Abstract
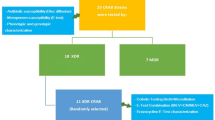
Healthcare-associated infections (HAIs) caused by carbapenem-resistant Pseudomonas aeruginosa (CRPA) are considered a major problem in hospitals due to their resistance to the most effective classes of antibiotics. The aim of the present study was to determine the antimicrobial susceptibility patterns of P. aeruginosa isolates from clinical specimens, and the combination effect of meropenem, amikacin, and ciprofloxacin against CRPA isolates. This cross-sectional study analyzed 70 P. aeruginosa isolates collected from hospitalized patients between August and November 2023. Antimicrobial susceptibility testing was performed on all isolates, and carbapenemase production was assessed in CRPA isolates. Five carbapenemase genes (blaIMP, blaVIM, blaKPC, blaSIM, and blaOXA-48) were screened. Meropenem minimum inhibitory concentrations (MICs) were determined via microbroth dilution method, and synergistic effects of meropenem, amikacin, and ciprofloxacin were evaluated using a three-dimensional checkerboard method. The highest resistance was to colistin 52/70 (74.3%), followed by cefotaxime 33/70 (47.1%) and tobramycin 28/70 (40%). Conversely, the lowest resistance was for aztreonam 19/70 (27.1%); 26/70 (37.1%) of isolates were CRPA. The frequency of genes blaIMP and blaOXA-48 were 4/70 (5.7%) and 1/70 (1.4%) in P. aeruginosa isolates, respectively; blaKPC, blaVIM, and blaSIM were not detected. The three-dimensional checkerboard method was established on 8 selected non-susceptible carbapenem P. aeruginosa isolates and the synergistic effect was recorded for 5 of these tested isolates. This study presents a positive correlation between CRPA and multidrug resistance (MDR), including resistance to all tested antibiotics. However, synergistic effects observed in vitro with meropenem, amikacin, and ciprofloxacin combinations suggest that combination therapy holds promise for treating these infections.
Similar content being viewed by others
Introduction
Hospital-acquired infections (HAIs) caused by pathogenic bacteria are considered significant contributors to mortality among hospitalized patients. These infections prolong hospital stays and increase treatment costs, placing a substantial economic burden on healthcare systems1. Pseudomonas aeruginosa (P. aeruginosa) is among the most important bacteria responsible for these infections. The remarkable adaptability of this bacterium to the environment and increasing antibiotic resistance, particularly multidrug-resistant (MDR) and extensively drug-resistant (XDR), has transformed the treatment of infections caused by P. aeruginosa into a global concern2,3.
P. aeruginosa is a serious opportunistic pathogen, particularly in immunocompromised individuals. It is responsible for acute and chronic infections in patients with conditions such as cystic fibrosis, cancer, trauma, burns, pneumonia, and sepsis4. Effective antibiotic selection is critical for optimizing clinical outcomes; however, the extensive drug resistance exhibited by P. aeruginosa complicates this process5. The bacterium’s ability to acquire MDR phenotypes, particularly in hospital settings, further exacerbates the challenge of treatment6.
Carbapenems are among the most potent antibiotics for managing severe infections caused by MDR Gram-negative bacteria. However, the emergence and spread of Carbapenem-resistant P. aeruginosa (CRPA) have substantially diminished the efficacy of this antibiotic class7,8. CRPA often exhibits cross-resistance to other commonly used antimicrobial agents, resulting in high mortality rates in infections caused by this isolate9.
Given the increasing difficulty of treating MDR and XDR P. aeruginosa infections, combination antibiotic therapy has emerged as a promising strategy. This approach enhances antimicrobial efficacy and mitigates the risk of further resistance development that is possible in monotherapy10. Recent studies have reported synergistic effects against P. aeruginosa isolates when exposed to the combinations of amikacin/ceftazidime, tobramycin/colistin, and ceftazidime/colistin11, amikacin/imipenem12, and ciprofloxacin/gentamycin13. However, there is no evidence of in vitro evaluation of the minimum inhibitory concentration (MIC) of three different antibiotic combinations against CRPA isolates.
Therefore, the current study aimed to demonstrate the in vitro MIC of the combination of meropenem, amikacin, and ciprofloxacin using a three-dimensional checkerboard method against CRPA isolates from hospitalized patients in Shiraz, Iran.
Materials and methods
Bacterial isolates
In this cross-sectional study, clinical specimens were collected for the possible isolation and identification of P. aeruginosa from patients admitted to three tertiary referral teaching hospitals in Shiraz, Iran (Namazi, Rajaee, and Amir Oncology Hospital) between August and November 2023. The Ethics Committee of Shiraz University of Medical Sciences gave their approval for this study with the code number IR.SUMS.REC.1402.466. The written informed consent was signed by each participant for the anonymous use of their data. Parents or legal guardians provided written informed permission for individuals under 16 years.
