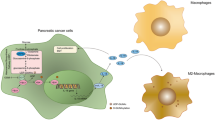
Abstract
GULP PTB domain containing engulfment adaptor 1 (GULP1) protein is involved in regulating biological processes such as endocytosis and apoptosis. The function of GULP1 in cancer, however, has been the subject of fewer studies; its importance as a potential prognostic factor in pancreatic cancer is still uncertain. Thus, the purpose of this work was to investigate GULP1’s immunologic and oncogenic activities in a range of malignancies, as well as any potential relevance to pancreatic cancer. Using multiple bioinformatic databases, GULP1 expression, prognostic significance, mutation status, methylation and phosphorylation levels, biological functions, immune cell infiltration and immunotherapeutic responses and drug sensitivity were comprehensively assessed in pan-cancer and functionally validated in pancreatic cancer. The results revealed that GULP1 was differentially expressed in most tumours and correlated with poor prognostic indicators in most tumours, which may be related to the fact that GULP1 is involved in the regulation of apoptotic pathways. In addition, the differential expression of GULP1 was linked to immune cell infiltration levels, immunotherapy response, and chemotherapy resistance. GULP1 could hinder the body’s ability to fight tumors and respond to immunotherapy by promoting the accumulation of immune cells and suppressing the activity of cytotoxic T lymphocytes. In pancreatic cancer, down-regulation of GULP1 expression inhibits proliferation, invasion and migration of pancreatic cancer cells. These phenotypic changes may be achieved by regulating HIPPO, mTOR, and RTK signaling pathways. Taken together, it makes sense to think that GULP1 could be a biomarker for immunotherapy and prognosis in pan- and pancreatic cancer.
Similar content being viewed by others
Introduction
Cancer is a serious global disease that causes enormous losses to both human health and socioeconomics1. During 2022, there were approximately 20 million new instances of cancer and over 9.7 million cancer-related deaths worldwide2. Cancer incidence and mortality rates are still continuing to rise due to a variety of factors such as population aging, lifestyle changes, environment, and diet3. The amount of new cancer cases globally is projected to exceed 35 million during the mid-century, a 77% jump from 20223. And there are notable variations in incidence and death rates among different regions and types of cancer4,5. Cancer has become one of the primary reasons for premature death worldwide and an important challenge to be overcome to extend human life span6,7. Therefore, identification of valuable cancer genes is essential to reveal the underlying mechanisms of different tumor formation and evolution.
GULP1 also known as CED6 or GULP, is a protein-coding gene located at 2q32.3-q33 and encodes its protein from 304 amino acids8. The GULP1 protein is required for phagocytosis of apoptotic cells by phagocytosis articulating protein8. GULP1 is widely expressed in many cell types and is involved in regulating cellular glucose, lipid, and cholesterol transport9,10. It may also play a critical role in the internalization and endosomal trafficking of multiple LRP1 ligands (e.g., PSAP) and increase the level of GTP-bound ARF6 in cells11. Research points out that GULP1 abnormalities are strongly linked to the development of ovarian cancer12, medulloblastoma13, and bladder cancer14,15. In bladder cancer, Masamichi Hayashi et al. found that GULP1 knockdown promotes tumor cell proliferation in vitro and in vivo and increases resistance to cisplatin drugs15. And low or no expression of GULP1 was also observed in most cases of cisplatin-naïve bladder cancer15. In addition, The varying levels of GULP1 expression were connected to lymph node metastasis in bladder cancer14. Methylation silencing of GULP1 was associated with ovarian cancer prognosis12. These results suggest that GULP1 may have potential in cancer prognosis and treatment. However, the biological functions and immunotherapeutic potential of GULP1 in different tumors are currently understudied.
With the continuous improvement of multi-omics data in cancer, comprehensive pan-cancer analysis will be more helpful to fully understand the molecular mechanisms and predictive value of oncogenes, thus maximizing clinical diagnosis and therapeutic choices16,17. Therefore, in this study, we comprehensively assessed GULP1 expression, prognosis, mutation, methylation, phosphorylation, biological function, immune microenvironment, immunotherapeutic response, and drug sensitivity in pan-cancer through multiple bioinformatic databases. Subsequently, we performed functional validation in pancreatic cancer. The results of our research demonstrated the value of GULP1 in pan-cancer prognosis and immunotherapy, which is expected to be a powerful biomarker for cancer.
Materials and methods
Expression analysis
The TCGA and GTEx pan-cancer transcriptome datasets were obtained from the UCSC (http://xena.ucsc.edu) database18, and the data were log2(x + 0.001) transformed using the Sangerbox (http://SangerBox.com)19 online tool to exclude cancer types with less than three specimens and to visualize the normal sample and tumor samples between GULP1 mRNA expression differences. Expression data in pancreatic cancer cell lines were obtained from the CCLE database (https://sites.broadinstitute.org/ccle/)20. Based on the CPTAC dataset, GULP1 total and phosphorylated protein expression levels in pan-cancer were analyzed by the UALCAN database (http://ualcan.path.uab.edu/)21. GULP1 immunohistochemical images and subcellular localization were obtained from the Human Protein Atlas (HPA, https://www.proteinatlas.org/)22. The relationship between GULP1 expression and molecular subtypes, pathological staging and grading was investigated by TISIDB online website (http://cis.hku.hk/TISIDB/index.php)23.
Survival analysis
Based on the TCGA and GTEx datasets, samples with a prognostic follow-up shorter than 30 days and different types of cancer with less than 10 samples were excluded. A univariate COX risk regression model was developed using the R package “survival” as a means to assess the relationship between GULP1 expression and prognostic characteristics (overall survival (OS), progression-free interval (PFI), disease-specific survival (DSS), and disease-free interval (DFI)) of patients with pan-cancer, and plotted the Kaplan-Meier curves. The assessment metrics included hazard ratios (HR), 95% confidence intervals, and P-values, and the significance threshold was observed at a significance level of P < 0.05.
Gene mutation and methylation analysis
Mutation types, frequencies, counts, mutation sites and three-dimensional (3D) structures of GULP1 gene in TCGA PanCancer Atlas Studies were obtained through cBioPortal online website (https://www.cbioportal.org/)24. Mutation and expression analysis of GULP1 gene in PanCancer was performed using Sangerbox web tool. Copy Number Variation (CNV) was normalized by GSCA online website (https://guolab.wchscu.cn/GSCA/)25. The correlation heatmap of GULP1 expression with Mismatch Repair (MMR) genes was visualized using the R package “ggplot2”.
GULP1 promoter methylation levels were explored in the TCGA dataset using the UALCAN website. Overall survival of GULP1 CpG loci in various types of cancer in the TCGA database was analyzed through the MethSurv website (http://biit.cs.ut.ee/methsurv/)26. Transcriptome data of 44 m1A (10), m5C (13), and m6A (21) modified marker genes were normalized through the Sangerbox online website and correlation heatmaps of GULP1 and the three classes of RNA methylation-modified marker genes were drawn.
Single cell analysis
The correlation between GULP1 expression at the single cell level and various functionality modes of multiple tumor cells was explored using the CancerSEA (http://biocc.hrbmu.edu.cn/CancerSEA/) database27. The T-SNE plots displayed the expression profiles of GULP1 in single cells. In addition, we used the TISCH web tool (http://tisch.comp-genomics.org/documentation/)28 to quantify and visualize GULP1 expression in different cell types.
Enrichment analysis
We obtained the top 50 experimentally determined interactions networks of GULP1-binding proteins from the STRING online website (https://string-db.org/)29. The top 100 genes corresponding to GULP1 expression were obtained from the GEPIA 2.0 online tool (http://gepia2.cancer-pku.cn/)30 and selected genes were subjected to Pearson correlation analysis. Then, GO (Gene Ontology) enrichment analysis and KEGG (Kyoto Encyclopedia of Genes and Genomes) signaling pathway analysis were performed on the top 100 GULP1-related genes to investigate the possible functions and pathways of GULP1 in tumors. Visualization was done by R packages “ggplot2” and “clusterProfiler”. p < 0.05 was considered to be statistically significant.
