Abstract
Nail diseases, including such common conditions as fungus, and more serious issues like melanoma, may be important clues to the overall health and require a clear diagnosis to be treated. The purpose of the paper is to create a nail disease detection system based on the advanced machine learning methods, including transfer learning and federated learning. The research seeks to show how machine learning and federated learning can be combined to detect nail disease performance with high accuracy without having to share data. The data include pictures of diverse nail conditions including Acral Lentiginous Melanoma, Onychogryphosis, and Pitting among others that are checked to maintain the quality of data in a uniform manner to facilitate the effective training of the models. The most common feature extraction models are ResNet152V2, DenseNet201, MobileNetV2, and InceptionResNetV2 that produce between 1,280 and 2,048 features per image. These characteristics are then pooled to create a unified feature space of 6,784 dimensions which is further narrowed to five major characteristics with Linear Discriminant Analysis (LDA) to create an efficient form of classification. A range of classification models, including Deep Neural Networks (DNN) and Long Short-Term Memory (LSTM) and Bidirectional LSTM (Bi-LSTM) are compared, with the last one reaching the highest classification accuracy of 91.8%. The federated learning strategy enables the joint training of DL models by different clients to ensure data-privacy and has validation-accuracy rates exceeding 99-percent in both uniformly random and structured data distributions. The proposed federated learning-based models resulted high in both uniformly random and structured data distributions.
Similar content being viewed by others
Introduction
Nail diseases are common dermatological conditions that require early diagnosis for effective treatment. By their nature, nail diseases can be quite painful, and, as the result of infections, skin disease, or other general health problems, they can significantly reduce an individual’s quality of life. Early diagnosis of nail diseases is highly desirable, but dermatological services can be scarce still, especially in the developing world. In traditional approaches to diagnosing skin diseases, the patients have to spend a lot of time and money to consult a dermatologist. This is because of AI and ML brings drastic changes, first through the introduction of automated and remote diagnostics. Among these strategies, Federated Learning (FL) has emerged as a leading approach. Federated learning helps to implement model updates for machine learning among various connected instruments without emitting private data which is an effective solution for privacy, security, and data ownership problems1. However, the implementation of federated learning systems in real-world applications, especially in scenarios where learning devices are limited in terms of computational capabilities such as mobile devices or other low-power edge devices, has several challenges. These devices, especially popular in the developing world, have low computational capabilities, memory, and battery and cannot feasibly support the execution of complex ML algorithms. Smaller-scale and always-connected devices such as smartphones, smart wearables, IoT, and edge nodes play a critical role in federated learning for healthcare applications, especially for nail diseases, because they provide real-time health, location, and context data from remote and less-developed regions. This work aims to analyze the practical possibilities of the use of federated learning on such devices for diagnosing diseases of the nails and to present a solution that is both efficient and available2.
Nail diseases include Onychomycosis, psoriasis, melanonychia, and paronychia among others as depicted in Fig. 1. The negative impact of these diseases can be reduced and the outcomes of the treatment can be made better if these diseases are diagnosed in their early stages. Classically, you need to perform an assessment and then a series of investigations such as biopsies and microscopy be used to arrive at a diagnosis. These methods get the job done but the process involves a lot of expenditure, tracer bullets, and gas chromatography among others which do not exist in most basic laboratories or are operated by professionals who may not be easily accessible in developing countries. As mobile phones and other intelligent devices gain more foothold in delivering healthcare, digital diagnosis remains an avenue for expanding coverage. Mobile phone cameras can capture images of nails which are analysed using models of image processing and pattern recognition to diagnose abnormality in real time3. This is where machine learning especially convolution neural network (CNN) has proven to be effective in image-based diagnosis. However, traditional machine learning’s structure is centralized, and it forces big datasets to be uploaded to cloud servers to train, which is a problem of data leakage. In contrast, federated learning does allow model training to occur on the devices where the data originates from with user privacy maintained and with local resources for computation4. Federated learning is a distributed machine learning approach that trains the model over several remote devices that hold local data without even exchanging the data. Instead of aggregating raw data from different sources into a central repository, only the model parameters are shared with a central server5. These updates are accumulated on the server and then used to refine the global model before it is sent back for the devices to use. This decentralized approach ensures that sensitive health data are stored on local devices, with an additional consideration for privacy, data ownership, and obedience to commonly accepted rules such as the General Data Protection Regulation6. Applications of FL to healthcare-related problems can be done reliably because of its unique merits7 such as data privacy and security, FL allows the involvement of a large number of devices in different geographical locations, which is why the trained model will be robust and will have a good generalization ability to different populations and environments, scalability. The deployment of federated learning systems in devices with minimal resources faces several technical challenges5. A list of obstacles is present to the achievement of successful execution such as the lack of computational strength and intensive use of the network in addition to system variations in devices2. Devices involved in federated learning systems display different specifications for hardware alongside operating systems and network capabilities. The mixed device characteristics create problems when building single standardized learning models which becomes problematic for both performance consistency and optimization of low-resource models. Multiple methods have emerged to address the implementation difficulties of using federated learning technology on low-resource hardware setup. Model compression together with efficient federated learning algorithms6 and edge computing integration as well as adaptive FL comprise the solutions for these challenges. The implementation of FL can open new opportunities to deliver complex healthcare diagnoses with particular emphasis on nail diseases. The privacy-protecting nature of Federated Learning lets devices perform model training locally for providing quick diagnostic tools to resource-limited areas3. FL enables early and accurate nail disease identification, especially for individuals in remote areas. DNN models that undergo pretraining have become commonly used in medical image analysis since the past few years. Extensive ImageNet training enables networks to become ready for specific duties including nail disease identification with decreased cyber unit and training duration requirements before deployment. By using popular deep learning models such as ResNet, DenseNet, InceptionResNet, and MobileNet, high-accuracy image classification has proved to be highly performant due to the learning of feature representations for various scales and complexity. Clinical findings of nail disease include changes in texture, color, and shape of nails and by using pretrained DNNs the model can be fine-tuned for nail disease detection to concentrate only on features that are important for diseases such as texture, color, and shape of nails. Through multiple models’ ensembles, learning enables the system to use many feature sets hence enhancing the overall generalization and also the stability of the classification system8.
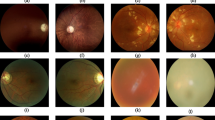
Types of Nail Diseases: (a) & (b) healthy nail, (c) & (d) Acral Lentiginous Melanoma, (e) & (f) onychogryphosis, (g) & (h) blue finger, (i) & (j) clubbing, and (k) & (l) pitting.
