Abstract
Accurate classification of brain tumors from Magnetic Resonance Imaging (MRI) images remains a significant technical challenge in medical image analysis. Recent advancements have primarily focused on developing automated image classification methods. However, traditional convolutional neural networks (CNNs) have limited feature extraction capabilities, leading to suboptimal recognition performance. To address this issue, this paper proposes a novel image classification framework named multi-scale channel attention CNN integrated with support vector machine (MCACNN-SVM). In the proposed MCACNN-SVM, hierarchical spatial features are extracted with multi-scale convolutional kernels, and then further adaptively enhanced by adopting the channel attention mechanism. Finally, an SVM classifier optimized by grid search algorithm is employed to optimize the decision boundaries and enhance classification accuracy. Furthermore, the cosine annealing with warm restarts strategy is adopted to accelerate convergence and improve generalization. Extensive experiments on the brain tumor MRI dataset demonstrate that the proposed framework achieves competitive performance in accuracy, precision, recall, and F1-score, exhibiting strong robustness and generalization ability for practical applications.
Similar content being viewed by others
Introduction
Accurate and automated classification of brain tumors from Magnetic Resonance Imaging (MRI) images presents a significant challenge in medical image analysis1,2. The complex anatomical structure of the brain and the considerable inter-subject variability render manual diagnosis time-consuming and prone to subjectivity3,4. While deep learning has shown promise in automating this task, models based on traditional convolutional neural networks (CNNs) often have limitations in capturing comprehensive features and constructing optimal decision boundaries5. Therefore, developing advanced computer-aided diagnostic frameworks that are both robust and highly accurate is of paramount importance for practical applications.
MRI has emerged as a primary modality for brain tumor characterization owing to its superior soft-tissue contrast. The precise analysis of MRI data now serves as a cornerstone for brain tumor classification. Driven by continuous developments in medical imaging, substantial research efforts have been directed toward automating the classification process to enhance diagnostic accuracy and reproducibility6. Early approaches frequently relied on traditional machine learning techniques. For example, Helen et al.7 implemented an automated preprocessing and feature extraction method combined with a Random Forest (RF) classifier for binary classification on the BraTS 2015 dataset, markedly improving the differentiation between benign and malignant tumors through block-wise features. Similarly, Soltaninejad et al.8 confirmed the effectiveness of RF in feature selection tasks, reporting an average classification accuracy of \(80.86\%\).
The evolution of computer-aided diagnosis (CAD) systems has incorporated a wider range of classifiers and methodologies. Sachdeva et al.9 developed an integrated framework that combined image segmentation, feature extraction, and multi-class classification, achieving \(85\%\) overall accuracy using artificial neural networks for six tumor types. Support Vector Machines (SVMs) have also demonstrated strong performance. Dandil et al.10 achieved a classification accuracy of \(91.05\%\) , with a sensitivity of \(90.8\%\) and specificity of \(94.7\%\) in medical imaging applications. Recognizing that feature extraction is pivotal for model performance, Gumaei et al.11 introduced a hybrid method based on a Regularized Extreme Learning Machine (RELM), while Paul et al.12 merged fully connected networks with Convolutional Neural Networks (CNNs) to devise a generalized classification technique, attaining an accuracy of \(91.43\%\). Other specialized approaches, such as the tumor resection technique by Hamamci et al.13 which processes multiple MRI modalities (T1, T2, T1-Gd, FLAIR) separately, and the voxel-level SVM classification utilizing intensity and spatial features by Havaei et al.14, further enriched the methodological landscape.
More recently, deep learning approaches have dominated the field, offering significant improvements in accuracy15. CNN-based models achieved \(97\%\) classification accuracy on the BraTS 2013 dataset16, and subsequent studies with data augmentation reported accuracies of \(95\%\) and \(98\%\) on other datasets17. Sliding window-based CNNs and other deep architectures have been widely employed for medical image classification, demonstrating notable advantages in both accuracy and stability over traditional machine learning methods. Despite their success, these techniques are often hampered by inherent limitations, including high model complexity, substantial computational demands, long training times, and a propensity for overfitting. For instance, Deepak et al.18 employed transfer learning with data augmentation to enhance model generalization, while Swati et al.19 demonstrated the effectiveness of fine-tuning pre-trained networks for brain tumor classification. Similarly, Díaz-Pernas et al.20 developed a multi-scale CNN architecture that captures discriminative features at different resolutions, achieving robust performance on MRI datasets.
Convolutional Neural Networks, while representing a cornerstone technology in image recognition and achieving remarkable success in classification tasks, are consequently often characterized by these operational drawbacks21,22,23. In parallel, Support Vector Machines (SVMs) have been extensively applied across various image processing tasks-including image recognition, object detection, and segmentation-due to their strong theoretical foundations and effectiveness in high-dimensional spaces24. Nevertheless, SVMs are also associated with challenges such as high computational complexity, poor scalability, and limited efficacy in handling non-linear data without appropriate kernel choices.
Motivated by the above discussion, this paper proposes a novel hybrid model named multi-scale channel attention CNN integrated with support vector machine (MCACNN-SVM)) for brain tumor image classification. This model leverages the powerful image feature extraction capability of MCACNN and the efficient classification performance of SVM, fully utilizing the strengths of both techniques. This multi-scale fusion approach provides a more efficient and flexible solution for brain tumor image classification, laying a foundation for further improving classification accuracy and robustness. The main contributions of this paper are summarized as follows: 1) A hybrid model named MCACNN-SVM is proposed for brain tumor MRI image classification, which integrates a multi-scale convolutional neural network with a support vector machine. The MCACNN extracts rich spatial features from the MRI images using convolution kernels of different scales, while the SVM effectively classifies the features, improving the model’s performance. 2) The introduction of a channel attention mechanism (SENet) optimizes the feature extraction process by enhancing the importance of critical feature channels, thus improving the robustness and classification accuracy of the model. 3) A comprehensive optimization strategy is implemented to enhance model performance: a cosine annealing with warm restarts strategy dynamically adjusts the learning rate to facilitate convergence and generalization, while the hyperparameters of the SVM classifier are systematically optimized via grid search with cross-validation to ensure the formation of optimal decision boundaries. 4) Extensive experiments on the Brain Tumor MRI Dataset demonstrate that the MCACNN-SVM model outperforms existing state-of-the-art methods in accuracy, recall, precision, and F1-score, confirming its high robustness and practical value for clinical diagnostic applications.
