Abstract
Breast cancer remains a critical global health challenge, with timely and reliable diagnosis being essential for improving clinical outcomes. Although recent advances in computer aided diagnosis (CAD) have increasingly adopted deep learning, many existing solutions rely on computationally intensive architectures such as Transformer based models, deep ensemble models, multi scale attention networks, DenseNet based frameworks that limit their practical utility. To overcome this limitation, we proposed a comparatively lightweight Reciprocal Gating Fusion framework that integrates two efficient convolutional neural networks, SqueezeNet and ShuffleNetV2, enabling high quality feature extraction with substantially reduced computational overhead. The proposed reciprocal gating mechanism facilitates structured bidirectional interaction between the networks, enhancing complementary feature exchange while suppressing redundant responses to produce a more informative fused representation. Extensive empirical evaluations on benchmark datasets demonstrate strong performance and generalization capability, achieving 97% multiclass accuracy and 99% binary accuracy on the ICIAR-2018 dataset, along with 99.72% accuracy on the BreakHis dataset at 100\(\times\) magnification. These results highlight the effectiveness of the proposed framework in delivering a precise and dependable CAD solution for breast cancer detection. The code is made available at: https://github.com/Cmatermedicalimageanalysis/RCG_ICIAR_Breakhis
Similar content being viewed by others
Introduction
Breast cancer remains one of the most common and life-threatening diseases that affect women worldwide. According to GLOBOCAN 2022, an estimated 20 million new cancer cases and 9.7 million deaths occurred worldwide in 20221. Among these, breast cancer was the most frequently diagnosed malignancy in women, accounting for nearly 15% of all cancers and causing approximately 670,000 deaths per year2.
Breast cancer develops when normal breast cells acquire genetic or molecular changes that lead to uncontrolled growth and tumor formation. The primary method for diagnosis is still histopathology3,4 examination, where stained biopsy and microscopic slides5 are studied under a microscope to evaluate tumor type, grade and structure. However, this manual approach is time consuming, subjective, and difficult to scale as the number of samples increases. In recent years, deep learning based computer aided diagnosis (CAD) systems have significantly advanced breast cancer image analysis. Across diverse medical imaging domains, Artificial Intelligence (AI)-based diagnostic systems6,7,8 and transformer based systems9,10 have demonstrated remarkable capabilities in analyzing complex pathological patterns11,12,13,14. These systems leverage fine-grained attention mechanisms and knowledge-based collaborative networks to enhance diagnostic accuracy and interpretability, particularly for grading tasks that require nuanced understanding of disease progression15. Convolutional neural networks (CNNs) now outperform traditional feature based techniques in tasks such as classification, segmentation16,17, and detection18, offering greater accuracy, consistency, and efficiency in clinical diagnosis. Moreover, spatial-spectral interactive learning approaches and deep unfolding frameworks have emerged as powerful paradigms for detecting small-scale targets in complex backgrounds, offering interpretability through the integration of optimization-based principles with neural network learning19,20.
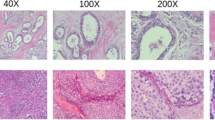
The ICIAR-2018 dataset presents several challenges that complicate the classification of automated breast cancer histopathological images. A major difficulty lies in the high inter-class visual overlap between Benign and Normal samples, as both exhibit similar epithelial and stromal patterns with only subtle morphological differences. Similarly, In Situ carcinoma often shows ambiguous ductal boundaries, which makes it visually comparable to Invasive carcinoma. These fine-grained variations necessitate highly discriminative feature extraction. Moreover, the dataset is affected by stain inconsistencies, illumination variations, and uneven contrast across image patches, arising from differences in image acquisition and colour processing. Such factors lead to visual heterogeneity that can hinder feature learning and model generalisation. Figure 1 demonstrates some sample histopathology images from the ICIAR-2018 dataset, highlighting the inherent challenges in the dataset.
Examples of histopathology images from the ICIAR-2018 dataset illustrating inter-class visual overlap.
Breast cancer remains a leading cause of cancer-related mortality among women worldwide, with histopathological analysis serving as the gold standard for diagnosis. Although recent deep learning–based computer-aided diagnosis systems have shown promise in automating breast cancer histopathology analysis, many existing methods struggle to effectively capture subtle inter-class differences and remain sensitive to stain variability and visual heterogeneity. In particular, conventional CNNs and attention mechanisms often rely on isolated or unidirectional feature interactions, limiting their ability to model fine-grained contextual dependencies in complex tissue structures. To address these limitations, we propose a Reciprocal Cooperative Gating (RCG) mechanism that enables bidirectional feature interaction and adaptive emphasis of discriminative regions, thereby enhancing robust feature representation and improving classification performance on challenging datasets such as ICIAR-2018.
Contributions
This study proposes a Reciprocal Cooperative Gating Fusion (RCG) framework elaborated in Section 3.3, that combines two lightweight CNNs, SqueezeNet 1.0 and ShuffleNetV2_X2.0, pre-trained on ImageNet and fine-tuned on breast cancer histopathological images. Highlighting points of our work are:
-
A novel Reciprocal Cooperative Gating module: It enhances feature fusion by amplifying informative, non-redundant channels and suppressing mutually redundant responses across backbones using learnable parameters (\(\alpha\) and \(\beta\)) and a fixed redundancy threshold \(\tau\).
-
A refined discriminative fused representation: The proposed approach applies learned gates to spatial features, followed by global pooling and vector fusion, to obtain a refined discriminative fused representation that improves robustness and accuracy on breast histopathology images.
