Abstract
Accurate detection of the histological grade of pancreatic neuroendocrine tumors (PanNETs) is important for patients’ prognoses and treatment. Traditional grading methods based on mitotic count and Ki-67 index are subjective and time-consuming. In this work, we developed and validated a pathomics model for accurate grading of PanNETs. Pathomics features were extracted from H&E whole slide images (WSIs) using Lasso regression to create a pathomics score. Its performance was evaluated in three cohorts involving 2 centers and 272 patients. This score was significantly associated with PanNET grade and could differentiate between high- and low-risk groups. In the validation and test cohorts, the pathomics model, which combined the pathomics score with clinical features, achieved AUCs of 0.85 and 0.93, respectively. In conclusion, our model enhances data processing efficiency and provides a quantitative assessment of histological features, holding promise for guiding individualized treatment and risk stratification in PanNET management.
This study was registered at ClinicalTrials. Trial number: gov ChiCTR2400090898
Similar content being viewed by others
Introduction
Pancreatic neuroendocrine neoplasms (PanNENs), rare tumors originating from peptidergic neurons and neuroendocrine cells within the pancreas, have experienced a notable surge in incidental detections, increasing 4-7-fold due to advancements in health screenings and diagnostic technologies1,2. They constitute about 10% of all pancreatic malignancies, and carry substantial clinical significance3.
The 2019 WHO classification categorizes PanNENs into three distinct types: pancreatic neuroendocrine tumors (PanNETs), pancreatic neuroendocrine carcinomas (PanNECs), and mixed neuroendocrine-non-neuroendocrine neoplasms (MiNENs)4. Recent research highlights significant molecular differences between PanNECs and PanNETs, necessitating their distinct consideration5,6,7. Further subclassification of PanNETs into grades G1, G2, and G3, based on mitotic index and Ki-67 index, is pivotal as the pathologic grade directly influences treatment strategies and correlates strongly with survival rates. Specifically, the 5-year overall survival rates for G1, G2, and G3 tumors are 77.33%, 63.06%, and 20.04%, respectively8. In contrast, PanNECs, which are high-grade malignancies with aggressive biological behavior, exhibit significantly poorer outcomes, with reported survival usually <1 year2.
According to European Neuroendocrine Tumor Society (ENETS) guidelines, radical resection is recommended for all PanNETs exceeding 2 cm and functional tumors9. G1 PanNETs can be managed with surgeries such as central pancreatectomy and enucleation, which offer favorable outcomes and extended post-operative survival. G2 PanNETs, which exhibit intermediate biological aggressiveness, typically require standard radical resection with lymphadenectomy, and adjuvant therapy may be considered in cases with high-risk features (e.g., lymphovascular invasion, tumor size >4 cm or lymphatic metastasis)10. Conversely, G3 PanNETs, characterized by a high risk of recurrence, require adjuvant chemotherapy following resection11,12.Post-operative management is further individualized based on tumor grade, TNM stage, functionality, somatostatin receptor (SSTR) expression, and patient comorbidities. For instance, SSTR-positive tumors may benefit from peptide receptor radionuclide therapy (PRRT) or somatostatin analogs, while advanced-stage or metastatic disease often requires multimodal approaches combining systemic therapy, targeted agents, or immunotherapy10.
Accurate grading of PanNETs depends on Ki-67 (MIB-1) immunohistochemistry, as stipulated by the WHO classification13. Precise assessment of the tumor proliferation index requires manual counting of at least 500–2000 tumor cells14. However, this method is labor-intensive, costly, and can produce inconsistent results due to IHC preparation variability and pathologist subjectivity, particularly in highly differentiated PanNETs where minor differences (1-5%) can significantly affect tumor grade and treatment decisions15,16.
Recently, pathomics, an approach utilizing digital pathology, has gained attention for its potential to analyze extensive data from digital pathology images, generating quantitative features to characterize tissue sample phenotypes17,18. Subsequent analysis of these data aids in diagnosis and survival prediction. Hence, this study aims to develop and validate a pathomics model for precise grading of pancreatic neuroendocrine tumors, utilizing digital histopathological features derived from H&E-stained sections.
Results
Patient demographics
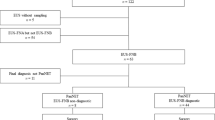
A total of 272 patients were finally enrolled. There was a significant difference in tumor size, the tumor burden score19 (TBS, which was defined according to the previously proposed calculation: TBS²= (maximum tumor diameter)²+ (number of tumors)²), N stage, M stage and disease progression between the G1, G2 and G3 groups in all sets. There was also a significant difference in the T stage, LVSI (lymph-vascular space invasion) and perineural invasion between the G1, G2 and G3 groups in the training and validation sets. The characteristics of the patients are detailed in Table 1. The inclusion and exclusion criteria for this study are shown in Fig. 1.
Flow chart visualizing the patient selection process.
The performance of the segmentation model
The DeepLabv3+ segmentation model exhibited exceptional proficiency in distinguishing PanNET regions from the adjacent pancreatic parenchyma (Fig. 2). The Tumor Intersection over Union (IoU) was 0.88 ± 0.02. Additionally, the Stroma IoU achieved a score of 0.89 ± 0.01.
A Original H&E-stained whole slide image (WSI). B Original H&E-stained WSI. C Model segmentation result (Tumor IoU: 0.90; Stroma IoU:0.92). D Model segmentation result (Tumor IoU: 0.89; Stroma IoU: 0.91). Segmented regions are color-coded: red (tumor) and blue (stroma).
Construction of the pathomics scores
We acquired a total of 427 pathomics features. First, we removed features with near-zero variance (coefficient of variation <0.1) across all WSIs, reducing features from 427 to 189. Then we liminated highly correlated features (Pearson’s |r | >0.8) using hierarchical clustering, retaining 63 features. After filtering, we applied 10-fold cross-validated LASSO with lambda.min criterion, selecting 6 features with non-zero coefficients (Fig. 2A, B). Then, the pathomics scores were calculated using the Lasso regression model. The pathomics scores revealed a notable downward trend from G1 to G3 tumors across all datasets, with G1 tumors showing consistently lower median scores (ranging from −0.67 to −0.45) compared to higher values in G2 (ranging from −0.27 to 0.07) and G3 (ranging from −0.45 to 0.08) tumors, and the P-values (<0.001 in the training and test sets, 0.011 in the validation set) underscored the statistically significant differences in pathomics profiles across these tumor grades. (Table 2 and Fig. 3C-E).
