Abstract
Chemotherapy-induced immunosuppression compromises therapeutic outcomes in oncology, particularly in non-small cell lung cancer (NSCLC). While natural polysaccharides have emerged as promising candidates to counteract drug-related immunosuppression, the therapeutic potential of riclin remains unexplored in chemotherapeutic contexts. Here, we systematically evaluated riclin’s immunoadjuvant efficacy in a murine NSCLC model treated with gemcitabine (GEM). Oral riclin modulated gut microbiota diversity and metabolite profiles while activating the immune-hematopoietic axis, thereby boosting immunity and hematopoiesis. Mechanistically, riclin counteracted GEM-induced immunosuppression through coordinated NF-κB and JAK-STAT activation, as evidenced by the restoration of circulating immune cells and splenic architecture, expansion of bone marrow mononuclear cells (BMNCs) and Lineage⁻Sca-1⁺Kit⁺ (LSK) cells, and suppression of apoptosis. Notably, riclin demonstrated synergistic effects with GEM, achieving 98% tumor burden reduction while concurrently alleviating systemic immunosuppression. We propose that riclin, a novel oral immunoadjuvant capable of enhancing chemotherapeutic efficacy in NSCLC, supports its clinical development as a promising combination strategy.

Similar content being viewed by others
Introduction
Chemotherapy remains a cornerstone of cancer treatment, yet its clinical efficacy is frequently limited by treatment-induced immunosuppression that compromises host immunity, reduces therapeutic durability, and deteriorates patient quality of life1. This necessitates the urgent development of strategies that mitigate immunosuppression while maintaining antitumor efficacy. Current approaches to counteract chemotherapy-induced immunosuppression typically involve combining cytotoxic agents with immune-enhancing therapies. For instance, cancer vaccines or adoptive cell therapies synergize with chemotherapy to activate or reprogram immune cells for tumor-specific recognition and elimination, thereby circumventing immune evasion mechanisms2. Concurrent administration of immune checkpoint inhibitors (e.g., anti-PD-1 or anti-CTLA-4 antibodies) with chemotherapy enhances antitumor responses by blocking inhibitory signals and restoring T-cell functionality3. Advances in drug delivery systems have further expanded therapeutic possibilities: CD44-targeted GEM nanotherapeutics demonstrate enhanced tumor growth inhibition with reduced systemic toxicity4, and RGDV-modified GEM nanomedicine improves drug stability, overcomes chemoresistance, and mitigates myelosuppression5. Complementary strategies such as lifestyle interventions, nutritional optimization, and prophylactic regimens may additionally alleviate immunosuppression and improve patient resilience6. However, clinical implementation of these approaches encounters substantial challenges, including high costs, treatment-related toxicities, interpatient response variability, and unresolved long-term safety concerns. Consequently, developing strategies that effectively balance immunosuppression alleviation with sustained antitumor activity remains a critical unmet need in oncology.
Natural products, characterized by their pleiotropic immunomodulatory properties, have emerged as compelling candidates to counteract chemotherapy-driven immune dysfunction7. For example, flavonoids such as UP446 (derived from Scutellaria baicalensis and Acacia catechu) and flavonols from Aronia berries demonstrate dual functionality in attenuating immunosuppression while enhancing both innate and adaptive immunity8,9. Similarly, ginsenosides modulate immune responses through macrophage activation and T-cell regulation10,11. Berberine has shown immunoregulatory potential in ameliorating immune disorders, although its role in alleviating chemotherapy-induced immunosuppression remains underexplored12. However, the clinical translation of many natural compounds with promising preclinical profiles remains hindered by challenges, including standardization, dose-limiting toxicity, variable bioavailability, and insufficient potency12,13,14,15. In contrast, natural polysaccharides have gained prominence as immunoadjuvants due to their favorable safety profile, structural accessibility, robust bioactivity, and extensive validation in preclinical studies16.
Natural polysaccharides, ubiquitously distributed in fungi, plants, microorganisms, and algae, exhibit well-documented immunomodulatory effects through intrinsic immune cell activation and lymphocyte proliferation regulation17,18,19,20. For instance, Tremella polysaccharides were shown to enhance immune function and attenuate cyclophosphamide (CTX)-induced hepatotoxicity during immunosuppression19. Similarly, polysaccharides from Menispermum dauricum DC rhizomes demonstrated synergistic antitumor effects with CTX while ameliorating myelosuppression and immunosuppression in Lewis tumor-bearing mice18. Complementary studies revealed that turmeric polysaccharides and Atractylodes lancea rhizome polysaccharides mitigate immunosuppression and intestinal damage20,21, whereas sodium alginate restored splenic architecture and optimized gut microbiota composition in immunosuppressed murine models22. Furthermore, the exopolysaccharide EPS103 was found to activate systemic immunity via modulation of short-chain fatty acids (SCFAs) production and gut microbial communities, thereby alleviating immunosuppression23. Among these diverse sources, microbial-derived polysaccharides present distinct advantages, including rapid production scalability, cost-effectiveness, structural stability, and functional versatility24. However, their potential as immunoadjuvants in tumor chemotherapy remains underexplored.
Riclin, a non-toxic fermentative exopolysaccharide produced by Agrobacterium sp. ZCC3656 exhibits biocompatibility with scalable production advantages, including short fermentation cycles, high yields (21 g/L), and cost-effectiveness25,26,27. Previous studies have characterized its immunogenic properties and glycemic modulation capabilities28,29,30, yet its adjuvant potential and mechanism of action in NSCLC chemotherapy remain uncharacterized. To address this knowledge gap, we systematically investigate the immune activation mechanisms of orally administered riclin and evaluate its therapeutic synergy with chemotherapy in NSCLC. This study seeks to develop a novel combinatorial strategy for optimizing chemotherapeutic efficacy while preserving systemic immune homeostasis.
Results
Riclin alters gut microbiota profiles and immune-metabolic pathways
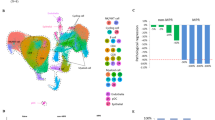
Sample adequacy was validated by observed species, rank abundance, and specaccum curves, confirming sufficient sequencing depth to characterize microbial community richness and evenness (Fig. S1A–C). Oral administration of riclin significantly remodeled the gut microbiota structure, manifested as a marked decrease in species richness (reduced ACE and Chao1 indices, P < 0.01) and distinct beta diversity separation from controls (Fig. 1A–B, E). However, the Shannon and Simpson indices reflecting species diversity showed no significant changes (Fig. 1C, D). This restructuring is marked by the selective enrichment of the Bacteroidota (phylum) and genera Muribaculaceae, Bacteroides, Alistipes, and Parabacteroides, concurrent with the suppression of Helicobacter (Fig. 1F–G and Fig. S1D, E). Untargeted liquid chromatography–mass spectrometry (LC–MS) metabolomic profiling demonstrated distinct intergroup metabolite clustering (Fig. 1H), identifying 31 downregulated and 102 upregulated metabolites post-riclin treatment (Figs. 1I and S1F). The variable importance in projection (VIP)-ranked heatmap of the top 50 differential metabolites highlighted upregulated amino acid metabolism intermediates, including L-histidinol, histidinal, N-acetyl-L-aspartate (NAA), and indolepyruvate (Fig. 1J–N and Fig. S1G). Pearson correlation analysis revealed robust associations between enriched Bacteroides and immunomodulatory metabolites (L-histidinol, NAA, riboflavin, liquoric acid; Figs. 1O and S1H–K), suggesting microbiota-metabolite interplay as a potential mechanism underlying riclin’s immune modulation.