Related work
Several research studies have been performed on various models and methods for the histopathological classification of breast cancer images using the ICIAR-2018 and BreakHis datasets. Garg et al.21 introduced a transfer learning-based lightweight ensemble model that integrates a pretrained MobileNetV2 with a custom shallow CNN, where features from both networks are fused using a multilayer perceptron, achieving accuracies above 96% across multiple datasets. Majumdar et al.22 introduced a Gamma function-based rank ensemble method that combines the confidence scores of GoogLeNet, VGG11, and MobileNetV3-Small using a non-linear gamma ranking function to enhance robustness and accuracy across magnifications. Bagchi et al.23 proposed a multi-stage deep learning framework that divides large histopathological images into stain-normalized patches, extracts features using fine-tuned CNN models (VGG16, VGG19, and Inception-ResNet v2), classifies patches through an ensemble of machine learning classifiers, and fuses patch-level predictions using a two-stage neural network for image-level classification.
Jothi et al.24 developed DIRXNet, a hybrid deep network that fuses DenseNet201, InceptionResNetV2, and Xception by concatenating global average pooled intermediate feature maps, enabling robust multi-level representation learning and improving interpretability. Kumar et al.25 proposed SPCZP-CNN, a Spatial Pyramid Complex Zernike Moments Pooling CNN built on DenseNet-121 with squeeze and excitation (SE) blocks, incorporating Zernike moments for rotation invariant texture representation and better morphological feature extraction. Murphy et al.26 presented ensemble frameworks combining DenseNet121, InceptionV3, and ResNet50 with traditional classifiers such as SVM, Random Forest, and LightGBM, showing that hybrid deep machine learning ensembles can outperform standalone CNNs in imbalanced data settings.
Jia et al.27 proposed a hybrid DenseNet LSTM network (DenLSNet) that integrates SE channel attention blocks, iterative convolutional feature fusion, and an LSTM-based classifier to capture contextual dependencies in sequential patch representations. Nair et al.28 developed a hybrid SE-Residual Convolutional and Conformer-based model, where SE Res blocks emphasize salient channel wise features while the Conformer captures long-range dependencies, improving spatial contextual learning. Yan et al.29 introduced DWNAT-Net, a hybrid framework that integrates the Discrete Wavelet Transform (DWT) for multi frequency decomposition with a Neighborhood Attention Transformer (NAT) to model local spatial attention, resulting in improved texture structure awareness. Kutluer et al.30 proposed a hybrid feature selection and deep learning framework combining deep feature extraction using ResNet-50, GoogLeNet, Inception-v3, and MobileNet-v2 with Gray Wolf Optimization (GWO) for optimal feature subset selection, achieving significant improvements in accuracy and feature compactness.
More studies in breast cancer histopathological image classification have introduced a variety of innovative methodologies and architectures. For instance, Gül et al.31 proposed a hybrid approach based on Local Binary Patterns (LBP), where texture descriptors are fused with a custom CNN to extract discriminative features from H&E stained breast histology images. Sreelekshmi et al.32 developed SwinCNN, a hybrid architecture combining a Swin Transformer and a CNN backbone to capture both global (transformer) and local (CNN) contextual information, thereby improving breast cancer grade classification on histopathological slides. Liang et al.33 introduced Brea-Net, an interpretable dual attention network employing spatial and channel attention modules to address class imbalance and enhance feature discrimination. In another line of research, Alshehri et al. proposed a Modality Specific CBAM-VGGNet model34, which integrates the Convolutional Block Attention Module (CBAM) with VGGNet via transfer learning for modality-specific feature extraction. Furthermore, they developed BreNet35, an attention-enhanced multi scale CNN framework designed to combine features across different spatial scales, highlighting the advantages of multi scale representation. Hossain et al.36 proposed an interpretable ensemble approach with threshold filtered Single Instance Evaluation (SIE) to improve histopathology-based HER2 breast cancer classification, achieving high accuracy while employing Grad-CAM for model interpretability. Collectively, these studies underscore the significant potential of histopathological image analysis for breast cancer diagnosis and classification.
Apart from histopathological image analysis, numerous other imaging modalities and learning strategies have been explored for breast cancer detection and classification. Qureshi et al.37 presented a comprehensive review of mammography based diagnostic techniques ranging from traditional image processing to advanced deep learning frameworks for the segmentation and classification of microcalcifications, highlighting how modern CNN architectures have significantly improved diagnostic accuracy. Strelcenia et al. 38 proposed a K-CGAN based generative framework that synthetically augments breast cancer datasets, thereby reducing class imbalance and enhancing classifier robustness. Zeng et al. 39 introduced the FastLeakyResNet-CIR framework, a residual network variant optimized with fast convergence and leaky activation to achieve superior performance in multiple imaging modalities, including mammography, ultrasound, and MRI. Hussein et al. 40 developed a CNN based on transfer learning that automatically detects and classifies breast cancer from mammogram images using pretrained networks such as VGG19 and ResNet50, achieving high accuracy with minimal training data. Similarly, Sharma et al. 41 proposed a multimodal fusion model combining MRI features with deep neural representations to enhance tumor localization and classification accuracy in magnetic resonance imaging. While these studies demonstrate the strong potential of modalities such as ultrasound, mammography, and MRI for early detection and classification, the present work particularly focuses on the analysis of histopathological images to achieve fine grained feature representation and improved interpretability of breast cancer subtypes.
Despite significant progress in breast cancer histopathological image classification, existing methods often rely on computationally heavy models including Transformer based models (e.g., Vision Transformers, Swin Transformers) or single backbone CNNs, which limit efficiency and generalizability across datasets. While lightweight CNNs and fusion strategies have been explored, effective multi backbone feature integration with attention or gating mechanisms remains under investigated, particularly in achieving a balance between accuracy, interpretability, and low computational cost. Second, existing multi backbone fusion strategies predominantly use simple concatenation or element wise operations that inadequately model complementary feature relationships, producing sub optimal representations with redundant or conflicting information. To bridge these gaps, our work introduces a comparatively lightweight RCG framework that integrates SqueezeNet and ShuffleNetV2 through a structured bidirectional gating mechanism, enabling adaptive complementary feature exchange while suppressing redundancies, thus achieving high classification accuracy with comparatively lower computational complexity.