Namazi Hospital is a general hospital encompassing various specialties and subspecialties, while Rajaee and Amir Hospitals are centers of trauma and oncology, respectively. The study included a variety of specimen types, such as blood, wound swabs, and sputum. All specimens underwent presumptive identification of P. aeruginosa isolates using biochemical standard tests, as previously described by Zomorodi et al.8. These tests included Gram staining, observation of pigmented colonies, catalase and oxidase tests, growth patterns on triple-sugar iron (TSI) agar, the oxidation-fermentation (OF) test (Merck, Germany), and the ability to exhibit growth at 42 °C, which is a criterion for differentiating P. aeruginosa from other lactose non-fermenting Gram-negative rods. One of the investigators (ZF) was notified by the laboratories of each of the three hospitals when P. aeruginosa was identified in any of the samples of interest, following which the sample would be kept for further evaluations. Presumptive P. aeruginosa isolates were stored in Tryptic Soy Broth (TSB) medium (Merck, Germany) supplemented with 30% glycerol at −70 °C for further experiments.
Molecular confirmation of P. aeruginosa
All primary identified P. aeruginosa isolates were confirmed through amplification of 16 S rRNA gene using the polymerase chain reaction (PCR). First, DNA of investigated isolates were extracted using the boiling method as previously explained by Zomorodi et al.14. The oligonucleotide primer sequences used were designed previously (Table 1). PCR amplification was conducted using the Veriti™ 96-Well Thermal Cycler (Applied Biosystems, USA) with an initial denaturation step at 95 °C for 2 min, followed by 25 cycles of denaturation at 95 °C for 20 s, annealing at 58 °C for 20 s, and extension at 72 °C for 40 s. A final extension step was conducted at 72 °C for 5 min. The DNA template from the P. aeruginosa ATCC27853 strain served as the positive control, while distilled water was used as the negative control8.
Antimicrobial susceptibility testing
The susceptibility of the P. aeruginosa isolates to 11 antibiotics was carried out through the Kirby–Bauer disk diffusion assay as recommended by the Clinical and Laboratory Standards Institute (CLSI, 2024)16. The tested antibiotics were as follow: ceftazidime (CAZ, 30 µg), cefotaxime (CTX, 30 µg), piperacillin (PIP, 100 µg) piperacillin-tazobactam (PTZ, 100/10 µg), meropenem (MRO, 10 µg), imipenem (IMP, 10 µg), aztreonam (AZT, 30 µg), Ciprofloxacin (CIP, 5 µg), tobramycin (TOB, 10 µg), amikacin (AMK, 30 µg), gentamicin (GEN, 10 µg) (Mast Group, UK). Additionally, susceptibility to colistin was assessed using the disc elution method as recommended by CLSI, 2024. Also, P. aeruginosa ATCC27853 were implied as quality control. Eventually, MDR isolates were defined as those resistant to at least one antibiotic in three or more antibiotic classes.
Carbapenemase production
CRPA isolates (those resistant to imipenem and/or meropenem; MIC of imipenem and/or meropenem ≥ 8 µg/mL) were subjected to determination of their carbapenemase production potential by the modified carbapenem inactivation method (mCIM)16.
Determination of the MIC of the meropenem
The microbroth dilution method was conducted to determine MIC of meropenem for P. aeruginosa isolates16. In this purpose, a range of MIC for meropenem (Sigma-Aldrich Co., St. Louis, MO, USA) was established from 0.125 to 128 µg/mL in 96-well U-shaped microplates containing cation-adjusted Mueller-Hinton broth (CAMHB) (Merck, Germany). The eleventh and twelfth columns served as the positive control (containing only CAMHB and the tested bacteria) and the negative control (which contained sterile distilled water in place of bacteria), respectively.
Three-dimensional checkerboard method
Based on the MIC results for meropenem, six CRPA isolates (MIC of meropenem ≥ 8 µg/mL) and two non-susceptible-carbapenem isolates (MIC of meropenem = 4 µg/mL) were randomly selected to investigate the in vitro antimicrobial effects of a combination of meropenem, amikacin, and ciprofloxacin using a three-dimensional checkerboard method. Briefly, the selected isolates were inoculated into sterile Falcon tubes containing 10 mL of Mueller-Hinton broth (MHB) (Merck, Germany) and incubated at 35 ± 2 °C with continuous shaking at 200 rpm for 18 to 20 h. Following incubation, the cultures were adjusted to the 0.5 McFarland standard, corresponding to 108 CFU/mL, and then diluted 1:100 with MHB to yield a suspension of 106 CFU/mL. The checkerboard assay was performed using a modified broth microdilution method that included additional concentrations of the antibiotics. A total of six 96-well U-shaped microplates were utilized, with 100 µL of the 106 CFU/mL bacterial suspension transferred into each well. The concentration range for each antibiotic was determined based on the MIC of each antibiotic when tested individually against the isolates. In this study, the MIC ranges for meropenem, amikacin, and ciprofloxacin were 0.125 to 256, 0.125 to 8, and 0.5 to 16 µg/mL, respectively. The fractional inhibitory concentration index (FICI) for the triple antibiotic combination was calculated as follows:
The fractional inhibitory concentration indices (FICIs) were calculated using the concentrations obtained from the first non-turbid well in each row and column along the interface of turbidity and non-turbidity, with results presented as median values. To avoid over-interpretation of synergistic interactions that could result from methodological errors associated with the double dilution method, we used the median FICI values rather than the lowest observed values. The FICI values were categorized as follows: FICI < 0.8 indicates synergy; 0.8 ≤ FICI < 4 reflects additive effects or indifference; and FICI ≥ 4 signifies antagonism17.