Alpha diversity indices: ACE (A), Chao1 (B), Shannon (C), and Simpson (D). PCA of microbial communities (E). Phylum-level microbiota composition (F). Genus-level microbiota heatmap (G). Metabolic profile PCA (H). Volcano plot (I) and KEGG-enriched pathways (J) of differential metabolites. Volcano plot threshold: variable importance in projection (VIP) > 1 and p-value < 0.05. Expression of key metabolites: L-histidinol (K), histidinal (L), indolepyruvate (M), and NAA (N). Pearson’s correlation analysis among altered metabolites and microbes (O). Absolute content of short-chain fatty acids (SCFAs): acetic acid (P), pentanoic acid (Q), isovaleric acid (R), isobutyric acid (S), hexanoic acid (T), and glutaric acid (U). Data represent mean ± SD (n = 8 per group); *p < 0.05, **p < 0.01 vs CK group.
To directly investigate whether riclin enhances the production of immunomodulatory gut microbial metabolites, we performed targeted quantitative analysis of SCFAs. We found that riclin treatment significantly altered the SCFA profile, marked by increased levels of acetic, pentanoic, isobutyric, isovaleric, hexanoic, and glutaric acids compared to the control group (Fig. 1P–U and Fig. S2A–E). This finding provides a functional link between riclin-induced microbiota remodeling and systemic immune activation.
Riclin activates RAW 264.7 macrophages in vitro
Building upon gut–immune axis interactions, we investigated riclin’s immunomodulatory potential using RAW 264.7 macrophages. Morphological assessment demonstrated riclin-induced cellular hypertrophy, nuclear expansion, and pseudopodia formation (Fig. 2A, B). Although riclin showed no proliferative effects (Fig. 2C), it significantly enhanced phagocytic capacity (P < 0.001) and NO production compared with the controls (Fig. 2D, E). ELISA quantification revealed elevated TNF-α, IL-1β, and IL-6 levels in riclin-treated cells compared with the control group (Fig. 2F–H). These findings indicate that riclin promotes macrophage activation, as evidenced by functional maturation and enhanced cytokine secretion.
Giemsa-stained morphology (scale bars: 20 μm, 5 μm) (A). Actin cytoskeleton visualization via Tracker Green-488 staining (scale bar: 20 μm) (B). Functional assessments: Cell viability (C), phagocytic activity (D), NO production (E), and secreted cytokines including TNF-α (F), IL-1β (G), and IL-6 (H). Data represent mean ± SD (n = 5–6 per group); **p < 0.01, ***p < 0.001 vs CK group.
Oral riclin enhances immune and hematopoietic functions in mice
Given that macrophage activation modulates systemic immunity through transcriptional regulation, we conducted RNA sequencing of spleen and bone marrow. Inter-sample correlation (Fig. S3A, C) and PCA revealed distinct transcriptional profiles between riclin-treated and CK (control) groups (Fig. 3A, B). Riclin administration induced 1164 spleen and 864 bone marrow differentially expressed genes (DEGs) (Figs. 3C, D and S3B, D), including upregulated proinflammatory cytokines (e.g., Il-6, Il-1β, Tnf-α), hematopoietic factors (e.g., GM-CSF, G-CSF, M-CSF), and immune regulators (e.g., Tlr2, Cd14, Relb) (Figs. 3E and S3E, F). Gene ontology (GO) analysis identified the top 30 terms enriched for immune/inflammatory responses and cytokine activity (Fig. S3G, H). Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways included TNF, Toll-like receptor, and NF-κB signaling, with concurrent enrichment of hematopoietic cell lineage pathways (Fig. 3F, G). These findings demonstrate that riclin enhances murine immune-hematopoietic functions, potentially mediated through NF-κB signaling.
PCA analysis of spleen (A) and bone marrow (B) transcriptomes. Volcano plots of DEGs in spleen (C) and bone marrow (D). Heatmap of DEGs in spleen and bone marrow (E). KEGG pathway enrichment analysis of spleen (F) and bone marrow (G) DEGs. Data derived from biological triplicates (n = 3/group).
Oral riclin engages the NF-κB/JAK-STAT axis to coordinate immune-hematopoietic regulation
Based on KEGG pathway enrichment, we validated NF-κB signaling in spleen and bone marrow. Compared with the controls, riclin significantly increased TNF-α, IL-1β, and IL-6 protein levels (Fig. 4A, D), indicating initial immune activation. This was accompanied by elevated p-NF-κB p65/NF-κB p65 ratios, reduced p-IκBα/IκBα ratios, and upregulated GM-CSF expression (Fig. 4B, D), consistent with NF-κB pathway activation and hematopoietic enhancement. Riclin further increased GM-CSF receptor (GM-CSFr) levels and phosphorylation ratios of JAK2 (p-JAK2/JAK2) and STAT5 (p-STAT5/STAT5) (Fig. 4C, D). These findings suggest concurrent activation of JAK-STAT signaling, mechanistically linking riclin-induced immune potentiation and hematopoietic promotion.
Representative immunoblots and densitometric quantification of key proteins in spleen and bone marrow: TNF-α, IL-1β, IL-6, p-IκBα/IκBα, p-NF-κB p65/NF-κB p65, GM-CSF, GM-CSFr, p-JAK2/JAK2, and p-STAT5/STAT5 (A–D). Bone marrow nucleated cell (BMNC) counts (E). Peripheral white blood cells (WBCs) (F) and lymphocyte (LYMPH) counts (G). Representative flow cytometry plots (H) and quantitative analysis of bone marrow LSK cells (I). Schematic of the riclin-mediated immune-hematopoietic axis (J). Data represent mean ± SD (n = 3–6); *p < 0.05, **p < 0.01, ***p < 0.001 vs CK group.
To functionally validate the requirement of the NF-κB/JAK-STAT axis in riclin-mediated hematopoietic recovery, we employed specific pharmacological inhibitors in vivo. As hypothesized, riclin monotherapy promoted hematopoietic recovery, evidenced by the restoration of BMNCs (Fig. 4E), peripheral white blood cells (WBCs) (Fig. 4F) and lymphocytes (LYMPH) counts (Fig. 4G), and expansion of the LSK progenitor pool (Figs. 4H, I and S4). These restorative effects were, however, substantially diminished upon co-administration of either the NF-κB inhibitor BAY 11-7082 or the JAK2 inhibitor Ruxolitinib (Fig. 4E–I). These results collectively demonstrate that riclin-mediated hematopoietic reconstitution involves the cooperative activation of NF-κB and JAK-STAT pathways (Fig. 4J).