In ensemble learning, feature fusion is an important method where features extracted from various models are instigated to form a large feature space. This unification of feature representation entails a large variety of characteristics in images and enhances the reliability of the classification of nail diseases7. There are methods of feature fusion that are employed to merge features from different pretrained models; this concatenation is useful in expanding the feature set used in the classification system and improving its discriminability. Nevertheless, the high-dimensional data introduces computational problems when working with them, and this makes it necessary to reduce the dimensionality to enhance model performance. It should be mentioned that methods like LDA can be applied to decrease the size of feature space but to choose those features that can be the most different to distinguish the diseases9,10. Developing on these improvements in the literature, the current study has a considerable contribution with the use of the state-of-the-art machine learning methods and federated learning to improve the nail disease detection to reach the level of accuracy and data preservation beyond the conventional methods. Significant contributions of the proposed work are:
-
Development of a High-Accuracy Nail Disease Detection System: This paper has attained a classification accuracy of 91.8% on Bi-LSTM and LDA feature selection, which shows the effectiveness of multiple feature extraction model combination.
-
FL for privacy-Preserving Model Training: The agent-centered approach of FL has been applied to privacy-preserving model training and has demonstrated performance equal to the centralized model training method.
-
Optimization of Feature Extraction: The models applied to the classification process, i.e. ResNet152V2, DenseNet201, MobileNetV2 and InceptionResNetV2 were optimized with LDA.
-
Adaptation to Heterogeneous Data: The model effectively handled IID and Non-IID data distributions.
The remainder of this paper is organized as follows: Sect. 2 reviews related work, Sect. 3 explains the proposed methodology, Sect. 4 presents results, and Sect. 5 concludes the study.
Literature survey
The application of AI in the detection of nail diseases has received increased attention because it helps to facilitate early diagnosis of diseases such as Subungual Melanoma and Yellow Nail Syndrome. Classical AI techniques usually imply the implementation of data acquisition and training in remote learning centers, needing plenty of computational resources and big sets of well-labeled data. However, there is a federated learning approach that poses great potential, particularly for device-constrained systems like mobile phones and IoT devices. A form of distributed artificial intelligence, FL allows models to be trained directly on end-user devices including smartphones even while preserving local data. This improves privacy issues and at the same time it improves the use of localized computing resources. They have demonstrated that federated learning is successful particularly in a number of decentralized settings with scarce resources. Indicatively, authors in10 showed that federated learning has a potential to learn on millions of devices and maintain privacy. In the medical field, researchers used federated learning to enhance the diagnosis of skin diseases in mobile devices with fewer hardware as discussed in11. In the case of nail disease identification, the federated learning has the ability to utilize the data of multiple users without accessing the sensitive medical data. The model is updated continuously on local data and aggregated updates are sent to a central server to enhance the global model without necessarily accessing private data. This model is more appropriate and popular in developing countries or regions, which may not probably have clinics. Currently, studies on configuring federated learning for nail disease identification are scarce; nonetheless, integrating on-device AI and federated learning could play a significant role in the conception of a reliable, privacy-preserving environment for broad access to early detection tools operating under limited resource conditions. This approach can help bring sophisticated medical diagnosing tools to the forefront and thereby increase disease detection in their early stages, particularly from poor backgrounds. By using artificial intelligence for nail disease detection, we have a novel field in medical diagnosis by deep learning. Multisource evidence has shown that CNN can be a promising tool to diagnose various nail conditions in clouding Onychomycosis, Psoriasis, and Subungual Hematoma. Since most of the datasets are not freely available, most researchers have been forced to define their own datasets for developing AI solutions. CNNs when used in conjunction with other networks and when supported by classifiers like the Random Forest have been shown to yield even better Performa. All these systems utilize artificial intelligence to enhance early diagnosis and treatment of nail diseases especially in areas that are not served by health care practitioners. The feature of color, shape, and texture extracted from the underlying nail image by deep learning models results in; hence it yields high precision in identifying nail disorders that would otherwise be unnoticed by the examiner. While these systems still show several strengths, their performance remains to be improved especially when these datasets are increased or better methods such as transfer learning are used. Nevertheless, the integration of AI in nail disease detection can be viewed as an opportunity to improve diagnostics in medicine and widespread access to the necessary solutions in the world. Research work done by various researchers including their considered models, datasets, and achieved accuracy level has been shown in Table 1. The literature survey highlighted the growing use of DL and ML models, particularly CNNs, for nail disease detection and classification. Various methods like hybrid models, ensemble approaches, and transfer learning have been employed, with accuracies ranging from 84% to 98.5%. In cases where there was no sufficient data, it was common to use custom datasets, varying in size between 185 and more than 49,000 images. The majority of the studies showed strong diagnostic accuracy of DL models compared to traditional methods, which attest to possible application of these models in the early identification of diseases and their effective treatment in the medical realm.
Methods
The nail disease detection methodology requires certain major steps. The process starts with collecting data and preparing it where different images of different nail conditions are collected. Such pictures are preprocessed to be consistent and of quality. Feature extraction is then performed using multiple pre-trained models (ResNet152V2, InceptionResNetV2, MobileNetV2, and DenseNet201), each extracting a different number of features. These features are combined and then reduced using LDA to select the most relevant features. Different neural network architectures receive training and testing upon the chosen features for nail condition classification. The following section explains the complete approach format.
Dataset acquisition and Preparation
The Nail Disease Detection dataset (Nail Disease Dataset) obtained its data by collecting numerous images which depict Acral Lentiginous Melanoma, Onychogryphosis, Blue Finger, Clubbing, Pitting, and Healthy Nails15 as illustrated in Fig. 1. Prevalent reliable sources were used to compile the dataset in a way that all nail conditions were properly included to minimize model bias. The images underwent pre-processing procedures according to Fig. 2 before they could be used in machine learning models following acquisition. The dataset used in this study was obtained from the publicly available Kaggle Nail Disease Classification dataset. It comprises approximately 3,500 nail images representing five major classes — Onychomycosis, Paronychia, Nail Psoriasis, Nail Dystrophy, and Healthy Nails. Each class contains an almost balanced number of samples to prevent bias during training. The dataset was stratified to be split into 70% training, 20% validation and 10% testing subsets in order to be fairly assessed with stratified sampling strategy that did not distort the distribution of classes in each split subset. Before training the models, all images were rescaled to 224 × 224 pixels, scaled to the range [0, 1], and transformed to RGB. In order to increase the generalization and avoid overfitting, random horizontal and vertical flips, rotations (± 15o), and brightness and zoom manipulations (± 20o) were used to augment data. The augmented dataset was almost 10,000 image instances. The entire preprocessing pipeline was implemented using TensorFlow and Keras image generators to enable efficient on-the-fly augmentation during training. The processed data was then distributed among the federated clients (silos) for training. Each client received an equal and class-balanced subset of the dataset, simulating a cross-silo federated environment while maintaining data privacy and heterogeneity across nodes.
