Abstract
Tumor ecosystem shapes cancer biology and potentially influence the response to immunotherapy, but there is a lack of direct clinical evidence. In this study, we utilized EcoTyper and publicly available scRNA-Seq cohorts from ICI-treated patients. We found a ecosystem subtype (ecotype) was linked to improved responses to immunotherapy. Then, a novel immunotherapy-responsive ecotype signature (IRE.Sig) was established and validated through the analysis of pan-cancer data. Utilizing IRE.Sig, machine learning models successfully predicted ICI responses in both validation and testing cohorts, achieving area under the curve (AUC) values of 0.72 and 0.71, respectively. Furthermore, using 5 CRISPR screening cohorts, we identified several potential drugs that may augment the efficacy of ICI. We also elucidated the candidate cellular biomarkers of response to the combined treatment of pembrolizumab plus eribulin in breast cancer. This signature has the potential to serve as a valuable tool for patients in selecting appropriate immunotherapy treatments.
Similar content being viewed by others
Introduction
Tumor is gradually recognized as an “ecosystem” consisting of a complex community of interacting cell types1. The powerful signaling networks formed by different cell types in the ecosystem affect tumorigenesis, cancer prognosis, and treatment response2,3.Targeted strategies that manipulate these networks are changing cancer treatment, such as the successful application of immune checkpoint inhibitor (ICI) based immunotherapy across a variety of cancer types4. Nevertheless, current response rate for ICI is highly heterogeneous, with only a relatively small proportion of patients obtaining a durable response5. Gaining a thorough comprehension of tumor related cell states, their interactive patterns, and their influence on clinical outcomes offers new opportunities for optimizing patient selection and combination strategies.
To improve cancer treatment and discover new therapeutic target, multiple broad phenotypic classes have been developed in human tumors. For example, according to the T cell infiltration, tumors are roughly classified into “hot” and “cold” tumors, while “hot” tumors likely respond better to immunotherapy than “cold” tumors6. Additionally, macrophages are divided into M1 and M2 macrophage based on their inflammatory activity, and the abundance of M2 phenotype corresponds to a poorer prognosis in most solid tumors7. Such classifications can contribute to cancer treatment strategies, including response to ICI. However, they lacked the comprehensive description of the tumor ecosystem, resulting in oversimplifying the cell states and complex interaction of different cell types.
Recently, the application of single-cell RNA sequencing (scRNA-Seq) has significantly transformed tumor research, as it uncovers the tumor heterogeneity and identifying rare cell subpopulations8. It offers profound understanding of tumor gene expression patterns, cell-cell interactions, and signaling pathways in tumors9. However, scRNA-Seq encounters several challenges, including technical noise, sample preparation complexity, and limited sample availability. Furthermore, in practice, scRNA-Seq were largely limited to single tumor types, medium sized sample cohorts.
Here, we utilized EcoTyper, a powerful machine learning framework developed by Luca et al.10, to discover and validate 14 cell states and 3 ecosystem subtypes (ecotypes) conserved in 5 scRNA-Seq ICI cohorts. During this process, we corroborated an ecotype associated with a better immunotherapy response. Afterward, a immunotherapy-responsive ecotype signature (IRE.Sig) was developed based on the comprehensive analysis of 34 scRNA-Seq cohorts. To further investigate and validate the predictive value of IRE.Sig, an integrated analysis was conducted on pan-cancer bulk-Seq cohorts (11001 patients; 32 cancer types), 12 independent ICI cohorts (921 patients; 5 cancer types) and 5 CRISPR screening cohorts(4 cancer types). We also explored the potential utility of IRE.Sig in treatment selection of specific cancer type. Together, our research revealed the potential of IRE.Sig in predicting immuotherapy outcomes, which is more accurate than previously identified pan-cancer signatures.
Methods
ICI scRNA-Seq cohorts
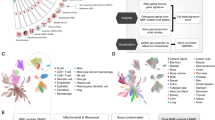
We downloaded 5 ICI scRNA-Seq cohorts from the TISCH2 portal (http://tisch.comp-genomics.org/)11. The standard scRNA-Seq workflow were already completed by TISCH2 portal including quality control, normalization, unsupervised clustering and cell type annotation. Detailed information of the 5 cohorts is shown in Fig. 1a. Ideally, it is better that all of the samples in ICI scRNA-Seq cohorts were responders (R) and non-responders (NR). However, the Jerby SKCM cohort had treatment naive patients which likely include both potential responders and non-responsers. IntegrateData function in Seurat R package was performed to integrate 2 discovery cohorts.
a Summary of ICI cohorts with scRNA-seq data. b UMAP of 5 ICI datasets profiled in this work by scRNA-seq. Cell state abundance patterns in the discovery (c) and validation (d) cohorts, with cell states organized into 3 ecotypes and tumor samples (columns) ordered by the most abundant ecotype per sample. NR non-responders, R responders, TN treatment naive patients.
Pan-cancer bulk-Seq datasets
The processed RNA-Seq data of 32 tumor types were downloaded from the USCS XENA portal (https://xena.ucsc.edu/) as TPM units. For those with processed data, these data were acquired directly for subsequent analysis. In total, 11,001 samples were included in analysis.
Pan-cancer scRNA-Seq datasets
To obtain immunotherapy-responsive ecotype signature (IRE.Sig), we collected 34 processed scRNA-Seq datasets and annotations from the TISCH2 portal (http://tisch.comp-genomics.org/). The process of quality control, normalization, unsupervised clustering and cell type annotation were already completed by TISCH2 portal.
ICI bulk-Seq cohorts
12 ICI bulk-Seq cohorts were acquired for validating the predictive value of IRE.Sig, including 6 SKCM cohorts (Van 201512, Abril-Rodriguez 202013, Puch 202114, Hugo 201615, Gide 201916, Auslander 201817), 2 urothelial carcinoma (UC) cohorts (Synder 201718, Mariathasan 201819), 1 renal cell carcinoma (RCC) cohort (Braun 202020), 1 non-small cell lung cancer (NSCLC) cohort (Wang 202221), 1 gastric cancer (GC) cohort (Kim 201822), 1 glioblastoma multiform (GBM) cohort (Zhao 201923). The details of each cohorts are summarized in Supplementary Table 5. Patients in these cohorts were categorized into two groups based on their response status. Specifically, complete response (CR) and partial response (PR) were designated as responders (R), or stable disease (SD) and progressive disease (PD) were labeled as non-responders (NR).
