Abstract
Mantle cell lymphoma (MCL) responds to frontline therapy but is susceptible to relapse. While Bruton’s tyrosine kinase inhibitors (BTKi) achieve high response rates, most patients eventually experience disease progression. Predicting responses to subsequent treatments remains challenging due to the lack of an established platform. Heterogeneity in gene alterations and cellular pathways contribute to resistance, complicating treatment approaches. Here, we present a multi-modal profiling platform, targeting key pathways rather than focusing on singular DNA-associated lesions. We identified dysregulated signaling pathways by performing gene expression profiling on 20 MCL samples using a custom MCL MATCH gene set and analyzed the data with gene-set variation analysis. We also screened 22 therapeutics in vitro to assess their efficacy. Whole exome sequencing was conducted to identify prevalent and actionable mutations linked to enriched pathways. Finally, we tested selected therapeutics in patient-derived xenograft mouse models to predict potential response in corresponding patients. Based on our integrative profiling, we identified the top therapeutic candidates for three patients in this study. This integrative platform may help identify targeted therapies for BTKi-relapsed/refractory MCL patients in clinical settings.
Similar content being viewed by others
Introduction
Mantle cell lymphoma (MCL) is an aggressive B-cell malignancy that remains incurable due to poor prognoses and frequent relapses following initial chemo-immunotherapy1. Targeted therapies such as Bruton’s tyrosine kinase inhibitors (BTKi), including ibrutinib, acalabrutinib, zanubrutinib, and pirtobrutinib, are commonly used as frontline treatments2,3 and are standard options for disease recurrence after chemo-immunotherapy4,5.
Despite the benefits of BTKi therapy, most MCL patients eventually relapse and face dismal outcomes6. After relapse, no reliable strategy exists to predict effective treatments, and current treatment decisions rely primarily on clinical factors such as age, metastatic stage, and histopathological subtype. This approach can lead to over-treatment or insufficient care for some patients. There is an urgent need for safer, more effective personalized therapies following BTKi therapy in the relapsed/refractory setting, or as alternative second-line treatments.
MCL is characterized by heterogeneous genomic alterations7, including a wide range in tumor mutational burden (TMB)8, dysregulated signaling pathways9 and a limited number of molecular targets. TP53 and ATM are frequently mutated and portend a poor prognosis10,11; however, there is not a viable strategy for targeting these genes. Gene expression signatures related to B-cell receptor (BCR) signaling in MCL cell lines are shown to identify effective targeted therapies12 and gene expression profiling in patients showed clustering related to BCR signaling and DNA damage repair pathways13.
Given that somatic mutations often lead to constitutive activation of signaling downstream of the affected gene14,15, we hypothesize that resistance-related DNA alterations drive disease progression through disrupted biological pathways. In this study, we aim to characterize cell signaling dysregulation using gene expression profiling (GEP) and pathway enrichment analysis in individual MCL patients and explore actionable targeted therapies based on dysregulated pathways rather than DNA-associated genetic lesions.
The NanoString nCounter® platform (NanoString, Seattle, WA) is FDA-approved for use in Clinical Laboratory Improvement Amendments (CLIA) compliant laboratories, signifying a use-case for treatment decisions in clinical trial settings16. We used the nCounter® platform with a curated MCL-specific gene panel (MCL MATCH) to identify differentially expressed genes (DEGs) and dysregulated pathways in silico with traditional statistical testing and algorithmic single-sample gene set variation analysis (GSVA)17. Correlative whole exome sequencing (WES) revealed specific mutations associated with these dysregulated pathways. We conducted in vitro drug screening, using 22 candidates targeting MCL-related pathways, in patient primary tumor cells and integrated molecular profiling data with drug response results to determine promising drug candidates. These candidates were further validated in vivo in corresponding patient-derived xenograft (PDX) mouse models mimicking clinical patient responses (Supplementary Fig. 1).
Results
Gene expression profiling provides pathway enrichment scores and differentially expressed genes
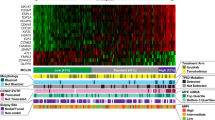
Our goal was to identify differences in GEP patterns and cellular pathways across patient samples (Supplementary Data 1). Using the GSVA algorithm, we visualized pathway enrichment patterns independent of BTKi response classification. These pathways represent key biological hallmarks of MCL and cancer, as well as potential therapeutic targets (Supplementary Data 2). Pathways with the highest GEP enrichment score were selected for correlative analyses and therapeutic exploration (Fig. 1). For each patient sample, the top three dysregulated pathways were identified based on GSVA enrichment score (ES) (Table 1).
Heatmap illustrating the ES of cellular pathways associated with the targets of in vitro drug panel across individual patient samples, calculated from gene expression values from the MCL MATCH gene set by GSVA.
All samples passed Nanostring quality control (Supplementary Fig. 2A–C). Multi-dimensional scaling (MDS) showed slight clustering based on tissue type and treatment status, but no major differences were observed related to sex or sample collection time points (baseline, during treatment or at recurrence) (Supplementary Fig. 2D). Samples were primarily purified CD19+/CD20+ B-cells, but immune cell profiling revealed differences in GEP-based immune signatures. Notably, BTKi responders showed enrichment of T cell- and other cytotoxic cell-associated pathways (Supplementary Fig. 3A).
Differential expression analysis identified 42 DEGs, with 27 genes significantly downregulated and 15 genes significantly upregulated in BTKi non-responders (MCL11-20) compared to responders (MCL1-10) (Fig. 2A, Supplementary Data 3). Hierarchical clustering of the DEGs and normalized expression levels among samples are shown in Fig. 2B.