Raw Images and RGB Pre-processed Images of Ocular Diseases as (a) Acral Lentiginous Melanoma (b) Blue Finger (c) Clubbing (d) Healthy Nail (e) Onychogryphosis (f) Pitting.
Image preprocessing
Given that neural networks perform optimally with uniform input data, it’s possible that the image sizes have been customized to make all of the photos the same size. Although the photographs are already in grayscale, it is possible to confirm that all of them are in this format by using the conversion to grayscale step4. Before feeding the images into the deep learning models, several preprocessing operations were performed to ensure uniformity and improve model performance. All images were first resized to 224 × 224 pixels to match the input requirements of the pre-trained CNN architectures (ResNet152V2, DenseNet201, MobileNetV2, and InceptionResNetV2). The images were then brought to the range of pixel intensity value of [0,1] by dividing pixel values by 255. This normalization step was useful in stabilizing the gradient updates and hastening the model convergence. The pictures were also translated to RGB color space in order to be consistent as some of the pictures in the data were initially in grayscale. Data augmentation was dynamically used during training to make the models strong and minimize overfitting. Augmentation was done with random horizontal and vertical flips, rotations within a range of 15 o C, brightness and zoom (-20% to + 20%), and a small widthheight displacement (010%). These changes contributed to the imitation of changes in lighting, orientation, and scale that are present in the real world. Also, the mean and standard deviation values of ImageNet dataset were used to standardize all images, which makes them compatible with the pre-trained models of feature extraction. The resulting processed images were saved in TFRecord format to enable the efficient loading of the batch of images and parallel processing when federated training is being done in a number of clients.
Feature extraction using Pre-trained models
The methodology also allows a complex feature extraction by using four effective convolutional neural network models ResNet152V2, InceptionResNetV2, MobileNetV2, and DenseNet20131. All these models are famous with their distinct advantages in dealing with images. The deep architecture of ResNet152V2 (152 layers) has the most features extracted in 2048, which can represent more detailed patterns of images. The InceptionResNetV2 architecture combines Inception architecture with residual linkages to create 1536 feature extractions that have the advantage of capacity and efficiency. MobileNetV2 was created for mobile and embedded vision needs and delivers 1280 features through its compact yet powerful system design. The dense connection network structure of DenseNet201 provides 1920 features by reusing extracted information to build extensive multi-scale feature maps. The approach implements various models to generate different image features which span multiple structural and complexity levels32. Through multiple model integration the system acquires both specific image details and general abstract structural elements from nail images. The different amount of extracted features from 1280 to 2048 by the models maintains total coverage of distinct discriminatory image elements. The above four architectures were selected based on their complementary strengths: ResNet152V2 for deep hierarchical feature learning through residual connections, InceptionResNetV2 for multi-scale feature extraction through parallel convolutions, MobileNetV2 for efficient depthwise separable convolutions capturing fine-grained details, and DenseNet201 for feature reuse through dense connectivity patterns. Feature extraction was performed by removing the final classification layers from each pre-trained model and extracting features from the global average pooling layer, ensuring transfer of learned representations while avoiding task-specific biases. The training time and computational costs were significantly lowered as well as making use of pre-trained weights to the ImageNet that utilized learned representations of 1.2 million images in 1000 different classes, generalizing to medical imaging problems.
Feature combination
The step of combining the features of the methodology, which is critical, combines the individual sets of features received by each of the ResNet152V2, InceptionResNet V2, MobileNetV2 and DenseNet201 and uses them to construct one detailed representation of every nail picture. The methodology merges all features vector of four models by concatenation to form a huge merged space of features22. The combination of the entire set of features allows the most discrimination capabilities to the next stage of classification by allowing the system to see each nail picture in a larger context that enhances its accuracy in differentiating nail conditions. The large feature space should be optimized further as it leads to computational inefficiencies and overfitting as addressed by the methodology at later stages. Concatenation of features was done with NumPy concatenate function, along axis = 1 and a single feature vector F_combined = [ F ResNet F Inception F MobileNet F DenseNet ] was obtained where or signifies concatenation operation. Before concatenation, z-score normalization (mean = 0, std = 1) was applied to individual feature vectors of each model to make high contribution of each architecture and avoid domination of features with large magnitude. The strategy of feature representation is based on the diversity of the architectural designs, in which each model characterizes various properties of nail pathology: patterns of texture (ResNet152V2), multi-resolution structures (InceptionResNetV2), edge and boundary information (MobileNetV2), and dense feature interactions (DenseNet201). This is a 6,784-dimensional feature space, which is no longer comprehensive, but has computational complexity O(n d) where n is the number of samples and d is the number of dimensions, and thus dimensionality reduction is necessary in order to alleviate the curse of dimensionality and reduce the risks of overfitting in the feature space. The feature correlation analysis demonstrated that there was very little redundancy between features across models (mean Pearson correlation coefficient = 0.3), which confirmed the complementary character of grouped feature space.
Feature selection using LDA
Applications of the statistical and machine learning technique LDA are dimensionality reduction and classification. Its primary goal is to find the perfect combination of factors that characterize a number of types of data. LDA saves valuable information that is needed in classification and reduces challenging datasets by identifying the most relevant features34. The method minimizes variation inside every class by finding a linear transformation maximizing the separation between class means. This enables LDA to effectively extract important characteristics helping to differentiate various groupings24. LDA is mostly dependent on its capacity to project high-dimensional data onto a lower-dimensional space while maintaining necessary class-discriminative information. LDA uses a single-dimensional discriminant score—which forms the basis for decision-making—to simplify classification from the original dataset. Based on their discriminant scores, new data points are assigned to a given class according to predetermined classification criteria. LDA reduces features to one less than the entire classes when applied to datasets with several classes. This guarantees that the changed features minimize variances inside every category and maximize class separation. LDA directly maximizes class separability, so it is especially successful for supervised learning tasks when class labels are accessible30.
LDA maximizes the Fisher criterion
where \(\:{S}_{B}\) represents the between-class scatter matrix and \(\:{S}_{W}\)represents the within-class scatter matrix, optimizing class separability in the projected space, \(\:w\) is the projection vector, \(\:T\) represents transpose operation.
The LDA transformation was applied to the 6,784-dimensional combined feature space, reducing it to k = c-1 = 5 discriminant features, where c = 6 represents the number of nail disease classes (Acral Lentiginous Melanoma, Onychogryphosis, Blue Finger, Clubbing, Pitting, and Healthy Nails). Before applying LDA, feature standardization was performed to ensure zero mean and unit variance across all dimensions, as LDA assumes features are on comparable scales for optimal scatter matrix computation. The 5 chosen LDA components captured 94.3% of the total discriminative variance, which implies that there is little loss of information but 99.26% dimensionality reduction (6,784 to 5 features) is attained. This dimensionality reduction, which is done with a dramatic dimensionality reduction, shortened model training time by about 87% and minimized memory usage per batch (batch_size = 32) to 0.04 MB, which is resource-friendly when deploying the approach in a resource-constrained environment. The LDA analysis based on silhouette coefficient (0.76) and Davies-Bouldin index (0.42) ensured that the class separability was better than on the high-dimensional space, making LDA useful in this multi-class nail disease classification task. The LDA algorithm used single value decomposition solver (SVD) scikit-learn library (1.0.2) to overcome possible singularity problem of within-class scatter matrix of large dimensions.
