Abstract
Ductal carcinoma in situ (DCIS) may progress to ipsilateral invasive breast cancer (iIBC), but often never will. Because DCIS is treated as early breast cancer, many women with harmless DCIS face overtreatment. To identify features associated with progression, we developed an artificial intelligence-based DCIS morphometric analysis pipeline (AIDmap) on hematoxylin-eosin-stained (H&E) tissue sections. We analyzed 689 digitized H&Es of pure primary DCIS of which 226 were diagnosed with subsequent iIBC and 463 were not. The distribution of 15 duct morphological measurements was summarized in 55 morphometric variables. A ridge regression classifier with cross validation predicted 5-years-free of iIBC with an area-under the curve of 0.67 (95% CI 0.57–0.77). A combined clinical-morphometric signature, characterized by small-sized ducts, a low number of cells and a low DCIS/stroma ratio, was associated with outcome (HR = 0.56; 95% CI 0.28–0.78). AIDmap has potential to identify harmless DCIS that may not need treatment.
Similar content being viewed by others
Introduction
Ductal carcinoma in situ (DCIS) may progress to invasive breast cancer (IBC), but 3 out of 4 patients never will if left untreated1,2,3. Since the introduction of population-based breast cancer screening, the incidence of DCIS has increased at least seven-fold4. In current practice, we are not able to distinguish the minority of DCIS that is prone to progress to IBC from those that never will5,6. Due to this uncertainty, almost all women with DCIS are treated with surgery, often followed by radiotherapy7. This implies that many women with harmless DCIS carry the burden of intensive treatment without any benefit8,9. Therefore, there is an urgent need to classify DCIS lesions that will remain indolent, and those that might progress to IBC to prevent overtreatment of low-risk DCIS.
Several studies explored classical morphological and molecular features to predict ipsilateral subsequent IBC (iIBC) after DCIS10. Although DCIS grade, expression of the estrogen receptor (ER), human epidermal growth factor receptor 2 (HER2) and prostaglandin-endoperoxide synthase 2 (COX-2) have shown an association with invasive progression10,11,12,13, the clinical utility of their prognostic value is a subject of ongoing debate9,10,11,12,13,14,15,16,17.
Conflicting results are published on the relationship between DCIS lesion size and risk of progression to iIBC, may be because assessment of DCIS size on macroscopic examination of a specimen is not highly accurate18,19,20. At the microscopy level, it is also notoriously difficult to estimate the extent of DCIS accurately after tissue has been sectioned, also because a standardized method measuring the DCIS lesion size is lacking19,21. In addition, there is also an obvious knowledge gap on the prognostic value of morphological measurements in hematoxylin-eosin-stained (H&E) slides22,23,24,25. To address this, we developed an artificial intelligence-based DCIS Morphometric Analysis Pipeline (AIDmap) to detect, measure and quantify DCIS features with high accuracy and reproducibility. Measurements were obtained from scanned whole-slide images (WSIs) of H&E sections from primary DCIS lesions of a large retrospective study in The Netherlands13,26.
We hypothesized that objective, reproducible, and accurate measurements of morphometric features of DCIS lesions using AIDmap could help to stratify risk of progression to IBC. Ultimately, this may aid DCIS management decisions. First, by sparing many women with low-risk DCIS the burden of potential over-treatment. Second, by adequate treatment of women with high-risk DCIS, thereby not compromising the excellent outcomes of DCIS management currently achieved.
Results
AIDmap development
We obtained H&E WSIs from a nested case-control study of patients diagnosed with primary DCIS, treated by breast-conserving surgery without radiotherapy (methods). Patients diagnosed with iIBC during follow-up were considered as “cases” and those with no invasive diagnosis considered as “controls”13. Pathologists digitally annotated stroma and DCIS regions in H&E WSIs. The region segmented as stroma corresponds to the eosin-stained structures in high density per pixels, corresponding to the extracellular matrix and fibroblasts areas, including blood vessels and terminal duct lobular units, while adipocyte-only areas were left outside (Supplementary Fig. 1). These annotations were used by the HALO AI module, a deep learning neural network that created a trained-by-example tissue classifiers to detect stroma areas and DCIS ducts. Additionally, we applied a nuclei segmentation to detect the nucleus of cells within the ducts. HALO platform estimated the area, the perimeter and the spatial coordinates of these stroma areas, DCIS ducts and cell nucleus (Fig. 1). Finally, we applied a computational filtering to improve the detection accuracy of DCIS ducts (Fig. 1, methods section). This resulted in our artificial intelligence-based DCIS morphometric analysis pipeline (AIDmap).
HALO deep learning neural network was trained to recognize morphological structures in H&E whole-slides images (WSIs) (details in methods section). 1: The first classifier was trained to annotate the fibroglandular tissue (stroma), leaving adipocytes outside (green line). 2: DCIS classification was applied within the annotated stroma, by detecting pixels that reached more than 90% of confidence of composing a DCIS duct (red areas in the image heatmap). 3: Next, a nuclear segmentation sensing hematoxylin staining was applied within the DCIS regions to detect all nuclear structures. After these three steps, HALO provided tables containing the area, perimeter and spatial coordinates of stroma, DCIS and nuclear objects that were imported to R studio. 4: A True/False computational filtering was applied according to the nuclear perimeter, area and circular shape factor in order to eliminate false nuclear objects. And a True/False filtering was applied on DCIS objects, according the density of cells and average minimal nuclear distance (min. nucl. dist.) within the duct, to eliminate false DCIS ducts detected by HALO. Finally, morphological measurements for each DCIS duct were obtained.