Methodology
Figure 2 illustrates the overall workflow of the proposed methodology. The framework integrates SqueezeNet42 and ShuffleNetV243 to extract features. These features are refined through the Reciprocal Cooperative Gating (RCG) module, as mentioned in Section 3.3, which allows mutual enhancement of features between the two streams. The gated feature representations are then concatenated and passed through fully connected layers to perform final classification into cancerous and non-cancerous categories.
Workflow of the proposed methodology. The equations corresponding to each step are described in Section 3.3.
SqueezeNet 1.0
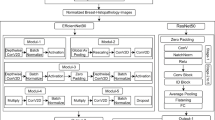
SqueezeNet, as the name suggests, is a deep neural architecture that has a “squeezed” design for training networks for image classification. SqueezeNet 1.042 is used as the extractor of the main characteristics of the proposed CAD model. The architecture44 is composed of an initial convolution maxpool layer followed by a sequence of Fire modules, each consisting of a squeeze layer (\(1\times\)1 convolutions for channel reduction) and an expand layer (a combination of \(1\times\)1 and \(3\times\)3 convolutions for feature expansion). This design strategy enables parameter efficiency while preserving representational power. Figure 3 illustrate the overall SqueezeNet architecture and the internal structure of the Fire module, respectively.
ShuffleNetV2_X2.0
ShuffleNetV243 is a lightweight CNN optimized for efficient computation on mobile devices and edge devices. It uses channel split, channel shuffle, and separable depthwise convolutions to balance accuracy and computational cost. The ShuffleNetV2_X2.0 architecture consists of an initial convolution layer and maxpooling, followed by three main stages of ShuffleNet units (Stage 2, Stage 3, and Stage 4) that gradually increase channel depth, and a final convolution \(1\times 1\) (Conv5) for high-level feature representation. The overall architecture of ShuffleNetV2 is illustrated in Fig. 4.
An illustration of the ShuffleNet architecture43.
Reciprocal cooperative gating mechanism
Our proposed Reciprocal Cooperative Gating (RCG) module is conceptually inspired by the Reciprocal Transformation Module (RTM) 45, particularly its reciprocal gating mechanism that balances bidirectional feature interactions between appearance and motion streams. While the original RTM was designed to improve spatio temporal correspondence between video frames through reciprocal scaling, transformation, and gating, our model adapts this principle to a static dual CNN framework for medical image classification. Specifically, we reformulate the concept of mutual feature refinement into a comparatively lightweight cooperative gating mechanism that dynamically modulates two feature streams originating from SqueezeNet and ShuffleNet through learnable parameters \(\alpha\) and \(\beta\), as well as contrast normalized responses controlled by a tunable threshold \(\tau\). This design enables bidirectional cooperation between the two networks without heavy attention computations, effectively reweighting and aligning feature representations to improve discriminative fusion while maintaining computational efficiency. The RCG module enhances multi backbone feature fusion by emphasizing complementary information and suppressing redundancy.
Let the feature maps extracted from the two backbones be denoted as \(f_1\) and \(f_2\), where each resides in a four dimensional tensor space characterized by batch size B, number of channels \(C_i\), and spatial dimensions \(H_i \times W_i\).
The RCG operation begins by passing each feature map through a lightweight \(1 \times 1\) convolution followed by batch normalization and a ReLU activation, producing intermediate representations \(z_i\) that capture channel wise interactions. These are then processed by global average pooling (GAP) to produce channel-wise intent vectors \(g_i\), each summarizing the semantic content of a channel for a given sample.
Next, each intent vector \(g_i\) is mean centered to compute deviation signals \(s_i\), which represent how much each channel deviates from the overall mean and thus indicate redundancy or channel imbalance. To regulate this redundancy, we introduce a scaling operation controlled by learnable parameters \(\gamma _i\), instantiated as \(\alpha\) and \(\beta\) for the two branches. The scaling involves a sigmoid transformation \(\sigma (\gamma _i)\) and a normalized deviation term adjusted by the threshold \(\tau\), ensuring that redundancy is adaptively suppressed during feature refinement.
Following this, each channel intent \(g_i\) is passed through a lightweight feedforward head (a linear layer \(W_i\)) followed by a sigmoid activation to produce the initial gate values \(h_i\). These gates are then refined cooperatively by incorporating redundancy feedback \(r_i\), resulting in final cooperative gates \(G_i\) defined through the interaction between \(h_i\) and \(r_i\). This cooperative gating allows the two branches to adjust their internal activations in a mutually beneficial way, ensuring that each stream enhances features that are complementary to the other.
Finally, the original feature maps \(f_i\) are modulated element wise by their corresponding gate values \(G_i\), producing gated maps \(f_i'\). Each gated map is subsequently globally pooled, concatenated across the two branches, and passed through a classifier to yield the final prediction. Through this reciprocal gating strategy, each backbone adaptively emphasizes salient information from the other, resulting in refined and discriminative feature representations that improve both robustness and interpretability.
The complete operational steps of the proposed RCG mechanism are summarized in Algorithm 1.
Reciprocal cooperative gating (RCG) mechanism.
Fusion
The proposed classifier leverages a dual-backbone fusion strategy by combining SqueezeNet 1.042 and ShuffleNetV2_X2.043. SqueezeNet contributes a compact 512 dimensional feature vector, while ShuffleNetV2_X2.0 provides a 2048 dimensional representation. These features are refined through the RCG module 3.3, which adaptively enhances informative channels and suppresses redundant ones via reciprocal interaction between the two networks. The gated outputs are globally pooled and concatenated into a unified 2560 dimensional vector, which passes through a fully connected head with batch normalization, ReLU activation, and dropout for final prediction. This design effectively combines the efficiency of SqueezeNet with the computational power of ShuffleNetV2, resulting in a lightweight yet robust model for medical image classification. Table 1 summarizes the quantitative resource metrics.
