Abstract
Brain MRI is the primary imaging modality for pediatric brain tumors, yet incomplete acquisitions are common, hindering the clinical utility of existing deep learning models for tumor segmentation and prognosis. These models are typically trained on complete MRI sequences and exhibit performance degradation when MRI sequences are missing at test time. In this retrospective study of 715 patients from the Children’s Brain Tumor Network and BraTS-PEDs, and 43 patients with 157 longitudinal MRIs from PNOC003/007 clinical trials, we developed strategies for handling missing sequences. Methods included a dropout-trained segmentation model that randomly omitted FLAIR and/or T1w inputs during training, a generative model for image synthesis, copy-substitution heuristics, and zeroed inputs. The dropout model achieved robust segmentation under missing MRI, with ≤0.04 Dice drop relative to complete-input and stable prognostic accuracy in survival analysis using model-derived tumor volumes and clinical covariates. Generative synthesis achieved high image quality (SSIM > 0.90) and removed artifacts, benefiting visual interpretability. Together, these approaches can facilitate broader deployment of AI tools in real-world pediatric neuro-oncology settings.
Similar content being viewed by others
Introduction
Multiparametric MRI plays a critical role in the evaluation of pediatric brain tumors (PBT), supporting comprehensive assessment for diagnosis and response monitoring1. These tasks depend on precise delineation of intratumoral subregions, including contrast-enhancing and non-enhancing tumor, cystic components, and peritumoral edema, because each must be selectively included or excluded according to Response Assessment in Pediatric Neuro-Oncology (RAPNO) guidelines across multiple PBT histologies2,3,4,5,6,7. Accurate delineation also supports treatment planning, by enabling radiation oncologists and neurosurgeons to better target diseased tissue while sparing healthy structures8. Integrating automated deep learning (DL)–based segmentation into clinical workflows may expedite diagnostic processes and reduce inter-observer variability, leading to more standardized and timely treatment decisions. Such efficiency is especially valuable in pediatric populations, where prompt interventions are critical to reducing long-term developmental impacts8. However, in clinical practice, complete MRI acquisitions are frequently unavailable due to protocol variations or imaging artifacts. This poses a significant challenge for DL segmentation models trained on fully sampled, curated datasets8,9,10,11. Protocols designed for clinical tasks such as surgical navigation often acquire only a limited subset of sequences, such as T1-weighted post-contrast and T2-weighted images. This is further exacerbated in pediatrics, where inherently small cohort sizes, due to the lower incidence of PBTs12, are further reduced by excluding patients with missing sequences, restricting sample size, and limiting model generalizability.
In recent years, a growing number of studies have addressed missing MRI sequences using generative DL models that synthesize missing scans from available inputs. A prominent approach involves Generative Adversarial Networks (GANs), which consist of a discriminator network trained to distinguish real from synthetic images and a generator that aims to fool the discriminator by producing progressively realistic scans. Through the adversarial training, the generator learns to synthesize high-fidelity MRI images13,14,15. Another line of work employs Denoising Diffusion Probabilistic Models (DDPMs), where Gaussian noise is progressively added to the target MRI sequence in a forward diffusion process. In the reverse process, a neural network conditioned on available MRI scans learns to iteratively remove the noise. At inference, the model can synthesize high-fidelity MRI scans starting from pure noise and the conditioning inputs16,17,18. Additionally, transformer-based frameworks synthesize missing sequences by capturing intra- and inter-contrast dependencies through attention, enabling global context modeling while preserving fine detail and providing interpretable maps of how each modality contributes to the synthesis19. Further architectural variants expanding upon these core paradigms have also been explored20,21,22.
An alternative class of methods includes segmentation models inherently robust to missing sequences, eliminating the need for synthesis. Strategies include disentangling modality-specific and shared features23, learning shared representations less dependent on any single input24,25 adversarial training to help networks learn effectively from both complete and incomplete MRI data26, style transfer techniques to bridge gaps between available and missing modalities27, and region-specific fusion methods guided by anatomical information28. However, many of these models are complex and computationally demanding, limiting their practical utility. A simpler and more effective inherently robust strategy was proposed in ref. 29 employing a dropout-like training mechanism that randomly removes input sequences during training to simulate missing data. Although effective, this model is not publicly available, hindering wider clinical adoption.
Most of these strategies have been developed and validated using adult cohorts, particularly the Brain Tumor Segmentation Challenge (BraTS) dataset30 which is highly curated and may not adequately reflect the complexities encountered in real-world pediatric clinical practice. In particular, PBT differ markedly from adult tumors in their prevalence, behavior, appearance, and histology8,11. Moreover, no systematic comparison of generative models versus robustness-based approaches has been performed in pediatric tumor segmentation.
In this study, we address these gaps, using a multi-institutional, multi-histology PBT cohort to develop strategies for handling missing sequences. Methods included a dropout-trained segmentation model that randomly omitted FLAIR and/or T1w inputs during training, a generative model for image synthesis, copy-substitution heuristics, and zeroed inputs. To evaluate real-world clinical utility, we systematically assessed their performance in tumor segmentation across realistic missing MRI scenarios using retrospective clinical trial data from the PNOC003 and PNOC007 cohorts. Tumor volumes derived from these segmentations are subsequently integrated with electronic health record (EHR) data to facilitate downstream risk stratification.