CRISPR screening data
A total of 5 CRISPR screens from publicly available datasets were used in this study24,25,26,27. The experimental design in these screens were shown in Fig. 5a which introduced immunotherapy into tumor/immune co-culture system. Details of these 5 CRISPR screens are given in Supplementary Table 7. We obtained these data from the supplementary information of corresponding publication. Several studies provided processed data, while other studies provided raw data. For those with processed data, these data were acquired directly for subsequent analysis; for those with raw data, MAGeCK pipeline (v0.5.9) was run with the default parameters to estimate the abundance of single guide RNAs (sgRNAs) targeting the corresponding genes. For those screens using mouse model, the R package biomaRt was utilized to map mouse genes to human genes. To quantify the effect of gene knockout on immunotherapy, we calculated the log-fold changes of sgRNAs counts between paired screens (Supplementary Table 8). Then z-score normalization was conducted for the log-fold changes to remove the batch effect among different CRISPR datasets. After that, genes were ranked by average z-score across 5 datasets (Fig. 5b). Ultimately, ICI enhancer genes were defined as those with log-fold change >0 and adj.P < 0.05 which represent the knockout of these genes were associated with ICI resistance. ICI suppressor genes were defined as those with log-fold change <0 and adj.P < 0.05 which represent the knockout of these genes were associated with ICI responsiveness.
Ecotype discovery
Ecotype discovery was performed with EcoTyper framework which is developed by Luca et al. to identify and characterize cell states and ecosystem subtypes from scRNA-Seq data or bulk RNA-Seq data10. EcoTyper implements a community detection algorithm that can discover robust collaborative network, termed ecosystem subtype or ecotype, in tissue samples. The discovery and recovery of cell states and ecotypes using EcoTyper framework need scRNA-Seq cohorts with the same cell types. Therefore, we choose the 5 cell types common to all 5 ICI scRNA-Seq cohorts, including B cells, CD4 T cells, CD8 T cells, macrophage, NK cells. Since the Sade et al. has largest sample number (n = 25) and the Caushi et al. Has highest cell number (n = 42,535), they were selected as discovery cohorts. To identify ecotype associated with better immunotherapy response, we picked out the immunotherapy responsive samples (n = 9) from discovery cohort. And then we applied EcoTyper scRNA-seq discovery framework with default parameters to identify cell states and ecotypes for these samples (n = 9). In total, we discovered 17 cell states while 3 of them were excluded from subsequent analysis due to not being assigned to any ecotypes. The remaining 14 cell states were assigned to 3 ecotypes (Supplementary Fig. 1).
Ecotype recovery
EcoTyper leverages the characteristics of Non-negative Matrix Factorization (NMF) to apply the discovered cell states and ecotypes to external cohorts10. The recovery of cell states and ecotypes in scRNA-Seq cohorts and bulk RNA-Seq cohorts adhered to the guidance provided in EcoTyper documentation (https://github.com/digitalcytometry/ecotyper), specifically the sections “Tutorial 2: Recovery of Cell States and Ecotypes in User-Provided scRNA-seq Data” and “Tutorial 1: Recovery of Cell States and Ecotypes in User-Provided Bulk Data”. After recovering the 3 ecotypes in TCGA pan-cancer samples, the patients of each cancer type were divided into 3 groups: ecotype1, ecotype2 and ecotype3 by EcoTyper.
Pathway analysis of the immunotherapy-responsive ecotype (ecotype2)
To investigate the enriched signaling pathways of the immunotherapy-responsive ecotype (ecotype2), we downloaded 27 functional gene expression signatures, which represent cellular phenotypes and signaling pathways, from a previous study28. The 27 signature scores across TCGA pan-cancer samples (n = 11,001) were computed by ssGSEA method using python29. tSNE projections was used to test intrinsic differences in 32 various cancers (Fig. 2a). To remove tissue-type-specific effects, we utilized median-score normalization for 27 signature scores of all samples. List of genes used in 27 functional gene expression signatures were shown in Supplementary Table 1.
a tSNE analysis of the samples across TCGA tumors. The dots represent individual tumor samples, and the colors represent the datasets (32 cancer types) from TCGA. b Bar graphs depicting recovery of the 3 ecotypes in TCGA datasets. c Enrichment analysis of 29 functional gene expression signatures across 11,001. TCGA carcinomas. d Enrichment analysis of several tumor associated pathways between tumor tissues with ecotype2 and other ecotype (ecotype1,3) across 32 cancer types in TCGA cohorts, NES normalized enrichment score in the GSEA algorithm, FDR false discovery rates.
Gene set variation analysis (GSVA) was leveraged to compute the enrichment of multiple tumor associated pathways in ecotype2. The R package GSVA v1.46.0 was used to conduct GSVA29. The gene sets used in GSVA were listed in Supplementary Table 2.
Gene ontology (GO) enrichment analysis was performed to identify whether known biological processes are enriched in IRE.Sig. The clusterProfiler v4.9.3R package was employed to perform this analysis and visualization30.
Gene Set Enrichment Analysis(GSEA) was applied to identify enriched biological function of B cell state6.
Construction of machine learning model for predicting ICI outcomes
To assess the predictive capability of IRE.Sig for response to ICI based immunotherapies, 12 ICI RNA-Seq cohorts were systemically collected. First, 7 ICI cohorts were integrated into a merged dataset (n = 741), including Abril-Rodriguez SKCM (n = 57), Kim GC (n = 45), Van SKCM (n = 36), Synder UC (n = 25), Mariathasan UC (n = 348), Puch SKCM (n = 49), Braun RCC (n = 181). The other five ICI RNA-Seq cohorts were consolidated as an independent testing set (n = 177). We employed ComBat algorithm to remove batch effects31. Subsequently, we randomly split the merged dataset into training (n = 593, 80%) and validation sets (n = 148, 20%). The genes in IRE.Sig was used to construct a prediction model in the training set. We integrated 10 machine learning algorithms: supervised principal components (SuperPC), random forest (RSF), gradient boosting machine (GBM), CoxBoost, elastic network (Enet), survival support vector machine (Survival-SVM), Stepwise Cox (StepCox), least absolute shrinkage and selection operator (LASSO), partial least squares regression for Cox (plsRcox) and ridge regression32. Among them, CoxBoost, Stepwise Cox, RSF and LASSO possess the capability of dimensionality reduction and variable screening. Consequently, they were combined with other algorithms, resulting in a total of 66 machine-learning algorithm combinations. To ensure the reliability of the algorithm combination, a rigorous validation procedure was followed, which involved performing 10 fold cross validation five times and utilizing validation set to compare their performance. Further, we employed testing set to evaluate the predictive ability. Utilizing the average C-index derived from validation set and testing set, we ultimately picked the best model.
