Abstract
Inflammation plays a crucial role in many chronic diseases with IL-6 and COX-2 being key mediators. The current article explored the anti-inflammatory potential of Convolvulus oxyphyllus extracts, combining phytochemical profiling, molecular docking, and gene expression analysis to elucidate their mechanisms of action. Total alcohol extract and extract-mediated Se-NPs of C. oxyphyllus were evaluated for their ability to modulate IL6 and COX2 expression using quantitative real-time PCR (qRT-PCR). LC–MS investigation was implemented to identify major phytoconstituents, followed by molecular docking to assess the binding interactions of selected most abundant flavonoid compounds (quercetin, daidzein-8-C-glucoside, kaempferol-7-neohesperidoside, and eriodictyol-7-O-neohesperidoside) with COX-2 (PDB ID: 8ET0) and IL-6 (PDB ID: 4N19). Both extract-mediated Se-NPs and total alcohol extract significantly down-regulated IL6 and COX2 expression (69% and 73%, respectively), comparable to Celecoxib. LC–MS profiling revealed the presence of flavonoids as dominant metabolites, alkaloids, coumarins and other constituents. Molecular docking demonstrated strong ligand–target interactions, with eriodictyol-7-O-neohesperidoside showing the highest binding affinity for COX-2 (− 10.2 kcal/mol) and kaempferol-7-neohesperidoside for IL-6 (− 7.4 kcal/mol), surpassing or matching the standard drug Celecoxib. These outcomes emphasize the therapeutic potential of C. oxyphyllus and its constituent phytochemicals as natural anti-inflammatory agents warranting further in vivo validation.
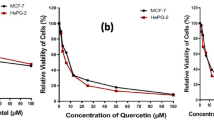
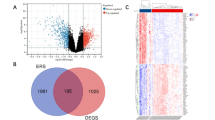
Similar content being viewed by others
Introduction
Medicinal plants constitute a major reservoir of bioactive compounds and continue to serve as a cornerstone for the discovery of novel therapeutic agents. Extensive phytochemical and pharmacological investigations have been conducted to evaluate their safety, efficacy, and mechanistic potential. Numerous plant families exhibit broad-spectrum biological activities, including anti-inflammatory, antidiabetic, antitumor, antiviral, and antimicrobial effects.
The genus Convolvulus is particularly notable for its rich phytochemical composition, including flavonoids, alkaloids, and phenolic compounds1. These constituents are widely recognized for their diverse pharmacological properties, especially their ability to modulate inflammatory signaling pathways2. Moreover, species within this genus have demonstrated significant antioxidant, antiulcer, and anti-inflammatory activities3 and have been traditionally employed in the management of respiratory conditions such as asthma and cough4.
Inflammation is a fundamental defense mechanism essential for host protection and tissue repair; however, when persistent or dysregulated, it contributes to the pathogenesis of numerous chronic diseases, including cardiovascular disorders, neurodegenerative conditions, and cancer5. Central inflammatory mediators, such as cyclooxygenase-2 (COX-2) and interleukin-6 (IL-6), play pivotal roles in propagating inflammatory cascades, and their sustained overexpression is closely associated with tissue damage and chronic pathological states6,7. Chronic low-grade inflammation is also recognized as a key contributor to cardiometabolic risk and age-related functional decline8,9.
Importantly, inflammation represents a modifiable biological target. Exercise-based interventions have been shown to attenuate inflammatory burden and improve health outcomes; for example, intermittent walking has been reported to improve cardiometabolic risk factors in postmenopausal women10, while resistance training enhances muscle strength and functional performance in older adults11. Collectively, these findings underscore the therapeutic relevance of targeting inflammatory pathways to promote cardiometabolic and functional health.
Tissue injury is a well-established trigger of inflammatory responses. Epidemiological studies in police cadets have demonstrated a high incidence of training-related musculoskeletal injuries12. Furthermore, mechanical overload and inadequate recovery may exacerbate tissue damage and prolong inflammatory responses13, emphasizing tissue injury as a modifiable contributor to overall inflammatory burden.
Nonsteroidal anti-inflammatory drugs (NSAIDs) and selective COX-2 inhibitors, such as celecoxib, exert their anti-inflammatory effects primarily by inhibiting prostaglandin synthesis and modulating cytokine signaling pathways. However, their long-term use is associated with significant adverse effects, including gastrointestinal, cardiovascular, and renal toxicity14. These limitations have stimulated increasing interest in the identification and development of natural anti-inflammatory agents, which may serve as promising alternatives or complementary therapies with potentially reduced side effects15.
Advances in analytical and computational methodologies now enable comprehensive characterization of plant-derived bioactive compounds. Liquid chromatography–mass spectrometry (LC–MS) is a powerful analytical technique for the precise identification and profiling of phytochemical constituents in complex plant extracts16. In parallel, molecular docking has emerged as a valuable computational approach that complements experimental analyses by predicting ligand–protein interactions. Specifically, it allows for the assessment of how phytoconstituents may bind to and potentially modulate key inflammatory targets such as COX-2 and IL-6, thereby providing mechanistic insight17,18.