Immunization and drug sensitivity analysis
The correlation between GULP1 expression and immune cell infiltration was explored through the TIMER 2.0 online website (http://timer.cistrome.org/)31, and stromal, immune, and estimate scores were computed using the R package “ESTIMATE”. The expression data of marker genes of five types of immune pathways(chemokine, receptor, MHC, immunoinhibitor, immunostimulator)were obtained from the pan-cancer dataset, and each expression value was transformed by log2(x + 0.001), and the pearson correlation analysis of the GULP1 and the five types of immune pathways was performed next. The next step was to analyze the pearson correlation between GULP1 and the marker genes of the five immune pathways. The relationship between GULP1 expression and TMB, MSI, Purity, HRD was visualized by Sangerbox tool. The association of GULP1 with immune subtypes was performed through TISIDB website. The association between GULP1 and CTL dysfunction was assessed through the TIDE website (http://tide.dfci.harvard.edu)32. GULP1 prediction of cancer immunotherapy efficacy was performed through the TISMO (http://tismo.cistrome.org)33 and ROC Plotter (https://www.rocplot.org/)34 online sites. Drug sensitivity data were obtained from the CTRP database (https://portals.broadinstitute.org/ctrp/)35 and the GDSC (https://www.cancerrxgene.org/)36 database and represented with the R package “ggplot2”.
Cell culture and transfection
The human pancreatic cancer cell lines ASPC1 and PANC1 (Servicebio, China) were characterized by short tandem repeat (STR). The cells were cultured in a cell culture incubator at 37 °C with 5% CO2 using DMEM medium supplemented with 10% fetal bovine serum (FBS; Gibco, USA). The vectors si-GULP1 were transfected into ASPC1 and PANC1 cells by Lipofectamine 2000 (Invitrogen, USA) according to the instructions. The three-in-one siRNA sequences were: siRNA-1: GAGATGCTGTAAGGAAACTAAAGTT; siRNA-2: ACACAATTGCCAGCTTCATAGAATA; siRNA-3: AAGTGTGCTGAAGAGAGATCACTTTAA;
Western blot
Total proteins in the cell lines were extracted using the protease inhibitor PMSF and RIPA lysate, and protein concentration was determined by bicinchoninic acid (BCA) kit (Beyotime, China). Proteins were transferred to PVDF membranes using 10% SDS-PAGE, sealed with 5% skimmed milk powder, and incubated overnight at 4 °C with the indicated primary antibodies, followed by 1 h of incubation with the secondary antibodies. Finally, after washing three times with TBST developed using an ECL kit (Invitrogen, USA).
Cell proliferation, migration and invasion assays
CCK-8 kit (Servicebio, China) was used for cell proliferation assay. Cells were inoculated at 4000/well in 96-well plates and incubated in an incubator. CCK-8 reagent was added at various time intervals and the absorbance at 450 nm was determined to assess the cell proliferation ability.
The wound healing assay was utilized to evaluate cell migration capability. Cells were inoculated in 6-well plates and incubated in an incubator, and when the cells were fused to 80%, they were scratched vertically with a lance tip, washed with phosphate buffer solution ( PBS), and the scratched area was photographed under a microscope at different time points.
Transwell assay was used to detect cell invasion. Cell suspensions were prepared with serum-free medium, and the suspensions containing 5 × 104 cells were inoculated in transwell chambers lined with matrix adhesive, and the lower chamber was added with medium containing 15% FBS, and the culture medium in the chambers was discarded after 24 h of incubation, washed with PBS, and fixed with 4% paraformaldehyde, and the non-migrated cells were wiped away from the chambers with a swab, and the cells were stained with 0.1% crystal violet and then imaged beneath the microscope.
Statistical analysis
Wilcoxon Rank Sum and Signed Rank Tests were used to detect differences in GULP1 expression. Logrank Tests were used for prognostic analysis. Correlation analysis between the two groups was performed using Spearman or Pearson. The analysis of experimental data was performed by GraphPad Prism 9. All tests were two-sided and were considered statistically significant when P < 0.05.
Results
Expression and subcellular localization of GULP1
See supplementary Table 1 for tumor abbreviations. We first analyzed GULP1 expression in normal tissues and found that GULP1 mRNA (Fig. 1a) and protein (Fig. 1f) were highly expressed in normal tissues of the epididymis. Next, we analyzed GULP1 mRNA expression in the TCGA dataset and showed that GULP1 mRNA was differentially expressed in 15 cancers (Supplementary Fig. 1a). Due to the limited amount of normal tissue samples in TCGA, we incorporated the TCGA_GTEx dataset, and the analysis indicated that GULP1 showed differential expression in 29 tumors, including PAAD (Fig. 1b). Further analysis revealed that GULP1 expression correlated with the molecular subtypes of BRCA, COAD, ESCA, HNSC, KIRP, LGG, LIHC, LUSC, PCPG, PRAD, STAD, and UCEC (Supplementary Fig. 1b, c). The GULP1 expression level was also positively correlated with the HNSC grading, and negatively correlated with the KIRC, OV, and UCEC grading (Fig. 1d, e). Higher GULP1 expression was associated with later pathologic staging in BLCA, COAD, ESCA, LIHC, PAAD, STAD, and TGCT, and the opposite was true in KIRC, UCEC (Supplementary Fig. 1d, e). We then evaluated GULP1 protein expression in cancers using the CPTAC database, which showed that total GULP1 protein was significantly under-expressed in BRCA, COAD, OV, KIRC, UCEC, LUAD, HNSC, and LIHC, and significantly over-expressed in PAAD (Fig. 1c). The immunohistochemistry (IHC) findings in the HPA database were in line with the same results (Supplementary Fig. 1f).
Differential expression and subcellular localization of GULP1. GULP1 mRNA expression in normal tissues (a). Differential expression of GULP1 mRNA between different tumors and normal tissues based on the TCGA_GTEx dataset, *p < 0.05, **p < 0.01, ****p < 0.0001, -, not significant (b). Differences in the expression of total GULP1 protein between cancerous and normal tissues (c). Based on CPTAC database. correlation between GULP1 expression level and tumor grade (d,e). Representative immunohistochemistry (IHC) images of GULP1 total protein in normal epididymal tissue (f).
Prognostic value of GULP1 in Pan-Cancer
We examined how GULP1 expression impacts the prognosis of cancer patients. Cox regression analysis identified that high GULP1 expression was a role as a risk factor for shorter OS, DSS, PFI, and short DFI in seven cancers, and low GULP1 expression was a risk factor for short OS in two cancers, short DSS in three cancers, and short PFI in four cancers (Fig. 2). Further Kaplan-Meier survival curves showed that GULP1 expression was associated with OS in 17 tumors. In BLCA, COAD, COADREAD, GBMLGG, KICH, LIHC, LUAD, OV, PAAD, STAD, and STES, high GULP1 expression implied poor OS, whereas in CHOL, GBM, KIPAN, KIRC, SARC, and WT, high GULP1 expression implied better OS (Supplementary Fig. 2 ). GULP1 expression was associated with DSS in 17 tumors. high GULP1 expression was associated with shorter DSS in BLCA, COAD, COADREAD, GBMLGG, KICH, LIHC, OV, PAAD, PCPG, STAD and STES, whereas high GULP1 expression was associated with longer DSS in CHOL, KIPAN, KIRC, THYM and longer DSS in UCS. (Supplementary Fig. 3). GULP1 expression was significantly associated with PFI in 18 tumors. upregulation of GULP1 resulted in shorter PFI in ACC, BLCA, CESC, COAD, COADREAD, GBMLGG, KICH, LIHC, LUAD, PAAD, STAD, STES, and CESC patients, and longer PFI in KIPAN, KIRC, THYM, CESC and UCS patients had prolonged PFI (Supplementary Fig. 4). GULP1 expression correlated with DFI in 14 tumors. In ACC, CESC, HNSC, KICH, LIHC, PAAD, STAD, STES and TGCT, GULP1 expression was negatively correlated with DFI, while it was positively correlated in COAD, GBMLGG, LGG, MESO and UCS (Supplementary Fig. 5a). The outcomes listed above suggest that GULP1 may be a potential prognostic marker gene for some cancers, such as COAD, GBMLGG, KICH, LIHC, PAAD, STAD, STES (Supplementary Fig. 5b).
Univariate COX regression analysis of GULP1 expression level and prognostic characteristics of pan-cancer patients with prognostic indicators including overall survival (OS), disease-specific survival (DSS), progression-free interval (PFI), and disease-free interval (DFI).
