Abstract
Cytotoxic drugs form a heterogeneous group of antineoplastic agents commonly employed in the management of cancer and other disorders but are commonly linked with limited therapeutic indices and severe side effects. It is important to learn their physicochemical properties thus making a prediction of absorption, permeability and distribution. One of such properties is the Topological Polar Surface Area (Top_PSA), an important property of membrane transport and a popular surrogate of passive diffusion and blood brain barrier permeability. We also explored in this study whether graph-theoretrical and molecular descriptors could be a consistent predictor of RDKit/Mordred-calculated Top_PSA values of a curated dataset of 156 structure-diverse cytotoxic agents. Fifty eight descriptors were calculated and preprocessed in five pre-processing schemes such as direct fitting, PCA, robust scaling, identification and elimination of outliers, and feature selection based on the VIF by using linear, LASSO and ridge regression model. K-fold cross-validation was strictly applied to all the models. The best predictive performance of robust scaling with LASSO was the largest \((R^{2} \approx 0.97)\), which proves the effectiveness of robust preprocessing and a sparsity-inducing regularization in the case of heteroscedasticity and multicollinearity in datasets with many descriptors. PCA performed similarly in terms of predictive accuracy but with a lower level of interpretability, and VIF-based pruning never did. The analysis of the non-zero LASSO coefficients indicated that the heteroatom content, the ability to form hydrogen-bonds, and a set of indices that are weighted by electronegativity were found to be significant factors contributing to Top_PSA, which is in line with the chemical definition of it being fragment-based. Overall, the given work provides valuable recommendations about preprocessing, feature selection, and model selection in QSAR working processes, and the importance of clear computational pipelines to gain correct Top_PSA predictions by the use of descriptors.
Similar content being viewed by others
Introduction
Cytotoxic drugs are seldom administered on their own, as these are usually combined with chemotherapy or radiotherapy. There is a rationale behind combining drugs rather than just using a single agent. This is not dissimilar to the rationale behind using two or more drugs in antibiotic treatment; it is a means of attempting to prevent the emergence of drug resistance. During initial diagnosis, many tumors are likely to contain cancer cells that have developed naturally without external factors and gained tolerance to cytotoxic treatment. This is distinct from antibiotic resistance, which eradicates the need for earlier exposure to the drug. Innate mutation rates are sufficiently high to give a possibility for partial cellular mutation that resists the drugs, and subsequently, these cells undergo massive proliferation. In the course of monodrug therapies, these drugs may enable additional growth of resilient sublineages of tumor cells, thus regulating the potential for a definitive cure. From the start of treatment, there is the option of combining drugs to alleviate the problem. The choice of drugs for combination therapy is guided by three basic principles : (1) engage the use of drugs that are active against diseases. (2) Consider the use of drugs that act in various ways. (3) Employ drugs that have different toxicities. By employing drugs with different biological effects, such as pairing an agent that works as an antimetabolite with one that actively damages DNA, we can achieve a truly synergistic effect. It is not a good idea to apply a combination of drugs with comparable adverse effects: Two aggressively myelodepressing drugs may induce an unacceptably large threat of neutropenic sepsis. Combination therapy, where feasible, should be dictated by the toxicity profiles of the drugs involved. If one thinks of radiotherapy as one among many other drugs, one can also consider radiotherapy in conjunction with chemotherapy.
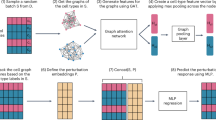
Layers of graph-based modeling and machine learning have recently been closely integrated in several biomedical as well as chemical analysis tasks. Network embedding and graph convolutional methods have been used to predict drug–disease1,2 and herb–disease associations3, they demonstrate that graph learning well aligns with complex biological interactions and can enhance the predictive power. Molecular networking workflows have also been developed to facilitate dereplication and identification of natural products, thus promoting systematic exploration of bioactive chemical space4. Simultaneously, deep learning methods have been applied for automatic feature selection, molecular generation and epigenetic prediction such as transporting gate based optimal variable selection technique5,6. Upcoming learning models such as the vision transformers and neuromorphic architectures, from where they are leveraged by the causal graph splitting system exploring seizure prediction, intelligent cell sorting or causality inference of complex biological networks7,8,9. Complementary to these computational advances, experimental work has elucidated mechanisms of drug-based control against inflammation, neuroprotection, cardiac fibrosis and endothelial dysfunction, thereby providing biological support for predictions made by computational models10,11,12. Together, these studies underscore the emerging importance of graph-based descriptors interfacing with machine learning as a powerful approach to quantitatively modeling both cytotoxic and therapeutic compounds that spans molecular structure through biological activity to data-driven drug discovery.
