Abstract
Distributed learning enables collaborative machine learning model training without requiring cross-institutional data sharing, thereby addressing privacy concerns. However, local quality control variability can negatively impact model performance while systematic human visual inspection is time-consuming and may violate the goal of keeping data inaccessible outside acquisition centers. This work proposes a novel self-supervised method to identify and eliminate harmful data during distributed learning model training fully-automatically. Harmful data is defined as samples that, when included in training, increase misdiagnosis rates. The method was tested using neuroimaging data from 83 centers for Parkinson’s disease classification with simulated inclusion of a few harmful data samples. The proposed method reliably identified harmful images, with centers providing only harmful datasets being easier to identify than single harmful images within otherwise good datasets. While only evaluated using neuroimaging data, the presented method is application-agnostic and presents a step towards automated quality control in distributed learning.
Similar content being viewed by others
Introduction
In the past decade, machine learning (ML) has become an essential tool for solving complex medical image analysis tasks, aiming to support clinical decision-making, reduce clinical workloads, and enable healthcare professionals to prioritize patient interactions1,2. The effectiveness of modern ML models depends heavily on three properties of the training datasets: quantity, diversity, and quality3. Within this context, models trained using high-quality, large, and diverse datasets capturing a broad representation of real-world cases are less likely to memorize (i.e., overfit) patterns that may be unrelated to the task at hand and more likely to learn features that are generalizable to new cases4. However, since suitable publicly available data is limited to only a few diseases or population-based studies and many medical centers do not have sufficient data to train disease-specific ML models alone, which is especially true for rare diseases or centers in remote or underserved areas, collaboration across multiple institutions is necessary to compile large data repositories. The establishment of the Alzheimer’s Disease Neuroimaging Initiative (ADNI)5 and the Parkinson’s Progression Markers Initiative (PPMI)6 repositories are great examples of collaborations that enabled impactful healthcare research advancement and training of advanced ML models. However, creating these repositories can be expensive, time-consuming, and often impractical due to privacy regulations prohibiting data sharing7,8.
While these regulations are crucial to protect patient-sensitive data, collecting sufficient multi-center data to develop ML models has become increasingly difficult. Within this context, distributed learning has emerged as an innovative approach to foster global collaboration, facilitate access to broader and more diverse datasets from various populations and demographics, and address privacy concerns associated with directly sharing data across institutions and borders9. The core idea behind distributed learning methods is to train ML models locally at each center without transferring data outside the digital borders of the data owner. Thus, privacy is better and easier preserved compared to the approach of creating a central repository of multi-center data as only the data-acquiring center has access to its own data10. The most widely implemented method for distributed learning is federated learning (FL). In the FL setup, separate ML models are trained in parallel at each participating center using local data only. Periodically, the local model parameters are sent to a central server, where they are aggregated and then redistributed back to the centers for further training11. An alternative to FL that has gained popularity lately is the travelling model (TM). In essence, the TM can be viewed as a serialized version of FL, where a single model is trained sequentially, moving from one location to another after each round of local training, either with or without an intermediary server. The TM has proven particularly effective in enabling centers with small datasets to participate in such distributed learning setups12,13. This strength stems from its iterative training approach, which helps mitigating the risk of local model overfitting - a common challenge when dealing with very small datasets within the FL setup14. As a result, the TM has a potential to further promote distributed learning frameworks internationally, specifically facilitating the participation of medical centers in low- and middle-income countries and remote areas. The COVID-19 pandemic is a good example showcasing the need for such international collaboration; if distributed learning frameworks had been already widely available and implemented at that time, clinical research and computer-aided diagnosis might have progressed more rapidly, potentially resulting in quicker disease understanding and accelerated development of preventive vaccines. Despite distributed learning not being widely explored during the pandemic, the corresponding data gathered by individual medical centers has played a crucial role in advancing distributed learning research in recent years within real-world healthcare settings15,16,17,18,19,20. Distributed learning has also been explored for several medical image analysis tasks in recent years, including adult brain cancer diagnosis12,21,22, breast cancer diagnosis23, brain age prediction14,24, Parkinson’s disease diagnosis25, retinal age prediction13, and many more26,27,28.