Among plant-derived compounds, flavonoids are particularly noteworthy due to their broad spectrum of biological activities, including antioxidant, anti-inflammatory, and analgesic effects19,20,21. Extensive reviews have highlighted the capacity of flavonoids, such as quercetin, to regulate multiple inflammatory signaling pathways and modulate immune responses22.
Convolvulus oxyphyllus (Convolvulaceae) is a relatively understudied medicinal plant traditionally used for its analgesic, anti-inflammatory, and antioxidant properties. Despite its ethnopharmacological relevance, the molecular mechanisms underlying its anti-inflammatory effects remain largely unexplored, representing a clear research gap. Moreover, the bioactive constituents of C. oxyphyllus may exhibit limited stability and bioavailability, potentially restricting their therapeutic efficacy.
To address these limitations, selenium extract-mediated selenium nanoparticles (Se-NPs) were employed as a nanoformulation strategy to enhance bioavailability and potentiate the activity of plant-derived compounds. Se-NPs are known to exert antioxidant and immunomodulatory effects and to modulate key inflammatory mediators, including interleukin-6 (IL-6) and cyclooxygenase-2 (COX-2). Compared with bulk selenium, Se-NPs demonstrate improved biocompatibility and therapeutic performance. Thus, combining C. oxyphyllus extract with Se-NPs represents a novel, mechanism-oriented approach to amplify anti-inflammatory activity while targeting central inflammatory pathways23,24,25.
In addition, molecular docking serves as a valuable in silico tool for predicting interactions between phytoconstituents and inflammatory targets, thereby providing mechanistic insight prior to extensive experimental validation. The integration of nanoformulation with computational modeling supports the central hypothesis that Se-NPs conjugated with C. oxyphyllus extract may offer an enhanced, mechanism-driven anti-inflammatory intervention.
Accordingly, this study investigated the anti-inflammatory potential of total alcoholic extract and extract-mediated Se-NPs of C. oxyphyllus using an integrated experimental–computational framework. Gene expression analysis was performed to assess the modulation of IL6 and COX2, while LC–MS profiling enabled identification of major phytoconstituents potentially responsible for these effects. Molecular docking simulations were subsequently conducted to evaluate the binding affinities and interaction modes of selected compounds with COX-2 and IL-6. This comprehensive approach aimed to elucidate the molecular basis of the anti-inflammatory activity of C. oxyphyllus, bridging phytochemical characterization with mechanistic and computational evidence.
Materials and methods
Plant material
The plant’s aerial parts were gathered in April 2024 from South Sinai (Egypt), plant sample was identified and authenticated by Dr. Ahmad el khouly at DRC Herbarium, Desert Research Center. And a comparison to flora of Egypt plant description (Boulos 2000)26. Voucher a specimen of the plant was deposited in the herbarium of Desert Research Center with the code number (DRC 20/850). Plant collection and experimental protocol were achieved after permission from the “Desert Research Center, Cairo, Egypt”. Experimental research and field studies on wild plant, including the collection of plant material, comply with relevant institutional, national, and international guidelines and legislation. The plant material was air-dried in a shaded area, and ground into powder.
Extraction
In 3 L conical flask, 350 gm from last prepared plant powder was macerated in 1.5 L of 70% ethanol for two days at room temperature. Under the same circumstances, this extraction process was carried out three times. The resulting ethanol extracts were combined, fitered through filter paper Whatman No. 1, then evaporated at 40 °C under decreased pressure in a rotavapor. After thorough drying, 40 g of solid extract was obtained from the crude alcohol extract.
LC/Mass/Mass analysis of the alcohol extract
Instrument
The analysis of the sample was carried out by using LC-ESI-MS/MS (liquid chromatography–electrospray ionization–tandem mass spectrometry) with an ExionLC AC system (USA) for isolation and SCIEX Triple Quad 5500 + Mass/Mass system (Singapore) for detection it was connected with ESI (electrospray ionization).For identification of compounds Using Mass-Dial version 4.70 with ResPect library. The detected compounds identifications were assigned using MS-DIAL (Mass Spectrometry Data Independent AnaLysis), version 4.70 (available at: https://systemsomicslab.github.io/compms/msdial/main.html) open-source software in combination with ReSpect positive (2737 records) or ReSpect negative (1573 records) databases were used as the reference database.27.
Negative ionization mode
The isolation was carried out with a Ascentis® Express 90 Å Column is C18 (2.1 × 150 mm, 2.7 μm). The contents of mobile phases were two eluents A: 5 mM ammonium formate pH 8; B: acetonitrile (LC grade). From 0 to 1 min the mobile phase started at 5% B, from 1 to 20 min increased from 5 to 100% B, from 20 to 25 min was held at 100% B, at 25.01 back to 5%, at 25.01–30 min mobile phase stays at 5% from. The rate of flow was 0.3 ml/min and the volume of injection was 5 µl.
Analyses were performed in negative electrospray ionization (ESI) mode using an EMS-IDA-EPI workflow. For MS¹, the scan range was set from 100 to 1000 Da with the following source parameters: curtain gas, 25 psi; IonSpray voltage, − 4500 V; source temperature, 500 °C; ion source gases 1 and 2, 45 psi. For MS², the scan range was 50 to 1000 Da with a declustering potential of − 80 V and a collision energy of − 35 eV.
