of the bread wheat genome using whole-genome shotgun sequencing. Nature 491 (7426), 705–710 (2012).
Sears, E. R. The aneuploids of common wheat. Res. Bull. Missouri Agricultural Exp. Stn. 572, 1–58 (1954).
Sears, E. R. & Sears, L. M. S. The telocentric chromosomes of common wheat. In: (Ramanujam S, ed.) Proc. of the Fifth International Wheat Genetics Symposium. Volume 1. Session IV. Cytogenetics. New Delhi: Indian Society of Genetics and Plant Breeding, 389–407 (1978).
Bhardwaj, S. C., Singh, G. P., Gangwar, O. P., Prasad, P. & Kumar, S. Status of wheat rust research and progress in rust management-indian context. Agronomy 9, 892 (2019b).
Schachermayr, G. M. et al. Identification of molecular markers linked to the Agropyron elongatum-derived leaf rust resistance gene Lr24 in wheat. Theor. Appl. Genet. 90, 982–990 (1995).
Gupta, S. K., Charpe, A., Koul, S., Haque, Q. M. R. & Prabhu, K. V. Development and validation of SCAR markers co-segregating with an Agropyron Elongatum derived leaf rust resistance gene Lr24 in wheat. Euphytica 150, 233–240 (2006).
Zhang, L. et al. Evaluation of the resistance to Chinese predominant races of Puccinia triticina and analysis of effective leaf rust resistance genes in wheat accessions from the U.S. National plant germplasm system. Front. Plant Sci. 13 (2022).
WANG, J., ZHU, S. H. I. L., LI, L., LIU, D. & X. & Genetic analysis and molecular mapping of leaf rust resistance genes in the wheat line 5R618. Czech J. Genet. Plant. Breed. 50, 262–267 (2014).
Randhawa, M. S. et al. Interactions among genes Sr2/Yr30, Lr34/Yr18/Sr57 and Lr68 confer enhanced adult plant resistance to rust diseases in common wheat (Triticum aestivum L.) line ‘arula’. Aust. J. Crop Sci. 12, 1023–1033 (2018).
Roelfs, A. P., Singh, R. P. & Saari, E. E. Rust Diseases of Wheat: Concepts and Methods of Disease Management (Cimmyt, 1992).
Stakman, E. C., Stewart, D. M. & Loegering, W. Q. Identification of physiologic races of Puccinia graminis Var. tritici. Identif. Physiologic Races Puccinia graminis Var. tritici (1962).
Peterson, R. F., Campbell, A. B. & Hannah, A. E. A diagrammatic scale for estimating rust intensity on leaves and stems of cereals. Can. J. Res. 26c, 496–500 (1948).
Saghai-Maroof, M. A., Soliman, K. M., Jorgensen, R. A. & Allard, R. W. Ribosomal DNA spacer-length polymorphisms in barley: Mendelian inheritance, chromosomal location, and population dynamics. Proc. Natl. Acad. Sci. 81, 8014–8018 (1984).
Acknowledgements
B.K.B is grateful to Indian Council of Agricultural Research for financial assistance given as JRF and SRF during study at Punjab Agricultural University, Ludhiana, Punjab. This work was supported by the Department of Biotechnology (DBT) under the grant number BT/PR43090/AGIII/103/1317/2021 and BT/AG/Network/Wheat/2019-20. TS also acknowledges the receipt of BioCARe grant and fellowship (BT/PR51286/BIC/101/1353/2023) from Department of Biotechnology (DBT), Govt. of India. The authors are grateful to the ICAR-Indian Institute of Wheat & Barley Research, Regional Station, Flowerdale, Shimla, for providing pure inoculum of P. triticina pathotypes and the Punjab Agricultural University, Ludhiana, for facilitating the experiments.
Author information
Authors and Affiliations
Contributions
B.K.B population development, rust screening, marker analysis, data acquisition, data analysis and interpretation, drafting of manuscript; B.K marker analysis and drafting of manuscript; S.K Identification of plant material and development of Introgression Line, study supervision, data interpretation and review of manuscript; T.S, J.K and A.S review of manuscript; P.C; concept and design of study, study supervision, data interpretation and review of manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Bai, B.K., Kaur, B., Kaur, S. et al. Unveiling leaf rust resistance introgressed from non-progenitor wild species Aegilops kotschyi into hexaploid wheat. Sci Rep 15, 44063 (2025). https://doi.org/10.1038/s41598-025-99077-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-99077-7