Another aspect of drug and radiation combinations that goes beyond synergy and toxicity is spatial cooperation. Chemotherapy is a highly invasive systemic treatment, while radiotherapy is not. But, radiotherapy can reach sites that drugs cannot, like the central nervous system and the testis. Prophylactic cranial irradiation, for instance, may be part of the treatment protocol in patients predominantly treated with chemotherapy in conditions such as leukaemias, lymphomas, and small-cell lung cancer. 38 pharmacological agents (chemotherapeutic and biological agents) currently used in anticancer formulations13. A few topological indices used in this study along with their mathematical definitions/ equations are given in the Table 1:
Rationale for predicting topological polar surface area
Topological Polar Surface Area (Top_PSA) is a physicochemical parameter that is presently largely used in drug-design studies because it has excellent correlations with molecular transport processes, such as absorption across the intestines, cell membrane permeability, passive diffusion, and with blood-brain barrier (BBB) penetration. The transport related properties are of particular interest to cytotoxic drugs with narrow therapeutic indices and with a requirement to be effectively transported to intracellular sites. Compounds of unduly high polar surface area usually have low membrane permeability and low oral bioavailability, and compounds of very low polar surface area may have undesired toxicity as a result of uncontrolled distribution. Top_PSA is thus an important parameter utilized when assessing the ease with which a cytotoxic agent can access its target site with the least amount of off-target exposure. In addition, Top_PSA is computationally friendly since it can be computed using 2-dimensional molecular topology, it does not need 3D conformer generation, and provides high-speed and reproducible screening of large virtual chemical libraries. In drug classes where extensive structural diversity is observed, i.e. antineoplastic agents, and early-stage filtering is essential (e.g. Top_PSA), accurate prediction can be used to give an efficient surrogate measure of the molecular behavior in terms of absorption and distribution. Our research question in this paper is whether Top_PSA can be predicted with the help of graph-theoretical measures and regression approaches, and how the preprocessing approaches can affect the predictor quality. Elucidating this correlation assists in the discovery of sets of descriptors that make the greatest contribution to the estimation of surface-polarity as well as aids the larger objective of comprehending the cytotoxic-compound behavior in terms of the topology-motivated modeling.
Data collection/ material
The dataset used in this study consists of 156 cytotoxic compounds whose structures were obtained from peer-reviewed literature and publicly accessible chemical databases, including PubChem and DrugBank. These compounds represent clinically relevant or experimentally validated cytotoxic agents used in anticancer therapy, ensuring that the dataset reflects molecules with established biological activity and curated structural information. All molecular structures were downloaded in standardized SMILES format to avoid transcription errors, and each entry was cross-checked against at least two independent chemical repositories to ensure structural accuracy and consistency.
In order to assess chemical quality and integrity, every compound was preprocessed by following several steps such as valence verification, inorganic species removal, and charge balance verification using RDKit. Ambiguous stereochemical molecules and those molecules that had no structural annotation were removed. Following these filters, 156 high-quality compounds were left, which gave a clean and chemically coherent dataset that could be used to compute descriptors and regression modelling.
These molecules have structural diversity across a range of chemotypes with a variety of common cytotoxic drugs classes, such as, alkylating agents, antimetabolites, topoisomerase, anthracyclines, vinca alkaloids, and antitumor antibiotics. Such diversity is manifested as a wide variety of molecular weights, aromaticity, ring systems, hydrogen-bonding capabilities etc. This type of structural heterogeneity makes sure that the space of descriptors is good enough to capture the various cytotoxic scaffolds, and the regression models can capture general trends instead of class effects.
Data availability and the need of structural reliability informed the selection of used compounds of 156. There is a paucity of cytotoxic drugs whose structure is well validated and a good number of compounds mentioned in the literature lack full or standardized digital structure to be able to compute descriptors. Rather than artificially expanding the dataset with uncertain or low-quality structures which could distort the regression analysis, we opted for a curated and chemically rigorous set of 156 molecules. This dataset size is consistent with previous topological-descriptor and QSAR studies where similar sample sizes have been used to benchmark regression performance and evaluate descriptor relevance.
Although the dataset is moderate in size, the study focuses on comparing preprocessing strategies and regularized regression methods, which are explicitly designed to handle high-dimensional descriptor spaces and smaller sample sizes. The aim is not to build a final deployable predictive model but to understand how modeling choices influence the predictability of Top_PSA when using graph-theoretical descriptors. Therefore, the chosen dataset (Fig. 1) is appropriate for a controlled methodological investigation and provides interpretable results within the scope of this work.
Dataset of 156 cytotoxic compounds.
Methods
In order to explore the effect of preprocessing, multicollinearity control, and model complexity on predicting the Topology Polar Surface Area (Top PSA), five different modeling schemes (Scheme #1- Scheme #5) were constructed and tested. Each scheme used identical curated set of 156 cytotoxic compounds and had a common training testing protocol to compare fairly.
In all schemes, k-fold cross-validation was rigorously used in the training of the model and in the selection of the features to ensure that overfitting was minimized and the predictive performance was also estimated strongly. The coefficient of determination (\(R^2\)) and mean squared error (MSE) were used to perform model evaluation on an independent test set.
Regression linear models
In this study, we have used three models to predict the topological polar surface area. Linear regression is a supervised machine learning algorithm that can be used to predict a numerical output. The loss function in the case of linear regression is
Here, Y is our matrix given for our target variable, \(\hat{Y}\) is a column matrix giving us the prediction. Linear regression is a regression model that tries to capture the relationship between independent variables and dependent variables using a linear equation. Let’s suppose we have the linear combination mentioned below:
Each \(w_i\) is the weight associated with the respective feature. Here, \(w_0\) is the intercept term. The weights for which the loss function gives the minimum value are used in the linear equation to predict the output. So, when we compute all the weights (including intercept), we can obtain an output for a particular input values of features. In our current problem, we initially have 57 independent features, and our goal is to predict the topological polar surface area. The linear equation becomes
Lasso regression is also a regression model that tries to find the best possible hyperplane (in higher dimensions) that can capture the relationship between input and output features. Here, a penalty (\(L^{1}\) norm) is added to the loss function. The loss function in the case of Lasso regression is
\(\alpha\) is the hyperparameter. w denotes the associated weights.
Ridge regression is another model that is being implemented here. It is also a regularized model that adds a penalty term to the loss function. But this penalty is known as the \(L^2\) norm.