Mass/Mass analyses for positive ionization mode were performed using a scan (EMS-IDA-EPI) from 100 to 1000 Da for MS1 with the following source parameters: curtain gas: 25 psi; IonSpray voltage: 5500; source temperature: 500 °C; ion source gases 1 and 2 were 45 psi and scan range was 50 to 1000 Da for MS2 with a declustering potential: 80 V and a collision energy: 35 eV.
Green synthesis of selenium extract-mediated Se-NPs (Se-NPs) using Convolvulus oxphylls extract
At room temperature, 0.2 g of C. oxyphyllus crude alcohol extract was dissolved in 20 milliliters of 70% methanol using a magnetic stirrer. Following full dissolution, 100 ml of the final volume was diluted with distilled water to reach a 2000 ppm extract concentration. In order to create Se-NPs from C. oxyphyllus aqueous extract28, procedures were modified to optimize the pH, temperature, incubation duration, and precursor concentration. 0.2 g of sodium selenite (Na₂SeO₃) was dissolved in 50 mL of room temperature deionized water and then added to 10 ml of C. oxyphyllus extract (filtrate) to create an environmentally friendly synthesis of Se-NPs. To create in-situ Se-NPs at pH 6.2, the mixture was heated to 65 °C for 1 h while being constantly agitated. Following the addition of 200 µL of 40 mM ascorbic acid a secondary reducing agent/catalyst to improve reaction yield and stabilize the particles, while the C. oxyphylls extract served as the primary green capping and stabilizing agent, ruby-red Se-NPs were produced, as previously reported29,30. Five milliliters of 1% NaOH were added to the solution dropwise so as to bring the pH down to a basic environment at pH 10. Ultimately, the reaction mixtures were supplemented with deionized water to achieve a 2000 ppm concentration. To improve nanoparticle dispersion and avoid agglomeration, the mixture was ultrasonically treated (sonicated) for 30 min at 35 °C after being incubated at room temperature for the entire night. Centrifugation was used to gather the resultant Se-NPs, which were then routinely cleaned with distilled water and kept at 4 °C until used.
Characterization of Se-NPs
Transmission electron microscopy (TEM)
The morphology, size, and distribution of synthesized Se-NPs were examined using a JEOL JEM-2100 F transmission electron microscope (JEOL Ltd., Tokyo) at 200 kV. Samples were prepared by placing a drop of diluted Se-NP suspension onto carbon-coated copper grids (300 mesh) and air-dried. Images were recorded with a Gatan Orius SC1000 CCD camera (Gatan Inc., Pleasanton, CA). Particle size distribution was determined by measuring over 200 particles from multiple images using ImageJ (version IJ 1.54; U.S. National Institutes of Health, Bethesda, Maryland, USA available at: https://imagej.net/ij/download.html).
Dynamic light scattering (DLS)
The hydrodynamic diameter, polydispersity index (PDI) of Se-NPs were determined using a Malvern Zetasizer Nano ZS (Malvern Panalytical, UK) with a 633 nm He-Ne laser at 25 °C and a 173° scattering angle. Deionized water was used to dilute the DLS samples before they were passed through 0.22 μm syringe filters. Every measurement was taken three times and averaged.
X-ray diffraction (XRD)
A Rigaku SmartLab X-ray diffractometer (Rigaku, Tokyo) using Cu Kα radiation (λ = 1.5406 Å) at 40 kV and 30 mA was used to examine the crystalline structure and phase of Se-NPs. With a 0.02° step size and a 2°/min scan speed, XRD patterns were captured between 10° and 80° 2θ. Concentrated Se-NP suspensions were drop-cast onto silicon zero-background holders to create the samples, which were then allowed to air dry. The Debye-Scherrer equation, D = 0.9λ/βcosθ, was used to determine the crystallite size.
Where D is the crystallite size (nm), λ is the X-ray wavelength (1.5406 Å), β is the full width at half maximum (FWHM) of the diffraction peak (in radians), and θ is the Bragg’s angle.130.
Anti-inflammatory activity
Effect on mRNA expression of inflammatory genes (IL6 and COX2) using one-step real-time quantitative RT-PCR
Cell treatment and sample preparation
RAW264.7 murine macrophage cells (Vacsera Cell Bank, Giza, Egypt) were used; cells were cultured under standard conditions (37 °C, 5% CO₂) in Dulbecco’s Modified Eagle Medium (DMEM) supplemented with 10% fetal bovine serum (FBS) and 1% penicillin–streptomycin. To establish an in vitro inflammatory model, cells were seeded into 6-well plates and allowed to adhere overnight. Inflammation was induced using lipopolysaccharide (LPS, Escherichia coli O111:B4) at a final concentration of 1 µg/mL for 24 h. Cells were pretreated with: C. oxyphyllus total alcohol extract, C. oxyphyllus nano-extract and Celecoxib (reference anti-inflammatory drug) for 1 h prior to LPS stimulation. Following pretreatment, LPS (1 µg/mL) was added without removing the test samples, and cells were incubated for 24 h. DMSO was used as vehicle control, with a final concentration maintained below 0.1% (v/v) in all treated groups. After 24 h incubation, cell lysates were collected for total RNA extraction.