Results
Study Design and Cohort Description
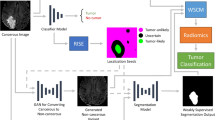
The study design focuses on an automated pipeline for assessing treatment response in PBT (Fig. 1A), in which a DL model segments key tumor subregions from multiparametric MRI and the resulting tumor volumes are combined with EHR covariates to inform a time-varying Cox model for survival risk estimation. We benchmarked four strategies for handling incomplete imaging: modality dropout, image synthesis, copy substitution, and a baseline method without imputation (Fig. 1B). To ensure a consistent ground truth when evaluating imputation strategies, analyses were restricted to patients with all four MRI sequences, with missing-sequence scenarios simulated during validation and testing. All imputation methods were trained and validated on the same random split of 340 and 85 CBTN/BraTS-PEDs patients, and segmentation performance was assessed on a held-out set of 290 CBTN/BraTS-PEDs patients. The external PNOC cohort of 43 patients was then used to evaluate segmentation generalization and perform downstream survival analysis.
A Multiparametric MRI scans are segmented at each available timepoint using a deep learning model to extract longitudinal tumor volumes. Combined with electronic health record data, they are used in a time-varying Cox model to estimate the patient’s survival risk. B Strategies for handling missing MRI sequences during segmentation: Dropout uses an nnU-Net model trained with randomly dropped sequences enabling inherent robustness to missing inputs. Synthesis, Copy, and Baseline use the out-of-the-box nnU-Net model downstream and impute missing sequences with synthetic scans, copies of closest physical counterparts, or zeros, respectively.
Figure 2 shows box-and-whisker plots of Dice scores for Whole Tumor (WT), Enhancing Tumor (ET), and Non-Enhancing Tumor (NET) in the CBTN/BraTS-PEDs held-out test set and the PNOC clinical trial data. Four test scenarios were simulated, where each of the CBTN/BraTS-PEDs and PNOC test patients had: (1) complete sequences, (2) FLAIR zeroed, (3) T1w-pre zeroed, and (4) both zeroed. Median Dice scores are annotated. Additional visualizations of Dice scores appear in Figs. S1–S3. Figure 3 presents qualitative comparisons for a representative PNOC patient with median cohort-level performance; Dice scores for each method are shown below the corresponding scans.
Box-and-whisker plots of Dice scores for WT, ET, and NET across imputation strategies and missing MRI scenarios in both the CBTN/BraTS-PEDs test set (top row) and PNOC clinical trial data (bottom row). Median Dice scores are annotated on each plot.
Segmentation outputs for a representative patient from the PNOC003 cohort under two scenarios: complete MRI input (left) and missing FLAIR and T1w-pre sequences (right). Segmentations are shown for each imputation method (Fig. 1), with Dice scores annotated below. Dice values reflect cases with performance closest to the cohort median in each setting. Tumor subregions are color-coded as follows: Green=NET, Red=ET, Blue=Cyst.
Figure 4 shows prognostic performance based on tumor volumes from manual, Baseline, and Dropout segmentations under complete input and missing FLAIR for the 43-patient (157 timepoint) external longitudinal PNOC cohort. Kaplan-Meier curves report the log-rank p-value and the corresponding forest plots show hazard ratios with 95% confidence intervals (CI) and indicators of statistical significance per covariate.
Top two rows: Kaplan–Meier survival curves and corresponding forest plots from Cox models using age, sex, treatment group, and tumor volumes derived from manual segmentations, Baseline, and Dropout models with complete MRI input. Bottom two rows: Corresponding Kaplan-Meier curves and forest plots under missing FLAIR, with segmentation handled via Dropout, Synthesis, or Baseline models. Kaplan–Meier curves are annotated with the concordance index (C-index) and log-rank p-value. Forest plots show hazard ratios with 95% confidence intervals and p-values for each covariate.
Segmentation under Complete MRI Sequences: Dropout Enhances Generalizability
On the CBTN/BraTS-PEDs test set (Fig. 2), Dropout and Baseline demonstrated comparable median/mean/std Dice scores (WT: 0.95/0.90/0.15 vs 0.95/0.90/0.13, NET: 0.91/0.82/0.22 vs 0.90/0.82/0.21, and ET: 0.81/0.73/0.24 vs 0.79/0.73/0.24) and interquartile ranges (IQRs). No significance was reached for ET (p adj. Bonferroni = 0.79, Table S1) and although statistical significance (p adj.<0.05, Table S1) was observed for WT and NET, the average per-patient Dice improvement for Baseline over Dropout was no more than 0.006 across the three regions (Table S1), suggesting limited clinical relevance. These trends indicate that incorporating dropout during training does not degrade segmentation performance when complete input sequences are available. In fact, the use of dropout appeared to enhance generalizability in the PNOC cohort reflected in increased Dice scores and reduced IQRs.