The remainder of this paper is organized as follows: Section 2 introduces the fundamental theories related to image classification algorithms, including the multi-scale convolutional layers, channel attention mechanism and support vector. Section 3 describes the model framework and image classification process. Section 4 analyses the experimental results and compares the performance of the proposed model with other leading methods. Finally, Section 5 concludes the paper and discusses potential future directions for enhancing the model’s efficiency and exploring its broader application in other medical image classification tasks.
Basic theories related to image classification algorithms
Convolutional neural network
Traditional CNN networks are fully convolutional networks . CNNs are well-known for their excellent image feature learning capabilities, as they rely on fully connected convolutional structures to extract feature vectors and classify targets25. Their multi-layer structure automatically learns hierarchical features of the data, from edges and textures to more complex patterns26,27. Moreover, due to the characteristics of parameter sharing and local connections, CNNs generally require fewer training parameters than fully connected networks for image recognition tasks, reducing the risk of overfitting and improving computational efficiency28. However, traditional CNNs also have limitations. When the network structure becomes deeper, issues such as vanishing or exploding gradients may occur during training, making the model difficult to train. Additionally, traditional CNNs may be limited when processing very large images or capturing global contextual information due to the restricted receptive field of their convolution kernels.
Channel attention mechanism
The attention mechanism works by generating and assigning weights to modulate the network parameters. After training, the network focuses on the key features, thus optimizing the learning capability of the CNN.
To improve feature discriminability in convolutional neural networks, where different channels often contribute unequally to task performance, this work incorporates and refines the channel attention mechanism established in29. This mechanism, known as SENet (Squeeze-and-Excitation Network), enhances representational power by explicitly modeling inter-channel relationships and adaptively emphasizing informative features. Its fundamental building block, the SE block, performs sequential squeezing and excitation operations to dynamically recalibrate channel-wise responses, as illustrated in Fig. 1.
Schematic diagram of SENet.
Brain tumor images typically exhibit complex textural and structural features. The channel attention mechanism can effectively assist the network in extracting and focusing on important feature channels. By enhancing the weights of critical features, it accelerates the extraction of key characteristics in brain tumor images while suppressing less relevant features, thereby reducing computational load and significantly improving recognition and classification performance. Furthermore, compared to other types of attention mechanisms, the channel attention mechanism offers lower computational complexity, making it easier to implement and train within networks. Thus, employing a channel attention mechanism represents a rational and efficient choice for brain tumor image recognition and classification tasks.
In a standard Convolutional Neural Network (CNN), Each channel processes information within its confined receptive area, lacking the capacity to incorporate contextual cues from neighboring channels. To address this limitation, the Squeeze network first extracts a feature map X through convolutional operations, then compresses it using global average pooling to capture global information z. The c element of z, denoted as \(z_c\), is calculated as follows.
where \(u_c\) denotes the c-th channel of the feature map X, while H and W represent its height and width, respectively. Following the Excitation step, the interdependencies between channels are modeled to produce the channel-wise weight coefficient s, computed as follows.
where \(\delta\) denotes the ReLU activation function, \(\sigma\) represents the Sigmoid function, and r is the reduction ratio (set to 16 in this study). The output feature map X is subsequently obtained by Eq.(3) .
where \(s_c\) corresponds to the c-th element of s.
Support vector machine
Support Vector Machines (SVMs) are a class of generalized linear classifiers that perform binary classification on data using supervised learning30,31. Their decision boundary is the maximum margin hyperplane derived from the training samples. Compared to neural networks, SVMs offer a clearer and more powerful approach when learning complex nonlinear equations. The core idea of the SVM algorithm is to identify a subset of data points located on the margins, known as support vectors, and use these points to determine a plane, referred to as the decision boundary, such that the distance from the support vectors to this plane is maximized
Schematic diagram of support vector machine.
Any hyperplane can be described by the following linear equation:
Where w is the weight matrix and b is the bias term.
The distance formula from a point (x, y) in two-dimensional space to a line \(Ax+By+C=0\) is as follows.
When extended to n-dimensional space, the distance from a point \(\textbf{x} = (x_1, x_2, \ldots , x_n)\) to the hyperplane \(\textbf{w}^T \textbf{x} + b = 0\) is given by Eq.(6).
Where \(\Vert w\Vert = \sqrt{w_1^2 + \ldots + w_n^2}\)
As illustrated, according to the definition of support vectors, it is known that the distance from the support vectors to the hyperplane is d, while the distance from other points to the hyperplane is strictly greater than d. Thus, the two parallel hyperplanes above and below the maximum-margin hyperplane can be obtained as shown in Fig. 2.
The proposed image classification framework of MCACNN-SVM
MCACNN-SVM model framework
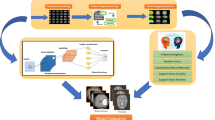
In order to overcome the limitations of traditional CNN feature extraction capabilities, this paper proposes the MCACNN-SVM model, with the overall framework shown in Fig. 3. The model integrates multi-scale convolutional neural networks with different scales to enhance the perception of information at varying scales within an image. In the MCACNN architecture, the traditional CNN network structure is carefully adjusted and extended to meet the specific needs of brain MRI image classification. By introducing multi-scale convolution kernels and pooling windows, the model captures richer image features, improving the network’s ability to recognize edges, textures, and more complex patterns. In addition, SVM serves as the final classifier responsible for optimizing the classification of high-level features extracted by multi-scale channel attention CNN (MCACNN). SVM achieves decision boundary partitioning in feature space by constructing the optimal hyperplane, which has the advantage of maintaining strong generalization ability when processing high-dimensional feature data. The synergy between MCACNN’s nuanced feature extraction and SVM’s discriminative classification creates a powerful framework for accurate image analysis.