The loss function mentioned in Lasso and ridge regression can be solved for a particular set of weights, such that the loss function gives the minimum value. In ridge and lasso regression, the \(\alpha\) hyperparameter basically helps in dealing with overfitting and underfitting phenomena as well. The lower the value of \(\alpha\), the higher the tendency of our model to overfit. In case of a higher value of \(\alpha\), the model will be exposed to underfitting. In our discussion, in subsequent sections, alpha’s value is 1, which is set in the sklearn library by default. For ridge regression ’cholesky’ solver has been used. The split ratio for our dataset is 80% for the training set and 20% for the testing set. That split has been done with a random state of 42.
Direct models fitting
In this scheme, all the features have been provided to the models. And after training the models on these features, we get the weights associated with each feature as mentioned in Fig. 2. We can see that the regularized models have more shrunk weights as compared to linear regression.
Weights assigned by each model to each of the features. The intercepts for linear regression, lasso, and ridge turned out to be -54.44, 12.17, and 28.82, respectively.
Linear regression gave the highest positive weight to the feature “Constitu_tional_1” while lasso and ridge gave the same highest positive weight to ’Atom_Count_4’. The Fig. 3 presents a comparative evaluation of three regression models i.e., linear regression, Lasso regression, and ridge regression using two key performance metrics: R2 Score and mean squared error (MSE). The R2 Score, shown in the top portion of the figure, reflects the proportion of variance in the target variable explained by each model. Among the three, ridge regression achieves the highest R2 score, indicating superior predictive capability, followed closely by Lasso, with linear regression performing the least effectively. The bottom portion of the figure illustrates the mean squared error, where lower values indicate better performance. Again, ridge regression yields the lowest MSE, suggesting more accurate predictions, while linear regression shows the highest MSE, reflecting greater average prediction error. Overall, the results clearly demonstrate that ridge regression provides the best balance of bias and variance for the dataset, outperforming both Lasso and standard linear regression in terms of both explained variance and prediction accuracy. The mean squared error for linear regression, lasso, and ridge turned out to be 3091.88, 148.40, and 623.80, respectively. The mean squared error was minimum for the ridge regression.
\(R^2\) scores and mean squared errors after applying models to the data directly.
Table 2 shows the regression equations of three regression models i.e., linear regression, ridge regression and Lasso regression. These equations describe the outcome \(\hat{y}\) as a linear combination of the predictor variables so that the coefficients of the predictor variables are interpreted to reflect the anticipated change in \(\hat{y}\) with all the other variables held constant. Linear Regression model incorporates all predictors without regularization with the members being given weights irrespective of their significance. Ridge Regression: Ridge Regression uses L2 regularization to shrink the coefficients, which are brought to zero, to minimize overfitting, but does not reduce the number of variables in the model. LASSO Regression employs L1 regularization, which is allowed to shrink certain coefficients to zero thereby performing variable selection and only the most significant predictors remaining. As a result, although the three models are all predicated on the same predictors, the coefficients in the models vary in magnitude and existence, which indicates the sensitivity of regularization on complexity and relevance of features in a model.
Linear models with principal component analysis
Principal component analysis has been used in this setup. First, the data has been scaled using standard scaling, the formula for which is given below:
Here \(X^{\prime }\) is the scaled feature, X is the original feature, and \(\mu\) and \(\sigma\) represent the mean and standard deviation of X, respectively. In this scheme, PCA has been applied to the features after standard scaling, and the dimensionality has been reduced to 10 features. When we analyze the number of components and the cumulative sum of variance explained by components, we see that when we use 10 components, it explains 91.16% of the variance. This relation has been depicted in Fig. 4, where a threshold of 90 % has also been mentioned.
Number of principal components vs percentage of explained variance.
The Fig. 5 illustrates the distribution of model coefficients (weights) across the first ten principal components (PCA0 to PCA9) for three regression methods: linear regression, Lasso, and ridge. The weight of a given principle component on a prediction is equal to the number of each bar and therefore gives a bar with its effect on the prediction. The existence of the relatively broad range of values of the weight, either of which is strongly positive or strongly negative in nature, is testified by the fact that the linear regression is responsive to all the elements when it is not regularized. The Lasso regression that favors sparsity that assigns weights that are non-positive or close to non-positive to a few components, i.e. the Lasso regression only employs the most informative components. The Ridge regression distributes the components that have moderate values more evenly as compared to the other models and thus, it is unlikely to shrink the coefficients and end up with all the features. Such comparison can show how various regularization methods influence the explainability of the model and the significance of the components of data transformed using PCA. The lasso and ridge intercepts obtained after the fit are equal to the results of linear regression (152.26).
Weights of features upon model fitting after applying standard scaling and principal component analysis.
Table 3 shows the regression equation of principal component analysis (PCA).
The Fig. 6 presents a performance comparison of three regression models-linear regression, Lasso, and ridge-applied to data transformed using principal component analysis (PCA). Two metrics are shown: the R2 Score (top panel) and the mean squared error (MSE, bottom panel). The R2 Score measures how well each model explains the variance in the target variable, with higher values indicating better performance.
The Fig. 6 of the paper provides the comparison of performance between three regression models i.e., linear regression, Lasso and ridge on the data transformed with the help of principal component analysis (PCA). Two figures are presented: the R2 Score (top panel) as well as the mean squared error (MSE, bottom panel). The R2 Score is a measure of how the model accounts the variance in the target variable with a high score meaning the model does a good job. Among the models, ridge regression achieves the highest R2 score, followed by Lasso and then linear regression, suggesting that ridge provides the best fit to the PCA-transformed data. The MSE, which quantifies prediction error, shows an inverse pattern: ridge has the lowest error, linear regression slightly higher, and lasso the highest. These results demonstrate that, even in a reduced-dimensionality space created by PCA, ridge regression maintains superior predictive performance in terms of both accuracy and explanatory power.