Comparing IRE.Sig with other predictive gene signatures
To evaluate the predictive power of IRE.Sig, we compared IRE.Sig with 6 previously published ICI response signatures (TIS.Sig33, C-ECM-up.Sig34, GEP.Sig35, NLRP3.Sig36, PD − L1.Sig37, IFNG.Sig35). All the codes and algorithmsutilized for the 6 signatures were obtained from relevant published studies, such as average gene expression for TIS.Sig, ssGSEA for C-ECM-up.Sig and so on. Detailed information of these signatures is provided in Supplementary Table 6.
Potential drug search against the CMAP database
To identify promising drugs that enhance immunotherapies, we employed Connectivity Map (CMap) database (https://clue.io)38. The CMap database is a tool to match query signature with drug signatures in the CMap database. The search of query signature within CMap database was conducted as a “rapid search” in the querying segment. And in the search results, only drugs that have been approved or have successfully completed phase I and II clinical trials were selected.
Analysis of pembrolizumab plus eribulin
We downloaded gene expression and clinical data of the NCT03051659 trial from the supplementary information of the study39. First, we recovered our cell states and ecotypes in this bulk RNA-Seq dataset. Each sample consisted of various cell states with different proportions. Then, we examined the relationships among 17 cell states and overall survival time using Spearman correlation test. Subsequently, the marker genes of switched memory B cells were obtained from literature40,41. The scRNA-Seq discovery cohort of Yost et al. was used to present the expression of these marker genes among different B cell states. Finally, all samples in NCT03051659 trial were assigned to B cell state6 high and low groups according to the median value of the abundance of B cell state6.
We used independent validation cohort from Liu et al.42. In this trial, patients with breast cancer received Camrelizumab plus Eribulin therapy. Because pembrolizumab and camrelizumab are all immune checkpoint PD-1 inhibition, we used this cohort as independent validation cohort. Data acquisition and analysis are the same as above.
Spatial transcriptomics
For Fig. 6D, F, two human tumor sections with tertiary lymphoid structures (TLS) profiled by 10x Visium were analyzed43. We used TransferData function of Seurat R package to transfer scRNA-seq cell types in Fig. 1b to spatial scRNA-seq. Then, we applied EcoTyper to recover our cell states in the Visium array. Specifically, we set the highest abundant cell state in each spatially-barcoded spot to 1, while the rest to 0. To annotate tertiary lymphoid structures in the sections, the location information of TLS was obtained from original research43. Finally, to construct the plots in Fig. 6e, g, we computed the mean Euclidean distance for each B cell state spot from the spot to the nearest three spots of tertiary lymphoid structures.
Ethics approval
Since the sequenced data generated from TCGA and GEO were publicly available, additional ethics committee approval was not necessary.
Statistical analysis
All statistical analyses were performed using R v4.3.2 (https://www.r-project.org). The Kaplan–Meier curves and the log-rank test were utilized to assess the disparities in survival rates between the two groups. For correlation analysis, the Pearson correlation coefficient was conducted for data exhibiting a normal distribution, whereas the Spearman correlation coefficient was applied to data with a non-normal distribution. To analyze the differences between two groups of data, unpaired Student’s t test was used for normally distributed variables and the Mann–Whitney U-test was performed for non-normally distributed variables. For comparisons involving more than two groups, we utilized the one-way analysis of variance (ANOVA) as the parametric method and the Kruskal–Wallis’s test as the non-parametric method. The figure legends provide the statistical details, including the statistical test employed for each dataset. Unless otherwise mentioned, a p-value of 0.05 was deemed statistically significant.
Results
Identification of a ecotype associated with better response to immunotherapy
Tumor transcriptomic profiling at the single-cell level enabled a deeper understanding of the diversity of tumor ecosystem44. And EcoTyper framework which is developed by Luca et al. allowed us to identify tumor ecosystem subtypes (termed ecotypes)10. These basis has made it possible for us to discover a ecotype with better response to ICI based immunotherapies. So, we collected 5 ICI scRNA-Seq cohorts to identify the tumor ecosystem of patients responding to immunotherapy (Fig. 1a). Owing to the requirement of EcoTyper framework that the cell types of the discovery cohort and the validation cohorts must be the same. We selected 5 cell types common to all 5 ICI scRNA-Seq cohorts, including B cells, CD4 T cells, CD8 T cells, macrophage, NK cells (Fig. 1b). Due to the sample number of Sade et al. cohort was largest (n = 25), and the cell number of Caushi et al. cohort was highest (n = 42535), we integrated them into a discovery cohort. By applying EcoTyper to decode cell types and ecotypes in the responder patients of discovery cohort, 3 ecotypes were revealed and most of the responders corresponded to ecotype2 (Supplementary Fig. 1a). After we recovered these 3 ecotypes in all samples of discovery cohort, we were surprised to find that the majority of ecotype2 samples were responders(70%) (Fig. 1c). To corroborate these findings, 3 independent validation cohorts were employed (Fig. 1d). Our results from the discovery cohort were highly reproducible, both for cell states and ecotypes. Statistical analysis also showed patients in ecotype2 exhibited a higher proportion of responders than those in ecotype1 or ecotype2.
Analysis of the potential links between ecotype2 and anti-tumor immune infiltration using pan-cancer TCGA cohort
To explore whether the 3 ecotypes are conserved across a broad array of cancers, we recovered the 3 ecotypes in a total of 11,001 TCGA tumors across 32 cancer types. The result showed the 3 ecotypes were reliably recoverable within 32 various cancers (Fig. 2b). Although the 3 ecotypes were recoverable across multiple cancers, further evaluation was required to evaluate the significance of this ecosystem subtyping; hence, the enriched signaling pathways of the 3 ecotypes, especially ecotype2, were assessed at the pan-cancer level. We downloaded 27 functional gene expression signatures from a previously published study28. And ssGSEA method was applied to calculate the 27 signature scores across 11,001 TCGA tumors. Owing to substantial inherent differences in cellular composition, the signature scores resulted in distinct nosology-specific subsets (Fig. 2a). Following normalization, tissue-type-specific effects were removed (Fig. 2a). A active association was observed between ecotype2 and anti-tumor immune infiltration, including antitumor cytokines, NK cells, checkpoint inhibition (Fig. 2c). Then we applied GSEA to evaluate the enrichment of several tumor associated pathways in each cancer type. We compared the normalized enrichment score (NES) between ecotype2 and remaining ecotypes (ecotype1, ecotype3). The results indicated that anti-tumorigenic signaling pathways were enriched in samples classified into ecotype2, such as T cell activation, T cell chemotaxis and antigen procession and presentation (Fig. 2d); while the pro-tumorigenic signaling pathways were decreased in these samples, like hypoxia, tumor proliferation rate and epithelial to mesenchymal transition (Fig. 2d).
