Abstract
Artificial intelligence (AI) is quickly becoming a revolutionary and game-changing tool in modern oncology, with promising uses in early diagnosis and drug discovery. Machine learning (ML), deep learning (DL), reinforcement learning (RL), natural language processing (NLP), and generative models are some of the AI methods that are becoming very important for cancer care. With an emphasis on early diagnosis, mutation mapping, and drug design, this article aims to review the existing literature and investigate the role of AI technologies in oncology.

Similar content being viewed by others
Introduction
Cancer continues to be one of the leading health problems, with increasing morbidity and mortality worldwide1,2. The IARC projects that in 2022, there will be 19.9 million new cancer cases and 9.7 million cancer-related deaths3. Several irreconcilable theories exist that explain the complex pathophysiology of cancer. However, the uncontrolled proliferation of cells is one of the leading and most accepted mechanisms4. According to cancer statistics, there are more than 200 distinct types of cancers, which are substantially different in their behaviour and response to treatment. Different cancers are diagnosed using different approaches or a combination of approaches, such as cytology, cytogenetics, histology, pathology, imaging, and other relevant clinical data that can help in the process of diagnosis5.
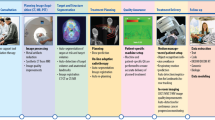
Artificial intelligence (AI) is a branch of computer science that deals with developing smart machines that can perform tasks that normally require human input and intelligence6. AI is an emerging and interdisciplinary science that was first introduced in 1956 by John McCarthy7. It has significantly improved performance in basic biology, pharmacology, and clinical medicine (Fig. 1), sometimes matching or even outperforming human experts in particular tasks. Over the years, AI has emerged as a key tool in healthcare, with applications such as diagnosis, prognosis, screening, and drug discovery8,9.
Demonstrates the various areas in which artificial intelligence (AI) aids in cancer research and treatment, from imaging, clinical decision support, and global data integration to mutation mapping, biomarker discovery, and treatment development.
Natural language processing (NLP), machine learning (ML), evolutionary algorithms, and deep learning (DL) are some of the subfields of artificial intelligence (AI)8,10. DL and ML are two of the most important since DL uses multilayered neural networks to identify patterns through hierarchical abstraction, while ML enables autonomous learning from datasets9,11,12. By finding patterns in existing datasets, supervised machine learning techniques like support vector machines and random forest algorithms have been used in oncology to classify tumours, predict prognoses, and produce data-driven details12. Similarly, to improve tumour detection in medical imaging, DL architectures such as convolutional neural networks (CNNs) have been used in radiology. Recurrent neural networks (RNNs), which recognise sequential patterns in speech or text, are increasingly being used in natural language processing (NLP) applications, particularly in the extraction of clinically relevant data from reports on cancer pathology11. All of these strategies have worked together to establish AI as a revolutionary tool for early cancer detection, treatment selection, and individualised care12,13.
Three major areas, early cancer diagnosis, therapy, and prediction, that span the domains of cancer incidence, recurrence, and survival, were highlighted by Sebastian and Peter as areas in oncology where AI holds particular promise14. AI is also crucial for cancer screening and clinical trials. Diagnostics, particularly radiology and pathology, account for 80% of AI applications in oncology. Artificial intelligence (AI) algorithms are often faster and more accurate than human experts at processing and interpreting complex imaging data from MRIs, CT scans, and X-rays. These tools enable earlier and more accurate diagnosis by spotting minute patterns and abnormalities that the human eye might miss15.
In recent years, electronic healthcare record (EHR) infrastructures have expanded globally, allowing large volumes of clinical data to be stored and accessed more efficiently16. Numerous digital collaborations are now using EHRs to advance research in the early diagnosis of cancer17. AI-based imaging algorithms are increasingly being applied in clinical practice to detect and monitor potentially cancerous lesions as well as to support management decisions18,19. For instance, an FDA-approved software platform enables comprehensive detection and longitudinal tracking of pulmonary nodules, prediction of lung malignancy in lesions identified on low-dose CT scans20, and integration of management guidelines21. Deep neural networks have also been developed for identifying enlarged lymph nodes and colonic polyps on CT images, and a real-time DL computer-aided detection system for colonoscopy has received FDA approval to enhance polyp detection22,23. Furthermore, AI-augmented interpretation of endoscopic images has been shown to consistently improve the accuracy of oesophageal cancer detection24,25.
With the rapid expansion of digital pathology, AI is being applied to pathological image analysis for diagnosis, grading, and prognostic biomarker assessment. Much of the progress has focused on automating labour-intensive tasks, thereby enhancing pathologists’ efficiency and allowing greater focus on complex, high-level decision-making associated with disease presentation26. One notable example is a DL system that uses whole-slide image analysis of radical prostatectomy specimens to assign Gleason scores, with an accuracy higher than that of general pathologists27. The identification of tumour-infiltrating lymphocytes in tissue slide images from The Cancer Genome Atlas, a prognostic indicator for clinical outcomes across 13 cancer subtypes, has also been automated using convolutional neural networks28. AI has also been effectively used to annotate skin lesions, including melanoma, and classify dermoscopy images with accuracy on par with that of skilled dermatologists29,30.
To fully characterise the tumour landscape and improve diagnostic accuracy, there is increasing emphasis on integrating multi-modal data, such as transcriptomic, radiomic, metabolomic, genomic, and clinical features, given the complexity of tumour biology. Cross-modality association studies are made easier, and model development is supported by large-scale resources like LinkedOmics, which includes multi-omic data from 11,158 patients with 32 different types of cancer31,32. When combined with clinically annotated cohorts, the application of AI to analyse large-scale multi-omics datasets has made it possible to predict RNA splice sites, enhance variant detection, uncover drug-susceptibility genes, and gain a better understanding of cancer biology33,34,35,36.
This review aims to evaluate the current literature on the role of AI in cancer with a focus on cancer diagnosis, prognosis, drug development, and effective treatment strategies. This will also analyse the emerging role of AI-based algorithms in radiology and imaging. The importance of several platforms, such as DL, ML, and RL, will also be evaluated in the context of cancer care. This review will also combine evidence from multiple omics with cancer genomics and personalised drug design to show how AI-driven approaches are transforming cancer research and clinical practice. Particular emphasis is placed on how these technologies improve diagnostic accuracy, make it easier to accurately identify therapeutic targets, and speed up the development of individualised treatment plans.
Methodological approaches
Literature search strategy
A range of electronic databases, including EMBASE, Web of Science, JSTOR, ScienceDirect, BioMed Central, Medline, PubMed, and Google Scholar, were searched to retrieve literature for this review article. Artificial intelligence (AI), cancer, oncology, cancer genomics, diagnosis, drug discovery, cancer epigenetics, machine learning, computational oncology, deep learning, biomarkers, big data, and precision medicine were among the search terms used. To generate various keyword combinations, logical operators (“AND,” “OR”) were used. Articles with clinical relevance to artificial intelligence and oncology, as well as those published in high-impact factor journals, were given preference. As shown in the flowchart (Fig. 2), out of the 641 articles that were retrieved, 310 were deemed suitable for inclusion.
Shows how research articles were screened and selected for this review article. Initially, 641 articles were retrieved from different databases, of which 310 articles were selected.
Eligibility criteria
Original research articles published in English that looked at the use of AI in oncology, with a focus on machine learning and deep learning techniques, were the main focus of the inclusion criteria. There were also other types of articles that summarised the use of AI in oncology, like meta-analyses and systematic reviews. Letters, editorials, commentaries, articles lacking primary data, and publications published in languages other than English were all excluded based on the exclusion criteria. Inadequately detailed review articles were also omitted (Table 1).
Overview of AI-driven technologies in healthcare
Over the last few years, AI technologies have been increasingly integrated into several sectors, including healthcare. These tools not only help in diagnosis, screening, treatment, and decision-making but also give a sense of hope to patients, clinicians, and caregivers with more accuracy and improved outcomes in the near future37. During the COVID-19 pandemic, several AI-assisted technologies were used to provide personalised assistance and recommendations to patients38. Similarly, several other AI-driven digital technologies, such as Google Assistant, Amazon Alexa, Apple Siri, and AI-driven robots and personal assistants, are now being used in a wide range of scenarios39.
In the last decade, we have effectively utilised machine learning techniques to analyse large volumes of medical data. This approach enables systems to learn independently, producing accurate predictions instead of depending on traditional rule-based techniques40,41,42. ML and DL algorithms powered by AI have been used to find intricate patterns in clinical and medical images. To enable meaningful decisions that are frequently difficult for humans, they try to translate images and support clinical judgments. AI improves the collection of diverse data streams into symptomatic systems that are dynamically integrated43,44,45,46. These techniques can help you extract important information from patient data found in electronic health records (EHRs)47,48,49,50 and can help diagnose diseases by uncovering risk factors and their underlying causes51,52,53.
As ML continues to evolve, DL has emerged as a significant subset that uses multiple layers of nonlinear modules54,55,56,57. These stacked modules work together to gradually convert raw data into more abstract representations56,57,58,59. DL offers intriguing possibilities for biomedical informatics and disease diagnosis, with numerous projects currently underway to apply these techniques in the medical field. For example, Google DeepMind has announced its plans to utilise its expertise for medical applications57. These days, convolutional neural networks (CNNs) are a popular deep learning technique for automatically detecting tumours in cancer patients’ medical images60,61,62,63,64.
The early onset of heart failure has been predicted using DL-based recurrent neural network (RNN) models65,66,67. Likewise, generative adversarial networks (GANs) are dynamic medical imaging tools that enhance training techniques, enhance image quality, and are crucial to the advancement of healthcare technologies68,69,70,71. Using ML and AI algorithms to efficiently interpret text data is known as natural language processing (NLP)72,73,74,75. Common NLP techniques include topic modelling, which creates topics of interest by identifying word frequencies and associations75,76; and determines whether a text accurately or negatively reflects attitudes, opinions, and feelings77,78.
Computer vision (CV) is another branch of artificial intelligence that enables computers to understand and interpret visual information from images and videos79,80,81,82,83,84. In the medical field, it supports a range of tasks and extracts clinically relevant information from medical images and videos, such as endoscopic recordings, CT scans, MRIs, and X-rays85,86. By examining CT and MRI scans, CV helps doctors pinpoint potentially life-threatening issues like internal bleeding and blocked blood vessels with greater accuracy. The use of automation, especially through robotic systems, enhances this precision by catching tiny details that might escape the human eye. This boost in diagnostic ability ultimately elevates the overall quality of patient care87,88,89. CV can be applied in a range of healthcare settings, from early disease detection to patient health monitoring during and after surgical procedures. Its integration into healthcare has developed into a ground-breaking tactic that is altering how medical data is viewed, assessed, and utilised90,91,92.
The role of artificial intelligence in cancer diagnosis
The AI-assisted technologies and algorithms have permeated the healthcare industry with great success. The use of these technologies in the diagnosis, screening, and treatment of different types of cancer is attracting enormous attention93. For instance, machine learning, a subset of AI that functions based on analysing large datasets, can be used in decision-making94. Similarly, deep learning, a subset of machine learning, is another key tool, based on an artificial neural network of interconnected layers called computing units95. Moreover, several deep learning computer vision (DLCV) models have been developed with important applications in healthcare96. These technologies are rapidly expanding and being applied in different phases of cancer diagnosis and treatment, with added benefits of accuracy, sensitivity, and improved outcomes.
AI in cancer imaging
AI has demonstrated a lot of promise in the modern era for tumour identification and description. AI is a first line of defence against omission errors and lowers observational errors, whereas detection involves locating objects of interest on radiographs14,97,98. Characterisation can include prognosticating diseases and forecasting results based on specific treatment modalities, in addition to staging and segmenting tumours (Fig. 3). This information is crucial for later diagnostic evaluations and for calculating radiation dosages during treatment planning99,100,101,102,103. Despite being widely used as the standard for evaluating automated algorithms, manually traced segmentations may overlook subclinical conditions and restrict analyses because of human biases98,104. Through automated segmentation, AI has the potential to significantly improve tumour measurement accuracy, efficiency, and reproducibility105,106,107.
Role of AI in the diagnosis of different types of cancers, which includes imaging, detection, characterisation, staging, and segmentation of tumours.
Given that the normal clinical course of a cancer patient identifies multiple crucial phases where AI systems can offer significant advantages, ML and DL present a variety of opportunities to improve cancer imaging98,108. For instance, one of the effective uses of imaging technologies is to perform cancer risk assessment. In the United States, several states require breast density assessment as part of cancer risk evaluation. DL systems have shown high accuracy in classifying breast density, enabling more consistent patient notification in breast cancer screening109,110. Studies have shown that the DL model integrating mammographic features with established risk factors outperformed conventional breast cancer risk models in identifying individuals at the highest risk of malignancy111,112. More recently, strong agreement in breast cancer risk assessment was reported when mammographic breast density was evaluated by a senior radiologist, a junior radiologist, and AI software113.