In more detail, in the PNOC cohort with complete sequences, dropout achieved notably smaller IQRs and improved median/mean/std Dice scores for WT (0.85/0.78/0.22 vs 0.84/0.69/0.31) and NET (0.76/0.71/0.23 vs 0.72/0.63/0.31) with statistical significance (p adj.=0.003 for WT and 0.006 for NET) and average per-patient Dice gain of approximately 0.1 over Baseline (Table S1). For ET, though Baseline demonstrated higher median (0.80 vs 0.77), no statistical significance was observed (p adj.=0.18), with the two methods achieving equal means (0.70) and the Dropout method demonstrating a narrower IQR (0.26 vs 0.31) suggesting improved robustness (Table S1). Altogether, these results suggest that training with modality dropout enhances model robustness and generalizability. Additional statistics can be found in Table S1.
Segmentation under Missing MRI Sequences: Dropout Yields Most Robust Performance
In most missing MRI scenarios (Fig. 2), the Dropout model achieved the highest median Dice scores and narrowest IQRs. This advantage was especially evident in the most clinically relevant cases: ET segmentation with missing T1w-pre and NET segmentation with missing FLAIR. For ET, Dropout outperformed the next-best method (Synthesis) in both cohorts: median/mean/std of 0.79/0.72/0.24 vs 0.71/0.64/0.27 (CBTN), and 0.78/0.69/0.28 vs 0.74/0.60/0.33 (PNOC). For NET with missing FLAIR, Dropout achieved 0.87/0.79/0.23 vs 0.85/0.74/0.26 (CBTN) and 0.74/0.68/0.23 versus 0.68/0.61/0.27 (PNOC). All differences were statistically significant (p adj. < 0.05). Across all remaining regions and MRI availability scenarios Dropout achieved average per-patient Dice gains up to 0.12 over the next-best method and up to 0.57 over all others (Tables S2, S3).
The Dropout model maintained segmentation performance under missing sequences comparable to its performance with complete inputs, both in median Dice and IQRs (Fig. 2). Specifically, in key clinical cases: For ET with missing T1w-pre, the median/mean/std Dice scores for CBTN were 0.79/0.73/0.24 (complete) vs 0.79/0.72/0.24 (missing), and for PNOC, 0.77/0.70/0.27 vs 0.78/0.69/0.28. For NET under missing FLAIR, scores were 0.90/0.82/0.21 vs 0.87/0.79/0.23 in CBTN, and 0.76/0.71/0.23 vs 0.74/0.68/0.23 in PNOC. Although these differences were statistically significant, the average per-patient Dice drop never exceeded 0.03 and 0.04 when considering all remaining regions and MRI availability scenarios indicating minimal practical impact in segmentation performance (Tables S4, S5).
The Dropout method was outperformed in median Dice only in two cases: Synthesis in ET segmentation with missing FLAIR (0.79 vs 0.78 for CBTN and 0.81 vs 0.78 for PNOC), and by Copy in NET with T1w-pre missing (0.91 vs 0.90 for CBTN). However, these differences were either not statistically significant or had average per-patient Dice difference ≤ 0.03 indicating no practical significance (Tables S2, S3).
Synthesis Models Add Information Beyond Source Scans
In most missing MRI scenarios (Fig. 2), the Synthesis approach outperformed the Copy method, achieving significantly (p adj.<0.05) higher median Dice and lower IQRs with average per-patient Dice gains of up to 0.26 (Tables S6, S7). For WT and NET segmentation with T1w-pre missing Copy performed comparably or slightly better, though gains over Synthesis did not exceed 0.03 on average (Tables S6, S7). These results suggest that the synthesis model generates anatomically meaningful information beyond what is present in nearest physical modality.
Sensitivity Analysis: Clinical Characteristics Affecting Segmentation Performance
To identify cases where missing T1w-pre or FLAIR sequences led to substantial segmentation degradation, we formed two targeted cohorts based on the Baseline method segmentation performance: Patients with ≥ 0.1 Dice drop in ET when T1w-pre was missing, and in NET when FLAIR was missing. Radiologist review revealed distinct patterns: the cohort affected most by missing T1w-pre was marked by patients with mildly enhancing tumors, diffuse edema, or hydrocephalus. The cases affected most by missing FLAIR sequence primarily tended to lack post-contrast enhancement on T1w-post. These patterns highlight clinical contexts where specific sequences are critical to accurate subregion delineation.
Under complete-sequence inputs, variance in the Dice scores naturally reflects the heterogeneous nature of the CBTN/BraTS-PEDs cohort: while the Baseline method performs well for WT segmentation, it shows greater variance for the smaller ET and NET subregions, whose size, and fragmentation, make their Dice scores more sensitive to minor errors. Manual review confirmed that low contrast, noise, irregular ground-truth labels, and very small or very large tumors further amplify this variability. On the external PNOC cohort, additional variance arises from domain shift, though the Dropout model shows reduced variability compared to the Baseline model. In incomplete-MRI scenarios, Dropout maintains relatively stable variance, whereas Synthesis, Copy, and Baseline, display progressively more unstable performance.