A The graph shows the LASSO coefficient profiles of 427 pathomics features. B The graph shows the selection of the tuning parameter (λ) in the LASSO model via 10-fold cross-validation based on the minimum criteria. Binomial deviances from the LASSO regression cross-validation procedure were plotted as a function of log(λ); the y-axis indicates binomial deviances. The lower x-axis indicates log(λ), and the upper x-axis shows the mean number of predictors. The red dots indicate the mean deviance values for each model with a given lambda, and the vertical bars through the red dots show the upper and lower values of deviances. The dotted lines define the optimal values of lambda, where the model provides the best fit to the data. The optimal lambda value of 0.056 with log(λ) = −1.252 was selected. C A Lasso regression path plot illustrates the varying coefficients of diverse pathomic features as the regularization parameter changes. D The bar chart illustrates the Pathomics scores for different patients. It distinguishes between two groups, G1 and G2/3, using red and blue colors respectively. E The combined boxplot and scatter plot illustrates the comparison of pathomics scores between G1 and G2/3. F The KM plot shows the progression-free survival (PFS) between the low-risk and high-risk groups according to the cutoff value of the pathomics score, which was 0.24.
Association between the pathomics score and PFS in the primary cohort
The cutoff value of the pathomics score was 0.24. The patients were divided into low-risk and high-risk group. Twelve progressions occurred in the low-risk group, and twenty-six progressions occurred in the high-risk group. The mean follow-up duration was 102.89 months (95% CI, 96.08–109.70 months) in the low-risk group and 71.52 months (95% CI, 58.06–84.98 months) in the high-risk group. The log-rank test revealed a longer survival duration in the low-risk group than in the high-risk group (P = 0.023) (Fig. 3F).
Univariable analysis
Tumor size, T stage, N stage, M stage, TBS, LVSI, Perineural invasion and pathomics score were significantly associated with G grades in the training set (all P < 0.05). The results of the univariable analysis are shown in Table 3.
Development, performance, and validation of clinical model
For the multivariate logistic regression analysis, tumor size, T stage, N stage, M stage, and TBS were selected for the clinical model. In the training set, 68 out of 82 patients (82.93%) were accurately identified in the G1 group, and 40 out of 67 patients (59.70%) were accurately identified in the G2/3 group using the clinical model. In contrast, in the validation set, 26 out of 38 patients (68.42%) were accurately identified in the G1 group, and 23 out of 27 patients (85.19%) were accurately identified in the G2/3 group using the clinical model. In the test set, 26 out of 40 patients (65.00%) were accurately identified in the G1 group, and 14 out of 18 patients (77.78%) were accurately identified in the G2/3 group using the clinical model. The clinical model demonstrated good discrimination, with an AUC of 0.80 (95% CI, 0.72–0.87) in the training set, 0.83 (95% CI: 0.73–0.93) in the validation set, and 0.77 (95% CI: 0.64–0.90) in the test set.
Development, performance, and validation of pathomics model
For the multivariate logistic regression analysis, tumor size, T stage, N stage, M stage, TBS, and pathomics score were selected for the pathomics model. In the training set, 66 out of 82 patients (80.49%) were accurately identified in the G1 group, and 58 out of 67 patients (86.57%) were accurately identified in the G2/3 group using the pathomics model. In contrast, in the validation set, 26 out of 38 patients (68.42%) were accurately identified in the G1 group, and 25 out of 27 patients (92.59%) were accurately identified in the G2/3 group using the pathomics model. In the test set, 31 out of 40 patients (77.50%) were accurately identified in the G1 group, and 17 out of 18 patients (94.44%) were accurately identified in the G2/3 group using the pathomics model.
The pathomics model demonstrated good discrimination, with an AUC of 0.91 (95% CI: 0.86–0.95) in the training set, 0.85 (95% CI: 0.75–0.94) in the validation set, and 0.93 (95% CI: 0.86–0.99) in the test set. The AUCs were significantly different between the pathomics model and the clinical model according to the DeLong test in the training set (P = 0.0002) and the test set (P = 0.03); however, they were not significantly different in the validation set (P = 0.66) (Table 4).
The Hosmer-Lemeshow test yielded p-values of 0.07, 0.38, and 0.26 in the training, validation, and test sets, respectively, suggesting no deviation from a good fit. Table 3 present the multivariate regression models. Table 5 and Fig. 4 show the performance of the clinical and pathomics models. Figure 5 illustrates two cases diagnosed using these models.
A–C The Confusion Matrix plots demonstrate the prediction outcomes of the clinical prediction model in the training set, validation set, and external test set. D–F The Confusion Matrix plots demonstrate the prediction outcomes of the pathomics prediction model in the training set, validation set, and external test set. G–I The ROC curves of the pathomics model and clinical model in the training set (AUC = 0.91 vs. AUC = 0.79), validation set (AUC = 0.85 vs. AUC = 0.83) and external test set (AUC = 0.93 vs. AUC = 0.77). J–L The decision curve analysis for the pathomics model and clinical model in the training, validation, and test sets, respectively. The x-axis represents the threshold probability. The y-axis represents the corresponding net benefit. The decision curves indicated that if the threshold probability fell between 0.05 and 0.95 (training set), between 0.21% and0.30, and 0.33 and 0.72 (validation set), and between 0.13 and 0.90 (Test set) in the pathomics model, respectively, using the nomogram to predict G1 and G2/3 of PanNET is more beneficial than the “treat-all-patients as G1” scheme or the “treat-all-patients as G2/3” scheme.
A–D A 45-year-old female patient has been diagnosed with PanNET G1, staged as T3N0M0. She remains alive, having achieved a progression-free survival (PFS) of 105 months. Notably, the clinical model misclassified her condition as G2/3, while the pathomics model accurately categorized it as G1. Furthermore, her pathomics score of −1.907 places her in the low-risk group, with a designated cut-off value of 0.24. A The figure shows the H&E slide at the whole-slide imaging (WSI) level. B The result from the semantic segmentation model applied to the WSI. C The sketch map shows the single-cell morphological and topological profiling. D Immunohistochemical staining for Ki-67, showing a Ki-67 proliferation index of 1%. E–H A 67-year-old female patient has been diagnosed with PanNET G2, staged as T2N0M0. Liver metastases were identified 14 months post surgery. Notably, the clinical model and the pathomics model also accurately classified it as G2/3. Additionally, her pathomics score of −0.208 places her in the low-risk group, with a designated cut-off value of 0.24. E The figure shows the H&E slide at the WSI level. F The result from the semantic segmentation model applied to the WSI. G The sketch map of single-cell morphological and topological profiling. H Immunohistochemical staining for Ki-67, showing a Ki-67 proliferation index of 18%. I Patient 1, diagnosed as G1 by the WHO G grading gold standard, received a total score of 106 based on model characteristics including TBS, Size, N, and M stages, yet the clinical nomogram incorrectly predicted a probability of 0.497 for a G2/3 diagnosis. Similarly, Patient 2, who was actually diagnosed as G2 by the gold standard, had a clinical nomogram incorrectly predicted a probability of 0.418 for a G1 diagnosis. These cases highlight errors in the clinical model’s diagnostic accuracy. This indicates that the clinical model made errors in the diagnosis of these two patients. J In contrast, the pathomics model, incorporating the combined pathomics score (−1.191), accurately predicted Patient 1’s diagnosis with a probability of 0.0257, aligning with the gold standard G1. For Patient 2, the inclusion of the pathomics score (−0.207) resulted in a predicted probability of 0.37, leading to a final diagnosis that concurred with the gold standard G2. This underscores the pathomics model’s diagnostic accuracy and highlights the pivotal role of the Pathomics score in enhancing model performance, effectively rectifying the clinical model’s diagnostic outcomes.