Despite the proven feasibility and general success of distributed learning in research settings, significant data-related challenges remain when applied in real clinical scenarios, particularly when building distributed ML models for medical image analysis. One of these problems is that many distributed learning methods simply assume that local centers provide high-quality data without providing any method to formally check or control for this. However, medical image data quality can significantly impact the performance of ML models, similar to how it can influence diagnostic accuracy by human experts. Image quality is a complex problem because even if individual centers conduct rigorous quality control, the definition of what constitutes a high-quality image is subjective and may differ across centers and observers. At the same time, a standard quality control process, where a single person visually inspects all data samples, conflicts with the distributed learning principle of limiting data access.
Some automatic image quality assessment tools have been developed previously to identify and filter out low-quality images during the image preprocessing of centralized data29,30. However, the quality assessment metrics used in those methods do not intrinsically relate an image’s quality to its ability to affect the diagnostic accuracy of an ML model. Conversely, these methods typically define quality thresholds based on parameters like signal-to-noise ratio or the presence of artifacts, but their optimal values may vary based on the dataset and task. In addition to that, there is no evidence that these metrics truly represent the best indicators of image quality. Therefore, there is still an urgent need for a method that can automatically identify potentially harmful data - such as low-quality images, incorrect image modalities, or images of the wrong anatomical regions - to successfully translate distributed learning methods from research into clinical practice. Failing to detect and remove such data samples from local training sets can significantly degrade the performance of the resulting ML models.
To date, the distributed learning community has not extensively explored how to systematically check for and ensure the quality of the distributed data. Instead, previous research has primarily focused on scenarios where malicious centers join distributed learning networks with the intent of altering the model behavior during training31,32,33,34,35,36,37,38. While this research offers valuable insights into the effects of a center exclusively contributing harmful data and towards identifying intentionally malicious centers, the methods used for that task cannot be applied to cases where centers include only a few harmful data samples in their training sets, whether intentionally or due to oversight. Additionally, these methods have been primarily designed for FL setups, where the server receives parameters updates from all centers simultaneously, enabling collective assessment and removal of potential malicious centers during the aggregation39. Within this context, previously proposed methods have utilized distance- or behaviour-based metrics for the identification of malicious centers. Distance-based approaches compute how far the centers’ updated parameters are from a reference. For instance, multi-Krum40 and the robustness of secure federated learning approach41 exclude centers if any parameter fails to meet the distance criteria. Conversely, self-centered clipping42 excludes only the parameters that do not meet the requirements, allowing the aggregation of valid parameters. Error rate-based rejection43 assesses the performance of a model by generating multiple versions of the aggregated model, each excluding one center at a time. These models are then evaluated on a hold-out test set to identify centers whose updates cause the model performance to drop. Centers with poor behaviour are excluded from the final aggregation. While these methods are well-suited for FL, they are impractical for TM since the server receives only a single version of the model containing updates from a specific center. By this point, the model has already incorporated updates from previous centers, making it difficult to determine whether any undesirable behaviour stems from the most recent update (i.e., that the last center is malicious) or from the cumulative effect of all prior updates. To the best of our knowledge, no prior work has addressed the problem of identifying malicious centers in the TM, and there are no prior works proposing methods for quality control of single data samples in distributed learning networks, either in FL or TM setups.
Therefore, this work introduces a novel self-supervised methodology to automatically identify and exclude harmful data samples from a center’s training set in the TM setup without relying on human inspection. Here, harmful data samples are defined as samples that, when included in the training, downgrade model accuracy and increase misdiagnosis rates. This is especially critical in healthcare applications, where incorrect diagnoses can have a significant impact on patient lives. We showcase the feasibility of this proposed method by evaluating its effectiveness using an established TM for Parkinson’s disease (PD) classification based on T1-weighted brain magnetic resonance imaging (MRI) data acquired in 83 different centers25. Therefore, the main contributions of this work are the development of the first automatic quality control method for decentralized data and the demonstration of its ability to detect cases when all local datasets from one center are harmful as well as cases where only a few data samples from a single center are harmful across several scenarios: (1) introducing three centers to the distributed data network, each with the entire local dataset composed of harmful data samples, (2) randomly incorporating a single harmful data sample into the datasets from nine of the 83 centers with large local datasets, and (3) employing the original 83 centers, providing only assumed correct MRI images to identify potentially harmful data that was missed during manual quality control.
