Abstract
Microscopic hyperspectral imaging (MHSI) of unstained tissue provides quantitative, label-free cues for pathology, but practical diagnosis is hindered by weak morphological contrast and high-dimensional spectra. Patch-wise classification is therefore unstable: discriminative spectral signatures are subtle, spatially sparse, and easily confounded by noise and tissue heterogeneity. To address this, we construct a new unstained breast MHSI dataset and formulate slice-level diagnosis as a multiple instance learning (MIL) problem. We propose a Multi-Scale Hierarchical Attention Network (MS-HAN) tailored to hyperspectral MIL. Each instance (patch) is encoded by an Inception-like multi-branch extractor that operates at a fixed spatial resolution using parallel convolution kernels to capture spectral–spatial patterns at different receptive fields. To reduce high intra-class spectral variability, we introduce a prototype-based clustering regularization that softly assigns instance embeddings to learnable centers and refines the representation. We then apply dual attention directly on the spatial feature map: channel (spectral) attention generates band-wise weights from global spatial descriptors, explicitly modeling inter-band dependencies, followed by spatial attention producing a 2D attention map to localize informative cellular regions. These modules are trained end-to-end with only slice-level labels. Finally, a hierarchical aggregator models inter-patch dependencies via self-attention and performs attention pooling to obtain the slice representation for classification. On a strictly patient-split cohort of 60 patients, MS-HAN achieved 86.7% accuracy and 0.92 AUC, outperforming strong MIL baselines (e.g., TransMIL and DS-MIL). McNemar’s test demonstrated statistically significant improvements over ABMIL (\(p=0.0251\)) and DS-MIL (\(p=0.0198\)), with marginal significance against CLAM and TransMIL (\(p<0.1\)). Ablations verified the necessity of the prototype regularization and hyperspectral-specific attention. Attention visualizations highlighted regions consistent with tumor-related morphology and emphasized informative spectral ranges without pixel-level annotations, pending expert validation. MS-HAN suggests that hyperspectral-specific feature refinement and hierarchical MIL aggregation may improve robust, stain-free breast cancer detection from microscopic MHSI. Further multi-center validation and expert review of attention explanations are needed to establish clinical utility.
Similar content being viewed by others
Introduction
Breast cancer continues to be one of the most widespread and lethal cancers globally, highlighting the pressing need for diagnostic tools that are not only accurate but also objective and reproducible1. The cornerstone of current diagnosis, histopathological analysis of stained tissue, relies on subjective interpretation by pathologists, leading to inter-observer variability that can affect diagnostic accuracy, particularly in complex cases2,3.
Microscopic Hyperspectral Imaging (MHSI) applied to unstained tissue sections emerges as a promising alternative, offering a pathway to quantitative, stain-free digital pathology4,5. By capturing hundreds of spectral bands for each pixel, MHSI can detect subtle biochemical shifts in endogenous molecules like proteins and nucleic acids, which are directly linked to neoplastic transformation6. However, this powerful approach introduces a unique set of computational challenges that limit its clinical translation. The absence of staining results in images with extremely low morphological contrast, where diagnostically critical nests of neoplastic cells are often obscured within a sea of structurally ambiguous stroma. The diagnostic information is thus encoded in faint spectral variations and subtle textural patterns, which are easily buried in noise and scattering effects7.
To capture sufficient cellular detail under these conditions, high-magnification microscopy is essential. This inherently limits the field of view, making a multi-point grid-scanning protocol necessary to ensure comprehensive coverage and capture tissue heterogeneity5,8. However, processing such data with standard supervised Deep Learning methods would require detailed pixel-level or patch-level annotations, which are labor-intensive, subjective, and often impractical for large-scale datasets9. Consequently, the field of computational pathology has increasingly shifted towards Weakly Supervised Learning (WSL), specifically Multiple Instance Learning (MIL), as the standard paradigm for analyzing Whole Slide Images (WSIs) without dense manual annotations9,10.
The complexity of unstained MHSI data demands an analytical tool that can account for both the rich spectral information within each patch and the overarching heterogeneity across a tissue slice. Standard CNNs, while effective for extracting features from individual patches, are not inherently designed to aggregate information from a collection of patches to make a single, slice-level diagnosis. This limitation can lead to noisy predictions, as the significance of a single patch is best understood in the context of its surrounding tissue.
To systematically address these challenges, this study presents a comprehensive framework tailored for stain-free computational pathology. First, we constructed a novel Microscopic Hyperspectral Breast Cancer Dataset (MHBCD) derived from unstained, paraffin-embedded tissue sections, filling a critical gap in data resources for spectral pathology. Second, unlike generic MIL models designed for standard RGB images, we propose the MS-HAN, a specialized architecture explicitly engineered for hyperspectral data. By integrating a multi-scale spectral feature extractor with a unique clustering constraint, MS-HAN is designed to capture fine-grained biochemical signatures that are often lost in low-contrast, unstained morphological structures. Finally, we shift the diagnostic paradigm from unreliable single-patch classification to a holistic slice-level assessment. This approach effectively mitigates the ambiguity of individual field-of-views by aggregating global context, providing a robust and objective diagnostic tool which we validate on a strictly split patient cohort.
Results
Dataset characteristics
In this study, we collected a novel Microscopic Hyperspectral Breast Cancer Dataset (MHBCD). This dataset was built from a cohort of 60 patients with breast cancer, prospectively collected between January 2022 and December 2023. The cohort’s mean age was \(53.55 \pm 13.23\) years and mean BMI was \(21.08 \pm 3.28\) kg/m\(^2\). Pathological and clinical characteristics are detailed in Table 1. Each MHSI data cube was acquired at a \(696\times 520\) spatial resolution, with 128 spectral bands from 397–1032 nm at a 5 nm resolution11.