Development of survival prediction models
Univariable analysis identified tumor size (HR = 1.19, P < 0.0001), advanced T/N/M stages (HR = 3.74–15.28, P ≤ 0.041), positive margins (HR = 2.23, P = 0.026), lymphovascular invasion (HR = 2.88, P = 0.002), and WHO grade (G3 vs G1: HR = 22.89, P < 0.0001) as significant prognostic factors.
Multivariable models demonstrated: Survival model comparisons demonstrated that the Clinical-Grade Prognostic Model (incorporating WHO grading) achieved time-dependent AUCs of 0.91 (95%CI: 0.85–0.96), 0.942 (0.90–0.98), and 0.948 (0.91–0.98) at 12-, 24-, and 36-month intervals, respectively. The Pathomics Prognostic Model (utilizing pathomics score instead of WHO grade) showed comparable performance with corresponding AUCs of 0.865 (0.79–0.94), 0.897 (0.83–0.96), and 0.90 (0.84–0.96). Statistical equivalence testing revealed no significant differences between models at any time point (Delong’s P = 0.22, 0.12, and 0.06 for 12/24/36 months respectively), further supported by concordance indices of 0.90 vs 0.88 (P = 0.22). Complete model parameters are detailed in Supplementary Tables 2-3 and Supplementary Fig. 5.
DCA for pathomics nomogram
The x-axis represents the threshold probability, which is when the expected benefit of G1 is equal to the expected benefit of G2/3. The y-axis represents the threshold probability, which is the point at which the expected benefit of G1 is equal to the expected benefit of G2/3. The decision curves indicated that if the threshold probability fell between 0.05 and 0.95 (training set), between 0.21% and 0.30, and 0.33 and 0.72 (validation set), and between 0.13 and 0.90 (test set) in the pathomics model, respectively, using the nomogram to predict G1 and G2/3 of PanNET was more beneficial than the “treat-all-patients as G1” scheme or the “treat-all-patients as G2/3” scheme (Fig. 4J-L).
Association between predicted G grades and PFS in the primary cohort
In the G1 group, there were two progressions of disease, whereas the G2/3 group experienced thirty progressions of disease. The average follow-up period was 103.88 months (95% CI: 101.10–106.65 months) for the G1 group and 81.17 months (95% CI: 70.50–91.84 months) for the G2/3 group. According to the log-rank test, the predicted G1 group exhibited a significantly longer PFS compared to the predicted G2/3 group (P < 0.0001) (Supplementary Fig. 1). Univariate analysis revealed that the pathomics model-predicted G2/3 was associated with a shorter PFS (hazard ratio [HR]: 20.50; 95% CI: 4.96–84.79; P < 0.001). Furthermore, multivariable Cox regression analysis confirmed that the pathomics model-predicted G2/3 was an independent predictor of shorter PFS (HR: 17.59; 95% CI: 1.99–155.83; P = 0.01).
Cost-benefit analysis
Retrospective analysis revealed that pathologists spent 12.5 ± 3.1 minutes solely on mitotic counting and Ki-67 assessment (manual hotspot identification and counting 500–2000 cells). This supports the AI’s efficiency in automating these specific tasks (4.99 minutes vs. 12.5 minutes). In terms of diagnostic costs, the total cost for AI diagnosis using HE-based G grade for the 272 patients was $17,634, while the cost for traditional pathological diagnosis was as high as $25,071. Paired T-test analysis revealed extremely significant differences between the two groups in both diagnostic time and diagnostic costs (P < 0.0001).
Discussion
The grading of PanNETs is crucial for prognosis and personalized treatment9. Traditionally, grading relies on the assessment of mitotic count and cellular proliferative activity by pathologists, a method that is costly, time-consuming, and prone to variability. Consequently, there is a growing demand for more objective and reproducible grading methods. In this retrospective study, we introduced several innovative advancements. Firstly, we achieved the first fully automated and precise segmentation and quantification of postoperative H&E-stained PanNET sections, revealing pathological features not visible to the naked eye and laying the foundation for exploring PanNET heterogeneity and novel targets. Secondly, we introduced the “pathomics score”, a novel biomarker based on precise pathological quantification and analysis of cell morphology and topology. This score not only correlated strongly with PanNET G-staging but also effectively stratified patients into high- and low-risk groups, with significant differences in PFS. Thirdly, our pathomics model, integrating pathomics scores with other pathological features, significantly outperformed the clinical model in predicting G stages. Furthermore, the pathomics model-predicted-WHO grade were associated with significant differences in PFS and served as an independent prognostic indicator. Importantly, our pathomics model, based on H&E-stained sections, offers simplicity, economy, and efficiency compared to traditional immunohistochemical diagnosis, making it more clinically feasible.
A key strength of this study is the meticulous segmentation and precise quantification of H&E-stained pathological slides of pancreatic neuroendocrine tumors (PanNETs), which are notable for their marked heterogeneity20,21. Traditional microscopic examination struggles with accurate quantification and classification of these features. To overcome this, we utilized AI technology for precise segmentation and quantification. PanNETs consist of two major components: tumor cells and stroma, each with distinct characteristics contributing to tumor heterogeneity. Well-differentiated PanNETs, for example, exhibit clear boundaries, uniform tumor cell arrangement, and a salt-and-pepper chromatin pattern under H&E staining20,21. Additionally, stromal components like fibrosis play a crucial role in tumor growth and invasion, with tumors with thick fibrotic septa demonstrating enhanced aggressiveness22,23,24. Therefore, precise segmentation of tumor cells and stroma was a primary objective. Another significant feature of PanNETs is the tumor bed. Tumor cells accumulate within a certain spatial range, forming a coherent area of the tumor bed. This not only helps assess the thoroughness of the tumor removal but also provides important reference for subsequent treatment decisions25. Quantitative analysis of this region aids in assessing the risk of recurrence after tumor resection. Furthermore, PanNET heterogeneity is particularly prominent at the single-cell level. Through single-cell analysis, we can uncover topological relationships between cells and characteristics of the tumor microenvironment, which jointly determine PanNET biological behavior and clinical prognosis26. In this study, we extracted 378 features from tumor, inflammatory, and stromal cells, categorizing them into morphological, texture, and topological features to analyze cell shape, pixel distribution, and cell-cell relationships. Overall, this study revealed PanNET heterogeneity from multiple dimensions, including the tumor, stroma, tumor bed, and single-cell levels.