Result and discussion
Datasets
The datasets used for evaluation of our proposed model are ICIAR-201846 and BreakHis47. The ICIAR-2018 dataset consists of Hematoxylin and Eosin (H&E) stained breast histopathology microscopy and whole-slide images. It contains a total of 400 microscopy images labeled as normal, benign, in situ carcinoma and invasive carcinoma, where each of the four classes has 100 images. We also evaluated the model on BreakHis dataset, a standard dataset used to study the breast cancer classification problem. It is a histopathological dataset containing 1176 samples (100x magnification) which fall under the two main categories: Benign and Malignant. The Benign class contains 588 samples and the Malignant class contains 588 samples.
The standard datasets analysed in this study are available from the following sources:
-
ICIAR-2018: https://iciar2018-challenge.grand-challenge.org/
-
BreaKHis: web.inf.ufpr.br
Data preprocessing
All images in both the ICIAR-2018 and BreakHis datasets are normalized and resized to a height and width of 256. From the ICIAR-2018 dataset, we take randomly 100 test images (25 images from each class), and the remaining 300 images are augmented to 3600 images, with random rotations of degree 5, horizontal flip, and vertical flip. The train-val dataset contains these augmented 3600 images and the original 300 images. The train-val dataset is then divided into training and validation sets at an 80:20 ratio22. We also perform a 2-class classification on this dataset, where normal and benign labels are taken as the in-carcinoma class, and in situ and invasive carcinoma labels are taken as the carcinoma class. Therefore, each of the two classes consists of 200 images each. Here, 100 random images are taken as test images (50 in each class) and the remaining 300 images are preprocessed similarly to the 4-class classification mentioned above. From the BreakHis dataset, we randomly take 352 images of the total samples as test images and the remaining 2081 samples are divided into training and validation sets in an 80:20 ratio. The detailed dataset splits and augmentations for both 4-class and 2-class classifications are summarized in Table 2.
Hyperparameters and evaluation metrics
Each of the CNN models and methods discussed in Section 3.1, Section 3.2, Section 3.3, and Section 3.4 are retrained on the training set for 100 epochs with early stopping by fine-tuning all layers using ReduceLROnPlateau learning rate scheduler and CrossEntropyLoss loss function. All relevant hyperparameters and training settings are summarized in Table 3.
Table 4 reports the ablation study conducted to analyze the influence of the learnable parameters \(\alpha\), \(\beta\), and \(\tau\) in the proposed RCG module. Multiple combinations were evaluated on the ICIAR-2018 and BreakHis datasets. The results indicate that balanced values of \(\alpha\) and \(\beta\), along with an intermediate threshold \(\tau\), lead to improved classification performance. The best results were achieved with \(\alpha =\beta =0.6\) and \(\tau =0.15\), demonstrating the effectiveness of the selected configuration.
To assess the performance of our multi-class image classification model, we employ a set of standard evaluation metrics, namely Accuracy, Precision, Recall, and F1-score. Additionally, we include the Confusion Matrix and the t-distributed Stochastic Neighbor Embedding (t-SNE) plot as performance visualization measures to provide a deeper understanding of class-level predictions and feature separability. The formal definitions of these metrics are as follows:
Accuracy. Accuracy quantifies the overall correctness of the model by measuring the ratio of correctly classified instances to the total number of samples:
where TP, TN, FP, and FN represent true positives, true negatives, false positives, and false negatives, respectively. In a multi-class setting, accuracy is computed as the proportion of correctly classified samples across all classes.
Precision. Precision reflects how many of the samples predicted as positive are truly positive, and is given by:
It indicates the trustworthiness of the model’s positive predictions.
Recall. Recall, also known as Sensitivity, measures the proportion of actual positive instances that are correctly identified by the model:
This metric highlights the model’s effectiveness in identifying all relevant positive cases.
F1-score. The F1-score provides a balanced measure between Precision and Recall by taking their harmonic mean:
It is particularly useful when there is an imbalance between classes.
Confusion Matrix. The confusion matrix presents a detailed breakdown of classification outcomes by comparing actual class labels with predicted ones. Each row corresponds to a true class, and each column represents a predicted class. The diagonal elements indicate correctly classified samples for each class, while the off-diagonal elements show the types and counts of misclassifications. This matrix helps in identifying specific classes where the model may be underperforming.
t-Distributed Stochastic Neighbor Embedding (t-SNE). To further investigate the model’s feature representation and its ability to separate distinct classes in the learned embedding space, we utilize the t-SNE technique. t-SNE is a non-linear dimensionality reduction method that projects high-dimensional feature vectors into a two-dimensional space while preserving local neighborhood structures. By visualizing the t-SNE plots of features extracted from the final layer, we can observe the degree of clustering among samples of the same class and the separation between different classes. A well-clustered and distinctly separated t-SNE plot indicates that the model has effectively learned discriminative feature representations, thereby reinforcing the quantitative evaluation metrics.
Heatmap. To further investigate the model’s decision making process and validate its focus on clinically relevant regions, we employ heatmap visualization techniques. Heatmaps generate spatial activation maps that highlight the most discriminative regions in histopathological images by identifying areas that contribute most significantly to the classification decision. By visualizing these activation maps overlaid on the original tissue samples, we can observe whether the model correctly attends to diagnostically meaningful features such as cellular morphology, nuclear characteristics, and tissue architecture. A well-localized heatmap that emphasizes pathologically significant regions indicates that the model has effectively learned to identify disease-specific patterns, thereby reinforcing confidence in the classification results and enhancing the interpretability of the proposed framework.