A Volcano plot showing differentially expressed genes (DEGs) in non-responders to BTKi treatment in comparison to responders. DEGs with an adjusted p value \(\le\) 0.10 (-log10 = 1) and a log2 fold-change \(\ge\) 1 are labeled on the plot. B A heatmap displaying the normalized expression levels of DEGs between BTKi-responders and non-responders.
Many downregulated DEGs in BTKi non-responders are associated with immune function and may indicate an immune “cold” signature, suggesting decreased immune-cell infiltration and activity. These genes include STAT4, CCL5, NKG7, GRAP2, KLRB1, CD247, CD4, STING1 and GZMH. Among upregulated genes in non-responders were CCNE1 and CDK2, which encode cyclin-dependent kinases involved in cell-cycle regulation, suggesting a potential increase in tumor proliferation and chromosomal instability. Additionally, Notch ligand DLL3 was also upregulated.
To further analyze pathway differences between non-responders and responders, we used GSVA with the Canonical Pathway (C2:CP) gene set from MSigDB18 (Broad Institute, v 7.0). This served as a confirmative analysis alongside the matched therapeutics gene set. In non-responders, pathways involved in immune activation were downregulated, including the BioCarta T Helper Pathway, BioCarta T Cell Apoptosis Pathway, BioCarta Cytotoxic T Lymphocyte (CTL) Pathway, and Reactome immunoregulatory interactions (RUNX1 and FOXP3 development of Tregs) (Supplementary Fig. 3B). Pathways in the Tumor 360 panel related to cell-cycle regulation (Cell-cycle), metabolism (Lipid Metabolism) and DNA damage repair were upregulated in non-responders, suggesting potential mechanisms of resistance (Supplemental Fig. 3C).
Somatic mutations and copy number features related to BTKi resistance
Whole exome sequencing was performed on DNA from 20 MCL samples from patients with diverse clinical backgrounds, identifying a median of 28.5 (range 2–183) somatic, nonsynonymous mutations per sample. Many detected mutations were known MCL-associated alterations including TP53 (70%), NOTCH2 (40%), CCND1 (25%), NSD2 (20%), ATM (20%) and BIRC6 (20%). While TP53 mutations were slightly more common in non-responders (8:6), and NOTCH2 mutations occurred in five non-responders and in three responders, these differences were not statistically significant (Fig. 3).
Oncoplot depicts mutated genes detected by whole exome sequencing (WES) related to cellular pathways and targeted therapeutics with accompanying clinical annotations and the highest three Gene Set Variation Analysis (GSVA) results. Copy number alterations indicated in this plot were determined by clinical cytogenetics. Histotype was determined by clinical histopathology.
Mutations in GSVA pathways and clinico-pathologic patient characteristics are shown in Fig. 3. While 18 patients had at least one mutation in a cancer-related pathway, the low frequency of many pathway-associated variants highlights the somatic genetic heterogeneity of MCL.
We also detected multiple copy number (CN) aberrations in our cohort (Supplementary Fig. 4). Abnormal (3 or more CN aberrations) and complex karyotype (5 or more CN aberrations) are prognostic features in MCL and associated with poor response and survival. We observed a greater number of copy number aberrations in non-responders, and this was confirmed by clinical cytogenetics. Two responders had complex karyotype, whereas four non-responders had complex karyotype. Additionally, four non-responders were found to have a TP53 deletion or whole chromosome 17 deletions, and only one responder had a chromosome 17 deletion. Mosaic somatic loss of the X and Y chromosomes (mLOX and mLOY) are commonly reported in MCL and other hematological malignancies. Clinical cytogenetic correlation reported the phenomena in 50% of the non-responders and 20% of the responders.
In vivo matched validation in corresponding PDX models
Given the heterogeneity of somatic mutational profiles and the lack of established associations between specific mutations and therapeutic efficacy, we validated treatment candidates in three PDX models derived from patients 17, 18, and 20. Treatment candidates were selected based on in vitro screening and in silico GEP evidence.
We propose that GEP should precede the in vitro screen to narrow down potential therapies for preclinical testing. Our in silico drug matching approach utilized the GSVA algorithm to conduct single-sample pathway enrichment analyses with a curated MCL MATCH gene set representing targeted therapies that may be effective in relapsed/refractory MCL. By identifying potential therapeutic matches for BTKi non-responders, this approach may help guide alternative treatment strategies in cases of relapse or resistance to BTK-targeted therapies.
Cell cycle-targeting agents and CAR T-cell therapy demonstrates potent activity in MCL17-PDX with cell cycle dysregulation and enhanced T-cell co-stimulation
The BTKi-relapsed sample from MCL patient 17 (MCL17) had a frame-shift deletion in TP53 and a nonsense mutation in NOTCH2 (Fig. 3). The sample was deemed to have a pleomorphic histotype. CN analysis showed multiple deletions and amplifications throughout the genome. However, there was no clinical cytogenetics to validate this apparent complex karyotype. GEP and GSVA in silico pathway analysis identified high scores in PLK1, cyclin-dependent kinases (CDKs) and CAR T related pathways (Fig. 1, Table 1).
MCL17 showed relative resistance to all drugs tested in vitro, with a maximum inhibition rate (MIR) of less than 80% (Fig. 4A). Therefore, we prioritized pathway ES over in vitro cytotoxicity when considering therapeutic options for this patient, leading to the identification of PLK1 and CDK as potential therapeutic targets. FDA-approved drugs and those in advanced clinical trials were given precedence. Accordingly, abemaciclib, a CDK4/6 inhibitor that completed phase II trial for relapsed/refractory MCL19, and volasertib, which has undergone a phase I study in leukemia20, were selected for in vivo validation.