Molecular detection of carbapenemase gene
The presence of 5 carbapenemase mediated genes including blaVIM, blaSIM, blaKPC, blaIMP, and blaOXA-48 were sought among all P. aeruginosa isolates using PCR methods. The PCR reactions were conducted in a final volume of 25 µL; Each reaction included 12.5 µL of 2× PCR Master Mix (Amplicon, Denmark), 1 µL of each primer (10 µM), 2 µL of template DNA (100 ng/µL), and 8.5 µL of nuclease-free water. An archived positive bacterial strain and distilled water were used as positive and negative controls, respectively. The oligonucleotide sequences of the primers used are listed in Table 1.
Statistical analysis
Statistical analysis for this study was performed using SPSS Statistics (Version 22.0). The evaluation employed both Chi-square and Fisher’s exact tests, considering a p-value of less than 0.05 as statistically significant. 3D Graph was plotted with Matplotlib (Version 3.9.1) in Python (Version 3.11.8).
Results
Demographic information
A total of 85 specimens were investigated, yielding 70 isolates of Pseudomonas aeruginosa. Among these isolates, 38 (54.3%) were from Namazi Hospital, 20 (28.6%) from Rajaee Hospital, and 12 (17.1%) from Amir Oncology Hospital. The participating patients included 45 men (64.3%) and 25 women (35.7%), with a mean age of 43.97 ± 23.68 years (range: < 1 to 89 years). The most common specimens collected were blood (n = 17, 24.3%), sputum (n = 14, 20%), and wound samples (n = 12, 17.1%), respectively. P. aeruginosa isolates were obtained from various wards, with 24 isolates (34.3%) collected from the ICU, 20 isolates (28.6%) from the emergency department, and 10 isolates (14.3%) from pediatric wards (Table 2).
Antibiotic susceptibility test results
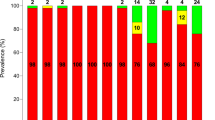
Table 3 presents the prevalence of resistant P. aeruginosa isolates across the included hospitals. For colistin, 18 isolates (25.7%) exhibited moderate sensitivity (MIC between 2 and 4 µg/mL), whereas 52 (74.3%) were resistant (MIC ≥ 4 µg/mL). After colistin, the highest resistance rates were observed against cefotaxime, tobramycin, and imipenem, observed in 33 (47.1%), 28 (40%), and 26 (37.1%) isolates, respectively. Conversely, the highest sensitivity was attributed to meropenem, with 44 (62.9%) of P. aeruginosa isolates showing sensitivity, as well as to ceftazidime and ciprofloxacin, both also exhibiting sensitivity in 44 (62.9%) of isolates (Fig. 1). Investigation of the MIC for meropenem revealed that MIC values of ≤ 0.5 µg/mL and 256 µg/mL were the most prevalent, with frequencies of 24 (34.2%) and 14 (20%) of P. aeruginosa isolates, respectively. Additionally, the MIC50 was found to be 4 µg/mL, while the MIC90 was 256 µg/mL. Furthermore, 26 (37.1%) of P. aeruginosa isolates were identified as MDR. The prevalence of MDR did not differ between the hospitals, wards, or specimens (p > 0.05).
Antimicrobial susceptibility pattern of P. aeruginosa isolates (N = 70). Abbreviation: CAZ: ceftazidime, CTX: cefotaxime, PIP: piperacillin, PTZ: piperacillin/tazobactam, AZT: aztreonam, MRO: meropenem, IMP: imipenem, CIP: ciprofloxacin, GEN: gentamicin, TOB: tobramycin, AMK: amikacin, COL: colistin, MDR: multi-drug resistant.
Frequency of CRPA isolates
In total, 26 (37.1%) of P. aeruginosa isolates were identified as CRPA. The prevalence of CRPA isolates was significantly higher among isolates obtained from Namazi Hospital than others (p = 0.004). Furthermore, a positive correlation was observed between CRPA isolates and MDR strains (p ≤ 0.001). Additionally, no significant differences were found between CRPA and carbapenem-susceptible P. aeruginosa isolates regarding ward or specimen type (p > 0.05).
Carbapenemase detection
A total of 26 CRPA were tested for the ability of carbapenemase production using the mCIM method, and the results were positive for 5 (20%) (Table 1). PCR products were visualized using the gel electrophoresis method (Fig. 2). Among the 70 investigated P. aeruginosa isolates, 4 (5.7%) and 1 (1.4%) tested positive for blaIMP and blaOXA-48 genes, respectively. The genes blaVIM, blaKPC, and blaSIM were not detected in any isolate.