AI algorithms are increasingly enhancing cancer screening by improving sensitivity and specificity through the analysis of large imaging and biomarker datasets114. In breast cancer screening, a study by McKinney et al. has revealed that AI models can match expert reader performance and, in some cases, exceed it by an absolute margin of 11.5%. These models can also support second readers during mammographic review, reducing their workload by up to 88%115. Additionally, AI-assisted triaging, which prioritises image interpretation, has been found acceptable by women undergoing mammographic screening116,117. In a recent study by Eisemann et al.118, 463,094 women underwent screening, of whom 260,739 were examined with AI support by 119 radiologists. The AI-supported group achieved a breast cancer detection rate of 6.7 per 1000, which was 17.6% higher and statistically superior to the rate of 5.7 per 1000 observed in the control group. In another study, Jiang et al.119 attempted to investigate whether the diagnostic performance of radiologists in differentiating cancer from noncancer on breast MRI improves when using an AI system compared with conventionally available software. The study concluded that the performance of radiologists to differentiate benign from malignant breast cancer significantly increased from 0.71 to 0.76 with the help of AI algorithms119.
Image quality and signal-to-noise ratio are inversely related to radiation dose, making it impossible to arbitrarily reduce the radiation exposure per examination. Deep neural networks can map data from one high-dimensional space to another120,121,122,123. For instance, transforming a low-dose, high-noise CT image into a high-dose, low-noise representation. Accordingly, novel approaches based on deep learning reconstruction (DLR) are being developed, with the potential to reduce radiation dose significantly. Two DLR solutions received FDA clearance and became clinically available in 2019124. DLR-based reconstruction methods have demonstrated lower radiation doses and/or improved image quality, while maintaining reasonably short reconstruction times. A pilot DLR study recently reported volumetric tomographic imaging generated from ultrasparse data sampling combined with a patient-specific prior125, which could further reduce radiation dose.
In DL applications, convolutional neural networks (CNNs) have been designed and validated for malignancy risk estimation. Using 16,077 lung nodules from the National Lung Screening Trial, one such algorithm by Venkadesh et al. demonstrated strong performance, achieving an area under the curve (AUC) of 0.93126. Zhang et al. developed an ML-based logistic regression model using CT images from 108 patients to predict and stratify pathological low and high-grade bladder cancer127. On a validation cohort of 37 patients, the model achieved an AUC of 0.86127. In another attempt, Fan et al. employed radiomics features from CT scans with 119 stage II colorectal cancer patients to train and test an ML model, which predicted microsatellite instability status with an AUC of 0.75128.
AI in cancer pathology
For many years, pathology, particularly histological evaluation, has been the gold standard for cancer diagnosis. Tissue-based analysis provides essential information on tumour type, grade, stage, and biomolecular characteristics to guide clinical decision-making129,130,131. However, there are several disadvantages to conventional pathology. Manual slide interpretation is time-consuming and often subject to inter-observer variability, particularly in complex situations132,133. In recent times, a variety of AI tools have emerged to assist with tumour evaluation. One notable example is Paige Prostate, which uses AI to identify prostate cancer by marking suspicious spots in biopsy samples. This support is invaluable for pathologists, ensuring that they don’t miss any potentially harmful areas134. Paige Lung is another AI tool for lung cancer detection that helps pathologists find early-stage cancers more precisely, including small nodules and subtle features that conventional analyses might miss135. A study by Eloy et al. found that Paige Prostate significantly reduced the need for immunohistochemistry (IHC) tests, second opinion requests, and reporting times while maintaining high diagnostic accuracy136.
In digital pathology, a wide range of AI technologies have been introduced in image processing and classification26,137,138. Several studies have reported the integration of AI in the identification, detection, and segmentation of tissues. Using a deep neural network (DNN), Chen and colleagues developed a model that aimed at automatic lymphocyte detection during H&E images. In another study, Ehteshami et al. designed a deep learning algorithm-based model to detect metastasis of lymph nodes in women with breast cancer139,140. Their model revealed comparable performance with an expert pathologist. Time is a critical factor in conventional pathology; however, several models in digital pathology have focused on automating the processes with reduced time consumption and better outcomes141,142,143.
PathAI assists in the identification and categorisation of various subtypes of breast cancer by precisely analysing histopathological images. Additionally, it has been used to ascertain if a patient has HER2144,145. AI-driven endoscopic lesion detection systems in gastrointestinal pathology improve diagnostic accuracy and reduce missed lesions by identifying and characterising lesions like polyps during colonoscopy through real-time image analysis146.
An important development in breast cancer grading is the application of DL models to improve the stratification of intermediate Nottingham Histological Grade (NHG) 2 cases, which have traditionally been challenging to diagnose due to their variability and unclear prognostic value. These models, which detect minute morphological patterns in whole-slide histopathology images, can be used to categorise NHG 2 tumours into lower- and higher-risk groups147. This method uses standard Haematoxylin and Eosin (H&E) slides and is quicker, less expensive, and yields prognostic data that is comparable to molecular assays. In a study, Mantrala et al.148 showed that AI could grade nuclear pleomorphism with performance on par with human experts, reducing pathologists’ inconsistencies148. Their research demonstrated that AI is capable of identifying important nuclear morphological characteristics that determine tumour grade and provide reliable survival stratification for a range of patient populations.
Without the need for labelled datasets, self-supervised learning (SSL) is an unsupervised method that aims to learn meaningful data representations. By comparing several representations of the same input, contrastive learning, a well-known SSL technique, trains models to distinguish between similar and dissimilar examples149,150,151,152,153,154. When Ciga et al. used contrastive learning on 57 different unlabelled breast cancer (BC) histopathology datasets, they discovered that their model performed better on datasets specific to BC than ImageNet-pretrained networks155,156. Similarly, Ashraf et al. classified BC histopathological images efficiently using contrastive learning and a new ResNet50-Inception hybrid architecture, reaching up to 98% accuracy with only 3.6 million parameters157.
Identifying small, occult tumour foci in lymph nodes remains challenging with conventional pathology and often necessitates additional confirmatory tests158,159. Promising approaches for more accurate lymph node metastasis detection are provided by recent developments in AI, especially DL, which may lessen the need for IHC and save time and money160,161,162,163,164. An algorithm created by Liu et al. can accurately detect metastatic cancer cells, including very tiny foci, in sentinel lymph node biopsies165. Their findings showed that even with typical variations in tissue samples, the algorithm continued to perform robustly. Steiner et al. assessed the function of DL support in pathologists’ examination of lymph nodes for metastases of breast cancer in a different study. Their results demonstrated that integrating AI decreased review time and error rates while increasing diagnostic accuracy, particularly for hard-to-detect micrometastases164. Bándi et al.166 investigated continual learning strategies for cancer-independent detection of lymph node metastases. This method eliminates the need for cancer-specific retraining by allowing AI models to generalise across a variety of cancer types, such as breast, colon, and head-and-neck cancers. Across a variety of datasets, their findings showed excellent accuracy and dependability166. Radiomics may be able to easily integrate into digital pathology workflows when paired with DL-based and cancer-independent AI models for lymph node metastasis detection.
Artificial intelligence in cancer genomics
Precision oncology is undergoing a dramatic change as a result of the integration of AI and genomics. By identifying somatic and germline pathogenic variants, advances in high-throughput sequencing technologies, especially next-generation sequencing (NGS), have completely changed our understanding of tumour biology167,168,169,170,171,172. Simultaneously, AI, particularly through ML, which finds patterns in data without explicit programming, and deep learning (DL), a more sophisticated method that uses artificial neural networks, are turning into a crucial tool for extracting clinically significant information from intricate, high-dimensional genomic datasets5,173.
Role of AI in mutation detection
Histopathological and genomic diagnostic techniques are used in precision oncology to help patients choose the best treatments for them. H&E staining is one of the most popular methods in histopathology, which analyses tumour morphology and phenotype and is still essential for cancer diagnosis and subtyping26,174. With their ability to predict patient survival and treatment response, genomic biomarkers are frequently used in advanced or metastatic cancers to supplement phenotypic assessment. Thus, more individualised treatment approaches have been made possible by genomics26. Over the past few decades, these developments have helped to significantly improve clinical results175,176.
Multiomics is the integrated analysis of multiple layers of biological information, including genomics, proteomics, and epigenetics, to achieve a more holistic understanding of biological systems177. Given that cancer is a complex, multifactorial disease with changes at several molecular levels, integrating these disparate datasets is especially crucial in precision oncology. Through the integration of knowledge from various omics layers, scientists and medical professionals can better describe tumour heterogeneity, find more accurate biomarkers, and establish customised therapeutic targets, ultimately leading to the development of more specialised cancer treatments178.
Structural variations (SVs) are genomic changes that involve large DNA segments (≥50 bp). These are referred to as copy-number variations (CNVs) and can be caused by a variety of mutational events, including duplications, insertions, and deletions, which alter the amount of genomic material179,180,181. Although CNVs have been analysed using a variety of techniques, whole-genome sequencing (WGS) is becoming more and more accepted as the preferred approach due to declining costs and ongoing advancements in variant detection technologies182.
On the other hand, small variants are defined as genomic changes that impact shorter DNA segments (<50 bp). These include small insertions or deletions (indels), which usually range from one to a few dozen base pairs, and single-nucleotide variants (SNVs), in which only one base is changed. Such variants, though smaller in scale, can have significant functional consequences, particularly when they occur within coding or regulatory regions183,184,185,186.
Cancer initiation and progression are greatly influenced by germline pathogenic variants (gPVs), which are inherited, and somatic pathogenic variants (sPVs), which develop within tumour cells187,188. The presence of gPVs in the BRCA1 and BRCA2 genes, which predispose carriers to breast and ovarian cancers, is one of the most well-characterised hereditary signatures in oncology189. High- and moderate-penetrance genes like TP53, PALB2, STK11, CHEK2, RAD51D, and ATM are among the other genes linked to hereditary breast cancer190. Cancer biology and treatment also heavily rely on somatic PVs in tumour suppressors and oncogenes191,192,193. For example, KRAS mutations are important in colorectal and pancreatic cancers because they promote uncontrolled cell growth, whereas TP53 mutations impair a vital tumour suppressor called the “guardian of the genome,” which leads to genomic instability, metastasis, and a poor prognosis194.
Recently, AI tools like ML, DL, CNNs, and MIL have shown promise in detecting genetic mutations in cancer195,196,197,198 (Fig. 4). MRI, PET, digital tomosynthesis, and mammography are among the imaging modalities for which researchers have created radiomic signatures that allow for the noninvasive prediction of genetic changes199,200,201. AI techniques for mutation prediction from image data are made possible by the growing digitisation of H&E slides in pathology. These methods are especially helpful when there is a shortage of tumour tissue and also cut down on the time and expense of genetic testing. Recent research using AI has demonstrated promising outcomes in the diagnosis, classification, and grading of various cancer types202,203,204,205,206,207 as well as in the detection of gene mutations, microsatellite instability, and treatment response208,209,210,211,212,213.
This figure demonstrates how AI supports cancer genomics by analysing sequencing data to detect cancer-associated mutations and integrating multi-omics data to discover clinically relevant biomarkers for diagnosis and therapy.
One prevalent and important pathophysiological characteristic of cancer is mutations in genes that control cell division and apoptosis. These mutations affect the course of the disease, patient survival, and treatment approaches in addition to aiding in the development of cancer214,215. Numerous AI-based models have demonstrated efficacy in detecting mutations in a variety of cancer types, such as gastrointestinal, brain, lung, and breast cancers216. A pre-trained 3D CNN was employed by Shao et al.217 to identify EGFR mutations in lung adenocarcinoma. After entering patients’ PET/CT scans into the model, they discovered that combining PET/CT scans with clinical information produced the best diagnostic results (AUROC: 0.73)217. Similarly, using DL methods, Dammak et al.218 aimed to identify high tumour mutation burden in lung squamous cell carcinoma using histopathological images, with their models achieving an AUC ranging from 0.6 to 0.8.
Liang et al.219 proposed a CNN model to identify PDGFRA and KIT mutations in gastrointestinal stromal tumours (GIST) from histological images. Using pre-trained models on ImageNet, their approach predicted these drug-sensitive mutations with an accuracy of 70–85%. In another study, Cao et al.220 trained their model using colorectal cancer histological images from the TCGA-COAD database and applied transfer learning (TL) to adapt it to the Asian-CRC dataset. Initially, the model trained on TCGA-COAD achieved an AUROC of 0.6497 on Asian-CRC. After implementing TL, performance improved to an AUROC of 0.8504, and further increasing the number of Asian-CRC cases used in fine-tuning boosted the AUROC to 0.9264220. Similarly, Zeng et al.220 investigated the identification of IDH mutation types in glioma patients using multi-modal MRI data. They employed a pre-trained ImageNet model for feature extraction, achieving an AUROC of 0.86 in predicting IDH status, with a sensitivity of 77.78% and a specificity of 75%220.