Distribution of key clinical and pathological characteristics in the MHBCD cohort (N=60). The charts display the distribution for (a) Molecular Subtype, (b) Molecular Subtype Count, (c) Menopausal Status, and (d) Tumor Grade.
Classification performance
The proposed MS-HAN model demonstrated high efficacy in differentiating between tumor and paracancerous tissue slices on the unseen test set, which comprised 94 slices. The model achieved a robust overall classification accuracy of 86.7% ± 1.2%. Its excellent discriminative power is further highlighted by an Area Under the Curve (AUC) of 0.92 ± 0.01, as depicted by the Receiver Operating Characteristic (ROC) curve in Fig. 2a. The detailed classification results are presented in the confusion matrix shown in Fig. 2b, which provides a comprehensive breakdown of true positives, true negatives, false positives, and false negatives across the 94 test slices.
From a clinical utility perspective, the model’s performance is highly encouraging, as detailed in Table 2. The model achieved a Sensitivity (Tumor Recall) of 84.0%, which is critical in a diagnostic setting as it indicates a relatively low false-negative rate (16%) for cancer detection. This high sensitivity ensures that the vast majority of malignant cases are correctly identified, minimizing the risk of missed diagnoses. Furthermore, the model demonstrated a Specificity (Paracancerous Recall) of 88.6%, showing a strong ability to correctly identify non-malignant tissue and thus reduce the likelihood of false positives, which could lead to unnecessary follow-up procedures and patient anxiety. The overall discriminative power of the model is underscored by an AUC of 0.92, signifying its excellent ability to distinguish between tumor and paracancerous tissue across all decision thresholds.
Performance metrics of the MS-HAN model on the unseen test set (94 slices). (a) Receiver Operating Characteristic (ROC) curve showing the Area Under the Curve (AUC) of 0.92. (b) Confusion matrix detailing the classification results, including true positives, true negatives, false positives, and false negatives.
Statistical significance analysis
To validate the robustness of our classification results, we conducted McNemar’s test to evaluate the statistical significance of the performance differences between MS-HAN and the baseline models on the test set. As shown in Table 5, MS-HAN demonstrated a statistically significant improvement in classification decisions compared to ABMIL (\(p=0.0251\)) and DS-MIL (\(p=0.0198\)). When compared to the strongest baselines, CLAM and TransMIL, our model achieved marginal significance (\(p < 0.1\)). This marginal statistical separation is likely attributable to the limited statistical power inherent in the current sample size. Despite this, the consistent trend of superior predictive capability suggests that MS-HAN captures discriminative features effectively. Future work will include multi-center validation to improve statistical power to expand the cohort size, which is expected to further substantiate these findings with higher statistical confidence.
Model interpretability: visualizing cellular-level spatial attention
To gain insight into the decision-making process of our MS-HAN model, we visualized the output of its spatial attention module. This mechanism allows the model to identify and weigh the importance of different spatial regions within an input image patch for classification12. The resulting attention maps, as shown in Fig. 3, highlight the areas the model focused on, where brighter, yellow regions indicate high importance and darker, blue regions indicate low importance. The spatial attention heatmaps are directly extracted from the SpatialAttention output \(\textbf{A}\in \mathbb {R}^{1\times H\times W}\) of the dual attention block (before global pooling), and visualized at the resolution of the intermediate feature map.
Visualization of the spatial attention mechanism at the cellular level. The heatmaps indicate the model’s regions of focus, with yellow representing high attention and blue representing low attention. (a) Paracancerous tissue sample, where attention is diffusely spread across stromal areas and sparsely distributed cells. (b) Tumor tissue sample, where attention is highly concentrated on a dense nest of malignant cells, suggesting that the model focuses on regions visually consistent with pathologically relevant cellular structures, pending expert validation.
Spectral attention analysis
To address the question of which spectral bands contribute most to classification, we further analyzed the weights produced by the channel-wise attention module (Fig. 4). To obtain a stable, cohort-level summary rather than a single-patch visualization, we aggregate spectral attention in two steps. First, within each sample group (bag), we select tumor instances based on labels and compute a group-level tumor spectral attention vector by taking a normalized MIL-pooling-weighted sum of the corresponding patch-level spectral attention vectors. Second, we average these group-level tumor vectors across all tumor groups from all patients, producing the final band-wise distribution over 397–1032 nm shown in Fig. 4.
The aggregated distribution is clearly non-uniform, indicating that MS-HAN does not treat all spectral bands equally. Instead, the channel-wise attention learns band-dependent reweighting, which is consistent with explicitly modeling inter-band dependencies. Importantly, these attention weights are learned end-to-end jointly with the slice-level classification objective (i.e., not post-hoc band selection), thereby providing quantitative, reviewer-requested interpretability regarding how the model emphasizes different parts of the spectrum when forming tumor predictions.
Visualization of spectral attention weights produced by the channel-wise attention module in MS-HAN for tumor tissue, aggregated across all patients. For each sample group (bag), patch-level spectral attention vectors are first aggregated within the group using normalized MIL pooling weights over tumor instances (identified by labels), yielding a group-level tumor spectral attention vector. The final curve is obtained by averaging these group-level vectors over all tumor groups. (a) Bar plot representation: Mean normalized attention weights for the 128 spectral bands spanning 397–1032 nm (color indicates magnitude; yellow: low, red: high). (b) Heatmap representation: A complementary heatmap visualization of the same aggregated weights. The resulting non-uniform band-wise distribution indicates that the model assigns systematically different importance to different spectral bands, providing quantitative and reviewer-requested interpretability of the learned spectral emphasis under weak (slice-level) supervision.