A In vitro cell viability assay of the designed drug panel on MCL17 primary cells after 24 h treatment. Each bar is the maximum inhibition rate for each agent. B–D MCL17-PDX mice were treated as indicated: vehicle (n = 5), abemaciclib (50 mg/kg, oral, daily; n = 5), and volasertib (5 mg/kg, intraperitoneal, weekly; n = 5). Tumor size was measured every 7 days during the treatment. Tumor volume = length x width2/2. Mice were euthanized after 21 days of treatment. Tumors were excised and weighed in comparison between each group. E Blood β2M tested on mouse plasma was plotted to monitor the tumor growth. F Western blot assay of 24 h treatment with abemaciclib on isolated tumor cells from vehicle-treated mice. G H&E and IHC staining of representative mouse tumor tissues dissected at the endpoint. Results were considered statistically significant at p < 0.05 (*); p < 0.01 (**); p < 0.001 (***), and p < 0.0001 (****).
To establish MCL17-PDX, we initially transplanted tumor cells into the subrenal capsule of first-generation mice and passaged subcutaneously in subsequent generations. Treatment with abemaciclib or volasertib significantly inhibited tumor growth (Fig. 4B). Both drugs reduced tumor size and prognostic indicator β2 microglobulin (β2M) serum levels at the study endpoint (Fig. 4C–E).
Immunohistochemical (IHC) analysis of abemaciclib-treated tumors confirmed cell cycle arrest and CDK inhibition, as shown by decreased p-Rb and TOPOIIα compared to vehicle-treated tumors (Fig. 4F). These findings were consistent with ex vivo treatment of isolated tumor cells with abemaciclib (Fig. 4G).
In addition, GEP showed high T-cell co-stimulation activity in both DEG and GSVA analyses, aligning with MCL17’s elevated CAR T pathway score (Fig. 1). To substantiate this, we established an orthotopic MCL17-PDX model to better assess CAR T-cell efficacy against metastasis-associated MCL features.
By intravenously injecting purified MCL17-PDX tumor cells into NSG mice, we generated a MCL17-PDX model that recapitulated the patient’s splenomegaly and hepatomegaly. To evaluate CAR T-cell therapy as a potential treatment, anti-CD19 CAR T cells were administered at a 3:1 ratio to tumor cells on day 4 post-xenografting. The CAR T cell-treated mice showed significant reductions in liver and spleen weight, tumor burden (Fig. 5A–D) and tumor involvement in bone marrow and blood compared with vehicle mice (Fig. 5E).
A, B CAR T cells were injected intravenously at a ratio of 3:1 to tumor cells on day 4 following tumor xenograft. All treated mice were euthanized on day 33, and spleen and liver were dissected and measured at the endpoint. C–E Tumor percentages in the spleen, liver, bone marrow, and blood were determined by detecting CD20+ and HLA+ populations using flow cytometry. F Body weights were measured and plotted during the treatment period. Results were considered statistically significant at p < 0.05 (*); p < 0.01 (**); p < 0.001 (***), and p < 0.0001 (****).
Neither group experienced weight loss during the first 20 days, though vehicle-treated mice began losing weight thereafter, likely due to tumor progression (Fig. 5F). These in vivo results support CAR T-cell therapy as a promising treatment option for MCL17.
The BCL-2 antagonist venetoclax demonstrates strong efficacy in MCL18-PDX with high expression of BCL-2 family genes
MCL patient 18 (MCL18) had lower CCND1 expression. Multiple somatic mutations were identified including TP53, MYC, NOTCH2, and NRAS along with a high TMB (Fig. 3). Clinical cytogenetics revealed an abnormal karyotype with loss of one X chromosome, which was confirmed by next-generation sequencing (NGS). GEP revealed the highest pathway enrichment in PLK1, CDK and Proteasome pathways utilizing the MATCH gene set (Table 1). Notably, individual expression levels of BCL2, BCL2A1 and BCL2L1 remained significantly upregulated, indicating a strong anti-apoptotic activity in these tumor cells (Supplementary Data 4).
In vitro screening showed MCL18 responded to 7 of 22 drugs, with MIRs ≥80% in these cases. Remarkably, the BCL-2 inhibitor venetoclax induced substantial tumor cell death (Fig. 6A). Given the observed upregulation of anti-apoptotic BCL-2 family members, we prioritized targeting these proteins for therapeutic intervention.
A In vitro cell viability assay of the designed drug panel on MCL18 primary patient cells. Each bar is the maximum inhibition rate for each agent. B Cell viability assay and apoptosis assay after 24 h treatment of BCL-2 inhibitor venetoclax, MCL-1 inhibitors S63845 and AZD5991 on MCL18-PDX isolated tumor cells ex vivo. C MCL18-PDX mice were treated as indicated: vehicle (n = 5) and venetoclax (50 mg/kg, oral, daily; n = 5). Tumor size was measured every ten days during the treatment. Tumor volume = length x width2/2. Mice were euthanized when one dimension of tumor reached 15 mm. Kaplan-Meier survival curve of PDX tumor–bearing mice from treatment start time in the treatment groups. Blood β2M tested from mouse plasma was plotted to monitor the tumor growth. D Tumor cells were isolated at the endpoint from each mouse. Cell viability was tested using the Trypan Blue dye exclusion method. Results were considered statistically significant at p < 0.05 (*); p < 0.01 (**); p < 0.001 (***), and p < 0.0001 (****).