Development of IRE.Sig through pan-cancer scRNA analysis
Next, we aimed to develop a pan-cancer gene signature that precisely reflects the features of ecotype2, which was named immunotherapy-responsive ecotype signature (IRE.Sig). Hence, 34 scRNA-Seq datasets across various cancer types were applied to develop the IRE.Sig (Fig. 3a). We recovered our ecotypes in these 34 datasets using EcoTyper, cells were sorted into 3 ecotypes. Genes that were upregulated in responders from 5 ICI scRNA-Seq cohorts of Fig. 1 (LogFc ≥ 0.25 & FDR < 0.05) were regarded as ‘Gx’. Genes that were upregulated in ecotype2 of each scRNA-Seq dataset (logFC ≥ 0.25 and FDR < 0.05) were considered as ‘Gy’. To collect IRE.Sig genes, ‘Gx’ and ‘Gy’ were intersected to give rise to ‘Gn’ (n = 1–34) for each dataset. For instance, G1 encompassed genes that resulted from the overlap of Gx and Gy in the first scRNA-Seq dataset. Finally, genes from G1 to G34 were filtered into IRE.Sig (Supplementary Table 3). We further examined the biological functions that were enriched in IRE.Sig. The enriched pathways were largely involved in the following: T cell chemotaxis, B cell activation, leukocyte activation and T cell migration (Fig. 3b, Supplementary Table 4). The cnetplot depicted specific genes of these pathways (Fig. 3c). Several of the genes are reported to be correlated with the better effect of immunotherapies, such as PARP3, TNFRSF9, DKK2, BCL6, CCL5 and PVRIG45,46,47,48,49,50.
a The circus plot shows the generation process of IRE.Sig. b Pathway enrichment analysis of IRE.Sig genes. The top 20 enriched GO terms are shown in the bar plot. c The cnetplot shows specifc gene networks from IRE.Sig genes.
Prediction of immunotherapy response using IRE.Sig
With respect to the high correlation between ecotype2 and immuntherapy response rate in ICI scRNA-Seq cohorts, we further investigated the prediction value of IRE.Sig. 12 ICI bulk RNA-Seq cohorts containing outcomes of immunotherapies were used. The flow diagram presenting the analysis process is outlined in Fig. 4a. Among these 12 cohorts, we split them into 3 groups: training cohort (n = 593), validation cohort (n = 148), and testing cohort (n = 177). In the training cohort, 66 machine-learning algorithm combinations were leveraged to develop prediction models, utilizing five repetitions of tenfold cross-validation. Then, the average C-index of each algorithm combination in the validation cohort and testing cohort were computed. Ultimately, we chose the ‘CoxBoost and Survival-SVM’ model which delivered the highest average C-index (0.718) (Fig. 4b). Associated AUC plots of validation cohort (0.72) and testing cohort (0.71) were shown in Supplementary Fig. 2b. To assess the predictive capability of the IRE.Sig model for overall survival, we categorized the patients of validation and testing sets into low-risk and high-risk groups derived from the predicted “R” and “NR”. The Kaplan-Meier survival curve is shown in Fig. 2d. In both validation and testing sets, the low-risk subgroup exhibited a statistically significantly longer overall survival (Fig. 4d).
a Flow chart of training, validating, and testing the IRE.Sig model constructed using machine learning process in ICI cohorts. b The C-indexes of 66 machine-learning algorithm combinations in the validation and testing cohorts. c Circos plot shows the comparison between the performance of the IRE.Sig model and previously published pan-cancer models for response to immunotherapy on different testing sets. d Kaplan–Meier curves comparing OS between High-risk and Low-risk patients in validation and testing set. “NR” and “R” predicted by the final IRE.Sig Model was defined as “High-risk” and “Low-risk” patients respectively.
We additionally obtained 6 pan-cancer predictive gene signatures for immunotherapies response prediction to compare the predictive preciseness of IRE.Sig and these signatures, including TIS.Sig33, C-ECM-up.Sig34, GEP.Sig35, NLRP3.Sig36, PD − L1.Sig37, and IFNG.Sig35. The results showed IRE.Sig exhibited best performance in the testing cohort, achieving an AUC of 0.71 (Fig. 4c, Supplementary Fig. S2A). Moreover, we also conducted subgroup analysis on five distinct cohorts that contributed to the construction of the testing set. The majority of the pan-cancer signatures manifested excellent performance within one or two cohorts. However, IRE.Sig demonstrated consistent and ideal performance across all cohorts, highlighting its promising potential as a predictive signature for the pan-cancer response to immunotherapy (Fig. 4c, Supplementary Fig. 2a).
Exploration of potential agents synergistic with immunotherapies using CRISPR screening data
A significant hurdle encountered in cancer immunotherapy lies in augmenting the tumor’s responsiveness towards ICI therapy. Prior experimental and clinical investigations have revealed that combining multiple therapeutic strategies can notably elevate the proportion of responders and yield substantial survival advantages51,52. We first aimed to gather immunotherapy-related genes for retrieving drug. Data from 5 CRISPR screening cohorts were systemically collected24,25,26,27, which introducing ICI treatment into the experimental design as shown in Fig. 5a. Hence, these screens could discern prospective regulators that either facilitate resistance (known as ICI enhancer genes) or responsiveness (termed ICI suppressor genes) to immunotherapy upon gene knockout. Totally, 2704 genes were recorded across these CRISPR datasets. Subsequently, we ranked the genes based on their average z scores. Genes with higher z-score were ICI enhancer genes, which means knockout of these genes is associated with ICI resistance. Genes with lower z-score were ICI suppressor genes, which means knockout of these genes is associated with ICI responsiveness. The process of gene ranking was depicted in Fig. 5b. The adj.P < 0.05 was considered statistically different. Therefore, we found 86 ICI enhancer genes and 103 ICI suppressor genes in total. The full gene list was listed in Supplementary Table 9. Then, considering that the absence of a particular gene might impact cancer cell survival directly without the involvement of affecting ICI treatment, we employed Cancer Dependency Map (DepMap) portal (https://depmap.org/portal/) to prevent potential bias introduced by essential genes53. Next, we calculated the average CERES scores across 1101 cancer cell lines (Supplementary Table 10), and genes with CERES score ranging from −0.25 to 0.25 were identified as tumor proliferation-independent genes (n = 10616) (Fig. 5c).
a Diagram summarizing the workflow of CRISPR screens focused on identifying potential regulator genes mediating response to cancer immunotherapy. The definitions of enhancer genes and suppressor genes were also illustrated. b Ranking of genes based on their knockout effects on immunotherapy across 5 CRISPR datasets. Negative (positive) z scores indicated ICI responsiveness (resistanse) after knockout of a specifc gene. Genes were ranked according to their mean z scores. Blank squares in the heatmap referred to missing values of gene data from the corresponding cohort. c Determination of proliferation-independent genes according to the CERES scores from CRISPR knockout screens across hundreds of cancer cell lines. d Intersections among proliferation-independent genes, IRE.Sig genes, ICI enhancer and suppressor genes. e Result of drug prediction. The top 10 drugs were labeled in the plot.
