Abstract
White blood cell (WBC) detection is pivotal in medical diagnostics, crucial for diagnosing infections, inflammations, and certain cancers. Traditional WBC detection methods are labor-intensive and time-consuming. Convolutional Neural Networks (CNNs) are widely used for cell detection due to their strong feature extraction capability. However, they struggle with global information and long-distance dependencies in WBC images. Transformers, on the other hand, excel at modeling long-range dependencies, which improves their performance in vision tasks. To tackle the large foreground-background differences in WBC images, this paper introduces a novel WBC detection method, named the Multi-Scale Cross-Deformation Attention Fusion Network (MCDAF-Net), which combines CNNs and Transformers. The Attention Multi-scale Sensing Module (AMSM) is designed to localize WBCs more accurately by fusing features at different scales and enhancing feature representation through a self-attention mechanism. The Cross-Deformation Convolution Module (CDCM) reduces feature correlation, aiding the model in capturing diverse aspects and patterns in images, thereby improving generalization. MCDAF-Net outperforms other models on public datasets (LISC, BCCD, and WBCDD), demonstrating its superiority in WBC detection. Our code and pretrained models: https://github.com/xqq777/MCDAF-Net.
Similar content being viewed by others
Introduction
White blood cell (WBC) detection is a cornerstone of modern medical diagnostics, providing critical insights into a wide range of health conditions, including anemia, infections, inflammation, and immune system disorders1,2. Despite its significance, WBC detection remains a formidable challenge due to the inherent complexities of medical imaging, such as variations in staining techniques, imaging conditions, and the frequent occurrence of multicellular adhesion phenomena3,4,5,6,7,8,9,10,11,12,13,14,15. Recent advancements in deep learning have revolutionized the field of medical image analysis, with Convolutional Neural Networks (CNNs) and Transformers emerging as the two dominant architectures. While CNNs excel in hierarchical feature extraction, Transformers leverage self-attention mechanisms to capture long-range dependencies, making them highly effective in complex vision tasks. However, both architectures have limitations when applied to WBC detection, particularly in handling scale variability, computational efficiency, and feature redundancy. This study addresses these challenges by proposing a novel hybrid model, MCDAF-Net, which integrates the strengths of CNNs and Transformers while introducing innovative modules to enhance performance.
CNNs have long been the backbone of medical image recognition due to their ability to construct hierarchical representations through convolution and pooling operations. Several studies have demonstrated their efficacy in WBC detection. For instance, Geng et al. 3 utilized the attention mechanism of Mask R-CNN4 to improve WBC segmentation accuracy. Zheng et al. 5 combined Itti’s visual attention model with an adaptive center-surround difference operator and an enhanced CenterNet model for WBC detection. Sivarao et al. 6 proposed a framework using SegNet for segmentation and EfficientNet for feature extraction, achieving state-of-the-art classification of WBC subtypes. Islam et al. 7 further advanced CNN-based methods by incorporating image pre-processing techniques and interpretability tools like SHapley Additive exPlanations(0.40.0) and Gradient-weighted Class Activation Mapping++(1.7.1), outperforming existing models.
Despite these advancements, CNN-based approaches often rely on a two-stage detection framework, which introduces parameter redundancy and increases computational overhead. To address these issues, researchers have proposed optimized architectures. Xu et al. 16 introduced TE-YOLOF, which incorporates depth-separable convolutions and EfficientNet17 as a backbone. Han et al. 18 developed MID-YOLO, integrating attention mechanisms to enhance contextual understanding. Wang et al. 19 enhanced YOLOv5 with coordinate attention to better handle large-scale WBC samples. Polejowska et al. 20 investigated the impact of image quality on detection performance using YOLOv5 and RetinaNet21. However, CNN-based detectors still struggle with scale variability and feature representation, particularly for small objects, highlighting the need for more robust solutions.
Transformers22, originally designed for natural language processing, have recently gained traction in vision tasks due to their self-attention mechanisms, which excel at capturing long-range dependencies. Carion et al. 23 redefined object detection as a set prediction problem, achieving competitive results on the COCO dataset24. Zhu et al. 25 introduced Deformable-DETR, which enhances multi-scale feature handling. Wang et al. 26 proposed PnP-DETR to address spatial redundancy, reducing computational load. Zhang et al. 27 improved DETR’s training convergence and query significance through contrastive denoising training. In medical imaging, Chen et al. 28 designed a Vision Transformer (ViT) with shifted windows and transfer learning for WBC classification on the BCCD dataset. Li et al. 29 improved the Detection Transformer’s residual module, while Dipto et al. 30 employed Explainable AI (XAI) with federated learning to accelerate ViT training. Katar et al. 31 developed an interpretable ViT model using self-attention and Score-CAM for clinical applications.
Despite their promise, Transformers face significant challenges in medical imaging. Their self-attention mechanisms incur high computational complexity, making them less efficient for high-resolution images. Additionally, Transformers require large datasets to perform optimally, a limitation in data-scarce medical domains. These challenges underscore the need for innovative approaches that can leverage the strengths of Transformers while mitigating their limitations.