To further assess efficacy of drugs targeting the BCL-2 family, MCL18-PDX was established by subcutaneous inoculation and isolated tumor cells were subsequently treated ex vivo with venetoclax and two MCL-1 inhibitors, S63845 and AZD5991. Venetoclax significantly reduced tumor cell viability and induced apoptosis more effectively than MCL-1 inhibitors (Fig. 6B), reinforcing its potential for in vivo validation.
MCL18-PDX tumor cells were passaged subcutaneously to next-generation mice, and treatment began once tumors became palpable. In vehicle-treated mice, tumors grew rapidly requiring euthanasia at a median survival of 32 days. In contrast, venetoclax completely abrogated tumor growth and eliminated β2M levels for up to 35 days without dramatically impacting body weight (Fig. 6C). Treatment was discontinued once all vehicle mice had succumbed, and venetoclax-treated mice were monitored for survival. Venetoclax significantly extended mouse survival, with a median of 92 days. At the study endpoint, tumor cells were isolated and assessed for viability using Trypan Blue staining. Venetoclax-treated cells showed markedly lower cell viability due to apoptosis induction (Fig. 6D). Together, these results support venetoclax as a potential therapy for MCL18.
The pan-PI3K inhibitor copanlisib suppresses tumor growth in MCL20-PDX with highest ES in proteasome, cell cycle, and thioredoxin reductase pathways
MCL patient 20 (MCL20) harbored somatic mutations in TP53, NOTCH2, and CCND1 (Fig. 3), indicating multiple hallmarks of tumorigenesis. The sample was assessed as having a classical histotype. Clinical cytogenetics and NGS indicated an abnormal karyotype, with deletion of chromosomes X and 17 (TP53) (Supplementary Fig. 4). GEP analysis identified the most dysregulated pathways as Proteasome, CDK and Thioredoxin Reductase (Table 1).
MCL20 demonstrated strong in vitro responses with nine drugs exceeding an 80% MIR threshold. Among them, PI3Kα inhibitor PIK75 exerted the most potent tumor-suppressive effects (Fig. 7A). Given that the PI3K pathway may interact with and enhance NF-κB21, and considering PI3K’s critical role in stimulating DNA synthesis and cell cycle progression22, we prioritized PI3K-targeted therapeutics for further investigation.
A In vitro cell viability of the designed drug panel on MCL20 primary patient cells. Each bar is the maximum inhibition rate for each agent. B Cell viability assay of copanlisib, vistusertib, and MK2206 in MCL20-PDX isolated tumor cells. C–E MCL20-PDX mice were treated with vehicle or copanlisib (10 mg/kg, intraperitoneal, every other day; n = 5) and mice were euthanized after 40 days of treatment. Spleen and liver were excised and measured at the endpoint, as well as the bone marrow involvement and serum β2M level. F Cells from vehicle-treated mice were treated with serial concentrations of copanlisib for 1 h and blotted for Akt signaling. G IHC staining for p-Akt on representative dissected tumors. Results were considered statistically significant at p < 0.05 (*); p < 0.01 (**); p < 0.001 (***), and p < 0.0001 (****).
An MCL20-PDX mouse model was established through intravenous injection, and PDX tumor cells were subsequently isolated for an ex vivo screen. Instead of PIK75, we selected the pan-PI3K inhibitor copanlisib, which has advanced to a phase I clinical trial in relapsed/refractory MCL, as well as the mTOR inhibitor vistusertib and the Akt inhibitor MK2206 for the testing. Compared to vistusertib and MK2206, copanlisib significantly reduced tumor cell viability at low nanomolar concentrations, supporting its potential for further in vivo evaluation (Fig. 7B).
The MCL20-PDX model mirrored the patient’s splenomegaly and hepatomegaly. Consistent with ex vivo findings, copanlisib effectively reduced spleen and liver size and decreased bone marrow involvement (Fig. 7C, D). Serum β2M levels were also decreased in copanlisib-treated mice compared to the vehicle-treated mice (Fig. 7E). We performed IHC staining for Akt phosphorylation on tumor tissues. Copanlisib-treated tumors showed a marked reduction in phosphorylated Akt levels, which was further confirmed by Western blot analysis and IHC staining (Fig. 7F, G). These results provide compelling evidence that copanlisib has potential as an appropriate therapeutic option for MCL20.
Discussion
We observed TP53, NOTCH2, NSD2, CCND1 and ATM somatic mutations in our MCL cohort. While TP53 and ATM have established prognostic significance5,7, no targeted therapies have been proven effective against these or other mutations. We also observed heterogeneous mutations related to signaling pathways within gene sets in the MCL MATCH panel but were unable to provide statistically significant results because of the heterogeneity of our small sample size. This limitation highlights the need to explore alternative therapeutic strategies based on transcriptomic data and in vitro drug screening. By analyzing gene expression dynamics, we can better understand the downstream effects of MCL’s heterogeneous mutational landscape.
Additionally, we observed increased copy number aberrations in BTKi resistant patients that has been linked to poor outcomes23,24,25. Our cohort was too small to formally assess the impact of these copy number features. Higher TMB has been associated with poorer overall survival but a positive response to immunotherapies in diffuse large B-cell lymphoma (DLBCL)26. There is some evidence to suggest that increased somatic mutations are indicative of aggressive MCL25, but more research is needed to understand TMB and therapeutic response.
To identify biologically relevant pathways at an individual level, we applied the GSVA algorithm, which evaluates GEP in a sample-specific manner rather than through group-based DEG analysis. This approach allows us to detect key pathways at baseline or throughout disease progression in individual patients, potentially predicting treatment response.