Since IRE.sig can predict immunotherapy response, the genes in IRE.sig may be involved in the regulation of immunotherapy. Combining IRE.sig with above genes is more helpful in exploring potential agents synergistic with immunotherapies. Subsequently, IRE.Signature genes and ICI enhancer genes were integrated into a meta-gene list, which was named positive regulators. Theoretically, augmenting the function of these genes could be linked to an enhanced responsiveness towards ICI based immunotherapies. The list of negative regulators was collected from ICI suppressor genes. Ultimately, we intersected positive, negative regulators and tumor proliferation-independent genes together to constitute the query signature (Fig. 5d). To identify potential agents synergistic with immunotherapies, we searched CMap database by inputting the query signature. In Fig. 5e and Supplementary Table 11, we demonstrated the results of drug prediction. Notably, of the top 10 drugs identified, 4 candidates, namely decitabine, tolazamide, tamoxifen, and sulfasalazine, were reported through experimental or clinical researches previously, indicating the reliability of our prediction. Moreover, a randomized phase II study has shown decitabine-plus-immunotherapy results in longer progression-free survival54.
Prediction of therapeutic benefit with immunotherapy-responsive ecotype
Despite recent progress, predicting the response to immunotherapy remains challenging and further research is needed. Since EcoTyper is able to quantitate pre-defined cell states and ecotypes in external cohorts, we next examined our immunotherapy-responsive ecotype in identifying predictive biomarkers in the context of a randomized clinical trial. The NCT03051659 trial aimed to investigate whether the combination of pembrolizumab and eribulin therapy could enhance overall survival (OS) among patients with breast carcinoma (BRCA) compared to eribulin therapy alone39. While preclinical and clinical studies suggested that eribulin inhibits TGF-β and the PD-1/PD-L1 axis55,56, eribulin plus pembrolizumab failed to improve survival. In order to determine whether our immunotherapy-responsive ecotype can provide insights into the clinical benefit of combining pembrolizumab with eribulin, we recovered our 17 cell states in this clinical trial as depicted in the flow chart (Fig. 6a). We assessed sample-level correlation between the proportions of these 17 cell states and overall survival (OS) (Fig. 6b). An extremely high correlation between B cell state6 (B.cells_S06) and better OS was observed. Across scRNA-seq profiles of discovery cohort, B cell state6 expressed higher levels of CD27 and IGHG1 than other B cell states (Fig. 6c). CD27 and IGHG1 are known to mark switched memory B cells40, which were enriched in the tumors of immunotherapy responders and have powerful anti-tumor functions41,57. In alignment with this observation, known markers of switched memory B cells were consistent with the B cell state6 expression profile, but not other B cell states (Fig. 6c). GSEA analysis similarly revealed that some known function of switched memory B cell were enriched in B cell state6, like antibody dependent cellular cytotoxicity, isotype switching to IgG isotypes, antigen processing and presentation(Supplementary Fig. 3a). Furthermore, switched memory B cells might function anti-tumor ability through tertiary lymphoid structures (TLS)41,43. Using two human tumor sections with tertiary lymphoid structures (TLS), we found B cell state6 rich zones localized significantly closer to tertiary lymphoid structures (TLS) than did other B cell states(Fig. 6d–g). Overall, using transcriptomic and spatial analysis, we linked B cell state6 to a previously described switched memory B cell state.
a Outline of the approach. b Correlation among 17 cell states in 88 samples based on their relative population abundance. c Expression of positive and negative marker genes of switched memory B cells, shown for breast carcinoma-associated B cells profiled by scRNA-seq and mapped to EcoTyper states. Circle color represents the mean log2 expression of each gene, while size represents the fraction of cells with higher expression in the indicated state. Localization of TLS in a frozen (d) and an FFPE (f) tumor profiled by spatial transcriptomics (ST). e, g Relative distance of each B cell state from spots annotated as TLS in the ST array (see d, f). h Differences in OS for patients stratified by treatment arm and by groups with high or low levels of B cells S6 (median split).
To investigate the effect of B cell state6 as a predictive biomarker, patients in the trial were divided into B cell state6 high and low groups according to the median value of the abundance of B cell state6. Notably, patients with higher abundance of B cell state6 in the eribulin and pembrolizumab arm exhibited significantly longer survival time than other groups (Fig. 6h). In independent validation cohort, the similar result were observed (Supplementary Fig. 4a).
In summary, these results indicate switched memory B cells in eribulin plus pembrolizumab efficacy and highlight the promise of immunotherapy-responsive ecotype in predicting clinical outcomes within multicellular communities.
Discussion
While there has been significant researches conducted on the correlation between tumor ecosystems and anti-tumor immunity, the identification of specific cellular states and ecosystem subtypes that respond to ICI based immunotherapy remains rarely reported. Although a previous study demonstrated the cell states were related to immunotherapy response, it didn’t produce a signature to evaluate the specific probability of immunotheapy response10. Here, we assessed the transcriptional profiles of various cell types by deploying EcoTyper, categorized them into ecosystem subtypes, and identified a strong correlation between a specific ecosystem subtype and ICI response, as corroborated by five ICI scRNA-Seq cohorts. This immunotherapy-responsive ecotype was found to be conserved in at least 11001 tumor samples from 32 cancer types. Building upon these findings, we performed a large-scale comprehensive analysis to identify significantly overexpressed genes linked to the immunotherapy-responsive ecotype. These genes comprising a pan-cancer immunotherapy-responsive ecotype signature, referred to as IRE.Sig. We rigorously validated the predictive efficacy of IRE.Sig, revealing its superior performance in forecasting ICI responses compared to previously published signatures across multiple independent ICI datasets utilizing bulk RNA-Seq. This research represents the initial investigation to establish a robust correlation between distinct ecotype and ICI outcomes through an exhaustive analysis of large-scale data. Significantly, we developed a gene expression signature, IRE.Sig, that effectively predicted the immunotherapeutic response across various cancer types.