The integration of CNNs and Transformers has emerged as a promising approach to leverage their complementary strengths. Marzahl et al. 32 proposed a leukocyte detection method combining region-based proposals with attention mechanisms. Huang et al. 33 introduced ARML, which enhances feature representation with adaptive attention-aware residuals. Nugraha et al. 34 combined YOLOv8 and DETR to enhance multi-object detection performance, particularly with insufficient datasets. Tarimo et al. 35 designed a 2-way-2-stage approach integrating YOLO for fast object detection and ViT for robust image representation. Zhang et al. 36 enhanced YOLO with multi-scale feature integration, while Bayat et al. 37 proposed a multi-attention framework for fine-grained WBC classification.
However, current hybrid models remain in their early developmental stages, struggling with complex, multi-scale features and channel feature redundancy. To address these challenges, this study introduces MCDAF-Net, a novel network model tailored for WBC detection. The key innovations of this work include:
-
Attention Multi-scale Sensing Module (AMSM): This module combines multi-scale dilation convolution with self-attention mechanisms to capture critical features effectively, addressing the limitations of traditional CNNs and Transformers in handling scale variability.
-
Cross-Deformation Convolution Module (CDCM): This module extends feature representation through a ‘Split-Crossover-Fusion Deformation’ strategy, reducing channel feature redundancy and enhancing the model’s ability to distinguish between closely adhered WBCs.
-
Multi-Scale Cross-Deformation Attention Fusion Network (MCDAF-Net): By integrating AMSM and CDCM, this module achieves superior performance on public datasets such as LISC, BCCD, and WBCDD, setting a new benchmark for WBC detection accuracy and efficiency.
The proposed MCDAF-Net represents a significant advancement in WBC detection by addressing the limitations of existing CNN and Transformer-based models. By combining the hierarchical feature extraction capabilities of CNNs with the long-range dependency modeling of Transformers, MCDAF-Net offers a robust and efficient solution for medical image analysis. The integration of AMSM and CDCM not only improves the model’s ability to handle scale variability and channel redundancy but also enhances its interpretability and generalizability. This makes MCDAF-Net particularly suitable for clinical applications, where accuracy and efficiency are paramount. Furthermore, the model’s performance on public datasets demonstrates its potential to set new standards in WBC detection, paving the way for future research in medical image analysis.
The rest of the paper is organized as follows: In section “Methods”, the MCDAF-Net is introduced. Section “Experiments and results” shows the experimental results on three different data sets. Finally, a summary is presented in section “Discussion”.
Methods
Architecture
In this section, we present the overall structure of MCDAF-Net shown in Fig. 1; it consists of three components. The components are as follows: the backbone network of ResNet5038 for extracting shallow features, the MCDAF module for refining feature extraction, and the Transformer structure for acquiring long-range context dependencies. Within the MCDAF module, there are three submodules: the AMSM, the ADP module which consists of Adaptive Average Pooling and a 1 \(\times\) 1 Depthwise Convolution (DWconv)39, and the CDCM. The AMSM and ADP modules run in parallel, while the CDCM module follows the AMSM in sequence. Concretely, for the ResNet50 output feature map X, we first made four copies of it. Next, we downsize X by the ADP module to obtain the feature map \(X_{1}\). This step aims to reduce the dimensionality of the feature map, thereby reducing the computational complexity while retaining the key information. We utilize the AMSM module to obtain features \(X_{2}\), \(X_{3}\), and \(X_{4}\) with a broader effective receptive field. Subsequently, we obtain the channel-refined features by using the CDCM module. Next, the refined feature maps, along with the position encoding, are fed into the Transformer structure to obtain the bounding box and classification results of the object via the feed-forward neural network. We exploit both a broader effective receptive field and channel feature reconstruction in our MCDAF module, which can adaptively adjust feature weights to capture key features and can be connected to any CNN architecture to improve feature representation.
The proposed model structure consists of a backbone network, a multi-scale feature reconstruction network, and a Transformer structure.
AMSM for features positioning
To take advantage of better positioning features and a broader effective receptive field, we introduce the AMSM, as shown in Fig. 2. This module utilizes different convolutional approaches and combines the self-attention mechanism. The purpose of these convolutional operations is to obtain a larger receptive field. The obtained features are concatenated and used as key features by utilizing the self-attention mechanism22. Specifically, given an input feature map \(\textit{X} \in R^{N \times C \times H \times W}\), where N is the number of batch-size, C is the number of channels, H, W are the height and width axes of feature map. We utilize horizontal, vertical40, and dilated convolutions41 in parallel with convolution kernels of 1\({\times }\)3, 3\({\times }\)1, and then concatenate them as follows:
where X is the feature extracted by ResNet50, \(Conv_{2}\), \(Conv_{3}\), and \(Conv_{5}\) are convolution kernels of 1\({\times }\)3, 3\({\times }\)1, 3\({\times }\)3 respectively. \(Conv_{4,n}\) is then the dilated convolution. We set the dilation rate n \(\in \left\{ 2,3,6\right\}\) for AMSM parallel processing features X and then obtain features \(X_{2}\), \(X_{3}\), and \(X_{4}\). The self-attention mechanism is employed afterward to assign attention weights to the concatenated features. Attention weights represent the importance or relevance of each key feature in the given context, serving as a score that indicates how much focus should be placed on each feature (Attention weights \(\in \left( 0,1\right)\)). The need for this self-attention mechanism arises from its ability to dynamically highlight the most relevant features, effectively improving feature selection and providing a more flexible, context-aware model. By combining these convolutional operations with self-attention, the AMSM is able to capture both local and global contextual information, enhancing the overall model performance.