Oral riclin ameliorates GEM-induced immunopathology
We next evaluated riclin’s immunoprotective efficacy in a GEM-induced immunosuppression model (Fig. 5A). Riclin pretreatment mitigated GEM-induced body weight loss and restored thymic/splenic indices (Fig. 5B–D). Peripheral hematological analysis revealed riclin-enhanced immune function and hematopoietic capacity through increased WBCs, LYMPH, neutrophil (NE), and platelet counts (Fig. 5E–I). Histopathological assessment demonstrated riclin’s tissue-protective effects. GEM treatment caused severe destruction of the splenic tissue structure, loose cellular arrangement, altered bone marrow sinusoids, and decreased myeloid/erythroid ratios. Riclin treatment preserved the number of cells in the white and red pulp, with a tightly ordered cell arrangement, and restored the GEM-affected bone marrow (Fig. 5J, K). In vitro, riclin counteracted GEM-mediated macrophage cytotoxicity, rescuing phagocytic function and cytokine secretion (Fig. S5A–F). Flow cytometry confirmed riclin-enhanced splenic immunity via increased total F4/80⁺ cells and expanded CD3⁺, CD4⁺, and CD8⁺ subsets (Figs. 5L–P and S6A–H), alongside elevated TNF-α, IL-6, and IL-1β levels compared to GEM-treated mice (Fig. 5Q, R). Additionally, riclin attenuated the GEM-induced oxidative stress imbalance (Tables S1, S2 and Fig. S7A–H). These data collectively establish riclin’s protective role against GEM-induced immunotoxicity.
Experimental timeline and dosing regimen (A). Body weight (B). Organ indices: Thymus (C) and Spleen (D). Peripheral blood counts: WBCs (E), RBCs (F), LYMPH (G), NE (H), and platelets (I). Histology of spleen (Scale bar: 100 μm, 50 μm) (J) and bone marrow (Scale bar: 20 μm) (K). Flow cytometric quantification of splenic total cells (L), CD3⁺ T cells (M), CD3⁺CD4⁺ T cells (N), CD3⁺CD8⁺ T cells (O), and CD11b⁺F4/80⁺ macrophages (P). Spleen (Q) and serum levels (R) of TNF-α, IL-1β, and IL-6. Data represent mean ± SD (n = 4–11); *p < 0.05, **p < 0.01, ***p < 0.001 vs. CK; #p < 0.05, ##p < 0.01, ###p < 0.001 vs. GEM group.
Oral riclin alleviates GEM-induced hematopoietic suppression
Given that GEM exerts toxicity via cell cycle blockade, we analyzed BMNCs at 24, 48, 72, and 120 h post-treatment. Riclin groups maintained higher BMNC counts versus GEM controls after 72 h (Fig. 6A). Cell cycle profiling at 72 h (peak divergence) revealed elevated S + G2 phase proportions in riclin groups (40.3% and 34.2%) compared to GEM (27.8%), indicating proliferative rescue (Figs. 6B, C and S8A–D). RT-qPCR confirmed riclin-induced significant upregulation of hematopoietic factors GM-CSF, G-CSF, and M-CSF in spleen and bone marrow (Fig. 6D, E). To definitively assess the impact of riclin on primitive hematopoietic cells, we next quantified LSK cells in the bone marrow. The LSK population was modestly increased in GEM-treated mice compared to controls, consistent with spontaneous hematopoietic recovery and our cell cycle data. In contrast, riclin combination therapy led to a more pronounced expansion of the LSK population (Fig. 6F, G). Additionally, riclin attenuated GEM-induced apoptosis, reducing splenic (24.35% → 6.65%) and bone marrow (21.18% → 7.56%) apoptotic rates (Fig. 6H–K). This anti-apoptotic effect correlated with downregulated Bax and Caspase3 expression (Fig. 6L, M). Collectively, riclin ameliorates GEM-induced hematopoietic injury through dual mechanisms: promoting BMNCs and LSK cells proliferation and suppressing apoptosis.
BMNCs counts (A) and cell cycle distribution (G1 phase (B), S + G2 phase (C)) following riclin treatment at 24, 48, 72, and 120 h post-second GEM (i.p.). mRNA expression of hematopoietic cytokines (GM-CSF, G-CSF, M-CSF) in spleen (D) and bone marrow (E). Representative flow cytometry plots (F) and (G) quantitative analysis of bone marrow LSK cells. Flow cytometric analysis of apoptosis: representative plots and quantification in spleen (H–I) and bone marrow (J, K). mRNA expression of Bax, Bcl2, and Caspase3 in spleen (L) and bone marrow (M). Data represent mean ± SD (n = 3–6); *p < 0.05, **p < 0.01, ***p < 0.001 vs. CK; #p < 0.05, ##p < 0.01, ###p < 0.001 vs. GEM group.
Oral riclin enhances chemotherapeutic efficacy against NSCLC
To assess riclin’s immunoadjuvant potential in chemotherapy, we established a Lewis lung cancer (LLC) mouse model with GEM co-treatment (Fig. 7A). Riclin synergistically enhanced GEM’s antitumor efficacy, significantly reducing tumor growth rates compared to GEM monotherapy (Fig. 7B–D) and demonstrating combinatorial tumor suppression (Fig. 7E–G). Notably, riclin ameliorated GEM-induced systemic toxicity, as evidenced by restored body weight, normalized organ indices (thymus/spleen), and improved peripheral blood cell counts (WBCs, lymphocytes, neutrophils, platelets; Figs. 7H–K and Table S3). Histopathological analysis further validated riclin’s protective effects, with preserved splenic architecture (clear red/white pulp boundaries and ordered cellular arrangement) in riclin-treated mice versus GEM-induced structural disorganization (Fig. 7L). Given the spleen’s central role in systemic immunity, we assessed splenic responses to elucidate the collaborative antitumor mechanisms. Flow cytometric analysis confirmed the expansion of CD4⁺ T cells (Figs. 7M, N and S9), without a significant change in the proportion of CD8⁺ T cells (Fig. 7O, P). We then further assessed T-cell functional status by immunofluorescence. While GEM treatment substantially suppressed the expression of the critical surface receptors CD4 and CD8—suggesting functional impairment—riclin co-treatment preserved their expression (Fig. 7Q–S). Correspondingly, the co-administration of riclin elevated proinflammatory cytokines (TNF-α, IL-1β, IL-6) in the serum and spleen, an effect that was associated with enhanced antitumor immunity (Fig. 7T, U). Collectively, our data suggest that riclin-GEM combination therapy achieves superior antitumor efficacy while mitigating chemotherapy- induced toxicity.