Our analysis revealed that the IRE.Sig genes were notably enriched in key biological functions such as T cell chemotaxis, B cell activation, white blood cell activation, and T cell migration. T cells play a pivotal role in anti-tumor immunity, as their chemotaxis directs them to the tumor site where they can identify and eliminate tumor cells58. Augmenting T cell chemotaxis helps to enhance anti-tumor efficacy59. The migration of T cells plays a crucial role in facilitating immune cells to execute their anti-tumor response function, with their infiltration into tumors being essential for an effective anti-tumor response60. Activation of B cells stimulates antibody production, neutralizes tumor-associated antigens, and induces direct cytotoxicity towards tumor cells through the secretion of granzyme B61. Previous study has also observed a notable augmentation in B cell density and activity among tumor patients exhibiting favorable responses to immunotherapy62.
TCGA pan-cancer transcriptome analysis revealed a significantly increase of anti-tumor immune infiltration in tumors with immunotherapy-responsive ecotype, including M1 macrophages, NK cells, and histocompatibility complex class 1 molecules (MHC-1). M1 macrophages play a crucial role in activating and enhancing the anti-tumor function of T cells through the secretion of inflammatory factors and antigen presentation7. NK cells contribute to the anti-tumor immune response by directly killing tumor cells and releasing cytokines63. Additionally, the MHC-1 molecule is implicated in presenting tumor antigens to T cells, triggering a specific immune response64. Tumors with immunotherapy-responsive ecotype exhibit high anti-tumor immune infiltration characteristics, confirming the predictive value of IRE.Sig. This conclusion applied to more than 18 cancer types, including liver hepatocellular carcinoma (LIHC), lung adenocarcinoma (LUAD), kidney renal clear cell carcinoma (KIRC), and colon adenocarcinoma (COAD) (Fig. 2d). Only in a few tumor types, like cholangiocarcinoma (CHOL), samples in ecotype2 group exhibited decreased anti-tumorigenic signaling pathways. This may be due to the small sample size.
IRE.Sig is a novel signature for predicting responses to immunotherapy and effectively stratifying patients who may benefit from improved survival outcomes. We further compared IRE.Sig with other six pan-cancer signatures. IRE.Sig exhibited better generalization capabilities and superior performance in forecasting ICI responses in different cohorts across diverse cancer types.
The combination of immunotherapy and other treatment methods, involving chemotherapy, and targeted therapy, may synergistically enhance tumor eradication. Considering the robust correlation between IRE.Sig and ICI outcomes, we further utilized the CRISPR screening data to investigate potential synergistic drug combinations with immunotherapy. Our study introduced an improved signature matching based computational method for the integrated analysis of diverse datasets, which calculated a score representing the probability of synergistic cancer immunotherapy drugs. It is worth noting that 4 of the top 10 drugs, including decitabine, tolazamide, tamoxifen, and sulfasalazine, have been shown to have immunomodulatory functions. It has been reported that decitabine, as a DNA methylation transferase inhibitor, could reactivate silenced tumor-related genes, thereby augmenting the recognition and attack capabilities of immune cells against tumors65. Additionally, tolazamide has been found to reverse the depletion of CD8+ T cell function caused by DOCK2 inactivation, and enhance the efficacy of the combined treatment of anti-PD-L1 antibody and apatinib for liver cancer66. For tamoxifen, a selective estrogen receptor modulator, can inhibit the stimulatory effects of estrogen on breast cells and enhance anti-tumor effects when used in conjunction with immunotherapy67. Lastly, sulfasalazine has been shown to increase the dependency on glutamine in cancer cells and inhibit tumor growth68. In conclusion, the evidence presented for the four drugs indirectly supported the validity of our calculation method and the dependability of the prediction results. Subsequent investigation into these highly ranked drugs will aid in a combined strategy for immunotherapy.
In addition, the results of this study hold considerable importance for clinical research beyond merely identifying subgroups with better survival after treatment with standard immunotherapy regimens. For instance, in a recent randomized trial involving breast cancer patients, the combination of pembrolizumab (an ICI drug) with eribulin failed to show clinical utility39. Nevertheless, upon reevaluation of the patient cohort, a distinct subgroup was identified that appeared to derive benefit from the addition of pembrolizumab. Interestingly, the identification of this subgroup was based on the prevalence of a distinct B cell state, but not malignant cells. Consequently, our findings suggested potential patient selection criteria for upcoming clinical trials involving eribulin and pembrolizumab.
Our study has some limitations. Firstly, our reliance on transcriptome data from published cohorts may not accurately reflect protein function. However, the utilization of NetBid2 for regulatory network analysis or metaVIPER for protein activity interference is restricted to specific cancer types. Furthermore, the absence of large-scale pan-cancer cohorts with proteomic data hindered comprehensive analysis. Moreover, more ex vivo, in vivo experiments need to be added to support our explored potential agents synergistic with immunotherapy. Lastly, our study only encompassed 10 ICI RNA-Seq cohorts, representing a limited scope of six cancer types (GC, SKCM, RCC, UC, NSCLC, and GBM). The correlation between IRE.Sig and anti-tumor immunity in 32 different cancer types may partially mitigate this issue. Nevertheless, the predictive value of IRE.Sig in a pan-cancer environment needs to be validated through future prospective ICI trials.
In summary, our study provided a pan-cancer clinical evidence that a specific tumor ecosystem subtype was linked to better immunotherapy response. Through analysis of pan-cancer scRNA-Seq data, we created a gene expression signature, IRE.Sig, that surpasses other established signatures in predicting outcomes of ICI treatment across multiple datasets. Additionally, further investigation of IRE.Sig determined several candidate drugs for immunotherapeutic combinations. Collectively, these findings expand our comprehension of tumor ecosystem, with potential implications for the development of biomarkers and personalized therapies.
Data availability
The datasets used in this study are available online, as described in the Supplementary Material. For additional information, inquiries can be directed to the corresponding author.
Code availability
This study utilized publicly available packages with R version 4.3.2. All software and algorithms used in this study are publicly available and listed in the Method section. All code generated for analysis is available from the authors upon request.
References
McAllister, S. S. & Weinberg, R. A. Tumor-host interactions: a far-reaching relationship. J. Clin. Oncol. 28, 4022–4028 (2010).
Schurch, C. M. et al. Coordinated Cellular Neighborhoods Orchestrate Antitumoral Immunity at the Colorectal Cancer Invasive Front. Cell 182, 1341–1359.e1319 (2020).