Quantitative analysis
Following the pre-processing and augmentation steps described earlier, experiments were conducted on both the ICIAR-2018 and BreakHis datasets. As summarized in Tables 7, 8 and 9, the proposed RCG-based fusion network demonstrates superior classification performance compared to all baseline models across both datasets. All models have been analyzed and compared on the basis of precision, recall, F1-score and accuracy.
Base model selection
The selection of base models has been performed based on experiments using different CNN backbones. Performances of different backbones, in terms of accuracy, on ICIAR 2018 and BreakHis datasets have been shown in Table 5. It is observed that SqueezeNet and ShuffleNetV2 give the best performance across all datasets, and hence these two models are selected as the two backbones of our dual-branch network.
Attention based gating methods
In addition to the proposed gating mechanism, we benchmark our model against established attention-based gating methods, namely SE48, CBAM49, and ECA50, which are widely used in modern deep learning architectures. The detailed quantitative results are summarized in Table 6. It is observed that the proposed RCG outperforms all other gating methods considered here for comparison in both ICIAR 2018 and BreakHis datasets. This comparison enables a fair assessment of the proposed design against both lightweight and expressive attention formulations.
ICIAR-2018 multiclass
For the 4-class ICIAR-2018 dataset, as shown in Table 7, the proposed model achieves an overall accuracy of 97%, surpassing the best-performing baseline (SqueezeNet 1.0\(+\)ShuffleNetV2_X2.0) by approximately 3%. The t-SNE visualization in Fig. 5 clearly illustrates the feature separability among the four histological subtypes. The training and validation curves shown in Fig. 5 depict the convergence behavior of the proposed model, further validating the discriminative strength of the fused representations. The confusion matrices in Fig. 8 illustrate the classification performance of (a) ShuffleNetV2, (b) SqueezeNet, (c) the Fusion model, and (d) the proposed RCG Fusion. The RCG framework demonstrates more balanced predictions with fewer misclassifications across all histological categories, highlighting its superior feature fusion and discriminative capability through reciprocal cooperative gating.
t-SNE representation and training curves for RCG on the 4-class ICIAR-2018 dataset.
ICIAR-2018 binary
For the 2-class ICIAR-2018 dataset, as shown in Table 8, the proposed model achieves an overall accuracy of 99% between the carcinoma and non-carcinoma classes. The t-SNE visualization in Fig. 6 provides a clear depiction of the feature separability achieved by the proposed approach. The loss and accuracy plots in Fig. 6 further validate the training stability of the RCG-based fusion model. These results highlight the robustness and generalizability of the proposed lightweight reciprocal gating fusion mechanism, which effectively enhances discriminative feature learning while preserving computational efficiency, making it suitable for practical CAD implementations in histopathological image analysis. The confusion matrices in Fig. 9 illustrate the classification performance of (a) ShuffleNetV2, (b) SqueezeNet, (c) Simple Fusion, and (d) Proposed RCG-based Fusion. The RCG framework exhibits highly precise and balanced predictions across carcinoma and non-carcinoma categories, confirming its superior discriminative power and robust cooperative gating mechanism.
t-SNE representation and training curves for RCG on the 2-class ICIAR-2018 dataset.
BreakHis binary
On the BreakHis dataset, as shown in Table 9, the proposed model for binary classification secures 99.72% accuracy, maintaining consistently high precision and recall values. Figure 7 provides the t-SNE visualization, demonstrating the clear feature separability achieved by the proposed model. Figure 7 indicate the training and validation performance curves. The confusion matrices in Fig. 10 illustrate the classification performance of (a) ShuffleNetV2, (b) SqueezeNet, (c) the Fusion model, and (d) the proposed RCG Fusion (Figs. 8, 9, 10).
t-SNE representation and training curves for RCG on the BreakHis dataset.
Confusion matrices generated by the proposed fusion model and base models on ICIAR-2018 multiclass dataset.
Confusion matrices generated by the proposed fusion model and base models on ICIAR-2018 binary dataset.
Confusion matrices generated by the proposed fusion model and base models on BreakHis dataset.
Statistical analysis
To further substantiate the superiority of the proposed model compared to its ablation variants, we conducted a Wilcoxon Rank-Sum test to assess statistical significance across binary and multiclass classification tasks in the ICIAR-2018 dataset. This non-parametric test was selected because it does not rely on the assumption of normal data distribution, making it appropriate for evaluating model accuracy values collected from multiple independent runs. The analysis was performed using accuracy scores from ten separate experimental trials for each model configuration.
As shown in Fig. 11 (multiclass classification) and Fig. 12 (binary classification), results of the Wilcoxon Rank-Sum test clearly indicate significant performance differences between the proposed framework and its ablation variants ShuffleNetV2, SqueezeNet1.0, and the simple fusion model. For the multiclass task, the obtained p-values were 0.0022, 0.0002, and 0.0257, respectively. Similarly, for the binary classification task, the p-values were 0.0003, 0.0002, and 0.0006, all of which are below the 0.05 significance level.
Results obtained using the Wilcoxon Rank-Sum Test on 4-class ICIAR-2018 dataset.
Results obtained using the Wilcoxon Rank-Sum Test on 2-class ICIAR-2018 dataset.
These outcomes demonstrate that the improvements achieved by the proposed model are statistically significant rather than occurring by chance. They also confirm the effectiveness of the RCG-based fusion mechanism in enhancing the interaction between backbone features, leading to better classification performance compared to both individual lightweight networks and their simple fusion versions.