Experimental design (A). Tumor progression metrics: in vivo bioluminescence imaging (B), tumor volume (C), total fluorescence intensity (D), gross tumor morphology (E), H&E-stained histopathology (Scale bar: 50 μm) (F), and tumor weight (G). Systemic effects: body weight (H); organ indices of thymus (I) and spleen (J). LYMPH count (K). Histopathology of spleen (Scale bars: 50 μm, 20 μm) (L). Flow cytometric analysis of splenic T-cell subsets: representative CD3⁺CD4⁺ (M) and CD3⁺CD8⁺ T-cell (O) profiles with quantification (N, P). Q Immunofluorescence staining of CD4 and CD8 in spleen tissue. Scale bar: 50 μm. Quantification of CD4 (R) and CD8 mean fluorescence intensity (S). Cytokine levels: splenic mRNA expression (T) and serum concentrations of TNF-α, IL-1β, and IL-6 (U). Data represent mean ± SD (n = 3–8); *p < 0.05, **p < 0.01, ***p < 0.001 vs. CK; #p < 0.05, ##p < 0.01, ###p < 0.001 vs. GEM group.
Discussion
This study identifies riclin, a microbial exopolysaccharide, as a highly promising oral immunoadjuvant that concurrently ameliorates chemotherapy-induced immunosuppression and enhances antitumor efficacy in NSCLC. Through coordinated modulation of gut microbiota homeostasis, dual activation of NF-κB and JAK-STAT signaling pathways, and hematopoietic system restoration, oral riclin fundamentally improves the therapeutic window of GEM-based chemotherapy regimens.
Host immunity is strongly associated with the gut microbial-metabolic axis31. The observed reduction in species richness (ACE/Chao1 indices) and pronounced β-diversity profile indicate that riclin promotes selective restructuring of the gut microbiota rather than ecological disruption, consistent with the effects of other bioactive polysaccharides32,33. While decreased diversity is often considered unfavorable, in the context of riclin treatment, it indicates a selective shift toward beneficial, immunomodulatory taxa. This ecological specialization was characterized by the enrichment of Muribaculaceae, which interacts with innate and adaptive immunity via Immunoglobulin A coating34, Alistipes, which promote intestinal immune maturation35, and Bacteroides, which contribute to immunomodulation and enhance the epithelial barrier function—alongside the suppression of the opportunistic pathogen Helicobacter35,36. This aligns with established mechanisms where dietary polysaccharides enhance immune homeostasis through microbiotic modulation37,38. Critically, this structural reorganization was coupled with a profound reprogramming of the gut metabolome. The concurrent increase in immunomodulatory metabolites (e.g., L-histidinol, NAA, indolepyruvate)39,40,41 and key microbially derived SCFAs (particularly acetate, pentanoic, and isovaleric acid)42,43,44, confirms that the riclin-remodeled microbiota exhibits enhanced functional capacity for immune regulation, demonstrating that microbial quality and functional orientation, rather than sheer diversity, govern host immunological outcomes. These multi-omics findings establish riclin’s dual mechanism of intestinal immunoprotection through coordinated microbiota remodeling and metabolic network regulation.
Macrophages serve as pivotal regulators of both intestinal and systemic immunity. In this study, riclin demonstrated robust macrophage activation through enhanced phagocytic activity, NO production, and elevated secretion of proinflammatory cytokines (TNF-α, IL-6, IL-1β)—functional hallmarks of M1 polarization. Phagocytic capacity, a key indicator of macrophage activation, reflects their functional maturation in immune surveillance45,46. NO, generated by activated macrophages, plays a critical role in scavenging viruses and bacteria47. The upregulated cytokines TNF-α, IL-6, and IL-1β, predominantly secreted by macrophages, are critical mediators of immune response signal transduction. This immunostimulatory pattern aligns with the activity of exopolysaccharides derived from Lactobacillus helveticus LZ-R-5, which similarly potentiate macrophage function48. Collectively, these findings indicate that riclin exhibits potent immunostimulatory activity via macrophage activation and hold promise as a candidate immunoadjuvant.
Macrophage activation modulates systemic immunity through transcriptional regulation. Transcriptomic profiling demonstrated riclin-induced upregulation of IL-6, TNF-α, and IL-1β, cytokines critical for lymphocyte proliferation, humoral immunity, and hematopoietic regulation49,50. These cytokines engage cognate receptors to activate downstream signaling cascades, including induction of the pleiotropic hematopoietic factor GM-CSF, which bridges immune and hematopoietic functions51. Although dysregulated NF-κB activation is implicated in autoimmunity52, riclin appears to harness this pathway constructively, mirroring the TLR4/NF-κB-mediated immunomodulation observed with sulfated glycosaminoglycans53. Mechanistically, riclin-induced inflammatory factors initiate immunity and activate the NF-κB pathway, which in turn triggers the GM-CSF-mediated JAK2/STAT5 pathway and promotes hematopoietic stem cell proliferation54,55.
GEM, a first-line antimetabolite chemotherapeutic agent for NSCLC, pancreatic cancer, and ovarian malignancies, is clinically limited by dose-dependent pancytopenia and immunosuppression that compromise patient quality of life7. This study demonstrates riclin’s dual therapeutic capacity to counteract GEM-induced immunotoxicity. Mechanistically, riclin preserves hematopoietic function by inhibiting organ atrophy and restoring peripheral leukocyte/platelet counts, thereby rescuing immune-hematopoietic homeostasis. Concurrently, riclin enhances splenic immunocompetence through structural remodeling (restored red/white pulp architecture), increased splenocyte populations, and amplified proinflammatory cytokine secretion. These coordinated actions collectively mitigate chemotherapy-associated immunosuppression.
GEM induces cytotoxicity primarily through cell cycle blockade56. Our data demonstrate that riclin ameliorates GEM-induced myelosuppression by rescuing BMNC proliferation and reversing S/G2 phase arrest, critical processes for maintaining bone marrow hematopoietic homeostasis. This proliferative recovery mechanistically accounts for the elevated peripheral blood cell counts in riclin-treated mice, which mitigates risks of immune injury and hematopoietic dysfunction. Notably, riclin markedly improved the proportion of LSK cells (a population enriched with hematopoietic stem and progenitor cells)57, providing direct evidence that riclin functionally rescues GEM-induced hematopoietic damage. Given that impaired hematopoietic factor expression (G-CSF, M-CSF, GM-CSF) drives chemotherapy-associated cytopenia58, riclin’s ability to upregulate these factors in murine spleen and bone marrow directly restores hematopoietic function. Furthermore, riclin suppresses apoptosis in hematopoietic organs, as evidenced by reduced Bax and Caspase3 expression59. These findings collectively establish riclin as a dual-mechanism therapeutic agent that concurrently addresses chemotherapy-induced immunosuppression and hematopoietic injury, providing a translational foundation for its clinical development.
Lung cancer remains the leading cause of cancer-related mortality globally, with NSCLC comprising 80–85% of cases and exhibiting poor prognosis and low survival rates60. Based on this, we evaluated the potential of riclin as an immunoadjuvant to enhance chemotherapy efficacy and counteract immunosuppression in a NSCLC murine model. The results indicated that riclin demonstrates synergistic antitumor activity with GEM, simultaneously reducing tumor burden and counteracting systemic immunosuppression, a dual advantage uncharacteristic of conventional adjuvants. Specifically, riclin effectively reversed chemotherapy-induced immunosuppression by restoring immune organ indices, peripheral blood counts, and splenic architecture in tumor-bearing mice. Mechanistically, riclin exerts its immunoenhancing effects through T-cell-dependent mechanisms, elevating the abundance of splenic CD4⁺ T cells and potently activating both CD4⁺ and CD8⁺ T-cell functions. This leads to the re-establishment of immune homeostasis, enabling the host to sustain an effective antitumor response. These findings align with established evidence that immune activation within secondary lymphoid organs can potently orchestrate antitumor immunity61,62. Collectively, these findings substantiate riclin’s translational potential as a novel immunoadjuvant to mitigate chemotherapy-induced immunosuppression and enhance therapeutic outcomes in NSCLC.
