Abstract
The 5′ untranslated region (5′UTR) plays a pivotal role in controlling translation efficiency and protein synthesis. However, existing models often struggle to jointly capture local regulatory motifs and long-range dependencies while effectively integrating multimodal biological features. We present UTR-DynaPro, a multimodal language model that combines a parallel CNN–Transformer architecture with a k-mer–specific mixture-of-experts module and a dynamic fusion mechanism. The CNN branch extracts contiguous motif patterns, while the Transformer branch models hierarchical long-range interactions. To address the complexity of 5’UTR regulation, a dynamic fusion gate is employed to integrate sequence-derived embeddings with key biophysical and structural determinants, including minimum free energy, CDS length, AT ratio, G/C content and upstream open reading frames (uORFs). Across translation efficiency (quantified by mean ribosome loading) and expression level prediction tasks, UTR-DynaPro achieves up to 3.3%, 2.2%, and 2.4% improvements over state-of-the-art methods, respectively. Attention-based motif analysis further identifies both known and novel regulatory elements with consistent performance across cell types, offering a generalizable framework for decoding complex 5′UTR regulation and guiding the design of high-performance regulatory sequences.
Similar content being viewed by others
Introduction
Messenger ribonucleic acid (mRNA), as the core carrier of genetic information transmission, is a single-stranded nucleotide molecule generated by DNA transcription1,2. It decodes the genetic information into protein synthesis instructions through translation and serves a fundamental function in the central dogma3,4. Mature mRNA contains two functional modules, the coding region (CDS) and the untranslated region (UTR), of which the 5’untranslated region (5'UTR) has become a focus area of gene expression regulation research due to its special sequence features and spatial conformation5.
As the core hub of mRNA translation regulation, the 5′ untranslated region (5′UTR) plays a crucial role in modulating gene expression and translation. It exerts its regulatory function through a complex, multi-layered mechanism that ensures precise control of gene expression at the translational level6,7,8. The efficiency of protein synthesis, crucial for mRNA vaccine development, is strongly influenced by the 5′UTR9,10,11. Though the same translational machinery is used by most cellular mRNAs, their translation kinetics often vary in function12,Shah et al.13). For example, strong secondary structures within the 5’UTR of mRNAs can impede ribosome scanning, thereby reducing the rate of protein synthesis14,15,16,17, Weinberg18 et al. proposed that the length of the CDS and 5’UTR, the G/C content of the 5’UTR, the number of uAUGs, and the ribosome density of different mRNAs, all affect the level of mRNA expression. Although there are known cases of regulation at each stage of translation, there are many factors that are largely under the control of translation at its onset, all of which affect mRNA translation rates19,Chu and von der Haar20,13.
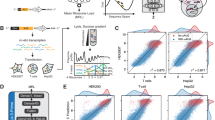
The 5′UTR regulates translation through diverse mechanisms21 is shown in Fig. 1, including modulation of ribosome recruitment and scanning, interaction with RNA-binding proteins22,23, and the influence of structural and sequence elements such as upstream AUGs, GC content, and secondary structure stability. While these individual determinants have been characterized, the quantitative and dynamic interplay between non-coding sequence features and protein outQ2put, particularly the synergistic regulation between the 5′UTR and coding sequence, remains largely unexplored. This gap in understanding limits our ability to rationally design mRNA constructs for applications such as vaccine development and gene therapy.
5'UTR sequence and protein molecule translation regulation mechanism map. The 5′UTR integrates sequence features (e.g., CDS length, nucleotide composition, AT ratio, and upstream open reading frames [uORFs]) and RNA secondary structures to regulate translation. These determinants collectively influence multiple quantitative traits, including mean ribosome load (MRL), translation efficiency (TE), and expression level (EL), which ultimately shape protein output, such as in the context of mRNA vaccines.
As the research on 5’UTR is gradually deepening, more and more algorithms combined with deep learning are being used in the prediction of 5’UTR function24,25,Li et al.26,27,28, such as predicting mRNA translation efficiency and mRNA expression level29. Advances in deep learning (DL) have prompted researchers to develop DL-based approaches to study RNA regulation; these include convolutional neural networks (CNNs),30 recurrent neural networks (RNNs)31, or integration of both architectures. These models have shown promise in predicting mRNA translation efficiency and expression levels32,33. However, these methods seem to have plateaued in performance, potentially missing key information and affecting prediction accuracy.
To address the technical bottlenecks in the analysis of the dynamic regulatory network mentioned above, this study innovatively proposes the UTR-DynaPro fusion architecture is shown in Fig. 2, which realizes the systematic decoding of the 5′UTR regulatory mechanism through multimodal feature fusion.
Conceptual framework of the UTR-Dynapro model for comprehensive prediction and rational design of 5′UTR functions. Endogenous data from multiple species, human cell line/tissue datasets, and independent test sets are used as input. The encoder integrates a mixture-of-experts gating module with parallel Transformer and CNN branches to capture both global dependencies and local sequence features, followed by dynamic fusion and nonlinear transformation. The model is pre-trained with minimum free energy (MFE) and masked nucleotide (MN) objectives, and subsequently fine-tuned on downstream tasks, including mean ribosome load (MRL), translation efficiency (TE), and expression level (EL). Model performance is evaluated with multiple statistical metrics (e.g., Pearson R, Spearman R, RMSE), with checkpoint saving at the best epoch.