In 2020, Yuhan Su et al.221 applied BERT to classify genetic mutations by analysing textual evidence from an annotated database, demonstrating improved capability in handling complex clinical text. Among the three BERT-based approaches tested, the BERT with the abstract shortness method performed best as a standalone model, achieving an F1 score of 0.705221. More recently, in 2023, Sanad Aburass and colleagues developed a hybrid ensemble model integrating GRU, LSTM, CNN BiLSTM, and GloVe embeddings. Using Kaggle’s Personalised Medicine dataset, this model achieved an F1 score of 0.831 for classifying gene mutations in cancer222. Sun et al.223 developed a deep neural network (DNN) model using genomic point mutations to classify tissues as either one of 12 TCGA cancer types or healthy controls from the 1000 Genomes Project. Trained on the most frequent cancer-specific point mutations derived from whole-exome sequencing data, the classifier achieved high accuracy in distinguishing healthy from tumour tissues (AUC = 0.94). However, its performance declined in the multi-class classification task of differentiating all 12 cancer types simultaneously, with an AUC of 0.70223.
In prostate cancer, one study combined ML techniques with features recommended by the National Institute for Health and Care Excellence to explore associations between prognosis and genetic mutation profiles224. ML has been used in other studies to predict the relationship between colorectal cancer and being overweight using healthy eating index scores225. Furthermore, by comparing the performance of various machine learning models, modelling techniques have shown the value of the exact binomial test for examining genome-wide somatic mutations226. Cho et al. used TCGA data from patients with lung adenocarcinoma to further demonstrate the value of machine learning in cancer genomics. These studies demonstrate how AI can enhance the detection of mutations in large cancer datasets and provide vital information on genetic alterations linked to the outcomes of patients with lung cancer227.
The role of AI in biomarker discovery
AI in oncology has revolutionised cancer diagnosis, prognosis, and treatment due to the increasing demand for precision medicine228. One of AI’s most important contributions has been the identification of biomarkers, which are crucial for precision oncology. Biomarkers, which are genes, proteins, or other measurable substances that indicate the occurrence or course of disease, have long been used in cancer diagnosis and treatment planning. If the complex nature of cancer necessitates finding novel therapeutic strategies in which important molecular pathways could be targeted, at the same time, it highlights the need for discovering biomarkers that can help in early detection and treatment selection229,230. Contrary to anatomic and qualitative-based markers, the focus has been shifted to find non-invasive and more specific biomarkers that could better characterise survival prediction, tumour phenotype, and prediction of treatment response231,232. The emergence of AI-assisted technologies and advanced deep learning approaches may mark a new era for non-invasive biomarkers in cancer care. AI improves diagnostic accuracy by evaluating intricate histopathological images233. These methods have proven effective in differentiating between cancer patients and healthy people, advancing precision medicine and the creation of tailored cancer treatments234.
Potential biomarkers and pathways in glioblastoma were investigated experimentally using blood-based liquid biopsies, with a focus on platelets235. This method found seven enriched pathways and 42 genes, including those linked to immunological regulation, neurodegenerative processes, and protein translation. A multi-step machine learning framework was used to relate the results to 12-month survival. Along with related proteins, the analysis also looked at CD133, a putative glioblastoma biomarker. Using a multi-step machine learning framework, it connected these results to 12-month survival. In a different investigation, Liu et al.236 developed a gene signature based on machine learning that included 12 T-cell exhaustion-related subtypes in glioblastoma. Using transcriptional profiling of T-cell immunity in the tumour microenvironment, the authors discovered two hitherto unknown subtypes, TEX-C1 and TEX-C2. These subtypes showed different immunological and clinical profiles, according to multi-omics analysis236.
Studies have suggested that AI-driven predictive frameworks can play a key role in the pre-treatment evaluation237,238. To create an ML-based predictive framework for pretreatment evaluation, Kawakami et al.239 divided 334 patients with ovarian epithelial carcinoma and 101 patients with benign ovarian tumours into training and test groups in 2019. The model incorporated 32 biomarkers and clinical parameters from standard blood tests using classifiers like RF. At 92.4%, the RF classifier successfully differentiated ovarian epithelial carcinoma from benign tumours; however, its accuracy in predicting clinical stage was lower at 69.0%. These studies confirm the importance of biomarkers in the early detection, diagnosis, and treatment of cancer.
Integration of AI into cancer drug discovery and development
Through ML and DL, AI improves compound optimisation, expedites target identification, enables rapid analysis of large biomedical datasets, and improves clinical outcome prediction240. AI assists with lead optimisation and de novo drug design in addition to screening by generating novel molecular structures and refining their properties to improve efficacy and safety241. Predictive modelling helps with preclinical development, especially when using in silico simulations to assess toxicity and pharmacokinetics. Additionally, AI speeds up the discovery of biomarkers, assisting in the identification of molecular signatures for patient stratification and individualised treatment plans. At the clinical stage, AI incorporates real-world data to improve patient recruitment, streamline trial design, and reduce turnaround times, all of which raise the success rate of developing cancer drugs241,242.
Advanced models that can learn from high-dimensional biological data are becoming more and more important in the integration of AI into cancer drug discovery and development. This change is best illustrated by transformer-based frameworks used in single-cell transcriptomics, which model tumour–immune interactions, reconstruct gene regulatory networks, and capture tumour heterogeneity243. By combining translational applications with basic cancer genomics, these methods speed up the discovery of new drug targets, forecast resistance mechanisms, and improve treatment approaches (Fig. 5).
In silico drug response prediction, regulatory network inference, and precise tumour clustering are made possible by the method.
AI in drug target identification and validation
Since AI integrates and analyses a variety of biological datasets, it is becoming more and more significant in drug target identification. In contrast to conventional methods, AI-driven models combine data from proteomics, genomics, and other high-throughput data sources to more effectively identify new targets244,245. For example, it is possible to identify candidate genes and their protein products as possible therapeutic targets by mining genomic data to find genetic variations linked to disease246. Similarly, proteomic analyses allow AI to examine protein functions, structures, and interaction networks, providing details about disease pathways and target druggability247. For better systematic evaluation and target identification, several platforms could be integrated with AI248,249, such as DrugBank250, Antibiotic Adjuvant DataBase251, PubChem252, and the Antibiotic Combination DataBase240.
Fahimian et al.253 developed a novel algorithm called RepCOOL to identify potential candidates for the treatment of stage II breast cancer253. The discovery of novel GSK-3 inhibitors for the treatment of acute myeloid leukaemia is another significant advancement254. At the same time, ligand-based virtual screening methods have employed isolation forests, artificial neural networks, and Gaussian mixture models to expedite drug identification and improve prediction accuracy255.
In 2020, Yin et al.256 introduced DeepDrugw, a deep learning framework for predicting drug-target interactions (DTI), and drug-drug interactions (DDI), using graph-based techniques257. Similarly, by employing a graph autoencoder based on GCN to capture higher-order structural relationships within heterogeneous networks, Qu et al.258 demonstrated graph–drug–target interactions, enabling a more accurate representation of drugs, proteins, and their surrounding contexts259. Recently, Yu et al.260 proposed a heterogeneous GNN that combines Bi-LSTM with an attention mechanism to improve heterogeneous data aggregation and node feature extraction. To further improve model performance, a negative sampling strategy was added.
Several approaches have been used in recent studies to predict unknown drug targets, such as disease-related molecular networks261, drug–drug similarity networks, protein–protein similarity networks262, and drug side-effect networks263. To predict DTIs based on drug-target similarity networks, Tian et al.264 Sadaqat et al.265 successfully identified important target proteins by using molecular modelling, network pharmacology, and sequencing data to examine the effects of Bacopa monnieri on hepatocellular carcinoma. However, these models often have drawbacks, such as unbalanced or incomplete datasets, a lack of validated drug–target data, and a limited capacity to incorporate intricate relationships between various databases.
Following identification, a therapeutic target needs to be thoroughly validated to ensure its applicability. Since target identification and validation are inherently complex and multidisciplinary processes, it is necessary to employ a range of validation techniques to increase confidence in the observed results266. Careful execution of these steps remains crucial for reducing attrition rates and avoiding ineffectiveness in later stages of development267. A recent study demonstrated the effective use of AI in locating and validating targets for the creation of cancer medications. Researchers used an AI-driven screening method to find Z29077885, a novel anticancer compound that targets STK33268. Both in vitro and in vivo tests verified the efficacy of Z29077885 after it was identified. Mechanistic analyses revealed that the compound inhibits the STAT3 signalling pathway, which results in cell cycle arrest at the S phase and apoptosis269.
AI in drug screening and lead optimisation
High throughput screening (HTS) employs automated systems, advanced detection tools, and microtiter plates to effectively gather and analyse data in its cellular or molecular scale experiments. This method enables the rapid evaluation of thousands to millions of compounds during the discovery phase, which is an essential strategy for identifying active compounds early in the drug development process256. AI-driven virtual screening, in particular, expedites the selection process and drastically reduces the number of compounds that require experimental validation by making it simpler to predict how molecules will interact with target proteins258.
AI can help overcome resistance to current treatments by facilitating the transformation of genes and proteins that are known or may be associated with resistance into novel therapeutic targets. To predict drug sensitivity, for instance, Xiao et al.270 used ridge regression models that were trained on transcriptomic profiles and drug response data. This approach was verified across multiple colorectal cancer cohorts. In vitro pharmacodynamic assays revealed that two BCL-XL inhibitors, WEHI-539 and navitoclax, exhibited selective sensitivity towards colorectal cancer cells with high chromosomal instability, confirming CIN as a promising therapeutic target in colorectal cancer270. Similarly, Zhang et al.271 integrated multi-omics datasets to predict patient responses to immune checkpoint inhibitors using a Bayesian model. They found that anti-tumour immunity was inversely correlated with a stemness signature271. The end-to-end DL framework developed by Wen et al.272 combines a transformer architecture with a self-supervised graph neural network. They filtered 4,527,000 compounds and found 50 candidate clusters by optimising the model on the BindingDB database272.
Synthetic lethality (SL) has gained more attention recently as a useful approach for identifying potential targets for anticancer drugs in cancer pateints273. However, high costs, batch variability, and off-target effects are frequently limited by experimental SL screening. To address these issues, Wang et al.274 presented KG4SL, a brand-new model based on graph neural networks (GNNs) that integrates knowledge graph (KG) messaging into the prediction procedure. Their findings showed that the accuracy and efficacy of SL prediction were significantly improved by incorporating KG information into the GNN framework274.
Docking and scoring methods are used in structure-based virtual screening to find molecules that have a high binding affinity for a target protein275. Traditional docking procedures are frequently computationally demanding and restrict the scalability of large-scale screening, even though this approach is a useful tool in anticancer drug design. In a study, a DL-based study tool was designed to predict the docking score for cancer patients276. In another study, Yasuo et al. introduced SIEVE-Score, an AI-driven structure-based virtual screening method for hit compound identification that demonstrated notable improvements over the most advanced techniques currently in use277.
An important factor in drug development is the possibility of strong interactions, which can be evaluated by using AI models to estimate the binding affinity between target proteins and candidate drug molecules278. A predictive model called DrugnomeAI was created by Raies et al.279 to tackle the problem of targeted drug synthesis. Targets’ druggability throughout the human exome was predicted using DrugnomeAI, which is based on a stochastic semi-supervised machine learning framework. The study was noteworthy for its ability to accurately predict the druggability of targets linked to oncological diseases.
AI in preclinical and clinical trials
AI has a lot of potential to improve clinical trial design and execution. Improvements in trial development can be directly attributed to developments in cancer imaging and diagnostics280. AI’s potential to expedite and simplify patient recruitment is yet another significant benefit280. 50,000 patients from 19 datasets participated in a systematic review and meta-analysis that showed AI-assisted approaches to cancer clinical trial enrolment performed on par with, and in some cases better than, conventional manual screening techniques281.
By predicting patient behaviour and clinical outcomes, AI technologies can help with patient retention in addition to trial enrolment282. One AI model, for example, was created to forecast grade 3 or 4 adverse events in patients with breast cancer undergoing neoadjuvant treatment283. Similarly, using information from a publicly accessible cancer centre database, an ML platform created outside of the trial context was used to estimate 30-day unplanned readmission rates in cancer patients284. These tools can help investigators implement dose adjustments, early interventions, or customised symptom management strategies by identifying individual risk profiles for adverse events, trial attrition, or hospital readmission.