Mutation analysis of the GULP1 gene in pan-cancer
In the pan-cancer analysis, we utilized the cBioPortal database to investigate the gene mutations of GULP1. The results indicated that the top three cancers were UCEC, OV, and SARC, with mutations predominating in UCEC, amplifications occurring most frequently in OV, and more mutation types in SARC (Fig. 3a). We further explored the mutational landscape of GULP1 in these three cancers, and showed that the top 15 oncogenes (e.g., ATRX) and anti-oncogenes (e.g., TP53 and PTEN) differed in mutation frequency between the GULP1 high- and low-expression groups (Supplementary Fig. 6a). Notably, missense mutations were the most common type of mutation in the GULP1 gene, and S130C/I/R changes were found in the PID structural domain (Fig. 3b). Figure 3c shows the S130C/I/R alterations in the 3D configuration of the GULP1 protein. Supplementary Fig. 6b demonstrates the mutation counts of GULP1 in different cancers. We explored the CNV of GULP1 in depth. the top three cancers in terms of GULP1 CNV percentage were OV, UCS and TGCT (Fig. 3d). GULP1’s CNV and mRNA expression were found to be positively correlated in KIRP, STAD, HNSC, ESCA, SKCM, OV, LGG, LUAD, PAAD, BRCA and LUSC (Supplementary Fig. 6c). GULP1 CNV correlated with OS, PFS, DSS, and DFI in UCEC, ACC, MESO, SARC, TGCT, COAD, DLBC, and PAAD (Supplementary Fig. 6d). Figure 3e demonstrates the results of the detailed Kaplan-Meier survival analysis. We also investigated the variations in GULP1 gene expression among the various mutation subgroups, resulting in statistically significant differences observed in eight tumors (Supplementary Fig. 6f). Finally, we assessed the relationship between GULP1 expression and MMRs genes in pan-cancer. GULP1 expression was noted to be remarkably positively correlated with MMRs gene expression in most cancers, and GULP1 may promote cancer cell growth by positively regulating MMRs gene expression (Supplementary Fig. 6e).
GULP1 mutations in cancer. The cBioPortal was used to show the frequency of different mutation types (a) and mutation sites (b) altered in the GULP1 gene in pan-cancer. S130C/I/R mutation site was shown in the 3D protein structure of GULP1 (c). Percentage of GULP1 CNV in cancer (d). Kaplan-Meier curves of GULP1 CNV with OS, PFS, DSS, and DFI in cancer patients, p < 0.05 (e).
Promoter methylation of GULP1 in pan-cancer
DNA methylation has been shown to be involved in tumor development, and aberrant methylation is one of the characteristics of cancer37. We explored the GULP1 methylation values between pan-cancer tissues and paired normal tissues through the UALCAN database. The results led to the conclusion that methylation levels of the GULP1 promoter were higher in BLCA, BRCA, CESC, UCEC, HNSC, KIRC, LIHC, LUAD, and LUSC tissues than in normal tissues, whereas GULP1 promoter methylation levels were significantly lower in PRAD, READ, STAD, and COAD tissues (Fig. 4a). DNA methylation is mainly directed at the CpG islands present in regulatory regions like gene promoters or enhancers38. We found that methylation of 29 CpG islands of GULP1 was associated with cancer prognosis through the Methsurv database, with a high proportion of cg13274254 (Supplementary Table 2). cg13274254 methylation level of GULP1 CpG islands was positively correlated with the LAML, BRCA, HNSC, KIRC, LUAD, MESO and SKCM prognosis positively, while it had an adverse impact on the prognosis of KIRP and LGG (Fig. 4b). Notably, GULP1 CpG loci cg13416310, cg16256414, and cg17141115 were positively correlated with the prognosis of PAAD, while GULP1 CpG locus cg09305069 was negatively correlated with the prognosis of PAAD (Supplementary Fig. 7b). In addition, we analyzed the correlation between GULP1 expression and m1A, m5C, and m6A methylation marker genes in pan-cancer. The results showed an overall positive trend of correlation in most cancers. However, a negative correlation was observed in NB and WT (Supplementary Fig. 7a). The results indicate that promoter methylation could influence the transcriptional expression of GULP1, impacting tumor advancement.
GULP1 DNA methylation in pan-cancer. Comparison of promoter methylation levels of GULP1 between pan-cancer and normal tissues by UALCAN (a). Correlation of GULP1 methylation levels at the CG13274254 locus with cancer prognosis (b). Results showing p < 0.05.
Phosphorylation analysis of GULP1 in pan-cancer
Abnormal protein phosphorylation can enhance cancer cell proliferation, invasion and metastasis, and restrain cancer cell apoptosis, leading to cancer development and progression39. Therefore, we compared GULP1 phosphorylation changes between primary tumor tissues and normal tissues by CPTAC database. Figure 5a shows the overall GULP1 phosphorylation. Compared with normal tissues, the phosphorylation levels of S206, OV and T208 in KIRC and PAAD were drastically decreased (Fig. 5b).The phosphorylation levels of S206 and T208 of GULP1 were also significantly reduced in HNSC and LUAD (Fig. 5b). However, in BRCA, the T208 phosphorylation level of GULP1 was increased (Fig. 5b).
GULP1 phosphorylation levels in pan-cancer. Overall GULP1 phosphorylation between primary tumor tissues and normal tissues based on CPTAC database (a). phosphorylation levels of S206 and T208 sites of GULP1 in KIRC, OV, PAAD, HNSC, LUAD and BRCA (b).
Single-cell analysis of GULP1
We analyzed the connections between GULP1 expression and 14 functional states in various cancers at the single-cell level. The results showed that in GBM, SKCM, RB and UVM, GULP1 expression levels were significantly connected with multiple functional states of cancer cells, including Apoptosis, CellCycle, Differentiation, Inflammation, and Proliferation (Fig. 6a). In addition, GULP1 expression was negatively correlated with Angiogenesis and Apoptosis in AML, CellCycle, Differentiation and Quiescence in BRCA, Metastasis in KIRC, Invasion status in OV, and Invasion in AML, the EMT in Glioma, Differentiation in LUAD, and Angiogenesis status in OV were positively correlated (Fig. 6a). Figure 6b, Supplementary Fig. 8a shows the cancer types with correlations above 0.3. T-SNE plots show the single-cell level expression profiles of GULP1 in AML, BRCA, KIRC, LUAD, OV, SKCM, RB, and UVM (Fig. 6c). We also analyzed GULP1 expression in the single-cell dataset by the TISCH database. The heatmap showed the expression levels of GULP1 in immune cells, malignant cells, stromal cells and other functional cell types (Supplementary Fig. 8b). It was found that GULP1 was relatively highly expressed in malignant cells in Glioma and SKCM microenvironments in GSE102130, GSE72056 datasets (Supplementary Fig. 8c). In the GSE146771 Smartseq2 dataset, GULP1 was relatively highly expressed in stromal cells in the CRC microenvironment (Supplementary Fig. 8c).
GULP1 expression at the single cell level. Correlation of GULP1 expression with different functional states of tumor cells at the single-cell level based on CancerSEA website analysis (a,b). GULP1 expression profiles were shown at single cells from AML, BRCA, KIRC, LUAD, OV, SKCM, RB and UVM by T-SNE diagram, colours from light to dark represent low to high expression levels of GULP1 (c).
Functional analysis of GULP1 in cancer
To delve deeper into the potential molecular mechanisms of GULP1 in cancer development, we obtained 50 experimentally confirmed interaction networks of GULP1-binding proteins through the STRING website (Fig. 7a). The top 100 genes associated with GULP1 expression were obtained using the GEPIA2 tool combined with TCGA expression data (Supplementary Table 3). A scatter plot of the correlation of the top 3 genes in pan-cancer was plotted (Fig. 7b). In the majority of tumors, GULP1 expression levels had a favorable relationship with ARFGAP3, FNIP2, HERPUD1, SH3RF1, SYNJ2BP, and UNC13B expression (Fig. 7c). GO and KEGG enrichment analyses pointed out that GULP1 was involved in phagocytosis, apoptotic cell clearance, Rac/Ras protein signal transduction, (plasma) membrane invagination, establishment or maintenance of cell polarity, vascular endothelial growth factor receptor signaling pathway, positive regulation of substrate adhesion-dependent cell spreading biological process (BP) (Fig. 7d), involving cellular components (CC) such as endocytic vesicle, trans-Golgi network, organelle subcompartment, and other cellular components (Fig. 7e), which are involved in cargo/scavenger/low-density lipoprotein particle/lipoprotein particle receptor activity, SH3 domain binding, Rac/Rho/Ras/small GTPase binding, and phospholipid binding, among other molecular functions (MF) (Fig. 7f). KEGG enrichment analysis recommended that GULP1 was linked to Bacterial invasion of epithelial cells, Endocytosis, Fc gamma R-mediated phagocytosis, Endocrine and other factor- regulated calcium reabsorption, Synaptic vesicle cycle, Chemokine signaling pathway, Cholesterol metabolism and other pathways (Fig. 7g).