Longitudinal Risk Stratification using Clinical Trial Data
Figure 4, top shows Kaplan-Meier curves and corresponding forest plots based on tumor volumes from manual, Baseline, and Dropout segmentations under complete input, showing progressively stronger separation between low/high-risk groups. The C-index modestly improves from 0.63 (manual), to 0.64 (Baseline), to 0.65 (Dropout). Hazard ratios (HRs) also increased: 1.38 (p = 0.0541) for manual, 1.64 (p = 0.0045) for Baseline, and 1.68 (p = 0.0036) for Dropout. Risk scores did not differ significantly across models (p adj. > 0.05, Table S8). The Dropout model yielded the most predictive combination, with tumor volume (HR = 1.68, p = 0.0036) and treatment group (HR = 2.57, p = 0.0076) emerging as significant covariates, indicating that both disease burden and therapeutic intervention play significant roles in determining patient’s survival. Overall, automated segmentation under complete input yields prognostic performance comparable to manual segmentation.
When FLAIR was missing (Fig. 4, bottom), the Dropout model maintained a C-index of 0.65, and a significant association between tumor volume and survival (HR = 1.57, 95% CI: 1.15–2.14, p = 0.0045), comparable to its performance with complete input. Risk scores from Dropout with missing FLAIR did not differ significantly from its complete-input counterparts (p > 0.05, Table S8). In comparison, the Synthesis method achieved weaker results (C-index = 0.63; HR = 1.44, p = 0.0082), and Baseline underperformed (C-index = 0.61; HR = 1.29, p = 0.106). Under missing FLAIR, risk scores from Dropout were statistically different (p adj.<0.05) from those produced by both the Synthesis and Baseline models. These findings demonstrate that dropout training provides the most robust imputation strategy of missing FLAIR, outperforming both explicit synthesis and naïve omission. Age and sex were not significantly associated with survival across models while treatment group remained a statistically significant predictor.
Perceptual Quality Analysis of Synthesized scans
Image synthesis quality for the held-out 290-patient CBTN/BraTS-PEDs cohort and 43-patient external PNOC cohort was evaluated using structural similarity index (SSIM), mean squared error (MSE), and peak signal-to-noise ratio (PSNR). The Synthesis model generated FLAIR and T1w-pre scans with high perceptual quality in both CBTN/BraTS-PEDs and PNOC cohorts. On CBTN/BraTS-PEDs, median SSI/MSE/PSNR along the axial dimension were 0.93/0.003/27.1 in FLAIR synthesis and 0.91/0.015/20.4 for T1w-pre synthesis. On PNOC, median metrics were 0.91/0.004/24.9 for FLAIR synthesis and 0.90/0.014/20.9 for T1w-pre. Representative examples with perceptual metrics are shown in Fig. 5, additional samples and metrics appear in Fig. S4 and Table S9.
Shown are examples of synthesized FLAIR MRIs from the CBTN/BraTS-PEDs and PNOC cohorts, along with a case demonstrating artifact removal via synthesis. For each example, the ground-truth scan (left) is shown next to its synthesized counterpart (right), with SSIM, MSE, and PSNR calculated for the displayed pair of slices. The corresponding metrics averaged across the full volume in the axial plane are provided in parentheses.
In some cases, the synthetic images appeared visually superior to the ground truth, as the Synthesis method removed artifacts. Although these scans received lower perceptual scores due to dissimilarity with the artifact-containing ground truth, they exhibit higher visual quality. For instance, (Fig. 5, right), a strong artifact in the ground truth FLAIR obscures part of the tumor, whereas the synthetic scan recovers its full extent.
DISCUSSION
State-of-the-art DL segmentation models utilize the four standard MRI sequences (T1w-pre, T1w-post, T2w, and FLAIR), each providing unique information for tumor subregion delineation11. However, in clinical practice, MRIs exams are often incomplete due to acquisition variability or artifacts, a challenge amplified in pediatrics, where small cohort sizes are further reduced by missing data, hindering model generalizability.
Existing strategies including MRI synthesis14 and robustness-focused segmentation29, are often unavailable to the public or validated exclusively on adult datasets such as BraTS30, which are highly curated and not representative of pediatric clinical settings. Moreover, no systematic comparison of generative imputation models versus robustness-based approaches has been conducted for pediatric brain tumor segmentation. To address these gaps, we leveraged a large, multi-institutional, multi-histology pediatric cohort of 715 patients from the CBTN31,32, BraTS-PEDs8 to develop and compare various imputation methods including a dropout-trained segmentation model, a generative model for image synthesis, and copy-substitution heuristics. To evaluate real-world clinical utility, we systematically assessed their performance in tumor segmentation across realistic missing MRI scenarios using retrospective clinical trial data from the PNOC003/00733,34 clinical trials.
The dropout-trained model outperformed the out-of-the-box nnU-Net (Baseline) even under complete sequences, likely due to its more challenging training regime, where randomly dropped inputs forced the model to learn more robust and generalizable feature representations. In survival analysis, models built from Dropout-derived tumor volumes performed statistically comparable to those based on manual segmentations. Under missing sequences, the dropout-trained model consistently outperformed other imputation strategies, maintaining segmentation accuracy with minimal degradation, even when both T1w-pre and FLAIR were absent. Furthermore, under missing FLAIR, the model preserved prognostic risk stratification compared to complete MRI. This is particularly important for DMG/DIPG, which are predominantly non-enhancing tumors, making FLAIR imaging essential for accurate NET segmentation. DL-based MRI synthesis achieved high perceptual image quality, with anatomically consistent outputs, effective for artifact removal and qualitative assessment.