ML can optimise the choice of treatment regimens during the design phase of clinical trials by using simulation techniques to analyse large datasets from previous studies. For instance, in a clinical study of advanced non-small cell lung cancer patients who had never received systemic therapy before, Zhao et al. used a reinforcement learning technique called Q-learning285. This technique identified individualised, optimal strategies for particular patient subgroups using patient data from clinical reinforcement trials. Likewise, meticulous patient cohort selection and clear recruitment standards are important factors that determine the success of clinical trials. Remarkably, 86% of trials fall short of their recruitment goals within the allotted time286. AI models have been used to assess and improve trial eligibility requirements to overcome this difficulty. For example, Liu et al.287 showed that many commonly used criteria, especially laboratory test thresholds, significantly limited enrolment by simulating clinical trials in a cohort of 61,094 patients with advanced non-small cell lung cancer using a computational framework.
Tumour heterogeneity continues to pose a major obstacle in assessing treatment responses. AI-driven models that integrate baseline and follow-up CT imaging provide a powerful means of predicting therapeutic efficacy and enhancing patient stratification in clinical trials287,288,289. More accurate immunotherapy outcome prediction in advanced non-small cell lung cancer (NSCLC) has been made possible by modelling based on multi-temporal CT image features, which supports better trial design and forecasting287. Better patient selection, more precise monitoring of treatment effects, and the development of individualised therapeutic approaches in clinical trials are all made possible by these methods. By enhancing patient recruitment, streamlining trial design, and facilitating real-time treatment response prediction, these studies show how AI is changing cancer clinical trials290. AI boosts productivity, lowers attrition, and aids in the creation of customised treatments by integrating multi-modal data and sophisticated modelling. Accelerated patient delivery of safer, more effective cancer medications will depend on the ongoing validation and uptake of AI tools269,291.
Challenges and opportunities
AI has a wide range of uses in oncology, as this review article discusses, from drug discovery and personalised treatment to genomics and diagnosis. Large patient data sets can be integrated by AI-driven tools, which can improve chemotherapy, enable more accurate and timely prognosis, and possibly lower the risk of cancer progression15,292. While we have made significant advancements, challenges like patient data privacy and algorithmic bias have likely prevented the full-scale adoption of AI technologies in healthcare (Fig. 6). So, it is essential to find a middle ground between these exciting prospects and a careful consideration of the technological, ethical, and practical challenges.
The figure also demonstrates the current challenges and limitations in these models.
Despite the potential benefits of AI systems in cancer, we cannot ignore the significant flaws and risks that come with the use of AI293,294,295. One major concern is racial bias in various medical studies. For instance, several AI tools have been known to reflect biases in how they assess the health of African Americans, like the incorrect assumption that they have thicker skin or lower lung capacity compared to white patients296,297,298,299.
There are important legal and regulatory issues with the use of AI in cancer treatment that require careful thought198,300,301. In a scoping review, conducted by Chamouni et al. concluded that the most frequent ethical issue found in 20 articles was data privacy, especially genomic information and sensitive imaging data302. The second crucial issue they reported was the principle of non-maleficence, or not hurting patients, because in certain settings, AI tools come with the risk of losing lives if the system is unable to differentiate between real and false-positive cancer lesions302. Another important question arises: who will take the responsibility, the AI tool developer, clinicians, or the hospital that used that system?
Future studies should focus on AI-assisted multivariable prediction models, which can minimise the overall error and may improve outcomes in cancer care303,304. Moreover, stakeholders, including AI-experts, clinicians, and organisations, must move towards multi-institutional collaborations, which may pave the way for establishing reliable networks, AI models, and databases305,306,307,308. Issues related to patients’ privacy underlined the need for collaborations with legislators to find solutions that put patients’ privacy and safety first to achieve better and long-lasting results309. Furthermore, large, high-quality datasets are necessary for AI models, particularly DL approaches, and institutions must collaborate to gather, curate, and compile these datasets310,311.
To minimise the patient’s privacy-related issues, leakage of data must be prevented by using fixed algorithms that keep the patient’s data locked. It is important to make sure that federated learning and safe data-sharing frameworks are implemented because they can promote inter-institutional cooperation without jeopardising patient confidentiality. While designing AI models, the large datasets must be properly validated, annotated, and tested to have an idea about the errors before applying them to the final models. Data generated as a result of pathology, imaging, or endoscopy is often not labelled properly, and therefore should not be considered suitable for AI-algorithm training. Additionally, to guarantee generalisability, AI models that exhibit strong performance on small, single-institution datasets need to be verified across a variety of populations and institutions. To conclude, the emergence of AI has revolutionised cancer care, but we are still short of successfully developed models with real-world applications. Collective efforts are needed among clinicians, AI model developers, and policymakers to assist in designing specialised models with robust policy and regulatory implementations. This will promote an atmosphere of trust accuracy and confidence in the use of AI tools in actual clinical settings.
Conclusion
Artificial intelligence and its associated algorithms, such as machine learning and deep learning, will continue to evolve in cancer care. Currently, AI algorithms such as ML, DL, GNN, NN, and NLP have already revolutionised the diagnosis, prognosis, screening, biomarker discovery, digital pathology, drug response, outcome prediction, and monitoring. The emergence of ML and DL-based AI algorithms has resulted in significant accuracy and improved outcomes in digital pathology. AI algorithms that combine genomics, radiomics, and other clinical parameters have yielded promising results. NN has been widely used in cancer imaging combined with other tools for better sensitivity, accuracy, and predictive values. AI-based CNN-CAD systems have demonstrated significantly improved efficacy and accuracy in clinical oncology312,313.
The studies we reviewed show that the use of AI has improved the radiologists’ performances, treatment response, diagnostic accuracy, and decision-making in handling complex cases. However, these AI algorithms have not yet fully evolved, so they have not been integrated into routine clinical practices. Most of the existing AI tools are not fully validated, and there is a space of mistrust between both patients and clinicians. To achieve greater outcomes, both patients and clinicians must know the credibility and capability of these systems. Furthermore, healthcare AI regulations are still developing, underscoring the critical need for robust governance frameworks that promote innovation while guaranteeing safety and efficacy for cancer patients.
Data availability
No datasets were generated or analysed during the current study.
References
Upadhyay, A. Cancer: An unknown territory; rethinking before going ahead. Genes Dis. 8, 655–661 (2021).
Bourgeois, A. et al. Barriers to cancer treatment for people experiencing socioeconomic disadvantage in high-income countries: a scoping review. BMC Health Serv. Res. 24, 670 (2024).
Bray, F. et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 74, 229–263 (2024).
Majérus, M.-A. The cause of cancer: The unifying theory. Adv. Cancer Biol.—Metastasis 4, 100034 (2022).
Song, Q., Merajver, S. D. & Li, J. Z. Cancer classification in the genomic era: five contemporary problems. Hum. Genom. 9, 27 (2015).
Geéron A. Hands-on Machine Learning with Scikit-Learn and Tensor Flow: Concepts, Tools, and Techniques to Build Intelligent Systems (E-book, O’Reilly, 2017).
Basu, K., Sinha, R., Ong, A. & Basu, T. Artificial intelligence: how is it changing medical sciences and its future? Indian J. Dermatol. 65, 365–370 (2020).
Xu, Y. et al. Artificial intelligence: a powerful paradigm for scientific research. Innovation 2, 100179 (2021).
Rajpurkar, P., Chen, E., Banerjee, O. & Topol, E. J. AI in health and medicine. Nat. Med. 28, 31–38 (2022).
Yan, K., Wang, Y., Shao, Y. & Xiao, T. Gene Instability-Related lncRNA Prognostic Model of Melanoma Patients via Machine Learning Strategy. J Oncol 2021, 5582920 (2021).
LeCun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature 521, 436–444 (2015).
Guo, L. et al. Random-forest algorithm based biomarkers in predicting prognosis in the patients with hepatocellular carcinoma. Cancer Cell Int. 20, 251 (2020).
Rajpurkar, P. et al. Deep learning for chest radiograph diagnosis: a retrospective comparison of the CheXNeXt algorithm to practicing radiologists. PLoS Med 15, e1002686 (2018).
Sebastian, A. M. & Peter, D. Artificial intelligence in cancer research: trends, challenges and future directions. Life 12, 1991 (2022).
Alowais, S. A. et al. Revolutionizing healthcare: the role of artificial intelligence in clinical practice. BMC Med. Educ. 23, 689 (2023).
Gillum, R. F. From papyrus to the electronic tablet: a brief history of the clinical medical record with lessons for the digital age. Am. J. Med. 126, 853–857 (2013).
Post, A. R., Burningham, Z. & Halwani, A. S. Electronic Health Record data in Cancer Learning Health Systems: challenges and opportunities. JCO Clin. Cancer Inf 6, e2100158 (2022).
Chen, X. et al. A CT-based radiomics nomogram for prediction of lung adenocarcinomas and granulomatous lesions in patient with solitary sub-centimeter solid nodules. Cancer Imaging 20, 45 (2020).
Beig, N. et al. Perinodular and intranodular radiomic features on lung CT images distinguish adenocarcinomas from granulomas. Radiology 290, 783–792 (2019).
Ardila, D. et al. End-to-end lung cancer screening with three-dimensional deep learning on low-dose chest computed tomography. Nat. Med. 25, 954–961 (2019).
Baldwin, D. R. et al. External validation of a convolutional neural network artificial intelligence tool to predict malignancy in pulmonary nodules. Thorax 75, 306–312 (2020).
Roth, H. R. et al. Improving computer-aided detection using convolutional neural networks and random view aggregation. IEEE Trans. Med. Imaging 35, 1170–1181 (2016).
Spadaccini, M. et al. Discovering the first US FDA-approved computer-aided polyp detection system. Future Oncol 18, 1405–1412 (2022).
Zhang, S. M., Wang, Y. J. & Zhang, S. T. Accuracy of artificial intelligence-assisted detection of esophageal cancer and neoplasms on endoscopic images: a systematic review and meta-analysis. J. Dig. Dis. 22, 318–328 (2021).
Baik, Y. S., Lee, H., Kim, Y. J., Chung, J. W. & Kim, K. G. Early detection of esophageal cancer: evaluating AI algorithms with multi-institutional narrowband and white-light imaging data. PLoS ONE 20, e0321092 (2025).
Bera, K., Schalper, K. A., Rimm, D. L., Velcheti, V. & Madabhushi, A. Artificial intelligence in digital pathology—new tools for diagnosis and precision oncology. Nat. Rev. Clin. Oncol. 16, 703–715 (2019).
Shebib, R. et al. Randomized controlled trial of a 12-week digital care program in improving low back pain. NPJ Digit. Med. 2, 1 (2019).
Saltz, J. et al. Spatial organization and molecular correlation of tumor-infiltrating lymphocytes using deep learning on pathology images. Cell Rep. 23, 181–93.e7 (2018).
Esteva, A. et al. Dermatologist-level classification of skin cancer with deep neural networks. Nature 542, 115–118 (2017).
Yu, L., Chen, H., Dou, Q., Qin, J. & Heng, P. A. Automated melanoma recognition in dermoscopy images via very deep residual networks. IEEE Trans. Med. Imaging 36, 994–1004 (2017).
Vasaikar, S. V., Straub, P., Wang, J. & Zhang, B. LinkedOmics: analyzing multi-omics data within and across 32 cancer types. Nucleic Acids Res. 46, D956–d63 (2018).
Wang, Y., Su, H., Lu, Y. & Li, H. Regulatory Role of Fatty Acid Metabolism-Related Long Noncoding RNA in Prostate Cancer: A Computational Biology Study Analysis. J Oncol 2023, 9736073 (2023).
Lee, J. S. et al. Harnessing synthetic lethality to predict the response to cancer treatment. Nat. Commun. 9, 2546 (2018).
Zhou, J. et al. Deep learning sequence-based ab initio prediction of variant effects on expression and disease risk. Nat. Genet. 50, 1171–1179 (2018).
Davis, R. J. et al. Pan-cancer transcriptional signatures predictive of oncogenic mutations reveal that Fbw7 regulates cancer cell oxidative metabolism. Proc. Natl. Acad. Sci. USA 115, 5462–5467 (2018).
Jaganathan, K. et al. Predicting splicing from primary sequence with deep learning. Cell 176, 535–48.e24 (2019).
Rahman, M. A. et al. Impact of artificial intelligence (AI) technology in healthcare sector: a critical evaluation of both sides of the coin. Clin. Pathol. 17, 2632010x241226887 (2024).
Sezgin, E., Huang, Y., Ramtekkar, U. & Lin, S. Readiness for voice assistants to support healthcare delivery during a health crisis and pandemic. NPJ Digit. Med. 3, 122 (2020).