The core innovation of our model (UTR-Dynapro) is the integration of the CNN branch and the Transformer branch for parallel feature extraction. This design is inspired by the success of models like the Nucleic Transformer34, which demonstrated that combining self-attention with convolutional operations achieves superior performance in tasks such as DNA and promoter classification. While traditional RNNs are limited by stepwise processing and standard CNNs primarily capture local features35, our parallel design leverages the complementary strengths of both: the CNN branch excels at distilling local regulatory motifs, while the Transformer branch, powered by its self-attention mechanism, captures global long-range interactions across the entire 5′UTR sequence. By merging these complementary strengths, UTR-DynaPro effectively decodes the complex, multi-scale regulatory signals inherent in 5′UTR sequences.
In particular, this model introduces a dedicated representation layer to integrate biophysical parameters (e.g., minimum free energy) with structural genomic determinants (e.g., CDS length, AT ratio, and upstream open reading frames) via high-order tensor fusion. This approach constructs a comprehensive feature space for 5′UTR regulation. Through systematic evaluation on multiple representative high-throughput sequencing datasets, UTR-DynaPro significantly outperforms existing state-of-the-art methods across key regulatory metrics, including mean ribosome loading (MRL), expression level (EL), and translation efficiency (TE), while demonstrating superior cross-dataset generalization. This study not only proposes a flexible and efficient framework for 5′UTR modeling but also provides a robust foundation for decoding post-transcriptional regulatory mechanisms, functional mining of non-coding regions, and sequence-based biological design.
Method
Overview of UTR-DynaPro
In this study, we propose UTR-DynaPro, a deep learning architecture for modeling and predicting 5′UTR sequences. The model uses UTR-LM, a pre-trained language model based on the Evolutionary Scale Modeling 2 (ESM-2) framework, as an encoder to obtain contextual embedding representations. In the feature Fusion Module, we introduce a Parallel Convolution–Transformer Layer, which jointly captures local k-mer patterns and long-range dependencies. The convolutional branch extracts contiguous sequence motifs, while the Transformer branch models hierarchical global interactions. Their outputs are adaptively integrated through a Dynamic Fusion Gate, which assigns position-specific weights to local and global features. Furthermore, a Mixture of Experts layer with k-mer specific experts is incorporated into the feed-forward network to enhance multi-scale feature representation. Our Dynamic Fusion Gate uses a multi-head attention-style mechanism to compute adaptive weights for local (CNN) and global (Transformer) representations, enabling the model to selectively prioritize regulatory motifs or long-range structures depending on the sequence context.
To accommodate 5′UTRs of varying lengths, the model integrates a Frame-Aware Pooling Layer that samples feature across the three translational frames. By aggregating both peak signals and average contexts from these frames, the model captures frame-specific regulatory patterns before combining them with experimental indicators for final prediction. The overall architecture is shown in Fig. 2.
The model adopts a customized UTR-LM encoder that re-implements the ESM-2 Transformer architecture specifically for nucleotide sequences. The backbone is initialized with random weights and trained from scratch on large-scale UTR data to capture RNA-specific regulatory logic. This encoder is integrated with a multi-scale Parallel Feature Fusion Module and fine-tuned for downstream regression tasks, including mean ribosome loading (MRL), translation efficiency (TE), and expression level (EL). By synergizing genomic representations with biophysical priors, UTR-DynaPro achieves superior predictive accuracy and robust generalization across diverse experimental conditions.
Feature fusion module: parallel convolution–transformer layers
The feature Fusion Module is composed of three stacked Parallel Convolution–Transformer Layers (PCTL). Each layer takes the input sequence representation \(X^{\left( l \right)} \in {\mathbb{R}}^{T \times d}\) as its starting point and first applies layer normalization to ensure numerical stability and consistent feature distribution. The normalized data is then fed simultaneously into two parallel branches:
In the Transformer branch, the input is first projected into query (Q), key (K), and value (V) spaces. Each attention head processes a subspace of dimension \({\raise0.5ex\hbox{$\scriptstyle d$} \kern-0.1em/\kern-0.15em \lower0.25ex\hbox{$\scriptstyle h$}}\), where the number of heads is fixed at h=8. Using the scaled dot-product attention mechanism, the model computes the pairwise relevance between any two positions and establishes global dependencies across the sequence. In a feature Fusion Module setting, if autoregressive generation is required, a lower-triangular mask is applied to the attention score matrix to ensure that each position attends only to past tokens. The multi-head outputs are concatenated and linearly transformed to produce the global branch output ZT, which has the same shape as the input.
In the convolution branch, a one-dimensional convolution kernel (with a default size of Kmer=7 is used to extract local patterns. Symmetric zero-padding is applied to ensure that the output length matches the input length. The convolution operates along the temporal dimension, attending only to the Kmer neighboring positions, thereby capturing local k-mer–level statistical features. A nonlinear activation function, such as GELU, is applied to introduce nonlinearity into the representation. The output of the convolution branch is denoted as Zc.
The outputs from the two branches are adaptively combined through a Dynamic Fusion Gate, whose gating weights are given by:
where \(\sigma \left( \cdot \right)\) denotes the Sigmoid activation function, \(W_{g}^{T}\) learns how much weight to assign to.
the global (Transformer) information.;\(W_{g}^{C}\) learns how much weight to assign to the local (convolution) information. The fusion is computed as
with \(\odot\) representing element-wise multiplication. This design ensures that the fusion is a convex combination of global and local information and can smoothly adjust the proportion of the two during the training process.
The fused result is added to the input through a residual connection to form an intermediate representation H, which is then passed through either a Feed-Forward Network (FFN) or a k-mer–specialized Mixture-of-Experts (MoE) network for nonlinear transformation and feature reconstruction. The FFN consists of two fully connected layers, typically with a higher-dimensional hidden layer in between to enhance representational capacity. The MoE, in contrast, employs a gating network to selectively activate a subset of expert sub-networks, thereby increasing the model’s diversity and capacity. Finally, the residual connection adds the FFN’s output back to H, producing the input X(l+1) for the next layer.