The framework of MCACNN-SVM for image classification.
In the proposed MCACNN-SVM hybrid model, through this parallel design, the multi-scale convolutional network layers can simultaneously extract features at various scales and fuse them together. This enhances the model’s ability to perceive multi-scale features, thereby improving its understanding and classification accuracy of brain tumor images. To extract fine-grained spatial features from MRI images and enhance spatial attention information, three parallel 2D convolutional pathways are designed, as illustrated in Fig. 3. Each pathway consists of three sequential layers: a Convolutional layer (Conv), a Batch Normalization layer (BN), and a Max Pooling layer (MaxPool). The original single-scale convolution kernels are replaced with \(3\times 3\), \(5\times 5\), and \(7\times 7\) kernels to capture features at different scales. After the three different-scale convolutions, a \(1\times 1\) convolution is applied to concatenate the inputs from the three scales and then compress the channels. This design enables the model to capture more comprehensive details and structural features in brain tumor images.
This parallel structure ensures that features extracted from convolution kernels of different scales can be learned and updated independently, thereby avoiding potential interference among multi-scale features. The outputs from multiple branches are merged through concatenation and then compressed with a \(1\times 1\) convolution, transforming the complex nonlinear features of brain tumors into a more linear feature space. Although the softmax layer in CNNs performs well for classifying linearly separable data, it is less effective in handling complex decision boundaries. In contrast, SVM offers a more distinct separation of the feature space. Unlike fully connected layers with softmax, SVM leverages maximum-margin separation to establish a more robust decision boundary in the high-dimensional feature space extracted by deep networks, thereby enhancing classification stability and generalization. Therefore, in this model, CNN is employed for feature extraction, and classification is ultimately performed by SVM, with the integration carried out at the feature level.
The improved MCACNN network, by introducing multi-scale convolution and pooling operations, is better equipped to capture multi-scale information in images. This enhances the network’s ability to perceive information at different scales, thereby improving the accuracy and robustness of image classification tasks. With the incorporation of multi-scale convolutions and pooling operations, the network gains a more comprehensive understanding of image features and can effectively process information at various scales. The key model parameters are summarized in Table 1.
Image classification process of MCACNN-SVM model
The full flowchart of the MCACNN-SVM model for MRI image classification is shown in Fig. 4.
The overall pipeline comprises three main stages: data partitioning, model training, and diagnostic classification. Initially, the dataset is partitioned into training, validation, and test subsets. The model is then initialized and trained on the training set, with hyperparameters iteratively optimized using the validation set. Upon reaching a predefined maximum number of iterations, the optimal model is automatically saved. Lastly, the trained model is utilized for performing classification and diagnostic tasks on the test set.
MCACNN-SVM model image classification process.
Experimental results and analysis
The experiments were conducted using the Keras deep learning framework with Python 3.7. The hardware environment included an Intel i7-13700KF CPU and a Colorful GeForce RTX 4060 Ti GPU with 16 GB VRAM, accelerated by the CUDA 10.1 parallel computing architecture on a Windows 11 operating system.
Experimental dataset and data preprocessing
To evaluate the effectiveness of the proposed MCACNN-SVM model, we employed the publicly available Brain Tumor MRI Dataset32. This dataset, also hosted on Kaggle, comprises a total of 7,023 T1-weighted contrast-enhanced magnetic resonance imaging (CE-MRI) slices. The images were collected from real clinical cases in Chinese hospitals and have been rigorously preprocessed, including anonymization, skull-stripping, and alignment, to ensure quality and consistency33,34. The dataset is categorized into four classes:glioma, meningioma , pituitary tumor , and no tumor, providing a approximately balanced multi-class classification task. The detailed distribution of the dataset for training and testing is summarized in Table 2.
Additionally, to enhance the model’s generalization capability and mitigate overfitting, a comprehensive data preprocessing and augmentation pipeline was applied exclusively to the training set. Each image was first normalized to a fixed size of \(224 \times 224\) pixels. Pixel values were then rescaled to the range [0, 1] by dividing by 255. During training, the following real-time data augmentation techniques were employed using the Keras ImageDataGenerator:
-
Random Rotation A range of \(\pm 15^\circ\).
-
Random Horizontal Flip With a probability of 0.5.
-
Random Zoom A zoom range of \(\pm 10\%\).
-
Grayscale Normalization Performed per-image by standardizing to zero mean and unit variance.
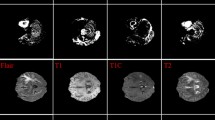
MRI images of four types of brain tumors. (a) Glioma, (b) Meningioma, (c) Pituitary, (d) No tumor.
The validation and test sets were only subjected to resizing and normalization, without any augmentation, to ensure a fair evaluation. Figure 5 shows example MRI images for each of the four tumor categories in the brain tumor MRI dataset.
Model performance metrics
In this paper, a four-class performance evaluation was conducted on the MCACNN-SVM hybrid model developed based on the brain tumor MRI dataset. The performance metrics used include precision, recall, accuracy, F1-score, as well as macro average and micro average.
Accuracy refers to the proportion of correctly predicted samples to the total number of samples. The formula is as follows.
Recall is the proportion of all actually positive samples that are correctly identified as positive. The formula is as follows.
Precision is the proportion of samples that are actually positive among all the samples predicted as positive by the model. The formula is as follows.
The F1-score is the harmonic mean of precision and recall. It aims to balance the two metrics, and is particularly meaningful in cases of imbalanced data. The formula is as follows
where TP, FP, TN, and FN represent true positive, false positive, true negative, and false negative, respectively. These metrics are derived by comparing the model’s predictions with the ground truth labels.
In addition, this paper introduces two summary metrics: macro average and micro average to provide a more comprehensive evaluation of the multi-class model’s overall performance. Due to the weak class balance present in the data, macro-average was specifically adopted as the model performance evaluation metric.