Brill, T. M., Munoz, L. & Miller, R. J. Siri, Alexa, and other digital assistants: a study of customer satisfaction with artificial intelligence applications. In The Role of Smart Technologies in Decision Making (eds. Eleonora P. and Francesca S.) 35–70 (Routledge, 2022).
Woodman, R. J. & Mangoni, A. A. A comprehensive review of machine learning algorithms and their application in geriatric medicine: present and future. Aging Clin. Exp. Res. 35, 2363–2397 (2023).
Janiesch, C., Zschech, P. & Heinrich, K. Machine learning and deep learning. Electron. Mark. 31, 685–695 (2021).
Taye, M. M. Understanding of machine learning with deep learning: architectures, workflow, applications and future directions. Computers 12, 91 (2023).
Patil, S. et al. Reviewing the role of artificial intelligence in cancer. Asian Pac. J. Cancer Biol. 5, 189–199 (2020).
Moeskops, P. et al. (eds) Deep learning for multi-task medical image segmentation in multiple modalities. In International Conference on Medical Image Computing and Computer-assisted Intervention (Springer, 2016).
Al Sharkawy M. et al. Breast cancer detection using support vector machine technique applied on extracted electromagnetic waves. Appl. Comput. Electromagn. Soc. J. 292–301 (2012).
Li, M., Jiang, Y., Zhang, Y. & Zhu, H. Medical image analysis using deep learning algorithms. Front. Public Health 11, 1273253 (2023).
Wong, J., Horwitz, M. M., Zhou, L. & Toh, S. Using machine learning to identify health outcomes from electronic health record data. Curr. Epidemiol. Rep. 5, 331–342 (2018).
Yang, S., Varghese, P., Stephenson, E., Tu, K. & Gronsbell, J. Machine learning approaches for electronic health records phenotyping: a methodical review. J. Am. Med. Inf. Assoc. 30, 367–381 (2023).
Javaid, M., Haleem, A., Pratap Singh, R., Suman, R. & Rab, S. Significance of machine learning in healthcare: features, pillars and applications. Int. J. Intell. Netw. 3, 58–73 (2022).
Tayefi, M. et al. Challenges and opportunities beyond structured data in analysis of electronic health records. Wiley Interdiscip. Rev. Comput. Stat. 13, e1549 (2021).
Ngiam, K. Y. & Khor, I. W. Big data and machine learning algorithms for health-care delivery. Lancet Oncol 20, e262–e273 (2019).
Ahsan, M. M., Luna, S. A. & Siddique, Z. Machine-learning-based disease diagnosis: a comprehensive review. Healthcare (Basel) 10, 541 (2022).
Qasrawi, R. et al. The role of machine learning in infectious disease early detection and prediction in the MENA region: a systematic review. Inform. Med. Unlocked 56, 101651 (2025).
Mehta, J. & Majumdar, A. Rodeo: robust de-aliasing autoencoder for real-time medical image reconstruction. Pattern Recognit 63, 499–510 (2017).
Choudhary, K. et al. Recent advances and applications of deep learning methods in materials science. npj Comput. Mater. 8, 59 (2022).
Alzubaidi, L. et al. Review of deep learning: concepts, CNN architectures, challenges, applications, future directions. J. Big Data 8, 53 (2021).
Ahmed, S. F. et al. Deep learning modelling techniques: current progress, applications, advantages, and challenges. Artif. Intell. Rev. 56, 13521–13617 (2023).
Bengio, Y., Courville, A. & Vincent, P. Representation learning: a review and new perspectives. IEEE Trans. Pattern Anal. Mach. Intell. 35, 1798–1828 (2013).
Erfanian, N. et al. Deep learning applications in single-cell genomics and transcriptomics data analysis. Biomed. Pharmacother. 165, 115077 (2023).
Rainio, O. & Klén, R. Convolutional neural networks for tumor segmentation by cancer type and imaging modality: a systematic review. Netw. Model. Anal. Health Inform. Bioinform. 14, 58 (2025).
Gull, S., Akbar, S. & Khan, H. U. Automated detection of brain tumor through magnetic resonance images using convolutional neural network. BioMed. Res. Int. 2021, 3365043 (2021).
Alqazzaz, S. et al. Combined features in region of interest for brain tumor segmentation. J. Digit. Imaging 35, 938–946 (2022).
Ranjbarzadeh, R., Zarbakhsh, P., Caputo, A., Tirkolaee, E. B. & Bendechache, M. Brain tumor segmentation based on optimized convolutional neural network and improved chimp optimization algorithm. Comput. Biol. Med. 168, 107723 (2024).
Lin, W.-W. et al. 3D brain tumor segmentation using a two-stage optimal mass transport algorithm. Sci. Rep. 11, 14686 (2021).
Choi, E., Schuetz, A., Stewart, W. F. & Sun, J. Using recurrent neural network models for early detection of heart failure onset. J. Am. Med. Inf. Assoc. 24, 361–370 (2017).
Raval, D. & Undavia, J. N. A Comprehensive assessment of Convolutional Neural Networks for skin and oral cancer detection using medical images. Healthc. Anal. 3, 100199 (2023).
Chen, R., Stewart, W. F., Sun, J., Ng, K. & Yan, X. Recurrent neural networks for early detection of heart failure from longitudinal Electronic Health Record data: implications for temporal modeling with respect to time before diagnosis, data density, data quantity, and data type. Circ. Cardiovasc. Qual. Outcomes 12, e005114 (2019).
Hussain, J., Båth, M. & Ivarsson, J. Generative adversarial networks in medical image reconstruction: a systematic literature review. Comput. Biol. Med. 191, 110094 (2025).
Makhlouf, A., Maayah, M., Abughanam, N. & Catal, C. The use of generative adversarial networks in medical image augmentation. Neural Comput. Appl. 35, 24055–24068 (2023).
Alajaji, S. A. et al. Generative adversarial networks in digital histopathology: current applications, limitations, ethical considerations, and future directions. Mod. Pathol. 37, 100369 (2024).
Koshino, K. et al. Narrative review of generative adversarial networks in medical and molecular imaging. Ann. Transl. Med. 9, 821 (2021).
Nawab, K., Ramsey, G. & Schreiber, R. Natural language processing to extract meaningful information from patient experience feedback. Appl. Clin. Inf. 11, 242–252 (2020).
Maleki Varnosfaderani, S. & Forouzanfar, M. The role of AI in hospitals and clinics: transforming healthcare in the 21st century. Bioengineering (Basel, Switzerland) 11, 337 (2024).
Khurana, D., Koli, A., Khatter, K. & Singh, S. Natural language processing: state of the art, current trends and challenges. Multimed. Tools Appl. 82, 3713–3744 (2023).
Supriyono, W., Suyono, A. P. & Kurniawan, F. Advancements in natural language processing: implications, challenges, and future directions. Telemat. Inform. Rep. 16, 100173 (2024).
Serrano-Guerrero, J., Bani-Doumi, M., Chiclana, F., Romero, F. P. & Olivas, J. A. How satisfied are patients with nursing care and why? A comprehensive study based on social media and opinion mining. Inf. Health Soc. Care 49, 14–27 (2024).
Scharkow, M. Thematic content analysis using supervised machine learning: an empirical evaluation using German online news. Qual. Quant. 47, 761–773 (2013).
van Buchem, M. M. et al. Analyzing patient experiences using natural language processing: development and validation of the Artificial Intelligence Patient Reported Experience Measure (AI-PREM). BMC Med. Inf. Decis. Mak. 22, 183 (2022).
Javaid, M., Haleem, A., Singh, R. P. & Ahmed, M. Computer vision to enhance healthcare domain: an overview of features, implementation, and opportunities. Intell. Pharm. 2, 792–803 (2024).
Al-Oraiqat, A. M. et al. Method for determining treated metal surface quality using computer vision technology. Sensors 22, 6223 (2022).
Gumbs, A. A. et al. The advances in computer vision that are enabling more autonomous actions in surgery: a systematic review of the literature. Sensors (Basel, Switzerland) 22, 4918 (2022).
Dudek, P. et al. Sensor-level computer vision with pixel processor arrays for agile robots. Sci. Robot. 7, eabl7755 (2022).
Hassan, H. et al. Review and classification of AI-enabled COVID-19 CT imaging models based on computer vision tasks. Comput. Biol. Med. 141, 105123 (2022).
Wu, Z., Chen, Y., Zhao, B., Kang, X. & Ding, Y. Review of weed detection methods based on computer vision. Sensors 21, 3647 (2021).
Lee, J. O., Zhou, H. Y., Berzin, T. M., Sodickson, D. K. & Rajpurkar, P. Multimodal generative AI for interpreting 3D medical images and videos. NPJ Digit. Med. 8, 273 (2025).
Rani, K., Kumar, A., Kumar, S., Gupta, A. & Singh, A. Medical imaging using machine learning, computer vision and applications. Int. J. Eng. Sci. Emerg. Technol. 11, 260–268 (2023).
Sharma, A. et al. Computer vision based healthcare system for identification of diabetes & its types using AI. Meas. Sens. 27, 100751 (2023).
Awwad, S., Tarvade, S., Piccardi, M. & Gattas, D. J. The use of privacy-protected computer vision to measure the quality of healthcare worker hand hygiene. Int. J. Qual. Health Care 31, 36–42 (2019).
Wang, D. et al. Real world validation of an AI-based CT hemorrhage detection tool. Front. Neurol. 14, 1177723 (2023).
Buaka, E. S. D. & Moid, M. Z. I. AI and medical imaging technology: evolution, impacts, and economic insights. J. Technol. Transf. 49, 2260–2272 (2024).
Ali, H., Mohsen, F. & Shah, Z. Improving diagnosis and prognosis of lung cancer using vision transformers: a scoping review. BMC Med. Imaging 23, 129 (2023).
Eswaran, U. & Khang, A. Artificial intelligence (AI)-aided computer vision (CV) in healthcare system. In Computer Vision and AI-Integrated IoT Technologies in the Medical Ecosystem (eds. Alex K., Vugar A., Olena H. and Arvind K. S.) 125–137 (CRC Press, 2024).
Najar Najafi, N., Hajihassani, H. & Azimzadeh Irani, M. The impact of artificial intelligence on cancer diagnosis and treatment: a review. Cancer Inform 24, 11769351251371273 (2025).
Surur, F. M. et al. Unlocking the power of machine learning in big data: a scoping survey. Data Sci. Manag. 8, 519–535 (2025).
Taherdoost, H. Deep learning and neural networks: decision-making implications. Symmetry 15, 1723 (2023).
Chai, J., Zeng, H., Li, A. & Ngai, E. W. T. Deep learning in computer vision: a critical review of emerging techniques and application scenarios. Mach. Learn. Appl. 6, 100134 (2021).
Liang, M. et al. Low-dose CT screening for lung cancer: computer-aided detection of missed lung cancers. Radiology 281, 279–288 (2016).
Bi, W. L. et al. Artificial intelligence in cancer imaging: clinical challenges and applications. CA Cancer J. Clin. 69, 127–157 (2019).
Rastogi, D. et al. Deep learning-integrated MRI brain tumor analysis: feature extraction, segmentation, and survival prediction using replicator and volumetric networks. Sci. Rep. 15, 1437 (2025).
Zhao, X. et al. Deep bone oncology diagnostics: computed tomography based Machine learning for detection of bone tumors from breast cancer metastasis. J. Bone Oncol. 48, 100638 (2024).
Yuan, X. et al. Systematic review and meta-analysis of artificial intelligence for image-based lung cancer classification and prognostic evaluation. npj Precis. Oncol. 9, 300 (2025).
Yao, I. Z., Dong, M. & Hwang, W. Y. K. Deep learning applications in clinical cancer detection: a review of implementation challenges and solutions. Mayo Clin. Proc.: Digit. Health 3, 100253 (2025).
Das, S., Dey, M. K., Devireddy, R. & Gartia, M. R. Biomarkers in cancer detection, diagnosis, and prognosis. Sensors 24, 37 (2024).
Rusanov, B. et al. Guidance on selecting and evaluating AI auto-segmentation systems in clinical radiotherapy: insights from a six-vendor analysis. Phys. Eng. Sci. Med. 48, 301–316 (2025).
Sarkar, S., Teo, P. T. & Abazeed, M. E. Deep learning for automated, motion-resolved tumor segmentation in radiotherapy. npj Precis. Oncol. 9, 173 (2025).
Huang, J. et al. Application of artificial intelligence in medical imaging for tumor diagnosis and treatment: a comprehensive approach. Discov. Oncol. 16, 1625 (2025).