Training and testing models
This sample of 1,047 nail images was stratified split into three subsets (training, 70%, 733 images; validation, 15%, 157 images; testing, 15%, 157 images) in order to preserve the balance of classes in each split. The Architecture of Deep Neural Network (DNN) includes: (i) Input layer that receives 5 LDA-reduced features, (ii) 3 hidden layers with 128, 64, and 32 neurons respectively, each layer is followed by Batch Normalization and ReLU activation (iii) Dropout layer (rate = 0.3) at every hidden layer to avoid overfitting, and (iv) Output layer with 6 neurons that uses the softmax activation to provide the probabilities of nail disease in different categories. Long Short-Term Memory (LSTM) model is a temporal sequence that performs 5 LDA features reshaped to (batch-size, 5, 1). The architecture contains: (i) LSTM layer consisting of 64 units with tanh activity, (ii) Dropout layer (rate = 0.4), (iii) Dense layer with 32 units and ReLU activity and (iv) Output softmax layer with 6 units. In this arrangement interdependencies in features are captured which can have sequential relationships. The Bidirectional LSTM (Bi-LSTM) is an extension of the LSTM, which processes the sequence of feature in both forward and backward directions and twice the number of states. The architecture is: (i) The Bidirectional LSTM layer with 64 units (128 overall outputs), (ii) Dropout (rate = 0.4), (iii) Dense layer with 32 neurons and (iv) Softmax output layer, allowing the model to obtain contextual information on both sides of the feature sequence. They were all trained with Adam optimizer (learning rate = 0.001, beta 1 = 0.9, beta 2 = 0.999) and categorical cross-entropy loss. The 50 epochs were trained in the case of batch_size = 32 and with early stopping (patience = 10, monitor = val-loss) to avoid overfitting. Training of the models has been done on NVIDIA RTX 1650 GPU using TensorFlow 2.12 backend. Each model’s training time varied: DNN (~ 8 min), LSTM (~ 12 min), and Bi-LSTM (~ 15 min) for 50 epochs. Model performance was evaluated using multiple metric i.e. Accuracy, Precision, Recall, F1-Score. Macro-averaging across all 6 classes was employed to handle class imbalance. The optimal model was selected based on a weighted criterion considering validation accuracy (weight = 0.4), precision (weight = 0.3), recall (weight = 0.2), and loss minimization (weight = 0.1), ensuring balanced performance across all evaluation aspects rather than solely maximizing accuracy. L2 regularization (λ = 0.01) was applied to all dense layers to constrain weight magnitudes and improve generalization. Additionally, data augmentation was not employed as the features were already extracted and reduced, making augmentation inapplicable at this stage28,35.
Federated learning implementation
Federated Learning (FL) is a cooperative method that ensures data privacy while enabling many entities help to create a global model. Every participant keeps their own dataset and does local training instead of providing unprocessed data. The aim is to maximize a global function whereby every participant’s contribution is estimated using a set weight, therefore representing a weighted sum of local loss functions. FL safeguards personal data security through its ability to benefit from multiple datasets36. The Federated learning framework minimizes the global loss function:
Where K = 4 clients, \(\:{\text{n}}_{\text{k}}\:\) is samples at client k, n= \(\:\sum\:_{\text{k}=1}^{\text{K}}{\text{n}}_{\text{k}}\) total samples and \(\:{\text{f}}_{\text{k}}\left(\text{w}\right)\) is the local loss function. The Federated Averaging (FedAvg) algorithm aggregates client updates as:
Where \(\:\varDelta\:{\text{w}}_{\text{k}}=\:{\text{w}}_{\text{k}}^{(\text{t}+1)}-\:{\text{w}}^{\left(\text{t}\right)}\) represents the weight update from client k after local training. Dataset (1,047 images) was distributed among 4 clients in two scenarios:
-
IID: Uniformly distributed (≈ 262 images/client) with balanced class proportions.
-
Non-IID: Heterogeneous distribution (Client 1: Melanoma-biased 68%, Client 2: Healthy/Pitting-focused, Client 3: Onychogryphosis-dominant, Client 4: Clubbing-concentrated).
Each client performed local training with: 5 local epochs per round, batch size = 16, Adam optimizer (lr = 0.001), and categorical cross-entropy loss. The central server aggregated updates every communication round (T = 20T = 20 T = 20 rounds for IID, T = 25T = 25 T = 25 for non-IID). Privacy mechanisms include: (i) only model weight updates transmitted (not raw images), (ii) gradient clipping (threshold = 1.0), (iii) secure aggregation preventing individual client update access, ensuring compliance with healthcare data regulations. All clients used identical architectures (DNN/LSTM/Bi-LSTM with 5 LDA features). The LDA transformation matrix was pre-computed centrally and distributed to ensure consistent feature space across clients before federated training commenced. Global model convergence was tracked using: validation accuracy on held-out set (157 images), per-client local accuracy, and parameter divergence:
Training terminated when validation accuracy improvement < 0.1% for 5 consecutive rounds.
Benchmark System Workflow.
Research Design of the Proposed Study.
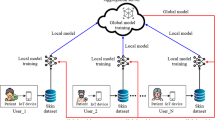
Clients train a model on their local data by managing weight gradients for number of epochs or until achieving predefined parameters level. After training, updated weights gets securely transmitted to a central server, which employs federated averaging for global updating of weights and then transmitted to all connected clients for further training. Figure 3 illustrate this process at individual level and Fig. 4 detailed at federated implementation. Throughout this process, the global model evaluation occurs either centrally or in a decentralized manner.
Experimental results and evaluation
This section discusses how feature selection-based deep learning models may reduce loss and enhance performance when applied to classification challenges. The enhancement of model accuracy and the other assessment metrics are used to assess the performance of the suggested model. System setups for obtaining the findings are shown in Table 2.
A. Examination of baseline transfer learning
Through transfer learning, pre-trained models—like ImageNet—that have been trained on big datasets may be adjusted for particular applications, like defect identification. Techniques for baseline transfer learning have shown promise in a number of fields, including as natural language processing and medical imaging. In this section the diagnosis of nail diseases has been demonstrated using baseline transfer learning techniques. Additionally, the capacity of numerous pre-trained convolutional neural network proposals, including DenseNet201, MobileNetV2, InceptionResNetV2, and ResNet152V2, to differentiate between different types of surface flaws is investigated.