Results
Overview
Briefly described, the proposed method for data quality control in distributed learning setups extends the TM concept by introducing three new steps: data verification, data revisit, and data elimination (Fig. 1 illustrates the overall process). In the verification step, a hold-out test set at each center is used to evaluate the model after training with each batch of local training data to identify any potentially harmful images to add to the revisit list for later review. In this case, harmful images are defined as those that increase misdiagnosis above an acceptable error threshold when used during training. Images on the revisit list are skipped in subsequent local training cycles until the revisit cycle is reached. The revisiting process is crucial to avoid unwanted elimination of images, for example, due to the sequence in which centers are visited during each cycle. During the revisit step, the verification process is repeated on those potentially harmful images. If an image still negatively impacts the model’s performance, it is completely eliminated from the pool of available training data for the corresponding center. Conversely, if an image is no longer harmful, it returns to the pool of training data for future cycles. While the revisit and elimination steps may not occur in every cycle, the verification step is performed in every regular and revisit cycle.
a In the traditional model, a solid arrow shows a single transfer of the trained model from centers to the server after training, using the entire local database. b In the travelling model with the data quality control mechanism, dashed arrows indicate multiple rounds of communication between centers and the server, as centers send the model around for local verification after each batch of training. c The bottom panel illustrates the verification step performed at the server. Created in BioRender: https://BioRender.com/m74p229.
To evaluate our proposed quality control method on a clinically relevant task, we adopted an established TM for Parkinson’s disease (PD) classification25. The PD data network available for training the TM includes data from 83 real centers, each providing varying quantities of T1-weighted brain MRI scans from healthy participants or patients with PD. In addition to the real-world application and evaluation, we simulated two scenarios to actively alter the model’s behavior during training by introducing three types of harmful data samples: inverted T1-weighted MRI, noise images, and chest computed tomography (CT) data (see Fig. 2). In the first scenario, three centers were added to the network, each contributing a local dataset consisting entirely of harmful image samples, simulating the case of a malicious center joining the network. In the second scenario, nine of the 83 centers, each with larger local datasets, included a single harmful image sample to simulate the presence of potentially harmful data samples within otherwise good datasets. These two scenarios allowed us to determine the acceptable error threshold and revisit cycle hyperparameters using a data-driven approach. The best hyperparameters were chosen based on the following metrics: models’ accuracy, sensitivity, specificity, F1-score, and area under the receiver operating characteristic curve (AUROC). The baselines for comparison included the performance of the pre-trained model used as the basis for every experiment (details in “Methods”), the “dirty” baseline, which refers to the model’s performance when harmful data samples are included, and the “clean” baseline, which refers to the model’s performance when the harmful data samples are excluded (i.e., when only the 83 centers providing T1-weighted brain MRI scans participate in the training). Thus, our methodology is successful when it identifies and eliminates harmful images, outperforming the “dirty” baseline for its scenario and the pre-trained model (accuracy: 67%, sensitivity: 72%, specificity: 63%, F1-score: 67%, and AUROC: 73%), and performing similarly to the “clean” baseline. The “clean” baseline performance and its lower and upper bounds were determined based on six TM runs using different seeds, achieving an accuracy of 75% (71–75%), sensitivity of 75% (71–75%), specificity of 75% (70–75%), F1-score of 73% (69–73%), and AUROC of 82% (79–82%).
The correct data type is represented with a green label while the harmful data types are illustrated with red labels. MRI magnetic resonance imaging, CT computed tomography.