Fountzilas, E., Pearce, T., Baysal, M. A., Chakraborty, A. & Tsimberidou, A. M. Convergence of evolving artificial intelligence and machine learning techniques in precision oncology. npj Digit. Med. 8, 75 (2025).
Zhang, B., Shi, H. & Wang, H. Machine learning and AI in cancer prognosis, prediction, and treatment selection: a critical approach. J. Multidiscip. Health 16, 1779–1791 (2023).
Mohamed, A. A. et al. A deep learning method for classifying mammographic breast density categories. Med. Phys. 45, 314–321 (2018).
Arieno, A., Chan, A. & Destounis, S. V. A review of the role of augmented intelligence in breast imaging: from automated breast density assessment to risk stratification. Am. J. Roentgenol. 212, 259–270 (2019).
Yala, A., Lehman, C., Schuster, T., Portnoi, T. & Barzilay, R. A deep learning mammography-based model for improved breast cancer risk prediction. Radiology 292, 60–66 (2019).
Dembrower, K. et al. Comparison of a deep learning risk score and standard mammographic density score for breast cancer risk prediction. Radiology 294, 265–272 (2020).
Le Boulc’h, M. et al. Comparison of breast density assessment between human eye and automated software on digital and synthetic mammography: Impact on breast cancer risk. Diagn. Int. Imaging 101, 811–819 (2020).
Wu, G., Bajestani, N., Pracha, N., Chen, C. & Makary, M. S. Hepatocellular carcinoma surveillance strategies: major guidelines and screening advances. Cancers (Basel) 16, 3933 (2024).
McKinney, S. M. et al. International evaluation of an AI system for breast cancer screening. Nature 577, 89–94 (2020).
Dembrower, K. et al. Effect of artificial intelligence-based triaging of breast cancer screening mammograms on cancer detection and radiologist workload: a retrospective simulation study. Lancet Digit. Health 2, e468–e474 (2020).
Lennox-Chhugani, N., Chen, Y., Pearson, V., Trzcinski, B. & James, J. Women’s attitudes to the use of AI image readers: a case study from a national breast screening programme. BMJ Health Care Inform 28, e100293 (2021).
Eisemann, N. et al. Nationwide real-world implementation of AI for cancer detection in population-based mammography screening. Nat. Med. 31, 917–924 (2025).
Jiang, Y., Edwards, A. V. & Newstead, G. M. Artificial intelligence applied to breast MRI for improved diagnosis. Radiology 298, 38–46 (2021).
Feng, Q., Liu, Z. & Chen, C. L. P. Broad and deep neural network for high-dimensional data representation learning. Inf. Sci. 599, 127–146 (2022).
Cui, Z. & Grindrod, P. Mappings, dimensionality and reversing out of deep neural networks. IMA J. Appl. Math. 89, 2–11 (2024).
Sarker, I. H. Deep learning: a comprehensive overview on techniques, taxonomy, applications and research directions. SN Comput. Sci. 2, 420 (2021).
Javid, A. M., Venkitaraman, A., Skoglund, M. & Chatterjee, S. High-dimensional neural feature design for layer-wise reduction of training cost. EURASIP J. Adv. Signal Process. 2020, 40 (2020).
Singh, R., Wu, W., Wang, G. & Kalra, M. K. Artificial intelligence in image reconstruction: the change is here. Phys. Med. 79, 113–125 (2020).
Shen, L., Zhao, W. & Xing, L. Patient-specific reconstruction of volumetric computed tomography images from a single projection view via deep learning. Nat. Biomed. Eng. 3, 880–888 (2019).
Venkadesh, K. V. et al. Deep learning for malignancy risk estimation of pulmonary nodules detected at low-dose screening CT. Radiology 300, 438–447 (2021).
Kniep, H. C. et al. Radiomics of brain MRI: utility in prediction of metastatic tumor type. Radiology 290, 479–487 (2019).
Fan, S. et al. Computed tomography-based radiomic features could potentially predict microsatellite instability status in stage II colorectal cancer: a preliminary study. Acad. Radiol. 26, 1633–1640 (2019).
Tseng, L. J., Matsuyama, A. & MacDonald-Dickinson, V. Histology: the gold standard for diagnosis? Can. Vet. J. 64, 389–391 (2023).
Aljehani, M. R. et al. The importance of histopathological evaluation in cancer diagnosis and treatment. Int. J. Health Sci. 7, 3614–3623 (2023).
Yang, Z. et al. A foundation model for generalizable cancer diagnosis and survival prediction from histopathological images. Nat. Commun. 16, 2366 (2025).
Singh, N. N., Tandon, A. & Jayasankar, P. Strength, weakness, opportunities and challenges (SWOC) experience of histopathology image analysis, enhanced by artificial intelligence. J. Oral. Biol. Craniofacial Res. 15, 1057–1063 (2025).
Fatima, G. et al. Transforming diagnostics: a comprehensive review of advances in digital pathology. Cureus 16, e71890 (2024).
Campanella, G. et al. Clinical-grade computational pathology using weakly supervised deep learning on whole slide images. Nat. Med. 25, 1301–1309 (2019).
Viswanathan, V. S., Toro, P., Corredor, G., Mukhopadhyay, S. & Madabhushi, A. The state of the art for artificial intelligence in lung digital pathology. J. Pathol. 257, 413–429 (2022).
Eloy, C. et al. Artificial intelligence-assisted cancer diagnosis improves the efficiency of pathologists in prostatic biopsies. Virchows Arch 482, 595–604 (2023).
Wong, A. N. N. et al. Current developments of artificial intelligence in digital pathology and its future clinical applications in gastrointestinal cancers. Cancers (Basel) 14, 3780 (2022).
Salvi, M., Acharya, U. R., Molinari, F. & Meiburger, K. M. The impact of pre- and post-image processing techniques on deep learning frameworks: a comprehensive review for digital pathology image analysis. Comput. Biol. Med. 128, 104129 (2021).
Ehteshami Bejnordi, B. et al. Diagnostic assessment of deep learning algorithms for detection of lymph node metastases in women with breast cancer. JAMA 318, 2199–2210 (2017).
Oiseth, S. J. & Aziz, M. S. Cancer immunotherapy: a brief review of the history, possibilities, and challenges ahead. J Cancer Metastasis Treat 3, 250–61 (2017).
Sirinukunwattana, K. et al. Gland segmentation in colon histology images: the glas challenge contest. Med. Image Anal. 35, 489–502 (2017).
Liu, Y. et al. Detecting cancer metastases on gigapixel pathology images. arXiv preprint arXiv:170302442 (2017).
Wang, H. et al. Mitosis detection in breast cancer pathology images by combining handcrafted and convolutional neural network features. J. Med. Imaging (Bellingham) 1, 034003 (2014).
Soliman, A., Li, Z. & Parwani, A. V. Artificial intelligence’s impact on breast cancer pathology: a literature review. Diagn. Pathol. 19, 38 (2024).
Sandbank, J. et al. Validation and real-world clinical application of an artificial intelligence algorithm for breast cancer detection in biopsies. NPJ Breast Cancer 8, 129 (2022).
Pannala, R. et al. Artificial intelligence in gastrointestinal endoscopy. VideoGIE 5, 598–613 (2020).
Wang, Y. et al. Improved breast cancer histological grading using deep learning. Ann. Oncol. 33, 89–98 (2022).
Mantrala, S. et al. Concordance in breast cancer grading by artificial intelligence on whole slide images compares with a multi-institutional cohort of breast pathologists. Arch. Pathol. Lab. Med. 146, 1369–1377 (2022).
Tian, Y. et al. What makes for good views for contrastive learning? Advances in Neural Information Processing Systems 33, 6827–6839 (2020).
Abdulrazzaq, M. M. et al. Consequential advancements of self-supervised learning (SSL) in deep learning contexts. Mathematics 12, 758 (2024).
Espis, A., Marzi, C. & Diciotti, S. Comparative analysis of supervised and self-supervised learning with small and imbalanced medical imaging datasets. Sci. Rep. 15, 32345 (2025).
Richter, T., Bahrami, M., Xia, Y., Fischer, D. S. & Theis, F. J. Delineating the effective use of self-supervised learning in single-cell genomics. Nat. Mach. Intell. 7, 68–78 (2025).
Penarrubia, C., Valero-Mas, J. J. & Calvo-Zaragoza, J. Self-supervised learning for text recognition: a critical survey. Int. J. Comput. Vis. 133, 6221–6250 (2025).
Zhang, X. & Han, L. A generic self-supervised learning (SSL) framework for representation learning from spectral–spatial features of unlabeled remote sensing imagery. Remote Sens 15, 5238 (2023).
Ciga, O., Xu, T. & Martel, A. L. Self supervised contrastive learning for digital histopathology. Mach. Learn. Appl. 7, 100198 (2022).
Kang, X., Li, D. & Sun, R. Nanotechnology and natural killer cell immunotherapy: synergistic approaches for precise immune system adjustment and targeted cancer treatment in gastrointestinal tumors. Front. Med. (Lausanne) 12, 1647737 (2025).
Ashraf, F. B., Alam, S. M. & Sakib, S. M. Enhancing breast cancer classification via histopathological image analysis: leveraging self-supervised contrastive learning and transfer learning. Heliyon 10, e24094 (2024).
Hu, A. et al. The diagnosis and management of small and indeterminate lymph nodes in papillary thyroid cancer: preoperatively and intraoperatively. Front. Endocrinol. (Lausanne) 15, 1484838 (2024).
Tang, Q. et al. Preoperative MRI and CA19-9 for predicting occult lymph node metastasis in small pancreatic ductal adenocarcinoma (≤2 cm). BMC Med. Imaging 25, 318 (2025).
Huang, S. C. et al. Deep neural network trained on gigapixel images improves lymph node metastasis detection in clinical settings. Nat. Commun. 13, 3347 (2022).
Wang, R., Gu, Y., Zhang, T. & Yang, J. Fast cancer metastasis location based on dual magnification hard example mining network in whole-slide images. Comput. Biol. Med. 158, 106880 (2023).
Challa, B. et al. Artificial intelligence-aided diagnosis of breast cancer lymph node metastasis on histologic slides in a digital workflow. Mod. Pathol. 36, 100216 (2023).
Caldonazzi, N. et al. Value of artificial intelligence in evaluating lymph node metastases. Cancers (Basel) 15, 2491 (2023).
Steiner, D. F. et al. Impact of deep learning assistance on the histopathologic review of lymph nodes for metastatic breast cancer. Am. J. Surg. Pathol. 42, 1636–1646 (2018).
Liu, Y. et al. Artificial intelligence-based breast cancer nodal metastasis detection: insights into the black box for pathologists. Arch. Pathol. Lab. Med. 143, 859–868 (2019).
Bándi, P. et al. Continual learning strategies for cancer-independent detection of lymph node metastases. Med. Image Anal. 85, 102755 (2023).
Bhinder, B., Gilvary, C., Madhukar, N. S. & Elemento, O. Artificial intelligence in cancer research and precision medicine. Cancer Discov 11, 900–915 (2021).
Satam, H. et al. Next-generation sequencing technology: current trends and advancements. Biology (Basel) 12, (2023).
Sahoo, O. S. et al. Role of next-generation sequencing in revolutionizing healthcare for cancer management. MedComm–Future Med. 3, e70001 (2024).
Vashisht, V., Vashisht, A., Mondal, A. K., Woodall, J. & Kolhe, R. From genomic exploration to personalized treatment: next-generation sequencing in oncology. Current Issues Mol. Biol 46, 12527–12549 (2024).
Yap, T. A., Stadler, Z. K., Stout, L. A. & Schneider, B. P. Aligning germline cancer predisposition with tumor-based next-generation sequencing for modern oncology diagnosis, interception, and therapeutic development. Am. Soc. Clin. Oncol. Educ. Book 43, e390738 (2023).
Fang, Y. et al. Systematic Investigation of Tumor Microenvironment and Antitumor Immunity With IOBR. Med Research 1, 136–140 (2025).
Zhang, J., Li, H., Tao, W. & Zhou, J. GseaVis: An R Package for Enhanced Visualization of Gene Set Enrichment Analysis in Biomedicine. Med Research 1, 131–135 (2025).
Djuric, U., Zadeh, G., Aldape, K. & Diamandis, P. Precision histology: how deep learning is poised to revitalize histomorphology for personalized cancer care. NPJ Precis. Oncol. 1, 22 (2017).
Andre, F. et al. Genomics to select treatment for patients with metastatic breast cancer. Nature 610, 343–348 (2022).
Kato, S. et al. Real-world data from a molecular tumor board demonstrates improved outcomes with a precision N-of-One strategy. Nat. Commun. 11, 4965 (2020).
Chen, C. et al. Applications of multi-omics analysis in human diseases. MedComm 4, e315 (2023). (2020).