\(R^2\) scores and mean squared error after standard scaling followed by principal component analysis.
Model fitting with robust scaling
In this scheme, the input features have been scaled using a robust scaling technique. It is considered robust to outliers. The formula for robust scaling is given below:
Here \(X^{\prime }\) will be the new scaled feature, \(X_0\) will be the unscaled feature of the data. \(X_{median}\) will be the median of particular feature X. IQR will be the interquartile range that is the difference of \(75^{th}\) percentile and \(25^{th}\) percentile. The scaled training data’s distribution has been given in Fig. 7. When the models were trained on this scaled data, the weights of features that were obtained are given in Fig. 8. linear regression, lasso, and ridge gave the highest positive weights to ’Atom_Count_1’. The intercepts were 125.95, 133.16, and 131.69 for linear regression, lasso, and ridge, respectively.
Distributions of training data post robust scaling.
Weights assigned by models to each feature after robust scaling.
The Fig. 9 compares the performance of three regression models-linear regression, Lasso, and ridge-using two common evaluation metrics: R2 score and mean squared error (MSE). The R2 score, which reflects the proportion of variance in the dependent variable explained by the model, is shown along the horizontal axis in the top portion of the figure. All three models exhibit similar R2 values, indicating comparable abilities to capture the underlying data variance. In contrast, the bottom portion of the figure presents the MSE values, where a clear difference in predictive accuracy emerges. Ridge regression achieves the lowest MSE, followed by Lasso, while linear regression shows the highest error. This suggests that although the models are similarly effective in explaining variance, ridge regression offers superior generalization performance by minimizing prediction error more effectively. We can see that Lasso’s mean squared error was very small compared to other models. Lasso also showed the highest R-squared score in this scheme as compared to other models.
\(R^2\) scores and mean squared error upon model fitting after robust scaling of feature.s
Regression model without outliers
In this scheme, the experimentation of the adjustment of outliers has been done.
Methodology adopted to adjust outliers of skewed distributions
The distributions of the features were checked for skewness with the criteria that the features having skewness greater than 1 or less than -1 were termed as skewed features. For skewed features, we can check their boxplots to find the outliers. So, the features whose skewness turned out to be greater than +1 or less than -1, their boxplots have been shown in Fig. 10.
Among the skewed features, the features that had values greater than \(Q3+1.5*IQR\) were assigned the value of \(Q3+1.5*IQR\), and the features having values less than \(Q1-1.5*IQR\) were assigned the value of \(Q1-1.5*IQR\). Here, Q1 and Q3 are the first and third quartiles, respectively. IQR is the interquartile range, which is the difference between the \(3^{rd}\) quartile and the \(1^{st}\) quartile. After the adjustment of outliers, the boxplots of the skewed features have been given in Fig. 11.
Boxplots of features having skewed distributions.
Boxplots of skewed distributions after adjustment of outliers.
Methodology adopted to adjust outliers of other non-skewed distributions
For the features that had skewness greater than -1 and less than +1, the scheme of Winsorization scheme has been used. The distributions of non-skewed distributions have been presented in Fig. 12. In this scheme, a threshold (5%) has been decided such that an upper bound has been fixed, like if there is any value in a feature greater than the \(95^{th}\) percentile, the value will be adjusted to the \(95^{th}\) percentile. And if the value is less than the \(5^{th}\) percentile, the value will be adjusted to \(5^{th}\) percentile. The distributions after the adjustment have been given in Fig. 13.
Distributions of non-skewed features.
Non-skewed distributions after adjustment of outliers.
Model fitting and evaluation
The Fig. 14 contains the weights assigned to each feature by models. Lasso and ridge regression gave most positive weight to ’Z1’ and ’Information_Content_4’ respectively, while linear regression gave most positive weight to ’Frame_work’. The intercepts of linear regression, lasso, and ridge obtained are 4.45, 1.75, and 0.31, respectively.
Weights assigned to features by respective models after adjustment of outliers.
This Fig. 15 also illustrates the comparative performance of three regression models-linear regression, Lasso, and ridge-based on two evaluation metrics: the R2 score and mean squared error (MSE). The R2 score, shown in the top part of the figure along the horizontal axis, measures the proportion of variance in the target variable that is explained by the model. All three models display relatively similar R2 values, indicating that they capture the underlying data patterns to a comparable extent. In contrast, the lower part of the figure presents the MSE values for each model. The ridge regression model achieves the lowest MSE, suggesting it makes the most accurate predictions among the three. Lasso follows with slightly higher error, while linear regression exhibits the highest MSE. This indicates that, despite their comparable ability to explain variance, ridge and Lasso models offer improved prediction accuracy, likely due to their regularization techniques that help reduce overfitting. MSE for linear regression, lasso, and ridge turned out to be 0.33, 0.18, and 0.24, respectively.
\(R^2\) scores and mean squared error of models after outliers adjustment.
Regression model with variance inflation factor
Multicollinearity is a problem in which our independent variables can have a high correlation with each other. So, in this scheme, the variance inflation factor has been used to detect the ability of our independent variables to predict each other. What happens here is that one of the independent variables becomes the target variable, and the other independent variables try to predict it. To be precise, a linear regression is used to see if an independent variable can be predicted by a linear combination of other independent variables or not. Variance inflation factor is given by
Variance inflation factor of our independent features on a logarithmic scale has been given in Fig. 16. We calculated the variance inflation factor for our variables and shortlisted a few features that had a variance inflation factor of less than 10. Generally, a threshold of 5 or 10 is chosen. We picked a threshold of 10, and the features that we have got are in Table 4.
Variance inflation factor of features on logarithmic scale.