Ablation studies
To systematically validate the effectiveness of the proposed architectural components in MS-HAN, we conducted a comprehensive ablation study. Specifically, we evaluated four model variants by removing key modules one at a time while keeping the rest of the network unchanged:
-
w/o Multi-scale Extractor: The multi-branch feature extractor was replaced with a standard single-scale convolutional layer (kernel size \(3 \times 3\)) to assess the importance of multi-granularity spectral-spatial feature capture.
-
w/o Clustering Constraint: The clustering-guided feature refinement module and its associated loss \(\mathcal {L}_{clust}\) were removed to verify the contribution of prototype learning in handling intra-class heterogeneity.
-
w/o Dual Attention: The channel and spatial attention mechanisms were disabled to evaluate their role in highlighting diagnostically relevant spectral bands and spatial regions.
-
w/o Dual Classifier: The auxiliary classification head and the dynamic fusion gate were removed, using only the final aggregated feature for prediction, to test the benefit of the dual-path decision mechanism.
Discussion
In this study, we addressed the inherent challenge of diagnostic ambiguity in unstained breast cancer sections by reframing the task from a local patch-level classification to a holistic slice-level diagnosis. We developed and validated MS-HAN, a specialized MIL architecture explicitly engineered for Microscopic Hyperspectral Imaging (MHSI). Evaluated on a strictly split test set from our newly constructed Microscopic Hyperspectral Breast Cancer Dataset (MHBCD), MS-HAN achieved a diagnostic accuracy of 86.7% and an AUC of 0.92. Crucially, it outperformed strong state-of-the-art (SOTA) baselines, including CLAM 10, DS-MIL 13 and TransMIL 14, underscoring the necessity for domain-specific architectural design for handling high-dimensional spectral data.
Our work shares conceptual similarities with other hierarchical attention models in computational pathology, such as HAMIL for colorectal cancer typing 15. However, while HAMIL operates on standard RGB histology images, MS-HAN is fundamentally designed to process the high-dimensional spectral-spatial information unique to MHSI. Similarly, while methods like DFLNet have shown success in breast cancer segmentation in ultrasound images 16, our approach focuses on classification in a different imaging modality and leverages the rich biochemical information encoded in hyperspectral data, which is absent in ultrasound.
It is worth clarifying that MS-HAN is fundamentally different from common multi-scale attention architectures used in medical imaging and hyperspectral analysis. Those models are typically designed for single-image (or single-volume) prediction and rely on multi-resolution feature pyramids with explicit cross-scale aggregation. In contrast, MS-HAN is formulated as a multiple-instance learning (MIL) problem for slice-level diagnosis: the input is a bag of N hyperspectral patches, and the model explicitly performs instance-to-bag aggregation to output a single prediction for the entire slice.
Accordingly, attention in MS-HAN serves two different roles from cross-scale attention. First, a within-instance dual attention module (channel/spectral attention followed by spatial attention) refines features at a fixed spatial resolution, emphasizing informative spectral bands and cellular regions within each patch. Second, a cross-instance self-attention module models dependencies among patches in the bag, and an attention-based pooling operation aggregates instance features into a robust bag representation. Moreover, the “multi-scale” design in MS-HAN is implemented as parallel convolution branches with different kernel sizes at the same resolution (Inception-like), rather than a multi-resolution pyramid. Finally, MS-HAN introduces a prototype-based clustering regularization and a dual-head (main/auxiliary) supervision with learnable fusion, which are not standard components in typical multi-scale attention backbones.
The superior performance of MS-HAN compared to general-purpose MIL models can be attributed to its targeted handling of spectral-spatial features. While baseline models are highly effective for extracting morphological features from standard RGB (H&E) images, they often struggle to capture the fine-grained spectral signatures present in unstained tissue. A critical design choice in our approach was treating spectral bands as channels in multi-scale 2D convolutions, rather than employing computationally intensive 3D-CNNs 17. While 3D-CNNs explicitly model spectral correlations, they often require significantly larger datasets to avoid overfitting 18; our results suggest that the proposed multi-scale 2D strategy offers an optimal balance between feature extraction capability and model generalization on clinical datasets.
Our ablation studies further underscored the contribution of each specific module to learning from unstained tissue. As detailed in Table 3, the removal of the Dual Classifier and Clustering Constraint resulted in the most significant performance drops, with accuracy decreasing to 74.0% and 74.8%, respectively. This finding indicates that the dual-path decision mechanism and prototype learning are fundamental requirements for stabilizing the feature space and handling the high intra-class heterogeneity of biological spectra. Additionally, the Dual Attention mechanism and Multi-Scale Extractor also proved beneficial, as their removal led to consistent declines in both Accuracy and AUC compared to the full model.