Functional enrichment analysis. STRING showing the top 50 GULP1 binding protein interaction networks (a). Correlation scatter plot of the top 3 genes associated with GULP1 expression in pan-cancer (b). Heatmap of correlation between GULP1 expression and ARFGAP3, FNIP2, HERPUD1, SH3RF1, SYNJ2BP, UNC13B in pan-cancer, *p < 0.05, **p < 0.01, ***p < 0.001 (c). GO (d–f) and KEGG enrichment (g) of the top 100 GULP1-related genes.
GULP1 is associated with cancer immune infiltration and Immunomodulatory genes
Immune cell infiltration is strongly associated with cancer progression40. Various algorithms in the TIMER2 database assessed the correlation between GULP1 expression and immune cell infiltration in cancers31. Figure 8 displayed a positive correlation between GULP1 expression and the infiltration levels of CAF, Endo, Macrophage, Monocyte, and Neutrophil in various cancers, particularly in BLCA, BRCA, COAD, HNSC, and READ. A negative correlation was observed with the degree of NKT cell infiltration. According to the XCELL algorithm, the infiltration levels of CD4 + T cells, DC were also negatively correlated with GULP1 expression in most cancers. In TGCT, GULP1 expression was negatively correlated with the infiltration level of CD8 + T cells, B cells. Tregs infiltration was negatively correlated with GULP1 in LUAD, UCS, KIRC, and GBM, and positively correlated in THCA. In addition, immunosubtyping results showed that GULP1 upregulation was associated with C1 isoforms (wound healing) in LIHC, HNSC and TGCT, and C2 isoforms (IFN-γdominant) in BLCA. Meanwhile, high GULP1 expression was also associated with C6 isoform (TGF-βdominant) in BRCA and C4 isoform (lymphocyte depleted) in KIRC (Supplementary Fig. 9a, b).
Correlation between GULP1 expression and immune cell infiltration in cancer. Positive correlation (p < 0.05, partial_cor > 0), negative correlation (p < 0.05, partial_cor < 0), no significance (p > 0.05).
We continued to investigate the correlation between GULP1 expression and the 3 immunescores, and the results demonstrated the highest correlation among the different scores (Supplementary Fig. 10a). StromalScore showed that the level of immune cell infiltration in COAD, COADREAD, READ, TGCT, DLBC, BRCA, THYM, BLCA, and UCS increased with high expression of GULP1. The level of GULP1 expression was positively correlated with the immunescores in COAD, COADREAD, READ. EstimateScore also showed a significant positive correlation between GULP1 expression and the level of immune cell infiltration in COAD, COADREAD, READ, BLCA, and DLBC. These results demonstrated the importance of GULP1 in immune infiltration of tumor cells.
We also analyzed the association of GULP1 with immunomodulatory genes including chemokine, receptor, MHC, Immunoinhibitor and Immunostimulator (Supplementary Fig. 9c). The findings demonstrated a positive correlation between GULP1 expression and the majority of immunoregulatory genes in cancers, particularly in READ, COAD, COADREAD, BRCA, BLCA, THCA, and LIHC. While in TGCT, KIPAN, GULP1 expression was negatively correlated with most Immunoinhibitor and Immunostimulator genes. In addition, GULP1 was positively correlated with chemokine genes CXCL12, CX3CL1, CXCL5, and receptor gene CX3CR1 in most cancers. In PAAD, GULP1 expression was significantly positively correlated with the expression of immunoinhibitor genes TGFBR1, CD274, and PDCD1LG2, as well as with immunostimulator genes RAET1E, HHLA2, TNFSF13, TNFSF15, NT5E, TNFSF4, ENTPD1, TNFSF18, MICB, CD80, IL2RA, and TNFRSF9 (Supplementary Fig. 9d). The correlation between GULP1 and immunomodulatory gene expression helps us comprehend intricate dynamics more effectively in the tumor microenvironment.
GULP1-based prediction of immunotherapeutic response and sensitive drugs
Since TMB and MSI are key predictors of immunotherapy efficacy41, we evaluated the correlation between GULP1 expression and pan-cancer TMB and MSI. TMB was positively correlated with GULP1 expression in LAML, THYM, and negatively correlated with GULP1 in LGG, CESC, BRCA, KIPAN, PRAD, HNSC, and KIRC (Supplementary Fig. 10b). GULP1 expression was positively correlated with MSI in KIPAN, KIRC, TGCT, and CHOL, and negatively correlated with MSI in LUAD, HNSC, and DLBC (Supplementary Fig. 10c). Tumor purity is thought to impact the effectiveness of immune checkpoint inhibitor therapy42, and we confirmed that GULP1 expression was significantly positively correlated with Purity in 10 tumors, more so in KIPAN, TGCT, and SKCM, but significantly negatively correlated in 10 tumors including COAD, COADREAD, READ, and BLCA (Supplementary Fig. 10d). In addition, HRD status is a key indicator of treatment selection and prognosis in a wide range of tumors42, and we observed a significant correlation between GULP1 expression and HRD status in nine tumors, with significant positive correlation in COAD, LAML, STES, HNSC, and LIHC, and significant negative correlation in BRCA, KIPAN, UCEC, and THYM (Supplementary Fig. 10e) .
CD8 cytotoxic T lymphocytes (CTLs) are immune cells that specifically attack tumors43. Impairment of CTLs promotes the emergence of adaptive immune resistance, resulting in tumor immune escape44. We investigated the relationship between GULP1 and CTL dysfunction through the TIDE database. We discovered a positive correlation between GULP1 and the degree of CTL dysfunction in BRCA, LUAD, SARC, and AML(Supplementary Table 4). High GULP1 expression resulted in decreased infiltration of CTLs, which was detrimental to antitumor immunity and reduced OS in these patients. Conversely, lowering GULP1 expression reversed this unfavorable outcome (Fig. 9c). GULP1 expression was negatively correlated with the degree of CTL dysfunction in DLBC, KIRC and SKCM (Supplementary Table 4). High expression of GULP1 instead led to increased infiltration of CTLs, favoring anti-tumor immunity and prolonging patient survival (Fig. 9c).
GULP1 predicts immunotherapy response and sensitive drugs. Based on the TISMO web tool, box line plots showing the expression of GULP1 in cancer cell lines before and after cytokine treatment (a), and the expression of GULP1 in cancer cell lines before and after PDL1 or CTLA4 treatment (b). *p < 0.05, **p < 0.01, ***p < 0.001. correlation analysis between GULP1 and CTL (c). ROC Plotter was applied to assess the predictive effect of GULP1 on the response to BRCA, OV, COADREAD, and GBM immunotherapy (d). The top 5 drugs showing positive or negative correlation with GULP1 expression based on drug sensitivity databases CTRP (e) and GDSC (f).
We explored the response to cytokine therapies and immunotherapy based on GULP1 expression in mouse models by TISMO online tools, and showed that IFNβ and IFNγ exhibited favorable therapeutic effects in high GULP1-expressing mouse models, especially in SKCM, BRCA, lung cancer, COADREAD and HNSC cell lines (Fig. 9a). GULP1 expression levels predicted the response of SKCM, BRCA, HNSC and gastric adenocarcinoma to PD1, PDL1 or CTLA4 immunotherapy in mouse models (Fig. 9b). We validated the predictive effect of GULP1 on treatment response to cancer immune-checkpoint blockers (ICBs) through the ROC Plotter database (Fig. 9d), and the results showed that BRCA patients who exhibited elevated levels of GULP1 expression showed a greater likelihood of responding positively to chemotherapy, while OVs with high GULP1 expression, COADREAD patients were more likely to develop chemoresistance. In addition, GULP1 expression in GBM showed no significant difference in immunotherapy response.