Figure 17 displays the feature weights assigned by three different regression models-linear regression, lasso, and ridge-to various input variables in the context of a dataset involving molecular descriptors. Each bar represents the coefficient (weight) associated with a specific feature, such as Wiener, Acid Base, Lipinski, and others, across the three modeling approaches.
In linear regression, the weights vary widely, including both large positive and negative values, reflecting its sensitivity to multicollinearity and the lack of regularization. lasso, which incorporates L1 regularization, sets several feature weights exactly to zero, effectively performing variable selection by excluding less relevant features. ridge, which applies L2 regularization, shrinks all coefficients but retains all features with reduced magnitude, striking a balance between coefficient size and model complexity.
This visualization underscores how regularization impacts model interpretability and robustness: Lasso offers a sparse solution that aids in feature selection, while ridge maintains all variables but mitigates overfitting through weight shrinkage.
Weights assigned by models after selecting features based on variance inflation threshold. Corresponding intercepts using linear regression, lasso and ridge came out to be 6822.73, 6808.45, and 6805.48, respectively.
The Fig. 18 presents a comparative assessment of three regression models-linear regression, lasso, and ridge-based on two key performance metrics: R2 score and mean squared error (MSE). The upper portion of the figure displays the R2 scores, which indicate how well each model explains the variance in the dependent variable. The R2 values are relatively similar across the three models, suggesting that they perform comparably in terms of capturing the underlying data structure, and they show that the model’s \(R^2\) scores are not good with these features. The lower portion of the figure shows the MSE values, which quantify the average squared difference between predicted and actual values. Here, a significant difference is evident: ridge regression yields the lowest MSE, followed by Lasso, while linear regression has the highest MSE and shows a drastic increase. This indicates that while all models explain a similar proportion of the variance, ridge and lasso-due to their regularization-provide more accurate and stable predictions, especially in the presence of multicollinearity.
\(R^2\) scores and mean squared error upon selecting features based on variance inflation factor.
Model performance evaluation using cross-validation
Table 5 presents the cross-validated performance of linear regression, ridge regression, and LASSO regression models for Top_PSA prediction, evaluated using multiple statistical metrics, including the coefficient of determination (\(R^2\)), mean squared error (MSE), root mean squared error (RMSE), and mean absolute error (MAE). All reported values represent mean scores across 10-fold cross-validation, ensuring robust and unbiased performance estimation.
Among the evaluated models, LASSO regression achieved the highest predictive performance, yielding the largest mean R2 value (0.93) and the lowest MSE (155.66) and RMSE (11.05). This indicates that the sparsity-inducing L1 regularization effectively balances model complexity and generalization while selecting chemically relevant descriptors. The slightly higher MAE observed for LASSO compared to ridge suggests that while overall prediction accuracy is high, ridge regression may provide marginally more stable average error behavior. Ridge regression also performed strongly, with a mean \(R^2\) of 0.92 and lower error metrics than ordinary linear regression. This improvement highlights the benefit of L2 regularization in mitigating multicollinearity and stabilizing coefficient estimates in descriptor-rich QSAR models. In contrast, linear regression, which lacks regularization, exhibited the lowest \(R^2\) and the highest error values, underscoring its sensitivity to correlated and heterogeneously scaled descriptors.
Overall, these results demonstrate that regularized regression models outperform unregularized linear regression for Top_PSA prediction, with LASSO offering the best trade-off between predictive accuracy and model interpretability. The superior performance of LASSO further supports its suitability for descriptor-driven QSAR modeling, where feature selection and robustness are critical.
Performance of nonlinear machine learning models
In order to evaluate the existence of extra predictive benefits of nonlinear learning algorithms as compared to linear and regularized regression models, a number of nonlinear machine learning algorithms were also tested, and they are decision trees, k-nearest neighbor (k-NN), support vector regression (SVR), random forest, gradient boosting, and a voting regressor ensemble. Table 6 is a summary of the results of the tree-based ensemble techniques, specifically random forest and gradient boosting, which performed the best in generalization, with the test-set \(R^2\) squared of 0.94 and 0.93, respectively. Single decision trees on the other hand had indications of overfitting and the SVR had low predictive ability given the existing set of descriptors. These findings suggest that nonlinear models have the ability to represent complicated structure-property dynamics in the data of Top_PSA; nevertheless, these models are more complex and less interpretable, and this led the researcher to rely primarily on linear and regularized regression models. The nonlinear performances thus, act as their supplementary checkpoints validating the strength and competitiveness of the suggested linear modeling structure.
Results and discussion
The comparison of the R² scores of linear regression, LASSO and ridges models under preprocessing strategies (direct fitting, PCA, robust scaling, outlier adjustment and K-fold Cross Validation) is given in Fig. 19. At these combinations of preprocessing models and models, clear patterns of performance are observed and these patterns illuminate not just the behavior of the methods but also indicate the chemical relevance of new descriptors.
Model performance comparison.
Model performance
In all the types of models, the best performance was always in robust scaling, which attained an R² of 0.97 using the LASSO regression model. It means that strategies of preprocessing which reduce extreme values and eliminate heteroscedasticity help model generalization significantly. The idea of robust scaling seems especially useful when the set of descriptors has skewed distributions, such as those found in graph-theoretical indices, since then the effect of unusual values of the descriptors, which tends to cause a skewed least squares and ridge estimates, is diminished.
PCA also worked well particularly in the LASSO and the ridge models indicating that dimensionality reduction can be used successfully to handle multicollinearity and noise but preserve most of the variance in the descriptor matrix. PCA models were, however, a little less accurate than robust-scaled models, which is a trade-off between interpretability and performance of using PCA: despite PCA generating a compact representation, it hides the original chemical meaning of the original descriptors in the transformed components.