Validation of DCIS detection
To validate the detection accuracy of the DCIS segmentation used in the AIDmap, we applied it on 20 H&E sections of DCIS lesions from an independent cohort (Translational Breast Cancer Research Consortium, TBCRC)27. All DCIS lesions within these 20 H&E slides were annotated by a pathologist, blinded to the AIDmap detection. The validation consisted of quantifying the spatial overlap of the computational prediction and the pathologist’s annotations using the intersection-over-union (IOU) score (Supplementary Fig. 2A–C)28. Spatial overlap between computational prediction and pathologist’s annotation had a median IOU score of 0.76 (interquartile range = 0.68–0.83; Supplementary Fig. 2D). Additionally, we compared the number of DCIS ducts detected by the pathologist and by AIDmap using Pearson’s correlation coefficient. The number of DCIS lesions assigned by the pathologist was strongly and significantly correlated with the number predicted by AIDmap: r = 0.79 (95% CI 0.44–0.89, p = 2.9 × 10−5; Supplementary Fig. 2E).
Spatial validation of DCIS segmentation
Given the highly heterogeneous architecture and growth of DCIS, we assessed the robustness of DCIS morphometry by comparing the measurements obtained from 3D image analysis of Formalin-Fixed Paraffin-Embedded (FFPE) DCIS blocks, with measurements obtained from AIDmap on H&E slides from the same FFPE DCIS blocks. We accessed 15 primary DCIS FFPE blocks from a living biobank of patient-derived DCIS mouse xenografts29. Details on the 3D-whole-mount imaging using immunolabeling and a modified FLASH protocol optimized for human archival tissue samples are described in the methods section)29,30. Measures of DCIS lesion volume, volume of individual DCIS structures, and distance between the ducts (crowding coefficient) were compared with the area and minimal distance between DCIS ducts obtained from AIDmap (Supplementary Fig. 3A, B). In order to avoid spatial bias, the H&E slide considered in the comparison, was the one with the largest area of DCIS ducts detected among all H&E slides available for each block. The average volume of DCIS structures in the 3D analysis of 15 primary DCIS FFPE blocks was positively correlated with the average area of DCIS duct areas per H&E slide measured by AIDmap (Pearson’s correlation: r = 0.76, p = 0.001, Supplementary Fig. 3C). When looking at the distribution of DCIS structures, the crowding coefficient obtained in 3D analysis was positively correlated with the average minimal distance between DCIS ducts obtained from AIDmap (Pearson’s correlation: r = 0.64, p = 0.009, Supplementary Fig. 3D).
Building a DCIS morphometrics classifier
In total, we uploaded 793 H&E WSIs of primary DCIS lesions from 793 patients treated with BCS only. However, after visual inspection, 104 did not achieve the minimal quality for computational segmentation due to weak hematoxylin or eosin staining, tissue section damage, artefacts, out-of-focus regions, or images scanned with scanner device out of the standard (Fig. 2A). We successfully applied AIDmap on 689 WSIs, in which 463 were assigned as controls and 226 assigned as cases (Fig. 2A and Table 1). We detected a total of 37,020 DCIS ducts in this dataset, with a median of 36 DCIS ducts per slide, ranging between 1 and 623.
A Flow chart of the study population of patients diagnosed with primary DCIS in the Netherlands between 1989 and 2004. For training in the HALO AI module, 57 H&E sections from DCIS treated with BCS alone were used for automated stroma and DCIS segmentation. In total, 689 H&E WSI were successfully analyzed and their images revealed the variability on the density of DCIS ducts: large (B) and small (C) duct size (mm2); density of DCIS cells within the ducts: high (D) and low (E) DCIS cells/μm2; and average size of DCIS nucleus of the cells within the DCIS ducts: large (F) and small (G) DCIS nucleus (μm2); among other morphometric variables that varied among the samples.
The area, perimeter and spatial coordinates of DCIS objects, stroma and cells were the basic measurements used to calculate all possible and reasonable geometric and spatial features for each DCIS duct. In summary, we obtained 15 morphological measurements (Supplementary Table 1).
DCIS duct morphological measurements displayed large intra- and inter-patient variability in the whole dataset of 689 WSIs (Supplementary Figs. 4–5). This variability, such as the area of DCIS ducts, the density of DCIS cells inside the ducts and the size of cell’s nucleus are examples of morphological structures in which the heterogeneity can also be observed by conventional microscopic examination of DCIS WSIs (Fig. 2B–G). AIDmap measured several of these morphological structures and their spatial arrangement observed in WSIs with high accuracy and precision.
Assessing the prognostic value of morphometric features
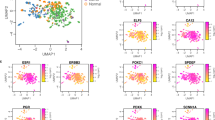
We further summarized the values of these morphological measurements in each WSI with the aim to reproduce the morphometric inter and intra-variability. In brief, we calculated 8 parameters of the distribution of the 15 morphological measurements in each WSI (Supplementary Fig. 6A), yielding in total 120 variables. Of these, 55 non-redundant variables were selected with representative potential to describe the morphometric heterogeneity of DCIS lesions (Supplementary Table 2 and Supplementary Fig. 6B). These 55 morphometric variables were mostly weakly correlated (85% had Spearman’s correlation coefficient between −0.5 and 0.5; see Fig. 3A).
A Heatmap with the Spearman’s rank correlation coefficients between the 55 variables obtained from the AIDmap in each H&E slide. Row side colors represent the parameters used to calculate each morphometric variable. Abbreviations are listed in Supplementary table 2. Receiver operating characteristic (ROC) curves and area under the curve (AUC) calculations from the generalized linear models to predict absence of iIBC event during follow up after 5, 10 or 15 years (B–D, respectively).
To assess the performance of the DCIS morphometric variables to predict 5, 10, and 15 years free of iIBC progression after primary DCIS diagnosis, we built a classifier using a logistic-ridge penalty regression based upon these 55 morphometric variables. After 10-fold double-loop cross validation, we evaluated the prediction accuracy in the test set from each loop. We obtained a median AUC of 0.67 (95% CI 0.57–0.77) to predict 5-years free of subsequent iIBC, 0.62 (95% CI 0.54–0.70) to predict 10-years and 0.62 (95% CI 0.54–0.69) to predict 15-years (Fig. 3B–D). In addition, we tried additional models to test the discriminant power of the 55 morphometric variables to predict iIBC progression: elastic nets in which AUCs were very similar to the ridge penalties (Supplementary Fig. 7A–C) and support vector machine, in which AUC values were lower and not statistically significant (Supplementary Fig. 7D–F). Finally, we evaluated the performance metrics based on the confusion matrix, by calculating the false negative rates (FNR), false positive rates (FPR), false detection rates (FDR) and false omission rates (FOR) for each loop of the cross-validation models (Supplementary Fig. 8). Values trend for a low FOR and high FDR towards the prediction for the first 5 years.