According to the correlation analysis between GULP1 expression and drug sensitivity based on the CTRP dataset, the top 5 drugs positively correlated with GULP1 expression were, in order, BRD-K30748066, dinaciclib, SNX-2112, PX-12, and narciclasine. In contrast, abiraterone, VAF-347, ML334 diastereomer, trametinib, and BRD-K99006945 were the top 5 drugs negatively associated with GULP1 expression (Fig. 9e). Based on the correlation analysis of GULP1 levels with drug sensitivity in the GDSC dataset, Methotrexate, NPK76-11-72-1, BX-912, PIK-93, and AT-7519 were the top 5 drugs positively correlated with GULP1 expression. In contrast, the top 5 drugs negatively correlated with GULP1 expression were, in order, trametinib, 17-AAG, selumetinib, PD-0325901, and RDEA119 (Fig. 9f). These results demonstrate the value of GULP1 in predicting the efficacy of cancer immunotherapy.
GULP1 enhances the proliferation, migration and invasion of PAAD cells
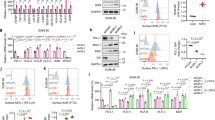
Given that GULP1 was significantly overexpressed in PAAD and correlated with poor OS, DSS, PFI, and DFI in this condition, we conducted further analysis and validation in PAAD. Interestingly, we found higher levels of GULP1 expression in male obese patients compared to female normal-weight pancreatic cancer patients, which also implies that male obese patients possess a worse prognosis (Supplementary Fig. 11a). In addition, the area under the ROC curve (AUC) reached 0.724, demonstrating the significant value of GULP1 expression for PAAD (Fig. 10a). Results from the CCLE database demonstrated the expression level of GULP1 in common pancreatic cancer cell lines (Fig. 10b). To explore the biological function of GULP1 in PAAD, we first performed a correlation analysis between GULP1 and marker genes. The results indicated that the expression level of GULP1 was profoundly positively correlated with the expression of proliferation marker genes PCNA and KI67, apoptosis inhibition marker gene BCL2, autophagy marker gene p62, cell cycle marker genes CDK1 and CDK6, and invasion marker gene KRAS (Supplementary Fig. 11b). Next, we knocked down GULP1 in ASPC1 and PANC1 cells for in vitro experiments, and Western blotting confirmed the transfection efficiency of si-GULP1 (Fig. 10c). CCK8 results indicated that the proliferation ability of ASPC1 and PANC1 cells was greatly decreased after GULP1 was knocked down (Fig. 10d). The results of wound healing assay (Fig. 10e, g) and transwell assay (Fig. 10f, h) also showed that GULP1 knockdown reduced the migration and invasion ability of ASPC1 and PANC1 cells. We analyzed the possible mechanisms by which GULP1 affects the biological functions of PAAD cells. GULP1 was found to be significantly enriched in the HIPPO, mTOR, and RTK signaling pathways (Supplementary Fig. 11c) and significantly positively correlated with key genes of these pathways (Fig. 10i). This suggests that GULP1 may regulate the proliferation, migration, and invasion of PAAD cells through HIPPO, mTOR, and RTK signaling pathways.
Biological functions of GULP1 in PAAD. ROC curve to assess the diagnostic value of GULP1 in PAAD (a). Relative mRNA expression of GULP1 in common PAAD cell lines from the CCLE database (b). Western blotting to verify the knockdown efficiency of GULP1 in PAAD cell lines (c). CCK8 was used to determine the proliferative capacity of PAAD cells (d). Wound healing assay (g) and transwell assay (h) to detect PAAD cell migration and invasion ability. Histograms show the relative area of migration (e) and the number of invaded cells (f), ***p < 0.001, ****p < 0.0001. Scatter plot of correlation between GULP1 expression and pathway marker genes (i).
Discussion
Cancer is a global public health problem. According to the assessment of the International Agency for Research on Cancer (IARC), cancer is projected to surpass cardiovascular disease as the primary cause of premature death in the majority of countries around the world in this century. The global cancer burden continues to increase, posing significant hazards to human life and health security45. As the world’s most populous country, China also has the largest aging population in the world, and the cancer data in China are not optimistic46. According to the latest statistics, China will have about 4,824,700 new cancer cases and 2,574,200 new cancer deaths in 2022. Both in terms of the number of new cases and deaths, China ranks first in the world46.
PAAD is one of the deadliest cancers among all solid tumors and the malignant tumor with mortality rate closest to the incidence rate, and has the title of “king of cancers“47. According to the latest data released this year, PAAD ranks 9th among malignant tumors in men and 13th in women in China, and ranks 6th and 7th in malignant tumor mortality, respectively46. It ranks 7th in incidence among women in the United States and is the 3rd most common cause of cancer-related deaths in women1. The geographic distribution of PAAD varies widely, being highest in North America and Western Europe and lowest in East Africa and Central and South Asia48. This is likely related to the higher prevalence of obesity and diabetes in economically developed regions48,49. In addition, several studies have shown that factors such as smoking, alcohol abuse, injury infection, chronic pancreatitis, and familial inheritance are strongly associated with the development of PAAD47. Surgical eradication is the only chance to cure PAAD so far. However, due to the subtle symptoms in the initial phase of PAAD, most of the patients are in the middle or late stage when they visit the clinic, and the opportunity of radical surgery is lost. And within 2 years after surgery, close to 80% of PAAD patients experience recurrence and unpredictable metastasis, which greatly increases the difficulty of treatment47. Therefore, exploring the molecular mechanisms of PAAD recurrence and metastasis and searching for potential biomarkers are of great significance in reducing PAAD recurrence and metastasis.
GULP1 acts as a specific articulating protein that promotes phagocytosis8. It is involved in various biological processes such as apoptosis, lipid transport, and phagocytosis10,11. Prior research has indicated that GULP1 can transactivate the KEAP1-NRF2 signaling pathway in BLCA, thereby inducing cancer cell proliferation, invasion, and resistance to cisplatin drugs15. In addition, GULP1 activates SMAD3 phosphorylation or inactivates AKT/PDK1 and MAPK in ovarian cancer cells, inhibiting cancer cell proliferation12. However, the biological function of GULP1 in cancer progression remains largely unknown.
In the present study, we systematically evaluated the value of the phagocytic bridging protein GULP1 in pan-cancer using multiple bioinformatics techniques. We found that GULP1 was tissue-specific in normal tissues of the epididymis. According to the pan-cancer expression results from the TGCA and GTEx databases, GULP1 was significantly down-regulated in most malignant tumors, but significantly highly expressed in GBM, GBMLGG, LGG, LUAD, KIRP, PRAD, SKCM, PAAD, KICH, and CHOL. This suggests that GULP1 is likely to become a new cancer biomarker. Tumor typing, grading and staging are important indicators for evaluating the biological behavior and invasive metastatic ability of tumors and the degree of clinical progression, especially molecular typing, which is important for the implementation of individualized tumor therapy50. We found that aberrant GULP1 expression was associated with molecular subtypes of BRCA, COAD, ESCA, HNSC, KIRP, LGG, LIHC, LUSC, PCPG, PRAD, STAD, and UCEC, tumor grades of HNSC, KIRC, OV, and UCEC, pathological stages of BLCA, COAD, ESCA, LIHC, PAAD, STAD, KIRC, UCEC and TGCT. This suggests that the potential functions and biological processes of GULP1 may differ in different tumor types. COX regression analysis showed that GULP1 is a risk factor for a variety of cancers, which further confirms to some extent that GULP1 shows promise as a reliable biomarker for cancer. Prior research has demonstrated that the expression of GULP1 is elevated in SKCM and promotes tumor metastasis51. low GULP1 expression is associated with a poor prognosis in BLCA patients14,15. These results are consistent with our findings. Notably, further Kaplan-Meier survival analysis showed that GULP1 expression level was significantly associated with four prognostic indicators, OS, DSS, DFI, and PFI, in COAD, GBMLGG, KICH, LIHC, PAAD, STAD, and STES. and the diagnostic value of GULP1 expression for PAAD reached more than 70%. This provides a reference direction for future studies.