The architecture of the AMSM consists of two parts: the horizontal-vertical-dilation convolution and the self-attention mechanism.
CDCM for channel-refined features
To leverage channel-wise refined features, we propose the CDCM, as shown in Fig. 3. The module implements a ‘Split Crossover-Fusion Deformation’ strategy that systematically processes channel-wise refined features through three coordinated phases. This architecture works in conjunction with the AMSM module to reconstruct the attention maps generated by AMSM using the channel features from the CDCM. The deformable kernel explicitly models channel interactions, while the proposed module amplifies cross-channel discriminative patterns and suppresses redundant feature responses.
Split crossover
For given multi-scale features \(X_{2}\), \(X_{3}\), and \(X_{4}\) \(\in R^{c \times h \times w}\), we first split them into two channels of size a according to the number of channels \(\frac{c}{2}\). In this way, the multi-scale feature map is split into six features named \(X^1_{2}\), \(X^2_{2}\), \(X^1_{3}\), \(X^2_{3}\), \(X^1_{4}\), and \(X^2_{4}\). Subsequently, we first use 1\({\times }\)1 Pointwise convolution operations to obtain features \(X^2_{2}\), \(X^2_{3}\), and \(X^2_{4}\). Compared to standard convolution, Pconv42 serves to extract and integrate representative information between channels and reduces the number of parameters and computational effort. For the remaining features, no operation is performed, and they are used to ensure the accuracy of the features. Thus we aggregate the features \(X^2_{2}\), \(X^2_{3}\), and \(X^2_{4}\) obtained after the Pconv operation with the split original features \(X^1_{2}\), \(X^1_{3}\), and \(X^1_{4}\) to form the merged representative shape features. Where Cross-Block(N) (with N \(\in \left\{ 1,2,3\right\}\)) represents the operations in Eqs. (5), (6), and (7), respectively. The ‘Split-Crossover’ stage described above can be formulated as follows:
where \(\oplus\) is element-wise summation, ‘Cat’ is concatenation, the split original is represented as feature \(X^1_{2}\) \(\in R^{\frac{c}{2} \times h \times w}\). We split the features for reconstruction, which not only reuses the original features, but also further refines the channel features and enhances the feature representation.
Fusion deformation
For the features obtained from the ‘Split Crossover’ operation, they are residually concatenated with the features from AMSM. Then, weight diffusion is performed again using DeformConv2d43, followed by the fusion of features at three different scales. The combination of these two modules not only fuses features across different scales but also enhances the model’s ability to capture multi-level contextual information, thereby improving its robustness and generalization. The CDCM, as a key component, leverages residual concatenation and weight diffusion through DeformConv2d to strengthen channel interactions, enabling the model to emphasize discriminative features while suppressing redundant ones. This leads to a more efficient and effective representation of the data. The formula is calculated as follows:
where \({X_{i}}^{\prime }\) is the feature obtained by residual concatenation, which is intended to improve model performance and promote feature reuse. The standard convolution operation uses a fixed convolution kernel, which cannot well model complex spatial transformations. Deformable convolution enhances the ability to model spatial transformations by learning additional offsets that allow the convolution kernel to adaptively adjust its position.
The architecture of the CDCM involves two main components. This module separates and cross-fuses multiple feature maps in order to facilitate the reconstruction of channel features.
Integration features
After the AMSM and CDCM modules are performed, the multi-scale feature \({X_{2,3,4}}^{\prime }\) is obtained, which removes the redundancy of channel information. Different from standard convolution, we use the ADP(yellow background in Fig. 1) module containing the \(1 \times 1\) DWconv to obtain the local features \(X_1\). This drastically reduces the model’s parameter count and preserves the network’s ability to learn cross-channel correlations and feature interactions. Meanwhile, we utilize adaptive average pooling to extract the global average information of the input features. Finally, we again use \(1 \times 1\) DWconv to integrate the two features that are fused together to obtain more accurate feature information, which is calculated as follows:
where \({X_1}\) is the feature obtained by ADP module. The combination of adaptive average pooling and DWconv in feature extraction leverages their respective strengths to preserve the spatial information of the inputs, reduce the risk of overfitting, and effectively reduce the number of parameters in the model, thereby enhancing the representation of the features.
In brief, we adopt the AMSM, CDCM, and ADP modules to obtain more comprehensive multi-scale features and eliminate channel redundancy. Overall, AMSM can be deployed standalone or integrated with the CDCM operation. By arranging the AMSM and CDCM modules in a sequential manner and then juxtaposing them with DWconv and adaptive average pooling, the proposed MCDAF module is established.