Constructing a morphometric signature
We applied univariate regression models on all the 55 morphometric variables to estimate their association with iIBC event status during follow-up in 5 levels: iIBC event diagnosed during the first 5 years (n = 83), between the 5th and 10th year (n = 90), between the 10th and 15th year (n = 33), later than the 15th year of follow-up after primary DCIS diagnosis (n = 7) or no iIBC event during follow-up (n = 476). We identified 30 variables significantly associated with iIBC status (Fig. 4A and Table 2).
A Volcano plot showing the odds ratios (OR) and the p values (p) of the 55 morphometric variables, obtained from linear regression analysis according the iIBC status during follow-up. B Heatmap of the hierarchical cluster analysis of the 30 morphometric variables statistically associated in the volcano plot. Row side colors in blue degrees represent the categories of iIBC events during follow-up. The dendrogram colors highlights the 4 groups sharing morphometric similarities.
Subsequently, hierarchical clustering of samples using these 30 morphometric variables identified four distinct morphometric signatures (Fig. 4B). One clear signature (1-blue) contained lesions with significant lower average levels of total DCIS area and DCIS/stroma ratio and lower number of cells in DCIS ducts, when compared to the other morphometric signatures: 2-red, 3-green and 4-orange (all p < 0.001; Fig. 6A–C). In addition, the 1-blue signature showed a higher proportion of clinging/FEA growth pattern, compared with the others (Supplementary Fig. 9A), as well as higher proportion of grade 1 DCIS (Supplementary Fig. 9B). Visually observing H&E slides from morphometric signature 1-blue, it is possible indeed to appreciate that DCIS ducts are not enlarged compared with the DCIS ducts from the other morphometric signatures (Fig. 5). Other differences between signature 1-blue and the other signatures are illustrated in Supplementary Figs. 9–10.
Morphometric signature of DCIS. Representative examples of H&E images with segmentation marks. A larger image and a magnification of the boxed area is shown for, respectively, the blue profile (A, B), the red profile (C, D), the green profile (E, F) and the orange profile (G, H).
Using a morphometric profile to identify low-risk DCIS lesions
By analyzing the 15 years iIBC cumulative risk curve, we observed that patients with lesions classified within the 1-Blue signature (containing lesions with small duct sizes, reduced number of cells, and lower DCIS/stroma ratio) had a significant favorable iIBC-free survival compared with the other signatures (p = 0.0001; Fig. 6C). The association with low-risk of iIBC events remained significant after multivariate Cox regression analysis including histopathological grade, ER, HER2 and COX-2 expression: hazard ratio (HR) = 0.56 (0.40–0.80 95%CI) (Fig. 6D). The 1-Blue signature also showed better iIBC-free survival among patients with DCIS lesions grade 1 or 2 (p = 0.014; Supplementary Fig. 11A), even after the multivariate Cox regression analysis including the same features: HR = 0.58 (95%CI 0.38–0.88) (Supplementary Fig. 11B).
Differences are illustrated in the violin plots of the distribution of 4 morphometric variables among the morphometric signatures: DCIS/stroma area ratio (A) and the total number of cells inside DCIS ducts (B). The iIBC risk curve for the patients classified with one of the morphometric signatures (C), and the forest plots from the Cox multivariate regression models estimating the risk of iIBC progression during follow-up (D).
Discussion
To the best of our knowledge, this is an original study indicating that automatically and objectively assessed microscopic morphometry of DCIS ducts in H&E whole slide images (WSIs) relate to the risk of progression of DCIS to subsequent ipsilateral invasive breast cancer (iIBC). This was done by developing, testing, and validating our AI-based DCIS Morphometric Analysis Pipeline (AIDmap). The main features related to low-risk DCIS were smaller ducts, lower DCIS/stroma ratio and lower number of DCIS cells per duct, whereas those with larger ducts and a higher cellularity are associated with a higher risk to progress to iIBC. This recently-designed tool has the potential to identify low-risk DCIS lesions that do not need surgical intervention and/or radiotherapy, saving many women the burden of such intensive treatment.
Using AIDmap, we achieved a high degree of agreement in DCIS area detection (IOU = 0.76) and high concordance in the count of DCIS ducts compared to the annotations of the breast pathologist (r = 0.79). This is highly relevant, as interobserver variability among breast pathologists is high when evaluating parameters that are based on morphological differences, such as grade and growth pattern15,17. Additionally, AIDmap morphological measurements in a single H&E section showed to represent the volume and spatial configuration of DCIS ducts in the whole FFPE block. It was demonstrated by the comparison of AIDmap results with a 3D whole-mount imaging analysis.
Of note, we applied AIDmap morphometric signature on H&E slides of subsequent sections from the same patient, in total: 326 H&E slides from 184 patients27 (142 two sections and 42 three sections per patient). As expected, due to the heterogeneity of DCIS morphology, the match rate of duplicated slides with the same morphometric profile was 54%. Still, 77% of the patients had duplicated H&E slides of 1-Blue signature or duplicated non-1-Blue signatures, revealing a higher match rate to distinguish 1-Blue signature from the others (Supplementary Fig. 12). By choosing the H&E slide with the most representative DCIS total area from each patient, we believe to reduce potential spatial bias when assigning an accurate morphometric profile using AIDmap. Importantly, we successfully performed AIDmap in H&E WSIs of primary DCIS lesions from patients treated in different hospitals, in a different continent, stained locally in their laboratory and scanned with a different scanner than used in our department. AIDmap was also performed in a few numbers of H&E sections from core biopsies, however, future research in a curated data set is necessary to design a signature considering the technical specificities of core biopsies, like the smaller stroma region and reduced DCIS total area.