Qualitative evaluation
The qualitative evaluation of model interpretability is illustrated through Grad-CAM and heatmap visualizations shown in Figs. 13, 14, 15 and 16. The comparison highlights activation regions obtained from different backbone models - ShuffleNet and SqueezeNet, and the proposed RCG-based fusion network. As observed, the proposed model produces more compact and discriminative activation maps, accurately focusing on diagnostically relevant tissue regions in histopathological images. This demonstrates that the cooperative gating mechanism enhances feature fusion and spatial attention, resulting in improved model interpretability and performance.
Grad-CAM visualizations for different backbone models in ICIAR-2018 dataset. Comparison among (a) Original image, and outputs of (b) SqueezeNet, (c) ShuffleNet, and (d) RCG; demonstrates that RCG-based fusion produces more focused and discriminative activation regions on histopathology images.
Grad-CAM visualizations for different backbone models in the BreakHis (100\(\times\)) dataset. Comparison among (a) Original image, and outputs of (b) SqueezeNet, (c) ShuffleNet, and (d) RCG; demonstrates that RCG-based fusion produces more focused and discriminative activation regions on histopathology images.
Heatmap visualizations in ICIAR-2018 dataset.
Heatmap visualizations in BreakHis (100x) dataset.
Discussion
The comparison in Table 10 clearly indicates that the proposed approach achieves substantial and consistent improvements in classification accuracy and F1-score. We have proposed an RCG-based fusion framework utilizing two light weight CNN models, ShuffleNetV2_X2.0 and SqueezeNet 1.0, making the overall model significantly lighter in terms of parameters compared to other related works on breast cancer histopathological image classification. The RCG module plays a crucial role in feature reshaping, enhancing the discriminative capability of the extracted features and thereby improving model robustness. As a result, the proposed method demonstrates superior generalization and consistent performance gains across evaluation metrics. Furthermore, the bar graph presented in Figure 17 visually compares the proposed model with existing methods, highlighting its clear advantage in accuracy across all evaluated datasets.
Comparison of the proposed model with existing models across datasets.
Strengths, weaknesses and future extension
In this section, we summarize the strengths and weaknesses of the proposed model. In addition, we also provide some suitable suggestions to tackle the problems faced by RCG.
Strengths
-
RCG dynamically suppresses noisy or less informative channels while enhancing discriminative ones through sample-wise, channel-wise gating. This is particularly beneficial in medical images where lesions occupy small regions or background tissue dominates the feature space.
-
RCG adds minimal overhead: only 1\(\times\)1 convolutions, channel-wise MLP heads, no spatial attention maps. This makes it suitable for: resource-constrained clinical settings and edge or real-time medical inference.
Weaknesses
-
Although the proposed RCG framework achieves strong performance on the BreakHis dataset, we note that BreakHis contains multiple images acquired from the same patient at different magnifications. In the current study, data splitting is performed at the image level, which may allow images from the same patient to appear in both training and test sets, potentially leading to optimistic performance estimates. A patient-wise split would provide a more stringent and clinically realistic evaluation. Future work will focus on conducting patient-level experiments and validating the proposed method under such protocols.
-
RCG is applied after backbone feature extraction, which means: earlier layers are not guided, low-level noise may propagate forward. Early or multi-stage gating could further improve robustness.
Future extension
-
The strength of reciprocal gating may be adjusted dynamically per sample or per training epoch using statistical measures such as feature variance, entropy, or confidence scores.
-
Reciprocal gating may be extended to multiple intermediate layers of the backbone networks to allow progressive feature cooperation rather than relying solely on late-stage gating.
-
Task-adaptive threshold scheduling strategies may be investigated, where \(\tau\) evolves during training to balance exploration and suppression more effectively across different classes.
Conclusions
We propose a Reciprocal Cooperative Gating–based fusion framework that integrates SqueezeNet 1.0 and ShuffleNetV2_X2.0, two lightweight pretrained CNNs fine-tuned for the classification of the histopathological image of breast cancer. The mutual gating mechanism in RCG enables reciprocal feature exchange between networks, enhancing discriminative power while maintaining low computational cost. The proposed model achieved 97% (multiclass) and 99% (binary) accuracy on the ICIAR-2018 dataset, and 99.72% on BreakHis (100\(\times\)), surpassing existing methods.
The framework offers a compact design and efficient feature fusion, establishing it as a robust and accurate CAD tool for breast cancer detection. Despite these advantages, the model has certain limitations: the gating mechanism, redundancy scaling, and cross-branch feedback require careful hyperparameter tuning (e.g. \(\tau\), \(\alpha\) and \(\beta\)) for stable training, and the current design is tailored for exactly two backbones, which limits scalability to architectures with more feature extractors.
Future work will explore class-imbalance–aware loss functions and adaptive gating or attention mechanisms to improve performance across multiple magnifications and enhance multiclass classification with minimal computational overhead.
Data availability
The public datasets used and analyzed during the current study are available in the following sources: ICIAR-2018 dataset: https://rdm.inesctec.pt/dataset/604dfdfa-1d37-41c6-8db1-e82683b8335a/resource/df04ea95-36a7-49a8-9b70-605798460c35/download/breasthistology.zip) and BreaKHis dataset: https://www.inf.ufpr.br/vri/databases/BreaKHis_v1.tar.gz.
Change history
01 April 2026
A Correction to this paper has been published: https://doi.org/10.1038/s41598-026-46426-9
References
Bray, F. et al. Global cancer statistics 2022: Globocan estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 74, 229–263. https://doi.org/10.3322/caac.21825 (2024).
World Health Organization. Global cancer burden growing, amidst mounting need for services. Online (2024). World Health Organization News Release.
Luo, J. et al. Dca-daffnet: An end-to-end network with deformable fusion attention and deep adaptive feature fusion for laryngeal tumor grading from histopathology images. IEEE Trans. Instrum. Meas. 72, 1–15 (2023).