This study introduces a novel pathological biomarker, the pathomics score, which offers a fresh perspective for treating neuroendocrine tumors. The score seamlessly combines six key features extracted from HE-stained slides, spanning quantitative, morphological, textural, and graphical dimensions. These features delve into the tumor ecosystem in WSIs, providing a holistic view of tumor characteristics. Specifically, “tumor bed solidity” assesses malignancy by reflecting the cell density of the tumor cell aggregation area. Morphological features like “I Morph Circularity min” and “S Morph Curv Max min” reveal irregular growth and invasion, while “Morph Extent min” evaluates tumor morphological variation. The textural feature “I Texture Homogeneity mean” quantifies uniformity of cell composition and arrangement within the inflammatory region. Lastly, the graph-based “Graph I.S min Edge Length max” feature describes stromal-inflammatory cell relationships, indicating immune infiltration levels27. Individually, these features have unique biological and clinical significance; collectively, they form the comprehensive pathomics score. This score not only refines neuroendocrine tumor treatment but also opens doors for multimodal data analysis and personalized cancer therapies in future research.
The third and notable advantage of this study lies in the creation of a predictive model for G grading that is not only accurate and efficient, but also cost-effective, precise, and straightforward. Previous research has predominantly concentrated on predicting pathological grade using preoperative imaging. For instance, Wang et al. analyzed three-phase CT images (plain scan, artery, and vein), constructed a model based on eight radiomic features, and combined these with T stage and dilated main pancreatic duct/bile duct status to create a nomogram, achieving an AUC of 0.875 in the test set. However, this model was limited by a small sample size of 139 patients and lacked external validation28. Similarly, our team previously investigated the correlation between MRI radioactive score (rad-score) and NF-PanNET grade using arterial and portal vein MRI, finding a significant association (AUC = 0.775); nonetheless, the model’s performance was lower in the validation set (AUC = 0.736)29. To the best of our knowledge, research based on deep learning for PanNETs H&E slides is extremely rare. Klimov et al. 30 utilized deep learning to predict metastasis risk in PanNETs based on histological image analysis. While their work highlighted the prognostic value of morphological features, our study extends this approach by developing a pathomics model for accurate grading of PanNETs. Our study marks the first pathomics grading investigation of H&E sections in PanNET patients, involving 272 patients from two clinical centers. Our model achieved robust and generalizable AUCs of 0.91 (95% CI: 0.86–0.95), 0.85 (95% CI: 0.75–0.94), and 0.93 (95% CI: 0.86–0.99) in the training, validation, and test sets, respectively. Compared to traditional grading, our pathomics model offers several advantages: it eliminates the need for time-consuming and costly Ki-67 immunohistochemistry, utilizing routine H&E sections for enhanced accessibility and cost-effectiveness; it provides more meaningful prognostic information by incorporating data from the entire tumor without sampling bias; and it enables real-time grading, streamlining diagnostics and facilitating timely treatment decisions. Critically, this model can be seamlessly integrated into existing clinical workflows through a two-step stratification strategy: initial WHO grading to guide systemic therapy selection (e.g., platinum-based regimens for G3/NEC9), followed by pathomics-guided risk refinement to personalize surveillance intensity for G1/G2 cases and molecular subtyping for G3 tumors.
Our head-to-head comparison between traditional WHO grading and the pathomics score in survival prediction demonstrates statistical non-inferiority of our computational approach. Rather than replacing manual Ki-67/mitotic counting, we propose the pathomics model as a complementary tool that can enhance workflow efficiency, particularly in resource-limited settings or borderline cases where subjective assessment may lead to inconsistent grading. The modest differences in AUC values (2.8–5.3%) between models suggest that while WHO grading remains a powerful prognosticator, pathomics analysis captures additional dimensions of tumor biology that may refine risk stratification within traditional grade categories.
While this study marks a significant advancement, it’s important to recognize its limitations. First, the pathomics model was developed using retrospectively collected data from surgically treated patients, potentially introducing selection bias and limiting generalizability. Second, the model requires access to digital pathology infrastructure, including WSI scanners, data storage, and computational capabilities, which may not be available in resource-limited settings. This may constrain the immediate applicability of our approach in certain clinical environments. Third, the use of a single scanner may restrict the results’ applicability across different optical platforms. Future research will incorporate multiple scanners for enhanced robustness. Fourth, this study has two key limitations: (1) patients with PanNECs and MiNENs were excluded, and (2) the model does not separately predict G2 vs. G3 PanNETs despite their divergent therapeutic implications (e.g., platinum-based chemotherapy for G3 vs. targeted therapy/PRRT for G2). Future studies should expand the cohort size to improve accuracy across all PanNEN subtypes and optimize model granularity to distinguish G2 from G3 cases, ensuring precision medicine tailored to biologically distinct high-grade entities. Fifth, despite a non-significant AUC difference (P = 0.66) between the pathomics (0.85) and clinical (0.83) models in the validation set (n = 65), both show high diagnostic capabilities. In decision curve analysis, using the pathomics model yields significantly greater benefits than the clinical model when predicted probabilities are 21–30% or 33–72%, highlighting the importance of model selection based on probability ranges to maximize patient benefits. Prospective clinical validation of the pathomics model is needed to confirm its efficacy and reliability, which we plan to undertake next. Lastly, the exclusion of SSTR expression and functional status from the model due to reliance on HE sections alone and incomplete documentation, respectively, though future work aims to integrate these features for comprehensive risk stratification.
In conclusion, our study successfully developed a pathomics model augmenting traditional grading by quantifying histologic heterogeneity and providing prognostic insights beyond Ki-67. It is designed to assist—not replace—pathologists, particularly in cases with ambiguous morphology or at the threshold boundary value. Its potential to guide individualized treatment and risk stratification underscores its clinical relevance and promises to revolutionize PanNET management. In future work, we aim to develop strategies to improve the accessibility of the model, such as optimizing it for lower computational resources or creating web-based tools that allow remote access to the model without the need for extensive local infrastructure. In future work, we aim to improve the model’s accessibility by optimizing it for lower computational resources and developing web-based tools for remote access; concurrently, prospective multicenter validation is underway (planned enrollment: 120 G3 PanNETs and 60 PanNECs) to refine a high-grade classifier integrating pathomics, radiomics, and targeted sequencing for systemic therapy guidance.