AI-tools have been developed in cancer pathology to make predictions beyond the subjective interpretation by the pathologist, including outcome and treatment response31,32,33,34. However, AI algorithms often lack interpretability regarding the predicting features that are evaluated and recognized35. For instance, Klimov et al.34 developed a machine learning-based model to predict recurrence risk in primary DCIS using H&E WSIs34. They reported promising accuracy values to identify high-risk DCIS (HR = 6.39, 95%CI 3.0–13.8). However, their model was built based on features that cannot be explained or related to our current clinical and biological knowledge on DCIS. This might hamper acceptance in the clinic as a risk stratification tool. Additionally, the number of DCIS samples analyzed was limited, the patients were not uniformly treated, and the test dataset was composed by grade 3 DCIS lesions only34.
In previous decades, the measurement of geometric features and spatial arrangement of DCIS lesions would not be feasible using classical microscopy and H&E glass slides only. Here, we developed a hand-crafted pathologist-trained AI-based tool that provides an output of explainable features relating to DCIS histology, which consequently contributes to developing trust and transparency into an AI decision making process36. We also used a large patient dataset (n = 689), treated with BCS only, and including DCIS lesions from various histologic grade and growth patterns. It is important to mention that the use of ground truth information to train the AI algorithm can be time consuming and might feed human bias into the models37. Since the calculation of the morphometric features and the segmentation learning process were blinded to the clinical outcome variables, AIDmap has a relatively simple architecture and lower computation cost37.
To develop AIDmap, we have summarized the distribution parameters of the morphological measurements of DCIS ducts in H&E WSIs to reflect the inter- and intra-variability of their morphology. The morphometric signature developed in this study can be interpreted based on the results from a recent study using patient-derived mouse intraductal DCIS models, that reflected the full spectrum of DCIS morphology observed in patients29. Hutten et al. observed that DCIS lesions more prone to progress to IBC had larger volume and more spherical morphology (called as expansive growth pattern); when compared with DCIS lesions with lower rate of invasive progression, which had a smaller volume and more elongated shape (replacement growth pattern)29. In the present study, as we hypothesized, lesions containing smaller DCIS areas, lower number of cells, lower DCIS stroma/ratio and less spherical ducts grouped together in the cluster analysis. This group of DCIS lesions showed lower events of subsequent iIBC. Altogether, clinical H&E slides and data in model systems show that smaller DCIS duct sizes and non-spherical duct shapes are associated with a low progression rate.
A well curated dataset is key for successfully developing AI tools. The patient samples used in this study constitute one of the largest DCIS patient series treated with BCS only, long-term follow-up, well annotated clinical-pathological variables and with well-preserved tissue material13. Moreover, we performed our analysis on a nested case-control study, based on a population-based DCIS cohort. As such, we have a strong enrichment for cases, in relation to other published hospital or cohort studies, in order to increase the power to find clinically relevant associations. A limitation of our study is that accurate external validation is very difficult to achieve, as large, well-curated, unbiased datasets, analogous to the dataset analyzed here on which AIDmap is based, are not available yet. Additionally, the accuracy of DCIS segmentation and detection in digital slides could still be optimized, as larger datasets come available.
Different approaches attempting to predict DCIS lesions according to risk of invasive progression have been proposed, including some using protein expression11,12,13,14, a panel of gene expression16,38, stromal expression patterns27, immune cell composition27,39,40 and clinical-pathological models2,10,12,15,18,19,20. While interesting associations were reported, their potential to be translated to the clinic is unknown due to the lack of proper prediction accuracy tests. Our study addressed accuracy and probability of risk classification by applying penalized logistic regression and double-loop cross validation. The AUC values we got so far suggest that the morphometric signatures have clinical validity. In the case of molecular studies, the cost of such technologies could be an extra limitation. AIDmap is relatively low-cost, since it only requires inexpensive H&E slides, already routinely used in daily clinical practice.
Since AIDmap was developed based on tissue sections from BCS, it is essential to confirm its clinical utility in biopsies taken prior to treatment in guiding DCIS management. This is not trivial, because biopsies contain limited tissue that might not be fully representative of the DCIS lesion20.
In conclusion, AIDmap is the first step on the road to a promising tool to identify women with indolent DCIS lesions and therefore may be spared surgery and/or radiotherapy, while not compromising the excellent outcomes of women with high-risk DCIS that do need treatment.
Methods
Study population
We obtained data from a population-based cohort in which all women with primary DCIS without IBC between 1989 and 2004 in the Netherlands were included26. Endocrine treatment is not recommended in the Netherlands for women with DCIS. To avoid any confounding effect of endocrine treatment, we also excluded the few patients that received endocrine therapy. From this cohort, H&E slides of FFPE tissue blocks, and well-annotated clinical and follow-up data were available13,26. Median follow-up time is 12 years (interquartile range 9.0–15.3 years). Based on this cohort, a nested case-control study including women treated with breast-conserving surgery (BCS) alone was designed. Patients diagnosed with iIBC during follow-up were considered as “cases” and those with no invasive diagnosis considered as “controls”13. Matching between cases and controls was based on age at diagnosis and follow-up time. In addition, we obtained the expression levels of a series of molecular markers performed by immunohistochemistry, scored by pathologists, including ER, progesterone receptor (PR), HER2, COX-2, p53 and p16, as described previously13. These samples have well-annotated morphological profile evaluated in previous studies13,15, such as histologic grade, necrosis, microcalcification, periductal fibrosis, periductal lymphocytes and DCIS growth patterns: a description of the proliferative architecture of DCIS defined as solid, cribriform, micropapillary, or clinging/flat epithelial atypia (FEA); all scored by pathologists, as previously described by Visser et al and Groen et al.13,15 The study was approved by the review boards of the NCR (request K12.281; January 3, 2013) and PALGA (LZV990; April 16, 2013). The secondary use of tissue and data was allowed under an opt-out regime in this study conform Dutch regulations and the Code of Conduct of Federa-COREON, and therefore the Institutional Review Board provided a waiver for informed consent. The study was performed in accordance with the Declaration of Helsinki, with institutional ethics and local research governance approval.