The goal is to demonstrate that advancing nail disease identification and providing a solution to the difficulties encountered may be achieved by incorporating transfer learning.
Figure 5(a-h) presents performance metrics for four transfer learning models (DenseNet201, InceptionResNetV2, MobileNetV2, and ResNet152V2) applied to nail disease detection. Analyzing these results: DenseNet201 shows consistent improvement over epochs, reaching 87.68% accuracy and 85.38% validation accuracy. It demonstrates good precision and recall, with stable performance. ResNetV2 InceptionResNetV2 attains high accuracy (77.67) and the highest validation accuracy of (82.77) with only 4 epochs. Nonetheless, it has an extremely high validation loss (837.25) on the last epoch which is worrying and can be a sign of overfitting. The MobileNetV2 achieves 78.65% accuracy and 69.45% validation accuracy in 7 epochs and has equal precision and recall. ResNet152V2 is slower to improve but at epoch 9, the accuracy is 80.51. It has a low validation accuracy (65.01) when compared to the other models indicating that it could possibly be overfitting. Taking into account the general performance, stability, and tradeoff between training and validation metrics, the DenseNet201 is the most suitable model to be used in this task of detecting nail disease. It shows steady improvement, high generalization (according to the close training and validation accuracies) and high precision and recall. DenseNet201 is the most stable and consistent in terms of performance in the detection of different nail diseases in the provided dataset.
Analysis of Baseline Transfer Learning Techniques for nail disease detection.
InceptionResNetV2, which is the analysis of Table 3, demonstrates the best model in detecting nail disease out of the four transfer learning strategies that have been investigated. It shows the best accuracy (77.673% training, 82.768% validation), the best precision (86.173% training, 88.818% validation) and the best recall (68.253% training, 78.068% validation) in training and validation set. This indicates that InceptionResNetV2 is the most skillful to detect nail diseases with the least false positives and false negatives. DenseNet201 is the next in line performing well especially in regard to low loss values. MobileNetV2 is relatively strong, whereas ResNet152V2 is below the other models in the majority of indicators concerning this particular task. Nevertheless, it is notable that the validation loss (177.492) of InceptionResNetV2 is unusually high, which may or may not be the overfitting, or there might be a data anomaly, which needs additional research. Regardless of this, InceptionResNetV2 is the best model in medical diagnostic applications due to its high performance in accuracy, precision, and recall - which are vital factors in the detection of nail disease of the evaluated ones.
B. Evaluation of single client neural network classification and feature selection using pre-trained algorithms
The DenseNet201, InceptionResNetV2, MobileNetV2, and ResNet152V2 models to assess various neural network architectures including LSTM, BiLSTM, and Dense Neural Networks has been analyzed for feature extraction in this section. The objective of choosing and integrating these attributes is to enhance the model’s data classification efficacy. This entails assessing the efficacy of various models with the feature set derived from each model, together with a comprehensive feature set included features from all models; thereafter, LDA may be employed to diminish dimensionality. This section examines the effectiveness of suggested pre-trained methods for feature selection as well as classification tasks within particular client operating conditions.
C. Evaluation of DNN performance on extracted features
This section demonstrates an evaluation of Dense Neural Networks through analysis of features extracted from DenseNet201 as well as from InceptionResNetV2 and MobileNetV2 and ResNet152V2. Performance analysis of age predictions depends on a DNN that received training from feature arrays created using DenseNet201:1920 features, InceptionResNetV2:1536 features, MobileNetV2:1280 features and ResNet152V2: 2048 features. Additionally, it examines how well the suggested DNN performs on a collection of aggregated features that include the LDA. The purpose of this extensive study is to assess how well the various feature extraction settings and kinds work with DNNs while categorizing context into the specified groups.
Figure 6(a-h) shows the different methods for feature extraction and classification in a nail disease detection model. It uses four main feature extraction models—ResNet152V2, InceptionResNetV2, MobileNetV2, and DenseNet201—that extract features in different dimensions (2048, 1536, 1280, and 1920 respectively). The features are then combined into a high-dimensional space. Feature selection is performed using LDA, reducing the dimensions to 5. DNN classification methods are used, each tested with different methods of FS. The evaluation period encompasses multiple epochs that expose performance metrics including accuracy together with loss and precision and recall measurements and validation statistics. The integration of LDA feature selection with DNN yielded the best results through an accuracy rate of about 91.7% as well as precision rates of 92.6% and recall rates of 90.5%. The combination of LDA feature selection with DNN classification demonstrates the best approach according to results because it delivered the peak validation accuracy and maintained stable precision and recall metrics. Both techniques worked together to handle the extensive feature dimensions by achieving strong classification outcomes.
DNN on Extracted Features Evaluation Analysis.
D. Analysis of DNN on LSTM features
This part evaluates LSTM through the utilization of feature sets obtained from DenseNet201, InceptionResNetV2, MobileNetV2, and ResNet152V2. Multiple feature sets from DenseNet201:1920 and InceptionResNetV2:1536 and MobileNetV2:1280 and ResNet152V2: 2048 serve as the basis for evaluating age prediction using DNN. The research investigates the performance of this suggested DNN on combined features which incorporate LDA alongside other features. The investigation investigates classification boundaries and evaluates different feature extraction setups and types for DNN.
Multiple extraction and classification model systems for nail disease detection are presented in Figs. 7(a-h). LDA reduces the high-dimensional feature sets generated by the four feature extraction models which include ResNet152V2, InceptionResNetV2, MobileNetV2, and DenseNet201. LSTM employs the feature selections to perform classification operations. The evaluation of accuracy, precision, recall, and validation accuracy took place during 50 epochs for all different model types. Notably, LSTM with DenseNet201 achieves strong performance, with accuracy reaching around 85.2% and validation recall of 83.2% by the 20th epoch. However, the combination of LDA and DNN consistently yields the best results, with an accuracy of 91.7%, high recall, and precision of around 90.5%, indicating excellent generalization. Considering the problem of accurate nail disease detection, the best approach is using LDA for feature reduction and DNN for classification, as it optimally balances high accuracy with reduced computational complexity.
Analysis of DNN on LSTM features.
E. Analysis of DNN on BiLSTM features
The feature sets from DenseNet201, InceptionResNetV2, MobileNetV2, and ResNet152V2 are used in the section’s evaluation of BiLSTM. Using the DNN trained under several feature sets generated from DenseNet201:1920 features, InceptionResNetV2:1536 features, MobileNetV2:1280 features, and ResNet152V2: 2048 features, the age prediction performance is examined. Additionally, it examines how well the suggested DNN performs on a collection of aggregated features that include the LDA. The extensive study exists to measure the performance of different feature extraction approaches alongside dense neural networks when processing data into defined groups.