RNA-aware architectural design principles
Although UTR-DynaPro builds upon established deep learning components, its methodological contribution lies in how these components are adapted and integrated to reflect the biological properties of 5′UTR -mediated translation regulation. Each nucleotide is treated as a biologically meaningful token rather than a generic character, with special tokens (BOS/EOS) delimiting regulatory regions and contextual embeddings from the pretrained encoder capturing biochemical and evolutionary semantics beyond simple one-hot encoding. This representation enables the model to reason over regulatory motifs and structural context.
5′UTRs exhibit substantial length variability (25–100 nt and beyond) and encode regulatory information in a frame-dependent manner. Instead of truncating sequences, we preserve full-length representations and employ masked attention and frame-aware pooling to obtain fixed-dimensional outputs, ensuring that both short and long regulatory elements are retained. Frame-aware pooling aggregates features separately over the three reading frames, allowing the model to explicitly capture frame-specific regulatory patterns such as upstream open reading frames (uORFs) and start-codon context.
Regulatory mechanisms in 5′UTRs arise from the interplay between short motifs (e.g., Kozak sequence, uAUGs) and long-range dependencies (e.g., secondary structure and CDS co-adaptivity), as well as heterogeneous regulatory programs across transcripts. The parallel CNN-Transformer design mirrors this biology by capturing local motifs and global interactions, while the Dynamic Fusion Gate enables position-wise adaptation between these two regimes. The \(\text{k}\)-mer-specific Mixture-of-Experts module further allows the model to route sequence segments to specialized experts, reflecting diverse regulatory mechanisms such as structure-driven versus motif-driven control.
Model workflow
Notation
Throughout this section, we use bold uppercase letters to denote tensors, bold lowercase letters for vectors, and italic letters for scalars. For example, \(H\in {R}^{B\times L\times d}\) denotes a tensor, \({h}_{i}\in {R}^{d}\) denotes a vector, and \(\text{d}\) denotes the embedding dimension.
Data representation and tokenization
Each 5′UTR sequence is represented as a nucleotide string \(\mathcal{S}=\left({s}_{1},{s}_{2},\dots ,{s}_{L}\right)\), where \({s}_{i}\in \{A,C,G,U\}\) and \(L\) denotes the sequence length. To facilitate model processing, we tokenize the sequence using a fixed vocabulary:
Special tokens are prepended and appended to the sequence to define the boundaries, resulting in the augmented sequence \(\widetilde{\mathcal{S}}=\left(BOS,{s}_{1},{s}_{2},\dots ,{s}_{L},EOS\right)\).
The tokenized sequence is then mapped into a high-dimensional space through an embedding layer. Specifically, a pretrained encoder (e.g., UTR-LM or ESM-2 backbone) is employed to transform \(\widetilde{\mathcal{S}}\) into contextualized representations:
where \(\text{B}\) is the batch size and $d$ represents the embedding dimension.
To focus on the biologically meaningful nucleotide information and eliminate the influence of auxiliary tokens, we strip the embeddings corresponding to the BOS and EOS positions. The resulting content embeddings \(E\) are defined as:
where \(H=\left({h}_{1},{h}_{2},\dots ,{h}_{L+2}\right)\). This refined representation \(E\) serves as the primary input for the subsequent parallel decoding architecture.
Feature fusion module input and processing
The feature Fusion Module consists of \({\text{L}}_{dec}=3\) stacked Parallel Convolution-Transformer Layers (PCTLs). The biologically relevant encoder output \(\text{E}\), obtained after removing special tokens, is directly used as the input to the feature Fusion Module.
Each PCTL contains a Transformer branch and a convolution branch operating in parallel. The Transformer branch models long-range dependencies using multi-head self-attention, while the convolution branch extracts local \(\text{k}-\text{mer}\) motifs using a one-dimensional convolution with kernel size \({\text{k}}_{mer}\).
Their outputs are adaptively fused by a Dynamic Fusion Gate:
where \(\upsigma \left(\cdot \right)\) denotes the sigmoid function and \(\odot\) denotes element-wise multiplication.
The fused representation is then passed through a feed-forward network or a \(\text{k}\)-mer specific Mixture-of-Experts (MoE) block, with residual connections, to produce the output of the layer.
To explicitly integrate biological priors during inference, we define a biophysical feature vector \({\text{v}}_{\text{bio}}\in {\mathbb{R}}^{n}\) containing attributes such as MFE and CDS co-adaptivity. The final fused representation \({\text{u}}_{\text{final}}\) is constructed by projecting and concatenating the sequence-derived features \(\text{p}\), the biological vector \({\text{v}}_{\text{bio}}\), and the experimental metadata \(\text{e}\):
where \({\text{W}}_{\text{p}}\), \({\text{W}}_{\text{bio}}\), and \({\text{W}}_{\text{e}}\) are learnable weight matrices used for dimensional alignment. This formulation ensures that the model actively incorporates heterogeneous biological information to refine the prediction of translation efficiency.
Frame-based pooling
The output of the final feature Fusion Module layer is a tensor \(Z\in {R}^{B\times L\times d}\). We split \(Z\) into three reading frames and apply masked global max-pooling and mean-pooling to each frame. The pooled features are concatenated to form a \(6d\)-dimensional vector \(\text{p}\), which summarizes translation-frame–aware sequence information.
Metadata fusion and prediction head
An experimental indicator vector \(e \in {\mathbb{R}}^{B \times 2}\) is linearly projected to the same hidden size as the pooled content features and fused additively:
Where \(W_{p} \in {\mathbb{R}}^{6d \times n}\), \(W_{e} \in {\mathbb{R}}^{2 \times n}\),and \(n\) is the hidden size of the prediction head.