Correa-Aguila, R., Alonso-Pupo, N. & Hernández-Rodríguez, E. W. Multi-omics data integration approaches for precision oncology. Mol. Omics 18, 469–479 (2022).
Escaramís, G., Docampo, E. & Rabionet, R. A decade of structural variants: description, history and methods to detect structural variation. Brief. Funct. Genom. 14, 305–314 (2015).
Li, Y. et al. Patterns of somatic structural variation in human cancer genomes. Nature 578, 112–121 (2020).
Ho, S. S., Urban, A. E. & Mills, R. E. Structural variation in the sequencing era. Nat. Rev. Genet. 21, 171–189 (2020).
Pös, O. et al. Copy number variation: methods and clinical applications. Appl. Sci. 11, 819 (2021).
Kosugi, S. & Terao, C. Comparative evaluation of SNVs, indels, and structural variations detected with short- and long-read sequencing data. Hum. Genome Var. 11, 18 (2024).
Yang, L. A practical guide for structural variation detection in the human genome. Curr. Protoc. Hum. Genet. 107, e103 (2020).
Mahmoud, M. et al. Structural variant calling: the long and the short of it. Genome Biol 20, 246 (2019).
Ferlaino, M. et al. An integrative approach to predicting the functional effects of small indels in non-coding regions of the human genome. BMC Bioinform 18, 442 (2017).
Garraway, L. A. & Lander, E. S. Lessons from the cancer genome. Cell 153, 17–37 (2013).
Horak, P. et al. Standards for the classification of pathogenicity of somatic variants in cancer (oncogenicity): Joint recommendations of Clinical Genome Resource (ClinGen), Cancer Genomics Consortium (CGC), and Variant Interpretation for Cancer Consortium (VICC). Genet. Med. 24, 986–998 (2022).
Sessa, C. et al. Risk reduction and screening of cancer in hereditary breast-ovarian cancer syndromes: ESMO Clinical Practice Guideline. Ann. Oncol. 34, 33–47 (2023).
Calabrese, A., Von Arx, C., Tafuti, A., Pensabene, M. & De Laurentiis, M. Prevention, diagnosis and clinical management of hereditary breast cancer beyond BRCA1/2 genes. Cancer Treat. Rev. 129, 102785 (2024).
Chandrashekar, P. et al. Somatic selection distinguishes oncogenes and tumor suppressor genes. Bioinformatics 36, 1712–1717 (2020).
Dakal, T. C. et al. Oncogenes and tumor suppressor genes: functions and roles in cancers. MedComm 5, e582 (2024). (2020).
Singh, S. R. et al. Exploring the genetic orchestra of cancer: the interplay between oncogenes and tumor-suppressor genes. Cancer 17, 1082 (2025).
Schon, K. & Tischkowitz, M. Clinical implications of germline mutations in breast cancer: TP53. Breast Cancer Res. Treat. 167, 417–423 (2018).
Shah, S. A. et al. Explainable AI-based skin cancer detection using CNN, particle swarm optimization and machine learning. J. Imaging 10, 332 (2024).
Vilhekar, R. S. & Rawekar, A. Artificial Intelligence in Genetics. Cureus 16, e52035 (2024).
Ashayeri, H. et al. Transfer learning in cancer genetics, mutation detection, gene expression analysis, and syndrome recognition. Cancers (Basel) 16, 2138 (2024).
Tiwari, A., Mishra, S. & Kuo, T.-R. Current AI technologies in cancer diagnostics and treatment. Mol. Cancer 24, 159 (2025).
Liu, Q. & Hu, P. Extendable and explainable deep learning for pan-cancer radiogenomics research. Curr. Opin. Chem. Biol. 66, 102111 (2022).
Qi, Y., Zhao, T. & Han, M. The application of radiomics in predicting gene mutations in cancer. Eur. Radiol. 32, 4014–4024 (2022).
Bodalal, Z., Trebeschi, S., Nguyen-Kim, T. D. L., Schats, W. & Beets-Tan, R. Radiogenomics: bridging imaging and genomics. Abdom. Radiol. 44, 1960–1984 (2019).
Alam, M. R. et al. Recent applications of artificial intelligence from histopathologic image-based prediction of microsatellite instability in solid cancers: a systematic review. Cancers 14, 2590 (2022).
Kather, J. N. et al. Pan-cancer image-based detection of clinically actionable genetic alterations. Nat. Cancer 1, 789–799 (2020).
Fu, Y. et al. Pan-cancer computational histopathology reveals mutations, tumor composition and prognosis. Nat. Cancer 1, 800–810 (2020).
Oh, J. M. et al. U1 snRNP regulates cancer cell migration and invasion in vitro. Nat. Commun. 11, 1 (2020).
Coudray, N. et al. Classification and mutation prediction from non-small cell lung cancer histopathology images using deep learning. Nat. Med. 24, 1559–1567 (2018).
Yu, B. H. et al. The clinicopathological relevance of uniform CD56 expression in anaplastic large cell lymphoma: a retrospective analysis of 18 cases. Diagn. Pathol. 16, 1 (2021).
Kather, J. N. et al. Deep learning can predict microsatellite instability directly from histology in gastrointestinal cancer. Nat. Med. 25, 1054–1056 (2019).
Moore, G. W. K., Howell, S. E. L., Brady, M., Xu, X. & McNeil, K. Anomalous collapses of Nares Strait ice arches leads to enhanced export of Arctic sea ice. Nat. Commun. 12, 1 (2021).
Sirinukunwattana, K. et al. Image-based consensus molecular subtype (imCMS) classification of colorectal cancer using deep learning. Gut 70, 544–554 (2021).
Skrede, O. J. et al. Deep learning for prediction of colorectal cancer outcome: a discovery and validation study. Lancet 395, 350–360 (2020).
Chong, Y. et al. Recommendations for pathologic practice using digital pathology: consensus report of the Korean Society of Pathologists. J. Pathol. Transl. Med. 54, 437–452 (2020).
Štorkánová, H. et al. Plasma Hsp90 levels in patients with systemic sclerosis and relation to lung and skin involvement: a cross-sectional and longitudinal study. Sci. Rep. 11, 1 (2021).
Waarts, M. R., Stonestrom, A. J., Park, Y. C. & Levine, R. L. Targeting mutations in cancer. J. Clin. Investig. 132, e154943 (2022).
Mendiratta, G. et al. Cancer gene mutation frequencies for the U.S. population. Nat. Commun. 12, 5961 (2021).
Kafieh, R. Artificial Intelligence in Cancer, Biology and Oncology (MDPI—Multidisciplinary Digital Publishing Institute, 2024).
Shao, X. et al. Transfer learning-based PET/CT three-dimensional convolutional neural network fusion of image and clinical information for prediction of EGFR mutation in lung adenocarcinoma. BMC Med. Imaging 24, 54 (2024).
Dammak, S., Cecchini, M. J., Breadner, D. & Ward, A. D. Using deep learning to predict tumor mutational burden from scans of H&E-stained multicenter slides of lung squamous cell carcinoma. J. Med. Imaging 10, 017502 (2023).
Liang, C. W., Fang, P. W., Huang, H. Y. & Lo, C. M. Deep convolutional neural networks detect tumor genotype from pathological tissue images in gastrointestinal stromal tumors. Cancers 13, 5787 (2021).
Cao, R. et al. Development and interpretation of a pathomics-based model for the prediction of microsatellite instability in colorectal cancer. Theranostics 10, 11080–11091 (2020).
Su, Y. et al. Application of BERT to enable gene classification based on clinical evidence. BioMed. Res. Int. 2020, 5491963 (2020).
Aburass, S., Dorgham, O. & Al Shaqsi, J. A hybrid machine learning model for classifying gene mutations in cancer using LSTM, BiLSTM, CNN, GRU, and GloVe. Syst. Soft Comput. 6, 200110 (2024).
Sun, Y. et al. Identification of 12 cancer types through genome deep learning. Sci. Rep. 9, 17256 (2019).
Zhang, S. et al. Improvement in prediction of prostate cancer prognosis with somatic mutational signatures. J. Cancer 8, 3261–3267 (2017).
Shiao, S. P. K., Grayson, J., Lie, A. & Yu, C. H. Personalized nutrition—genes, diet, and related interactive parameters as predictors of cancer in multiethnic colorectal cancer families. Nutrients 10, 795 (2018).
Hui, X. et al. EBT: a statistic test identifying moderate size of significant features with balanced power and precision for genome-wide rate comparisons. Bioinformatics 33, 2631–2641 (2017).
Cho, H.-J., Lee, S., Ji, Y. G. & Lee, D. H. Association of specific gene mutations derived from machine learning with survival in lung adenocarcinoma. PLoS ONE 13, e0207204 (2018).
Farina, E., Nabhen, J. J., Dacoregio, M. I., Batalini, F. & Moraes, F. Y. An overview of artificial intelligence in oncology. Futur. Sci. OA 8, Fso787 (2022).
Passaro, A. et al. Cancer biomarkers: emerging trends and clinical implications for personalized treatment. Cell 187, 1617–1635 (2024).
Tufail, M., Jiang, C.-H. & Li, N. Wnt signaling in cancer: from biomarkers to targeted therapies and clinical translation. Mol. Cancer 24, 107 (2025).
Forghani, R. Precision digital oncology: emerging role of radiomics-based biomarkers and artificial intelligence for advanced imaging and characterization of brain tumors. Radiol. Imaging Cancer 2, e190047 (2020).
Chiu, F.-Y. & Yen, Y. Imaging biomarkers for clinical applications in neuro-oncology: current status and future perspectives. Biomark. Res. 11, 35 (2023).
Chen, M. M. et al. Artificial intelligence in oncologic imaging. Eur. J. Radiol. Open 9, 100441 (2022).
Pandey, P., Mayank, K. & Sharma, S. (eds) Bio-Marker Cancer Prediction System Using Artificial Intelligence. 2023 International Conference on Integration of Computational Intelligent System (ICICIS), 1–4 November 2023 (2023).
Quddusi, D. M. & Bajcinca, N. Identification of genomic biomarkers and their pathway crosstalks for deciphering mechanistic links in glioblastoma. IET Syst. Biol. 17, 143–161 (2023).
Liu, Y. H., Jin, H. Q. & Liu, H. P. Identification of T-cell exhaustion-related gene signature for predicting prognosis in glioblastoma multiforme. J. Cell. Mol. Med. 27, 3503–3513 (2023).
Zhou, J., Zeng, Z. Y. & Li, L. Progress of artificial intelligence in gynecological malignant tumors. Cancer Manag. Res. 12, 12823–12840 (2020).
Kourou, K., Exarchos, T. P., Exarchos, K. P., Karamouzis, M. V. & Fotiadis, D. I. Machine learning applications in cancer prognosis and prediction. Comput. Struct. Biotechnol. J. 13, 8–17 (2015).
Kawakami, E. et al. Application of artificial intelligence for preoperative diagnostic and prognostic prediction in epithelial ovarian cancer based on blood biomarkers. Clin. Cancer Res. 25, 3006–3015 (2019).
Lv, J., Liu, G., Dong, W., Ju, Y. & Sun, Y. ACDB: An Antibiotic Combination DataBase. Front. Pharm. 13, 869983 (2022).
Sarvepalli, S. & Vadarevu, S. Role of artificial intelligence in cancer drug discovery and development. Cancer Lett 627, 217821 (2025).
Bassey, G. E., Daniel, E. A., Okesina, K. B. & Odetayo, A. F. Transformative role of artificial intelligence in drug discovery and translational medicine: innovations, challenges, and future prospects. Drug Des. Dev. Ther. 19, 7493–7502 (2025).
Long, X. et al. Artificial intelligence and anti-cancer drugs’ response. Acta Pharm. Sin. B 15, 3355–3371 (2025).
Abou Hajal, A. & Al Meslamani, A. Z. Insights into artificial intelligence utilisation in drug discovery. J. Med. Econ. 27, 304–308 (2024).
Serrano, D. R. et al. Artificial intelligence (AI) applications in drug discovery and drug delivery: revolutionizing personalized medicine. Pharmaceutics 16, 1328 (2024).
Hopkins, A. L. & Groom, C. R. The druggable genome. Nat. Rev. Drug Discov. 1, 727–730 (2002).
Lin, Q., Tam, P. K. & Tang, C. S. Artificial intelligence-based approaches for the detection and prioritization of genomic mutations in congenital surgical diseases. Front. Pediatr. 11, 1203289 (2023).
Chen, W., Liu, X., Zhang, S. & Chen, S. Artificial intelligence for drug discovery: resources, methods, and applications. Mol. Ther. Nucleic Acids 31, 691–702 (2023).