RNA extraction
Total RNA was isolated from cell lysates applying the Qiagen RNeasy Mini Kit (Qiagen, Germany) following the manufacturer’s instructions. RNA purity and concentration were determined spectrophotometrically (A₂₆₀/A₂₈₀ ratio) using a NanoDrop™ spectrophotometer (Thermo Fisher Scientific). Only RNA samples with A₂₆₀/A₂₈₀ ratios between 1.8 and 2.0 were used for downstream analysis to ensure quality and purity.
Quantitative real-time PCR (qPCR)
Quantitative one-step real-time PCR was achieved using the iScript™ One-Step RT-PCR Kit with SYBR® Green Master Mix (Bio-Rad, USA) on a Rotor-Gene™ Real-Time PCR System (Qiagen, Germany). Thermal cycling conditions were as follows: reverse transcription at 50 °C for 10 min, initial denaturation at 95 °C for 5 min, followed by 40 cycles of denaturation at 95 °C for 10 s and annealing/extension at 60 °C for 30 s. A melt-curve analysis (65–95 °C) was performed at the end of each run to verify the specificity of amplification.
Gene | Forward Primer (5′→3′) | Reverse Primer (5′→3′) | Reference |
|---|---|---|---|
IL6 | GAGGATACCACTCCCAACAGACC | AAGTGCATCATCGTTGTTCATACA | NCBI RefSeq |
COX2 | TGAGCAACTATTCCAAACCAGC | GCACGTAGTCTTCGATCACTATC | NCBI RefSeq |
GAPDH | AGGTCGGTGTGAACGGATTTG | TGTAGACCATGTAGTTGAGGTCA | Housekeeping gene |
Data analysis
Cycle threshold (Ct) values were automatically calculated using Rotor-Gene 6000 Series Software, version 1.7 (Build 87) (Qiagen, Hilden, Germany; available at https://rotor-gene.software.informer.com/1.7/#google_vignette). Expression levels of target genes (IL6 and COX2) were normalized to the housekeeping gene (GAPDH) and expressed relative to the LPS-stimulated control group using the 2⁻ΔΔCt method, where:
PCR efficiency (E = 1.903) was determined using standard curves and incorporated into fold change calculations to improve accuracy.
Molecular docking
The three-D structures of the target receptor proteins (cycloxygenae-2 and interleukin-6) were downloaded from the Protein Data Bank (PDB) (https://www.rcsb.org/). Ligands (quercetin, daidzein-8-C-glucoside, kaempferol-7-neohesperidoside and Eeriodictyol-7-O-neohesperidoside) three-dimensional (3D) structures were downloaded from the PubChem database in SDF format and then optimized. Molecular docking was conducted using PyRx Virtual Screening Tool, version 0.9.8 (available at: https://pyrx.sourceforge.io/) with AutoDock Vina (version 1.2.3) as the docking engine to predict the binding interactions of Convolvulus oxyphyllus bioactive compounds with inflammatory targets such as COX-2 and IL-6. Protein structures were retrieved from the Protein Data Bank (PDB) and prepared by removing water molecules, co-crystallized ligands, and adding polar hydrogens. Ligand structures were energy-minimized using the Universal Force Field (UFF) and converted to pdbqt format31,32,33. Docking simulations were performed within a grid box encompassing the active site of each protein; the box dimensions and center coordinates were optimized to fully cover the binding pocket. The default scoring function of AutoDock Vina was used to estimate binding affinities (kcal/mol) and predict binding poses. Redocking of the co-crystallized ligands was performed to validate the docking protocol, ensuring that the predicted binding pose closely reproduced the experimentally observed conformation (RMSD < 2 Å). Key docking parameters included exhaustiveness = 8, number of modes = 10, and energy range = 3 kcal/mol. The resulting docking interactions and binding conformations was visualized and analysed using Discovery Studio client, version 2025 (BIOVIA, Dassault Systèmes; available at: https://www.3ds.com/products/biovia/discovery-studio/visualization). It should be emphasized that these results predict binding potential rather than confirm enzymatic inhibition, and further experimental validation is necessary to substantiate the anti-inflammatory activity.
Results and discussion
LC/Mass/Mass analysis of Convolvulus oxyphyllus alcohol extract
Negative and positive modes of ESI masses were used in the identification of main 54 compounds in Convolvulus oxyphyllus alcohol extract. The compounds included flavonoids (28), phenolic acids and their derivatives (7), coumarine derivatives (2), terpens (2), carboxylic acids and their derivatives (5), alkaloids (2), Purines and pyrimidine derivatives (4), Amino acids derivatives (1) and others (3).
Presented data revealed that flavonoid compounds were represented the major abundance. Results also revealed that most abundant flavonoids in Convolvulus oxyphyllus were Quercetin (9.92) followed by Kaempferol-7-neohesperidoside (9.89), Daidzein-8-C-glucoside (9.80) and Eriodictyol-7-O-neohesperidoside (8.4) (Table 1).
The detected compounds identifications were assigned using MS-DIAL version 4.70 open-source software in combination with ReSpect positive (2737 records) or ReSpect negative (1573 records) databases were used as the reference database.27.