In summary, this study establishes that riclin mitigates GEM-induced immunosuppression and hematopoietic injury in preclinical models, providing a mechanistic rationale for its development as an immunoadjuvant to optimize chemotherapy. While its action through the gut–immune–bone marrow axis represents a distinct mode of action, it should be noted that our evaluation was confined to GEM combination therapy. Future work should systematically assess its efficacy with other standard agents (e.g., platinum-based drugs or taxanes) in immunologically heterogeneous tumor models and combination immunotherapy settings. Clinical translation will require comprehensive safety profiling, pharmacokinetic studies, and dose optimization. Addressing these areas will be crucial to define riclin’s broader potential as a combinatory agent in oncology.
Methods
Preparation and structural information of riclin
Riclin was extracted and purified from Agrobacterium sp. ZCC3656 following established methods25,30, ensuring batch-to-batch consistency. Briefly, crude riclin was solubilized in ultrapure water (1% w/v) and subsequently purified via a heat-alkaline plus Sevag method, yielding a final purity of 97.5%. With a molecular weight of approximately 2.5 × 106 Da, riclin’s structure has been elucidated by methylation and nuclear magnetic resonance (NMR) analyses, which confirmed it is mainly composed of 1,3-linked Glcp, 1,4-linked Glcp, 1,3-linked Galp, 1,6-linked Glcp, 1,4,6-linked Glcp in a ratio of 2:2:1:1:2 as previously described (Fig. S10)30.
Cell lines and animals
Luc-LLC cells (murine Lewis lung carcinoma, luciferase-labeled) and RAW 264.7 macrophages (Cell Bank of Chinese Academy of Sciences, Shanghai) were cultured in Dulbecco’s Modified Eagle Medium (DMEM, Gibco) supplemented with 10% FBS and 1% penicillin/streptomycin (Thermo Fisher Scientific, Waltham, MA, USA). Cells were maintained at 37 °C in a 5% CO2 incubator during the logarithmic growth phase for experiments.
Male BALB/c and C57BL/6 mice (6-8 weeks old) from Yangzhou University Comparative Medicine Center were housed in specific pathogen-free (SPF) conditions (22 ± 1 °C, 12 h light/dark cycle) with ad libitum access to food/water. All procedures strictly adhered to the ARRIVE guidelines and were approved by the Animal Care Ethics Committee of Bengbu Medical University (No. BBMUEC2022022). Humane endpoints were defined as >25% body weight loss, severe lethargy or impaired mobility, or spontaneous bleeding. At the end of the experiment, the animals were anesthetized and euthanized by cervical dislocation.
Microbiota profile of 16S rRNA
Male BALB/c mice were orally administered riclin (20 mg/kg, n = 8) or saline (control, also called CK) daily for 14 days. Fresh fecal samples were collected aseptically, snap-frozen at −80 °C, and stored for intestinal microbiota and metabolomic profiling. Genomic DNA was extracted using the MagPure Soil DNA LQ Kit (Magen Biotech, Guangzhou, China). The V3-V4 hypervariable region of bacterial 16S rRNA was amplified via PCR and sequenced on the Illumina NovaSeq 6000 platform (OE Biotech, Inc., Shanghai, China). Raw data were generated using Illumina MiSeq or NovaSeq (Illumina, Inc., San Diego, CA, USA) sequencing. After quality control analyses, including denoising and chimera removal, performed with DADA2 in QIIME2 (2020.11), representative sequences and amplicon sequence variant (ASV) abundance tables were obtained. Subsequently, all representative sequences were annotated by alignment against the Silva database (version 138). A p-value < 0.05 indicates differentially abundant taxa.
Metabolomics analysis
Metabolomic profiling was conducted by OE Biotech Co., Ltd. using an ACQUITY UPLC I-Class Plus system (Waters Corporation, Milford, MA, USA) coupled to a Q Exactive mass spectrometer (Thermo Fisher Scientific). Raw data were processed using Progenesis QI v3.0 software (Nonlinear Dynamics, Newcastle, UK) and analyzed for identification via databases such as The Human Metabolome Database (HMDB). The principal component analysis (PCA) of the metabolomic data matrix was performed using the R package. Differential metabolites were defined by VIP > 1.0 and p < 0.05, followed by KEGG pathway enrichment analysis. Pearson correlation analysis was applied to assess associations between 20 differentially abundant microbial species and metabolites.
To quantitatively assess the levels of microbially derived SCFAs, fecal samples collected from BALB/c mice after 14 days of saline (CK) or riclin (20 mg/kg, Ric) treatment (n = 8 per group) were analyzed. Targeted SCFA profiling was performed using Liquid Chromatography-Tandem Mass Spectrometry (LC–MS/MS). Briefly, fecal samples were homogenized, acidified, and extracted. The peak areas of metabolites were substituted into the linear equation of the standard curve for calculation, ultimately yielding the absolute content data for each metabolite in the actual samples. A total of 11 SCFA were detected.
In vitro experiments
RAW 264.7 macrophages (5 × 105 cells/mL) were plated in 6-well plates and treated with riclin (20 μg/mL) for 24 h. Morphological changes were analyzed using Giemsa and Actin-Tracker Green-488 staining (Beyotime Biotechnology Co., Ltd., Shanghai, China) under an Olympus BX53 microscope. For functional assays, cells (5 × 105 cells/mL) in 96-well plates were divided into: (1) control (CK, blank DMEM), (2) LPS (10 μg/mL, L970739, Cas: 93572-42-0, Macklin), (3) GEM (10 μg/mL, G810678, 122111-03-9, 99%, Macklin), and (4) riclin (LR/HR: 10/20 μg/mL). After 24 h, proliferation and phagocytosis were assessed via CCK-8 and neutral red staining63. Supernatants were analyzed for nitric oxide (NO) (Beyotime Nitric Oxide Assay Kit) and cytokines (TNF-α, IL-1β, IL-6) using ELISA (MultiSciences Biotech).
RNA sequencing analysis
BALB/c mice were administered riclin (20 mg/kg, Ric, n = 3) or saline (CK) via oral gavage. Splenocytes and bone marrow-derived cells were harvested 6 h post-treatment. Total RNA was isolated using TRIzol reagent, and transcriptome sequencing was performed by OE Biotech Co., Ltd. DESeq2 software (v1.30.1) was used to normalize gene counts across samples and perform significance testing for differential expression. DEGs were defined by an absolute log2 Fold Change >1.0 and a false discovery rate (FDR)-adjusted p-value (Q-value) <0.05. The hypergeometric distribution test was employed to assess the enrichment significance of GO and KEGG entries, thereby identifying critical biological functions and pathways associated with differentially expressed genes. The GO and KEGG Databases used are linked as https://ftp.ncbi.nih.gov/gene/DATA/gene2go.gz and http://www.genome.jp/kegg/.