Huang, P. et al. A vit-amc network with adaptive model fusion and multiobjective optimization for interpretable laryngeal tumor grading from histopathological images. IEEE Trans. Med. Imaging 42, 15–28 (2022).
Pan, H. et al. Breast tumor grading network based on adaptive fusion and microscopic imaging. Opto-Electron. Eng. 50, 220158–1 (2022).
Mo, C. et al. Harnessing artificial intelligence for accurate diagnosis and radiomics analysis of combined pulmonary fibrosis and emphysema: Insights from a multicenter cohort study. medRxiv (2025).
Chen, J. et al. Interpretable and reproducible machine learning model for coronary calcification and segment-level stenoses stratification on computed tomography angiography. BMC Med. 23, 657 (2025).
Wu, Y. et al. An eyecare foundation model for clinical assistance: A randomized controlled trial. Nat. Med. 1–10 (2025).
Wang, Y. et al. The swin-transformer network based on focal loss is used to identify images of pathological subtypes of lung adenocarcinoma with high similarity and class imbalance. J. Cancer Res. Clin. Oncol. 149, 8581–8592 (2023).
Huang, P. & Luo, X. Fdts: A feature disentangled transformer for interpretable squamous cell carcinoma grading. IEEE/CAA J. Automatica Sinica (2025).
Qu, Y. et al. Cgam: An end-to-end causality graph attention mamba network for esophageal pathology grading. Biomed. Signal Process. Control 103, 107452 (2025).
Li, C., Bozorgtabar, B., Ping, Y., Huang, P. & Qin, J. Positive semi-definite latent factor grouping-boosted cluster-reasoning instance disentangled learning for wsi representation. arXiv preprint arXiv:2511.01304 (2025).
Li, C., Huang, P., Qin, J. & Luo, X. Knowledge-driven multiple instance learning with hierarchical cluster-incorporated aware filtering for larynx pathological grading. IEEE J. Biomed. Health Inform. (2025).
Ma, M. et al. A multi-instance learning network with prototype-instance adversarial contrastive for cervix pathology grading. Med. Image Anal. 103880 (2025).
Tian, M. et al. Fine-grained attention & knowledge-based collaborative network for diabetic retinopathy grading. Heliyon 9 (2023).
Zhu, M., Cheng, D., Mao, Y., Sun, L. & Jing, W. Local-global multi-scale attention network for medical image segmentation. PeerJ Computer Sci. 11, e3033 (2025).
Fan, Y., Song, J., Yuan, L. & Jia, Y. Hct-unet: Multi-target medical image segmentation via a hybrid cnn-transformer unet incorporating multi-axis gated multi-layer perceptron. Visual Computer 41, 3457–3472 (2025).
Arun Kumar, S. & Sasikala, S. Review on deep learning-based cad systems for breast cancer diagnosis. Technol. Cancer Res. Treatment 22, 15330338231177976 (2023).
Li, Y., Wang, L. & Chen, S. Smile: Spatial-spectral mamba interactive learning for infrared small target detection. IEEE Trans. Geosci. Remote Sensing. (2025).
Li, Y., Wang, L. & Chen, S. From optimization to network: A low-rank and sparse-aware deep unfolding framework for infrared small target detection. Adv. Eng. Inform. 69, 103991 (2026).
Garg, S. & Singh, P. Transfer learning based lightweight ensemble model for imbalanced breast cancer classification. IEEE/ACM Trans. Comput. Biol. Bioinform. 20, 1529–1539 (2022).
Majumdar, S., Pramanik, P. & Sarkar, R. Gamma function based ensemble of cnn models for breast cancer detection in histopathology images. Expert Syst. Appl. 213, 119022 (2023).
Bagchi, A., Pramanik, P. & Sarkar, R. A multi-stage approach to breast cancer classification using histopathology images. Diagnostics 13, 126 (2022).
Jothi, J. A. A. & Damania, K. Dirxnet: A hybrid deep network for classification of breast histopathology images. SN Computer Sci. 5, 77 (2023).
Kumar, A., Singh, C. & Sachan, M. K. A moment-based pooling approach in convolutional neural networks for breast cancer histopathology image classification. Neural Comput. Appl. 37, 1127–1156 (2025).
Murphy, G. & Singh, R. Comparative analysis and ensemble enhancement of leading cnn architectures for breast cancer classification. arXiv preprint arXiv:2410.03333 (2024).
Jia, Y. et al. Denlsnet-c: A novel model for breast cancer classification in pathology images based on densenet and lstm. J. Supercomput. 81, 934 (2025).
Nair, L. S., Amarnath, K. & Nair, J. J. Advancing breast cancer detection: Se-conformer framework for malignancy detection in histopathology images. IEEE Access (2025).
Yan, Y., Lu, R., Sun, J., Zhang, J. & Zhang, Q. Breast cancer histopathology image classification using transformer with discrete wavelet transform. Med. Eng. Phys. 138, 104317 (2025).
Kutluer, N., Solmaz, O. A., Yamacli, V., Eristi, B. & Eristi, H. Classification of breast tumors by using a novel approach based on deep learning methods and feature selection. Breast Cancer Res. Treatment 200, 183–192 (2023).
Gül, M. A novel local binary patterns-based approach and proposed cnn model to diagnose breast cancer by analyzing histopathology images. IEEE Access 13, 39610–39620. https://doi.org/10.1109/ACCESS.2025.3545052 (2025).
Sreelekshmi, V., Pavithran, K. & Nair, J. J. Swincnn: An integrated swin transformer and cnn for improved breast cancer grade classification. IEEE Access https://doi.org/10.1109/ACCESS.2024.3397667 (2024).
Liang, Y. & Meng, Z. Brea-net: An interpretable dual-attention network for imbalanced breast cancer classification. IEEE Access 11, 100508–100517. https://doi.org/10.1109/ACCESS.2023.3314978 (2023).