Methods
Ethics approval
The research received clearance from the Ethics Committee of Changhai Hospital, and adhered to stringent ethical standards (Approval No. CHEC-Y2024-011). Data in external test set was approved by the Ethics Committee of Shanghai 411 Hospital (Approval No. AH411-2024-07). Given the study’s retrospective framework and reliance on standard clinical practices, the need to obtain informed consent was waived by the Institutional Review Board. The study was conducted in accordance with the Declaration of Helsinki. This retrospective study followed the STARD and TRIPOD guidelines for individual prognosis or diagnosis31,32.
Patients’ data acquisition
The study enrolled patients who underwent surgical resections of PanNET between April 2014 and January 2024. There were 160 consecutive patients (mean age \(\pm\) standard deviation, 54 years \(\pm\) 11; 90 men) with G1, and 112 consecutive patients (mean age \(\pm\) standard deviation, 52 years \(\pm\) 12; 55 men) with G2/G3. The patients’ records were reviewed to obtain demographic and clinicopathological data. This study was registered at ClinicalTrials.gov (trial number: ChiCTR2400090898). The study design is illustrated in Fig. 6.
A Development of the pathological segmentation model. B Multi-granularity feature extraction. C Feature selection. D Prediction model for G grading of pancreatic neuroendocrine tumors (PNET).
The inclusion criteria were patients with confirmed non-functional neuroendocrine tumors of the pancreas through pathological examination. The exclusion criteria were: (a) patients with pathological types of pancreatic neuroendocrine carcinoma (NEC) or mixed pancreatic neuroendocrine neoplasms (MiNEN); (b) patients with concurrent other malignant tumors; (c) patients with few(tumor tissue comprising <10% of the WSI) or no tumor tissues in the whole slide images; (d) patients who were lost to follow-up.
Following the TRIPOD guidelines, the patient cohort, named the primary cohort, from XX Hospital was split into two groups: a training set consisting of individuals enrolled between April 2014 and October 2021, and a validation set comprising patients enrolled from November 2021 to January 2024. Additionally, an external test set was from the XXX Hospital between May 2019 and December 2021.
Sample size estimation
It was expected that a multivariate prediction model would be developed using five candidate prediction parameters. Based on previous evidence, assuming a 2-year PFS rate of 83%, a mean follow-up time of 3.5 years, and an area under the curve (AUC) of 0.8 for the predictive model, the minimum sample size required was 66, based on pre-set parameters. The sample size of this study met these requirements.
Endpoints
The primary outcome of this study was the histopathologic grade of PanNETs, while the secondary outcome was progression-free survival (PFS), measured from surgery to disease progression or last follow-up. PFS encompassed disease control and survival status. Patients underwent follow-up imaging with dynamic enhanced CT or MRI every 3-6 months for the first 2-3 years, then annually. Disease progression was defined as local recurrence, distant metastases, or enlargement of existing metastatic lesions9. Follow-up concluded on July 17, 2024.
Pathological features
Pathological findings were initially extracted from institutional diagnostic reports. To ensure diagnostic accuracy, all cases underwent secondary review by a pancreatic pathologist (J.H.) with two decades of subspecialty experience. Discordant interpretations between primary and secondary assessments were adjudicated through consensus review with an independent senior consultant (JM.Z) possessing 30 years of pancreatic pathology expertise. The histopathological grading, conducted in strict accordance with the WHO 2019 classification criteria, serves as the gold standard for establishing our prediction model. Mitotic figures and Ki-67 proliferation indices were quantitatively assessed through dual independent evaluations by blinded pathologists, with interobserver discrepancies resolved by a third impartial reviewer.
The following pathological features were documented: (1) the size of the tumor; (2) the location of the pancreatic tumor; (3) the number of tumors; (4)TBS, the previously proposed calculation: TBS²= (maximum tumor diameter)²+ (number of tumors)²; (5) TNM categories (determined by pathologists according to the American Joint Committee on Cancer TNM Staging Manual, 9th Edition)33; (6) tumor texture (solid or cystic, cystic was defined as the pathological appearance of any containing cystic component)34; (7) LVSI; (8) Perineural invasion; (9) Surgical margin; (10) G grades according to the WHO 2019 criteria4. Patients were categorized into two groups: Grades 1 and Grades 2/3.
Pathomics workflow
One pathology image containing the largest tumor region was selected from all digital pathology images of the patient by a pathologist with 5 years of experience. Extract WSIs in the above training set for segmentation model development and the validation set and external test set for segmentation model validation.
WSIs preparation
H&E-stained histopathological slides of surgically resected tumors were retrieved from the archives of the pathology department. These slides were then scanned into WSIs using a Hamamatsu NanoZoomer S60 scanner (Hamamatsu Healthcare, Tokyo, Japan) at a magnification of 40×. To ensure image quality, sections that did not meet our standards were either rescanned or excluded from subsequent analyses. Specifically, any slides exhibiting defects such as air bubbles beneath the coverslip, suboptimal staining, noticeable fading, tissue folding, or scanning artifacts (such as out-of-focus regions) were not used. Furthermore, we ensured that all patients included in the study had complete clinical and pathological data, with no instances of missing information.
Pathological segmentation model development
The workflow comprised four key stages: image preprocessing, histopathological annotation, patch-level segmentation model development, and WSI-level segmentation integration.
Image preprocessing
WSIs, often several gigabytes in size, presented significant challenges for comprehensive annotation. To overcome this, the WSIs were divided into smaller, manageable patches of 256 × 256 pixels. Image enhancement techniques such as normalization, denoising, and histogram equalization were applied to optimize model performance. Additionally, rigorous data validation was performed to ensure consistent label counts and precise alignment between masks and image dimensions.
Histopathological annotation
A pathologist selected 70 WSIs from the training set by simple random sampling and identified representative regions of interest (ROIs) containing tumors and stroma for slicing. This process yielded a total of 9200 patches. Two junior pathologists used the Labelme tool (version 5.0.1) [available at https://github.com/wkentaro/Labelme] to annotate the tumor boundaries, ensuring that the annotations tightly enclosed the tumor cells without overlapping into the tumor-stroma regions. Each pathologist annotated a total of 3890 patches. An experienced pathologist with over 15 years of expertise reviewed the annotations, particularly for cases with ambiguous tumor boundaries. After the annotation process, multi-class masks were generated, assigning a class value to each pixel: 1 for tumor, 0 for background, and 2 for stroma. The annotated patches and their corresponding masks were exported and stored in PNG format, with separate folders for the original patches and mask files.