Macro average is the simple average of metrics, such as precision, recall, F1-score for each class, assigning equal weight to each class without considering the differences in the number of samples across classes. Macro average emphasizes the balanced performance of the model across classes and is suitable for performance comparison in imbalanced class situations. The calculation equations for the Macro average of Recall , Precision and F1 are shown as follows.
where \(P_i\) is the precision of category i, \(R_i\) is the Recall of category i, \(F1_i = 2 \cdot \frac{P_i \cdot R_i}{P_i + R_i}\) is the F1 value of category i, and C is the total number of categories.
Micro average is calculated by accumulating the true positives, false positives, and false negatives for all classes and then computing the metrics. It gives equal weight to all samples, and is more reflective of the model’s overall classification ability across the entire dataset. The calculation equations for the Micro average of Recall , Precision and F1 are shown in Eq.(14) , Eq.(15) and Eq.(16), respectively.
Model parameter settings
The configuration of hyperparameters, such as learning rate, optimizer, and batch size-critically influences model training outcomes. In particular, the learning rate and optimizer settings significantly affect convergence behavior and overall performance, ultimately determining the effectiveness of the training procedure. Therefore, meticulous tuning of these parameters is essential.
Optimizer and learning rate parameters
Based on the Stochastic Gradient Descent with Warm Restarts (SGDR) algorithm35, which combines the SGD optimizer with a warm restart mechanism, significant improvements have been achieved in multiple classification tasks. Taking inspiration from the above, this paper further improves the performance of the MCACNN-SVM model by applying cosine annealing and warm restart strateg to the training process. The core idea of the warm restart mechanism is to set a periodic cycle during training. once this cycle is completed, the learning rate is reset to its initial value and training continues. Moreover, after each restart, the model uses the parameters from the previous restart as its initial state. This mechanism essentially constitutes a method for dynamically adjusting the learning rate.
The accuracy of the model varies with the number of iterations under different configurations of the optimizer and learning rate schedulers. Notably, the warm restart strategy combined with cosine annealing cyclically adjusts the learning rate, beginning from a low value, rising to a peak, and then decaying gradually following a cosine schedule. After each annealing cycle, the warm restart mechanism resets the learning rate to its initial value and begins a new annealing process. The purpose of this strategy is to prevent the model from being trapped in local optima by allowing the periodically increased learning rate to escape from local minima, thereby improving convergence performance. The learning rate update formula based on warm restarts mechanism for each iteration is shown in Eq.(17).
Among these, i denotes the number of iterations, \(\eta _{\max }^{i}\) and \(\eta _{\min }^{i}\) represent the upper and lower bounds of the learning rate adjustment, espectively, \(T_{cur}\) indicates the number of epoch changes during the interval from the start to the end of each restart, and \(T_{i}\) stands for the restart cycle. The learning rate adaptively adjusts according to the number of iterations, as illustrated in Fig. 6.
The learning rate variation curve of Cosineannealing with WarmRestart mechanism.
To facilitate rapid training of the MCACNN-SVM model based on the warm restart mechanism and further improve its performance, the initial cycle a is typically set to a relatively small value36. Therefore, the maximum initial learning rate for the cosine annealing with warm restart learning rate scheduler is set at \(\eta _{\max }^{i}\) = 1e-2, and the minimum initial learning rate is set at \(\eta _{\min }^{i}\) = 1e-5 in this paper. Before each subsequent restart, it is multiplied by a scaling factor b to adjust the restart cycle c. This dynamic adjustment of the restart cycle enables the model to explore different local minima, efficiently converge to the optimal solution, and reduce training costs.
Different warm restarts mechanism parameter settings have a significant impact on the performance of the network model. Additionally, the number of epochs required for convergence varies depending on the configuration. To find the best configuration that promotes model convergence while minimizing training costs, an ablation study was conducted to analyze the effect of different warm restarts mechanism settings on model performance.
As shown in Table 3, although the training cost is the lowest under the configuration \(T_0 =20, T_{mult} =1\), its classification error rate on the MRI image dataset is not the best. In contrast, with the \(T_0 =10, T_{mult} =2\) parameter setting, the training cost increases by approximately 5 epochs, but the lowest test error rate of only \(2.06\%\) is achieved on the same dataset. Therefore, this configuration demonstrates a better balance between training cost and model performance. Based on this observation, all subsequent experiments in this study adopt the \(T_0 =10, T_{mult} =2\) configuration for model training.
Additionally, in this paper, the impact of five commonly used optimizers and learning rate combinations on model accuracy was evaluated, and the average results of five independent experiments are shown in Fig. 7. These configurations encompass: the classical stochastic gradient descent (SGD), famous Adam optimizer, SGD implemented with a cosine annealing learning rate scheduler incorporating warm restarts (SGDR), the Adam optimizer augmented with a warm restart learning rate scheduling mechanism, and the AdamW optimizer integrated with a similar warm restart learning rate adjustment protocol. Among them, the learning rates of SGD and Adam optimizers are fixed at 1e-3. AdamW adds weight decay on the basis of Adam, improving the problem of incompatibility between regularization and adaptive learning rates.
The impact curve of different optimizer and learning rate scheduler combinations on model performance.
AdamW with Cosine Annealing Warm Restarts learning rate scheduler demonstrated high training and validation accuracy in this experiment, especially achieving an accuracy of \(97.94\%\) on the validation set. This indicates that AdamW with Cosine Annealing Warm Restarts mechanism performs well in preventing overfitting and accelerating model convergence. Considering that AdamW with Cosine Annealing Warm Restarts periodically increases the learning rate, warm restarts can help escape these mundane local optima and continue to search for better solutions, potentially achieving better performance and generalization ability. Although the Adam optimizer is adaptive and converges quickly, it occasionally settles into suboptimal local minima. Therefore, when facing complex image classification tasks, it is appropriate to choose AdamW optimizer with the Warm Restarts learning rate scheduler.