Alshehri, H. M. Modality specific cbam-vggnet model for the classification of breast histopathology images via transfer learning. IEEE Access 13, 143377–143391. https://doi.org/10.1109/ACCESS.2025.xxxxxx (2025).
Alshehri, H. M. Brenet: Attention-enhanced multi-scale cnn framework for breast cancer classification in histopathological images. IEEE Access 13, 143392–143405. https://doi.org/10.1109/ACCESS.2025.xxxxxx (2025).
Hossain, M. S. S. et al. Addressing uncertainty in imbalanced histopathology image classification of her2 breast cancer: An interpretable ensemble approach with threshold filtered single instance evaluation (sie). IEEE Access 11, 122238–122251. https://doi.org/10.1109/ACCESS.2023.3278923 (2023).
Qureshi, S. A. et al. Breast cancer detection using mammography: Image processing to deep learning. IEEE Access 13, 60776–60801. https://doi.org/10.1109/ACCESS.2024.3523745 (2025).
Strelcenia, E. & Prakoonwit, S. Improving cancer detection classification performance using gans in breast cancer data. IEEE Access 11, 71594–71612. https://doi.org/10.1109/ACCESS.2023.3291336 (2023).
Zeng, R. et al. Fastleakyresnet-cir: A novel deep learning framework for breast cancer detection and classification. IEEE Access 12, 70825–70836. https://doi.org/10.1109/ACCESS.2024.3401729 (2024).
Hussein, A. et al. A novel deep-learning model for automatic detection and classification of breast cancer using the transfer-learning technique. IEEE Access. https://doi.org/10.1109/ACCESS.2025.1234567 (2025).
Sharma, R., Verma, A. & Kaur, S. Classification of breast cancer in mri with multimodal fusion. IEEE Conf. Biomed. Imaging https://doi.org/10.1109/BIOMED.2025.10230686 (2025) ((IEEE)).
Liu, Y., Li, Z., Chen, X., Gong, G. & Lu, H. Improving the accuracy of squeezenet with negligible extra computational cost. In 2020 International Conference on High Performance Big Data and Intelligent Systems (HPBD &IS), 1–6 (IEEE, 2020).
Ma, N., Zhang, X., Zheng, H.-T. & Sun, J. Shufflenet v2: Practical guidelines for efficient cnn architecture design. In Proceedings of the European conference on computer vision (ECCV), 116–131 (Springer, 2018).
Iandola, F. N. et al. Squeezenet: Alexnet-level accuracy with 50x fewer parameters and< 0.5 mb model size. arXiv preprint arXiv:1602.07360 (2016).
Ren, S. et al. Reciprocal transformations for unsupervised video object segmentation. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 15455–15464, https://doi.org/10.1109/CVPR46437.2021.01519 (2021).
Grand challenge on breast cancer histology images. Aresta, G., Araújo, T., Kwok, S., Chennamsetty, S. S. & Safwan, M. e. a. Bach. Medical Image Analysis 56, 122–139. https://doi.org/10.1016/j.media.2019.05.010 (2019).
Spanhol, F. A., Oliveira, L. S., Petitjean, C. & Heutte, L. A dataset for breast cancer histopathological image classification. IEEE Trans. Biomed. Eng. 63, 1455–1462. https://doi.org/10.1109/TBME.2015.2496264 (2015).
Hu, J., Shen, L. & Sun, G. Squeeze-and-excitation networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR) (2018).
Woo, S., Park, J., Lee, J.-Y. & Kweon, I. S. Cbam: Convolutional block attention module. In Proceedings of the European Conference on Computer Vision (ECCV) (2018).
Wang, Q. et al. Eca-net: Efficient channel attention for deep convolutional neural networks. In Proceedings of the IEEE/CVF conference on computer vision and pattern recognition, 11534–11542 (2020).
Wang, G. et al. Multi-classification of breast cancer pathology images based on a two-stage hybrid network. J. Cancer Res. Clin. Oncol. 150, 505 (2024).
Guzel, K. & Bilgin, G. Hft-net: Hybrid fusion transformer network for multi-source breast cancer classification. IEEE Access 13, 170126–170146. https://doi.org/10.1109/ACCESS.2025.3615654 (2025).
Alzoubi, H. et al. Enhancing breast cancer diagnosis with resnet50 and salp swarm-based feature reduction on breakhis dataset. Biomed. Signal Process. Control 114, 109319. https://doi.org/10.1016/j.bspc.2025.109319 (2026).
Acknowledgements
The study was supported by the Ministry of Economic Development of the Russian Federation (agreement No. 139-10-2025-034 dd. 19.06.2025, IGK 000000C313925P4D0002).
Author information
Authors and Affiliations
Contributions
Conceptualization: Britika Khati, Ram Sarkar and Dmitrii Kaplun; Investigation: Britika Khati, Sayan Mukherjee, Aleksandr Sinitca; Methodology: Britika Khati, Sayan Mukherjee and Ram Sarkar; Project administration: Ram Sarkar and Dmitrii Kaplun; Resources: Dmitrii Kaplun; Software: Britika Khati, Sayan Mukherjee, Aleksandr Sinitca; Supervision: Ram Sarkar and Dmitrii Kaplun; Validation: Britika Khati, Sayan Mukherjee and Aleksandr Sinitca; Writing: All authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this Article was revised: The original version of this Article contained an error in the spelling of the author Dmitrii Kaplun which was incorrectly given as Dmtrii Kaplun.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Khati, B., Mukherjee, S., Sinitca, A. et al. Reciprocal cooperative gating fusion of SqueezeNet and ShuffleNetV2 for breast cancer detection in histopathology images. Sci Rep 16, 5904 (2026). https://doi.org/10.1038/s41598-026-36375-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-36375-8