Functionally, the hierarchical attention mechanism of MS-HAN mimics the workflow of a pathologist: scanning the whole slide to identify and weigh the most salient regions of interest (ROIs). As visualized in our interpretability analysis, the model successfully learned to disregard non-informative stromal regions and focus its attention on diagnostically critical cell nests, achieving robust classification without pixel-level annotations. Importantly, this hierarchical attention design is particularly well-suited to microscopic hyperspectral imaging. First, in stain-free MHSI the morphological contrast within a single field-of-view is often weak and the diagnostically relevant cues are encoded as subtle, high-dimensional spectral variations that may occupy only a small fraction of a patch. As a result, single-patch predictions can be unstable and sensitive to noise, scattering, and local acquisition variability. Second, spectral signatures exhibit substantial intra-class heterogeneity across regions within the same slice due to mixed tissue components and micro-environmental differences. By hierarchically attending (i) within each patch to emphasize pathology-relevant cellular structures and (ii) across patches to down-weight non-informative or noisy fields-of-view while modeling inter-patch context, MS-HAN produces a more robust slice-level decision than treating patches independently.
Despite these promising results, this study is subject to several limitations. First, our dataset was limited to 60 patients from a single center. This relatively small sample size constrained the statistical power of our comparative analysis, which likely accounts for the marginal statistical significance (\(p<0.1\)) observed against strong baselines like TransMIL, despite the consistent improvement in mean performance metrics. However, it is noteworthy that acquiring high-resolution microscopic hyperspectral data is significantly more time-intensive than standard digital pathology scanning, making large-scale cohorts challenging to assemble. Nevertheless, while our strict patient-level splitting ensures internal validity, larger multi-institutional datasets will be required to establish external generalizability and validate the model’s robustness across different scanning systems and patient populations 19. Second, the computational cost of processing high-dimensional hyperspectral cubes is higher than that of traditional RGB analysis, which may pose challenges for deployment in resource-limited clinical settings 20.
Furthermore, the model’s performance may be compromised in scenarios with significant imaging artifacts, extremely low cellularity, or rare tumor subtypes not well-represented in the training data. As detailed in Table 1 and Fig. 1, our dataset exhibits a moderate class imbalance in molecular subtypes, with a 6.25:1 ratio between the most frequent (Luminal A) and least frequent (Triple-negative) classes. This distribution, however, is not a result of sampling bias but rather reflects the real-world prevalence of breast cancer subtypes in the clinical population. The model was trained on this imbalanced data without specific re-sampling techniques to ensure it learns from a distribution representative of clinical reality. While this approach risks a bias towards the majority classes, we observed stable overall performance on the held-out test set; subgroup robustness remains to be validated due to limited sample sizes. Future work could explore stratified sampling or cost-sensitive learning to further enhance performance for the minority classes. An analysis of the confusion matrix reveals that the model is more prone to misclassifying paracancerous tissue as tumor, a conservative error pattern that is preferable in a clinical screening context but indicates a need for improved feature specificity. The interpretability of the model, while promising, also requires further validation. Although the attention maps highlight regions consistent with tumor morphology, a formal validation by certified pathologists is a critical next step to confirm the clinical relevance of the model’s learned features. In particular, while the attention maps provide intuitive, qualitative cues for model interpretation, we did not perform region-level ground-truth annotation or independent pathologist validation of the highlighted hotspots in this study; therefore, these visualizations should not be interpreted as verified tumor localization.
While a full quantitative cost-benefit analysis is beyond the scope of this study, it is important to consider the potential advantages of a stain-free digital pathology workflow. Traditional histopathology, while the gold standard, involves material costs for reagents and stains, as well as significant labor costs and turnaround time for slide preparation and pathologist review 21,22,23,24. Our proposed MHSI-based approach eliminates the need for chemical staining and coverslipping steps; for example, automated H&E workflows can have a time to first result on the order of 40–45 minutes 25, reducing material costs and removing a time-consuming step from the laboratory workflow. The automated analysis provided by MS-HAN could further reduce the time required for diagnosis, particularly in high-throughput settings. By serving as a pre-screening tool, it could help prioritize cases for expert review, optimizing pathologists’ workload. Although the initial capital investment for hyperspectral imaging equipment is higher than for a standard brightfield microscope, the long-term savings in reagents, labor, and time, coupled with the potential for improved diagnostic objectivity and consistency, present a compelling case for its clinical consideration. Future studies should aim to conduct a formal health-economic evaluation to quantify these benefits.
Regarding clinical integration, MS-HAN is not intended to replace but rather to augment the existing pathology workflow. It could serve as a powerful pre-screening tool, automatically flagging suspicious slides or regions of interest for pathologists to review, thereby prioritizing workload and reducing diagnostic time. In settings like intraoperative frozen section analysis, a rapid, stain-free method could provide valuable preliminary diagnoses.
Future work will be directed towards expanding our cohort through multi-center collaborations to validate the model’s robustness across different scanning systems. Concurrently, we will explore model compression techniques and spectral channel selection to reduce computational demands. Additionally, integrating MHSI with other modalities, such as quantitative phase imaging or digital pathology, represents a promising avenue for developing a comprehensive, multi-modal diagnostic platform26. This advantage is particularly critical in time-sensitive scenarios such as intraoperative frozen section analysis, where turnaround time is often expected to be on the order of tens of minutes 27,28. Currently, this procedure represents a bottleneck where patients remain under anesthesia while surgeons wait for the pathology lab to process and stain tissue. By enabling diagnosis directly on unstained tissue, MS-HAN essentially eliminates the physical and chemical processing time, accelerating the ’biopsy-to-diagnosis’ turnover. In such high-stakes environments, the system’s value is measured not merely in reagent cost savings, but in the reduction of surgical wait times and anesthesia duration, justifying the capital investment through enhanced patient safety and operating room efficiency.