Characterization of Rh-Se-NPs
According to TEM analysis in . 1, the green-synthesized selenium extract-mediated Se-NPs generated with aqueous extract of C. oxyphyllus were primarily spherical morphology. The consistent, black circular shapes of the extract-mediated Se-NPs on a lighter backdrop suggested that the selenium ions were effectively reduced and that stable particles were formed. Indicating effective surface capping by C. oxyphyllus molecules, minimal aggregation was seen. TEM images displaying the major series of particle size that were between 3.18 and 17 nm. The lack of asymmetrical shapes or sizable clusters bolsters the function of the hydroxyl and carbonyl groups in C. oxyphyllus molecules in maintaining the stability of the extract-mediated Se-NPs surface. DLS analysis in . 2 showed that Se-NPs had an intensity-weighted mean hydrodynamic diameter of 88.6 nm. The polydispersity index (PDI) was 0.09, indicating a highly monodisperse extract-mediated Se-NPs population with a unimodal size distribution and narrow peak. X-ray diffraction analysis (XRD), shown in . 3, confirmed the crystalline nature and phase purity of Se-NPs, showing distinct peaks at 2θ values of 23.51°, 29.70°,41.32o, 43.65o, 45.36o and 51.71°corresponding to the Miller indices assignments (-100), (-10-1), (-210), (-102), (-21-1), and (-20-1) confirms the successful formation of trigonal selenium (t-Se) with hexagonal crystal structure. The intense peak at 29.70° ((-10-1) plane) indicates a preferred crystallographic orientation during biosynthesis. Sharp, well-defined peaks verify high crystallinity. The excellent match between experimental peak positions and reference data (COD 9012501)34 validates the hexagonal crystal structure with space group P 3■ 2 1. The lattice parameters calculated from the peak positions (a = 4.366 Å, c = 4.954 Å) are in excellent agreement with literature values for trigonal selenium, demonstrating the structural integrity of the synthesized extract-mediated Se-NPs. the significant difference between the TEM core size (17 nm) and the DLS hydrodynamic size (88.6 nm). This discrepancy is likely due to the formation of a dense bio-organic capping layer from the plant extract and a degree of agglomeration in the aqueous dispersion medium, which DLS is highly sensitive.
TEM analysis.
DLS analysis.
X-ray diffraction analysis (XRD).
Anti-inflammatory activity
Effect on mRNA expression of inflammatory genes (IL6 and COX2) using one-step real-time quantitative RT-PCR
Quantitative real-time PCR analysis showed that treatment of RAW264.7 macrophages with C. oxyphyllus total alcohol extract, C. oxyphyllus extract-mediated Se-NPs, and celecoxib obviously downregulated the mRNA expression of the pro-inflammatory genes IL6 and COX2 compared with the LPS-stimulated control (Table 2). Using GAPDH as the internal reference housekeeping gene, relative quantification by the 2⁻ΔΔCt method revealed that IL6 expression was declined to 0.44 fold in C. oxyphyllus total alcohol extract-treated cells, 0.31 fold in C. oxyphyllus extract-mediated Se-NPs -treated cells, and 0.28 fold in celecoxib-treated cells. Furthermore, COX2 expression decreased to 0.36 fold, 0.27 fold, and 0.20 fold for C. oxyphyllus total alcohol extract, C. oxyphyllus extract-mediated Se-NPs and celecoxib-treated cells respectively, relative to LPS-stimulated control (set to 1.0).
In addition, results showed that C. oxyphyllus extract-mediated Se-NPs exhibited excellent suppression of both IL6 and COX2, indicating enhanced anti-inflammatory efficacy compared to C. oxyphyllus total alcohol extract and a response comparable to the standard anti-inflammatory drug celecoxib. These findings suggest that both the C. oxyphyllus total alcohol extract and its extract-mediated Se-NPs possess These findings suggest anti-inflammatory potential through transcriptional modulation of inflammatory mediators in macrophages (Fig. 4).
Data are normalized to GAPDH and expressed relative to the LPS-stimulated control using the 2⁻ΔΔCt method. PCR efficiency (E) = 1.903 was applied. Lower fold change (< 1) indicates downregulation relative to LPS-stimulated control.
In the present study, treatment of RAW264.7 macrophages with C. oxyphyllus total alcohol extract, C. oxyphyllus extract-mediated Se-NPs and celecoxib showed significant suppression of both IL6 and COX2 expression, key mediators of inflammation. These findings validate the anti-inflammatory capacity of the plant’s bioactive compounds at the transcriptional level. Similar findings have been reported for other medicinal plants, where phytochemicals could significantly inhibit both cytokine expression and prostaglandin biosynthesis through suppression of COX-2 and IL-6 pathways35,36. Likewise, both leaf and fruit extracts of Terminalia ferdinandiana showed potent inhibition of COX-2 protein levels in macrophages37.
Moreover, the extract-mediated Se-NPs extract achieved up to a 69% reduction in IL6 expression whereas the total alcohol extract reduced it by approximately 56%. These results suggested that nanoscale delivery enhances bioavailability and cellular uptake of active phytoconstituents an effect consistent with other nano phytotherapy studies38.