To illustrate the feasibility of our approach, we performed the proposed integrative analysis to guide therapeutic selection in three patients: MCL17, MCL18 and MCL20. MCL17’s tumor cells exhibited in vitro resistance to most targeted therapies, so we prioritized GEP signaling ES to propose the potential sensitivity to the PLK1 inhibitor volasertib and the CDK4/6 inhibitor abemaciclib, which was successfully validated in vivo. Retrospectively, this finding did not contradict the in vitro screen results, as in vitro proliferation assays do not always accurately reflect in vivo tumor biology, particularly because primary patient cells often show poor proliferation, a characteristic that can render them insensitive to cell cycle-targeting drugs.
Gene expression analysis of MCL18 revealed upregulation of anti-apoptotic protein-related genes, including BCL2, BCL2A1, and BCL2L1, and identified the highest efficacy of venetoclax in the in vitro drug screen, supporting BCL-2 targeting as a viable approach for this patient. Regarding MCL20, proteasome and CDK pathways were the most dysregulated; however, the PI3Kα inhibitor PIK75 emerged as the most potent agent in vitro. Considering PI3K’s role in NF-κB/proteasome signaling and cell cycle progression, the pan-PI3K inhibitor was finally selected and validated for this patient.
Interestingly, BTKi non-responders exhibited enrichment in immune “cold” signatures, characterized by downregulated immune-associated pathways, consistent with previous reports27,28,29,30. BTKi therapy may deplete the tumor microenvironment of immune cells31, potentially impacting treatment response. CAR T- and NK-cell therapies offer alternative strategies that could bypass these immune microenvironment limitations32. Additionally, several genes linked to key cancer hallmarks33, including DNA repair, cell-cycle regulation, cytogenetic instability and metabolic processes, were differentially expressed between responders and non-responders. These pathways present viable targets for both existing and emerging therapeutics.
A limitation of our study is the small, heterogeneous sample size. Samples were collected at various clinical stages (naïve, during treatment, or at relapse), which may influence mutational profiles and gene expression patterns. Additionally, our preclinical screening focused solely on monotherapies, whereas evaluating combination therapies could potentially enhance treatment efficacy. Despite these limitations, we anticipate that our early findings will contribute to the development of a more precise treatment-matching algorithm. As we expand our PDX models and drug screening to include more patients, broader statistical analyses with help refine our approach and improve clinical relevance.
PDX are clinically relevant models for precision therapy matching, allowing assessment of drug response and potential toxicities while providing insight into compound behavior in biological systems. However, PDX models have limitations, including differences in tumor microenvironments between patient tumors and their corresponding PDX models. Additionally, they cannot fully predict individual patient responses or adverse events. This highlights the importance of incorporating oncologists’ domain knowledge and engaging in shared decision making with patients.
Integrating multi-omics with clinical response data can enhance prognostic modeling using classical statistical or machine learning approaches, as demonstrated in our previous study25. These techniques could be useful in future evaluation of combination therapies and forecasting adverse events34. Further integration of germline genomics, metabolomics, epigenomics and clinical-exposome features could provide a more comprehensive understanding of therapeutic responses.
Our strategy to combine in vitro drug screening, genomics and transcriptomics identified potential precision therapeutics by elucidating the complex systems biology underlying MCL physiopathology. Our first-in-class pilot study supports the value of multimodality platforms, particularly for MCL patients who relapse or progress after BTKi therapy. Building on this work, the MCL MATCH trial (NCT04872413) was initiated at MD Anderson Cancer Center.
Methods
Patient samples
Patient samples were acquired from patients after obtaining written informed consent following the University of Texas MD Anderson Cancer Center Institutional Review Board-approved protocols and in accordance with the Declaration of Helsinki. We collected MCL patient samples (n = 20, Supplementary Data 1) from clinical apheresis, excisional biopsies, bone marrow, and peripheral blood. These samples included both treatment-naïve cases and those obtained during treatment or relapse. Germline DNA from saliva were available for 17 patients. DNA and RNA extraction followed standard protocols at the MD Anderson Core Facility. All samples were collected from patients with or without prior treatment with covalent BTK inhibitors at the time of study enrollment. The BTKi responses reported in this study refer exclusively to covalent BTKi, and response evaluation was conducted according to the Lugano classification criteria35.
Whole exome sequencing (WES) and data processing
Genomic DNA was sheared, and libraries were prepared, sequenced and assessed for quality control (for specifications, please see the supplementary methods). Reads were aligned to the hg38 reference genome and resulting BAM files were merged, deduplicated and sorted prior to variant calling. Somatic nonsynonymous point mutations and small insertions and deletions were called with Mutect2 (GATK, Broad Institute, Boston, MA, v4.4.0.0) with matched germline DNA available for 17 samples. For three samples without matched germline DNA, a panel of normal constructed built from the 1000 Genomes36 and gnomAD37 was used to exclude likely germline variants. Mutect2 calls labeled with “KEEP” were retained for further analysis and were filtered using GATK (Broad Institute, Boston, MA, v4.4.0.0) and annotated with Funcotator generating VCF and MAF files. A multi-tiered approach was used to identify high-confidence somatic variants.
Copy number was analyzed by creating a pooled normal coverage file from the cohort’s germline BAM files and utilizing package CNVkit38 which implements binned coverage ratios and the circular binary segmentation algorithm to estimate copy number and identify amplifications and deletions. Orthogonal clinical comparison was done by data extraction from clinical records resulting from fluorescent in-situ hybridization (FISH), comparative genomic hybridization (CGH) and other cytogenetic methods.