Conclusions
In conclusion, this study presents a comprehensive framework for stain-free computational pathology, comprising a novel hyperspectral dataset and a specialized deep learning architecture. By reframing the diagnostic task as a Multiple Instance Learning problem, our proposed MS-HAN model demonstrated superior performance compared to state-of-the-art MIL methods (including TransMIL and CLAM) on unstained tissue sections. The critical role of the clustering constraint and dual classification mechanism, as evidenced by our ablation studies, highlights the unique requirements of modeling high-dimensional spectral data. While validation on larger, multi-center cohorts remains a necessary next step, our results support the feasibility of stain-free breast cancer detection and suggest that domain-specific MIL models may contribute to more objective and reproducible decision support, pending further external and prospective validation.
Methods
Patient cohort and ethical approval
This study was conducted in accordance with the Declaration of Helsinki29 and received approval from the Institutional Review Board (IRB) of the Sixth Affiliated Hospital of Sun Yat-sen University (approval number: 2022ZSLYEC-241). Informed consent was obtained from all participants prior to their inclusion.
The study cohort consisted of 60 female patients diagnosed with breast cancer at the collaborating hospital between January 2022 and December 2023. As part of their standard clinical care, all patients underwent an initial core needle biopsy to confirm the cancer diagnosis and determine its histological and molecular subtype. These diagnostic assessments included hormone receptor status (ER, PR), HER2 status via immunohistochemistry and/or FISH, and classification into subtypes such as luminal A/B, HER2-enriched, or triple-negative, following established clinical protocols30. For this research, tissue specimens used for hyperspectral imaging were obtained from the surgical resection performed after the initial diagnosis.
Dataset splitting and evaluation protocol
To ensure a robust and unbiased evaluation of the model’s generalization capability, the patient cohort was strictly partitioned at the patient level into training, validation, and testing sets. The 60 patients were randomly allocated into a training set of 42 patients (70%), a validation set of 6 patients (10%), and a test set of 12 patients (20%). This patient-level separation guarantees that all tissue slides from a single patient belong exclusively to one set, preventing any data leakage between the training and evaluation phases and providing a more realistic assessment of the model’s performance on unseen cases.
Hyperspectral data acquisition and characteristics
Following surgical resection, the collected tissue specimens were processed by a certified pathology laboratory according to standard histopathological procedures21. The tissues were fixed, paraffin-embedded, and sectioned into 4-micron thick slices, following standard protocols. For each patient, sections containing both tumor regions and paracancerous tissue were mounted on glass slides for imaging.
Microscopic hyperspectral imaging (MHSI) was performed using a custom-assembled system in our laboratory, comprising an Olympus CX31RTSF biological microscope coupled with an SOC-710 hyperspectral camera (Surface Optics Corporation)11. The system was calibrated to capture data across a spectral range of 397–1032 nm with a spectral resolution of 5 nm. This process generated hyperspectral data cubes, with each cube representing a three-dimensional dataset (spatial \(\times\) spatial \(\times\) spectral), as illustrated in Fig. 5a. Each cube has a spatial resolution of \(696 \times 520\) pixels and contains 128 spectral bands.
This high-dimensional structure provides a complete spectral profile for every pixel (Fig. 5b) and reveals unique tissue information at different wavelengths, which can be visualized as individual grayscale images (Fig. 6). The rich spectral data reflects variations in tissue composition, such as protein, lipid, and nucleic acid content, which are essential for distinguishing malignant from paracancerous tissue31. All raw data cubes underwent a preprocessing pipeline to correct for illumination variations and remove noise and artifacts, ensuring data quality for subsequent analysis.
Visualization of the hyperspectral data cube structure and spectral characteristics. (a) A 3D representation of the hyperspectral data cube (696\(\times\)520\(\times\)128), with a pseudo-color image on its face and a central pixel marked in red for spectral analysis. This illustrates the data’s three-dimensional structure across spatial (Width, Height) and spectral (Depth) dimensions, where each pixel contains a complete spectral signature. (b) The spectral profile of the marked pixel, plotting its reflectance intensity across the full wavelength range (397–1032 nm, 128 bands at 5 nm resolution). The vertical dashed lines indicate the specific bands (Red: 640 nm, Green: 550 nm, Blue: 460 nm) used to generate the RGB pseudo-color reference image, demonstrating how different wavelengths reveal distinct tissue information.
Multi-wavelength visualization of hyperspectral tissue data. (a) The 2D pseudo-color reference image, generated by combining three spectral bands (Red at 640 nm, Green at 550 nm, and Blue at 460 nm), clearly shows the sample’s morphology and the location of the analyzed pixel. This composite image provides an intuitive representation of tissue structure. (b) Grayscale images from four individual spectral bands (460, 550, 640, and 850 nm), each revealing different tissue characteristics at specific wavelengths. The visible variation in contrast and texture across wavelengths demonstrates the rich, multi-dimensional information contained within the hyperspectral data cube, where each spectral ’layer’ captures unique biochemical signatures that are invisible in conventional RGB imaging.
Implementation details
All models were implemented using PyTorch version 2.1.0 on a system with CUDA 12.1, and trained on a single NVIDIA GeForce RTX 4090 GPU with 24GB of memory. The operating system was Ubuntu 20.04. For our MS-HAN model, the training process for the entire dataset took approximately 12 hours to converge over 200 epochs. During inference, the processing time for a single tissue section (a bag of 20 patches) was approximately 120 seconds, demonstrating the potential for rapid diagnostic support.