The fused representation is passed through a three-layer MLP with ReLU activation and dropout, producing the final scalar regression output.
Operational mode
This feature Fusion Module operates in fully parallel mode for both training and inference; there is no auto-regressive decoding. Self-attention layers employ padding masks but no causal masks, as the task does not involve next-step prediction.
Experiments
Dataset
The primary data used in this study were derived from multiple collections of publicly available experiments22,36,33, which measured the translation efficiency (TE) of a large number of synthesized 5′UTR sequences under different experimental conditions, Mean Ribosome Load (MRL), Transcription Efficiency(TE)and Expression Level (EL) under different experimental conditions. Each sample contains the nucleotide sequence itself, the corresponding experiment condition, and the experimentally determined phenotypic values (e.g., TE, MRL, and EL).
MRL dataset
For MRL prediction, we used a dataset from33 containing 5′UTR sequences and corresponding MRL values. This dataset is split into two subsets: Random_Vary, with 83,919 random 5′UTR sequences (25–100 nucleotides), and Human_Vary, with 15,555 human 5′UTR sequences of similar length.
The second dataset contains endogenous 5′UTR sequences derived primarily from the NCBI RefSeq database36 as well as mammalian sequences obtained through the UCSC Genome Browser’s Table Browser tool. The dataset contains 323,153 sequences from seven species: human (51,245), chimpanzee (68,511), bonobo (58,825), rhesus (56,902), macaque (53,638), mouse (32,678), and virus (1,354). A stringent cross-species deduplication was applied to the combined dataset to ensure each 5′UTR entry is a unique sequence-phenotype pair. By integrating sequences from seven evolutionary distant clades (ranging from humans to viruses), the resulting 323,153 non-redundant samples provide a diverse sequence space that minimizes the impact of lineage-specific conservation on model evaluation. Details of the species distribution can be found in Supplementary Table 1.
TE and EL dataset
In this study, we utilized three endogenous human 5′UTR datasets analyzed by22 et al. to predict EL and TE. These datasets contain the corresponding 5′UTR sequences along with their associated EL and TE values, which were measured using Ribo-seq RPKM. The datasets came from Human embryonic kidney cell line HEK293T, Human prostate cancer cell line PC3, and human muscle tissue, containing 14,410, 12,579, and 1,257 sequences, respectively.
For all downstream analyses, the data were partitioned into training (80%), validation (10%), and test (10%) sets, ensuring that the distribution of experimental conditions remained consistent across subsets. To improve the model’s adaptability to varying experimental conditions, we introduced an experiment indicator, encoded using one-hot encoding, as an additional input for each sample. This allows the model to explicitly incorporate experimental contextual information during multitask training.
Training setup
All models were implemented in PyTorch and trained on a single NVIDIA GeForce RTX 4090 GPU (24 GB), and each method was run 6 times to obtain average results. The encoder was initialized with pretrained. The optimizer was SGD with an initial learning rate of \(1 \times 10^{ - 4}\) momentum 0.9, and weight decay \(1 \times 10^{ - 4}\), and the loss function was the Huber loss. The feature Fusion Module consisted of three stacked Parallel Convolution–Transformer Layers, each using 16 attention heads in the Transformer branch and a 1D convolution branch with kernel size \(k_{mer} = 7\) and symmetric padding.
Implementation of benchmarks
We replicated and evaluated several representative benchmark methods under a unified training and evaluation framework, including Optimus33, FramePool37, MTtrans, RNABERT38, RNA-FM39, as well as two standard architectures: a one-dimensional convolutional neural network (CNN) and a vanilla Transformer. The basic CNN consists of multiple convolution–activation–pooling blocks to capture local kmer patterns, while the Transformer models long-range relationships using multi-head self-attention.
We re-implemented or adapted all methods from their official repositories, using hyperparameters from the original publications to maintain fairness. The same data preprocessing pipeline, training/validation/test splits, and downstream task definitions as in UTR-DynaPro were applied.
The pretrained RNA-FM and RNABERT models output embeddings of (sequence length, 640) and (sequence length, 120). We apply mean pooling along the sequence dimension to obtain fixed-size feature vectors (640 and 120 dimensions), which are then passed through a single-layer MLP to produce predictions for the downstream tasks.
For all baseline models, we use the same number of training epochs and the same evaluation metrics as UTR-DynaPro across three downstream prediction tasks: TE, MRL, EL. This unified setup allows us to systematically assess the generalization performance and robustness of each method in a multi-task prediction setting.
Results and analysis
Predicting mean ribosome load
MRL, defined as the mean number of ribosomes bound to an mRNA, serves as an important indicator of translation rate and efficiency40. This indicator is affected by various factors such as 5′ UTR sequence characteristics and regulation of translation initiation28. Accurate prediction of MRL not only helps to analyze the translation regulation mechanism of 5′ UTR, but also provides theoretical support for protein synthesis optimization.
In the Random_Vary dataset, UTR-DynaPro achieves excellent performance on 5′ UTR in all length intervals (Fig. 3a), with Spearman R of 0.913 (25-44nt), 0.924 (45-64nt), 0.924 (65-84nt) and 0.915 (85-100nt), reaching 0.925 overall, which is comprehensively better than baseline models such as UTR-Insight and UTR-LM. In contrast, UTR-Insight and UTR-LM performed closely behind with slightly lower correlation coefficients, while other methods such as MTtrans and Optimus were relatively weak, and RNAFM_MLP performed the worst.
Performance of UTR-Dynapro in predicting MRL for sequences of varying lengths. (a) Independent evaluation on random 5′ UTRs. (b) Independent evaluation on human 5′ UTRs.