Smith, E. A. & Hodges, H. C. The Spatial and Genomic Hierarchy of Tumor Ecosystems Revealed by Single-Cell Technologies. Trends Cancer 5, 411–425 (2019).
Ribas, A. & Wolchok, J. D. Cancer immunotherapy using checkpoint blockade. Science 359, 1350–1355 (2018).
Sharma, P. et al. The Next Decade of Immune Checkpoint Therapy. Cancer Discov. 11, 838–857 (2021).
Galon, J. & Bruni, D. Approaches to treat immune hot, altered and cold tumours with combination immunotherapies. Nat. Rev. Drug Discov. 18, 197–218 (2019).
Yang, M., McKay, D., Pollard, J. W. & Lewis, C. E. Diverse Functions of Macrophages in Different Tumor Microenvironments. Cancer Res. 78, 5492–5503 (2018).
Suva, M. L. & Tirosh, I. Single-Cell RNA Sequencing in Cancer: Lessons Learned and Emerging Challenges. Mol. Cell 75, 7–12 (2019).
Aoki, T. et al. Single-Cell Transcriptome Analysis Reveals Disease-Defining T-cell Subsets in the Tumor Microenvironment of Classic Hodgkin Lymphoma. Cancer Discov. 10, 406–421 (2020).
Luca, B. A. et al. Atlas of clinically distinct cell states and ecosystems across human solid tumors. Cell 184, 5482–5496.e5428 (2021).
Sun, D. et al. TISCH: a comprehensive web resource enabling interactive single-cell transcriptome visualization of tumor microenvironment. Nucleic Acids Res. 49, D1420–D1430 (2021).
Van Allen, E. M. et al. Genomic correlates of response to CTLA-4 blockade in metastatic melanoma. Science 350, 207–211 (2015).
Abril-Rodriguez, G. et al. PAK4 inhibition improves PD-1 blockade immunotherapy. Nat. Cancer 1, 46–58 (2020).
Cui, C. et al. Ratio of the interferon-gamma signature to the immunosuppression signature predicts anti-PD-1 therapy response in melanoma. NPJ Genom. Med. 6, 7 (2021).
Hugo, W. et al. Genomic and Transcriptomic Features of Response to Anti-PD-1 Therapy in Metastatic Melanoma. Cell 165, 35–44 (2016).
Gide, T. N. et al. Distinct Immune Cell Populations Define Response to Anti-PD-1 Monotherapy and Anti-PD-1/Anti-CTLA-4 Combined Therapy. Cancer Cell 35, 238–255.e236 (2019).
Auslander, N. et al. Robust prediction of response to immune checkpoint blockade therapy in metastatic melanoma. Nat. Med. 24, 1545–1549 (2018).
Snyder, A. et al. Contribution of systemic and somatic factors to clinical response and resistance to PD-L1 blockade in urothelial cancer: An exploratory multi-omic analysis. PLoS Med. 14, e1002309 (2017).
Mariathasan, S. et al. TGFbeta attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature 554, 544–548 (2018).
Braun, D. A. et al. Interplay of somatic alterations and immune infiltration modulates response to PD-1 blockade in advanced clear cell renal cell carcinoma. Nat. Med. 26, 909–918 (2020).
Zhao, J. et al. Multi-omics indicators of long-term survival benefits after immune checkpoint inhibitor therapy. Cell Rep. Methods 3, 100596 (2023).
Kim, S. T. et al. Comprehensive molecular characterization of clinical responses to PD-1 inhibition in metastatic gastric cancer. Nat. Med. 24, 1449–1458 (2018).
Zhao, J. et al. Immune and genomic correlates of response to anti-PD-1 immunotherapy in glioblastoma. Nat. Med. 25, 462–469 (2019).
Wang, G. et al. CRISPR-GEMM Pooled Mutagenic Screening Identifies KMT2D as a Major Modulator of Immune Checkpoint Blockade. Cancer Discov. 10, 1912–1933 (2020).
Li, J. et al. Epigenetic and Transcriptional Control of the Epidermal Growth Factor Receptor Regulates the Tumor Immune Microenvironment in Pancreatic Cancer. Cancer Discov. 11, 736–753 (2021).
Li, F. et al. In Vivo Epigenetic CRISPR Screen Identifies Asf1a as an Immunotherapeutic Target in Kras-Mutant Lung Adenocarcinoma. Cancer Discov. 10, 270–287 (2020).
Dubrot, J. et al. In vivo screens using a selective CRISPR antigen removal lentiviral vector system reveal immune dependencies in renal cell carcinoma. Immunity 54, 571–585.e576 (2021).
Bagaev, A. et al. Conserved pan-cancer microenvironment subtypes predict response to immunotherapy. Cancer Cell 39, 845–865.e847 (2021).
Hanzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinforma. 14, 7 (2013).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007).
Vougas, K. et al. Machine learning and data mining frameworks for predicting drug response in cancer: An overview and a novel in silico screening process based on association rule mining. Pharm. Ther. 203, 107395 (2019).
Senbabaoglu, Y. et al. Tumor immune microenvironment characterization in clear cell renal cell carcinoma identifies prognostic and immunotherapeutically relevant messenger RNA signatures. Genome Biol. 17, 231 (2016).
Chakravarthy, A., Khan, L., Bensler, N. P., Bose, P. & De Carvalho, D. D. TGF-beta-associated extracellular matrix genes link cancer-associated fibroblasts to immune evasion and immunotherapy failure. Nat. Commun. 9, 4692 (2018).
Ayers, M. et al. IFN-gamma-related mRNA profile predicts clinical response to PD-1 blockade. J. Clin. Invest. 127, 2930–2940 (2017).
Rooney, M. S., Shukla, S. A., Wu, C. J., Getz, G. & Hacohen, N. Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell 160, 48–61 (2015).
Topalian, S. L. et al. Safety, activity, and immune correlates of anti-PD-1 antibody in cancer. N. Engl. J. Med. 366, 2443–2454 (2012).
Lamb, J. et al. The Connectivity Map: using gene-expression signatures to connect small molecules, genes, and disease. Science 313, 1929–1935 (2006).
Tolaney, S. M. et al. Effect of Eribulin With or Without Pembrolizumab on Progression-Free Survival for Patients With Hormone Receptor-Positive, ERBB2-Negative Metastatic Breast Cancer: A Randomized Clinical Trial. JAMA Oncol. 6, 1598–1605 (2020).
Fridman, W. H. et al. B cells and tertiary lymphoid structures as determinants of tumour immune contexture and clinical outcome. Nat. Rev. Clin. Oncol. 19, 441–457 (2022).