Patch-level segmentation model development
Leveraging the data derived from the previously described WSIs, we developed a semantic segmentation model based on DeepLab-v3 + 35. The training process utilized the Stochastic Gradient Descent (SGD) optimizer. The initial learning rate was set to 0.01, and a polynomial learning rate decay strategy was applied, progressively reducing the learning rate to converge to 0. The batch size was set to 16. To enhance generalization, data augmentation techniques such as random flipping, random scaling (0.5x to 2x range), and random cropping were implemented. The pixel-wise classification was optimized using the cross-entropy loss function, and the mean Intersection over Union (MIoU) was computed on the validation set to evaluate segmentation performance. The calculation of MIoU involves the following key steps: When calculating IoU, first determine the intersection and union areas between the predicted segmentation result and the ground truth segmentation result. Then, divide the intersection area by the union area to obtain the IoU value, which reflects the degree of overlap between the two. For multi-class segmentation tasks, calculate the IoU for each class, then average these IoU values to get the MIoU. MIoU is used to assess the overall performance of the model across all categories.
If the MIoU falls below 0.80, a senior pathologist will correct the dataset. The model will be retrained iteratively until the MIoU reaches or exceeds 0.80. Once this threshold is achieved, further dataset corrections will cease, and the model will be designated as the optimal semantic segmentation model. To quantitatively evaluate the segmentation model’s performance, we computed pixel-wise Intersection-over-Union (IoU) for tumor regions and stroma regions on 15 WSIs in the additional test set.
After the tumor segmentation results were obtained, the region of the tumor bed was determined on the basis. Generating a tumor bed from scattered tumor cell regions involves aggregating these cells within a defined space to create a coherent area. This method first merges neighboring tumor cell regions based on a distance threshold to form an initial outline. It then uses morphological dilation to expand boundaries and further merge close but unconnected regions, while controlling the dilation extent to prevent over-expansion. Finally, it employs edge smoothing and erosion-opening operations for post-processing to remove noise and optimize the shape of the tumor bed (Supplementary Fig. 4).
WSI-level segmentation integration
The individual patch segmentations were integrated into a unified segmentation map for the entire whole slide image (WSI). (a) The location information for each patch was obtained by recording the starting coordinates, as the patch size is fixed at 256 × 256 pixels. (b) The patient WSI metadata was extracted using the OpenSlicer software package (https://github.com/OpenSlicer/OpenSlicer). After obtaining the necessary metadata, a blank matrix matching the WSI size was created. This matrix will be used to store the processing results of each patch, ultimately constructing a comprehensive view of the entire WSI. (c) Patches were mapped to the WSI matrix based onto their coordinate information. Each patch was accurately aligned, ensuring no overlap or omission, ultimately forming a complete image. (d) All patches were integrated to achieve the overall segmentation of the target tissue components. The classification and segmentation results from the different patches were combined to identify specific tissue structures. Additional data processing techniques were applied to address boundary ambiguity or classification inconsistencies, ensuring the accuracy and reliability of the final results.
Quantitative feature extraction
Utilizing the segmentation outcomes, we proceed to quantify the size and area of tumor tissue, tumor bed, and stromal tissue. The area was calculated using the following formula: S = P-i,*,S-x, *,S-y. Here, P-i represents the total number of pixels in the target component, S-x represents the pixel spacing in the x-axis direction, and S-y represents the pixel spacing in the y-axis direction. In essence, the area of the target component was determined by multiplying its total number of pixels by the actual space size occupied by each pixel in the x and y directions. The image processing tool used was OpenCV 4.8.0 (https://github.com/opencv/opencv/releases/tag/4.8.0). This process facilitates the extraction of 49 quantifiable parameters, categorized into three groups: tumor related parameters, tumor bed-related parameters, and tumor-stroma ratio-related parameters. The proportion of neoplastic cell areas to stroma within the tumor bed, calculated as: (Tumor Area/(Tumor Area + Stroma Area)) × 100%. Specifically, there are eight features related to tumor, 26 features related to the tumor bed, and 15 features related to the tumor-stroma ratio (Supplementary Table 1).
Single-cell profiling with morphology and topology insights
Given the complex tissue distribution within WSIs and the weak correlation between cell distribution in the background, normal acini, and the tumor microenvironment, segmenting all regions for cell semantics would result in significant redundant calculations. Therefore, this study primarily focuses on regions of medical interest, namely the tumor, stroma, and lymphocytes. The process consists of three steps: (a) Nuclear segmentation and classification; (b) Cell-level feature extraction; (c) Visualization. All steps are based on the methodology outlined on the following website: https://github.com/fuscc-deep-path/sc_MTOP.
Nuclear segmentation and classification
We employ a pre-trained HoVer-Net model for simultaneous nuclear segmentation and classification, including tumor cells, connective cells, inflammatory cells, and other cell types36. The model is pre-trained on the PanNuke dataset and utilizes the horizontal and vertical distances of nuclear pixels to their center of mass to separate clustered units37. By inputting the WSI file, the model outputs the corresponding .json file, which contains all the information regarding nuclear segmentation and classification for each sample. The color of the nuclear boundary indicates the type of nucleus: green represents inflammatory cells, red represents tumor cells, and blue represents connective cells.
Cell-level feature extraction
Subsequently, we performed feature analysis on tumor cells, inflammatory cells, and stromal cells, which are the primary components of the tumor ecosystem. A pathologist delineated the ROI using ImageScope software (version 12.4.6) and generated the corresponding .xml file. The WSI files, .xml files, and the corresponding .json files generated during the segmentation step were then used as input. For each example, features were extracted using WSI Graph, resulting in a folder containing four .csv data files38. For each cell type (tumor cells, inflammatory cells, and stromal cells), three .csv files store the properties of all cells of that type, with each cell identified by a unique cell ID and its centroid coordinates. Another .csv file stored the edge information for the example, representing each edge by the connected cell IDs. Finally, we extracted 378 pathological features from each WSI. These features were categorized into three groups: (a) morphological features, describing the shape and contour of the nuclei; (b) texture features, characterizing the local pixel distribution patterns within the nuclear contour; and (c) topological features, depicting cell-cell relationships based on graph algorithm principles.
Visualization of the nuclear segmentation results and nuclear graph
Finally, based on the provided WSI files, the feature files generated from the feature extraction step, and the .xml files defining the ROI, the resulting visualization will be saved in the annotation file and can be viewed using ImageScope software.
In this process, we extracted 378 pathological features from each WSI.