Moreover, nano-formulations designed to target inflammation have been shown to reprogram macrophage phenotype and reduce pro-inflammatory gene expression39. By using nano-carriers, the extract’s components are protected and can be delivered more effectively to target sites, leading to enhanced therapeutic effects, thus increasing the activity40,41.
Furthermore, the recorded more noticeable suppression of COX2 compared with IL6 (up to 80% reduction for Celecoxib) is consistent with the hierarchical role of COX-2 in prostaglandin-mediated inflammation downstream of IL-6 signaling7. Moreover, recent studied indicates that plant-derived polyphenols can modulate both NF-κB and AP-1 transcriptional activity, producing broad suppression of inflammatory mediators42.
Relative mRNA expression of IL6 and COX2 in LPS-stimulated RAW264.7 macrophages treated with Convolvulus oxyphyllus extracts and celecoxib. Expression levels were normalized to GAPDH and calculated using the efficiency-corrected 2⁻ΔΔCt method (E = 1.903).
Molecular docking of Convolvulus oxyphyllus constituents targeting cycloxygenae-2 and interleukin-6
Molecular docking is a valuable computational approach in natural product research, enabling prediction of ligand–protein interactions and estimation of binding affinity. The calculated binding energy reflects the predicted stability of the ligand–target complex, with more negative values generally indicating stronger interactions¹⁷˒¹⁸. However, docking results represent theoretical models based on static protein structures and should be interpreted cautiously.
Interleukin-6 (IL-6) is a multifunctional cytokine that mediates inflammation through protein–protein interactions with its receptor complex (IL-6R/gp130), activating downstream JAK/STAT signaling⁹. Because IL-6 is not a classical enzyme with a well-defined catalytic pocket, docking predictions may not directly translate into functional inhibition. While predicted binding poses can identify potential interaction sites and guide compound prioritization, they do not confirm suppression of IL-6 signaling. Therefore, experimental validation through cytokine quantification, receptor-binding assays, and cell-based functional studies remains essential⁹.
In order to examine the possible compound/enzyme-binding relationships, compounds quercetin, daidzein-8-C-glucoside, kaempferol-7-neohesperidoside and Eeriodictyol-7-O-neohesperidosid were docked into the active site of cycloxygenae-2 (PDB code 8ET0) (Fig. 5) and interleukin-6 (PDB code 4N19) (Fig. 6). Table 3 and Figs. 7, 8a–d, 9 and 10a–d emphasize 3D and 2D interactions representations of the docked poses. The energies showed by ligands for cycloxygenae-2 range from (-10.2 to -9.6 Kcal/mol); the highest binding affinity Eeriodictyol-7-O-neohesperidoside (-10.2 Kcal/mol) with interacting amino acid residues (CYS15, PRO122, CYS4, GLN430, VAL124, GLY93, GLN296, ALA125, SER17, MET16 and GLY104); followed by Quercetin (-9.9 Kcal/mol) with interacting amino acid residues (THR175, HIS355, HIS176, ASP101 and HIS357) then Daidzein-8-C-glucoside and kaempferol-7-neohesperidoside (-9.7 Kcal/mol and − 9.6 Kcal/mol, respectively). Concerning the energies for interleukin-6 (-7.4 to -6.9 Kcal/mol); the highest binding affinity kaempferol-7-neohesperidoside (-7.4 Kcal/mol) with interacting amino acid residues such as (MET67, GLU172, LYS171, GLN175, ARG179 and PHE74); then Eeriodictyol-7-O-neohesperidoside (-7.2 Kcal/mol) with interacting amino acid residues (LYS120, ARG113, GLU109, ARG113, GLU109 and GLN102) followed by both Quercetin and Daidzein-8-C-glucoside (-6.9 Kcal/mol). However, the standard Celecoxib binding affinity was (-8.7 Kcal/mol and − 6.8 Kcal/mol) for both enzymes, respectively; which lower than that exerted by the tested compounds.
The analysis of these results suggests that the compounds may have potential as anti-inflammatory agents, warranting further in vitro and in vivo studies to validate their activity and facilitate their future development. While the docking results provide insights into possible interactions with inflammatory targets, they predict binding potential rather than confirm actual enzymatic inhibition, and experimental validation (e.g., COX-2 inhibition assays and cytokine or cell-based assays) is required to substantiate these findings. In the context of this study, docking to IL-6 serves as a mechanistic hypothesis generator, helping to prioritize bioactive compounds from Convolvulus oxyphyllus for future experimental investigation, while emphasizing that conclusions regarding IL-6 inhibition must remain tentative until validated.
Cycloxygenae-2 (8ET0).
Interleukin-6 (4N19).
3D and 2D interactions of Celecoxib within cycloxygenae-2 active site.


3D and 2D interactions of the isolated compounds from Convolvulus oxyphyllus Boiss within cycloxygenae-2 active site. (a) Eeriodictyol-7-O-neohesperidoside, (b) Quercetin, (c) Daidzein-8-C-glucoside, and (d) Kaempferol-7-neohesperidoside.
3D and 2D interactions of Celecoxib within interleukin-6 active site.