In the Human_Vary dataset, UTR-DynaPro showed superior MRL prediction performance on 5′UTR sequences of different length intervals (Fig. 3b), with correlation coefficients of 0.833 (25–44 nt), 0.874 (45–64 nt), 0.864 (65-84nt) and 0.856 (85-100nt), and overall reaches 0.868, which are all ahead of UTR-Insight, UTR-LM, and other comparative models.The performance of UTR-Insight and UTR-LM is closely followed by slightly lower correlation coefficients, while the performance of MTtrans, Optimus and RNAFM_ResNet models have relatively weak performance, and RNAFM_MLP has the worst performance. This result indicates that it can stably capture the regulatory features of 5′ UTR sequences under different species and length conditions, which significantly improves the prediction effect of MRL.
Analysis of ablation experiments
On the Random_Vary dataset (Fig. 4a–f), UTR-DynaPro performed the best in all five metrics (R2, Pearson R, Spearman R, RMSE, MAE) (R2 = 0.908, Pearson R = 0.952, Spearman R = 0.937). Removing either the multi-scale fusion module (w/o Fusion) or the multi-expert gating mechanism (w/o MOE) significantly reduces the performance (R2 down to 0.877 and 0.868, respectively), suggesting that both components are critical for modelling complex structural features. Removing the experimental condition features (w/o ExpInfo) showed a significant increase in error although there was little change in correlation, suggesting that it helps to reduce prediction bias. Retaining only a single structure (Transformer Only or CNN Only) resulted in a significant decrease in performance (R2 = 0.735 and 0.703), further validating the importance of multimodal fusion. The performance gradually increases but saturates when the number of attentional headss increased from k = 3 to k = 7.
Ablation study of UTR-Dynapro on MRL prediction using the Random_Vary dataset (random 5′ UTRs). Performance was evaluated under different ablation settings, including removing specific modules (w/o Fusion, w/o MoE, w/o ExpInfo), restricting to Transformer-only or CNN-only architectures, and varying kernel sizes. Evaluation metrics include: (a) coefficient of determination (R2), (b) Pearson correlation coefficient, (c) Spearman correlation coefficient, (d) root mean square error (RMSE), and (e) mean absolute error (MAE). (f) Radar chart summarizing overall performance across all metrics. Results indicate that each module contributes to predictive accuracy, with the full UTR-Dynapro model achieving the best overall balance.
In the Human_Vary dataset (Fig. 5a–f), UTR-DynaPro was also optimal (R2 = 0.836, Pearson R = 0.918, Spearman R = 0.887). Removal of the Multiscale Fusion module (w/o Fusion) and the Multi-Expert Mechanism (w/o MOE) still caused performance degradation, with R2 dropping to 0.805 and 0.794. The non-use of experimental conditional features (w/o ExpInfo) slightly affects accuracy, but the RMSE and MAE are still smaller than the other ablation structures, suggesting that the module enhances the model’s ability to generalise. Transformer Only and CNN Only perform the worst, especially the MAE rises to 0.623 and 0.761 respectively, verifying the importance of the fused structure. The results of the settings with different numbers of attention kernels show that the best R2 = 0.749 and the smallest MAE = 0.468 are achieved at k = 7, again verifying the enhancement of the model modelling ability by the multi-head attention design.
Ablation study of UTR-Dynapro on MRL prediction using the Human_Vary dataset (human 5′ UTRs). Performance was evaluated under different ablation settings, including removing specific modules (w/o Fusion, w/o MoE, w/o ExpInfo), restricting to Transformer-only or CNN-only architectures, and varying kernel sizes. Evaluation metrics include: (a) coefficient of determination, (b) Pearson correlation coefficient, (c) Spearman correlation coefficient, (d) root mean square error, and (e) mean absolute error. (f) Radar chart summarizing overall performance across all metrics. Results indicate that each module contributes to predictive accuracy, with the full UTR-Dynapro model achieving the best overall balance.
MRL prediction performance compared with baseline methods
In the Random_Vary dataset (Fig. 6a–f), UTR-DynaPro achieved the optimal results for all five evaluation metrics (R2 = 0.908, Pearson R = 0.952, Spearman R = 0.937, RMSE = 0.462, MAE = 0.332). Compared with the closest performing UTR-Insight, the present model improves about 0.01–0.02 in correlation, while the RMSE and MAE are reduced by 7.8% and 6.3%, respectively, reflecting stronger error suppression. Informer slightly outperforms the UTR-LM in correlation, but still lags significantly behind in error control. UTR-DynaPro. There is a significant performance gap between UTR-LM based on general language model and Optimus and FramePool based on artificial features, especially on Spearman R, which is more than 0.05, indicating that there is a bottleneck in capturing the 5′UTR long-range modulation mode.
Prediction of MRL on the Random_Vary dataset (random 5′ UTRs) by UTR-Dynapro and baseline models (UTR-Insight, Informer, UTR-LM, Optimus, and FramePool). Performance comparison across multiple evaluation metrics: (a) coefficient of determination, (b) Pearson correlation coefficient, (c) Spearman correlation coefficient, (d) root mean square error, and (e) mean absolute error. (f) Radar chart summarizing overall performance across all metrics. UTR-Dynapro consistently achieves the best or near-best results, highlighting its superior accuracy and robustness compared to existing methods.
In the Human_Vary dataset (Fig. 7a–f), UTR-DynaPro again maintains the lead (R2 = 0.836, Pearson R = 0.918, Spearman R = 0.887, RMSE = 0.472, MAE = 0.336). Compared to UTR-Insight, the correlation improvement is relatively limited (~ 0.005–0.01), but the error is significantly reduced (RMSE by 6.1%, MAE by 8.2%), which is particularly critical for the interpretability and stability of applications with real human data. The rest of the methods, especially FramePool and Optimus, perform significantly worse in terms of correlation and error, which verifies the advantage of fusing multi-scale structural features with experimental condition information.