Identification of centers that provide exclusively harmful data samples
The results of the first scenario experiment demonstrated that all combinations of the two hyperparameters effectively identified the three centers contributing only harmful data samples immediately (Table 1), surpassing the performance of its “dirty” baseline (accuracy: 54%, sensitivity: 0%, specificity: 100%, F1-score: 0%, and AUROC: 49%) and pre-trained model (accuracy: 67%, sensitivity: 72%, specificity: 63%, F1-score: 67%, and AUROC: 73%). Figure 3a supports these findings by illustrating that every harmful data sample intentionally introduced by the “malicious” centers was flagged the first time it was used for training, regardless of the error threshold employed. Moreover, the bottom of panel (a) shows that every flagged sample was eliminated in the first revisit cycle. Detailed proportions of data samples that were eliminated across all centers per training cycle for every hyperparameter combination can be found in the heatmaps in the Supplementary Fig. 1, which also shows that the datasets from the three centers named “malicious-X” (marked in red) were identified and excluded early on. Although all experiments demonstrate that the TM with quality control is able to successfully remove all harmful images, upon closer examination of the models’ performance (Table 1), it becomes evident that setting the acceptable error threshold to 2% and reviewing potentially harmful data after two cycles yielded the best performance across all metrics (accuracy: 73%, sensitivity: 70%, specificity: 75%, F1-score: 70%, and AUROC: 80%), achieving performance results comparable to the “clean” baseline, with performance metrics falling within its lower and upper bounds.
The x-axis shows an example of each harmful data type with noise images, computed tomography data, and inverted magnetic resonance imaging being presented from left to right. a The first scenario, where the entire local datasets are harmful. b The second scenario, where a single harmful data sample is added to an otherwise good local database.
Identification of harmful data within otherwise good local datasets
The second scenario experiment highlighted the difficulty of identifying single harmful data samples within a larger pool of otherwise good local datasets. Figure 3b illustrates that smaller acceptable error thresholds of 2% and 3% were required to effectively flag and remove all harmful image samples right away. Furthermore, 4% and 5% error thresholds missed more inverted MRI samples than noise and chest CT samples, potentially due to the brain structure still being present in this harmful image type. When analyzing the proportion of flagged images that were eliminated during the revisit cycle (bottom of panel (b)), it becomes evident that the 5% error threshold failed to remove harmful data samples in this scenario, as none of the flagged images were excluded. Figure 4 shows the cycle where each harmful data sample was eliminated for every hyperparameter combination. Here, it can be observed that reviewing potentially harmful data samples after two cycles successfully eliminated most harmful image samples, even with larger error thresholds of 4% and 5%. These findings are further supported by the heatmaps in Supplementary Fig. 2, which illustrate the proportion of harmful data eliminated per center and cycle. As in the first scenario experiment, setting a lower acceptable error threshold, specifically 3%, and reviewing potentially harmful data after two cycles yielded the best performance across all metrics (accuracy: 72%, sensitivity: 72%, specificity: 72%, F1-score: 70%, and AUROC: 78%), achieving results comparable to the “clean” baseline, with performance metrics falling within its bounds. In contrast to the first scenario experiment, where most configurations led to improvements over its “dirty” baseline and pre-trained model, in the second scenario, larger error thresholds (4% and 5%) led to a performance on par with its “dirty” baseline (accuracy: 54%, sensitivity: 0%, specificity: 100%, F1-score: 0%, and AUROC: 45%), which was inferior to the pre-trained model’s performance (Table 1).
The two graphs show the eight combinations of error and revisit cycle hyperparameters. The x-axis shows each harmful data sample included in these experiments while the y-axis shows the cycle that each data sample was eliminated. CT computed tomography. INV inverted magnetic resonance imaging.