3D and 2D interactions of the isolated compounds from Convolvulus oxyphyllus Boiss within interleukin-6 active site. (a) Eeriodictyol-7-O-neohesperidoside, (b) Quercetin, (c) Daidzein-8-C-glucoside, and (d) Kaempferol-7-neohesperidoside.
Conclusion
This study demonstrates that C. oxyphyllus extracts, particularly in selenium nanoparticle (Se-NP) form, effectively downregulate IL-6 and COX-2 transcription in LPS-stimulated RAW264.7 macrophages, highlighting their potential as anti-inflammatory agents. The nanoformulation likely enhances the stability, bioavailability, and cellular uptake of the plant’s bioactives, contributing to the observed efficacy. Molecular docking further supports these findings by predicting favorable interactions of key flavonoid glycosides—eriodictyol-7-O-neohesperidoside with COX-2 (− 10.2 kcal/mol) and kaempferol-7-neohesperidoside with IL-6 (− 7.4 kcal/mol)—with critical residues of the inflammatory targets.
The combined downregulation of IL-6 and COX-2, alongside these predicted ligand–target interactions, suggests that C. oxyphyllus extracts in nanoform may offer a natural alternative or adjunct to conventional NSAID therapy, potentially driven by their flavonoid content. However, as docking results are predictive, further experimental validation is essential. Future studies should include protein-level assays (ELISA or Western blot), dose–response and cytotoxicity evaluations, and in vivo assessments to fully establish their therapeutic potential and mechanism of action.
Data availability
The datasets used and/or analyzed during the current study available from the corresponding author (R.A.L.) on reasonable request.
References
Awaad, A. S., Al-Rifai, A. A., El-Meligy, R. M., Alafeefy, A. M. & Zain, M. E. New activities for isolated compounds from Convolvulus austro-aegyptiacus as anti-ulcerogenic, anti-Helicobacter pylori and their mimic synthesis using bio-guided fractionation. Phytother. Res. 29, (2015).
Abdelhafez, O. M. et al. Phytochemistry and pharmacological properties of Convolvulus species: An updated review. Arab. J. Chem. 16, 104519 (2023).
Sethiya, N. K., Trivedi, A., Patel, M. B. & Mishra, S. H. Comparative pharmacognostical investigation on four ethnobotanicals traditionally used as Shankhpushpi in India. J. Adv. Pharm. Technol. Res. 1, 388–395 (2010).
Donia, A. M., Alqasoumi, S. I., Awaad, A. S. & Craker, L. Antioxidant activity of Convolvulus hystrix Vahl and its chemical constituents. Pak J. Pharm. Sci. 24, 143–147 (2011).
Medzhitov, R. The spectrum of inflammatory responses. Science 374, 1070–1075 (2021).
Hunter, C. A. & Jones, S. A. IL-6 as a keystone cytokine in health and disease. Nat. Immunol. 16, 448–457 (2015).
Ricciotti, E. & FitzGerald, G. A. Prostaglandins and inflammation. Arterioscler. Thromb. Vasc Biol. 31, 986–1000 (2011).
Ridker, P. M., Rifai, N., Stampfer, M. J. & Hennekens, C. H. Plasma concentration of interleukin-6 and the risk of future myocardial infarction among apparently healthy men. Circulation 101, 1767–1772 (2000).
Tanaka, T., Narazaki, M. & Kishimoto, T. IL-6 in inflammation, immunity, and disease. Cold Spring Harb Perspect. Biol. 6, a016295 (2014).
Abassi, W. et al. Intermittent walking training improves thyroid function and cardiometabolic risk factors in postmenopausal women. Climacteric 2026 1–9 (2026).
Chaabene, H. et al. The effects of eccentric versus traditional resistance training on muscle strength, power, hypertrophy, and functional performance in older adults: A systematic review with multilevel meta-analysis of randomized controlled trials. Ageing Res. Rev. 102933 (2025).
Dhahbi, W., Ben Saad, H., Dergaa, I., Souaifi, M. & Chamari, K. Injury profiling in male police cadets during initial training phase: a retrospective cohort study. Am. J. Mens Health. 18, 15579883241304584 (2024).
Dhahbi, W. Advancing biomechanics: enhancing sports performance, mitigating injury risks, and optimizing athlete rehabilitation. Front. Sports Act. Living. 7, 1556024 (2025).
O’Connor, J. P., Larkin, P. J. & Jones, P. W. Safety and efficacy of COX-2 inhibitors: A critical review. Pharmacol. Ther. 237, 108110 (2022).
Yatoo, M. I. et al. Bioactive natural products and their role in anti-inflammatory drug discovery. Front. Pharmacol. 12, 720002 (2021).
Deng, J. et al. Recent advances in LC–MS-based metabolomics for natural product research. J. Pharm. Biomed. Anal. 191, 113604 (2020).
Pinzi, L. & Rastelli, G. Molecular docking: Shifting paradigms in drug discovery. Int. J. Mol. Sci. 20, 4331 (2019).
Sadybekov, A. & Katritch, V. Computational approaches for structure-based drug discovery. Nat. Chem. Biol. 19, 1070–1080 (2023).
Ferraz, C. R. et al. Therapeutic potential of flavonoids in pain and inflammation: Mechanisms of action, pre-clinical and clinical data, and pharmaceutical development. Molecules 25, 762 (2020).