Final dataset composition for model training
The final dataset for our analysis was composed of 468 tissue sections from the 60 patients. In our previous work, each captured hyperspectral cube (patch) was treated as an independent sample. To better address tissue heterogeneity and model the diagnostic task at a more clinically relevant level, we adopted a MIL framework for this study.
Under the MIL paradigm, each tissue section (slide) is considered a ”bag,” and the 20 distinct, non-overlapping hyperspectral data cubes acquired from it are treated as its ”instances.” This resulted in a dataset of 468 bags, where each bag contains 20 instances. The label (Tumor or Paracancerous) assigned to a bag corresponds to the pathologist-delineated region from which its instances were sampled. This slice-level approach requires the model to aggregate evidence from all 20 patches to make a single, robust prediction for the entire tissue section, providing a more comprehensive assessment. This robust dataset structure supported the training and evaluation of our MS-HAN model.
Data preprocessing
Prior to model training, all acquired hyperspectral data cubes underwent a standardized preprocessing pipeline. To retain greater spatial detail from the original high-resolution images while managing computational load, the spatial resolution of each data cube, originally \(696\times 520\) pixels, was center-cropped to \(128\times 128\) pixels. The spectral dimension, containing 128 bands, was kept intact. Consequently, the final input tensor for each patch is a preprocessed data cube of shape \(\textbf{X}\in \mathbb {R}^{128\times 128\times 128}\) (interpreted here as \(C\times H\times W\)), consistent with the notation used in Fig. 7.
Model description
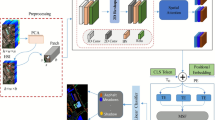
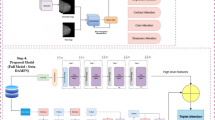
To analyze slice-level MHSI data, we propose a deep multiple-instance learning (MIL) model, MS-HAN, that takes an entire slice (20 hyperspectral patches) as a bag and outputs a single diagnosis. As illustrated in Fig. 7, the architecture comprises four stages:
Overview of MS-HAN in a MIL framework. A tissue section is partitioned into a bag of \(N{=}20\) patches and batched as \(\textbf{X}\!\in \!\mathbb {R}^{B\times N\times C\times H\times W}\).
MS-HAN pipeline. Given a bag \(\{\textbf{x}_i\}_{i=1}^{N}\) of patch tensors, the model outputs a slide-level prediction \(\textbf{z}\in \mathbb {R}^{C_{\text {cls}}}\) through the following components.
-
(1) Multi-scale patch feature maps. Each hyperspectral patch is processed by a multi-branch convolutional extractor (kernel sizes \(\{1,3,5,7\}\)) to produce an intermediate spatial feature map \(\textbf{F}_i\in \mathbb {R}^{C\times H\times W}\).
-
(2) Dual attention on feature maps + prototype-based clustering constraint on instance embeddings. We apply a CBAM-style dual attention block directly on the spatial feature map (before pooling). Channel (spectral) attention generates a band-wise weight vector \(\textbf{w}_i\in \mathbb {R}^{C\times 1\times 1}\) from global spatial descriptors of \(\textbf{F}_i\) (e.g., global pooling followed by an MLP and a sigmoid gate), and reweights the feature map as \(\textbf{F}_i^{(c)}=\textbf{w}_i\odot \textbf{F}_i\). This models inter-band dependencies by producing each band weight conditioned on the aggregated response across all bands. Next, spatial attention produces a 2D attention map \(\textbf{A}_i\in \mathbb {R}^{1\times H\times W}\) and refines the representation as \(\textbf{F}_i^{(cs)}=\textbf{A}_i\odot \textbf{F}_i^{(c)}\). We then perform global average pooling to obtain an instance embedding \(\textbf{f}_i\in \mathbb {R}^{C}\). To mitigate high intra-class spectral variability, we further apply a prototype-based clustering constraint on \(\textbf{f}_i\). The module computes a soft assignment vector \(\textbf{q}_i\in \mathbb {R}^{K}\) over K learnable prototypes \(\textbf{P}\in \mathbb {R}^{K\times C}\), reconstructs an embedding \(\hat{\textbf{f}}_i=\textbf{q}_i^\top \textbf{P}\), and uses the reconstruction residual \((\textbf{f}_i-\hat{\textbf{f}}_i)\) to refine the instance embedding, producing \(\textbf{v}_i\) for downstream MIL aggregation. Importantly, while reconstruction is used inside the module for feature refinement, we do not add an explicit L2/MSE reconstruction loss; instead, we regularize the assignment distribution to prevent prototype collapse. Both the attention modules and the clustering module are differentiable and trained end-to-end with the slice-level classification objective (i.e., not post-hoc).
-
3) Bag-level aggregation. The sequence of instance embeddings \([\textbf{v}_1,\dots ,\textbf{v}_N]\) is fed to a Transformer-style multi-head self-attention module (with LayerNorm and residual connections) to model inter-patch dependencies. The bag representation \(\textbf{s}\in \mathbb {R}^{d_s}\) is obtained by attention pooling over the instance tokens.
-
4) Dual classification head and fusion. Two linear heads map \(\textbf{s}\) to logits \(\textbf{z}_{\text {main}}\) and \(\textbf{z}_{\text {aux}}\) over \(C_{\text {cls}}\) classes. A learnable scalar gate \(\alpha \in [0,1]\) forms the fused logits
$$\textbf{z}=\alpha \,\textbf{z}_{\text {main}}+(1-\alpha )\,\textbf{z}_{\text {aux}},$$which drive the primary cross-entropy.