Despite these advances, this study has limitations. We selected ResViT for its strong performance in image synthesis in adult studies, but its 2D architecture lacks volumetric context and requires extensive training and preprocessing, including skull-stripping and 2D slicing. While the dropout-based segmentation model demonstrated robustness to missing FLAIR and/or T1w-pre, its performance under additional missing MRI scenarios was not evaluated and warrants future investigation. Future work could explore more advanced generative architectures, such as 3D denoising diffusion probabilistic models (DDPMs), or many-to-one synthesis strategies that leverage multiple available sequences to reconstruct missing modalities.
In conclusion, we developed a robust modality imputation strategy using a dropout-trained segmentation model that performs reliably under both complete and incomplete MRI conditions. This approach offers a generalizable and practical solution for handling missing data in pediatric brain tumor segmentation and survival analysis, supporting more reliable AI deployment in real-world pediatric neuro-oncology settings.
METHODS
Datasets
We assembled a multi-institutional, multi-histology retrospective cohort of 715 pediatric patients from the Children’s Brain Tumor Network (CBTN)31,32 and the Brain Tumor Segmentation in Pediatrics (BraTS-PEDs)8 datasets. Additionally, we curated a retrospective longitudinal cohort of 43 patients with diffuse midline glioma (DMG) from the PNOC003 and PNOC007 clinical trials33,34 comprising 157 imaging timepoints. Each of the 715 CBTN/BraTS-PEDs and 43 PNOC patients had availability of standard MRI consisting of pre- and post-contrast T1-weighted (T1w-pre, T1w-post), T2-weighted (T2w), and fluid-attenuated inversion recovery (FLAIR) images and expert manual segmentations. Additional inclusion criteria for the PNOC003/007 cohorts included a confirmed diagnosis of diffuse intrinsic pontine glioma (DIPG), exclusion of post-contrast FLAIR scans, and removal of scans affected by imaging artifacts. All images were co-registered and resampled to an isotropic resolution of 1 mm³ based on the anatomical SRI24 atlas, resulting in MRI volumes of size 240 × 240 x 155 using CaPTk software35,36. Demographics are provided in Tables 1,2.
DL-Based Tumor Segmentation
We select nn-UNet37 as the backbone for DL-based tumor segmentation (Fig. 1A), as its open-source availability and out-of-the-box usability make it highly suitable for clinical translation. By automatically configuring data preprocessing, network architecture, training, and postprocessing based on available hardware, nn-UNet removes the need for specialized DL expertise required in the complex task of building and optimizing segmentation pipelines. The framework has achieved state-of-the-art performance in both adult and pediatric brain tumor segmentation11,37. Here, the segmentation model is used to delineate key tumor subregions, including Whole Tumor (WT), Enhancing (ET), Non-Enhancing tumors (NET), cyst, and edema using standard MRI sequences (T1w-pre, T1w-post, T2w, and FLAIR). However, incomplete imaging is common due to protocols variability, artifacts, or in surgical navigation protocols. To address frequent absence of T1w-pre and/or FLAIR, we augmented the nnU-Net backbone resulting in four imputation strategies (Fig. 1B): modality dropout, image synthesis, copy substitution, and a baseline method without imputation. Each of the models was trained and validated on the same random split of 340 and 85 CBTN/BraTS-PEDs patients.
Modality Dropout: Training for Robustness to Missing Inputs
Inspired by the dropout mechanism used in neural networks to prevent overfitting38, our “modality dropout” strategy randomly removes MRI sequences during training to simulate incomplete inputs, encouraging the model to learn representations robust to missing data. We integrated this into the nnU-Net framework as a data augmentation step, preserving its automated configuration. For each training sample (comprising four MRI sequences as tensor channels), FLAIR and T1w-pre sequences are independently set to zero with probability p ∈ {0, 0.1, 0.2, …, 1}, treated as a tunable hyperparameter. For example, with \(p=0.5\) each sequence is dropped in 50% of samples, and both are dropped simultaneously in about 25% of cases, assuming independence. Due to the stochastic nature of this approach, the specific dropout patterns for a given patient vary across epochs, providing a diverse range of input combinations, helping the model generalize to real-world missing data scenarios. We optimized p by averaging WT Dice across four simulated validation scenarios, where each of the 85 validation patients had: (1) all sequences available, (2) FLAIR zeroed, (3) T1w-pre zeroed, and (4) both zeroed. The model trained with p = 0.4 achieved the best average performance and was selected for inference on the test set. Visualization of the described dropout patterns during training and validation Dice scores for each scenario are shown in Fig. S5, S6.