Prediction of MRL on the Human_Vary dataset ((human 5′ UTRs) by UTR-Dynapro and baseline models (UTR-Insight, Informer, UTR-LM, Optimus, and FramePool). Performance comparison across multiple evaluation metrics: (a) coefficient of determination, (b) Pearson correlation coefficient, (c) Spearman correlation coefficient, (d) root mean square error, and (e) mean absolute error. (f) Radar chart summarizing overall performance across all metrics. UTR-Dynapro consistently achieves the best or near-best results, highlighting its superior accuracy and robustness compared to existing methods.
Performance of UTR-DynaPro on TE and EL prediction
Translation Efficiency (TE) reflects the protein production per mRNA unit and serves as a key marker of post-transcriptional regulation. Expression Level (EL) is a central indicator of the overall production of target proteins in a cell, and is affected by multiple factors such as transcription level, translation efficiency, and protein stability32,11. The 5′UTR is crucial for regulating translation initiation and ribosome attachment, and its sequence identity has a significant effect on both EL and TE. Accurate prediction of these two indicators not only helps to understand the mechanism of gene expression regulation but also provides an important reference for sequence design in biotechnology and therapeutic protein production.
Analysis of ablation experiments
Analysis was performed on three endogenous human datasets, including muscle tissue, the PC3 prostate cancer cell line, and the HEK293T embryonic kidney cell line- UTR-DynaPro achieved Spearman R of 0.65, 0.66 and 0.58 respectively (Fig. 8a–c), all of which were optimal or tied for the best. The ablation experiments show that removing either the multi-scale fusion module (w/o Fusion) or the multi-expert mechanism (w/o MOE) leads to a performance degradation, validating its role in integrating multi-scale features and capturing complex modulation signals. Removal of experimental condition features (w/o ExpInfo) similarly degraded performance, suggesting that experimental background information helps to model UTR modulation patterns. A single architecture (Transformer Only or CNN Only) significantly outperforms the hybrid structure, suggesting that the two are complementary in capturing features at different levels. Replacing the dynamic convolutional kernel with a fixed convolutional kernel (k = 3, 5, 7) did not achieve the performance of the dynamic scheme, biao the multi-scale dynamic receptive field is more advantageous in adapting to modulation patterns of different lengths and complexities.
Prediction of mRNA translational efficiency (TE) and expression level (EL) on endogenous datasets. (a–c) Ablation studies of UTR-Dynapro on TE prediction for three cell types: muscle, PC3, and HEK, evaluated by Spearman correlation under different ablation settings (w/o Dynamic Fusion, w/o MoE, w/o ExpInfo, Transformer-only, CNN-only, and kernel sizes. (d) Comparative performance of UTR-Dynapro and baseline models (UTR-LM, Optimus, Cao-RF, Kipoi, MTtrans, RNAFM_MLP, and RNABERT_MLP) on TE prediction across the three cell types. (e) Comparative performance on EL prediction across the same cell types. UTR-Dynapro consistently achieves higher correlation values compared to baseline methods, highlighting its robustness and generalizability in modeling endogenous regulatory mechanisms.
TE prediction performance compared with baseline methods
We systematically compare UTR-DynaPro with multiple classes of representative methods (Fig. 8d), covering the language model classes (UTR-LM, RNAFM_MLP, RNABERT_MLP), generalised deep learning models (Optimus, MTtrans, Kipoi), and traditional machine learning methods (Cao-RF). In the Muscle and PC3 datasets, UTR-DynaPro achieves a Spearman R of 0.65 and 0.66, outperforming all the compared methods; in the HEK dataset, it is slightly lower than Optimus (0.58 vs. 0.60), but has the most stable performance overall. The performance of the traditional method Cao-RF was extremely low on all datasets (up to 0.04), relying on artificial features (e.g., k-mer frequencies, secondary structure energies, UTR lengths, and start codon positions, etc.), which is not scalable on large-scale and cross-conditional data, etc. The performance of UTR-LM was close to that of RNAFM_MLP in some conditions (e.g., Muscle: 0.63, PC3: 0.60, PC3: 0.60, PC3: 0.60, PC3: 0.60). UTR-LM and RNAFM_MLP were close to each other in some conditions (e.g. Muscle: 0.63, PC3: 0.63), but fluctuated a lot in the HEK dataset, suggesting that they have limited generalisation ability; Optimus and Kipoi could reach a moderate level in some conditions, but the consistency across conditions was poor; The Spearman R for MTtrans and RNABERT_MLP was below 0.5 across most datasets, suggesting weak performance in capturing 5′UTR regulation.
EL Prediction performance compared with baseline methods
Based on three independent sets of EL sequencing data (Muscle, PC3, HEK), we compared the final optimised UTR-Dynapro with the existing state-of-the-art model UTR-LM under the same evaluation conditions (Fig. 8e). UTR-Dynapro achieved Spearman correlation coefficients of 0.63, 0.60, and 0.59 on Muscle, PC3, and HEK datasets, respectively, which were stable and outperformed all the compared models on PC3 and HEK datasets. These results show that the proposed base model has excellent generalisation ability and robustness in the context of real cell lines, especially under different experimental conditions and significant differences in data distribution.
Model interpretability and visualization
Figure 9 present heatmaps of the Dynamic Fusion Gate distribution for samples from different batches, derived from the translation efficiency (TE) prediction dataset, covering two cell lines: HEK and PC3. Each input sample corresponds to an encoded mRNA sequence (with special padding tokens removed), where each position (“Position, valid”) represents a valid nucleotide token. The horizontal axis index 0 denotes the first valid token in the sequence, followed sequentially thereafter.