Support vector machine parameters
To optimize the performance of the SVM classifier, this research employs GridSearchCV algorithm combined with 5-fold cross-validation to systematically adjust its key hyperparameters. Based on the theoretical foundation and practical experience of SVM, a search space for three core hyperparameters was established. Regularization parameter C: set values include 1, 10, 100, used to control model complexity and overfitting penalty strength. Kernel function parameter gamma: set values include 0.1, 0.01, 0.001, affecting the influence range of samples in the Radial Basis Function (RBF) kernel. Kernel function type: evaluating three commonly used kernel functions, including radial basis function (RBF), linear, and sigmoid, in order to adapt to diverse feature distribution characteristics.
The SVM hyperparameter optimization process is as follows: Firstly, the trained MCACNN model is used to extract feature representations from the training set as inputs for SVM. After initializing the SVM classifier, the GridSearchCV algorithm is used to traverse all hyperparameter combinations, with a total of 27 configurations of 3 \(\times\) 3 \(\times\) 3. Use 5-fold cross validation to evaluate the performance of each set of parameters, with classification accuracy as the evaluation metric, to ensure the robustness of parameter selection. The hyperparameter combination that performs the best on the validation set is ultimately selected to construct the final SVM model.
After the above hyperparameter optimization verification experiments, the SVM parameters ultimately used in this study are: [’C’=1, ’gamma’=0.01, ’kernel’=rbf]. This study significantly improved the classification accuracy of the hybrid model on the test set, verifying the effectiveness of parameter optimization. Through the above systematic hyperparameter tuning, SVM can fully utilize the discriminative features extracted by CNN, achieving more accurate brain tumor MRI image classification, while avoiding the subjectivity of manual parameter selection.
Analysis of model performance and visualization
Analysis of the model convergence process
Batch normalization is integrated into the MCACNN-SVM model to accelerate the training process and enhance generalization capability. By normalizing the input data, batch normalization effectively enhances the training speed, stability, and generalization performance of the model.
Model convergence process. (a) Accuracy curve, (b) Loss curve.
To evaluate the convergence behavior of the model across both training and validation sets, a 100-epoch experiment was carried out to enhance performance and prevent underfitting or overfitting. Following several refinements, the convergence behavior of the MCACNN-SVM model is depicted in Figure 8. The results reveal a steady decline in loss values for both sets as iterations proceed. Around epoch 70, the training and validation loss values stabilize, suggesting successful convergence of both loss and accuracy curves.
Visualization of the model training process
This paper applied t-SNE to visualize the training dynamics of the model. The technique projects high-dimensional data into a two-dimensional space, offering an intuitive depiction of feature distribution. Figure 9 illustrates the evolution of feature representations before and after training.
Visualization of the Model Training Process. (a) Original data feature distribution, (b) Channel attention layer feature distribution, (c) Fusion layer feature distribution, (d) Output layer feature distribution.
The initial feature distribution of the test set, illustrated in Fig. 9a, shows complete overlap among the four classes, indicating no separability at the input stage. After processing by the channel attention mechanism, the features exhibit reduced inter-class separation, as visible in Fig. 9b. Subsequent feature maps following the fusion layer, shown in Fig. 9c , reveal initial clustering tendencies, though with limited margin between classes. As the network deepens, the model progressively learns more discriminative feature representations, ultimately forming well-separated class boundaries in the output layer which is shown in Fig. 9d. Notably, the final features demonstrate large inter-class margins and compact intra-class clustering, confirming the ability of the MCACNN-SVM model to achieve effective and robust MRI image recognition.
Visualization of model cassification results
Fig. 10 presents the confusion matrix for the four-class classification results of the MSCNN-SVM model on the test set. The numbers on the matrix diagonal represent the correctly classified samples, while those off the diagonal indicate the number of misclassified samples. As shown, the classifier used performs well in classifying the sample data.
Confusion matrix for MRI image classification.
Table 4 provides the detailed performance of the MCACNN-SVM model on the four-class brain tumor dataset. Clearly, the MCACNN-SVM model achieves a classification accuracy of \(96.9\%\) across all test samples. This demonstrates that the model has strong generalization ability in distinguishing different types of brain tumors and can accurately differentiate normal tissue from various tumor types. it is worth noting that compared to other categories, the recall rate for the category of meningioma is \(90.79\%\) , which is slightly lower. This performance gap may be partially attributed to its relatively smaller sample size in the training dataset.
ROC curves for MRI image classification.
Figure 11 illustrates the ROC curves for the four-class classification task, where the model demonstrates outstanding ROC-AUC performance. The AUC values for all categories exceed 0.99, with no tumor, pituitary, as well as the micro- and macro-averages achieving an AUC of 1.00, indicating virtually no errors in distinguishing these classes. The ROC curves are almost adjacent to the top-left corner of the figure, suggesting that the model possesses very strong classification capability. Although there is a slight decline in recall for the specific class of meningioma, the model generally performs at a high level across most categories, with particularly excellent performance in classifying no tumor and pituitary. Both the macro-average and micro-average AUC values reach 1.0, demonstrating that the model achieves an ideal balance between overall and class-specific performance. The ROC curves further confirm that the MCACNN-SVM model exhibits exceptional performance in the classification of brain tumor MRI images.
Although the slight class imbalance present in the dataset, the model maintains robust performance across most categories. This resilience can be attributed to two key factors: first, the incorporation of the channel attention mechanism (SENet) enables the model to adaptively focus on the most discriminative features for each class, effectively mitigating the bias towards majority classes; second, the adoption of macro-averaged evaluation metrics such as Macro F1-score and AUC ensures that each class contributes equally to the overall performance assessment, providing a more balanced view of the model’s capability.
Comparative analysis with other image classification methods
In order to comprehensively evaluate the performance of the proposed model algorithm, a comparative analysis of the MACACNN-SVM model with several well-established and high-performance modern CNN architectures in the field of computer vision, including the classic deep network VGG16 proposed by Simonyan et al.38, ResNet18 proposed by He et al. to solve the gradient vanishing problem39, DenseNet121 proposed by Huang et al. with dense connections40, and EfficientNet proposed by Tan et al. to achieve high parameter efficiency through composite scaling41. These pre trained model weights were loaded and fine tuned. In addition, MSCNN16 which is the multi-scale CNN structures applied to brain tumor classification were also compared.