Image Synthesis: Generating Missing Sequences with ResViT
As an alternative, we employed a generative model to synthesize missing MRI sequences, imputing absent inputs during nnU-Net test-time inference. We selected ResViT14, a GAN-based publicly available model that has demonstrated strong performance in MRI synthesis on the adult BraTS dataset30. ResViT combines the sensitivity to global context of vision transformers, the local feature extraction capabilities of convolutional networks, and the image synthesis realism of adversarial learning strategies. We trained two ResViT models to respectively synthesize missing FLAIR from T2w and T1w-pre from T1w-post scans (Synth FLAIR and Synth T1, respectively, in Fig. 1B). In addition to standard image preprocessing (co-registration and resampling), each MRI volume was skull-stripped39 and padded to a uniform shape of 256 × 256 × 155 using its background voxel intensity. Volumes were then normalized to [-1, 1]. To accommodate the 2D input format required by ResViT, each volume was sliced along the axial plane, resulting in 155 slices of size 1 × 256 × 256 per MRI sequence. We followed the training protocol detailed in ref. 14, performing a 100-epoch grid search over learning rates {\(1{0}^{-5},\,1{0}^{-4},\,2\times 1{0}^{-4},\,5\times 1{0}^{-4},\,1{0}^{-3}\}\) and loss weights {20, 50, 100,150, 200, 250} using the 85-patient validation cohort. The hyperparameters that maximized PSNR on the validation set were a learning rate of 1e-4 and a loss weight of 200, differing from those reported as optimal in adult cohorts14. Following14 transformer modules were then inserted into the pre-trained ART blocks of ResViT and fine-tuned for 50 epochs. The described validation curves for ResViT training are shown in Figs. S7, S8.
Copy Substitution: a simple imputation strategy
As a simple imputation strategy, missing sequences were replaced with their closest physical counterparts13: missing FLAIR with T2w, and missing T1w-pre with T1w-post. This provides a benchmark to assess the added value of learned synthesis models, which are trained on the same input-output pairs. This comparison is critical, as DL-based synthesis models could converge to suboptimal solutions by simply replicating anatomical structures from the input without recovering the intended contrast, especially when source and target sequences are structurally similar. Superior performance by ResViT would suggest it captures contrast-specific features not present in the nearest physical modality.
Baseline Model: Standard nnU-Net with Zeroed Inputs
As a final baseline, we evaluate the standard nnU-Net by running inference with missing sequences set to zero. This reflects the model’s default behavior without any imputation or robustness strategy and helps quantify the value added by the proposed imputation strategies. The self-configured nnU-Net network adopts a six-stage encoder-decoder 3D full-resolution convolutional U-Net with skip connections and Leaky ReLU activations. Training was performed for 1000 epochs using stochastic gradient descent (SGD) with an initial learning rate of 0.01, momentum of 0.99, weight decay of 3e-5, a polynomial learning rate scheduler, and a batch size of 2, where each sample comprised z-score normalized patches of size 96 × 160 × 160 from a patient’s four MRI sequences, concatenated as input channels. The loss function combined Dice and cross-entropy components, with deep supervision enabled throughout training. Inference was performed using a sliding window, to segment the entire volume. For fair comparison, all imputation strategies (Fig. 1B) used the same training hyperparameters for the nnU-Net backbone.
Time Varying Cox-Regression
To model overall survival (OS), we implemented a time-varying Cox regression framework, appropriate for longitudinal clinical data where tumor burden may evolve over time. Following emerging evidence that volumetric measures better capture tumor growth and response than traditional bidimensional measurements40,41 and in accordance with DIPG/DMG RAPNO guidelines7, whole tumor volume derived from expert manual segmentations and each of the imputation strategies of Fig. 1B, served as a time-varying covariate, while age, sex, and treatment group were modeled as fixed effects. Survival time was defined from the day of diagnosis. Kaplan–Meier curves reporting the log-rank p-value, were used to quantify survival separation between high- and low-risk groups. Patients were dichotomized using baseline risk at diagnosis, reflecting an interpretable clinical scenario in which decisions are made at presentation. C-index is computed using risk predictions from each patient’s last imaging timepoint, capturing dynamic discrimination based on accumulated longitudinal information. Differences in risk scores across Cox models were tested using the Wilcoxon signed-rank test with Bonferroni corrections (significance threshold, p = 0.05).
Ethics statement
All research involving human participants, clinical data, or biospecimens was conducted in accordance with the Declaration of Helsinki and relevant institutional guidelines. CBTN data were obtained from the Children’s Brain Tumor Network (cbtn.org), which provides de-identified imaging, clinical, and biospecimen data collected under IRB approval at the Children’s Hospital of Philadelphia (CHOP). Informed consent has been obtained from participants or their legal guardians at the time of enrollment to CBTN. BraTS-PEDs data are publicly available, fully deidentified, and contain no information that enables the identification of individual participants; therefore, additional consent was not required for their use in this study. Data from the PNOC003 and PNOC007 clinical trials were accessed under a data transfer and usage agreement. These datasets were provided in fully anonymized form, and the CHOP IRB granted a waiver of informed consent for this retrospective secondary analysis because the research involved no direct participant contact and posed minimal risk.
Data availability
Raw CBTN files can be accessed through https://cbtn.org/. CBTN processed and annotated data can be made available upon reasonable request to the corresponding author. The PNOC003/007 data can’t be shared. BraTS-PEDs data can be accessed through The Cancer Imaging Archive at https://www.cancerimagingarchive.net/collection/brats-peds/ (https://doi.org/10.7937/DX5C-TJ86).