Dynamic fusion gate heat maps across cell lines. (a) Dynamic fusion gate heat map of HEK cell line samples. (b) Dynamic fusion gate heat map of PC3 cell line samples.
To standardize input lengths, sequences were padded during both training and inference; therefore, the absolute horizontal coordinate values are not directly aligned to the same biological positions across samples. A batch simply refers to a set of samples processed together in a single forward pass, where samples may come from different cell lines and exhibit substantial variation in sequence lengths.
The color scale indicates the relative contribution of the Transformer branch in the fused output: a value of 0 corresponds to a fully CNN-dominated contribution, while a value of 1 corresponds to a fully Transformer-dominated contribution. Intermediate values represent mixed contributions from both branches. To ensure comparability across samples, the color scale range is fixed at [0, 1]. This visualization reveals the model’s positional preference patterns between global context modeling (Transformer) and local pattern extraction (CNN), facilitating the analysis of feature fusion strategies across different cell lines and sequence structures.
Discussion
Deep learning has advanced the functional prediction of 5′ untranslated regions (5′UTRs), yet models often struggle with cross-condition generalization and interpretability. For example, UTR-LM32 captures large-scale sequence patterns but loses accuracy under varying experimental conditions, while CNN/RNN-based Optimus33 is limited in modeling long-range dependencies. Pretrained models such as RNAFM and RNABERT38,39 improve general RNA representation but remain suboptimal for translation-specific tasks.
UTR-DynaPro addresses these limitations with a multi-branch architecture combining CNNs and Transformers, enabling simultaneous modeling of local motifs and global dependencies. Its mixture-of-experts design and integration of experimental condition features further enhance adaptability across datasets and cell types. Our ablation studies confirm the necessity of these modules, particularly the role of contextual features in capturing regulatory signals. Compared with state-of-the-art models, UTR-DynaPro achieves superior cross-condition generalization and predictive stability, as shown in the Human_Vary dataset.
Beyond methodological advances, UTR-DynaPro offers practical value in rational 5′UTR design for applications such as protein expression optimization, gene therapy, and mRNA vaccine development. Future work could integrate richer experimental contexts, RNA structural and modification data, and multi-omics with causal inference to move from prediction toward mechanistic insights. With these extensions, UTR-DynaPro may serve as a versatile tool for both understanding translational regulation and advancing personalized medicine.
Data availability
The datasets utilized in this study are publicly available. The datasets for the translation efficiency (TE) and expression level (EL) tasks can be accessed at ttps://codeocean.com/capsule/671182211, which includes both the training data for the pretrained model and datasets for various downstream tasks. The data for the mean ribosome load (MRL) task have been deposited in the OMIX database (https://ngdc.cncb.ac.cn/omix), part of the Genome Sequence Archive (GSA) at the China National Center for Bioinformation41,42, and are available under the accession number OMIX008723.
The datasets utilized in this study are publicly available. The datasets for the translation efficiency (TE) and expression level (EL) tasks can be accessed at https://codeocean.com/capsule/6711822(Chu et al., 2024), which includes both the training data for the pretrained model and datasets for various downstream tasks. The data for the mean ribosome load (MRL) task have been deposited in the OMIX database (https://ngdc.cncb.ac.cn/omix), part of the Genome Sequence Archive (GSA) at the China National Center for Bioinformation(Chen et al., 2021; CNCB-NGDC Members and Partners, 2024), and are available under the accession number OMIX008723.
Code availability
The source code for UTR-DynaPro is available at https://github.com/shenhaoye8-dotcom/UTR-DynaPro. This repository includes the implementation of our Parallel Feature Fusion Module, k-mer specific MoE, and the full training pipeline to ensure reproducibility.
References
Araujo, P. R. et al. Before It Gets Started: Regulating Translation at the 5’ UTR. Comp. Funct. Genom. 2012, 475731 (2012).
Galloway, A. & Cowling, V. H. mRNA cap regulation in mammalian cell function and fate. Biochim. Biophys. Acta Gene Regul. Mech. 1862, 270–279 (2019).
Li, X. et al. Finding the target sites of RNA-binding proteins. Wiley Interdiscip. Rev. RNA 5, 111–130 (2014).
Miao, Z. et al. Secondary structure of the SARS-CoV-2 5’-UTR. RNA Biol. 18, 447–456 (2021).
Zeraati, M. et al. Cancer-associated noncoding mutations affect RNA G-quadruplex-mediated regulation of gene expression. Sci. Rep. 7, 708 (2017).
Li, G. et al. Predicting functional UTR variants by integrating region-specific features. Brief. Bioinform. 25, bbae248 (2024).
Schwanhäusser, B. et al. Corrigendum: Global quantification of mammalian gene expression control. Nature 495, 126–127 (2013).
Zheng, W. et al. Discovery of regulatory motifs in 5’ untranslated regions using interpretable multi-task learning models. Cell Syst. 14, 1103-1112.e6 (2023).
Dvir, S. et al. Deciphering the rules by which 5’-UTR sequences affect protein expression in yeast. Proc. Natl. Acad. Sci. U. S. A. 110, E2792-2801 (2013).
Pan, X. et al. 5’-UTR SNP of FGF13 causes translational defect and intellectual disability. Elife 10, e63021 (2021).
Schlusser, N. et al. Current limitations in predicting mRNA translation with deep learning models. Genome Biol. 25, 227 (2024).