Validation of experiments and identification of challenging harmful data type
Given that 2% and 3% acceptable error thresholds and a data revisit after two cycles led to the best-performing models for the first and second scenarios, we conducted six ablation studies to determine if one of the harmful data types is more challenging than others and to validate our findings in different scenarios. To do so, we examined scenarios where the three centers denoted as “malicious-X” and the nine centers with large local databases contributed the same harmful data type. In the first scenario, for each harmful data type, malicious #1 contributed three images, malicious #2 contributed seven images, and malicious #3 contributed five images. In contrast, in the second scenario, each of the nine centers’ local datasets included just one image. Our findings revealed that every harmful data sample included in the six ablation studies was identified and eliminated right away (see Supplementary Fig. 3), supporting our previous findings that a small error threshold is effective in identifying harmful data samples. Supplementary Tables 1 and 2 also show that the TM with quality control improved the model performance, achieving metrics within the lower and upper bounds of the “clean” baseline for all experiments.
Identification of harmful data within the real-word Parkinson’s disease network
Given that 2% and 3% acceptable error thresholds and a data revisit after two cycles led to the best-performing models for all experiments, we examined whether these parameters are also effective in identifying potentially harmful data among the 83 centers providing the correct data modality (T1-weighted brain MRI) of assumed good quality. The findings of this analysis showed that these parameter settings successfully identified and removed potentially harmful images and even entire centers (Fig. 5) in the PD data network. However, the 2% error threshold was found to be excessively restrictive, leading to insufficient training data, which poses a significant challenge for ML development. In contrast, a 3% threshold achieved a good balance, effectively removing harmful data samples while maintaining a model performance (accuracy: 72%, sensitivity: 71%, specificity: 73%, F1-score: 72%, and AUROC: 78%) within the bounds of the “clean” baseline and reducing the number of required training cycles from 26 to 22 (Table 1). Additionally, we conducted a detailed visual inspection of the data from centers whose entire datasets were excluded after one cycle employing the 3% acceptable error threshold, marked in red in Fig. 5. This review revealed significant issues within the excluded datasets, including generally poor image quality, blurring, low spatial resolution, microhemorrhages, and severely enlarged ventricles, as illustrated in Fig. 6. Notably, Fig. 6b shows a case of a scan of a subject who is considered healthy but displays considerable microhemorrhages. While this individual may be considered healthy from a PD perspective, the brain certainly shows pathologies that may confuse the model and explain why this case was excluded. These findings underscore the success of our proposed method in effectively identifying and addressing data quality issues in distributed learning setups, as these images are typical examples of low-quality and potentially harmful data that may still survive manual local quality control, especially if no comparison data is available.
Each row on the y-axis corresponds to a specific center, and each column on the x-axis represents a training cycle. Centers that had their dataset visually inspected are highlighted in red.
a Three examples of scans with poor quality. b llustrates an example of microhemorrhages in a healthy (i.e., no Parkinson’s disease) participant. c An example of a scan with considerably enlarged ventricles.
Discussion
This work introduced the first self-supervised and fully-data driven method for identifying single-case and whole-center harmful data samples, such as low-quality images, incorrect image modalities, or images of the wrong anatomical regions during distributed ML training, which could potentially degrade model performance. This approach marks a significant advancement toward decentralized data quality control. By incorporating three key steps - data verification, data revisit, and data elimination—into the travelling model approach, we are able to efficiently identify and remove harmful data, which if not removed leads to an increased rate of Parkinson’s disease (PD) misdiagnosis. As a result, the proposed quality control method ensures that only high-quality and accurate data from contributing centers in the distributed learning network are used to train and improve the final model’s performance.