In vivo pathway inhibition assay
To validate the necessity of the NF-κB and JAK-STAT signaling pathways in riclin-mediated hematopoietic function, an additional in vivo experiment was conducted. Male BALB/c mice (8 weeks) were randomly divided into four groups: Control (CK, saline, n = 6), riclin (20 mg/kg, p.o.), riclin + BAY 11-7082 (NF-κB inhibitor, HY-13453, MedChemExpress LLC, 5 mg/kg, i.p.), and riclin + Ruxolitinib (JAK2 inhibitor, HY-50856, MedChemExpress LLC, 60 mg/kg, p.o.). The inhibitors were administered 30 min prior to riclin gavage. All treatments were performed daily for 3 consecutive days. 24 h after the final administration, mice were euthanized. Bone marrow cells and peripheral blood were collected for flow cytometric analysis of LSK cells and complete blood count analysis.
Immunosuppressed mouse model
An immunosuppression model was established in BALB/c mice via intraperitoneal gemcitabine (GEM, 100 mg/kg every 3 days, n = 11)55. Mice were randomly grouped and co-administered riclin (20 or 40 mg/kg/day, oral) or vehicle, with riclin pretreatment initiated 7 days prior to GEM. Animals were euthanized on day 11 of the regimen. Body weight and immune organ indices were recorded. Blood, spleen, and femur samples were collected for complete blood counts, histopathology (H&E staining), and downstream assays. Bone marrow and splenic single-cell suspensions were prepared, quantified, and analyzed for cell cycle and apoptosis using commercial kits (Beyotime).
Murine tumor model
To generate the murine LLC tumor model, male C57BL/6 J mice received subcutaneous implantation of 1 × 106 Luc-LLC cells in the right axillary region. Tumor-bearing mice (50–100 mm3 at day 10) were randomized into four treatment groups: Model control (saline, n = 8), GEM monotherapy (80 mg/kg, i.p.), GEM + low-dose riclin (20 mg/kg, p.o.), and GEM + high-dose riclin (40 mg/kg, p.o.). Tumor volume (0.5 × length × width2) and bioluminescence intensity were monitored until Day 21. Mice were euthanized on Day 22 for necropsy. Body weight, tumor weight, gross morphology, and spleen/thymus indices were recorded. Complete blood counts were analyzed using a Mindray hematology analyzer. While blinding during treatment wasn’t feasible, all endpoint analyses were performed by blinded personnel.
Bioluminescence imaging
Longitudinal tumor bioluminescence in LLC-bearing mice was quantified every 3 days using Living Image 4.4 software. The main steps were as follows: mice were anesthetized with 2–3% isoflurane, injected intraperitoneally with D-Luciferin (potassium salt, APEXBIO, 15 mg/mL), and placed in BRUKER small animal live imaging system (Bruker Corporation, MA, USA) for photographs within 15–30 min.
Immunofluorescence analysis
Paraffin sections of spleens from tumor-bearing mice were deparaffinized, rehydrated, and subjected to heat-induced antigen retrieval using EDTA buffer (pH 8.0, G1206, Servicebio Biotechnology Co., Ltd., Hubei, China). After serum blocking, sections were co-incubated with primary antibodies, including rabbit anti-CD4 (GB15064, Servicevio) and rabbit anti-CD8 (GB15068, Servicevio) at 4 °C overnight. After washing, sections were incubated with HRP-conjugated secondary antibody (HRP-labeled goat anti-rabbit IgG) and then subjected to tyramide signal amplification: CD4 was visualized with iF555-Tyramide (red fluorescence, G1233, Servicevio) and CD8 with iF647-Tyramide (yellow fluorescence, G1232, Servicevio). Nuclei were counterstained with DAPI (G1012, Servicevio). All images were acquired under standardized parameters using a Nikon Eclipse C1 fluorescence microscope (Nikon, Japan). The mean fluorescence density for CD4⁺ and CD8⁺ cells from five randomly selected fields per sample was quantified using ImageJ software (1.49).
Flow cytometry
Splenic and bone marrow single-cell suspensions were analyzed for cell phenotypes using fluorochrome-conjugated antibodies. CD45-APC-Cy7, CD3-PerCP-Cy5.5, CD4-FITC, CD8-PE-Cy7, and F4/80-APC antibodies are from BD Biosciences (San Diego, CA, USA). CD11b-FITC, CD86-PE-Cy,7 and CD206-PE antibodies are from BioLegend, Inc. (San Diego, CA, USA). Mouse Hematopoietic Lineage Antibody Cocktail (Lin, FITC), c-Kit (CD117) Monoclonal Antibody (PE), and Ly-6A/E (Sca-1) Monoclonal Antibody (PerCP-Cyanine5.5) are from eBioscience (San Diego, CA, USA). Flow cytometry was performed on a Beckman Coulter CytoFLEX S system (CA, USA), with data analyzed using FlowJo v10.8.
Real-time quantitative PCR (RT-qPCR)
Total RNA was isolated using TRIzol, followed by reverse transcription with Thermo Fisher Scientific M-MLV reverse transcriptase and random primers. RT-qPCR was performed with SYBR Green PCR Mix (TOYOBO). Gene expression analysis employed the 2 − ΔΔCT method, normalized to β-actin. Primer sequences are listed in Table S4.
Western blotting
Mice received three consecutive treatments as per the “RNA sequencing analysis” section. Splenic and bone marrow proteins were extracted, quantified via BCA assay, and electrophoresed onto PVDF membranes (Merck, Germany). Membranes underwent sequential incubation with primary and HRP-conjugated secondary antibodies (Cell Signaling Technology, MA, USA). Protein bands were visualized using BeyECL Moon (Beyotime) and quantified with ImageJ. Following antibodies were used: β-actin (3700, monoclonal), TNF-α (11948, monoclonal), IL-6 (12912s, monoclonal), and p-NF-κB p65 (3033T, monoclonal) from Cell Signaling Technology, IL-1β (sc-52012, monoclonal), granulocyte-macrophage colony-stimulating factor (GM-CSF, sc-32753, monoclonal), NF-κB p65 (sc-8008, monoclonal), phosphor (p)-IκB-α (sc-8404, monoclonal), GM-CSFr (sc-393281, monoclonal), JAK2 (sc-390539, monoclonal), STAT5 (sc-74442, monoclonal), and p-STAT5a/b (sc-81524, monoclonal) from Santa Cruz Biotechnology (Santa Cruz, CA, USA), and p-JAK2 (AF3024, polyclonal) from Affinity Biosciences (Cincinnati, OH, USA).
Statistical analysis
Statistical analyses were performed in GraphPad Prism 8 (v8.0.1) using one-way ANOVA/Tukey’s test or Student’s t-test, as appropriate. Data are presented as mean ± SD.