DCIS tissue and cell segmentation
Pathologists selected one H&E slide for each patient diagnosed with primary DCIS lesion, in which DCIS ducts were most representative in number and size. We scanned H&E whole-slide tissue sections using the scanner Pannoramic P1000 of 3D Histech at the Core Facility of Molecular Pathology and Biobanking (CFMPB) in the Netherlands Cancer Institute (NKI) with a 20x objective and a 0.24 microns/pixel resolution. We uploaded all WSIs in the HALO platform developed by IndicaLabs (https://indicalab.com/).
Stroma areas were annotated in 98 H&E whole-slides images (WSIs) as the region corresponding to the extracellular matrix and fibroblasts (fibroglandular tissue), including blood vessels and terminal duct lobular units, while adipocyte-only areas were left outside. As illustrated in Supplementary Fig. 1, large areas with adipocytes were not considered as stroma region, however, when few adipocytes are embedded in the collagen and fibroblasts, they are considered as part of the stroma region. The criteria used by pathologists to digitally annotate DCIS ducts in 57 H&E WSIs, followed the one used to detect of DCIS lesions in the clinical pathology, characterized by the expansion of monomorphic appearing epithelial cells within mammary ducts. Each individual duct was annotated separately. The annotations followed the basal membrane of affected ducts. If one duct was partially populated by malignant cells, the whole duct was annotated, regardless of the size of the duct. During the development of the artificial intelligence-based DCIS morphometric analysis pipeline (AIDmap), stroma and DCIS ground truth annotations were used by the HALO AI module (https://indicalab.com/), a deep learning neural network that created a trained-by-example tissue classifiers to detect stroma areas and DCIS ducts respectively. During the fine tuning to optimize DCIS detection, we set the DCIS classifier to apply the segmentation only when the analyzed pixel reached detection confidence higher than 90% (Fig. 1). Additionally, we applied a nuclear segmentation classifier available in HALO, designed to sense areas stained with hematoxylin and identify the nucleus of cells within DCIS ducts. HALO platform estimated the area, the perimeter and the spatial coordinates of these stroma areas, DCIS ducts and cell nucleus. These measurements were exported as tab-delimited text files (Fig. 1).
Computational DCIS filtering
Due to the heterogeneity of DCIS morphology, H&E section artefacts and non-DCIS ducts similar to DCIS, we observed that HALO alone detected more DCIS objects than by pathologists. Based on the nature of these false positive detections, we applied a computational filtering to label some regions as non-DCIS ducts as described next. After segmentation, HALO output table was exported to Rstudio environment version 4.2.2 (2022-10-31). First, we calculated the circular shape factor of each object: [(4 x perimeter x area) / perimeter2]41. Due to some H&E artifacts, nuclear segmentation detected regions that were very big for a cell nucleus. After visual inspection, all nuclear objects with a perimeter larger than 125 μm, area larger than 455 μm2 or shape factor lower than 0.1, were not considered true nuclear objects and were removed from the data (corresponding to 0.2% of the total number of nuclear objects detected). Next, we calculated the number of cells inside each detected duct and its cell density (cellularity: cells/mm2). After rigorous visual inspection, DCIS ducts were defined as false detection when measurements reached the following criteria: duct detected with less than 5 cells, or with less than 26 cells and cellularity below 11 × 103 cells/mm2, or with less than 70 cells and cellularity below 800 cells/mm2, or with an average minimal distance between nuclear objects larger than 11 μm (Fig. 1). Detection of false DCIS ducts had an average of 45% (7–85%) among WSIs. The remaining true-labeled DCIS objects were the ones considered during external validation of the segmentation accuracy and for the morphometric analysis (Fig. 1).
3D analysis of human FFPE DCIS resections
We applied a spatial validation of DCIS detection. Once we are analyzing tissue sections in two dimensions (2D), but the DCIS lesion in embedded in a tissue with 3 dimensions (3D), we measured morphological structures of 15 DCIS lesions using a 3D imaging approach. FFPE blocks were immunolabelled and tissue-cleared with a modified FLASH protocol optimized for human archival tissue samples29,30. In brief, the FFPE tissue pieces were removed from the histology cassettes with a razor blade and deparaffinized in HistoChoice® for 2 h at 54 °C, after which the samples were 3 times washed in 100% MetOH, followed by incubation in dichlormethane for 3 h. After a second incubation in dichlormethane overnight, the tissue pieces were twice washed in 100% MetOH and bleached in 15% DMSO, 15% H2O2 in MetOH. The bleaching solution was refreshed for an extended incubation overnight. Before antigen retrieval, the pieces were rehydrated by a graded MetOH series (75% and 30% MetOH in dH2O), followed by 2 washes in dH2O, all for 1 h. We used FLASH reagent 2 (200 mM boric acid, 4 M urea and 8% 3-(Decyldimethylammonio)propanesulfonate inner salt (CAS 15163-36-7) in dH2O (pH ~7)) for antigen retrieval. Tissue pieces were equilibrated in FLASH reagent at room temperature for 1 h before the temperature was increased to 37 °C for overnight incubation. Next day, the solution was refreshed and temperature increased to 54 °C for 24 h. Excess retrieval solution was removed by repeated washes in PBT (0,2% Triton X-100 in PBS) at room temperature. For immunolabelling, samples were first washed in blocking buffer (10% FBS, 1% BSA, 5% DMSO, 0,2% Triton X-100, 0,02% NaAzide in PBS), before incubation in fresh blocking buffer with addition of mouse αSMA antibody, clone 1A4 (Sigma) diluted 1:1000 for 3 nights at room temperature. Excess antibody was removed by washes in PBS, after which fresh blocking buffer including AlexaFluor™-568 conjugated secondary donkey anti mouse IgG (Invitrogen) antibody 1:1000 and 1:1000 Hoechst 33342 was added and the samples incubated in the dark for 3 nights. Excess labels were removed by washes in PBS, and the samples dehydrated through a graded MetOH series of 30%, 50%, 75% and twice 100% MetOH in dH2O. For optical clearing, the tissue pieces were incubated in 30%, 70% and twice 100% methyl salicylate in MetOH for up to 6 h per incubation. After 2 days, methyl salicylate was replaced with a 2:1 mixture of benzyl benzoate and benzyl alcohol. The cleared tissues were imaged on an inverted multiphoton confocal microscope (Leica TCS SP8 MP) with a 25X water immersion objective (Fluotar VISIR 25x/0.95). Tiled z-scans capturing the entire FFPE blocks were acquired in Resonant Mode (8-bit) with 512 × 512 or 256 × 256 pixel format, 8000 Hz scan speed, 1.25 zoom, 2x line average and 5–15 μm z-steps. Fluorophores were excited simultaneously with an Insight X3 tunable two-photon laser at 800 nm. Three HyD-RLD detectors were used to simultaneously acquire SHG (390–410 nm), Hoechst emission (420–500 nm) and Alexa Fluor™ 568 emission (580–620 nm). Z-compensation of the detector gains was used to correct for lower detection levels in deeper tissue layers due to scattering of the emitted fluorescence. Imaris Viewer (9.7.2) was used for 3D visualization of the datasets and measurements were taken manually in Imaris (9.6.0). Ducts were revealed by the distribution of nuclear and αSMA staining. The line/polygon tool was used to measure the tissue sample size, DCIS lesion size, size of individual structures and distance between them within the lesion in three directions.