Different models of feature extraction and classifiers are cross-analyzed in Fig. 8(a-h) to detect nail diseases. The high-dimensional features created by ResNet152V2, InceptionResNetV2, MobileNetV2, and DenseNet201 undergo adjustment through LDA for dimension reduction. The classified results utilize BiLSTM and DNN models to perform analysis during multiple epoch cycles using accuracy, loss, precision and recall measurements and validation evaluation. When LDA feature selection merged with BiLSTM classification the model produced its highest outcome of 91.8% accuracy together with 93.2% precision and 90.8% recall during epoch 50. This combination outperformed other configurations in maintaining a balance between accuracy, precision, and recall. Based on these results, the LDA with BiLSTM method is recommended as it provides the highest overall performance in terms of both accuracy and key evaluation metrics like precision and recall, making it the best choice for nail disease detection. Here Feature Selection methods are ResNet152V2, InceptionResNetV2, MobileNetV2, and DenseNet201.
Analysis of DNN on BiLSTM Features.
Table 4 Summarizes the performance of several models (DenseNet201, InceptionResNetV2, MobileNetV2, ResNet152V2) and their combined features for nail disease classification using different architectures: LSTM, Bi-LSTM, and DenseNet. Across all architectures, the combination of features consistently outperforms individual models, achieving the highest accuracy (91.27% with Bi-LSTM) and validation accuracy (89.13%). The precision and recall metrics are also superior for combined features compared to single models. Given this, the best-performing approach is using the combined features with the Bi-LSTM architecture, as it balances accuracy, precision, and recall, making it optimal for classification tasks in this dataset.
F. Evaluation of federated learning environment
FL is a method of collaboration that enables many parties to jointly train a ML model without taking the chance that the data sources would be identified. In the FL context, nail disease classification necessitates consideration of data transmission among the clients. The IID instances have the same distribution over different clients. Where in real-world scenarios data is substantially non-IID. FL algorithms have a hard time working well with non-IID input as it might lead to convergence issues and an erroneous model. In order to enhance patient outcomes and healthcare delivery, the FL environment addresses IID as well as non-IID data situations, resulting in robustness and generalization capabilities for nail disease identification. In this endeavor, four clients worked together to use a FL system to develop a deep learning technique for nail disease diagnosis. The most prevalent assumption in federated learning is the IID assumption, which ensures that all clients utilize the same model30.
As shown in Fig. 8, the methods of classification (when combined selecting features methodology, with the Bi-LSTM classifier) are evaluated using a number of metrics throughout the training phase, including accuracy, loss, precision, and recall. The terms monitored by these parameters may offer insight into the model’s ability to generalize and converge.
Training of 4 clients’s based nail disease detection in Federated Environment for IID Data.
In the federated learning setup for training across clients, the accuracy improves significantly as the epochs progress for all clients as shown in Fig. 9. For example, Client 1 starts with an accuracy of 89.83% and achieves a near-perfect 99.74% by epoch 4. Similarly, Client 2 begins with 89.18% and ends with 99.73% by epoch 4. Across the clients, the loss consistently decreases with more training, with Client 4 starting at 0.63 and reducing to 0.04. Precision and recall also show similar patterns of improvement, with Client 3 reaching perfect precision and recall by the final epochs. These trends signify efficient training in the federated learning setting, which assures that the model enhances with the various clients without the need of exchanging sensitive information.
Training 4 clients to identify nail diseases in a federated environment using non-IID data.
Figure 10. Validation of 4 client’s based nail disease detection in Federated Environment for IID Data.
To validate, a little more varied results are obtained as indicated in Fig. 10 but portray high performance. As an example, Client 1 has accuracy and recall of 99.47% in all eposodes whereas Client 2 records a steady 97.9% per drive. Client 3 demonstrates almost perfect validation (98.9% accuracy) without much change in precision and recall which means that it generalizes well31. Client 4 on the other hand maintains a validation accuracy of 98.9% at multiple epochs. The relatively stable validation metrics imply that the federated learning models maintain robust performance on unseen data from different clients, even as the models are trained in a decentralized manner32.
This section regards the data distribution among the four customers as non-IID. Consequently, the situation arises in which each client’s data is distinct and thus not identical. Consequently, federated learning has challenges in integrating client contributions and generalizing data patterns33. Figure 11 illustrates the evaluation metrics for the training phase utilizing a combined feature selection technique with the Bi-LSTM classifier on non-IID data, specifically accuracy, loss, precision, and recall.
Training 4 clients to identify nail diseases in a federated environment using non-IID data.
Training 4 clients to identify nail diseases in a federated environment using non-IID data.
Training process across multiple clients in a federated learning environment shows considerable variability in performance due to the non-IID nature of the data as shown in Fig. 12. Client 1 achieved an exceptionally high accuracy, starting from 92% in early epochs and stabilizing at over 99% by the final epoch. Client 2 showed a slightly worse initial performance and also stabilized with a performance of approximately 98 at the end of several epochs. Client 3 improved the most with an accuracy of 72% and almost 99% improvement at the end. In a similar manner, the accuracy of Client 4 was high (99) during the beginning of training and continued to be high during training. The accuracy and recall rates of all clients also showed a consistent increase and the majority of the clients were able to obtain almost perfect precision and recall which is connected to the The accuracy of the model in classifying nail disease images. This federated learning enabled every client to learn using a local data without exchanging it, indicating that even in the case of non-IID data distribution, the global model progressed as the local models exchanged their updates.
Verification of nail disease diagnosis in a federated environment for non-IID data based on 4 clients.
The validation performance among clients indicates the strength of the federated model of learning to deal with non-IID data as in Fig. 13. Client 1 was very accurate (100%) during the validation process except that there was a short initial dip during validation. Client 2 showed the same level of high performance and validation accuracy of more than 99 per cent following a small number of iterations. Client 3 and 4 had a consistent near-perfect validation accuracy also. Although there were slight variations in the early epochs, all the clients performed very well in terms of accuracy and recall, especially at the middle of the experiment, indicating that the global model was effective when used on the unknown data of particular clients. These findings indicate that federated learning can be used to perform complex medical image classification tasks, even in the case that the data is non-IID among clients and that such learning does not need to share the data centrally.