Data availability
The omics datasets generated in this study have been deposited in public repositories. Specifically, 16S rRNA sequencing data are available in the NCBI Sequence Read Archive (SRA) under accession number PRJNA1364046; RNA-seq data are available in the SRA under accession number PRJNA1364311; Untargeted and Targeted Metabolomics datasets are available in the MetaboLights database under accession numbers MTBLS13320 and MTBLS13326, respectively.
References
Sharma, A., Jasrotia, S. & Kumar, A. Effects of chemotherapy on the immune system: implications for cancer treatment and patient outcomes. Naunyn Schmiedebergs Arch. Pharmacol. 397, 2551–2566 (2024).
Farhangnia, P., Khorramdelazad, H., Nickho, H. & Delbandi, A. A. Current and future immunotherapeutic approaches in pancreatic cancer treatment. J. Hematol. Oncol. 17, 40 (2024).
Salas-Benito, D. et al. Paradigms on immunotherapy combinations with chemotherapy. Cancer Discov. 11, 1353–1367 (2021).
Guo, B. et al. CD44-targeting hydrophobic phosphorylated gemcitabine prodrug nanotherapeutics augment lung cancer therapy. Acta Biomater. 145, 200–209 (2022).
Liu, W. et al. RGDV-modified gemcitabine: a nano-medicine capable of prolonging half-life, overcoming resistance and eliminating bone marrow toxicity of gemcitabine. Int. J. Nanomed. 14, 7263–7279 (2019).
Silk, A. W. et al. Society for Immunotherapy of Cancer (SITC) clinical practice guideline on immunotherapy for the treatment of nonmelanoma skin cancer. J. Immunother. Cancer 10, e004434 (2022).
Chae, J. S. et al. Yeast (1→3)-(1→6)-β-d-glucan alleviates immunosuppression in gemcitabine-treated mice. Int. J. Biol. Macromol. 136, 1169–1175 (2019).
Bushmeleva, K. et al. Effect of flavonols of Aronia melanocarpa fruits on morphofunctional state of immunocompetent organs of rats under cyclophosphamide-induced immunosuppression. Biomolecules 14, 578 (2024).
Yimam, M. et al. Botanical bioflavonoid composition from Scutellaria baicalensis- and Acacia catechu-protected mice against D-galactose-induced immunosenescence, and cyclophosphamide induced immune suppression. Nutrients 16, 3144 (2024).
Liu, X. et al. Ginsenoside Rg3 improves cyclophosphamide-induced immunocompetence in Balb/c mice. Int. Immunopharmacol. 72, 98–111 (2019).
Qian, Y. et al. Ginsenoside Rh2 reverses cyclophosphamide-induced immune deficiency by regulating fatty acid metabolism. J. Leukoc. Biol. 106, 1089–1100 (2019).
Shakeri, F., Kiani, S., Rahimi, G. & Boskabady, M. H. Anti-inflammatory, antioxidant, and immunomodulatory effects of Berberis vulgaris and its constituent berberine, experimental and clinical, a review. Phytother. Res. 38, 1882–1902 (2024).
Wang, B. et al. Design of self-assembled micelles based on natural dual-targeting strategies and evaluation of their anti-liver cancer effects as drug delivery systems. NPJ Precis. Oncol. 9, 82 (2025).
Duffy, C. et al. Honeybee venom and melittin suppress growth factor receptor activation in HER2-enriched and triple-negative breast cancer. NPJ Precis. Oncol. 4, 24 (2020).
Wang, X. et al. Natural product pharmacology: the British Journal of Pharmacology perspective. Br. J. Pharmacol. 18, 3547–3555 (2024).
Zhang, H. et al. Prevention effect of total ginsenosides and ginseng extract from Panax ginseng on cyclophosphamide-induced immunosuppression in mice. Phytother. Res. 37, 3583–3601 (2023).
Li, X. et al. Polysaccharide isolated from Grifola frondose eliminates myeloid-derived suppressor cells and inhibits tumor growth by enhancing T cells responses. Int. J. Biol. Sci. 20, 664–679 (2024).
Yang, P. et al. The ameliorative effect on chemotherapy-induced injury and tumor immunosuppressive microenvironment of the polysaccharide from the rhizome of Menispermum dauricum DC. Int. J. Biol. Macromol. 268, 131828 (2024).
Zhou, Y. et al. Immunomodulatory effect of tremella polysaccharides against cyclophosphamide-induced immunosuppression in mice. Molecules 23, 239 (2018).
Zhu, Z. et al. Modulating effects of turmeric polysaccharides on immune response and gut microbiota in cyclophosphamide-treated mice. J. Agric Food Chem. 72, 3469–3482 (2024).
Wang, D. et al. Atractylodes lancea rhizome polysaccharide alleviates immunosuppression and intestinal mucosal injury in mice treated with cyclophosphamide. J. Agric Food Chem. 71, 17112–17129 (2023).
Huang, J. et al. Sodium alginate modulates immunity, intestinal mucosal barrier function, and gut microbiota in cyclophosphamide-induced immunosuppressed BALB/c mice. J. Agric Food Chem. 69, 7064–7073 (2021).
Wang, J. et al. Effects of exopolysaccharides from Lactiplantibacillus plantarum JLAU103 on intestinal immune response, oxidative stress, and microbial communities in cyclophosphamide-induced immunosuppressed mice. J. Agric Food Chem. 70, 2197–2210 (2022).
Yang, Y., Jiang, G. & Tian, Y. Biological activities and applications of exopolysaccharides produced by lactic acid bacteria: a mini-review. World J. Microbiol. Biotechnol. 39, 155 (2023).
Cheng, R. et al. In vitro and in vivo anti-inflammatory activity of a succinoglycan Riclin from Agrobacterium sp. ZCC3656. J. Appl Microbiol. 127, 1716–1726 (2019).
Wang, L. et al. Safety assessment of functional oligooctasaccharide riclinoctaose: A pilot study of genotoxicity, acute toxicity, and subchronic toxicity. J. Food Sci. 87, 1306–1318 (2022).
Miao, Y. et al. Exopolysaccharide riclin and anthocyanin-based composite colorimetric indicator film for food freshness monitoring. Carbohydr. Polym. 314, 120882 (2023).
Ding, Z. et al. Dietary succinoglycan riclin improves glycemia control in mice with type 2 diabetes. J. Agric Food Chem. 70, 1819–1829 (2022).
Lu, W., Kong, C., Cheng, S., Xu, X. & Zhang, J. Succinoglycan riclin relieves UVB-induced skin injury with anti-oxidant and anti-inflammatory properties. Int. J. Biol. Macromol. 235, 123717 (2023).
Yang, Y. et al. Anti-tumor activity and immunogenicity of a succinoglycan riclin. Carbohydr. Polym. 255, 117370 (2021).
Kau, A. L., Ahern, P. P., Griffin, N. W., Goodman, A. L. & Gordon, J. I. Human nutrition, the gut microbiome and the immune system. Nature 474, 327–336 (2011).