Summary of DCIS morphometric variability
We calculated the values for 8 parameters of the distribution of the 15 morphological measurements in each H&E WSI (Supplementary Fig. 6A). These parameters and abbreviations are:
-
1.
total [to]: sum of all the values
-
2.
mean [me]: average mean of the morphometric distribution
-
3.
median [md]: average median of the morphometric distribution
-
4.
range [rg]: the difference between maximal and minimal values of the morphometric distribution
-
5.
mode [mo]: the mode of the morphometric distribution
-
6.
Kernel density [kd]: bandwidth obtained from Kernel density
-
7.
standard deviation [sd]: standard deviation of the distribution
-
8.
skewness [sk]: skewness from normal distribution
The value of these parameters from each morphological measurements generated 120 morphometric variables for each H&E WSI. After Spearman’s correlation tests and linear regression analyses, we filtered out the variables highly correlated with each other (correlation > 0.85), always deciding to keep the variable with strongest association with outcome. We ended up with 50 of these variables. We included 5 additional variables that capture important DCIS microenvironment characteristics: the number of DCIS ducts detected in the H&E, the number of DCIS ducts per mm2, DCIS/stroma area ratio (calculated by dividing the total DCIS area by the total stroma area), total stroma area (area corresponding to the extra cellular matrix, excluding adipocyte-only areas), total area of non-DCIS ducts (calculated based on the area non-DCIS regions detected during the computational filtering). The complete computational process of H&E WSIs, including the segmentation obtained from HALO, the filtering developed in RStudio and the summary of the morphometric variability in each image, composes the AIDmap, which provides the morphometric profile and signature of the slide analyzed.
Statistical analyses
All statistical analyses were performed using R version 4.2.2 (2022-10-31) and RStudio as integrated development environment, including base packages and others as package stats (v4_4.2.2), rgeos (v0.6-4) and OptimalCutpoints (v1.1-5). Generalized linear models with ridge penalties, elastic net and Support vector machines (package caret_v6.0-94), measuring likelihood within a 10-fold double-loop cross-validation, were applied to evaluate the performance of the morphometric variables to predict classes with no iIBC progression after 5, 10 or 15 years of the primary DCIS diagnosis. A proportion of 2/3 training set and 1/3 test set was used for all prediction loops. Each classification model used the same random train/test data set in each of the 10 loops. After training, receiver operating characteristic (ROC) curve and area-under the curves (AUCs) were calculated in the test set of each loop. Besides AUC, the performance of the prediction models were evaluated with metrics derived from the confusion matrix, such as, the false negative rate (FNR), the false positive rate (FPR), the false detection rate (FDR), and the false omission rate (FOR)42. Odds ratios (OR) with their respective 95% confidence intervals (95% CI) were calculated using binomial and Gaussian logistic regression models to measure the association between morphometric variables and outcome status. We applied hierarchical clustering and Euclidean dissimilarity (package heatmaply v1.4.0) to stratify the morphometric variables (morphometric signature). Weighted Cox regression analysis was done to access associations between morphometric signatures and time to iIBC event43. Multivariate regression models were applied using clinical-pathological variables as described during the results section. Due to the large-scale range of the values of the morphometric measurements, their distributions were log-scaled in the regression models, and percentile-transformed in the cluster analyses. We assessed correlations between morphometric variables, signatures and clinical-pathological variables of DCIS primary lesions using Spearman’s rank correlation analysis. Distribution differences of clinical-pathological variables among different groups were tested using analysis of variance (ANOVA), Student’s t test, chi-squared test and linear regression models. All P values reported were calculated by two-sided tests and the threshold for significance set at p = 0.05. Other RStudio packages used to graphic representation: kableExtra (v1.3.4), ggplot2 (v3.3.6), pROC (v1.18.0), forestplot (v3.1.3) and survival (v3.4-0).
Data availability
The data used in this study is not publicly available due to privacy and ethical considerations. Researchers interested in accessing the data may contact the corresponding author. Data access is subject to request and approval. Any use of the data will be subject to compliance with relevant ethical and legal standards.
Code availability
Codes associated with this manuscript are enrolled to potential legal protection under patent law and, therefore, cannot be disclosed at this time due to the need to preserve intellectual property rights. We recognize the importance of reproducibility and transparency in scientific research. Access may be requested by contacting the corresponding author.