Loss function
To better predict the classification of WBC and localize the bounding box, in this paper, we use the \(\textit{DIOU}\) loss44 to replace the \(\textit{GIOU}\) loss45 in the baseline model. This change can better address the WBC overlap problem and make the bounding box regression more stable. The loss function of our model consists of two parts: the cross-entropy loss for object classification and the sum of \(\textit{L1}\) Loss and \(\textit{GIOU}\) loss for bounding box regression. All of them are defined as follows:
where \(y_i\) in Eq. (11) denotes the value of i in the ground-truth labels, which takes the values of 0 or 1; \(p_i\) denotes the probability that the model predicts category i; \(\sum\) denotes the summation over all categories. The smaller the value of the cross-entropy loss function, the smaller the discrepancy between the model predictions and the ground-truth labels, i.e., the better the performance of the model. \(\varepsilon\) and \(\theta\) are hyperparameters that can be tuned to the relevant training data. Where \(\hat{y_j}\) in Eq. (12) is the true value, and \(y_j\) is the predicted value. The \(\rho ^{2} (G,F)\) in Eq. (12) represents the squared Euclidean distance between the coordinates of the center points of the ground-truth and predicted boxes, and c is the diagonal distance of the smallest box that contains them. \(\varepsilon\) and \(\theta\) are hyperparameters that can be tuned according to the relevant training data.
Experiments and results
Datasets
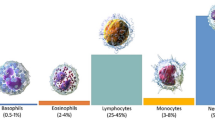
To validate the model, we used four datasets in total, three of which are widely used public datasets: the LISC46, the BCCD47, and the WBCDD48. The RSLI49 is a private dataset. The adopted datasets respectively contain 250, 364, 684, and 288 samples. Both the LISC and WBCDD datasets contain five types of cells: Neutrophil, Monocyte, Eosinophil, Lymphocyte, and Basophil. The various cell types and the number of cells in the dataset are shown in Table 1, where \(\times\) means not categorized. The details of these four datasets are as follows:
RSLI: This dataset consists of hematological images obtained from peripheral blood smears. The images were captured using a Motic Moticam Pro 252A microscope camera with an N800-D motorized autofocus microscope. The dataset contains 288 rapidly stained peripheral blood smear images, including 1170 white blood cells (WBCs)-comprising 1016 neutrophils, 171 monocytes, 123 lymphocytes, and 61 eosinophils. The spatial resolution of each image is \(2048 \times 1536\).
LISC: This dataset is a collection of hematological images obtained from peripheral blood of healthy subjects. Smears were stained by the Gismo-right technique and observations were captured on an Axioskope 40 microscope at 100X magnification using a Sony model SSCDC50AP camera. The spatial resolution of each image is \(720 \times 576\).
BCCD: This dataset contains 364 WBC images taken from peripheral blood and annotated by experts. The smears were stained using the Giemsa-staining technique, and observations were captured using a CCD color camera with 100X conventional light microscopy. Each WBC image extracted from the smear images was annotated by an expert into one of five categories. Each with a resolution of \(640 \times 480\).
WBCDD: This dataset, labeled by a doctor who observes the patient’s blood images through a microscope, contains 684 images with a resolution size of \(4000 \times 3000\).
Data augmentation
To verify the validity of the proposed method, we perform data augmentation on four public datasets to assess the robustness of the trained model. We augmented all training datasets, increasing their size by a factor of ten. This was achieved through techniques such as rotating the images at multiple angles, flipping them symmetrically, adjusting their contrast and brightness, and adding Gaussian noise. To address category imbalance issues, such as the low numbers of eosinophils and basophils in the RSLI dataset, we applied enhancement weighting to balance the classes. The same approach was applied to the other datasets.
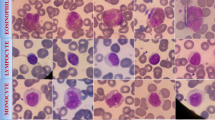
These are the results of the different methods of detecting LISC. Ground Truth is the truth label, (A–D) are the DETR, Deformable-DETR, DINO-DETR and our results respectively, the first to the fifth row are the results of Basophils, Eosinophils, Lymphocytes, Monocytes and Neutrophils each with the ground truth (black) and the detection frame, as well as the category and the confidence level, which red boxes represent missed or incorrect detections.
Implement details
We implemented the model based on the pytorch deep learning framework on Windows10 OS with a 2.50GHz \({\text {Inter}}^{(R)}\) \({\text {Core}}^{(TM)}\) i7 CPU, 64 GB RAM, and an NVIDIA RTX 3090 GPU (with 24 GB memory). The backbone in our model is initialized by a pre-trained ResNet50 network and then fine-tuned in its weights using migration learning. To prevent overfitting, we employed an early stopping strategy, monitoring the validation loss, and halting training if no improvement was observed for a specified number of epochs.
For the four datasets, we trained the model for 300 epochs at a size of 4 batches, with a learning rate of 0.00001 for the backbone network and 0.0001 for the model. the StepLR strategy is used to decay the learning rate to 0.1 times the original value every 200 epochs; the detection network is optimized using the AdamW optimizer, with the hyper-parameters \(\beta _{1}\) and \(\beta _{2}\) set to 0.9 and 0.999, respectively, and the weights decayed to 0.0001. The composition of the training set, validation set, and test set follows a ratio of 8:1:1 for each dataset.