In the case of immediate fitting strategy, the performance improved linear to ridge regression with the ridge attaining an R² of 0.94. The significance of regularization is indicated by this trend when dealing with descriptor sets that have correlated features and different scales. Penalty in Ridge regression stabilizes the coefficient estimates when there is multicollinearity and it provides superior generalization compared to unregularized linear regression.
In contrast, outlier adjustment showed a high level of variability. Although fairly decent with LASSO \((R^{2} \approx 0.91)\) it degraded significantly in ridge \((R^{2} \approx 0.76)\) and less so in linear regression \((R^{2} \approx 0.41)\). This implies that outlier removal can only be useful when sparse regularization is combined with outlier removal; otherwise, outlier removal can make the reduced dataset more unstable or wipe out informative chemical variation.
Selection of feature using variance inflation factor (VIF) was mostly linked with inferior predictive performance in comparison to other preprocessing approaches. It would imply that loss of features based on multicollinearity only might remove descriptors that represent chemically meaningful polar surface data. In QSAR studies with very many descriptors, correlation between the features does not always indicate redundancy and in practice the regularization based methods are much more flexible and effective in dealing with multicollinearity than hard-threshold feature pruning.
The benchmarks of nonlinear models were used to estimate the possible gains over and above linear regression. The random forest and gradient boosting tree-based models demonstrated good generalization behavior (\((R^{2} \textit{test} \approx 0.93-0.94\)), and decision trees and SVR did not, probably because these methods are hyperparameter sensitive and a high dimensional descriptor space. These more complex and less interpretable nonlinear methods were only compared, but interpretable regularized and linear models were primarily used in the study.
Descriptor importance and chemical interpretation
Examination of non-zero LASSO coefficients indicates obvious trends in which the significance of descriptors is associated with the chemical drivers of Top_PSA. In all folds of cross-validation, LASSO was always able to keep descriptors that encode heteroatom content (e.g., numbers of N, O, S atoms), hydrogen-bond donor and acceptor capacity and electronegativity-weighted topological indices. These characteristics are directly associated with the chemical determinants of polar surface area controlled by the polar atoms and functional groups number, identity, and spatial exposure.
Another choice of ring-centric descriptors by LASSO was to include heteroatom-substituted aromatic systems, which is an indication that the distribution of polar fragments over ring structures is a significant contributor to calculated Top_PSA. Conversely, descriptors related mainly to hydrocarbon topology like adjacency indices, carbon-centric Zagreb indices and branching descriptors had coefficients that were close to zero. Their subduction supports the chemical anticipation that changes in the skeleton of hydrocarbons do not significantly affect polar surface area relative to those of the heteroatomic functional groups.
Therefore, regularized models do not only provide strong predictions but also provide chemically intuitive results which are in agreement with the fragment-based Top_PSA algorithm created by Ertl and colleagues.
Connections to prior QSAR and TOP_PSA literature
The obtained results are consistent with the known Top_PSA theory and serve different contributions to the current work on QSAR. Previous studies have been done to determine the usefulness of Top_PSA as a predictor of permeability and ADME-practice32,33 and most recently the behaviour of individual descriptors families has been explorated by descriptor-comparison research45. Nonetheless, contrary to that, the present studies provide a methodological study of how preprocessing strategies impact the model performance and interpretability, which are insufficiently reported in the QSAR processes, however, that have a significant impact on the stability of the coefficients and the relevance of the features.
We have discovered that robust scaling and LASSO are especially useful as a combination in cases where we possess skewed or correlated variables in a collection of descriptors- a phenomenon that is frequently seen with graph-theoretical descriptor matrices. Meanwhile, PCA is more robust regarding the level of its performance regarding dimensionality reduction, whereas it may be less interpretable. The application of VIF-based features reduction, as it was commonly applied, proved harmful to the modeling of Top_PSA, which states that naive collinearity-based pruning can possibly drop down the descriptors describing significant polar-surface features.
Limitations and future directions
RDKit and Mordred were used to compute the Top_PSA labels in this study43,44. Computational labels may be used, although they are not a replacement of experimentally found PSA values, whereas they are appropriate to methodological benchmarking. Future studies should consider the possibility that the recommendations found in future studies apply to larger datasets, other QSAR endpoints, and models trained with experimental PSAs measurements.
The 156 cytotoxic compounds curated dataset afforded adequate chemical diversity to study interactions with preprocessing-model, but further and more diverse datasets will be necessary to construct highly generalisable predictive models. Also, one application of this analysis to non-linear model families (e.g., random forests, gradient boosting machines, neural networks) can potentially reveal such interactions not modeled by linear models, but interpretability concerns will become of greater concern. Though the dataset includes compounds related to various cancer types and is not evenly distributed across therapeutic classes, their disproportion does not have a direct impact on the current analysis since the endpoint of interest, Topological Polar Surface Area (Top_PSA) is an intrinsic property of a molecule that does not have biological or disease-specific connotation. Therefore, the predictive modeling structure is neither particularly therapeutically indicated. However, future research can build on this study by adding stratified analyses of clinically active compounds by classes or therapeutic targets, especially in the process of simulating biologically-relevant endpoints such as cytotoxic activity or response to treatment.