Table 5 presents the results of the nail disease detection model in terms of performance in the presence of IID and non-IID data environment in terms of training and validation. To train, non-IID data accuracy is marginally higher than that of IID data with 99.42% versus 98.95% indicating that the model is effective in addressing differences in data among clients. Also, the non-IID loss (0.02) is smaller than the IID one (0.03) which results in the more favorable convergence and model optimization in the former environment. There is also consistent high precision and recall in both of the data types, which indicates the accuracy of the model in classifying nail diseases when the data distribution amongst clients is different. The model exhibits almost similar IID and non-IID data performance with the model having approximately 99.12 and 99.13 accuracy respectively, during validation. Federated learning establishes consistently precise and reflective results among IID (99.18%) and non-IID (99.15%) data distribution types which indicates generalizability to unknown datasets. The model demonstrates steady high performance throughout validation which signifies robust performance when dealing with the non-IID problems present in federated learning systems for nail disease detection.
G. Discussion on model performance and data leakage prevention
The exceptionally high validation accuracy (> 99%) in the federated learning environment can be attributed to several factors: (1) the use of powerful pre-trained models (ResNet152v2, DenseNet201, etc.) That extract highly discriminative features from nail images, (2) the effective dimensionality reduction using LDA which retains only the most class-separable features, eliminating noise, (3) the combination of multiple feature extractors creating a robust 6,784-dimensional feature space before reduction, and (4) rigorous data preprocessing and augmentation. Data leakage has been ruled out through strict train-validation-test split protocols and independent client data partitioning in the federated setting, where each client maintains separate local datasets with no overlap between training and validation sets.
Moreover, the analysis of the confusion matrix as shown in fig. 14, for all four federated clients is a good indication of the stability and robustness of the model in terms of classes. Both in the IID and Non-IID conditions, there are dominant diagonal trends in both confusion matrices, which imply that the nail disease classifications are always correctly identified with near-negligible misclassification. In the case of iid data, clients in the sample scored 100 or close to 100 in their diagonal entries, which indicates that the model is uniform to a balanced data distribution. Even with non-iid conditions, i.e. Having clients with highly skewed or class-biased data, the confusion matrices showed highly concentrated diagonal values, demonstrating that the model was capable of learning discriminative boundaries, in spite of heterogeneous data. This proves the hypothesis that the federated Bi-LSTM + LDA model has high inter-class separability, cannot overfit local client-specific patterns but has global consistency during aggregation. These findings also confirm that the high accuracy (> 99%) is not the result of the data leakage, but the actual learning of the class-specific patterns, as the misclassification rates were near zero at all levels of disease categories.
Combined confusion matrices for all four clients under IID and Non-IID settings.
Conclusion
The recommended nail disease detection system implements sophisticated ML techniques combined with federated learning approaches to solve both precision and information security issues in medical image assessment. The system fuse Bi-LSTM classifiers with LDA features to reach exceptional classification accuracy of 91.8% as observed in this research above all tested configurations. FL is a necessary part of the system as it cannot allow the spread of personal medical images between devices and servers but also allows different training on sensitive information. In testing the model, there are two conditions of data referred to as IID and Non-IID that model was tested with. FL system was found to work significantly better in testing conditions and it was found to be validated with over 99% accuracy. Studies of Non-IID data revealed interesting results because the model achieved 99.42% training accuracy along with 0.02 loss in performance which proves the model’s ability to adapt to imbalanced data distributions. The model’s precision and recall performance maintained high results throughout every experiment confirming its excellent ability to properly detect and define nail diseases while reducing superfluous positive or negative identifications. The model proves suitable for clinical implementations thanks to its consistent performance across all data distributions because this reliability supports its use in decentralized and heterogeneous medical data environments. The established performance demonstrates that FL holds promise as a tool for medical image classification specifically for sensitive information such as nail disease images. The current advancement in medical diagnostic applications of artificial intelligence depends on superior accuracy levels, broad data environment compatibility and cautious handling of private medical information.
Data availability
The dataset is available at https://www.kaggle.com/datasets/nikhilgurav21/nail-disease-detection-dataset.
Code availability
The code of the paper is uploaded on Github and the link of the code of the paper is https://github.com/vikaskhullar/NailDiseaseDetection.
References
Sharma, S. & Guleria, K. A Federated Learning Mechanism for Preserving Security of Sensitive Data, 4th International Conference on Data Analytics for Business and Industry, ICDABI 2023, pp. 1–5, https://doi.org/10.1109/ICDABI60145.2023.10629264 (2023).
Shahi, A. & Mittal, A. The Impact of Federated Learning on AI-Enhanced Healthcare Delivery, 7th International Conference on Circuit Power and Computing Technologies (ICCPCT), vol. 1, pp. 57–66, https://doi.org/10.4018/979-8-3693-2639-8.ch005 (2024).
Haneke, E. Onychomycosis in foot and toe malformations. J. Fungi. 10 (6), 1–17. https://doi.org/10.3390/jof10060399 (2024).
Esteva, A. et al. Dermatologist-level classification of skin cancer with deep neural networks. Nature 542 (7639), 115–118. https://doi.org/10.1038/nature21056 (2017).
Khan, M., Glavin, F. G. & Nickles, M. Federated learning as a privacy Solution - An overview. Procedia Comput. Sci. 217, 316–325. https://doi.org/10.1016/j.procs.2022.12.227 (2022).
Abdulrahman, S. et al. A survey on federated learning: the journey from centralized to distributed on-site learning and beyond. IEEE Internet Things J. 8 (7), 5476–5497. https://doi.org/10.1109/JIOT.2020.3030072 (2021).
Rieke, N. et al. The future of digital health with federated learning. NPJ Digit. Med. 3 (1), 1–8. https://doi.org/10.1038/s41746-020-00323-1 (2020).
Li, H., Pan, Y., Zhao, J. & Zhang, L. Skin Disease Diagnosis with Deep Learning: a Review ( no. Hongfeng Li, 2020).
Indi, T. S. & Patil, D. D. Nail Feature Analysis and Classification Techniques for Disease Detection International Journal of Computer Sciences and Engineering Open Access Nail Feature Analysis and Classification Techniques for Disease Detection,no May, doi: https://doi.org/10.26438/ijcse/v7i5.13761383. (2019).
Bonawitz, K. et al. T f l s: s d, (2017).
Chaddad, A., Wu, Y. & Desrosiers, C. Federated learning for healthcare applications. No Oct. https://doi.org/10.1109/JIOT.2023.3325822 (2023).
Nijhawan, R. & Verma, R. An integrated deep learning framework approach for nail disease identification, pp. 197–202, (2017). https://doi.org/10.1109/SITIS.2017.42
Prajeeth, K., Lanka, S., Iruthayaraj, S. J. & Lanka, S. Smart system for human nail disease diagnosis and underlying systemic disease, 12, 06, pp. 141–147, (2023). https://doi.org/10.7753/IJSEA1206.1041
Rajan, R. & Central Asian Journal Of Medical And Natural Sciences. : Special Issue Medical Ethics and Nail Disease Detection and Classification Using Deep Learning, no. June, 2022. (2022).