Gene expression profiling (GEP), differentially expressed gene (DEG) and pathway analyses
We used the NanoString Tumor Signaling 360™ panel (760 genes across 40+ tumor-related pathways) and added 14 MCL-relevant B-cell genes with associated pathways, totaling 774 genes plus 20 reference genes (Supplementary Data 5–6). RNA samples were prepared per NanoString protocols. Quality control, normalization, DEG, and pathway analyses were performed using nSolver (RUO) and the ROSALIND cloud platform. Additional details and QC visuals are in the supplementary methods and Supplementary Figs. 2A–C.
DEGs comparing BTKi responders and non-responders are listed in Supplementary Data 7. Log2 fold-changes (lfc) were calculated by subtracting the arithmetic mean expression of responders from that of non-responders for each gene (Supplementary Data 4). Statistical significance was assessed using Welch’s t-test for unequal variances, with multiple testing corrections applied using the Benjamini-Yekutieli False Discovery Rate (FDR)39 method. DEGs were reported based on adjusted p-values (q) \(\le \,\)0.10 and lfc of 1 corresponding to a fold-change of 2. DEGs were visualized with the Enhanced Volcano40 and Complex Heatmap41 packages.
Gene set variation analysis (GSVA) and in silico drug matching
We constructed the MCL MATCH gene set based on pathways targeted by therapies tested in our in vitro cell viability assay and PDX models (Supplementary Data 5). These pathways were sourced from Hallmark42, KEGG43, Biocarta, and the Tumor Signaling 360TM gene sets. The CAR T pathway was defined by the expression of T-cell receptor genes and co-stimulatory markers44.
Gene Set Variation Analysis (GSVA) was performed using the GSVA R package17 according to the methodology of Hänzelmann et al.11. The sample-wise algorithm applied a matrix of normalized expression values to predefined gene sets, generating pathway enrichment scores. Pathways were visualized using the pheatmap R package, and differential enrichment was assessed with GSVA and the limma package. Linear models with empirical Bayes moderation identified significantly enriched pathways (adjusted p ≤ 0.10). For each sample, the top three MCL MATCH pathways were ranked by enrichment score (Table 1), with additional pathway scores from the original Tumor 360 panel shown for comparison (Supplementary Fig. 3C).
Reagents
All drugs were purchased from MedChemExpress (NJ, USA) and solubilized in dimethyl sulfoxide (DMSO) at an indicated dose for in vitro studies. CD19 directed CAR T cells were manufactured in-house. More information, including FDA approval status and use in MCL, pertaining to the therapies chosen is shown in Supplementary Data 8.
Cell viability assay
Purified MCL cells (CD19+/CD20+ >95%) were cultured in RPMI-1640 (Life Technologies, Grand Island, NY) supplemented with 10% heat-inactivated fetal bovine serum, penicillin (10,000 units/mL; Sigma), streptomycin (10 mg/mL; Sigma). Cells were seeded in 384-well plates at a density of 5 × 104 cells/well and treated with serial concentrations of indicated drugs for 24 h. Cytotoxicity was assessed by the CellTiter-Glo® Luminescent Assay (Promega). Data are expressed as a percentage of DMSO control values.
Histopathological analysis
Fresh excised PDX tumor tissues were fixed in 10% neutral-buffered formalin and paraffin-embedded, sectioned (5 μm), and stained with H&E and indicated antibodies as previously described. CD20 was detected with human anti-CD20 (Dako, CA, USA) and visualized with a ZEISS microscope.
Western blot analysis
Western blotting was performed as described previously45. β-actin was obtained from Santa Cruz Biotechnology (TX, USA), all other antibodies were obtained from Cell Signaling Technology (MA, USA). Film scans were shown in Supplementary Fig. 5.
In vivo PDX models
The Institutional Animal Care and Use Committee of The University of Texas MD Anderson Cancer Center approved all the experimental protocols and procedures. The NOD SCID gamma (NSG) mice (Jackson Laboratory, ME, USA) were housed and monitored in the animal research facility at MD Anderson Cancer Center. Patient-derived xenograft (PDX) models were established by inoculating freshly isolated MCL cells subcutaneously, intravenously or into subrenal capsules of 6- to 8-week-old NSG mice. Subrenal capsule implantation was performed under anesthesia using a 5% isoflurane vaporizer, whereas subcutaneous and intravenous implantations were conducted without anesthesia. For in vivo efficacy studies, mice with established tumors were randomly assigned to treatment groups (5 mice per cage) once tumors became palpable. The treatment routes and dosages for each drug candidate are provided in the corresponding figure legends. Tumor size was measured once tumor was palpable and circulating human β2 microglobulin (β2M) levels in mouse serum were used to monitor tumor growth using the β2m ELISA kit (Abnova, Walnut, CA). All mice were euthanized by carbon dioxide inhalation followed by cervical dislocation upon reaching the humane endpoint. Tumor masses were isolated, as well as the spleen and liver if hepatosplenomegaly was observed.
Additional statistical analyses
Cell viability assays were performed in triplicate and reported as mean ± SEM or SD. Statistical analyses were conducted in R (v4.2.2), with methods detailed in figure legends or relevant sections. Normality was assessed using the Shapiro-Wilk test. For continuous variables, t tests or Kruskal-Wallis tests were used based on data distribution; Fisher’s Exact test was used for categorical variables. P values were adjusted for multiple testing using the Benjamini-Hochberg method (FDR). Two-sided tests were used, with p ≤ 0.05 and adjusted p ≤ 0.10 considered significant unless otherwise noted.