Practical recommendations
The practical implications that this research has on the QSAR professionals are a number based on the synthesized results of this research. Where the task is on the use of descriptor matrices having extreme observations or correlated variables, robust scaling with regularized regression, including LASSO or ridge, are the most preferable and should therefore be prioritized. PCA is a friendly fallback, in which dimensionality reduction or noise compression is required but the loss of interpretability in the individual descriptors is then revealed to result in the requirement to adopt loading analyses were a mechanistic understanding of the resultant descriptors to be sought. On the other hand, a strict VIF-based pruning should not be used in the high-descriptor QSAR workflows, where it may remove the orthogonal features, which may have a potentially significant predictive value by chance. Finally, all the computational information surrounding the Top_PSA generation including the toolkit, the version and implementation parameters should be clearly stated by the researchers to ensure the reproducibility and comparability of modeling studies. Taken together, these recommendations are helpful in the best practices of predictive models of QSAR that are based on descriptors and offer sensible guidance towards the establishment of predictive models based on moderate-sized curated chemical data.
Conclusion
This study demonstrates the effectiveness of using topological and molecular descriptors combined with machine learning models to predict the Topological Polar Surface Area (Top_PSA) of cytotoxic compounds. It also tested several modeling methods, such as linear regression, lasso, and ridge regression, as well as preprocessing methods, such as principal component analysis (PCA), robust scaling, and outlier adjustments. Findings show that the regularized regression models especially lasso regression showed constant improvement over the simple linear regression in all the preprocessing schemes. Lasso regression using robust scaling performed best in terms of predictive accuracy \((R^{2} \approx 0.97)\) and the most chemically interpretable feature selection, whereas ridge regression excelled in initial direct model fitting. PCA was good at both dimensionality reduction and predictive performance, outlier adjustments were found to be greatly beneficial to model performance. Though feature selection based on variance inflation factor (VIF) was used to reduce multicollinearity, the accuracy of the model decreased, which suggests that the trade-off between predictive power and interpretability exists. These findings are also substantiated by cross-validated error measures, in which LASSO regression had the highest mean of the \(R^{2}\) and lowest errors of the predictions compared to any other tested regression model. On the whole, this study highlights the importance of integrating domain-specific descriptions with powerful machine learning pipelines to make effective predictions of the properties of molecules, which can be used in drug development and design. This work should be extended to more extensive and varied chemical datasets, and experimentally-determined PSA values should be used in future research, as well as non-linear machine-learning approaches that can describe more complicated structure-property relationships.
Data availability
The dataset used and/or analysed during the current study, python code files and some additional information related to this manuscript are available in our public GitHub repository at https://github.com/Shabbir-Ahmad-1/A-Quantitative-Study-of-Cytotoxic-Compounds.
References
Hu, X. et al. Predicting herb disease associations through graph convolutional network. Curr. Bioinform. 18, 610–619 (2023).
Zhou, R. et al. NEDD: A network embedding based method for predicting drug disease associations. BMC Bioinform. 21(13), 387 (2020).
Meng, Y. et al. Drug repositioning based on weighted local information augmented graph neural network. Brief. Bioinform. 25(1), bbad431 (2023).
Sheng, Y., Wang, J., Liu, S. & Jiang, Y. IMN4NPD: An integrated molecular networking workflow for natural product dereplication. Anal. Chem. 96(7), 2990–2997 (2024).
Feng, X. et al. AutoFE-Pointer: Auto-weighted feature extractor based on pointer network for DNA methylation prediction. Int. J. Biol. Macromol. 311, 143668 (2025).
Zheng, W. et al. GEP-DNN4Mol: Automatic chemical molecular design based on deep neural networks and gene expression programming. Health Inf. Sci. Syst. 13(1), 31 (2025).
He, W. et al. Neuromorphic-enabled video-activated cell sorting. Nat. Commun. 15(1), 10792 (2024).
Zhang, H., Ren, Y., Xia, Y., Zhou, S. & Guan, J. Towards effective causal partitioning by edge cutting of adjoint graph. IEEE Trans. Pattern Anal. Mach. Intell. 46(12), 10259–10271 (2024).
Shi, S. & Liu, W. B2-ViT Net: Broad vision transformer network with broad attention for seizure prediction. IEEE Trans. Neural Syst. Rehabil. Eng. 32, 178–188 (2024).
Jiang, C. et al. Xanthohumol inhibits TGF-\(\beta\)1-induced cardiac fibroblasts activation via mediating PTEN/Akt/mTOR signaling pathway. Drug Des. Dev. Ther. 14, 5431–5439 (2020).
Lan, Z. et al. Curcumin-primed olfactory mucosa-derived mesenchymal stem cells mitigate cerebral ischemia/reperfusion injury-induced neuronal PANoptosis by modulating microglial polarization. Phytomedicine 129, 155635 (2024).
Wang, J. et al. SIRT6 protects against lipopolysaccharide-induced inflammation in human pulmonary lung microvascular endothelial cells. Inflammation 47(1), 323–332 (2024).
P. Ronan O’Connell et al. Bailey & Love’s Short Practice of Surgery, CRC Press; 27th edition. (2018).
I. Gutman et al. Novel Molecular Structure Descriptors - Theory and Applications II, MCM Vol. 9, University of Kragujevac, Kragujevac, pp. 139–168 (2010).
Nilakantan, Ramaswamy et al. A amily of ring system-based structural fragments for use in structure activity studies: Database mining and recursive partitioning. J. Chem. Inf. Model. 46(3), 1069–1077 (2006).
Guy, W. B. & Mark A. M. The properties of known drugs. 1. molecular frameworks. J. Med. Chem. 39(15), 2887–2893 (1996).
Dahmani, Rahma, Manachou, Marwa, Belaidi, Salah, Chtita, Samir & Boughdiri, Salima. Structural characterization and QSAR modeling of 1,2,4 triazole derivatives as glucosidase inhibitors. New J. Chem. 45, 1253–1262 (2021).
Magnuson et al. Studies in Physical and Theoretical Chemistry, pp. 178–191 (Elsevier, 1983).