References
Ringberg, A., Palmer, B., Linell, F., Rychterova, V. & Ljungberg, O. Bilateral and multifocal breast carcinoma. A clinical and autopsy study with special emphasis on carcinoma in situ. Eur. J. Surg. Oncol 17, 20–29 (1991).
Maxwell, A. J. et al. Risk factors for the development of invasive cancer in unresected ductal carcinoma in situ. Eur. J. Surg. Oncol. 44, 429–435 (2018).
Ryser, M. D. et al. Cancer outcomes in DCIS patients without locoregional treatment. J. Natl Cancer Inst. 111, 952–960 (2019).
Myers, E. R. et al. Benefits and harms of breast cancer screening: a systematic review. JAMA 314, 1615–1634 (2015).
Falk, R. S., Hofvind, S., Skaane, P. & Haldorsen, T. Second events following ductal carcinoma in situ of the breast: a register-based cohort study. Breast cancer Res. Treat. 129, 929–938 (2011).
Rakovitch, E. et al. Refined estimates of local recurrence risks by DCIS score adjusting for clinicopathological features: a combined analysis of ECOG-ACRIN E5194 and Ontario DCIS cohort studies. Breast cancer Res. Treat. 169, 359–369 (2018).
Lazzeroni, M. et al. Adjuvant therapy in patients with ductal carcinoma in situ of the breast: the Pandora’s box. Cancer Treat. Rev. 55, 1–9 (2017).
King, M. T. et al. Patient-reported outcomes in ductal carcinoma in situ: a systematic review. Eur. J. cancer 71, 95–108 (2017).
Groen, E. J. et al. Finding the balance between over- and under-treatment of ductal carcinoma in situ (DCIS). Breast 31, 274–283 (2017).
Sanati, S. Morphologic and molecular features of breast ductal carcinoma in situ. Am. J. Pathol. 189, 946–955 (2019).
Glover, J. A., Hughes, C. M., Cantwell, M. M. & Murray, L. J. A systematic review to establish the frequency of cyclooxygenase-2 expression in normal breast epithelium, ductal carcinoma in situ, microinvasive carcinoma of the breast and invasive breast cancer. Br. J. Cancer 105, 13–17 (2011).
Visser, L. L. et al. Predictors of an invasive breast cancer recurrence after DCIS: a systematic review and meta-analyses. Cancer Epidemiol. Biomark. Prev. 28, 835–845 (2019).
Visser, L. L. et al. Clinicopathological risk factors for an invasive breast cancer recurrence after ductal carcinoma in situ-a nested case-control study. Clin. Cancer Res. 24, 3593–3601 (2018).
Weinmann, S. et al. Validation of a Ductal carcinoma in situ biomarker profile for risk of recurrence after breast-conserving surgery with and without radiotherapy. Clin. Cancer Res. 26, 4054–4063 (2020).
Groen, E. J. et al. Prognostic value of histopathological DCIS features in a large-scale international interrater reliability study. Breast Cancer Res. Treat. 183, 759–770 (2020).
Dettogni, R. S. et al. Potential biomarkers of ductal carcinoma in situ progression. BMC Cancer 20, 119 (2020).
van Seijen, M. et al. Variability in grading of ductal carcinoma in situ among an international group of pathologists. J. Pathol. Clin. Res. 7, 233–242 (2021).
Maffuz, A. et al. Tumor size as predictor of microinvasion, invasion, and axillary metastasis in ductal carcinoma in situ. J. Exp. Clin. Cancer Res. 25, 223–227 (2006).
Vicini, F. A. et al. Relationship between excision volume, margin status, and tumor size with the development of local recurrence in patients with ductal carcinoma-in-situ treated with breast-conserving therapy. J. Surg. Oncol. 76, 245–254 (2001).
Dillon, M. F. et al. Predictors of invasive disease in breast cancer when core biopsy demonstrates DCIS only. J. Surg. Oncol. 93, 559–563 (2006).
Dadmanesh, F., Fan, X., Dastane, A., Amin, M. B. & Bose, S. Comparative analysis of size estimation by mapping and counting number of blocks with ductal carcinoma in situ in breast excision specimens. Arch. Pathol. Lab. Med. 133, 26–30 (2009).
Norris, H. J., Bahr, G. F. & Mikel, U. V. A comparative morphometric and cytophotometric study of intraductal hyperplasia and intraductal carcinoma of the breast. Anal. Quant. Cytol. Histol. 10, 1–9 (1988).
Maiorana, A., Nigrisoli, E., Sannicola, C., Maiorana, M. C. & Cavallari, V. Proliferative lesions of the breast lobules. A morphometric analysis. Pathologica 82, 271–277 (1990).
Mayr, N. A., Staples, J. J., Robinson, R. A. & VanMetre, J. E. Intraductal breast carcinoma: initial results of a morphometric study using computerized digital image analysis. Clin. Oncol. 2, 66–70 (1990).
Ladekarl, M. & Sørensen, F. B. Quantitative histopathological variables in in situ and invasive ductal and lobular carcinomas of the breast. Apmis 101, 895–903 (1993).
Elshof, L. E. et al. Subsequent risk of ipsilateral and contralateral invasive breast cancer after treatment for ductal carcinoma in situ: incidence and the effect of radiotherapy in a population-based cohort of 10,090 women. Breast cancer Res. Treat. 159, 553–563 (2016).
Strand, S. H. et al. Molecular classification and biomarkers of clinical outcome in breast ductal carcinoma in situ: Analysis of TBCRC 038 and RAHBT cohorts. Cancer Cell 40, 1521–36.e7 (2022).
Rhee, D. J. et al. Automatic detection of contouring errors using convolutional neural networks. Med. Phys. 46, 5086–5097 (2019).
Hutten, S. J. et al. A living biobank of patient-derived ductal carcinoma in situ mouse-intraductal xenografts identifies risk factors for invasive progression. Cancer Cell 41, 986–1002.e9 (2023).