The PCR amplified products displayed with agarose gel electrophoresis (1.5% agarose). (A) Lane C-: negative control, Lane M: DNA marker (100 bp plus DNA Ladder), Lane: C+: control positive, Lanes 1,3, 5, and 6: Detection of 16 S rRNA gene for confirmation of P. aeruginosa isolates. (B) Lane C-: negative control, Lane M: DNA marker (100 bp plus DNA Ladder), Lane C+: positive control, Lanes 1 and 2: detection of blaIMP gene in P. aeruginosa isolates. (C) Lane C-: negative control, Lane M: DNA marker (100 bp plus DNA Ladder), Lane C+: positive control. (D) Lane C-: negative control, Lane M: DNA marker (100 bp plus DNA Ladder), Lane C+: positive control, Lane 5: Detection of blaOX-48 gene in P. aeruginosa isolate.
Three-dimensional checkerboard test
Eight non-susceptible P. aeruginosa isolates, including 6 CRPA and 2 intermediates, were selected based on their MIC values for meropenem. Table 4 presents the characteristics of the selected isolates. None of the included isolates were positive for carbapenemase genes. In general, a synergistic effect was observed for 5 out of the 8 tested P. aeruginosa isolates. Among the 48 different statuses of varying concentrations of three antibiotics, 7 synergistic effects and 6 antagonistic effects were identified (S1 Table). Figure 3 depicts the MIC of amikacin, ciprofloxacin, and meropenem in the combination setting in synergy, additive, and antagonism groups. Figure 4 demonstrates the decision tree classifier for the synergy class, suggesting that synergy is likely when the MICs of amikacin, ciprofloxacin, and meropenem in the combination setting are ≤ 2, 6, and 24 µg/mL, respectively.
3D scatterplot of the cases based on the Minimum Inhibitory Concentration (MIC) of the antibiotics in the combination setting. Labeled values indicate the overlap of a case in the additive category with a case in the synergy (A), antagonism (B), or additive (C) categories. Cases farther along the axes are displayed with reduced opacity for better visualization. The axes show the absolute MICs of each drug in the combination setting, whereas the color-coding reflects the interaction category based on the fractional inhibitory concentration index (FICI), which normalizes each combination MIC to the isolate’s corresponding single-agent MIC. Therefore, cases can share very similar combination MIC triplets yet fall into different categories if their single-agent MICs differ (e.g., lower baseline MICs yield larger fractional values at the same MIC in combination). Antagonism points correspond to cases in which the required concentrations in combination are close to or higher than the single-agent MIC(s), resulting in FICI ≥ 4.
Decision tree based on the Minimum Inhibitory Concentration (MIC) of the antibiotics in the combination setting. Percentages displayed in terminal nodes indicate the proportion of observations in that node in favor of the proposed condition (i.e., synergy expected versus not expected). The number of instances in each terminal node is as follows: (1) 21; (2) 9; (3) 3; (4) 15.
Discussion
P. aeruginosa is a leading cause of HAIs, morbidity, and mortality, with its virulence factors and extensive antibiotic resistance contributing to its clinical impact18. Recent studies indicate that P. aeruginosa is responsible for approximately 4.7–8.9% of hospital-acquired bloodstream infections19, 10–20% of ventilator-associated pneumonia (VAP)20, 16% of urinary tract infections in ICU patients21, and 26% of infections in hospitalized burn victims22. In the present study, 70 P. aeruginosa isolates were obtained from various specimens collected from hospitalized patients in different wards of Namazi, Rajaee, and Amir Hospitals. The prevalence of isolated P. aeruginosa was higher in men than in women (64.3% vs. 35.7%), consistent with previous studies conducted in Iran and other countries23,24,25,26.
The most prevalent specimens yielding P. aeruginosa were blood (24.3%), sputum (20%), and wound (17.1%). Elahi et al.. reported the prevalence of P. aeruginosa from sputum and wound specimens as 24.3% and 23.1%, respectively, consistent with our results24. However, the prevalence of blood cultures positive with P. aeruginosa ranges from 3.2% to 19.8% in previous studies conducted in Iran24,27,28,29. The discrepancy in the prevalence of the blood specimens is also observed among other studies. Chand et al.. reported the highest prevalence of P. aeruginosa isolates in urine specimens, while the lowest was found in blood specimens25. The prevalence of P. aeruginosa isolates in China was reported as 28% in sputum, 21.6% in wounds, and 13.7% in blood23, while in Ethiopia, the frequencies were 17.6% in blood and 35.3% in wounds26. These differences may be attributed to several factors, such as variations in sample sizes or specific inclusion and exclusion criteria used during sampling.
Over one-third of P. aeruginosa isolates were identified in the ICU, followed by the emergency (28.6%) and pediatric (14.3%) departments. The high prevalence of P. aeruginosa isolates in the ICU has also been documented in previous studies26,30,31. Additionally, previous systematic reviews and meta-analyses reported a significant association between MDR P. aeruginosa and ICU admission32,33. Contrastingly, the prevalence of MDR isolates did not differ between different wards in this study.