Confidence threshold selection
We compared our method with a range of previous methods on various leukocyte object detection datasets, including Faster R-CNN50, RetinaNet21, SSD51, TE-YOLOF16, DETR23, Deformable-DETR25, DINO-DETR27, YOLOV1152, and D-FINE53. To achieve the best detection results for each method, we tested them at different confidence thresholds. Finally, we set the confidence threshold to 0.5, which achieved the best performance for most methods.
Comparison of other methods
To better compare the performance of each detection model, our comparative results are based on the object detection results provided in the relevant papers and are computed using the same evaluation metrics. We report metrics including \({\text{AP}}_{50}\) (Average Precision at 50), \({\text{AP}}_{75}\) (Average Precision at 75), and AP (Average Precision), for individual cells.
The results of our model on the LISC leukocyte dataset are shown in Table 2. Our method achieves 80.7%, 100% and 99.5% on the LISC dataset for AP, \({\text{AP}}_{50}\), and \({\text{AP}}_{75}\) respectively. The experimental results show that our model can effectively improve the accuracy of leukocyte object detection by using different sets of convolutional operations to expand receptive fields and focus on feature manipulation. Compared to the end-to-end object detection model DETR, our model shows year-on-year growth of 5.2%, 0.8%, and 0.3% on AP, \({\text{AP}}_{50}\) and \({\text{AP}}_{75}\), respectively. The AP, \({\text{AP}}_{50}\), and \({\text{AP}}_{75}\) of the model are improved by 10.4%, 3.9%, and 6.8%, respectively, compared to the conventional one-stage detection model SSD based on multi-level feature extraction. The performance metrics of our model compared to the two-stage object detection model, Faster R-CNN, increase by 4.2% and 2.6% on AP, \({\text{AP}}_{75}\), respectively. In Table 2, AP values were calculated for each category of leukocytes, which helped to assess the performance of the model. In the LISC dataset, we observed significantly higher improvements in the AP values for monocyte and lymphocyte cells compared to other models. These results offer an important reference for further evaluating model performance.
To further evaluate the effectiveness of the model, we conducted the same experiments on two other public datasets, BCCD and WBCDD, and the private dataset RSLI. The results are shown in Tables 2 and 3. Our model performs well on all datasets and essentially obtains the best detection results. In the WBCDD dataset evaluation, our model achieves the best AP metric for the Basophils category, likely due to their unique morphological features, which our multi-scale feature fusion and attention mechanism effectively capture. More importantly, our model outperforms others in overall metrics (AP, AP50, AP75), demonstrating its global robustness and generalizability across all categories, rather than overfitting to a single class. Furthermore, to ensure transparency and fairness in our comparison, Table 4 provides a detailed overview of the hyperparameters for each of the models under consideration.
These are the results of the different methods of detecting RSLI. Ground Truth is the truth label, (A–D) are the DETR, Deformable-DETR, DINO-DETR, and our results respectively, the first to the fifth row are the results of Eosinophils, Lymphocytes, Monocytes, and Neutrophils each with the ground truth (black) and the detection frame, as well as the category and the confidence level, which red boxes represent missed or incorrect detections.
These are the results of the different methods of detecting WBCDD. Ground Truth is the truth label, (A–D) are the DETR, Deformable-DETR, DINO-DETR and our results respectively, the first to the fifth row are the results of Neutrophils, Lymphocytes, Basophils, Monocytes, and Eosinophils, each with the ground truth (black) and the detection frame, as well as the category and the confidence level, which red boxes represent missed or incorrect detections.
Ablation studies
Considering the methodological integrity required for comprehensive model assessment, we strategically conducted a systematic ablation study on all benchmark datasets to rigorously validate the efficacy of individual modules. Although we mitigated the category imbalance problem as much as possible through data enhancement techniques, to ensure the scientific validity of the study, we further merged three datasets with the same number of categories (RSLI, LISC, and WBCDD) and conducted ablation experiments. Such pan-dataset ablation methodology not only ensures the generalizability of our findings but also prevents overfitting to dataset-specific artifacts or annotation biases, thereby establishing a scientifically sound basis for interpreting the technical contributions.
Module effectiveness analysis
The proposed MCDAF module integrates three core components: the AMSM, the CDCM, and the ADP module, complemented by the DIoU loss function to specifically address cell overlap challenges. It should be emphasized that the sequential architecture inherently creates operational dependencies between AMSM and CDCM.Specifically, the CDCM module requires feature maps generated by AMSM as prerequisite inputs, making their combined implementation mandatory rather than optional. As quantitatively demonstrated in Table 5 through comprehensive ablation studies across all benchmark datasets, each constituent module exhibits statistically significant performance contributions. Notably, the progressive integration of these components systematically enhances detection accuracy, with the complete MCDAF configuration achieving optimal performance metrics, thereby confirming both the individual efficacy and synergistic value of our architectural design.
In particular, although the BCCD dataset has more detailed annotations covering white blood cells and various blood cells compared to other datasets, this breadth brings another challenge: the close arrangement of different blood cell types often leads to frequent object adhesion and occlusion, which increases the complexity of analysis. Therefore, in Table 5, the DIOU loss function used to solve cell overlap is of great help to it.