Specifically, Pre trained weights are used and fine tuned for fair experimentation. Ensure that all comparison models are fine tuned and tested under identical training conditions such as data augmentation, optimizer, learning rate strategy, and training cycle. Specifically,the feature maps are first converted into single-channel images in MSCNN, and then features are extracted using convolutional kernels of sizes 3\(\times\)3, 7\(\times\)7, and 11\(\times\)11.
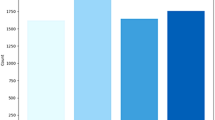
In order to accurately evaluate the performance of the MCACNN-SVM fusion model in brain tumor MRI image classification tasks, this paper conducted ten fold cross validation evaluations on the comparative models, and the experimental average results are shown in Table 5. The experiment compared the accuracy, precision, recall, F1 score and the number of parameters of the algorithm separately. The testing accuracy of DenseNet121 model is the lowest, with an mean value of\(91.2\%\), indicating its limited feature representation capability. This performance gap may stem from its relatively shallow depth in this specific task or suboptimal adaptation to the dataset’s characteristics. MSCNN, with larger convolutional kernels and pooling sizes, not only increases the computational complexity of the model but also leads to a rapid decrease in the spatial resolution of the feature maps, causing some loss of image detail37. Although the model achieves an accuracy of \(96.68\%\) , its recall rate is only \(83.4\%\). The VGG16, ResNet18 and EfficientNet-B3 models have slightly improved performance, but their parameters are relatively the largest.
The performance metrics distribution diagram of different comparison models. (a) Accuracy distribution diagram, (b) Precision distribution diagram, (c) Recall distribution diagram, (d) F1 value distribution diagram.
Compared with other models, The evaluation metrics and the number of parameters of the MCACNN-SVM model are all optimal, as this hybrid model combines the powerful feature extraction capability of the multiscale CNN with the efficient classification performance of SVM, further enhanced by the introduction of a channel attention mechanism to improve feature representation. The model extracts rich image features through parallel multiscale convolutional layers, while the channel attention mechanism dynamically adjusts the weights of different channels to emphasize key features and suppress redundant information. Subsequently, SVM performs efficient classification decisions using these optimized features. This combined model significantly improves performance, extracting image features more precisely while maintaining computational efficiency, which is particularly crucial for the accurate classification of brain tumors.
Further examination of the accuracy distribution of each comparative model is shown in Fig. 12a , where the red dots represent the mean values. The results indicate that the median and mean of DenseNet121 are nearly identical, suggesting a relatively symmetric distribution of experimental results with good overall stability, although its accuracy level is comparatively lower. For MSCNN, the mean is slightly higher than the median, implying a slight right-skewed distribution in which a few higher accuracy values pull the mean upward. The VGG16 model demonstrates stronger performance with a mean accuracy near 97.0%, benefiting from its deep hierarchical architecture. however, its relatively large variance indicates sensitivity to data distribution and potential overfitting. In contrast, the performance distribution of ResNet18 is more concentrated, reflecting greater robustness. The EfficientNet model slighty outperforms the others, achieving a median accuracy close to 98.0% with a tight interquartile range, highlighting its superior efficiency and robustness in extracting discriminative features across multiple scales. Notably, the median and mean of MCACNN-SVM almost completely overlap, demonstrating both high accuracy and optimal stability. This performance can be primarily attributed to the integration of an SVM classifier with multi-scale feature extraction, which enhances both feature representation and classification robustness. Therefore, MCACNN-SVM can be considered superior to the other models in terms of both accuracy and stability. Moreover, Fig. 12b, c, and d present the distributions of Precision, Recall, and F1 for the five comparative models, respectively. In comparison, MCACNN-SVM exhibits consistent results with higher values across all performance metrics, further validating its generalization and effectiveness.
Ablation study
To verify the contributions of each component in the proposed model, an ablation study was conducted on each module of the model architecture. Explore the multi-scale CNN feature extraction module, channel attention mechanism, and SVM of the model for classifying brain tumor MRI images separately. The algorithm in the MACNN-SVM model that removes the convolutional layers of the 5 \(\times\) 5 and 7 \(\times\) 7 parallel branches and only retains one branch of the 3 \(\times\) 3 convolutional layer architecture is denoted as Baseline, and the algorithm that removes the channel attention mechanism module and SVM module in the MACNN-SVM model is denoted as MACNN-SVM without Multi-scale (w./o. Multi-scale). The algorithm that removes the SENet module in the MACNN-SVM model is denoted as MACNN-SVM without SENet (w./o. SENet), while the model with the SVM replaced by a fully connected layer with Softmax activation is denoted as MCACNN-SVM without SVM (w./o. SVM). In order to ensure the fairness of the experiment, the experimental settings for all four groups used identical configurations. Each component was analyzed to determine its impact on the overall performance, and the experimental results are shown in Table 6.
As shown in Table 6, the removal of Multi-scale CNN module resulted in a comprehensive decline across all performance metrics. Specifically, the accuracy decreased by \(2.44\%\), and other performance indicators also decreased synchronously. This indicates that the multi-scale CNN fusion architecture can effectively capture discriminative features at varying receptive field, which plays a key role in improving the model’s ability to distinguish diverse fault modes. Excluding the SENet module caused a \(1.07\%\) decrease in accuracy, underscoring the importance of channel-wise attention in adaptively recalibrating feature responses and enhancing inter-channel dependencies. Furthermore, removing the SVM resulted in a marginal accuracy decrease of \(0.91\%\), but it led to relatively larger reductions in precision, recall, and F1-score. This indicates SVM’s crucial role in enhancing classification robustness, particularly in handling ambiguously classified cases near decision boundaries. These findings confirm that SVM provides more reliable decision boundaries in the high-dimensional feature space, which is critical for clinical applications where both false positives and false negatives carry serious consequences. In the end, our complete model achieved the best performance in all evaluation metrics, verifying the effectiveness of the collaborative design of multi-scale feature extraction, attention mechanism, and SVM classifier. This architecture not only achieves multi-scale and adaptive enhancement at the feature representation level, but also further optimizes the partitioning of the feature space in the classification decision stage, thereby exhibiting higher comprehensive performance and robustness in fault diagnosis tasks.