P. aeruginosa exhibits intrinsic resistance to penicillin G, aminopenicillins, 1 st - and 2nd -generation cephalosporins, macrolides, tetracyclines, chloramphenicol, sulphonamides, and trimethoprim34. Our findings revealed the second-highest resistance frequencies to cefotaxime (a 3rd -generation cephalosporin) and tobramycin (an aminoglycoside), with frequencies of 47.1% and 40%, respectively. Previous studies have reported resistance rates of P. aeruginosaisolates to ceftazidime (63%) and tobramycin (50%) by Abd El-Baky et al35., and to ceftazidime (57.1%) and tobramycin (38.1%) by Bazghandi et al28.. These findings underscore the critical need for robust antibiotic stewardship programs to mitigate resistance development and preserve the efficacy of available antimicrobial agents against P. aeruginosa.
Notably, the highest resistance in our study was observed against colistin, with 52 isolates (74.3%) being resistant. This finding contrasts with previous reports, which documented lower frequencies of colistin-resistant P. aeruginosa isolates from clinical samples. A recent systematic review and meta-analysis revealed that the global resistance rate to colistin has increased from 2% to 5% from 2020 to 2023, with the highest resistance rate (11%) observed among XDR isolates36. Another high resistance rate was 21.1% in the study by Abd El-Baky et al35., which was significantly lower than the prevalence observed in our study. This discrepancy may be attributed to using an alternative method (colistin disc elution) rather than the commonly used method (broth microdilution) to determine colistin susceptibility. This discrepancy in the reported resistance rates from different assessment methods has also been observed in other studies36,37,38. Nevertheless, the mentioned high resistance rate to colistin necessitates exploring alternative last-resort treatment options.
Along with colistin, carbapenems are considered a last-resort treatment option for infections caused by MDR Gram-negative bacteria, such as P. aeruginosa. Unfortunately, the number of CRPA infections is increasing globally7. Among the various mechanisms conferring carbapenem resistance, the production of carbapenemases stands out as a significant contributor9. In the current study, 37.1% of P. aeruginosa isolates were identified as CRPA, with 5 of these (19.2%) being carbapenemase producers. Amanati et al. reported the prevalence of CRPA in blood cultures as 71.6%, which was significantly higher than the prevalence observed in the present study29. However, the discovered prevalence was consistent with reported frequencies of 46.7% by Park et al39., 52.2% by Heidari et al31., and 41.9% by Xie et al40.. The increasing prevalence of CRPA further necessitates the implementation of robust antibiotic stewardship programs.
The frequency of carbapenemase-mediated genes was four (5.7%) for blaIMP and one (1.4%) for blaOXA-48. Additionally, blaKPC, blaVIM, and blaSIM were not detected. This finding aligns with the number of detected carbapenemase-producing P. aeruginosa isolates. However, Park et al.. reported a frequency of 35% for blaIMP, while they did not detect blaKPC, blaVIM, or blaOXA-4839. Additionally, other studies recorded frequencies of 75%, 62.5%, and 37.5% for the presence of blaIMP, blaVIM, and blaKPC genes among 16 carbapenemase-producing P. aeruginosa isolates, respectively40. This comparison suggests that the distribution of carbapenemase-mediated genes is widely different. Thus, further local epidemiological investigations could provide valuable information for determining antimicrobial stewardship and policy development.
The mortality rate stemming from CRPA infections remains high while exhibiting cross-resistance towards other common antimicrobial agents41. Combination therapy serves as an effective strategy for managing infections caused by MDR and XDR bacteria. This strategy has become attractive alongside increasing carbapenem- and colistin-resistant bacteria42. The literature indicates that most previous studies have focused on combinations of two antibiotics or antibiotics combined with oil extractions. Furthermore, articles evaluating combinations of three antibiotics have not extensively tested P. aeruginosa isolates. However, EL-Azizi determined a 100% synergistic effect of the combination of amikacin, ciprofloxacin, and ceftazidime against P. aeruginosa isolates43.
Our in vitro experiments revealed a synergistic effect between meropenem, amikacin, and ciprofloxacin at certain concentrations against CRPA isolates. Specifically, a synergistic effect was observed for 5 out of 8 tested isolates; however, it was not detected in two isolates with a high MIC of meropenem (512 µg/mL). Figures 3 and 4 demonstrate that a synergistic effect was likely only when the MIC values of amikacin, ciprofloxacin, and meropenem in the combination setting did not exceed 2, 6, and 24 µg/mL. This finding suggests that the combination therapy of meropenem, amikacin, and ciprofloxacin should be avoided when the MIC value of any of these antibiotics in the combination setting exceeds the mentioned value. However, if this criteria is met, there is a chance of a synergistic effect on the CRPA isolates. This could be particularly practical in resource-limited areas, where limited access to novel antibiotics may further give rise to MDR44. However, considering the small sample size and the number of instances for which the synergistic effect was observed, further studies are required to confirm this finding.