Kobayashi, et al. Prediction of soil adsorption coefficient in pesticides using physicochemical properties and molecular descriptors by machine learning models. Environ. Toxicol. Chem. 39(7), 1451–1459 (2020).
Abraham, M. H. & McGowan, J. C. The use of characteristic volumes to measure cavity terms in reversed phase liquid chromatography. Chromatographia 23, 243–246 (1987).
Labute, P. Derivation and applications of molecular descriptors based on approximate surface area. Methods Mol. Biol. 275, 261–278 (2004).
Todeschini, R. & Consonni, V. Handbook of Molecular Descriptors (John Wiley & Sons, 2008).
Estrada, E., Torres, L., Rodriguez, L. & Gutman, I. An atom-bond connectivity index: Modelling the enthalpy of formation of alkanes, Indian. J. Chem. 37A, 849–855 (1998).
Gao, W., Wang, W. F., Jamil, M. K., Farooq, R. & Farahani, M. R. Generalized atom-bond connectivity analysis of several chemical molecular graphs. Bulg. Chem. Commun. 48(3), 543–549 (2016).
Iqbal, Z., Ishaq, M. & Farooq, R. Computing different versions of atom-bond connectivity index of dendrimers. J. Inform. Math. Sci. 9(1), 217–229 (2017).
Shao, Z., Wu, P., Zhang, X., Dimitrov, D. & Liu, J. B. On the maximum ABC index of graphs with prescribed size and without pendent vertices. IEEE Access 6, 27604–27616 (2018).
Shao, Z., Wu, P., Gao, Y., Gutman, I. & Zhang, X. On the maximum ABC index of graphs without pendent vertices. Appl. Math. Comput. 315, 298–312 (2017).
Choudhary, S., Ranjan, P. & Chakraborty, T. Atomic polarizability: A periodic descriptor. J. Chem. Res. 44(3–4), 227–234 (2020).
Lipkus, Alan H. Exploring chemical rings in a simple topological-descriptor space. J. Chem. Inf. Comp. Sci. 41(2), 430–438 (2001).
Gerta, R. & Christoph, R. Counts of all walks as atomic and molecular descriptors. J. Chem. Inf. Comp. Sci. 33(5), 683–695 (1993).
Todeschini, R. & Consonni, V. Handbook of Molecular Descriptors, Wiley-VCH, Methods and Principles in Medicinal Chemistry. Volume 11 (2000).
Ertl, P., Rohde, B. & Selzer, P. Fast calculation of molecular polar surface area as a sum of fragment-based contributions and its application to the prediction of drug transport properties. J. Med. Chem. 43(20), 3714–3717 (2000).
Prasanna, S. & Doerksen, R. J. Topological polar surface area: A useful descriptor in 2D-QSAR. Curr. Top. Med. Chem. 16(1), 21–41 (2009).
Mauri, A. alvaDesc: A tool to calculate and analyze molecular descriptors and fingerprints. Methods Pharmacol. Toxicol. 32, 801–820 (2020).
Wildman, Scott A. & Crippen, Gordon M. Prediction of physicochemical parameters by atomic contributions. J. Chem. Inf. Comp. Sci. 39(5), 868–873 (1999).
Veber, Daniel F. et al. Molecular properties that influence the oral bioavailability of drug candidates. J. Med. Chem. 45(12), 2615–2623 (2002).
Das, K. C. & Gutman, I. Some properties of the second Zagreb index. MATCH Commun. Math. Comput. Chem. 52(1), 3–11 (2004).
Furtula, B., Gutman, I. & Ediz, S. On difference of Zagreb indices. Discrete Appl. Math. 178, 83–88 (2014).
Vukičević, D. & Furtula, B. Topological index based on the ratios of geometrical and arithmetical means of end-vertex degrees of edges. J. Math. Chem. 46(4), 1369–1376 (2009).
Randić, M. & Zupan, J. On interpretation of well-known topological indices. J. Chem. Inf. Comp. Sci. 41(3), 550–560 (2001).
Todeschini, R., Consonni, V., Ballabio, D. & Grisoni, F. Comprehensive Chemometrics (Chemometrics for QSAR Modeling), pp. 599–634 (2020).
Pölsterl, S. & Wachinger, C. Likelihood-Free Inference and Generation of Molecular Graphs (2019).
Moriwaki, H., Tian, Y.-S., Kawashita, N. & Takagi, T. Mordred: A molecular descriptor calculator. J. Cheminformatics 10, 4 (2018).
Landrum, G. (2006–present). RDKit: Open-source cheminformatics; rdMolDescriptors TPSA implementation. Available at: https://www.rdkit.org/
Lange, J. J. et al. Comparative analysis of chemical descriptors by machine learning methods. J. Chem. Inf. Model. 21(7), 3343–3355 (2024).
Funding
There is no funding to support this article.
Author information
Authors and Affiliations
Contributions
Shabbir Ahmad provided assistance with the data curation analysis, experiment design, and inquiry. In addition to providing computational support, Sana Javed reviewed and approved the final version of the manuscript. Sadia Khalid assists with computation, data analysis, funding resources, calculation validation, and enhancements to Matlab and Maple graphs. Muhammad Kamran Siddiqui helped with project management and conceptualisation in addition to writing the article’s initial draft. The approach, oversight, and resource collection were handled by Hassan Aftab. Brima Gegbe worked on software development, validation, funding acquisition, and formal data analysis. All authors read and approved the final draft.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ahmad, S., Javed, S., Khalid, S. et al. A quantitative study of cytotoxic compounds using graph based descriptors and machine learning. Sci Rep 16, 5076 (2026). https://doi.org/10.1038/s41598-026-35728-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-35728-7