Analysis on dilation rate
To explore the effect of different dilation rates on the AMSM module, we gradually adjusted its size during training on the WBCDD dataset from 2, 3, and 4 to 4, 5, and 6 to compare the AP, AP50, and AP75 metrics, as shown in Table 6. We find that these metrics perform best when the dilation rate is 2, 3, and 6. This superiority is largely attributed to the power of the Transformer structure in understanding context, while the local information we need is complemented by AMSM. However, a smaller receptive field may affect the ability to pinpoint regions of interest, which is the reason why dilation rates of 2, 3, and 6 perform better than dilation rates of 2, 3, and 4.
These are the results of the different methods of detecting BCCD. Ground Truth is the truth label, (A–D) are the DETR, Deformable-DETR ,DINO-DETR and our results respectively, the first to the fifth row are the results of Basophils, Eosinophils, Lymphocytes, Monocytes and Neutrophils each with the ground truth (black) and the detection frame, as well as the category and the confidence level, which red boxes represent missed or incorrect detections.
Model visualization analysis
In order to visually and powerfully highlight the accuracy and usefulness of the model predictions, we chose competitive models with Transformer structures for comparative visualisation. Figures 4, 5, 6, and 7 carefully present the cell categories and their locations predicted by the model, while superimposing the original images from the LISC, RSLI, WBCDD, and BCCD datasets and their ground truth and bounding boxes. Specifically, the red boxes in the figures clearly mark the ground truth, while the differently colored bounding boxes vividly illustrate the predictions made by our model and their corresponding confidence levels. These prediction bounding boxes accurately correspond to the detected cells, demonstrating the model’s ability to recognize and localize leukocyte images with high accuracy. Regarding the presentation of the BCCD dataset in Fig. 7, the original BCCD dataset comprises three categories: ‘Platelets’, ‘RBC’, and ‘WBC’. Given that the number of ‘RBC’ categories is significantly large, labelling each of them would impair the visualization. Therefore, we have chosen to display only the visualization results for the ‘RBC’ and ‘WBC’ categories.
The detailed display in the figure clearly shows that our model exhibits high prediction confidence and near-perfect positional accuracy in predicting these five key leukocyte categories. This not only confirms the model’s remarkable validity but also underscores its potential and value in medical image analysis, disease diagnosis, and other fields.
Discussion
This paper introduces a WBC detection method aimed at improving cell classification by integrating multi-scale features and eliminating channel feature redundancy. The method’s good classification performance is also demonstrated through extensive experiments. However, there are still two aspects that require further discussion.
Integration of multi-scale features
The proposed method integrates multi-scale features to enhance WBC detection by capturing both local details (e.g., texture, morphology) and global context (e.g., shape, spatial relationships). Local details help identify microstructural features, while global context provides broader spatial information, ensuring more accurate detection. This approach is particularly effective for WBC, which have significant variability in size, morphology, and structure, and are susceptible to background clutter.
Unlike traditional Transformer-based methods that focus on long-range dependencies, our method emphasizes the fusion of multi-scale information, extracting richer features at different scales. This allows us to better handle complex backgrounds and improve both detection and classification accuracy, especially in the presence of significant background noise.
Elimination of channel feature redundancy
In this paper, we propose a module designed to address the problem of channel feature redundancy by reducing redundancy through the interaction of multi-scale features. While multi-scale features are effective in enhancing local information extraction in Transformer-based models, they tend to introduce a large number of redundant features, which can be challenging to manage in high-dimensional feature spaces. To address this, we incorporate the CDCM module and sequentially integrate it with the multi-scale feature extraction module. This not only improves computational efficiency but also enhances the discriminative power of the model by focusing on the most salient features for WBC identification. This approach maintains the model’s lightness and versatility, effectively managing feature redundancy without compromising performance, especially in complex, high-dimensional feature spaces.
Conclusions
In this paper, we introduce MCDAF-Net, a novel architecture designed to enhance receptive field efficiency through the integration of multi-scale expansion convolutions and horizontal-vertical convolutions. This design not only broadens the receptive field but also preserves detailed local information. The incorporation of the self-attention mechanism further optimizes feature extraction by selectively focusing on salient regions, thereby improving the precision of feature representation. Additionally, the Channel Dependency and Correlation Modulation (CDCM) module effectively reconstructs channel features, significantly reducing redundancy and enhancing feature distinctiveness. The MCDAF module, being plug-and-play, offers unparalleled flexibility, allowing seamless integration into various convolutional operations without modifying the underlying network structure. To tackle the prevalent issue of cell overlap in leukocyte detection, we employ the DIOU loss function, which markedly improves detection accuracy. Our model’s robust performance across multiple datasets-LISC, BCCD, WBCDD, and the RSLI private dataset-demonstrates its practical applicability.
Future research will focus on two primary directions. First, despite our efforts to reduce the number of parameters, the current model still has a substantial parameter count. Therefore, we plan to develop lightweight variants of our model to enhance computational efficiency and scalability. Second, when integrating the attention mechanism with dilated convolution in other datasets, the dilation rate often needs to be adjusted manually. To address this, we aim to develop adaptive modules that can automatically adjust the dilation rate based on the dataset characteristics, thereby improving the model’s adaptability and generalization ability.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Ware, A. D. The complete blood count and white blood cell differential. In Contemporary Practice in Clinical Chemistry 429–444 (Elsevier, 2020).