Gu, S. et al. Significance of intratumoral infiltration of B cells in cancer immunotherapy: From a single cell perspective. Biochim. Biophys. Acta Rev. Cancer 1876, 188632 (2021).
Liu, J. et al. Multicenter phase II trial of Camrelizumab combined with Apatinib and Eribulin in heavily pretreated patients with advanced triple-negative breast cancer. Nat. Commun. 13, 3011 (2022).
Meylan, M. et al. Tertiary lymphoid structures generate and propagate anti-tumor antibody-producing plasma cells in renal cell cancer. Immunity 55, 527–541.e525 (2022).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Yang, C. et al. Nonredundant Upregulation of CD112R (PVRIG) and PD-1 on Cytotoxic T Lymphocytes Located in T Cell Nests of Colorectal Cancer. Mod. Pathol. 36, 100089 (2023).
Han, X. et al. Ginseng-derived nanoparticles potentiate immune checkpoint antibody efficacy by reprogramming the cold tumor microenvironment. Mol. Ther. 30, 327–340 (2022).
Sun, Q. et al. BCL6 promotes a stem-like CD8(+) T cell program in cancer via antagonizing BLIMP1. Sci. Immunol. 8, eadh1306 (2023).
Xiao, Q. et al. DKK2 imparts tumor immunity evasion through beta-catenin-independent suppression of cytotoxic immune-cell activation. Nat. Med. 24, 262–270 (2018).
Li, H. et al. Identification of lysosomal genes associated with prognosis in lung adenocarcinoma. Transl. Lung Cancer Res. 12, 1477–1495 (2023).
Jin, J. et al. Landscape of DNA damage response gene alterations in breast cancer: A comprehensive investigation. Cancer 129, 845–859 (2023).
Finn, R. S. et al. Atezolizumab plus Bevacizumab in Unresectable Hepatocellular Carcinoma. N. Engl. J. Med 382, 1894–1905 (2020).
Yu, W. D., Sun, G., Li, J., Xu, J. & Wang, X. Mechanisms and therapeutic potentials of cancer immunotherapy in combination with radiotherapy and/or chemotherapy. Cancer Lett. 452, 66–70 (2019).
Tsherniak, A. et al. Defining a Cancer Dependency Map. Cell 170, 564–576.e516 (2017).
Liu, Y. et al. Improved clinical outcome in a randomized phase II study of anti-PD-1 camrelizumab plus decitabine in relapsed/refractory Hodgkin lymphoma. J. Immunother. Cancer 9, e002347 (2021).
Yoshida, T. et al. Eribulin mesilate suppresses experimental metastasis of breast cancer cells by reversing phenotype from epithelial-mesenchymal transition (EMT) to mesenchymal-epithelial transition (MET) states. Br. J. Cancer 110, 1497–1505 (2014).
Ueda, S. et al. In vivo imaging of eribulin-induced reoxygenation in advanced breast cancer patients: a comparison to bevacizumab. Br. J. Cancer 114, 1212–1218 (2016).
Helmink, B. A. et al. B cells and tertiary lymphoid structures promote immunotherapy response. Nature 577, 549–555 (2020).
Zhang, Z. et al. Pan-cancer landscape of T-cell exhaustion heterogeneity within the tumor microenvironment revealed a progressive roadmap of hierarchical dysfunction associated with prognosis and therapeutic efficacy. EBioMedicine 83, 104207 (2022).
Yi, M. et al. Combination of oral STING agonist MSA-2 and anti-TGF-beta/PD-L1 bispecific antibody YM101: a novel immune cocktail therapy for non-inflamed tumors. J. Hematol. Oncol. 15, 142 (2022).
Mastrogiovanni, M. et al. The tumor suppressor adenomatous polyposis coli regulates T lymphocyte migration. Sci. Adv. 8, eabl5942 (2022).
Wang, S. S. et al. Tumor-infiltrating B cells: their role and application in anti-tumor immunity in lung cancer. Cell Mol. Immunol. 16, 6–18 (2019).
Zhang, E. et al. Roles and mechanisms of tumour-infiltrating B cells in human cancer: a new force in immunotherapy. Biomark. Res. 11, 28 (2023).
Nicolai, C. J. et al. NK cells mediate clearance of CD8(+) T cell-resistant tumors in response to STING agonists. Sci. Immunol. 5, eaaz2738 (2020).
DuPage, M., Mazumdar, C., Schmidt, L. M., Cheung, A. F. & Jacks, T. Expression of tumour-specific antigens underlies cancer immunoediting. Nature 482, 405–409 (2012).
Nie, J. et al. Addition of Low-Dose Decitabine to Anti-PD-1 Antibody Camrelizumab in Relapsed/Refractory Classical Hodgkin Lymphoma. J. Clin. Oncol. 37, 1479–1489 (2019).
Wang, S. et al. SULT2B1-CS-DOCK2 axis regulates effector T-cell exhaustion in HCC microenvironment. Hepatology 78, 1064–1078 (2023).
Terranova-Barberio, M. et al. Exhausted T cell signature predicts immunotherapy response in ER-positive breast cancer. Nat. Commun. 11, 3584 (2020).
Wang, X. et al. SNHG3 could promote prostate cancer progression through reducing methionine dependence of PCa cells. Cell Mol. Biol. Lett. 27, 13 (2022).
Acknowledgements
This study was supported by Chongqing Natural Science Foundation Project (cstc2021jcyj-msxmX0204). We express our gratitude to the personnel of the TCGA Research Network, the EcoTyper tool, the UCSC Xena data portal, the DepMap portal, the CMap portal, and the TISCH2 data portal, as well as to all contributing authors for their efforts in sharing their valuable research data with the public.
Author information
Authors and Affiliations
Contributions
S.Z., H.W. and C.L. were responsible for the conceptualization and design of the study. Data collection was carried out by S.Z., L.C. and J.T. Data analysis was conducted by S.Z. and X.L. Manuscript writing and contribution were completed by S.Z., X.L. and Z.L. L.C. and H.T. revised the manuscript. All authors made contributions to the article and approved the submitted version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zeng, S., Chen, L., Tian, J. et al. Integrative analysis of pan-cancer single-cell data reveals a tumor ecosystem subtype predicting immunotherapy response. npj Precis. Onc. 8, 205 (2024). https://doi.org/10.1038/s41698-024-00703-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41698-024-00703-w
This article is cited by
-
Fibroblast TGF-β signaling defines spatial tumor ecosystems linked to immune checkpoint blockade resistance
Communications Biology (2025)
-
Single-cell and bulk transcriptome analysis identifies B-cell subpopulations and associated cancer subtypes with distinct clinical and molecular characteristics
Cellular Oncology (2025)