Development of pathomics scores and prediction model
We conducted correlation analysis on the 427 extracted pathomic features, and those with significant correlations were further screened using LASSO (Least Absolute Shrinkage and Selection Operator) regression. Finally, a pathomics score was generated through the LASSO regression equation. In this study, two models were developed to predict G grades: the clinical model and the pathomics model. The clinical model was built using only routine pathological characteristics (for example: TBS, TNM). These parameters are mandatorily reported per the Chinese Society of Clinical Oncology (CSCO) protocols39, requiring no additional testing. The pathomics model was developed using all clinical model parameters and the pathomics score.
Cost-benefit analysis
Regarding diagnostic duration, we recorded the entire process time from the input of HE slides into the model to the output of G staging results, with separate timing for each patient. Meanwhile, we also sought the expertise of two pathologists (H.J. and YS.Z) who have been diagnosing pancreatic pathology for 20 and 10 years to review the pathology slides after gaining a comprehensive understanding of the clinical information. The evaluation was limited to mitotic figures and the Ki-67 index, and the final grade is based on whichever of the two proliferation indexes places the neoplasm in the higher-grade category4. Discordant cases (n = 15, 5.5%) underwent arbitration by a third expert to reach a final consensus. The entire evaluation process was completed under a microscope and documented the time each doctor spent on diagnosing each patient. No other pathological diagnostic steps were involved in this process.
In terms of diagnostic costs, the fee for HE slides is $64 per patient. An additional charge of $27 is required if pathologists need to perform G staging diagnoses in conjunction with Ki-67. Cost calculations for AI-based grading included H&E slide preparation ($64/patient) and cloud computational fees (maximum $2.10/patient using conservative AWS pricing estimates). Traditional grading costs included H&E preparation and Ki-67 IHC reagents/labor ($91/patient total). Pathologist salaries were excluded as fixed institutional costs. Conservative cloud cost estimates prevent underestimation concerns while maintaining 27% cost advantage over traditional methods.
Development of survival prediction models
In this study, we established two distinct prognostic models using the primary cohort. The Clinical-Grade Prognostic Model integrated clinicopathological features significantly associated with PFS and WHO grading, while the Pathomics Prognostic Model systematically replaced the WHO grading in CGPM with a pathomics-derived quantitative score.
Statistical analysis
Continuous variables were reported as median and range, qualitative variables as frequencies and percentages. Initially, differences between G1 and G2/3 groups were examined using Student’s t-test, Kruskal-Wallis H test, and chi-square test. Then, univariate regression analysis estimated effect sizes. A multivariate logistic regression model, using a stepwise method based on AIC, was developed for differential diagnosis and a nomogram constructed. Model performance was evaluated with ROC curves and AUC, compared using the DeLong test, and calibration assessed with bootstrapping and the Hosmer-Lemeshow test. DCA estimated clinical utility. Using the surv_cutpoint function, enter the dataset containing the patient pathomics score and the corresponding survival time (time-to-event) and disease progression, scan all possible cut-offs, and select the value leading to the maximum log-rank test (log-rank test) statistic as the best cut-off, stratifying patients into high- and low-risk groups. Survival differences were visualized with Kaplan-Meier curves and assessed with the log-rank test. Cost-benefit analysis used the paired T-test. A univariate Cox proportional hazards regression model was employed to identify potential prognostic factors for patients with PanNETs. A multivariate Cox proportional hazards regression model was subsequently used to develop a prediction for 1-, 2-, and 3-year Progression-Free Survival (PFS) for patients with PanNETs. The stepwise regression method, based on the Akaike information criterion, was used to identify the best-fitting model. To quantify the discrimination performance, the concordance index (C-index) and the 1-, 2-, and 3-year area under the receiver operating characteristic curve (AUROC) were calculated.
Statistical significance was set at a two-tailed p value of < 0.05. All analyses were performed using R software (version 3.3.3; R Foundation for Statistical Computing; https://www.r-project.org/).
Data availability
The original data are not publicly available, but are available from the corresponding author upon reasonable request.
Code availability
The prediction AI models and inference code were made publicly available on GitHub (https://github.com/CHANGHAI-AILab/NET).
References
Öberg, K., Knigge, U., Kwekkeboom, D., Perren, A. & Group, E. G. W. Neuroendocrine gastro-entero-pancreatic tumors: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann. Oncol. 23, vii124-vii130 (2012).
Dasari, A. et al. Trends in the Incidence, Prevalence, and Survival Outcomes in Patients With Neuroendocrine Tumors in the United States. JAMA Oncol. 3, 1335–1342 (2017).
Yao, J. C. et al. One hundred years after “carcinoid”: epidemiology of and prognostic factors for neuroendocrine tumors in 35,825 cases in the United States. J. Clin. Oncol. 26, 3063–3072 (2008).
Nagtegaal, I. D. et al. The 2019 WHO classification of tumours of the digestive system. Histopathology 76, 182–188 (2020).
Uccella, S. et al. Genomics of high-grade neuroendocrine neoplasms: well-differentiated neuroendocrine tumor with high-grade features (G3 NET) and Neuroendocrine Carcinomas (NEC) of various anatomic sites. Endocr. Pathol. 32, 192–210 (2021).
Yachida, S. et al. Small cell and large cell neuroendocrine carcinomas of the pancreas are genetically similar and distinct from well-differentiated pancreatic neuroendocrine tumors. Am. J. Surg. Pathol. 36, 173–184 (2012).
Simon, T. et al. DNA methylation reveals distinct cells of origin for pancreatic neuroendocrine carcinomas and pancreatic neuroendocrine tumors. Genome Med. 14, 24 (2022).
Sonbol, M. B. et al. Survival and incidence patterns of pancreatic neuroendocrine tumors over the last 2 decades: A SEER Database Analysis. Oncologist 27, 573–578 (2022).
Falconi, M. et al. ENETS Consensus Guidelines Update for the Management of Patients with Functional Pancreatic Neuroendocrine Tumors and Non-Functional Pancreatic Neuroendocrine Tumors. Neuroendocrinology 103, 153–171 (2016).
Wu, W. et al. The Chinese Guidelines for the Diagnosis and Treatment of Pancreatic Neuroendocrine Neoplasms (2020). Med. J. Peking. Union Med. Coll. Hosp.12, 460–480 (2021).
Akirov, A., Larouche, V., Alshehri, S., Asa, S. L. & Ezzat, S. Treatment options for pancreatic neuroendocrine tumors. Cancers 11 https://doi.org/10.3390/cancers11060828 (2019).
Hain, E. et al. Surgical management of pancreatic neuroendocrine tumors: an introduction. Expert Rev. Anticancer Ther. 19, 1089–1100 (2019).
Rindi, G. et al. A common classification framework for neuroendocrine neoplasms: an International Agency for Research on Cancer (IARC) and World Health Organization (WHO) expert consensus proposal. Mod. Pathol. 31, 1770–1786 (2018).