Hegde, R. B., Prasad, K., Hebbar, H. & Sandhya, I. Peripheral blood smear analysis using image processing approach for diagnostic purposes: A review. Biocybern. Biomed. Eng. 38, 467–480 (2018).
Geng, L. et al. White blood cell detection and segmentation combined with the channel space weighted feature pyramid networks. J. Comput.-Aid. Des. Comput. Graph. 33, 1418–1427 (2021).
He, K., Gkioxari, G., Dollár, P. & Girshick, R. Mask r-cnn. In Proceedings of the IEEE International Conference on Computer Vision, 2961–2969 (2017).
Zheng, X. et al. White blood cell detection using saliency detection and centernet: A two-stage approach. J. Biophoton. 16, e202200174 (2023).
SivaRao, B. S. S. & Rao, B. S. EfficientNet-XGBoost: An effective white-blood-cell segmentation and classification framework. Nano Biomed. Eng. 15, 126–135 (2023).
Islam, O., Assaduzzaman, M. & Hasan, M. Z. An explainable AI-based blood cell classification using optimized convolutional neural network. J. Pathol. Inform. 15, 100389 (2024).
Kouzehkanan, Z. M. et al. A large dataset of white blood cells containing cell locations and types, along with segmented nuclei and cytoplasm. Sci. Rep. 12, 1123 (2022).
Hou, Q., Zhou, D. & Feng, J. Coordinate attention for efficient mobile network design. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 13713–13722 (2021).
Kanavos, A., Papadimitriou, O., Al-Hussaeni, K., Maragoudakis, M. & Karamitsos, I. Advanced convolutional neural networks for precise white blood cell subtype classification in medical diagnostics. Electronics 13, 2818 (2024).
Yao, X., Sun, K., Bu, X., Zhao, C. & Jin, Y. Classification of white blood cells using weighted optimized deformable convolutional neural networks. Artif. Cells Nanomed. Biotechnol. 49, 147–155 (2021).
Tan, C. et al. A survey on deep transfer learning. In Artificial Neural Networks and Machine Learning–ICANN 2018: 27th International Conference on Artificial Neural Networks, Rhodes, Greece, October 4–7, 2018, Proceedings, Part III 27, 270–279 (Springer, 2018).
Huang, S., Lu, Z., Cheng, R. & He, C. FaPN: Feature-aligned pyramid network for dense image prediction. In Proceedings of the IEEE/CVF International Conference on Computer Vision, 864–873 (2021).
Li, X. et al. Detection of cervical cancer cells in whole slide images using deformable and global context aware faster RCNN-FPN. Curr. Oncol. 28, 3585–3601 (2021).
Lin, T.-Y. et al. Feature pyramid networks for object detection. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 2117–2125 (2017).
Xu, F., Li, X., Yang, H., Wang, Y. & Xiang, W. TE-YOLOF: Tiny and efficient YOLOF for blood cell detection. Biomed. Signal Process. Control 73, 103416 (2022).
Tan, M. & Le, Q. EfficientNet: Rethinking model scaling for convolutional neural networks. In International Conference on Machine Learning, 6105–6114 (PMLR, 2019).
Han, Z. et al. One-stage and lightweight CNN detection approach with attention: Application to WBC detection of microscopic images. Comput. Biol. Med. 154, 106606 (2023).
Wang, J., Sun, Z., Guo, P. & Zhang, L. Improved leukocyte detection algorithm of YOLOv5. Comput. Eng. Appl. 58, 134–142 (2022).
Polejowska, A., Sobotka, M., Kalinowski, M., Kordowski, M. & Neumann, T. Impact of visual image quality on lymphocyte detection using YOLOv5 and RetinaNet algorithms. In Polish Conference on Biocybernetics and Biomedical Engineering, 41–53 (Springer, 2023).
Lin, T. Focal loss for dense object detection. arXiv preprint arXiv:1708.02002 (2017).
Vaswani, A. et al. Attention is all you need. Adv. Neural Inf. Process. Syst. 30 (2017).
Carion, N. et al. End-to-end object detection with transformers. In European Conference on Computer Vision, 213–229 (Springer, 2020).
Lin, T.-Y. et al. Microsoft COCO: Common objects in context (2015).
Zhu, X. et al. Deformable DETR: Deformable transformers for end-to-end object detection. arXiv preprint arXiv:2010.04159 (2020).
Wang, T., Yuan, L., Chen, Y., Feng, J. & Yan, S. PnP-DETR: Towards efficient visual analysis with transformers. In Proceedings of the IEEE/CVF International Conference on Computer Vision, 4661–4670 (2021).
Zhang, H. et al. DINO: DETR with improved denoising anchor boxes for end-to-end object detection. arXiv preprint arXiv:2203.03605 (2022).
Chen, S., Lu, S., Wang, S., Ni, Y. & Zhang, Y. Shifted window vision transformer for blood cell classification. Electronics 12, 2442 (2023).