Data availability
The datasets generated and/or analyzed during the current study are not publicly available due to patient privacy concerns but are available from the corresponding author on reasonable request.
Code availability
The analysis code to generate the figures is available upon request from the corresponding author.
References
Jain, P. & Wang, M. Mantle cell lymphoma: 2019 update on the diagnosis, pathogenesis, prognostication, and management. Am. J. Hematol. 94, 710–725 (2019).
Wang, M. L. et al. Ibrutinib–rituximab followed by R-HCVAD as frontline treatment for young patients (≤65 years) with mantle cell lymphoma (WINDOW-1): a single-arm, phase 2 trial. Lancet Oncol. 23, 406–415 (2022).
Dreyling, M. et al. Ibrutinib combined with immunochemotherapy with or without autologous stem-cell transplantation versus immunochemotherapy and autologous stem-cell transplantation in previously untreated patients with mantle cell lymphoma (TRIANGLE): a three-arm, randomised, open-label, phase 3 superiority trial of the European Mantle Cell Lymphoma Network. Lancet 403, 2293–2306 (2024).
Song, Y. et al. Zanubrutinib in relapsed/refractory mantle cell lymphoma: long-term efficacy and safety results from a phase 2 study. Blood 139, 3148–3158 (2022).
Cheah, C. et al. Pirtobrutinib in covalent BTK-inhibitor pre-treated mantle cell lymphoma: Updated results, subgroup analysis from bruin with >3 years follow-up from start of enrollment. Hematol. Oncol. 41, 153–154 (2023).
Rule, S. et al. Ibrutinib for the treatment of relapsed/refractory mantle cell lymphoma: extended 3.5-year follow up from a pooled analysis. Haematologica 104, e211–e214 (2019).
Hill, H. A. et al. Genetic mutations and features of mantle cell lymphoma: a systematic review and meta-analysis. Blood Adv. 4, 2927–2938 (2020).
Jeong, S. et al. Genetic heterogeneity and prognostic impact of recurrent ANK2 and TP53 mutations in mantle cell lymphoma: A multi-centre cohort study. Sci. Rep. 10, 13359 (2020).
Jares, P., Colomer, D. & Campo, E. Molecular pathogenesis of mantle cell lymphoma. J. Clin. Invest 122, 3416–3423 (2012).
Eskelund, C. W. et al. TP53 mutations identify younger mantle cell lymphoma patients who do not benefit from intensive chemoimmunotherapy. Blood 130, 1903–1910 (2017).
Koff, J. L. et al. Mutations of ATM confer a risk of inferior survival in patients with TP53-wild type Mantle cell lymphoma. Blood 140, 3500–3503 (2022).
D’Agaro, T. et al. A B-cell receptor-related gene signature predicts response to ibrutinib treatment in mantle cell lymphoma cell lines. Haematologica 104, e410–e414 (2019).
Yi, S. et al. Genomic and transcriptomic profiling reveals distinct molecular subsets associated with outcomes in mantle cell lymphoma. J. Clin. Investig. 132, 1–8 (2022).
Fernàndez, V., Hartmann, E., Ott, G., Campo, E. & Rosenwald, A. Pathogenesis of mantle-cell lymphoma: All oncogenic roads lead to dysregulation of cell cycle and DNA damage response pathways. J. Clin. Oncol. 23, 6364–6369 (2005).
Krysiak, K. et al. Recurrent somatic mutations affecting B-cell receptor signaling pathway genes in follicular lymphoma. Blood 129, 473–483 (2017).
Morris, J. S. et al. Development and validation of a gene signature classifier for consensus molecular subtyping of colorectal carcinoma in a CLIA-certified setting. Clin. Cancer Res. 27, 120–130 (2021).
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: Gene set variation analysis for microarray and RNA-Seq data. BMC Bioinforma. 14, 7 (2013).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Morschhauser, F. et al. Clinical activity of abemaciclib in patients with relapsed or refractory mantle cell lymphoma - a phase II study. Haematologica 106, 859–862 (2021).
Cortes, J. et al. Phase 1 dose escalation trial of volasertib in combination with decitabine in patients with acute myeloid leukemia. Int J. Hematol. 113, 92–99 (2021).
Bai, D., Ueno, L. & Vogt, P. K. Akt-mediated regulation of NFkappaB and the essentialness of NFkappaB for the oncogenicity of PI3K and Akt. Int. J. Cancer 125, 2863–2870 (2009).
Wang, G., et al. PI3K stimulates DNA synthesis and cell-cycle progression via its p55PIK regulatory subunit interaction with PCNA. Mol. Cancer Ther. 12, 2100–2109 (2013).
Sarkozy, C. et al. Complex karyotype in mantle cell lymphoma is a strong prognostic factor for the time to treatment and overall survival, independent of the MCL international prognostic index. Genes Chromosomes Cancer 53, 106–116 (2014).
Khouja, M. et al. Comprehensive genetic analysis by targeted sequencing identifies risk factors and predicts patient outcome in Mantle Cell Lymphoma: Results from the EU-MCL network trials. Leukemia 38, 2675–2684 (2024).
Hill, H. A. et al. Integrative prognostic machine-learning models in Mantle cell lymphoma. Cancer Res. Commun. 3, 1435–1446 (2023).
Chen, C. et al. Tumor mutation burden estimated by a 69-gene-panel is associated with overall survival in patients with diffuse large B-cell lymphoma. Exp. Hematol. Oncol. 10, 20 (2021).