Schmitt, A. M. et al. WHO 2004 criteria and CK19 are reliable prognostic markers in pancreatic endocrine tumors. Am. J. Surg. Pathol. 31, 1677–1682 (2007).
Reid, M. D. et al. Calculation of the Ki67 index in pancreatic neuroendocrine tumors: a comparative analysis of four counting methodologies. Mod. Pathol. 28, 686–694 (2015).
Tang, L. H., Gonen, M., Hedvat, C., Modlin, I. M. & Klimstra, D. S. Objective quantification of the Ki67 proliferative index in neuroendocrine tumors of the gastroenteropancreatic system: a comparison of digital image analysis with manual methods. Am. J. Surg. Pathol. 36, 1761–1770 (2012).
Escobar Díaz Guerrero, R., Carvalho, L., Bocklitz, T., Popp, J. & Oliveira, J. L. Software tools and platforms in Digital Pathology: a review for clinicians and computer scientists. J. Pathol. Inf. 13, 100103 (2022).
Rodriguez, J. P. M. et al. Artificial intelligence as a tool for diagnosis in digital pathology whole slide images: A systematic review. J. Pathol. Inf. 13, 100138 (2022).
Dong, D. H. et al. Tumor burden score predicts tumor recurrence of non-functional pancreatic neuroendocrine tumors after curative resection. HPB 22, 1149–1157 (2020).
Capelli, P. et al. Endocrine neoplasms of the pancreas: pathologic and genetic features. Arch. Pathol. Lab. Med. 133, 350–364 (2009).
Salaria, S. N. & Shi, C. Pancreatic neuroendocrine tumors. Surg. Pathol. Clin. 9, 595–617 (2016).
Johnson, A. et al. Cadherin 17 is frequently expressed by ‘sclerosing variant’ pancreatic neuroendocrine tumour. Histopathology 66, 225–233 (2015).
Xue, Y. et al. Morphologic variants of pancreatic neuroendocrine tumors: clinicopathologic analysis and prognostic stratification. Endocr. Pathol. 31, 239–253 (2020).
Druce, M., Rockall, A. & Grossman, A. B. Fibrosis and carcinoid syndrome: from causation to future therapy. Nat. Rev. Endocrinol. 5, 276–283 (2009).
Ganss, R. Tumor stroma fosters neovascularization by recruitment of progenitor cells into the tumor bed. J. Cell Mol. Med 10, 857–865 (2006).
Cai, L. et al. Role of tumor-associated macrophages in the clinical course of Pancreatic Neuroendocrine Tumors (PanNETs). Clin. Cancer Res. 25, 2644–2655 (2019).
Takahashi, D. et al. Profiling the tumour immune microenvironment in pancreatic neuroendocrine neoplasms with multispectral imaging indicates distinct subpopulation characteristics concordant with WHO 2017 Classification. Sci. Rep. 8, 13166 (2018).
Wang, X. et al. Development and validation of a novel radiomics-based nomogram with machine learning to preoperatively predict histologic grade in pancreatic neuroendocrine tumors. Front. Oncol. 12. https://doi.org/10.3389/fonc.2022.843376 (2022).
Bian, Y. et al. Magnetic resonance imaging radiomic analysis can preoperatively predict G1 and G2/3 grades in patients with NF-pNETs. Abdom. Radio.46, 667–680 (2021).
Klimov, S. et al. Predicting Metastasis Risk in Pancreatic Neuroendocrine Tumors Using Deep Learning Image Analysis. Front Oncol. 10, 593211 (2020).
Bossuyt, P. M. et al. STARD 2015: An Updated List of Essential Items for Reporting Diagnostic Accuracy Studies. Clin. Chem. 61, 1446–1452 (2015).
Collins, G. S., Reitsma, J. B., Altman, D. G. & Moons, K. G. Transparent reporting of a multivariable prediction model for individual prognosis or diagnosis (TRIPOD): the TRIPOD statement. Bmj 350, g7594 (2015).
Chauhan, A. et al. Critical updates in neuroendocrine tumors: Version 9 American Joint Committee on Cancer staging system for gastroenteropancreatic neuroendocrine tumors. CA Cancer J. Clin. 74, 359–367 (2024).
Singhi, A. D. et al. Cystic pancreatic neuroendocrine tumors: a clinicopathologic study. Am. J. Surg. Pathol. 36, 1666–1673 (2012).
Chen L. C., P. G., Schroff F. Rethinking Atrous Convolution for Semantic Image Segmentation. arXiv e-prints, arXiv.1706.05587. https://doi.org/10.48550/arXiv.1706.05587 (2017).
Graham, S. et al. Hover-Net: Simultaneous segmentation and classification of nuclei in multi-tissue histology images. Med Image Anal. 58, 101563 (2019).
Gamper, J., A. Koohbanani. PanNuke Dataset Extension, Insights and Baselines. arXiv e-prints, arXiv:2003.10778. https://doi.org/10.48550/arXiv.2003.10778 (2020).
Zhao, S. et al. Single-cell morphological and topological atlas reveals the ecosystem diversity of human breast cancer. Nat. Commun. 14, 6796 (2023).
[Chinese expert consensus on gastroenteropancreatic neuroendocrine neoplasms (2022 edition)]. Zhonghua Zhong Liu Za Zhi 44, 1305–1329 (2022).
Acknowledgements
This work was supported in part by the National Science Foundation for Scientists of China (81871352, 82171915, 82171930, 82271972, 62402501 82202125 and 82371955), The Natural Science Foundation of Shanghai Science and Technology Innovation Action Plan (21ZR1478500 and 21Y11910300), China Postdoctoral Science Foundation Funded Project (2023M734270) and Clinical Research Plan of SHDC (SHDC2022CRD028). The funders played no role in study design, data collection, analysis and interpretation of data, or the writing of this manuscript.
Author information
Authors and Affiliations
Contributions
Y.L., C.C., Y.Z., Y.B. and H.J. designed the study. Y.L., C.C. and Y.Z. integrated and analyzed the data. J.Y., F.L. and T.W. collected the clinical data and analysis. Y.Z. and H.J. curated the pathological examinations. C.C. trained and developed the AI model. L.W., J.L., C.S. and H.J. obtained funding. C.C., X.F., Y.Z., H.J. and Y.B. revised the manuscript. Y.B. and H.J. supervised the study. All authors had full access to all the data. All authors were involved in drafting the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
C.W.C., Y.B., and C.W.S. are filing a patent for the prediction AI model in this study. All other authors declare they have no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, Y., Chen, C., Zhang, Y. et al. Development and validation of a pathomics model for accurate grading of pancreatic neuroendocrine tumors. npj Precis. Onc. 9, 235 (2025). https://doi.org/10.1038/s41698-025-01032-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41698-025-01032-2