Gene mutations disrupt the integrity and stability of the normal cellular genome and are closely associated with the development of cancer52. The study of gene mutations is significant in exploring the pathogenesis of cancer. In this study, we found that GULP1 mutations were associated with expression in a variety of cancers. And GULP1 mutations were also associated with the prognosis of UCEC, ACC, MESO, SARC, TGCT, COAD, DLBC, and PAAD. GULP1 expression was significantly positively linked to MMRs gene mutations in nearly all types of cancer. Post-translational modifications play a key role in regulating cell signaling in cancer cells53. DNA methylation is a crucial epigenetic alteration that has the potential to influence gene expression54. In this study, we observed GULP1 promoter hypermethylation in BLCA, BRCA, CESC, UCEC, HNSC, KIRC, LIHC, LUAD, and LUSC compared to normal tissues. GULP1 was reported to be highly hypermethylated in HNSC, LIHC tissues55,56. GULP1 hypermethylation in BLCA was negatively correlated with expression15. This is in keeping with our results. DNA methylation dysregulation is associated with cancer development and can affect cancer prognosis57. In this study, we identified 29 CpG islands of GULP1 whose methylation affects cancer prognosis, with cg13274254 accounting for the largest proportion. cg13274254 site methylation levels affect LAML, BRCA, HNSC, KIRC, LUAD, MESO, SKCM, KIRP, and LGG prognosis. Activation of many cancer-related signaling pathways is associated with protein phosphorylation58. Significant progress has been made in the development of targeted drugs against specific phosphorylation sites in cancer therapy59. Therefore, studying the mechanism of protein phosphorylation in cancer is important for revealing the molecular mechanism of cancer development, discovering new therapeutic targets, and developing more effective anticancer drugs. Our study showed that phosphorylation of S206 and T208 residues of GULP1 exhibited significant differences in a variety of cancers. including KIRC, OV, PAAD, HNSC, LUAD, and BRCA. mutations and epigenetic alterations allow tumor cells to acquire features such as uncontrolled proliferation, differentiation, and immune escape60. Our results suggest that GULP1 plays an important role in tumorigenesis at both genetic and epigenetic levels.
The single-cell sequencing results demonstrated multiple biological behaviors of cancer that may be regulated by GULP1, such as cell differentiation, proliferation, apoptosis, cell cycle, angiogenesis, EMT, invasion and metastasis. In contrast, functional enrichment analysis revealed the potential mechanisms by which GULP1 regulates the functional phenotypes of cancer, such as endocytosis, Rac/Ras protein signaling and chemokine signaling pathways. Our experimental results confirmed that GULP1 could regulate PAAD cell proliferation, migration and invasion, possibly by affecting HIPPO, mTOR and RTK signaling pathways.
Since the immune system was identified in the nineteenth century as a useful tool for targeting neoplastic diseases, more and more researchers have been keen on exploring the mechanisms of tumor immunity and developing tumor immunotherapies61. Some of these infiltrating immune cells are responsible for the ability of cancer cells to evade the immune response and establish their own tumor microenvironment (TME)62. Immune cells are crucial in identifying cancer cells and controlling tumor development63. Tumor-associated macrophages (TAMs) are a prominent cell type in TME that regulate cancer-associated inflammation64. High infiltration of TAM has been linked to a negative outcome in cancer65. TAM can mediate tumor immunity by modulating the function of other immune cells or by manufacturing diverse cytokines64,65. CAF not only provides mesenchymal support to tumor cells, but also plays a vital part in regulating cancer cell growth and metastasis66,67. Endothelial cells are cells that form the lining of blood vessels and can form new blood vessels to provide nutrients for cancer growth and metastasis68. Monocytes in the tumor microenvironment can be activated by signaling molecules released by tumor cells and other cells, resulting in polarization and transformation into macrophages that exert both anti-tumor and pro-tumor effects69. Tregs can maintain immune homeostasis through multiple pathways, including expression of CD25 and FOXP370,71. T lymphocytes participate in tumor immunity by secreting a variety of cytokines72. CD8 + T cells improves the prognosis of most cancer patients72. B cells are mainly responsible for the manufacture of antibodies, such as IgE, IgG, IgA, and IgM, to exhibit antitumor and immunomodulatory effects73. Our study identified significant correlations between GULP1 expression and CAF, Endo, Macrophage, Monocyte, Neutrophil, NKT, CD4 + T cells, CD8 + T cells, Tregs, DCs, and B cell infiltration in TME. Similarly, we analyzed the correlation between GULP1 expression and three immunity scores and immunomodulatory genes. The results also confirmed the association of GULP1 with tumor immunity. In most cancers, GULP1 expression was positively correlated with most immunomodulatory genes. For example, the chemokine genes CXCL12, CX3CL1, and CXCL5, and the receptor gene CX3CR1. We hypothesized that it is possible that GULP1 could form positive feedback with certain immune-regulated genes to enhance the malignant behavior of tumors and cause tumor cells to evade immune surveillance. Notably, in PAAD, GULP1 expression was associated with the Immunoinhibitor genes TGFBR1, CD274, PDCD1LG2 and the Immunostimulator genes RAET1E, HHLA2, TNFSF13, TNFSF15, NT5E, TNFSF4, ENTPD1, TNFSF18, MICB, CD80, IL2RA, and TNFRSF9. Analyzing the relationship between GULP1 expression and cancer immune cell infiltration, immune scores, and immunoregulatory genes will enhance our comprehension of how tumors escape the immune system, and provide an important reference for the development of new immunotherapeutic strategies.
In the past few years, high-throughput sequencing has led to the discovery of numerous novel immune checkpoints74,75. ICBs have improved the landscape of cancer treatment and have become one of the most popular tumor immunotherapies76,77. ICB includes anti-PD-1, anti-PD-L1, and anti-CTLA4 therapies, and durable anticancer responses lead to longer-lasting disease remission76,78. However, anti-tumor immunity is hampered by a variety of reasons, including tumor heterogeneity, intra-tumor immune cell crosstalk, and dysfunction of CTLs, leading to the development of drug resistance and severely limiting the number of patients who can benefit from ICB79,80,81. TMB, MSI, tumor purity, and HDR are thought to influence the efficacy of ICB41,42. We analyzed the correlation between GULP1 and the above four ICB metrics. The results showed that, TMB was positively correlated with GULP1 expression in LAML, THYM and negatively correlated in LGG, CESC, BRCA, KIPAN, PRAD, HNSC, KIRC. GULP1 expression was positively correlated with MSI in KIPAN, KIRC, TGCT, and CHOL, and negatively correlated with MSI in LUAD, HNSC, and DLBC. GULP1 expression was significantly positively correlated with Purity in 10 tumors, more significantly in KIPAN, TGCT, and SKCM, but significantly negatively correlated in 10 tumors including COAD, COADREAD, READ, and BLCA. In addition, a notable correlation was detected between the expression of GULP1 and HRD status in 9 tumors, which were significantly positively correlated in COAD, LAML, STES, HNSC, LIHC, and significantly negatively correlated in BRCA, KIPAN, UCEC, and THYM. The above results suggest the potential of classification based on these four ICB metrics for specific tumor immunotherapy. Our research showed a intriguing connection between GULP1 and CAFs, Macrophage and Tregs cell infiltration in multiple tumors. Previous studies have shown that Macrophage, Tregs, and CAFs regulate the activation and infiltration of CTLs in TME82. Dysfunction of CTLs promotes tumor immune escape, which leads to tumor growth, metastasis, and drug resistance43. We found an association between GULP1 and CTLs dysfunction and prognosis in BRCA, LUAD, SARC, AML, DLBC, KIRC, and SKCM. We also found that high GULP1-expressing BRCA patients were more likely to benefit from chemotherapy, and conversely, high GULP1-expressing OV and COADREAD patients were more likely to develop chemoresistance. In addition, the response of SKCM, BRCA, HNSC, and gastric adenocarcinoma to PD1, PD-L1, or CTLA4 immunotherapy could be effectively predicted based on the expression level of GULP1 in mouse models. Cytokines have been widely recognized as cancer therapeutic targets83. In the present study, we found that cytokine therapies (IFNβ and IFNγ) exhibited favorable therapeutic effects in high GULP1-expressing SKCM, BRCA, lung, COADREAD, and HNSC mouse models. Predicting individualized drug treatment response in advance would greatly improve the therapeutic efficacy and reduce the therapeutic risk84. In this study, we predicted sensitive small molecule drugs from the CTRP and GDSC databases based on GULP1 expression. This will provide a reference for drug development targeting GULP1. In summary, we believe that GULP1 has great potential as an immunoprediction or therapy-related biomarker. However, the current study lacks relevant experiments to verify the specific relationship between GULP1 and TME, which suggests the need for further experimental work.
In conclusion, we comprehensively analyzed GULP1 in pan-cancer by multiple bioinformatics techniques and validated its biological function in PAAD for the first time. We found that GULP1 may be a potentially powerful biomarker for cancer and can effectively predict cancer immunotherapy response. However, more experiments must take place to demonstrate our findings and elucidate the detailed mechanisms.