Kim, H. P., Son, K. H., Chang, H. W. & Kang, S. S. Antiinflammatory plant flavonoids and cellular action mechanisms. J. Pharmacol. Sci. 96, 229–245 (2004).
Scalbert, A., Johnson, I. T., Saltmarsh, M. & Polyphenols Antioxidants and beyond. Am. J. Clin. Nutr. 81, 215S–217S (2005).
Nair, H. K. et al. Inhibition of prostate cancer cell colony formation by the flavonoid quercetin correlates with modulation of specific regulatory genes. Clin. Diagn. Lab. Immunol. 11, 63–69 (2004).
Song, X. et al. Preparation, characterization, and in vivo evaluation of anti-inflammatory activities of selenium extract-mediated Se-NPs synthesized by Kluyveromyces lactis GG799. Food Funct. 12, 6403–6415 (2021).
Al-Anbar, Z. A., Mohammed, A. M. & Khalaf, Y. H. Selenium extract-mediated Se-NPs: a review of plant-mediated synthesis and their biomedical applications. J. Kufa Chem. Sci. 7, 16933 (2025).
Zhang, J. et al. Selenium extract-mediated Se-NPs attenuate inflammation and oxidative stress via NF-κB pathway modulation. Molecules 28, 8125 (2023).
Boulos, L. Flora of Egypt II (AlHadara Publishing, 2000).
Tsugawa, H. et al. MS-DIAL: Data-independent MS/MS deconvolution for comprehensive metabolome analysis. Nat. Methods 12, 523–526 (2015).
Wen, S., Hui, Y. & Chuang, W. Biosynthesis and antioxidation of nano-selenium using lemon juice as a reducing agent. Green. Process. Synth. 10, 178–188 (2021).
Boroumand, S. et al. Selenium extract-mediated Se-NPs: Synthesis, characterization and study of their cytotoxicity, antioxidant and antibacterial activity. Mater. Res. Express. 6, 0850d8 (2019).
Mourad, A. N. et al. Green synthesis of selenium extract-mediated Se-NPs, and its conjugation with antibacterial proteins for enhancing their antibacterial activity. Bioorg. Chem. 160, 108443 (2025).
Trott, O., Olson, A. J., AutoDock & Vina Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31, 455–461 (2010).
Dallakyan, S. & Olson, A. J. Small-molecule library screening by docking with PyRx. In Hempel, J.E., Williams, C. & Hong, C. (Eds.) Chem. Biol. Methods Protoc. Methods Mol. Biol. 1263, 243–250. https://doi.org/10.1007/978-1-4939-2269-7_19 (2015).
Eberhardt, J., Santos-Martins, D., Tillack, A. F. & Forli, S. AutoDock Vina 1.2.0: New docking methods, expanded force field, and Python bindings. J. Chem. Inf. Model. 61, 3891–3898 (2021).
Crystallography Open Database (COD). Entry 9012501: Selenium (trigonal). http://www.crystallography.net/cod/9012501.html.
Alotaibi, B. S. et al. Tamarix aphylla extract attenuates inflammation by suppressing COX-2 and pro-inflammatory cytokines in vitro and in vivo. BMC Complement. Med. Ther. 24, 359 (2024).
Akhtar, S. et al. Anti-inflammatory and antioxidant potential of Terminalia ferdinandiana extracts in macrophage models. Inflammopharmacology 32, 1473–1486 (2024).
Cock, I. E. Terminalia ferdinandiana Exell. extracts reduce pro-inflammatory cytokine and PGE2 secretion, decrease COX-2 expression and down-regulate cytosolic NF-κB levels. Inflammopharmacology 32, 1839–1853 (2024).
Patel, D. et al. Nanoformulation strategies for enhancing anti-inflammatory efficacy of plant bioactives: A review. Front. Pharmacol. 14, 1156734 (2023).
Hu, D. et al. Inflammation-targeted nanomedicines alleviate oxidative stress and reprogram macrophage polarization for myocardial infarction treatment. Adv. Sci. (Weinh. 11, e2308910 (2024).
Yang, B., Dong, Y., Wang, F. & Zhang, Y. Nanoformulations to enhance the bioavailability and physiological functions of polyphenols. Molecules 25, 4613 (2020).
Filho, S. A. et al. Biosynthesis of extract-mediated Se-NPs using plant extracts and essential oils. Molecules 28, 3060 (2023).
Jung, W. K. et al. Bamboo extract suppresses NF-κB activation and IL-6 production in LPS-stimulated macrophages. J. Ethnopharmacol. 137, 361–367 (2011).
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB).
Author information
Authors and Affiliations
Contributions
M.D.A.E., N.H.M., R.M.E. and R.A.L.: contributed to Conceptualization, Methodology, Investigation, analysis—discussing the data and Writing—reviewing the original article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
El-Halim, M.D.A., Mohamed, N.H., El-Meligy, R.M. et al. Biogenic selenium extract-mediated and total Convolvulus oxyphyllus extracts suppress IL6 and COX2 expression: insights from LC–MS metabolite profiling and molecular docking. Sci Rep 16, 14967 (2026). https://doi.org/10.1038/s41598-026-49047-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-49047-4