Messal, H. A. et al. Antigen retrieval and clearing for whole-organ immunofluorescence by FLASH. Nat. Protoc. 16, 239–262 (2021).
Han, Z. et al. Breast cancer multi-classification from histopathological images with structured deep learning model. Sci. Rep. 7, 4172 (2017).
Mercan, E. et al. Assessment of machine learning of breast pathology structures for automated differentiation of breast cancer and high-risk proliferative lesions. JAMA Netw. open 2, e198777 (2019).
Polónia, A. et al. Artificial intelligence improves the accuracy in histologic classification of breast lesions. Am. J. Clin. Pathol. 155, 527–536 (2021).
Klimov, S. et al. A whole slide image-based machine learning approach to predict ductal carcinoma in situ (DCIS) recurrence risk. Breast Cancer Res. 21, 83 (2019).
Haug, C. J. & Drazen, J. M. Artificial intelligence and machine learning in clinical medicine, 2023. N. Engl. J. Med. 388, 1201–1208 (2023).
Mandair, D., Reis-Filho, J. S. & Ashworth, A. Biological insights and novel biomarker discovery through deep learning approaches in breast cancer histopathology. NPJ Breast Cancer 9, 21 (2023).
Lipkova, J. et al. Artificial intelligence for multimodal data integration in oncology. Cancer Cell 40, 1095–1110 (2022).
Lin, C. Y. et al. Will oncotype DX DCIS testing guide therapy? A single-institution correlation of oncotype DX DCIS results with histopathologic findings and clinical management decisions. Mod. Pathol. 31, 562–568 (2018).
Narayanan, P. L. et al. Unmasking the immune microecology of ductal carcinoma in situ with deep learning. NPJ Breast Cancer 7, 19 (2021).
Almekinders, M. M. et al. Comprehensive multiplexed immune profiling of the ductal carcinoma in situ immune microenvironment regarding subsequent ipsilateral invasive breast cancer risk. Br. J. Cancer, 127, 1201–1213 (2022).
Exner, H. H. Quantitative description of microstructures by image analysis. Mater. Sci. Technol. https://doi.org/10.1002/9783527603978.mst0024 (2006).
Gonzales Martinez, R. & van Dongen, D.-M. Deep learning algorithms for the early detection of breast cancer: a comparative study with traditional machine learning. Inform. Med. Unlocked 41, 101317 (2023).
Schemper M., Wakounig S. & Heinze G. The estimation of average hazard ratios by weighted Cox regression. Stat. Med. 28, 2473–2489 (2009).
Acknowledgements
This work was supported by Cancer Research UK and by the Dutch Cancer Society (KWF) (ref. C38317/A24043). Research at the Netherlands Cancer Institute is supported by institutional grants from the KWF and the Dutch Ministry of Health, Welfare and Sport. Additional contributions from Breast Cancer Now (2015NovPR638), NIH R01 CA185138, CDMRP Breast Cancer Research Program Award (BC132057) and NWO–Veni (09150161910151). We thank all patients in the US and the Netherlands who have donated their data and tissue for this work. We also wish to thank all the collaborating hospitals, and in particular, pathology departments, and all persons who have helped in the process of data collection and analysis. The authors thank the registration team of the Netherlands Comprehensive Cancer Organization (IKNL) for the collection of data for the Netherlands Cancer Registry. We thank PALGA, the nationwide network and registry of histo- and cytopathology in the Netherlands, for providing pathology data and for their help in the collection of the residual patient material. We acknowledge the staff of the NKI Core Facility Molecular Pathology & Biobanking (CFMPB) for providing the tissue samples. Finally, we thank the Grand Challenge PRECISION Consortium Steering Group: Jelle Wesseling, Jos Jonkers, Jacco van Rheenen, Esther H. Lips, Marjanka Schmidt, Lodewyk F.A. Wessels and Proteeti Bhattacharjee (Netherlands Cancer Institute, Amsterdam, The Netherlands), Alastair Thompson (Baylor College of Medicine, Houston, Texas, USA), Serena Nik-Zainal, Helen Davies (University of Cambridge, Cambridge, UK), Elinor J. Sawyer (King’s College London, London, UK), Andrew Futreal, Nicholas Navin (MD Anderson Cancer Center, Houston, TX, USA), E. Shelley Hwang (Duke University School of Medicine, Durham, NC, USA), Fariba Behbod (Kansas University Medical Center, KS, USA), Daniel Rea (University of Birmingham, Birmingham, UK), Hilary Stobart (Independent Cancer Patients’ Voice, UK), Deborah Collyar (Patient Advocates in Research, USA), Donna Pinto (dcis411, USA), Ellen Verschuur, Marja van Oirsouw (Borstkanker Vereniging Nederland, the Netherlands).
Author information
Authors and Affiliations
Consortia
Contributions
Conceptualization: M.S.L., E.H.L., J.W. Resources: M.S.L., S.V., J.S. Data curation: M.A., L.V., E.G. and M.S.L. Software: S.V., X.M., N.L. and M.S.L. Formal analysis: M.S.L. and R.M. Supervision: Y.Y., R.M., E.H.L., J.V.R. and J.W. Funding acquisition: E.H.L., J.W., G.C.P.C. Validation: S.P.C., Y.B.H., C.E., H.A.M., E.S.H., M.vd.B., J.v.R. and Y.Y. Investigation: M.S.L., R.M., E.H.L., M.vd.B., H.A.M. and J.W. Visualization: M.S.L. Methodology: L.A., L.V., E.G., S.V., X.M., N.L., B.B., P.K., M.S.L, and H.A.M. Writing ± original draft: M.S.L. Project administration: S.V., P.K. Writing ± review & editing: All.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sobral-Leite, M., Castillo, S.P., Vonk, S. et al. A morphometric signature to identify ductal carcinoma in situ with a low risk of progression. npj Precis. Onc. 9, 25 (2025). https://doi.org/10.1038/s41698-024-00769-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41698-024-00769-6