This study examined the resistance patterns of P. aeruginosa isolates from various wards across three hospitals, focusing on the synergistic effects of amikacin, ciprofloxacin, and meropenem. However, limitations exist. Notably, we did not test for NDM among class B carbapenemases (MBLs), which are known to be prevalent in P. aeruginosa. The low frequency of carbapenemase genes suggests that resistance may primarily stem from non-carbapenemase mechanisms, such as porin loss and efflux pump overexpression. Future studies should address this gap by assessing NDM and other types of carbapenemases to better elucidate resistance mechanisms. Additionally, limited variability in the MIC values for individual antibiotics restricted our analysis to combination settings. We recommend future research investigate the impact of individual MIC values on combination outcomes. Finally, given the role of biofilm production as a resistance mechanism, evaluating the antibiofilm activity of various antibiotic combinations, alongside larger sample sizes and time-kill assays, will provide more clinically relevant insights for improving treatment strategies against P. aeruginosa infections.
Conclusion
In this study, P. aeruginosa isolates exhibited a high prevalence of carbapenem resistance, with CRPA accounting for 26 of 70 (37.1%) samples. A significant correlation was observed between CRPA and MDR, further highlighting the therapeutic challenges associated with these infections. The three-dimensional checkerboard assay demonstrated a synergistic effect when the combination MIC values of meropenem, amikacin, and ciprofloxacin were at ≤ 24, ≤2, and ≤ 6 µg/mL, respectively. These findings highilight potential areas where combination therapy could be beneficial and suggest that this approach might be a promising strategy for managing CRAP infections, particularly in resource-limited settings. However, it is important to note that the sample size in this study is small, and these findings should not be interpreted as clinical recommendations at this stage. Further investigations are necessary to validate the efficacy of this combination and optimize dosing strategies, as well as to assess the in vivo effects of the suggested therapeutic combinations. Therefore, these results should be viewed as hypotheses for future research rather than definitive clinical guidelines.
Data availability
The datasets used and/or analysed during the current study available from the corresponding author on reasonable request.
Abbreviations
- FICI:
-
Fractional inhibitory concentration index
- HAIs:
-
Hospital-acquired infections
- P. aeruginosa :
-
Pseudomonas aeruginosa
- CRPA:
-
Carbapenem-resistant P. aeruginosa
- MDR:
-
Multi-drug resistant
- XDR:
-
Extensively-drug resistant
- MIC:
-
Minimum inhibitory concentration
- TSI:
-
Triple sugar iron
- OF:
-
Oxidation-fermentation
- TSB:
-
Tryptic soy broth
- PCR:
-
Polymerase chain reaction
- CLSI:
-
Clinical of Laboratory Standard Institute
- CAMHB:
-
Cation-adjusted Muller-Hinton broth
- mCIM:
-
Modified carbapenem inactivation method
References
Meshkat, Z. et al. Detection of efflux pump genes in multiresistant Acinetobacter baumannii ST2 in Iran. Acta Microbiol. Immunol. Hung. 68(2), 113–20 (2021).
Ahmadi, N. et al. Genomic diversity of β-lactamase producing Pseudomonas aeruginosa in Iran; the impact of global high-risk clones. Ann. Clin. Microbiol. Antimicrob. 23(1), 5 (2024).
Folic, M. M., Djordjevic, Z., Folic, N., Radojevic, M. Z. & Jankovic, S. M. Epidemiology and risk factors for healthcare-associated infections caused by Pseudomonas aeruginosa. J. Chemother. 33(5), 294–301 (2021).
Sathe, N. et al. Pseudomonas aeruginosa: Infections and novel approaches to treatment “Knowing the enemy” the threat of Pseudomonas aeruginosa and exploring novel approaches to treatment. Infect. Med. 2(3), 178–94 (2023).
Thomsen, J. et al. Evolving trends among Pseudomonas aeruginosa: A 12-year retrospective study from the United Arab Emirates. Front. Public Health 11, 1243973 (2023).
Flynn, C. E. & Guarner, J. Emerging antimicrobial resistance. Mod. Pathol. 36(9), 100249 (2023).
Jean, S.-S., Harnod, D. & Hsueh, P.-R. Global threat of carbapenem-resistant gram-negative bacteria. Front. Cell. Infect. Microbiol. 12, 823684 (2022).
Zomorodi, A. R. et al. Investigation of mobile colistin resistance (mcr) genes among carbapenem resistance Pseudomonas aeruginosa isolates from bovine mastitis in Mashhad, Iran. Gene Rep. 29, 101695 (2022).
Stanton, R. A. et al. Whole-genome sequencing reveals diversity of carbapenem-resistant Pseudomonas aeruginosa collected through CDC’s emerging infections program, United States, 2016–2018. Antimicrob. Agents Chemother. 66(9), e00496-22 (2022).
Liu, Y. et al. Drug repurposing for next-generation combination therapies against multidrug-resistant bacteria. Theranostics 11 (10), 4910 (2021).
Ghorbani, H., Memar, M. Y., Sefidan, F. Y., Yekani, M. & Ghotaslou, R. In vitro synergy of antibiotic combinations against planktonic and biofilm Pseudomonas aeruginosa. GMS Hyg. Infect. Control 12, Doc17. https://doi.org/10.3205/dgkh000302 (2017).