Training. We optimized with AdamW 32 (weight decay 0.01). The learning rate linearly warmed up for 5 epochs to \(3{\times }10^{-4}\), then followed a ReduceLROnPlateau scheduler. The objective combined the main cross-entropy, an auxiliary cross-entropy, and a clustering diversity regularizer:
Let \(\textbf{q}_i\in \mathbb {R}^{K}\) denote the soft assignment of instance i to K prototypes, and let \(\bar{\textbf{q}}=\frac{1}{M}\sum _{i=1}^{M}\textbf{q}_i\) be the average assignment distribution over the M instance embeddings in the current minibatch. We define the entropy \(H(\bar{\textbf{q}})=-\sum _{k=1}^{K}\bar{q}_k\log \bar{q}_k\) and set a target entropy \(H_{\text {target}}=\rho \log K\) with \(\rho =0.8\). The diversity regularizer is a hinge loss:
which prevents prototype collapse by encouraging sufficiently diverse prototype usage. We trained for 200 epochs with a batch size of 4 slices, using gradient accumulation and automatic mixed precision.
Baselines. For comparison, we implemented one non-MIL and four state-of-the-art MIL baselines under a unified setup. All MIL baselines treated each slice as a bag of 20 instances (standardized by subsampling or repetition) and shared the same 2D-CNN instance encoder that mapped a \(128{\times }128\) patch with 128 spectral channels to a 512-d feature:
-
CNN + Max Pooling (non-MIL): Element-wise max pooling across the 20 instance features to obtain a single bag vector, followed by a linear classifier.
-
Attention-based MIL (ABMIL) 33: A fully connected attention mechanism scores instance features; the bag representation is the attention-weighted average fed to a linear classifier.
-
Clustering-constrained Attention MIL (CLAM) 10: A two-level attention MIL with an explicit clustering constraint that encourages the discovery of subtypes within positive bags.
-
Dual-Stream MIL (DS-MIL) 13: A dual-stream architecture that combines a max-pooling stream to identify the most critical instance and an attention-weighted averaging stream to capture global characteristics.
-
Transformer-based MIL (TransMIL) 14: Uses positional encodings and Transformer encoders to capture global dependencies among instances; context-aware outputs are aggregated by attention for classification.
Baseline training. We used AdamW 32 for all baselines with hyperparameters tuned per model: ABMIL used LR \(3{\times }10^{-4}\), weight decay 0.01; CLAM used LR \(2{\times }10^{-4}\), weight decay 0.01; TransMIL used LR \(1{\times }10^{-4}\), weight decay 0.005. All models were trained for up to 200 epochs with batch size 4 using cross-entropy; CLAM additionally included its clustering loss. A ReduceLROnPlateau scheduler and early stopping (by validation accuracy) were applied.
Statistical analysis
To ensure statistical rigor, performance metrics (Accuracy, AUC, F1-Score) are reported as Mean ± Standard Deviation (SD), derived from non-parametric bootstrap resampling on the test set with \(B=2,000\) iterations. Results are summarized in Table 4.
To statistically compare the classification performance of MS-HAN against baseline models, we employed McNemar’s Test. This test evaluates the significance of the discordance between the classifiers’ prediction errors on the paired test samples. A two-sided p-value \(< 0.05\) was considered statistically significant, while \(p < 0.1\) was considered marginally significant. All analyses were performed using Python (SciPy library).
Data availability
The datasets generated and/or analysed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- MHSI::
-
Microscopic Hyperspectral Imaging
- MIL::
-
Multiple Instance Learning
- CNN::
-
Convolutional Neural Network
- WSI::
-
Whole Slide Image
- ROI::
-
Region of Interest
- AUC::
-
Area Under the Curve
- SD::
-
Standard Deviation
References
Sung, H. et al. Global Cancer Statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 74(3), 209–49 (2024).
Gurcan, M. N. et al. Histopathological image analysis: A review. IEEE Rev. Biomed. Eng. 2, 147–71 (2009).
Elmore, J. G. et al. Diagnostic concordance among pathologists interpreting breast biopsy specimens. JAMA 313(11), 1122–32 (2015).
Roth, H.R., Lu, L., Seff, A., Cherry, K.M., Wu, J., Lu, S., et al. A new 2.5D representation for lymph node detection using random sets of deep convolutional neural network observations. In: Medical Image Computing and Computer-Assisted Intervention-MICCAI 2014. Springer; p. 520–7, (2014).
de Oliveira, A. M. D. C. F., da Mota, P. A. F. C., de Albuquerque, V. H. C. & de Farias, T. M. P. Microscopic hyperspectral imaging: a new tool for pathology. Expert Rev. Med. Devices 19(10), 837–51 (2022).
Shahshahan, M., Tsen, S. D. W., Kj, Bae, Peng, T. Q. & Esfandyarpour, R. Label-free hyperspectral imaging and deep learning for tracking the metabolic response of single cancer cells to chemotherapy. Sci. Rep. 9(1), 18663 (2019).
Li, S. et al. Hyperspectral image classification: A review of recent advances. IEEE Geosci. Remote Sens. Magaz. 8(3), 25–43 (2020).
Lu, G. & Fei, B. Medical hyperspectral imaging: A review. J. Biomed. Opt. 19(1), 010901 (2014).
Campanella, G. et al. Clinical-grade computational pathology using weakly supervised deep learning on whole slide images. Nat. Med. 25(8), 1301–9 (2019).
Lu, M. Y. et al. Data-efficient and weakly supervised computational pathology on whole-slide images. Nat. Biomed. Eng. 5(6), 555–70 (2021).