Conclusion
GULP1 is differentially expressed in most tumors and is associated with negative prognosis, which may be related to the involvement of GULP1 in the regulation of apoptotic pathways. GULP1 can reduce the effect of anti-tumor immunotherapy by increasing the infiltration of immune cells and inhibiting the function of cytotoxic T lymphocytes. In addition, the significantly high expression of GULP1 in pancreatic cancer suggests a poor prognosis. GULP1 regulates the proliferation, invasion, and metastasis of pancreatic cancer cells possibly by affecting the HIPPO, mTOR, and RTK signaling pathways. GULP1 has the potential to be a potential biomarker for prognosis and immunotherapy in pan- and pancreatic cancer.
Data availability
The original contributions presented in the study are included in the article/Supplementary Material. Further inquiries can be directed to the corresponding author.
Change history
26 September 2025
The original online version of this Article was revised: In this article author Wangbiao Wang was incorrectly denoted as the corresponding author, also the following information was missing from the Funding section: Tianshan Talents - Leading Talents in Science and Technology Innovation of Xinjiang Uygur Autonomous Region (2023TSLJ0038). The original article has been corrected.
References
Siegel, R. L., Giaquinto, A. N. & Jemal, A. Cancer statistics, 2024. CA Cancer J. Clin. 74, 12–49 (2024).
Xia, C. et al. Cancer statistics in China and united States, 2022: profiles, trends, and determinants. Chin. Med. J. (Engl). 135, 584–590 (2022).
Fitzgerald, R. C., Antoniou, A. C., Fruk, L. & Rosenfeld, N. The future of early cancer detection. Nat. Med. 28, 666–677 (2022).
Fitzmaurice, C. et al. Global, regional, and National Cancer incidence, mortality, years of life lost, years lived with disability, and Disability-Adjusted life-Years for 29 Cancer groups, 1990 to 2017: A systematic analysis for the global burden of disease study. JAMA Oncol. 5, 1749–1768 (2019).
Lin, L. et al. Global, regional, and National cancer incidence and death for 29 cancer groups in 2019 and trends analysis of the global cancer burden, 1990–2019. J. Hematol. Oncol. 14, 197 (2021).
Soerjomataram, I. & Bray, F. Planning for tomorrow: global cancer incidence and the role of prevention 2020–2070. Nat. Rev. Clin. Oncol. 18, 663–672 (2021).
Bray, F., Laversanne, M., Weiderpass, E. & Soerjomataram, I. The ever-increasing importance of cancer as a leading cause of premature death worldwide. Cancer 127, 3029–3030 (2021).
Smits, E., Van Criekinge, W., Plaetinck, G. & Bogaert, T. The human homologue of Caenorhabditis elegans CED-6 specifically promotes phagocytosis of apoptotic cells. Curr. Biol. 9, 1351–1354 (1999).
Liu, Q. A. & Hengartner, M. O. Human CED-6 encodes a functional homologue of the Caenorhabditis elegans engulfment protein CED-6. Curr. Biol. 9, 1347–1350 (1999).
Kiss, R. S. et al. The lipoprotein receptor-related protein-1 (LRP) adapter protein GULP mediates trafficking of the LRP ligand prosaposin, leading to sphingolipid and free cholesterol accumulation in late endosomes and impaired efflux. J. Biol. Chem. 281, 12081–12092 (2006).
Ma, Z., Nie, Z., Luo, R., Casanova, J. E. & Ravichandran, K. S. Regulation of Arf6 and ACAP1 signaling by the PTB-domain-containing adaptor protein GULP. Curr. Biol. 17, 722–727 (2007).
Maldonado, L. et al. Integrated transcriptomic and epigenomic analysis of ovarian cancer reveals epigenetically silenced GULP1. Cancer Lett. 433, 242–251 (2018).
Paret, C. et al. Identification of an Immunogenic Medulloblastoma-Specific fusion involving EPC2 and GULP1. Cancers (Basel). 13, 2072–6694 (2021).
Yan, D. et al. The APC/C E3 ligase subunit ANAPC11 mediates FOXO3 protein degradation to promote cell proliferation and lymph node metastasis in urothelial bladder cancer. Cell. Death Dis. 14, 516 (2023).
Hayashi, M. et al. GULP1 regulates the NRF2-KEAP1 signaling axis in urothelial carcinoma. Sci. Signal. 13, eaba0443 (2020).
Saidak, Z. et al. A pan-cancer analysis of the human tumor coagulome and its link to the tumor immune microenvironment. Cancer Immunol. Immunother. 70, 923–933 (2021).
Chen, H. et al. A Pan-Cancer analysis of enhancer expression in nearly 9000 patient samples. Cell 173, 386–399e312 (2018).
Goldman, M. J. et al. Visualizing and interpreting cancer genomics data via the Xena platform. Nat. Biotechnol. 38, 675–678 (2020).
Shen, W. et al. Sangerbox: A comprehensive, interaction-friendly clinical bioinformatics analysis platform. iMeta 1, e36 (2022).
Nusinow, D. P. et al. Quantitative proteomics of the Cancer cell line encyclopedia. Cell 180, 387–402e316 (2020).
Chandrashekar, D. S. et al. UALCAN: an update to the integrated cancer data analysis platform. Neoplasia (New York NY). 25, 18–27 (2022).
Colwill, K. & Gräslund, S. A roadmap to generate renewable protein binders to the human proteome. Nat. Methods. 8, 551–558 (2011).
Ru, B. et al. TISIDB: an integrated repository portal for tumor-immune system interactions. Bioinformatics 35, 4200–4202 (2019).
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
Liu, C. J. et al. GSCA: an integrated platform for gene set cancer analysis at genomic, Pharmacogenomic and Immunogenomic levels. Brief. Bioinform. 24, bbac558 (2023).
Modhukur, V. et al. MethSurv: a web tool to perform multivariable survival analysis using DNA methylation data. Epigenomics 10, 277–288 (2018).
Yuan, H. et al. CancerSEA: a cancer single-cell state atlas. Nucleic Acids Res. 47, D900–d908 (2019).
Sun, D. et al. TISCH: a comprehensive web resource enabling interactive single-cell transcriptome visualization of tumor microenvironment. Nucleic Acids Res. 49, D1420–d1430 (2021).
Szklarczyk, D. et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 47, D607–d613 (2019).
Tang, Z., Kang, B., Li, C., Chen, T. & Zhang, Z. GEPIA2: an enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Res. 47, W556–w560 (2019).
Li, T. et al. TIMER2.0 for analysis of tumor-infiltrating immune cells. Nucleic Acids Res. 48, W509–w514 (2020).
Fu, J. et al. Large-scale public data reuse to model immunotherapy response and resistance. Genome Med. 12, 21 (2020).
Zeng, Z. et al. TISMO: syngeneic mouse tumor database to model tumor immunity and immunotherapy response. Nucleic Acids Res. 50, D1391–d1397 (2022).
Kovács, S. A., Fekete, J. T. & Győrffy, B. Predictive biomarkers of immunotherapy response with Pharmacological applications in solid tumors. Acta Pharmacol. Sin. 44, 1879–1889 (2023).
Rees, M. G. et al. Correlating chemical sensitivity and basal gene expression reveals mechanism of action. Nat. Chem. Biol. 12, 109–116 (2016).
Yang, W. et al. Genomics of drug sensitivity in Cancer (GDSC): a resource for therapeutic biomarker discovery in cancer cells. Nucleic Acids Res. 41, D955–961 (2013).
Smith, J., Sen, S., Weeks, R. J., Eccles, M. R. & Chatterjee, A. Promoter DNA hypermethylation and Paradoxical gene activation. Trends cancer. 6, 392–406 (2020).
Nishiyama, A. & Nakanishi, M. Navigating the DNA methylation landscape of cancer. Trends Genet. 37, 1012–1027 (2021).
Liu, J., Peng, Y. & Wei, W. Cell cycle on the crossroad of tumorigenesis and cancer therapy. Trends Cell. Biol. 32, 30–44 (2022).
Ren, L. et al. Systematic pan-cancer analysis identifies APOC1 as an immunological biomarker which regulates macrophage polarization and promotes tumor metastasis. Pharmacol. Res. 183, 106376 (2022).
Cristescu, R. et al. Tumor mutational burden predicts the efficacy of pembrolizumab monotherapy: a pan-tumor retrospective analysis of participants with advanced solid tumors. J. Immunother. Cancer. 10, e003091 (2022).