Li, M. et al. Peripheral blood leukocyte detection based on an improved detection transformer algorithm. Sensors 23, 7226 (2023).
Dipto, S. M. et al. An XAI integrated identification system of white blood cell type using variants of vision transformer. In Proceedings of the Second International Conference on Innovations in Computing Research (ICR’23), 303–315 (Springer, 2023).
Katar, O. & Yildirim, O. An explainable vision transformer model based white blood cells classification and localization. Diagnostics 13, 2459 (2023).
Marzahl, C., Aubreville, M., Voigt, J. & Maier, A. Classification of leukemic b-lymphoblast cells from blood smear microscopic images with an attention-based deep learning method and advanced augmentation techniques. In ISBI 2019 C-NMC Challenge: Classification in Cancer Cell Imaging: Select Proceedings, 13–22 (Springer, 2019).
Huang, P. et al. Attention-aware residual network based manifold learning for white blood cells classification. IEEE J. Biomed. Health Inform. 25, 1206–1214 (2020).
Nugraha, S. J. A. & Erfianto, B. White blood cell detection using YOLOv8 integration with DETR to improve accuracy. Sinkron: jurnal dan penelitian teknik informatika 7, 1908–1916 (2023).
Tarimo, S. A. et al. WBC YOLO-ViT: 2 way-2 stage white blood cell detection and classification with a combination of YOLOv5 and vision transformer. Comput. Biol. Med. 169, 107875 (2024).
Zhang, D., Bu, Y., Chen, Q., Cai, S. & Zhang, Y. TW-YOLO: An innovative blood cell detection model based on multi-scale feature fusion. Sensors 24, 6168 (2024).
Bayat, N., Davey, D. D., Coathup, M. & Park, J.-H. White blood cell classification using multi-attention data augmentation and regularization. Big Data Cogn. Comput. 6, 122 (2022).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 770–778 (2016).
Chollet, F. Xception: Deep learning with depthwise separable convolutions. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 1251–1258 (2017).
Ding, X., Guo, Y., Ding, G. & Han, J. ACNet: Strengthening the kernel skeletons for powerful cnn via asymmetric convolution blocks. In Proceedings of the IEEE/CVF International Conference on Computer Vision, 1911–1920 (2019).
Yu, F., Koltun, V. & Funkhouser, T. Dilated residual networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 472–480 (2017).
Hua, B.-S., Tran, M.-K. & Yeung, S.-K. Pointwise convolutional neural networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 984–993 (2018).
Zhu, X., Hu, H., Lin, S. & Dai, J. Deformable ConvNets v2: More deformable, better results. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 9308–9316 (2019).
Zheng, Z. et al. Distance-IoU loss: Faster and better learning for bounding box regression. In Proceedings of the AAAI Conference on Artificial Intelligence, vol. 34, 12993–13000 (2020).
Rezatofighi, H. et al. Generalized intersection over union: A metric and a loss for bounding box regression. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 658–666 (2019).
Rezatofighi, S. H. & Soltanian-Zadeh, H. Automatic recognition of five types of white blood cells in peripheral blood. Comput. Med. Imaging Graph. 35, 333–343 (2011).
Vatathanavaro, S., Tungjitnob, S. & Pasupa, K. White blood cell classification: A comparison between vgg16 and resnet50 model. In The 6th Joint Symposium on Computational Intelligence (JSCI6), 4–5 (2018).
Chen, Y. et al. Accurate leukocyte detection based on deformable-DETR and multi-level feature fusion for aiding diagnosis of blood diseases. Comput. Biol. Med. 170, 107917 (2024).
Zheng, X., Wang, Y., Wang, G. & Chen, Z. A novel algorithm based on visual saliency attention for localization and segmentation in rapidly-stained leukocyte images. Micron 56, 17–28 (2014).
Ren, S., He, K., Girshick, R. & Sun, J. Faster R-CNN: Towards real-time object detection with region proposal networks. Adv. Neural Inf. Process. Syst. 28 (2015).
Liu, W. et al. SSD: Single shot multibox detector. In Computer Vision–ECCV 2016: 14th European Conference, Amsterdam, The Netherlands, October 11–14, 2016, Proceedings, Part I 14, 21–37 (Springer, 2016).
Khanam, R. & Hussain, M. YOLOv11: An overview of the key architectural enhancements. arXiv preprint arXiv:2410.17725 (2024).
Peng, Y. et al. D-fine: Redefine regression task in DETRs as fine-grained distribution refinement. arxiv 2024. arXiv preprint arXiv:2410.13842
Acknowledgements
This work was supported in part by the National Natural Science Foundation of China under Grants 62171002, 62302014, in part by the Scientific Research Foundation of Education Department of Anhui Province of China under Grant 2023AH050489, 2023AH050492.
Author information
Authors and Affiliations
Contributions
Z.X. and X.Q. wrote the main manuscript text, Z.S and Z.L. performed the data analysis, L.D. and Z.L. performed the data analysis. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zheng, X., Xu, Q., Zheng, S. et al. Multiscale deformed attention networks for white blood cell detection. Sci Rep 15, 14591 (2025). https://doi.org/10.1038/s41598-025-99165-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-99165-8