Farhan, S. M. et al. Effect of imipenem and amikacin combination against multi-drug resistant Pseudomonas aeruginosa. Antibiotics 10 (11), 1429 (2021).
Tarazi, Y. H., Abu-Basha, E., Ismail, Z. B. & Al-Jawasreh, S. I. Antimicrobial susceptibility of multidrug-resistant Pseudomonas aeruginosa isolated from drinking water and hospitalized patients in Jordan. Acta Trop. 217, 105859 (2021).
Rafati Zomorodi, A., Rad, M., Hashemitabar, G. R. & Salimizand, H. Molecular typing of cephalosporin resistant serovars of Salmonella enterica from poultry and farm animals. Bulg. J. Vet. Med. https://doi.org/10.15547/bjvm.2196 (2020).
Poirel, L., Walsh, T. R., Cuvillier, V. & Nordmann, P. Multiplex PCR for detection of acquired carbapenemase genes. Diagn. Microbiol. Infect. Dis. 70(1), 119–23 (2011).
CLSI. Performance Standards for Antimicrobial Susceptibility Testing. 34th ed. CLSI supplement M100. Clinical and Laboratory Standards Institute. 2024. (2024).
Stein, C. et al. Three dimensional checkerboard synergy analysis of colistin, meropenem, tigecycline against multidrug-resistant clinical Klebsiella pneumonia isolates. PloS one. 10 (6), e0126479 (2015).
Schwartz, B., Klamer, K., Zimmerman, J., Kale-Pradhan, P. B. & Bhargava, A. Multidrug resistant Pseudomonas aeruginosa in clinical settings: A review of resistance mechanisms and treatment strategies. Pathogens 13(11), 975 (2024).
Sendra, E. et al. Impact of multidrug resistance on the virulence and fitness of Pseudomonas aeruginosa: A microbiological and clinical perspective. Infection 52(4), 1235–68 (2024).
Yusuf, E., Zavascki, A. P., Endeman, H. & Kalil, A. C. Which trial do we need? One or two antimicrobials with anti-pseudomonal activity for the empirical treatment of ventilator-associated pneumonia due to Gram-negative bacteria. Clin. Microbiol. Infect. 29(11), 1364–6 (2023).
Reynolds, D. & Kollef, M. The epidemiology and pathogenesis and treatment of Pseudomonas aeruginosa infections: An update. Drugs 81(18), 2117–2131 (2021).
Corcione, S. et al. Epidemiology, prevalence and risk factors for infections in burn patients: results from a regional burn centre’s analysis. J. Chemother. 33 (1), 62–66 (2021).
Wang, W. & Wang, X. Prevalence of metallo-β-lactamase genes among Pseudomonas aeruginosa isolated from various clinical samples in China. J. Lab. Med. 44(4), 197–203 (2020).
Elahi, G., Goli, H. R., Shafiei, M., Nikbin, V. S. & Gholami, M. Antimicrobial resistance, virulence gene profiling, and genetic diversity of multidrug-resistant Pseudomonas aeruginosa isolates in Mazandaran, Iran. BMC Microbiol. 24(1), 546 (2024).
Chand, Y. et al. Clinical specimens are the pool of multidrug-resistant Pseudomonas aeruginosa harbouring oprL and toxA virulence genes: Findings from a Tertiary Hospital of Nepal. Emerg. Med. Int. 2021(1), 4120697 (2021).
Mekonnen, H., Seid, A., Molla Fenta, G. & Gebrecherkos, T. Antimicrobial resistance profiles and associated factors of Acinetobacter and Pseudomonas aeruginosa nosocomial infection among patients admitted at Dessie comprehensive specialized Hospital, North-East Ethiopia. A cross-sectional study. PLoS One 16(11), e0257272 (2021).
Mirzaei, B. et al. Prevalence of multi-drug resistant (MDR) and extensively drug-resistant (XDR) phenotypes of Pseudomonas aeruginosa and Acinetobacter baumannii isolated in clinical samples from Northeast of Iran. BMC Res. Notes 13, 1–6 (2020).
Bazghandi, S. A. et al. Prevalence of virulence genes and drug resistance profiles of Pseudomonas aeruginosa isolated from clinical specimens. Jundishapur J. Microbiol. 14(8), e118452. https://doi.org/10.5812/jjm.118452 (2021).
Amanati, A. et al. Bloodstream infections in adult patients with malignancy, epidemiology, microbiology, and risk factors associated with mortality and multi-drug resistance. BMC Infect. Dis. 21 (1), 636 (2021).
Brzozowski, M., Krukowska, Ż, Galant, K., Jursa-Kulesza, J. & Kosik-Bogacka, D. Genotypic characterisation and antimicrobial resistance of Pseudomonas aeruginosa strains isolated from patients of different hospitals and medical centres in Poland. BMC Infect. Dis. 20, 1–9 (2020).