Code availability
The code for the Modality Dropout segmentation model is publicly available at https://github.com/d3b-center/peds-brain-auto-seg-public/tree/main/Dropout_nnUNet_2_sequences. The remaining methods are publicly available from refs. 14,37.
References
Jung, A. Y. Basics for pediatric brain tumor imaging: techniques and protocol recommendations. Brain Tumor Res. Treat. 12(1), 1 (2024).
Warren, K. E. et al. Response assessment in medulloblastoma and leptomeningeal seeding tumors: recommendations from the response assessment in pediatric neuro-oncology committee. Neuro Oncol. 20(1), 13–23 (2018).
Hoffman, L. M. et al. Response assessment in pediatric craniopharyngioma: recommendations from the Response Assessment in Pediatric Neuro-Oncology (RAPNO) Working Group. Neuro Oncol. Oxf. Univ. Press 25, 224–233 (2023).
Lindsay B, et al. Response Assessment in Paediatric Intracranial Ependymoma: Recommendations from the Response Assessment in Pediatric Neuro-Oncology (RAPNO) Working Group. Vol 23.; 2022. www.thelancet.com/oncology.
Poussaint TY, et al. Title: Response Assessment in Pediatric High-Grade Glioma: Recommendations from the Response Assessment in Pediatric Neuro-Oncology Working Group.
Fangusaro, J. et al. Response assessment in paediatric low-grade glioma: recommendations from the response assessment in pediatric neuro-oncology (RAPNO) working group. Lancet Oncol. Lancet Publ. Group 21, e305–e316 (2020).
Cooney TM, et al. Series Imaging Guidelines for Paediatric Brain Tumours 3 Response Assessment in Diffuse Intrinsic Pontine Glioma: Recommendations from the Response Assessment in Pediatric Neuro-Oncology (RAPNO) Working Group.; 2020. www.thelancet.com/oncology.
Fathi Kazerooni, A. et al. BraTS-PEDs: results of the multi-consortium international pediatric brain tumor segmentation challenge 2023. Mach. Learn. Biomed. Imaging 3, 72–87 (2025).
Fathi Kazerooni, A. et al. Automated tumor segmentation and brain tissue extraction from multiparametric MRI of pediatric brain tumors: a multi-institutional study. Neurooncol Adv. 5(1), 1–12 (2023).
Familiar, A. M. et al. Towards consistency in pediatric brain tumor measurements: challenges, solutions, and the role of artificial intelligence-based segmentation. Neuro Oncol. 26(9), 1557–1571 (2024).
Vossough, A. et al. Training and comparison of nnu-net and deepmedic methods for autosegmentation of pediatric brain tumors. Am. J. Neuroradiol. 45(8), 1081–1089 (2024).
Madhogarhia R, et al. Radiomics and radiogenomics in pediatric neuro-oncology: a review. Neurooncol Adv.Oxford University Press. 2022;4. https://doi.org/10.1093/noajnl/vdac083.
Conte, G. M. et al. Generative adversarial networks to synthesize missing T1 and FLAIR MRI sequences for use in a multisequence brain tumor segmentation model. Radiology 299(2), 313–323 (2021).
Dalmaz, O., Yurt, M. & Cukur, T. ResViT: residual vision transformers for multimodal medical image synthesis. IEEE Trans. Med Imaging 41(10), 2598–2614 (2022).
Cao, B. et al. Autoencoder-driven multimodal collaborative learning for medical image synthesis. Int J. Comput Vis. 131(8), 1995–2014 (2023).
Li Y, et al. Zero-shot medical image translation via frequency-guided diffusion models. Published online April 5, 2023. https://doi.org/10.1109/TMI.2023.3325703.
Jiang L, Mao Y, Chen X, Wang X, Li C. CoLa-Diff: conditional latent diffusion model for multi-modal mri synthesis. Published online March 24, 2023. http://arxiv.org/abs/2303.14081.
Özbey, M. et al. Unsupervised medical image translation with adversarial diffusion models. IEEE Trans. Med Imaging 42(12), 3524–3539 (2023).
Liu, J. et al. One model to synthesize them all: multi-contrast multi-scale transformer for missing data imputation. IEEE Trans. Med. Imaging 42, 2577–2591 (2023).
Atli OF, et al. I2I-Mamba: multi-modal medical image synthesis via selective state space modeling. Published online May 22, 2024. http://arxiv.org/abs/2405.14022.
Li Y, Zhou T, He K, Zhou Y, Shen D. Multi-scale transformer network with edge-aware pre-training for cross-modality mr image synthesis. Published online December 2, 2022. https://doi.org/10.1109/TMI.2023.3288001.
Arslan F, Kabas B, Dalmaz O, Ozbey M, Çukur T. Self-consistent recursive diffusion bridge for medical image translation. Published online May 10, 2024. http://arxiv.org/abs/2405.06789.
Chen C, et al., ed. Medical Image Computing and Computer Assisted Intervention – MICCAI 2019. 447-456 (Springer, 2019).