Harmful datasets can be included intentionally, when centers join the network with the aim of manipulating the training process, or unintentionally, for example when centers mistakenly use the wrong image repository path or DICOM tags, include incorrect data modalities, or introduce low-quality data into a pool of otherwise good datasets. While both cases are harmful and impact the training process negatively, our findings revealed that identifying centers that provide exclusively harmful data samples is considerably easier than detecting single harmful data samples within a pool of otherwise good data. This is evident as any combination of hyperparameters for the quality control used to train the travelling model in the first scenario performed better than the pre-trained model and its “dirty” baseline, preventing undesired effects of the three centers named as “malicious-X”. In contrast, successfully removing single harmful data samples within otherwise good data requires smaller acceptable error thresholds and more frequent revisiting cycles. This becomes apparent when the method fails to identify inverted MRI samples as harmful when a larger error threshold is used. While a good quality control method should be able to identify and remove all types of harmful data samples, it may be expected that inverted MRIs are the most challenging to identify among the three harmful image types investigated in this work. More precisely, inverted MRI scans still contain all brain structures that may be detectable by some convolution filters that perhaps focus on shapes. While inverted MRIs are probably the most realistic harmful data example, corresponding to cases where simply a wrong MRI contrast (e.g., T2-weighetd vs. T1-weighted) is contributed locally, it is essential to highlight that our analysis of a single harmful data type at a time, revealed that small thresholds are effective in all cases, including inverted MRI.
The travelling model with the proposed quality control successfully identified and eliminated images with poor quality, features unrelated to PD (e.g., microhemorrhages), and potentially out-of-distribution images (e.g., extremely enlarged ventricles). However, it is important to note that using very small thresholds may be overly restrictive. This problem become evident in the real-world experiment involving data from the original 83 centers in the PD data network, which were assumed to have provided correct image modality (i.e., T1-weighted MRI) and high-quality images. The excessive elimination of data resulted in an insufficient amount of representative data for effectively training ML models. These findings underscore that selecting the appropriate hyperparameters depends on the overall reliability and quality of the contributed data. Therefore, a grid search or alternative techniques may be necessary when applying this methodology in other contexts.
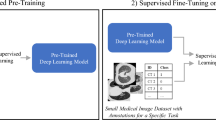
Despite self-supervising methods demonstrating promising performance in several medical imaging analysis tasks44, it is important to highlight some of the differences between other self-supervised methods proposed in the literature and this work. Self-supervision is usually attributed to tasks that have a portion of the data that is unlabeled. One example of self-supervised learning would be the pre-training of a ML model on a large unlabeled database of MRI scans and fine-tuning the model for a specific task (e.g., disease classification) on a smaller labeled database. In such cases, the goal is either to take advantage of publicly available data or to avoid random initialization of the model. On the contrary, the self-supervised quality control method proposed in this work never utilized labeled data for determining imaging quality. Instead, during a supervised learning task of PD classification, this work investigated misdiagnosis as a mediator for imaging quality. More precisely, this work evaluated the false-positive and false-negative rates and assumed that if an image sample leads to downgrades in these metrics, it harms the model. Thus, the quality control method considers that data sample as low-quality, flagging it as a candidate to be eliminated.
Importantly, this work offers insights beyond decentralized data quality control and could significantly contribute to developing more honest reward mechanisms. More precisely, numerous researchers have proposed methods to recognize centers for their contributions for distributed ML model development45,46,47. The goal of reward mechanisms is to establish a method for compensating center contributions, which can include monetary incentives or privileged access to the final model. Traditionally, quantifying the contribution of a medical center for its data contribution often makes use of the raw number of datasets provided for training, simply assuming that increasing data volume enhances the value added to ML models. However, the relationship between data quantity and value for a trained model is complex48. For example, while a large center might contribute several times more data than a small center, its data variability could be minimal, whereas the small center may offer fewer but more unique cases. As a result, data from the smaller center may hold greater relevance for the intended task and value for the ML model development. If center contributions are solely judged by dataset quantity, smaller centers might be undervalued compared to larger ones. Therefore, using the proposed method as the basis, researchers may be able to assess each dataset’s relevance to the intended task, paving the way for more refined and equitable reward mechanisms.