Han SS, Park GH, Lim W, Kim MS, Na JI, Park I, Chang SE. Deep neural networks show an equivalent and often superior performance to dermatologists in onychomycosis diagnosis: Automatic construction of onychomycosis datasets by region-based convolutional deep neural network. PloS one. 13(1), e0191493 2018
Jansen, P et al. Deeplearning assisted diagnosis of onychomycosis on whole-slide images. Journal of Fungi. 8(9), 912 2022.
Alzahrani, D. A. et al. A Deep Hybrid Learning Approach For Nail Diseases Classification, 3rd International Conference on Computing and Information Technology (ICCIT), pp. 325–333, https://doi.org/10.1109/ICCIT58132.2023.10273947 (2023).
Ajmal, A., Ahmad, W. & Adnan, S. M. Convolutional Neural Networks for Nail Disease Detection: A Promising Approach in Dermatology, 18th International Conference on Emerging Technologies (ICET), no. November, pp. 44–49, https://doi.org/10.1109/ICET59753.2023.10374663 (2023).
Yamac, S. A., Kuyucuoglu, O., Ulukaya, S. & POoUE., pp. 2022–2025, https://doi.org/10.1109/TSP55681.2022.9851300 (2022).
Vlv, C. Q. D. O., Qjlqhhulqj, R., Qjlqhhulqj, R., Dgkduvklql, S. I. & Qjlqhhulqj, R. ’ LVHDVH Disease ’ HWHFWLRQ Detection % DVHG Based RQ on 1DLO Nail & RORU Using, PDJH Image 3URFHVVLQJ, 1st International Conference on Computational Science and Technology (ICCST), pp. 1–5, https://doi.org/10.1109/ICCST55948.2022.10040425 (2022).
Maniyan, P. & Shivakumar, B. L. Early disease detection through nail image processing based on ensemble Of KNN classifier and image features. 20(3), 14–25 https://doi.org/10.9790/0661-2003021425 (2018).
Jarallah, J. A., Al-dujaili, A. & Fadhel, M. A. Human nail diseases classification based on transfer learning. No November. https://doi.org/10.24507/icicel.15.12.1271 (2021).
Roy, P. S. Emperical analysis of nail diseases through using hybrid algorithms of LSTM and CNN. 2024 IEEE Int. Conf. Comput. Power Communication Technol. (IC2PCT). 5 (1), pp54–59. https://doi.org/10.1109/IC2PCT60090.2024.10486393 (2024).
Mehra, P. M., Costa, S. D., Mello, R. D., George, J. & Stage, E. Leveraging Deep Learning for Nail Disease Diagnostic, 4th Biennial International Conference on Nascent Technologies in Engineering (ICNTE), no. Icnte, pp. 1–5, https://doi.org/10.1109/ICNTE51185.2021.9487709 (2021).
Marulkar, S. & Narain, B. Nail Disease Prediction using a Deep Learning Integrated Framework, 3rd International Conference on Intelligent Technologies (CONIT), pp. 1–6, https://doi.org/10.1109/CONIT59222.2023.10205721 (2023).
Pathan, S. K. Nail Insight: Enhanced Nail Image Analysis for Early Disease Detection, 5th International Conference for Emerging Technology (INCET), pp. 1–9, https://doi.org/10.1109/INCET61516.2024.10593189 (2024).
Yani, M., Irawan, B. & Setiningsih, C. Application of transfer learning using convolutional neural network method for early detection of Terry ’ s nail application of transfer learning using convolutional neural network method for early detection of Terry ’ s nail, https://doi.org/10.1088/1742-6596/1201/1/012052
Li, S., Lu, X., Sakai, S., Mimura, M. & Kawahara, T. Semi-supervised ensemble DNN acoustic model training, ICASSP, IEEE International Conference on Acoustics, Speech and Signal Processing - Proceedings, pp. 5270–5274, https://doi.org/10.1109/ICASSP.2017.7953162 (2017).
Siami-Namini, S., Tavakoli, N. & Namin, A. S. The Performance of LSTM and BiLSTM in Forecasting Time Series, Proceedings – 2019 IEEE International Conference on Big Data, Big Data 2019, pp. 3285–3292, https://doi.org/10.1109/BigData47090.2019.9005997 (2019).
Abdulrahman, S. et al. A survey on federated learning: the journey from centralized to distributed on-site learning and beyond. IEEE Internet Things J. 8 (7), 5476–5497. https://doi.org/10.1109/JIOT.2020.3030072 (2021b).
Marulkar, S. & Narain, B. Nail Disease Prediction using a Deep Learning Integrated Framework. 2023 3rd International Conference on Intelligent Technologies (CONIT), 1–6. https://doi.org/10.1109/CONIT59222.2023.10205721 (2023).
Li, X., Yang, W., Zhang, Z., Huang, K. & Wang, S. on the Convergence of Fedavg on Non-Iid Data. 8th International Conference on Learning Representations, ICLR 2020, 2019, 1–26. (2020).
Blanco-Justicia, A. et al. Achieving security and privacy in federated learning systems: Survey, research challenges and future directions. Eng. Appl. Artif. Intell. 106 (September), 104468. https://doi.org/10.1016/j.engappai.2021.104468 (2021).
Jaiyeoba, O., Ogbuju, E., Yomi, O. T. & Oladipo, F. Development of a model to classify skin diseases using stacking ensemble machine learning techniques. J. Comput. Theor. Appl. 2 (1), 22–38 (2024).
Nail Disease DAtaset. (n.d.) https://www.kaggle.com/datasets/nikhilgurav21/nail-disease-detection-dataset
Shandilya, G. et al. Autonomous detection of nail disorders using a hybrid capsule CNN: a novel deep learning approach for early diagnosis. BMC Med. Inf. Decis. Mak. 24 (1), 414 (2024).
Acknowledgements
“The authors extend their appreciation to the Deanship of Research and Graduate Studies at King Khalid University for funding this work through Large Research Project under grant number RGP2/585/45”. This study was funded by Princess Nourah bint Abdulrahman University Researchers Supporting Project number (PNURSP2026R759), Princess Nourah bint Abdulrahman University,Riyadh, Saudi Arabia.
Author information
Authors and Affiliations
Contributions
Conceptualization - VK, MA, IK and GG, Methodology - VK, IK, SJ and AK, Software - VK, Investigation - VK, MA, AK, DG and AN, Validation - IK, GG, SJ and AN, Writing original draft - VK, IK, GG, AN and DG, Writing – review & editing - MA, IK, AK, SJ, and AN, Funding - MA, AK, and AN.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Khullar, V., Abbas, M., Kansal, I. et al. Low resource federated learning for classification of nail disease by deploying cross-silo and heterogeneously dataset distributions. Sci Rep 16, 7707 (2026). https://doi.org/10.1038/s41598-026-36848-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-36848-w