Havaei M, Guizard N, Chapados N, Bengio Y. HeMIS: Hetero-Modal Image Segmentation. In: Medical Image Computing and Computer-Assisted Intervention – MICCAI 2016, Part II. 469-477 (Springer, 2016).
Dorent R, Joutard S, Modat M, Ourselin S, Vercauteren T. Hetero-Modal Variational Encoder-Decoder for Joint Modality Completion and Segmentation. In: Medical Image Computing and Computer Assisted Intervention – MICCAI 2019. 225-233 (Springer, 2019).
Wang Y et al. ACN: Adversarial Co-training Network for Brain Tumor Segmentation with Missing Modalities. In: Medical Image Computing and Computer Assisted Intervention – MICCAI 2021. 415-426 (Springer, 2021).
Azad, R., Khosravi, N. & Merhof, D. SMU-Net: Style matching U-Net for brain tumor segmentation with missing modalities. Int. Conf. Med. Imaging Deep Learn. (MIDL 2022) 172, 24–45 (2025).
Ding Y, Yu X, Yang Y. RFNet: Region-aware Fusion Network for Incomplete Multi-modal Brain Tumor Segmentation. In: Proceedings of the IEEE/CVF International Conference on Computer Vision (ICCV). 3975-3984 (IEEE, 2021).
Feng, X. et al. Brain tumor segmentation for multi-modal MRI with missing information. J. Digit Imaging 36(5), 2075–2087 (2023).
Baid U, et al. The RSNA-ASNR-MICCAI BraTS 2021 benchmark on brain tumor segmentation and radiogenomic classification. Published online July 5, 2021. http://arxiv.org/abs/2107.02314.
Lilly J V., et al. The children’s brain tumor network (CBTN)-Accelerating research in pediatric central nervous system tumors through collaboration and open science. Neoplasia 35, 100846 (2023).
Familiar AM, et al. A Multi-Institutional Pediatric Dataset of Clinical Radiology MRIs by the Children’s Brain Tumor Network.
Kline, C. et al. Upfront biology-guided therapy in diffuse intrinsic pontine glioma: therapeutic, molecular, and biomarker outcomes from PNOC003. Clin. Cancer Res. 28(18), 3965–3978 (2022).
Mueller, S. et al. A pilot precision medicine trial for children with diffuse intrinsic pontine glioma—PNOC003: a report from the Pacific Pediatric Neuro-Oncology Consortium. Int J. Cancer 145(7), 1889–1901 (2019).
Davatzikos, C. et al. Cancer imaging phenomics toolkit: quantitative imaging analytics for precision diagnostics and predictive modeling of clinical outcome. J. Med. Imaging 5(01), 1 (2018).
Pati S, et al. The cancer imaging phenomics toolkit (CaPTk): Technical overview. In: Lecture Notes in Computer Science (Including Subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics). Vol 11993 LNCS. 380-394 (Springer, 2020).
Isensee, F., Jaeger, P. F., Kohl, S. A. A., Petersen, J. & Maier-Hein, K. H. nnU-Net: a self-configuring method for deep learning-based biomedical image segmentation. Nat. Methods 18(2), 203–211 (2021).
Goodfellow I, Bengio Y, Courville A. Deep Learning.
Gandhi DB, et al. Automated pediatric brain tumor imaging assessment tool from CBTN: Enhancing suprasellar region inclusion and managing limited data with deep learning. Neurooncol Adv. 2024;6(1). https://doi.org/10.1093/noajnl/vdae190.
Lazow, M. A. et al. Volumsetric endpoints in diffuse intrinsic pontine glioma: p correlations in the International DIPG/DMG Registry. Neuro Oncol. 24(9), 1598–1608 (2022).
Von Reppert M, et al. Comparison of volumetric and 2D-based response methods in the PNOC-001 pediatric low-grade glioma clinical trial. Neurooncol Adv. 2024;6(1). https://doi.org/10.1093/noajnl/vdad172.
Acknowledgements
National Institute of Health (NIH) grants 75N91019D00024, Supplement 3U2CHL156291-03S2, and 75N91019D00024, awarded to A.C.R.
Author information
Authors and Affiliations
Contributions
A.F.K., A.N., and C.D. conceptualized the study, D.C. and D.B.G., designed and implemented the methodology and performed data analysis, D.C. and A.F.K. wrote the manuscript, D.B.G., S.A., A.M.F., B.V., S.V., N.K., N.K., S.H., P.J., H.A., W.T., J.B.W., S.M., C.K., A.F., M.P., and A.V. contributed to data curation and data generation, D.C., A.F.K., S.A., and A.V. contributed to the interpretation of data, P.B.S., A.N., and A.R. provided resources.
Corresponding author
Ethics declarations
Competing interests
Authors A.F.K. and A.N. served as guest editors for this journal's “Noninvasive biomarkers: bridging real-world evidence and clinical trial data” collection.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chrysochoou, D., Gandhi, D.B., Adib, S. et al. AI-powered segmentation and prognosis with missing MRI in pediatric brain tumors. npj Precis. Onc. 10, 63 (2026). https://doi.org/10.1038/s41698-025-01269-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41698-025-01269-x