Conclusions
This paper proposes a hybrid MCACNN-SVM model by comprehensively analyzing the strengths and limitations of deep learning techniques in brain tumor MRI image classification. The model first employs a multiscale CNN fusion strategy to capture richer image features, and then applies SVM for classification, with its key hyperparameters tuned using a grid search algorithm to improve performance. The CNN component is responsible for automatically extracting informative features, while the SVM focuses on learning decision boundaries, achieving a clear task separation that facilitates both optimization and interpretability. The model was optimized using AdamW, which decouples weight decay from gradient updates, leading to improved generalization. The learning rate was scheduled using a cosine annealing strategy with warm restarts. Experimental results show that the MCACNN-SVM model performs excellently in terms of accuracy, AUC, and confusion matrix, confirming that the CNN-SVM combination indeed enhances overall performance. The findings demonstrate the effectiveness of the MCACNN-SVM model, addressing issues such as limited data and overfitting that may arise in single-model approaches. By integrating the advantages of different architectures, multiscale fusion provides medical professionals with a more accurate and comprehensive diagnostic tool.
Despite the promising potential of multiscale fusion in brain tumor MRI image classification, several challenges remain, including computational complexity, the choice of fusion strategies, and large-scale data processing. Furthermore, while the model demonstrates robust performance using metrics that are less sensitive to class imbalance, the inherent slight class distribution differences in the datasets as observed in the current dataset, especially between glioma and no tumor categories, warrant further investigation. Additionally, The model was trained and evaluated solely on high-quality, preprocessed contrast-enhanced MRI slices from a single source, which may compromise its robustness on low-quality, noisy, or unprocessed clinical scans. Moreover, the consistent imaging protocol limits our understanding of its sensitivity to variations in MRI scanners, field strengths, or acquisition sequences. While the framework achieves strong performance across four brain tumor categories, its generalizability to other tumor types or broader medical imaging tasksremains unverified.
Future work will prioritize lightweight model design without sacrificing accuracy, multi-center and multi-protocol data collection, domain adaptation, and cross-disease transfer learning. To ensure uniformly high sensitivity across all tumor types, this work explores advanced class-imbalance learning techniques, such as weighted loss functions, data resampling, or focal loss, to explicitly address this issue. In addition, Deeper algorithmic optimization coupled with clinical feedback is expected to enhance both accuracy and real-world usability, ultimately advancing the model’s value in clinical practice.
Data availability
The datasets used and/or analyzed during the current study available from the corresponding author on reasonable request.
References
Sultan, H. H., Salem, N. M. & Al-Atabany, W. Multi-stage brain tumor classification using MRI based on deep convolutional neural networks. Appl. Sci.13, 4161 (2023).
Lilhore, U. K. et al. AG-MS3D-CNN multiscale attention guided 3D convolutional neural network for robust brain tumor segmentation across MRI protocols. Sci Rep15, 24306 (2025).
Rehman, A., Saba, T., Rahman, S. I. U., Al-Turjman, F. & Alabrah, A. Brain tumor classification from MRI images using a convolutional neural network with a modified ReLU activation function. Diagnostics12, 532 (2022).
Zhong, W., Huang, Z. Z. & Tang, X. A study of brain MRI characteristics and clinical features in 76 cases of Wilson’s disease. J. Clin. Neurosci.59, 167–174 (2018).
Latif, G., Brahim, G. B., Iskandar, D. A., Bashar, A. & Alghazo, J. M. Glioma tumors’ classification using deep-neural-network-based features with SVM classifier. Diagnostics12, 1018 (2022).
Ranjbarzadeh, R. et al. Brain tumor segmentation based on deep learning and an attention mechanism using MRI multi-modalities brain images. Sci. Rep.11, 10930 (2021).
Helen, R. & Kamaraj, N. CAD scheme to detect brain tumour in MR images using active contour models and tree classifiers. J. Electr. Eng. Technol.10, 670–675 (2015).
Soltaninejad, M. et al. Supervised learning based multimodal MRI brain tumour segmentation using texture features from supervoxels. Comput. Methods Progr Biomed.157, 69–84 (2018).
Sachdeva, J., Kumar, V., Gupta, I., Khandelwal, N. & Ahuja, C. K. Segmentation, feature extraction, and multiclass brain tumor classification. J. Digit. Imaging26, 1141–1150 (2013).
Dandıl, E., Çakıroğlu, M. & Ekşi, Z. Computer-aided diagnosis of malign and benign brain tumors on MR images. In Proceedings of the International Conference on ICT Innovations, 157-166 (Springer, Berlin/Heidelberg, 2014).
Gumaei, A., Hassan, M. M., Hassan, M. R., Alelaiwi, A. & Fortino, G. A hybrid feature extraction method with regularized extreme learning machine for brain tumor classification. IEEE Access7, 36266–36273 (2019).
Paul, J. S., Plassard, A. J., Landman, B. A. & Fabbri, D. Deep learning for brain tumor classification. In Proceedings of the Medical Imaging 2017: Biomedical Applications in Molecular, Structural, and Functional Imaging (SPIE), 253-268 (Orlando, FL, USA, 2017).
Hamamci, A., Kucuk, N., Karaman, K., Engin, K. & Unal, G. Tumor-cut: Segmentation of brain tumors on contrast enhanced MR images for radiosurgery applications. IEEE Trans. Med. Imaging31, 790–804 (2011).
Havaei, M., Larochelle, H., Poulin, P. & Jodoin, P. M. Within-brain classification for brain tumor segmentation. Int. J. Comput. Assist. Radiol. Surg.11, 777–788 (2016).
Yang, K. et al. SiamCorners: Siamese corner networks for visual tracking. IEEE Trans. Multimed.24, 1956–1967 (2022).
Díaz-Pernas, F. J., Martínez-Zarzuela, M., Antón-Rodríguez, M. & González-Ortega, D. A deep learning approach for brain tumor classification and segmentation using a multiscale convolutional neural network. Healthcare9, 153 (2021).