Surface Optics Corporation. SOC-710 Hyperspectral Imager Datasheet. Surface Optics Corporation; 2024. Accessed: 2025–06-18. Available from: https://www.surfaceoptics.com/products/hyperspectral-imaging-systems/soc-710-hyperspectral-imager/.
Woo, S., Park, J., Lee, J.Y., Kweon, I.S. (2018) CBAM: Convolutional Block Attention Module. In: Proceedings of the European conference on computer vision (ECCV); p. 3–19.
Li, B., Li, Y., Eliceiri, K.W. (2021) Dual-stream multiple instance learning network for whole slide image classification with self-supervised contrastive learning. In: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition; p. 14318–28.
Shao, Z. et al. TransMIL: Transformer based correlated multiple instance learning for whole slide image classification. Adv. Neural. Inf. Process. Syst. 34, 2136–47 (2021).
Ye, Z. et al. HAMIL: Hierarchical attention multi-instance learning for label-free colorectal cancer typing. Laser Photon. Rev. 18(6), 2301072 (2024).
Ye, Z. et al. DFLNet: Disentangled feature learning network for breast cancer ultrasound image segmentation. Digital Signal Process. 145, 104331 (2024).
Li, Y., Zhang, H. & Shen, Q. Spectral-spatial classification of hyperspectral imagery with 3D convolutional neural networks. Remote Sens. 9(1), 67 (2017).
Ahmad, J., Farman, H. & Jan, Z. 3D-convolutional neural networks for medical imaging analysis: A systematic review. Multim. Tools Appl. 82, 27373–415 (2023).
Rajpurkar, P., Chen, E., Banerjee, O. & Chen, Y. Appraising the state of the art in medical AI. Nat. Med. 30(5), 1264–74 (2024).
Kelly, C. J., Karthikesalingam, A., Suleyman, M., Corrado, G. & King, D. Key challenges for delivering clinical impact with artificial intelligence. BMC Med. 17(1), 195 (2019).
Bancroft, J. D. & Gamble, M. Bancroft’s Theory and Practice of Histological Techniques 8th edn. (Elsevier, 2019).
Swanson Biotechnology Center, MIT. Pricing | Histology. Massachusetts Institute of Technology; n.d. Available from: https://ki-sbc.mit.edu/histology/pricing.
MUSC College of Medicine. Services & Prices | Histology & Immunohistochemistry. Medical University of South Carolina (MUSC); n.d. https://medicine.musc.edu/departments/pathology-laboratory-medicine/divisions/experimental-pathology/cores/histology-immunohistochemistry/services-prices.
UC Davis Health. Research Histology: Rates. UC Davis Health; n.d. https://health.ucdavis.edu/pathology/research/research_labs/histology/rates.html.
Roche Diagnostics. VENTANA HE 600 automated staining system. Roche Diagnostics; n.d. https://diagnostics.roche.com/us/en/products/instruments/ventana-he-600-ins-4090.html.
Vale-Silva, L. & Rohr, K. A review on deep learning-based methods for multi-modal medical imaging fusion. Expert Syst. Appl. 239, 121542 (2024).
Novis, D. A. & Zarbo, R. J. Interinstitutional comparison of frozen section turnaround time. A College of American Pathologists Q-Probes study of 32868 frozen sections in 700 hospitals. Arch. Patholo. Lab. Med. 121(6), 559–67 (1997).
Chen, Y., Anderson, K. R., Xu, J., Goldsmith, J. D. & Heher, Y. K. Frozen-section checklist implementation improves quality and patient safety. Am. J. Clin. Pathol. 151(6), 607–12 (2019).
World Medical Association. World Medical Association Declaration of Helsinki: Ethical principles for medical research involving human subjects. JAMA 310(20), 2191–4 (2013).
Wolff, A. C. et al. Human Epidermal Growth Factor Receptor 2 Testing in Breast Cancer: American Society of Clinical Oncology/College of American Pathologists Clinical Practice Guideline Focused Update. J. Clin. Oncol. 36(20), 2105–22 (2018).
Vo-Dinh, T. editor. Biomedical Photonics Handbook, Second Edition: Biomedical Diagnostics. CRC Press; (2014).
Loshchilov, I., Hutter, F. Decoupled Weight Decay Regularization; (2019). OpenReview paper. International Conference on Learning Representations (ICLR). Available from: https://openreview.net/forum?id=Bkg6RiCqY7.
Ilse, M., Tomczak, J., Welling, M. Attention-based Deep Multiple Instance Learning. In: International Conference on Machine Learning (ICML). PMLR; p. 2127–36, (2018).
Author information
Authors and Affiliations
Contributions
Z.C. and Q.Y. contributed equally to this work. Z.C. and Q.Y. performed the data analysis, developed the model, and wrote the main body of the manuscript. G.Q., X.M., and Z.L. conducted the data acquisition. H.L. and B.S. provided guidance and supervision for the project. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
This study was conducted in accordance with the Declaration of Helsinki29 and received approval from the Institutional Review Board (IRB) of the Sixth Affiliated Hospital of Sun Yat-sen University (approval number: 2022ZSLYEC-241). Informed consent was obtained from all participants prior to their inclusion.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chen, Z., Yang, Q., Qin, G. et al. MultiScale hierarchical attention network for stain free breast cancer detection in microscopic hyperspectral imaging. Sci Rep 16, 9404 (2026). https://doi.org/10.1038/s41598-026-39267-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-39267-z