Heidari, R. et al. Antibiotic resistance, biofilm production ability and genetic diversity of carbapenem-resistant Pseudomonas aeruginosa strains isolated from nosocomial infections in southwestern Iran. Mol. Biol. Rep. 49(5), 3811–22 (2022).
Asmare, Z. et al. Antimicrobial resistance profile of Pseudomonas aeruginosa clinical isolates from healthcare-associated infections in Ethiopia: A systematic review and meta-analysis. PLoS One 19(8), e0308946 (2024).
Raman, G., Avendano, E. E., Chan, J., Merchant, S. & Puzniak, L. Risk factors for hospitalized patients with resistant or multidrug-resistant Pseudomonas aeruginosa infections: A systematic review and meta-analysis. Antimicrob. Resist. Infect. Control 7, 1–14 (2018).
Mączyńska, B. et al. Changes in antibiotic resistance of Acinetobacter baumannii and Pseudomonas aeruginosa clinical isolates in a multi-profile hospital in years 2017–2022 in Wroclaw, Poland. J. Clin. Med. 12(15), 5020 (2023).
Abd El-Baky, R. M. et al. Prevalence and some possible mechanisms of colistin resistance among multidrug-resistant and extensively drug-resistant Pseudomonas aeruginosa. Infect. Drug Resist. https://doi.org/10.2147/IDR.S238811 (2020).
Narimisa, N. et al. Prevalence of colistin resistance in clinical isolates of Pseudomonas aeruginosa: A systematic review and meta-analysis. Front. Microbiol. https://doi.org/10.3389/fmicb.2024.1477836 (2024).
Yadav, K. S., Pawar, S., Yadav, S. A. & Patil, S. Comparative analysis of colistin resistance in Pseudomonas aeruginosa: VITEK® 2 compact versus broth microdilution method. Cureus 16(12), e76646 (2024).
Banerjee, T. et al. Comparative evaluation of colistin broth disc elution (CBDE) and broth microdilution (BMD) in clinical isolates of Pseudomonas aeruginosa with special reference to heteroresistance. Indian J. Med. Microbiol. 47, 100494 (2024).
Park, Y. & Koo, S. H. Epidemiology, molecular characteristics, and virulence factors of carbapenem-resistant Pseudomonas aeruginosa isolated from patients with urinary tract infections. Infect. Drug Resist. https://doi.org/10.2147/IDR.S346313 (2022).
Xie, X. et al. Molecular epidemiology and carbapenem resistance mechanisms of Pseudomonas aeruginosa isolated from a hospital in Fujian, China. Front. Microbiol. 15, 1431154 (2024).
Zhou, R. et al. Impact of carbapenem resistance on mortality in patients infected with Enterobacteriaceae: A systematic review and meta-analysis. BMJ Open 11(12), e054971 (2021).
Coates, A. R., Hu, Y., Holt, J. & Yeh, P. Antibiotic combination therapy against resistant bacterial infections: Synergy, rejuvenation and resistance reduction. Expert Rev. Anti-Infect. Ther. 18(1), 5–15 (2020).
El-Azizi, M. Novel microdilution method to assess double and triple antibiotic combination therapy in vitro. Int. J. Microbiol. 2016(1), 4612021 (2016).
Cohn, J. et al. Accelerating antibiotic access and stewardship: A new model to safeguard public health. Lancet Infect. Dis. 24(9), e584–e90 (2024).
Acknowledgements
We thank the “Clinical Research Development Center, Amir Oncology Teaching Hospital, Shiraz University of Medical Sciences” for granting us access to the cancer registry center database, officially known as the Amir Hospital-Based Cancer Registry..
Funding
This work, as part of Zahra Fooladfar’s MSc. thesis, was made possible by the Vice-Chancellor for Research at Shiraz University of Medical Sciences. The grant number is IR.SUMS.REC.01-27785.
Author information
Authors and Affiliations
Contributions
ZF was responsible for conceptualization, data curation, and investigation. ARZ performed the data curation, formal analysis, and writing of the original draft. MHPM curated the data and performed the formal analysis. EM was responsible for formal analysis, visualization, and writing – review & editing. SRAM collaborated in visualization and writing – review & editing. MM and AA were responsible for conceptualization and writing – review & editing. Additionally, MM validated and supervised the study. All authors have critically reviewed and approved the final version of the manuscript for submission.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
The Ethics Committee of Shiraz University of Medical Sciences gave their approval for this study with the code number IR.SUMS.REC.1402.466. The written informed consent was signed by each participant for the anonymous use of their data. Parents or legal guardians provided written informed permission for individuals under 16 years. We adhered to all of the World Medical Association Declaration of Helsinki’s ethical guidelines for doing research on human subjects in medicine.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Fooladfar, Z., Rafati Zomorodi, A., Parvizi Mashhadi, M.H. et al. Synergistic effects of meropenem, amikacin, and ciprofloxacin against carbapenem resistant Pseudomonas aeruginosa isolates. Sci Rep 16, 10044 (2026). https://doi.org/10.1038/s41598-026-40988-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-40988-4