Ayadi, W., Elhamzi, W., Charfi, I. & Atri, M. Deep CNN for brain tumor classification. Neural Process. Lett.53, 671–700 (2021).
Deepak, S. & Ameer, P. M. Brain tumor classification using deep CNN features via transfer learning. Computers10, 18 (2021).
Swati, Z. N. K. et al. Brain tumor classification for MR images using transfer learning and fine-tuning. Comput. Med. Imaging Graph.75, 34–46 (2019).
Díaz-Pernas, F. J., Martínez-Zarzuela, M., Antón-Rodríguez, M. & González-Ortega, D. A deep learning approach for brain tumor classification and segmentation using a multiscale convolutional neural network. Healthcare9, 153 (2021).
Yang, Y., Gao, S., Ke, L. & Liu, X. Attention-edge-assisted neural HDRI based on registered extreme-exposure-ratio images. Symmetry17, 1381 (2025).
Ke, L., Liu, Y. & Yang, Y. Compound fault diagnosis method of modular multilevel converter based on improved capsule network. IEEE Access10, 41201–41214 (2022).
Yang, K. et al. DeforT: Deformable transformer for visual tracking. Neural Netw.24, 106380 (2024).
Wu, Z., Li, C., Li, H., Li, C. & Liu, Y. Intelligent fault diagnosis of rolling bearings using a SVM-based deep stacking network. Sensors22, 1943 (2022).
Wang, L., Lin, Z. Q. & Wong, A. COVID-Net: A tailored deep convolutional neural network design for detection of COVID-19 cases from chest X-ray images. Sci Rep10, 19549 (2020).
Wang, Z. et al. Meta-learning based prototype-relation network for few-shot image classification. Neurocomputing560, 126809 (2023).
Too, E. C., Yujian, L., Njuki, S. & Yingchun, L. A comparative study of fine-tuning deep learning models for plant disease identification. Comput. Electron. Agric.161, 105272 (2021).
Ke, L., Hu, G., Yang, Y. & Liu, Y. Fault diagnosis for modular multilevel converter switching devices via multimodal attention fusion. IEEE Access11, 135035–135048 (2023).
Hu, J., Li, S., Samuel, A. & Sun, G. Squeeze-and-excitation networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 7132-7141 (Salt Lake City, UT, USA, 2018).
Wang, Y. et al. A novel fault diagnosis method of rolling bearings based on lightweight neural network and support vector machine. Adv. Eng. Inform.62, 102448 (2024).
Chen, X. et al. Fusing vision transformer and support vector machine for few-shot brain tumor classification. Comput. Biol. Med.165, 107234 (2023).
Cheng, J. et al. Enhanced performance of brain tumor classification via tumor region augmentation and partition. PLoS ONE10, e0140381 (2015).
Saba, T., Mohamed, A. S., El-Affendi, M. A., Amin, J. & Sharif, M. Brain tumor detection using fusion of hand crafted and deep learning features. Cogn. Syst. Res.59, 221–230 (2020).
Anilkumar, B. & Kumar, P. R. Tumor classification using block wise fine tuning and transfer learning of deep neural network and KNN classifier on MR brain images. Int. J. Emerg. Trends Eng. Res.8, 574–583 (2020).
Loshchilov, I. & Hutter, F. SGDR: stochastic gradient descent with warm restarts. In Proceedings of the International Conference on Learning Representations (ICLR 2017), (Toulon, France, 2017).
Donti, P., Amos, B. & Kolter, J. Z. Task-based end-to-end model learning in stochastic optimization. In Advances in Neural Information Processing Systems, 5484-5494 (Long Beach, CA, USA, 2017).
Ke, L., Zhang, Y., Yang, B., Luo, Z. & Liu, Z. Fault diagnosis with synchrosqueezing transform and optimized deep convolutional neural network: An application in modular multilevel converters. Neurocomputing430, 24–33 (2021).
Simonyan, K. & Zisserman, A. Very deep convolutional networks for large-scale image recognition. arXiv preprint arXiv:1409.1556 (2015).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 770–778 (2016).
Huang, G., Liu, Z., van der Maaten, L. & Weinberger, K. Q. Densely connected convolutional networks. Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 4700–4708 (2017).
Tan, M. & Le, Q. V. EfficientNet: Rethinking model scaling for convolutional neural networks. Proceedings of the 36th International Conference on Machine Learning (ICML) 6105–6114 (2019).
Acknowledgements
We would like to thank the Open Fund of Engineering Research Center for Metallurgical Automation and Measurement Technology of Ministry of Education of China (Grant No. MADTOF2024B01), the Excellent Young and Middle-aged Science and Technology Innovation Team Plan Program of Hubei Higher Education of China (Grant No. T2022033), and the Science and Technology Innovation Talent Program of Hubei Province of China (Grant No. 2024DJC093 and Grant No. 2023DJC060).
Funding
This research was supported in part by the Open Fund of Engineering Research Center for Metallurgical Automation and Measurement Technology of Ministry of Education of China (Grant No. MADTOP2024B01), and in part by the Excellent Young and Middle-aged Science and Technology Innovation Team Plan Program of Hubei Higher Education of China (Grant No. T2022033), and in part by the Science and Technology Innovation Talent Program of Hubei Province of China (Grant No. 2024DJC093 and Grant No. 2023DJC060).
Author information
Authors and Affiliations
Contributions
L.K., G.H. and M.Z. conceived the research and designed the methodology. L.K. and Z. Liu developed the software and conducted the experiments. M.Z. and L.K. performed validation and formal analysis. Y.Y., Z. Liu and L.K. conducted data curation and investigation. L.K. wrote the original draft. L.K., Y.Y. and Z. Lv reviewed and edited the manuscript. L.K. and Liu acquired funding. All authors reviewed and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ke, L., Hu, G., Zhao, M. et al. Brain tumor classification from MRI images using a multi-scale channel attention CNN integrated with SVM. Sci Rep 16, 6297 (2026). https://doi.org/10.1038/s41598-026-36164-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-36164-3