Hao, Y., Liao, X., Wang, X., Lao, S. & Liao, W. The biological regulatory activities of Flammulina velutipes polysaccharide in mice intestinal microbiota, immune repertoire and heart transcriptome. Int. J. Biol. Macromol. 185, 582–591 (2021).
Wen, Z. et al. Moringa oleifera polysaccharide regulates colonic microbiota and immune repertoire in C57BL/6 mice. Int. J. Biol. Macromol. 198, 135–146 (2022).
Bunker, J. J. et al. Innate and adaptive humoral responses coat distinct commensal bacteria with immunoglobulin A. Immunity 43, 541–553 (2015).
Ma, T. et al. Effects of co-fermented collagen peptide-jackfruit juice on the immune response and gut microbiota in immunosuppressed mice. Food Chem. 365, 130487 (2021).
Chung, H. et al. Gut immune maturation depends on colonization with a host-specific microbiota. Cell 149, 1578–1593 (2012).
Song, W. et al. Modulating the gut microbiota is involved in the effect of low-molecular-weight Glycyrrhiza polysaccharide on immune function. Gut Microbes 15, 2276814 (2023).
Yu, B. et al. A new polysaccharide from Hawk tea: structural characterization and immunomodulatory activity associated with regulating gut microbiota. Food Chem. 418, 135917 (2023).
Jin, M. et al. Response of intestinal metabolome to polysaccharides from mycelia of Ganoderma lucidum. Int. J. Biol. Macromol. 122, 723–731 (2019).
Qiao, Y. et al. Fluoride induces immunotoxicity by regulating riboflavin transport and metabolism partly through IL-17A in the spleen. J. Hazard Mater. 476, 135085 (2024).
Zhang, X. et al. Endogenous indole pyruvate pathway for tryptophan metabolism mediated by IL4I1. J. Agric Food Chem. 68, 10678–10684 (2020).
Sun, M. J. et al. The acetic acid produced by lactobacillus species regulates immune function to alleviate PEDV infection in piglets. Probiotics Antimicrob. Proteins 17, 2962–2979 (2025).
Luu, M. et al. The short-chain fatty acid pentanoate suppresses autoimmunity by modulating the metabolic-epigenetic crosstalk in lymphocytes. Nat. Commun. 10, 760 (2019).
Wang, X. K. et al. Bacteroides-derived isovaleric acid enhances mucosal immunity by facilitating intestinal IgA response in broilers. J. Anim. Sci. Biotechnol. 14, 4 (2023).
Liao, Y. et al. Structural characterization and immunomodulatory activity of exopolysaccharide from Aureobasidium pullulans CGMCC 23063. Carbohydr. Polym. 288, 119366 (2022).
Yu, Y. et al. Chemistry and immunostimulatory activity of a polysaccharide from Undaria pinnatifida. Food Chem. Toxicol. 128, 119–128 (2019).
Rajoka, M. S. R. et al. Techno-functional properties and immunomodulatory potential of exopolysaccharide from Lactiplantibacillus plantarum MM89 isolated from human breast milk. Food Chem. 377, 131954 (2022).
You, X. et al. Structural characterization and immunomodulatory activity of an exopolysaccharide produced by Lactobacillus helveticus LZ-R-5. Carbohydr. Polym. 235, 115977 (2020).
Liu, Y., Ye, Y., Hu, X. & Wang, J. Structural characterization and anti-inflammatory activity of a polysaccharide from the lignified okra. Carbohydr. Polym. 265, 118081 (2021).
Zhu, S. et al. Structure elucidation and immunological activity of a novel exopolysaccharide from Paenibacillus bovis sp. nov BD3526. Carbohydr. Polym. 282, 119103 (2022).
Ingelfinger, F., De Feo, D. & Becher, B. G. M. - CSF: master regulator of the T cell-phagocyte interface during inflammation. Semin. Immunol. 54, 101518 (2021).
Sun, S. C. The non-canonical NF-κB pathway in immunity and inflammation. Nat. Rev. Immunol. 17, 545–558 (2017).
Yang, K. et al. Sulfate glycosaminoglycan from swim bladder exerts immunomodulatory potential on macrophages via toll-like receptor 4 mediated NF-κB signaling pathways. Int. J. Biol. Macromol. 271, 132439 (2024).
Hamilton, J. A. GM-CSF in inflammation. J. Exp. Med. 217, e20190945 (2020).
Lv, K. et al. CBL family E3 ubiquitin ligases control JAK2 ubiquitination and stability in hematopoietic stem cells and myeloid malignancies. Genes Dev. 31, 1007–1023 (2017).
Liu, Y. et al. Danggui Buxue decoction enhances the anticancer activity of gemcitabine and alleviates gemcitabine-induced myelosuppression. J. Ethnopharmacol. 273, 113965 (2021).
Magidey, K. et al. A unique crosstalk between tumor cells and hematopoietic stem cells reveals a myeloid differentiation pattern signature contributing to metastasis. Blood 134, 2465 (2019).
Sun, C. et al. Improvement of icaritin on hematopoietic function in cyclophosphamide-induced myelosuppression mice. Immunopharmacol. Immunotoxicol. 40, 25–34 (2018).
Liao, Y. et al. Genetically engineered cellular nanoparticles loaded with curcuminoids for cancer immunotherapy. Theranostics 14, 6409–6425 (2024).
Thai, A. A., Solomon, B. J., Sequist, L. V., Gainor, J. F. & Heist, R. S. Lung cancer. Lancet 398, 535–554 (2021).
Yu, X. et al. Novel formulation of c-di-GMP with cytidinyl/cationic lipid reverses T cell exhaustion and activates stronger anti-tumor immunity. Theranostics 12, 6723–6739 (2022).
Zhang, J. et al. Lymph node-targeted delivery of Lonicera japonica thunb. polysaccharides for enhancing antitumor immunotherapy. Mater. Today Bio 31, 101559 (2025).
Chen, S. et al. A new polysaccharide platform constructs self-adjuvant nanovaccines to enhance immune responses. J. Nanobiotechnology 20, 320 (2022).
Acknowledgements
We are grateful to Prof. Jianfa Zhang for providing exopolysaccharide riclin.
Author information
Authors and Affiliations
Contributions
Y.M., X.L., and J.T. contributed equally to this work. Y.M: Investigation; Data curation; Writing-original draft; Visualization; Methodology; Formal analysis. X.L. and J.T.: Investigation; Data curation; Writing-original draft; Methodology; Funding acquisition. Y.J. and Y.F.: Investigation; Data curation; Visualization. X.S, J.L, Z.D., and J.Z.: Writing-review & editing; Resources. Q.G.: Writing-review & editing; Visualization; Methodology; Validation; Formal analysis. Q.S.: Supervision; Writing-review & editing; Visualization; Methodology; Validation; Funding acquisition; Conceptualization; Project administration.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Miao, Y., Liu, X., Tao, J. et al. Natural polysaccharide riclin acts as an immune adjuvant to enhance chemotherapy efficacy in NSCLC. npj Precis. Onc. 10, 108 (2026). https://doi.org/10.1038/s41698-026-01318-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41698-026-01318-z