Pilié, P. G., George, A. & Yap, T. A. Patient selection biomarker strategies for PARP inhibitor therapy. Ann. Oncol. 31, 1603–1605 (2020).
Monteran, L. & Erez, N. The dark side of fibroblasts: Cancer-Associated fibroblasts as mediators of immunosuppression in the tumor microenvironment. Front. Immunol. 10, 1835 (2019).
Farhood, B., Najafi, M. & Mortezaee, K. CD8(+) cytotoxic T lymphocytes in cancer immunotherapy: A review. J. Cell. Physiol. 234, 8509–8521 (2019).
Dagenais, G. R. et al. Variations in common diseases, hospital admissions, and deaths in middle-aged adults in 21 countries from five continents (PURE): a prospective cohort study. Lancet 395, 785–794 (2020).
Han, B. et al. Cancer Incidence and Mortality in China, 2022 (Journal of the National Cancer Center, 2024).
Khalaf, N., El-Serag, H. B., Abrams, H. R. & Thrift, A. P. Burden of pancreatic cancer: from epidemiology to practice. Clin. Gastroenterol. Hepatol. 19, 876–884 (2021).
Wild, C., Weiderpass, E. & Stewart, B. W. World cancer Report: cancer Research for cancer Prevention (International Agency for Research on Cancer, 2020).
Arjani, S., Saint-Maurice, P. F., Julián-Serrano, S., Eibl, G. & Stolzenberg-Solomon, R. Body mass index trajectories across the adult life course and pancreatic Cancer risk. JNCI cancer Spectr. 6, pkac066 (2022).
Bignold, L. P. Typing, grading, and staging of cases of tumor. Principles of Tumors 279–315 (2020).
Marie, K. L. et al. Melanoblast transcriptome analysis reveals pathways promoting melanoma metastasis. Nat. Commun. 11, 333 (2020).
Pirozzi, C. J. & Yan, H. The implications of IDH mutations for cancer development and therapy. Nat. Rev. Clin. Oncol. 18, 645–661 (2021).
Geffen, Y. et al. Pan-cancer analysis of post-translational modifications reveals shared patterns of protein regulation. Cell 186, 3945–3967e3926 (2023).
Martisova, A. et al. DNA methylation in solid tumors: functions and methods of detection. Int. J. Mol. Sci. 22, 4247 (2021).
Inokawa, Y. et al. High-risk HPV infection-associated hypermethylated genes in oropharyngeal squamous cell carcinomas. BMC Cancer. 22, 1146 (2022).
Du, B., Wang, F., Jarad, B., Wang, Z. & Zhang, Y. A novel signature based on microvascular invasion predicts the recurrence of HCC. J. Transl Med. 18, 272 (2020).
Koch, A. et al. Analysis of DNA methylation in cancer: location revisited. Nat. Rev. Clin. Oncol. 15, 459–466 (2018).
Bugter, J. M., Fenderico, N. & Maurice, M. M. Mutations and mechanisms of WNT pathway tumour suppressors in cancer. Nat. Rev. Cancer. 21, 5–21 (2021).
Yip, H. Y. K. & Papa, A. Signaling pathways in cancer: therapeutic targets, combinatorial treatments, and new developments. Cells 10, 659 (2021).
Hanahan, D. Hallmarks of cancer: new dimensions. Cancer Discov. 12, 31–46 (2022).
Yang, K. et al. Glioma targeted therapy: insight into future of molecular approaches. Mol. Cancer. 21, 39 (2022).
Hiam-Galvez, K. J., Allen, B. M. & Spitzer, M. H. Systemic immunity in cancer. Nat. Rev. Cancer. 21, 345–359 (2021).
Talty, R. & Olino, K. Metabolism of innate immune cells in Cancer. Cancers (Basel). 13, 904 (2021).
Chen, Y. et al. Tumor-associated macrophages: an accomplice in solid tumor progression. J. Biomed. Sci. 26, 78 (2019).
Mantovani, A., Marchesi, F., Malesci, A., Laghi, L. & Allavena, P. Tumour-associated macrophages as treatment targets in oncology. Nat. Rev. Clin. Oncol. 14, 399–416 (2017).
Min, K. W. et al. Cancer-associated fibroblasts are associated with poor prognosis in solid type of lung adenocarcinoma in a machine learning analysis. Sci. Rep. 11, 16779 (2021).
Liu, T. et al. Cancer-associated fibroblasts: an emerging target of anti-cancer immunotherapy. J. Hematol. Oncol. 12, 86 (2019).
Zeng, Q. et al. Understanding tumour endothelial cell heterogeneity and function from single-cell omics. Nat. Rev. Cancer. 23, 544–564 (2023).
Ugel, S., Canè, S., De Sanctis, F. & Bronte, V. Monocytes in the tumor microenvironment. Annu. Rev. Pathol. 16, 93–122 (2021).
Wang, L. et al. Blood transfusion practices affect CD4(+) CD25(+) FOXP3(+) regulatory T cells/t helper-17 cells and the clinical outcome of geriatric patients with hip fracture. Aging (Albany N Y). 13, 21408–21420 (2021).
Sakaguchi, S. et al. Regulatory T cells and human disease. Annu. Rev. Immunol. 38, 541–566 (2020).
St Paul, M. & Ohashi, P. S. The roles of CD8(+) T cell subsets in antitumor immunity. Trends Cell. Biol. 30, 695–704 (2020).
Liu, Y. et al. High-plex protein and whole transcriptome co-mapping at cellular resolution with Spatial CITE-seq. Nat. Biotechnol. 41, 1405–1409 (2023).
Wang, Z. et al. The CXCL family contributes to immunosuppressive microenvironment in gliomas and assists in gliomas chemotherapy. Front. Immunol. 12, 731751 (2021).
Zhang, H. et al. Novel immune infiltrating cell signature based on cell pair algorithm is a prognostic marker in Cancer. Front. Immunol. 12, 694490 (2021).
Morad, G., Helmink, B. A., Sharma, P. & Wargo, J. A. Hallmarks of response, resistance, and toxicity to immune checkpoint Blockade. Cell 185, 576 (2022).
Jhunjhunwala, S., Hammer, C. & Delamarre, L. Antigen presentation in cancer: insights into tumour immunogenicity and immune evasion. Nat. Rev. Cancer. 21, 298–312 (2021).
Zhu, S. et al. Combination strategies to maximize the benefits of cancer immunotherapy. J. Hematol. Oncol. 14, 156 (2021).
Garner, H. & de Visser, K. E. Immune crosstalk in cancer progression and metastatic spread: a complex conversation. Nat. Rev. Immunol. 20, 483–497 (2020).
de Visser, K. E. & Joyce, J. A. The evolving tumor microenvironment: from cancer initiation to metastatic outgrowth. Cancer Cell. 41, 374–403 (2023).
Vitale, I., Shema, E., Loi, S. & Galluzzi, L. Intratumoral heterogeneity in cancer progression and response to immunotherapy. Nat. Med. 27, 212–224 (2021).
Liu, K. et al. Pan-cancer analysis of the prognostic and immunological role of ANLN: an onco-immunological biomarker. Front. Genet. 13, 922472 (2022).
Propper, D. J. & Balkwill, F. R. Harnessing cytokines and chemokines for cancer therapy. Nat. Rev. Clin. Oncol. 19, 237–253 (2022).
Adam, G. et al. Machine learning approaches to drug response prediction: challenges and recent progress. NPJ Precision Oncol. 4, 19 (2020).
Funding
This study was supported by the National Natural Science Foundation of China (82060520), the Guiding Funds of Central Government for Supporting the Development of the Local Science and Technology (ZYYD2022). Tianshan Talents - Leading Talents in Science and Technology Innovation of Xinjiang Uygur Autonomous Region (2023TSLJ0038).
Author information
Authors and Affiliations
Contributions
WBW, GS, and YXL conceived and designed the study. WBW and YXL conducted the experiments. SAL, FL, JG, FZL, GA, and JYL analyzed and organized the data. WBW wrote the original manuscript. GS revised the manuscript. All authors have read and agreed to the publication of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, W., Li, Y., Li, S. et al. The value of GULP1 in cancer prognosis and immunotherapy, validated from pan-cancer analysis to pancreatic cancer. Sci Rep 15, 19021 (2025). https://doi.org/10.1038/s41598-025-99909-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-99909-6