Nomie, K. et al. Tumor microenvironment molecular signatures that define therapeutic resistance in Mantle cell lymphoma. Blood 134, 2762–2762 (2019).
Nomie, K. et al. Multi-omics analysis of mantle cell lymphoma reveals an immune-cold tumor microenvironment associated with ibrutinib resistance. J. Clin. Oncol. 38, 8055–8055 (2020).
Zhang, W. et al. Identification of immune-related genes and development of a prognostic model in mantle cell lymphoma. Ann. Transl. Med. 10, 1323 (2022).
Jain, P. et al. Immune-depleted tumor microenvironment is associated with poor outcomes and BTK inhibitor resistance in mantle cell lymphoma. Blood Cancer J. 13, 156 (2023).
Niemann, C. U. et al. Disruption of in vivo chronic lymphocytic leukemia tumor-microenvironment interactions by ibrutinib-findings from an investigator-initiated phase II study. Clin. Cancer Res 22, 1572–1582 (2016).
Liu, Y. T. & Sun, Z. J. Turning cold tumors into hot tumors by improving T-cell infiltration. Theranostics 11, 5365–5386 (2021).
Hanahan, D. Hallmarks of cancer: New dimensions. Cancer Discov. 12, 31–46 (2022).
Abd El-Hafeez, T., Shams, M. Y., Elshaier, Y. A. M. M., Farghaly, H. M. & Hassanien, A. E. Harnessing machine learning to find synergistic combinations for FDA-approved cancer drugs. Sci. Rep. 14, 2428 (2024).
Cheson, B. D. et al. Recommendations for initial evaluation, staging, and response assessment of Hodgkin and non-Hodgkin lymphoma: The Lugano classification. J. Clin. Oncol. 32, 3059–3068 (2014).
Abecasis, G. R. et al. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Chen, S. et al. A genome-wide mutational constraint map quantified from variation in 76,156 human genomes. bioRxiv 2022: 2022.2003.2020.485034.
Talevich, E., Shain, A. H., Botton, T. & Bastian, B. C. CNVkit: Genome-Wide Copy Number Detection and Visualization from Targeted DNA Sequencing. PLOS Comput. Biol. 12, e1004873 (2016).
Benjamini Y., Yekutieli D. The control of the false discovery rate in multiple testing under dependency. Annals Stat. 29, 1165–1188 (2001).
Blighe, K., Rana, S. & Lewis, M. EnhancedVolcano: Publication-ready volcano plots with enhanced colouring and labeling. R. Package version 1, 10–18129 (2019).
Gu, Z., Eils, R. & Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32, 2847–2849 (2016).
Liberzon, A. et al. The molecular signatures database hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Kanehisa, M., Sato, Y., Kawashima, M., Furumichi, M. & Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 44, D457–D462 (2015).
Chen, L. & Flies, D. B. Molecular mechanisms of T cell co-stimulation and co-inhibition. Nat. Rev. Immunol. 13, 227–242 (2013).
Li, Y. et al. Potentiation of apoptosis in drug-resistant mantle cell lymphoma cells by MCL-1 inhibitor involves downregulation of inhibitor of apoptosis proteins. Cell Death Dis. 14, 714 (2023).
Acknowledgements
This project was supported by NIH/NCI Grant R21CA202104. HAH was supported by NIH/NCI Grant F99CA274686 and K00CA274686. Sequencing for this study was supported by the University of Texas MD Anderson Cancer Center Advanced Technology Genomics Core NIH/NCI Grant CA016672(ATGC). We thank Tracey Baas for critical reading and technical editing of the manuscript.
Author information
Authors and Affiliations
Contributions
M.W. and Y.L. conceived and designed the study, Y.L., Y.L., and J.M. performed the in vitro experiments, Y.L. established the PDX models and conducted the in vivo study, H.A.H. conducted the bioinformatic analyses, Y.L. and H.A.H. performed data analysis and wrote the manuscript, V.J., F.Y., Y.Y., Y.F., J.Z., L.Q., J.Y., P.J., K.C., and M.W. edited and reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
M.W. is consultant to AbbVie, Acerta Pharma, AstraZeneca, BeiGene, BioInvent, Deciphera, InnoCare, Janssen, Kite Pharma, Leukemia & Lymphoma Society, Lilly, Merck, Milken Institute, Oncternal, Parexel, Pepromene Bio, Pharmacyclics, and VelosBio, and he has received research support from Acerta Pharma, AstraZeneca, BeiGene, BioInvent, Celgene, Genmab, Genentech, Innocare, Janssen, Juno Therapeutics, Kite Pharma, Lilly, Loxo Oncology, Molecular Templates, Oncternal, Pharmacyclics, VelosBio, Vincerx. M.W. also received a speaker honorarium from AbbVie, Acerta Pharma, AstraZeneca, BeiGene, BioInvent, Dava Oncology, Eastern Virginia Medical School, IDEOlogy Health, Janssen, Kite Pharma, Leukemia & Lymphoma Society, LLC TS Oncology, Medscape, Meeting Minds Experts, MJH Life Sciences, Merck, Moffit Cancer Center, Oncology Specialty Group, OncLive, Pharmacyclics, Physicians Education Resources (PER), Practice Point Communications (PPC), and Studio ER Congressi. The remaining authors have no conflicts of interest to declare.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, Y., Hill, H.A., Li, Y. et al. Integrative profiling strategies to guide personalized therapy in mantle cell lymphoma: a pilot study. npj Precis. Onc. 9, 373 (2025). https://doi.org/10.1038/s41698-025-01158-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41698-025-01158-3