Qiu, X., Li, H., Ver Steeg, G. & Godzik, A. Advances in AI for protein structure prediction: implications for cancer drug discovery and development. Biomolecules 14, 339 (2024).
Knox, C. et al. DrugBank 6.0: the DrugBank knowledgebase for 2024. Nucleic Acids Res 52, D1265–D1275 (2024).
Lv, J., Liu, G., Ju, Y., Huang, H. & Sun, Y. AADB: a manually collected database for combinations of antibiotics with adjuvants. IEEE/ACM Trans. Comput. Biol. Bioinform. 20, 2827–2836 (2023).
Kim, S. et al. PubChem 2025 update. Nucleic Acids Res 53, D1516–d25 (2025).
Fahimian, G., Zahiri, J., Arab, S. S. & Sajedi, R. H. RepCOOL: computational drug repositioning via integrating heterogeneous biological networks. J. Transl. Med. 18, 375 (2020).
Wang, Y. et al. Discovery of novel glycogen synthase kinase-3α inhibitors: structure-based virtual screening, preliminary SAR and biological evaluation for treatment of acute myeloid leukemia. Eur. J. Med. Chem. 171, 221–234 (2019).
Gupta, R. et al. Artificial intelligence to deep learning: machine intelligence approach for drug discovery. Mol. Divers. 25, 1315–1360 (2021).
Dueñas, M. E. et al. Advances in high-throughput mass spectrometry in drug discovery. EMBO Mol. Med. 15, e14850 (2023).
Yin, Q. et al. DeepDrug: a general graph-based deep learning framework for drug–drug interactions and drug–target interactions prediction. Quant. Biol. 11, 260–274 (2023).
Zhavoronkov, A. et al. Deep learning enables rapid identification of potent DDR1 kinase inhibitors. Nat. Biotechnol. 37, 1038–1040 (2019).
Qu, X., Du, G., Hu, J. & Cai, Y. Graph-DTI: a new model for drug-target interaction prediction based on heterogenous network graph embedding. Curr. Comput.-Aided Drug Des. 20, 1013–1024 (2024).
Yu, L. et al. HGDTI: predicting drug-target interaction by using information aggregation based on heterogeneous graph neural network. BMC Bioinform 23, 126 (2022).
Yang, K., Bai, H., Ouyang, Q., Lai, L. & Tang, C. Finding multiple target optimal intervention in disease-related molecular network. Mol. Syst. Biol. 4, 228 (2008).
Cheng, F. et al. Prediction of drug–target interactions and drug repositioning via network-based inference. PLoS Comput. Biol. 8, e1002503 (2012).
Campillos, M., Kuhn, M., Gavin, A.-C., Jensen, L. J. & Bork, P. Drug target identification using side-effect similarity. Science 321, 263–266 (2008).
Tian, Z. et al. MHADTI: predicting drug–target interactions via multiview heterogeneous information network embedding with hierarchical attention mechanisms. Brief. Bioinform. 23, bbac434 (2022).
Sadaqat, M. et al. Advanced network pharmacology study reveals multi-pathway and multi-gene regulatory molecular mechanism of Bacopa monnieri in liver cancer based on data mining, molecular modeling, and microarray data analysis. Comput. Biol. Med. 161, 107059 (2023).
Emmerich, C. H. et al. Improving target assessment in biomedical research: the GOT-IT recommendations. Nat. Rev. Drug Discov. 20, 64–81 (2021).
Kiriiri, G., Njogu, P. & Mwangi, A. Exploring different approaches to improve the success of drug discovery and development projects: a review. Future J. Pharm. Sci. 6, 1–12 (2020).
Tran, N. L., Kim, H., Shin, C. H., Ko, E. & Oh, S. J. Artificial intelligence-driven new drug discovery targeting serine/threonine kinase 33 for cancer treatment. Cancer Cell Int 23, 321 (2023).
Albani, F. G., Alghamdi, S. S., Almutairi, M. M. & Alqahtani, T. Artificial intelligence-driven innovations in oncology drug discovery: transforming traditional pipelines and enhancing drug design. Drug Des. Dev. Ther. 19, 5685–5707 (2025).
Fang, X. et al. Chromosome instability functions as a potential therapeutic reference by enhancing chemosensitivity to BCL-XL inhibitors in colorectal carcinoma. Acta Pharmacol. Sin. 45, 2420–2431 (2024).
Zhang, Z. et al. Integrated analysis of single-cell and bulk RNA sequencing data reveals a pan-cancer stemness signature predicting immunotherapy response. Genome Med 14, 45 (2022).
Wen, T. et al. A Deep Learning Approach to Discover Cyclin-dependent Kinases 12 (CDK12) Inhibitors in Breast Cancer (American Society of Clinical Oncology, 2022).
Huang, A., Garraway, L. A., Ashworth, A. & Weber, B. Synthetic lethality as an engine for cancer drug target discovery. Nat. Rev. Drug Discov. 19, 23–38 (2020).
Wang, S. et al. KG4SL: knowledge graph neural network for synthetic lethality prediction in human cancers. Bioinformatics 37, i418–i425 (2021).
Sheng, C., Dong, G., Miao, Z., Zhang, W. & Wang, W. State-of-the-art strategies for targeting protein–protein interactions by small-molecule inhibitors. Chem. Soc. Rev. 44, 8238–8259 (2015).
Lu, C. et al. Systemic evolutionary chemical space exploration for drug discovery. J. Cheminform. 14, 19 (2022).
Yasuo, N. & Sekijima, M. Improved method of structure-based virtual screening via interaction-energy-based learning. J. Chem. Inf. Model. 59, 1050–1061 (2019).
Singh, S., Gupta, H., Sharma, P. & Sahi, S. Advances in artificial intelligence (AI)-assisted approaches in drug screening. Artif. Intell. Chem. 2, 100039 (2024).
Raies, A. et al. DrugnomeAI is an ensemble machine-learning framework for predicting druggability of candidate drug targets. Commun. Biol. 5, 1291 (2022).
Bailleux, C., Gal, J., Chamorey, E., Mograbi, B. & Milano, G. Artificial intelligence and anticancer drug development—keep a cool head. Pharmaceutics 16, 211 (2024).
Chow, R. et al. Use of artificial intelligence for cancer clinical trial enrollment: a systematic review and meta-analysis. J. Natl. Cancer Inst. 115, 365–374 (2023).
Azenkot, T., Rivera, D. R., Stewart, M. D. & Patel, S. P. Artificial intelligence and machine learning innovations to improve design and representativeness in oncology clinical trials. Am. Soc. Clin. Oncol. Educ. Book 45, e473590 (2025).
Cai, L. et al. Machine learning to predict the individual risk of treatment-relevant toxicity for patients with breast cancer undergoing neoadjuvant systemic treatment. JCO Clin. Cancer Inform 8, e2400010 (2024).
Stabellini, N. et al. Thirty-day unplanned hospital readmissions in patients with cancer and the impact of social determinants of health: a machine learning approach. JCO Clin. Cancer Inform 7, e2200143 (2023).
Dowden, H. & Munro, J. Trends in clinical success rates and therapeutic focus. Nat. Rev. Drug Discov. 18, 495–496 (2019).
Hu, Y. et al. In silico prediction of human organ toxicity via artificial intelligence methods. Chem. Res. Toxicol. 36, 1044–1054 (2023).
Liu, R. et al. Evaluating eligibility criteria of oncology trials using real-world data and AI. Nature 592, 629–633 (2021).
Zhan, Y., Hao, Y., Wang, X. & Guo, D. Advances of artificial intelligence in clinical application and scientific research of neuro-oncology: current knowledge and future perspectives. Crit. Rev. Oncol./Hematol. 209, 104682 (2025).
Zhao, X., Xiong, J., Li, D. & Zhang, Y. Clinical trials of nanoparticle-enhanced CAR-T and NK cell therapies in oncology: overcoming translational and clinical challenges - a mini review. Front Med (Lausanne) 12, 1655693 (2025).
Chopra et al. Revolutionizing clinical trials: the role of AI in accelerating medical breakthroughs. Int J. Surg. 109, 4211–4220 (2023).
Chong, P. L. et al. Integrating artificial intelligence in healthcare: applications, challenges, and future directions. Future Sci. OA 11, 2527505 (2025).
Moon, H., Nguyen, P. N., Park, J., Lee, M. & Ahn, S. AI-guided chemotherapy optimization in lung cancer using genomic and survival data. J. Pers. Med. 15, 218 (2025).
Liao, J. et al. Artificial intelligence assists precision medicine in cancer treatment. Front. Oncol. 12, 998222 (2022).
AlSamhori, J. F. et al. Artificial intelligence for breast cancer: implications for diagnosis and management. J. Med. Surg. Public Health 3, 100120 (2024).
Wilhelm, C., Steckelberg, A. & Rebitschek, F. G. Benefits and harms associated with the use of AI-related algorithmic decision-making systems by healthcare professionals: a systematic review. Lancet Reg. Health—Eur 48, 101145 (2025).
Singh, Y. et al. Beyond the hype: navigating bias in AI-driven cancer detection. Oncotarget 15, 764–766 (2024).
Morales-Forero, A., Rueda, L. J., Herrera, R., Bassetto, S. & Coatanea, E. Predictive representativity: uncovering racial bias in AI-based skin cancer detection. arXiv preprint arXiv:250714176 (2025).
Ganta, T. et al. Fairness in predicting cancer mortality across racial subgroups. JAMA Netw. Open 7, e2421290 (2024).
Preetam, S. et al. Next-gen diagnostics: artificial intelligence-powered imaging in breast cancer care. J Cancer Metastasis Treat 11, 29 (2025).
Derraz, B. et al. New regulatory thinking is needed for AI-based personalised drug and cell therapies in precision oncology. npj Precis. Oncol. 8, 23 (2024).
Chan, J., Parker, L., Carter, S., Nickel, B. & Carroll, S. Radiation oncology patients’ perceptions of artificial intelligence and machine learning in cancer care: a multi-centre cross-sectional study. Radiother. Oncol. 207, 110891 (2025).
Chamouni, G. et al. Ethical and legal concerns in artificial intelligence applications for the diagnosis and treatment of lung cancer: a scoping review. Front. Public Health 13, 1663298 (2025).
Collins, G. S. et al. Clinical prediction models using machine learning in oncology: challenges and recommendations. BMJ Oncol 4, e000914 (2025).
Adeoye, J., Akinshipo, A., Koohi-Moghadam, M., Thomson, P. & Su, Y. X. Construction of machine learning-based models for cancer outcomes in low and lower-middle income countries: a scoping review. Front. Oncol. 12, 976168 (2022).
Hogg, H. D. J. et al. Stakeholder perspectives of clinical artificial intelligence implementation: systematic review of qualitative evidence. J. Med. Internet Res. 25, e39742 (2023).
Yun, T. & Zhang, L. International partnerships in AI-driven healthcare: opportunities and challenges for advancing the UN Sustainable Development Goals—a perspective. Healthcare 13, 2053 (2025).
Roppelt, J. S., Kanbach, D. K. & Kraus, S. Artificial intelligence in healthcare institutions: a systematic literature review on influencing factors. Technol. Soc. 76, 102443 (2024).
Maleki Varnosfaderani, S. & Forouzanfar, M. The role of AI in hospitals and clinics: transforming healthcare in the 21st century. Bioengineering 11, 337 (2024).
Goh, S. et al. Challenges in implementing artificial intelligence in breast cancer screening programs: systematic review and framework for safe adoption. J. Med. Internet Res. 27, e62941 (2025).
Hutter, C. & Zenklusen, J. C. The Cancer Genome Atlas: creating lasting value beyond its data. Cell 173, 283–285 (2018).
Aldoseri, A., Al-Khalifa, K. N. & Hamouda, A. M. Re-thinking data strategy and integration for artificial intelligence: concepts, opportunities, and challenges. Appl. Sci. 13, 7082 (2023).
Saxena, S. et al. Role of artificial intelligence in radiogenomics for cancers in the era of precision medicine. Cancers (Basel) 14, 2860 (2022).
Uwimana, A., Gnecco, G. & Riccaboni, M. Artificial intelligence for breast cancer detection and its health technology assessment: a scoping review. Comput. Biol. Med. 184, 109391 (2025).
Author information
Authors and Affiliations
Contributions
Jun Li and Danyang Li: Conceptualisation, methodology, writing—original draft; Writing—review. Lei Zhang. Zhenglun Yu: Data curation, formal analysis, writing—review and editing. Zhiye Bao and Liming Wang: Supervision and writing—review and editing. All authors reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, J., Zhang, L., Yu, Z. et al. The impact of AI on modern oncology from early detection to personalized cancer treatment. npj Precis. Onc. 10, 69 (2026). https://doi.org/10.1038/s41698-026-01276-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41698-026-01276-6