Godefroy-Colburn, T. et al. Cap accessibility correlates with the initiation efficiency of alfalfa mosaic virus RNAs. Eur. J. Biochem. 147, 549–552 (1985).
Shah, P. et al. Rate-limiting steps in yeast protein translation. Cell 153, 1589–1601 (2013).
Bentele, K. et al. Efficient translation initiation dictates codon usage at gene start. Mol. Syst. Biol. 9, 675 (2013).
Ding, Y. et al. Weak 5’-mRNA secondary structures in short eukaryotic genes. Genome Biol. Evol. 4, 1046–1053 (2012).
Kertesz, M. et al. Genome-wide measurement of RNA secondary structure in yeast. Nature 467, 103–107 (2010).
Tuller, T. et al. An evolutionarily conserved mechanism for controlling the efficiency of protein translation. Cell 141, 344–354 (2010).
Weinberg, D. E. et al. Improved ribosome-footprint and mRNA measurements provide insights into dynamics and regulation of yeast translation. Cell Rep. 14, 1787–1799 (2016).
Andersson, S. G. & Kurland, C. G. Codon preferences in free-living microorganisms. Microbiol. Rev. 54, 198–210 (1990).
Chu, D. & von der Haar, T. The architecture of eukaryotic translation. Nucleic Acids Res. 40, 10098–10106 (2012).
La Fleur, A. et al. Decoding biology with massively parallel reporter assays and machine learning. Genes Dev. 38, 843–865 (2024).
Cao, J. et al. High-throughput 5’ UTR engineering for enhanced protein production in non-viral gene therapies. Nat. Commun. 12, 4138 (2021).
Ingolia, N. T. et al. Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324, 218–223 (2009).
England, W. E. et al. An atlas of posttranslational modifications on RNA binding proteins. Nucl. Acids Res. 50, 4329–4339 (2022).
Jin, B. T. et al. Mian: Interactive web-based microbiome data table visualization and machine learning platform. Bioinformatics 38, 1176–1178 (2022).
Li,G. et al. Predicting functional UTR variants by integrating region-specific features. Brief. Bioinform., 25, bbae248. (2024b).
Castillo-Hair, S. et al. Optimizing 5’UTRs for mRNA-delivered gene editing using deep learning. Nat. Commun. 15, 5284 (2024).
Pan, S. et al. UTR-Insight: Integrating deep learning for efficient 5’ UTR discovery and design. BMC Genom. 26, 107 (2025).
Wang, D. et al. DM3Loc: Multi-label mRNA subcellular localization prediction and analysis based on multi-head self-attention mechanism. Nucleic Acids Res. 49, e46 (2021).
Alipanahi, B. et al. Predicting the sequence specificities of DNA- and RNA-binding proteins by deep learning. Nat. Biotechnol. 33, 831–838 (2015).
Hill, S. T. et al. A deep recurrent neural network discovers complex biological rules to decipher RNA protein-coding potential. Nucleic Acids Res. 46, 8105–8113 (2018).
Chu, Y. et al. A 5′ UTR language model for decoding untranslated regions of mRNA and function predictions. Nat. Mach. Intell. 6, 449–460 (2024).
Sample, P. J. et al. Human 5′ UTR design and variant effect prediction from a massively parallel translation assay. Nat. Biotechnol. 37, 803–809 (2019).
He, S. et al. Nucleic transformer: Classifying DNA sequences with self-attention and convolutions. ACS Synth. Biol. 12, 3205–3214 (2023).
Chen, J. et al. QuoteTarget: A sequence-based transformer protein language model to identify potentially druggable protein targets. Protein Sci. Publ. Protein Soc. 32, e4555 (2023).
Na,O. et al. Reference sequence (RefSeq) database at NCBI: current status, taxonomic expansion, and functional annotation. Nucleic Acids Res., 44. (2016)
Karollus, A. et al. Predicting mean ribosome load for 5’UTR of any length using deep learning. PLoS Comput. Biol. 17, e1008982 (2021).
Akiyama, M. & Sakakibara, Y. Informative RNA base embedding for RNA structural alignment and clustering by deep representation learning. NAR Genom. Bioinform. 4, lqac012 (2022).
Chen,J. et al. Interpretable RNA Foundation Model from Unannotated Data for Highly Accurate RNA Structure and Function Predictions. (2022).
Korbel, F. et al. Interpreting deep neural networks for the prediction of translation rates. BMC Genomics 25, 1061 (2024).
Chen, T. et al. The Genome Sequence Archive Family: Toward explosive data growth and diverse data types. Genom. Proteom. Bioinform. 19, 578–583 (2021).
CNCB-NGDC Members and Partners. Database resources of the National Genomics Data Center, China National Center for Bioinformation in 2024. Nucleic Acids Res. 52, D18–D32 (2024).
Acknowledgements
We sincerely thank the developers of UTR-LM and UTR-Insight for making their code publicly available. Their open-source contributions have greatly facilitated our research and reproducibility effort
Author information
Authors and Affiliations
Contributions
S: Conceptualization, Methodology, Software Programming, Visualization, Writing- Original Draft, Writing-Reviewing and Editing draft preparation; L: Conceptualization, Methodology, formal analysis application of statistical; G: Conceptualization, Methodology, Writing-Reviewing and Editing draft preparation; Z: Data Curation, Software Assistance, Figure Preparation, Result Visualization; M: Conceptualization, Methodology, Data Curation; C: Methodology, Writing-Reviewing and Editing draft juntaochen066@gmail.compreparation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Shen, H., Liu, S., Guo, F. et al. UTR-DynaPro: a CNN–transformer multimodal language model for decoding 5′UTR regulatory mechanisms. Sci Rep 16, 10779 (2026). https://doi.org/10.1038/s41598-026-42175-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-42175-x