There are several important limitations that should be discussed regarding this work. First, the addition of three steps for the data quality control increases training time due to the verification step required after the training of each batch. Moreover, the verification step requires every center to have a hold-out set that can be used to monitor the model performance. Thus, future research should aim to reduce these additional communication costs and hold-out set requirements. Potential strategies could involve establishing a centralized hold-out set on the server to eliminate the need for distributing the model to every center for verification and eliminating the requirement of the hold-out set per center. Alternatively, implementing strategies to assess the potential negative impact of a batch on the model before training could reduce both, training and verification times. Second, the investigated scenarios only considered three relevant but exemplary types of harmful data. Further investigation could explore additional types or low-quality data that can potentially affect ML models, such as the same imaging modality but different organs, inclusion of severe imaging and motion artifacts, and others. However, the third experiment already showed that the proposed method can identify low-quality data samples that may even pass a manual quality control. Third, this work exclusively utilized a single established travelling model for PD diagnosis as a first proof-of-concept. Therefore, additional research is needed to determine if our findings hold true when employing different distributed learning approaches, ML architectures, or clinical tasks. However, it is worth noting that the proposed distributed network involved an extensive number of centers with some of them only contributing very few datasets, surpassing many datasets used in other federated learning and travelling model studies. This aspect potentially enhances the generalizability of our results. Fourth, it may be argued that sequential training is more susceptible to catastrophic forgetting. This work addressed this problem by employing multiple training cycles and cycle-to-cycle variability. However, it is important to highlight the need for future research to establish methods to quantify catastrophic forgetting. Fifth, while this work was conducted using a single computer, it is anticipated that the reported findings would remain consistent if each center employed computers with similar specifications. In this case, the physical network implementation may not affect the results. Nevertheless, future research could explore strategies for deploying this system, such as setting up cloud computing or intranet-based computer networks with appropriate connection protocols. Finally, while this work focused on evaluating the proposed decentralized quality control method for medical image analysis, it is important to note that this method is data-type and task agnostic. As such, it can even be applied to non-medical image analysis in distributed learning environments, offering benefits for ensuring data quality across various domains.
Methods
Parkinson’s disease classification model
The deep learning architecture used in this work for PD classification is the same as used in ref. 25, consisting of eight blocks: The initial five blocks feature a 3D convolutional layer with 3 × 3 × 3 kernel filters, batch normalization, 2 × 2 × 2 max pooling, and ReLU activations. The sixth block includes a 3D convolutional layer with 1 × 1 × 1 kernel filters, batch normalization, and ReLU activations. The seventh block incorporates a 3D average pooling layer, a dropout layer with a 0.2 rate, and a flattening layer with 768 features. The eighth block consists of a single dense layer with a sigmoid activation function for the binary classification output, distinguishing between patients with PD and healthy participants.
Travelling model for distributed learning
The core distributed learning concept implemented in this work is the travelling model (TM) for PD classification, as initially described in ref. 25, with the additional integration of the data quality control method described below. In standard TM training, a model is initialized at a server and then transmitted to each participating center according to a pre-defined travelling sequence. After the first center has completed training the model with local data, the updated model is returned to the server, which then sends it to the next center. This sequential process continues until the model has visited all centers, completing one cycle. To prevent the model from forgetting patterns learned at centers in the beginning of the sequence, a phenomenon known as catastrophic forgetting49, a new travelling sequence is defined after each cycle, and multiple cycles are executed, which also improves the model’s overall performance12,25. Training the model locally at each center ensures compliance with data-sharing regulations by keeping the data securely stored at its point of acquisition. This approach restricts data access to authorized personnel within the specific center, preventing other centers or ML developers from accessing or receiving any information about it.
In this work, a batch size of five was used when centers had five or more locally available data samples. For centers with less than five data samples, all of the locally available data was used in a single batch. Training employed the Adam optimizer, starting with an initial learning rate of 0.0001 and applied exponential decay after each cycle. The training was conducted on a single computer equipped with an NVIDIA GeForce RTX 3090 GPU, adhering strictly to the TM concept by fetching data from one center at a time.
Distributed learning training with data quality control
The data quality control method proposed in this work involves flagging and potentially removing images that could negatively impact the ability of the model to accurately learn the task at hand (here, PD classification). Therefore, a pre-trained model is required prior to this step to ensure that images are not incorrectly flagged as harmful while the model is learning its basic knowledge from scratch. In this work, we utilized the TM approach without data quality control to pre-train the model for ten cycles, utilizing the training data available at the 83 centers in the PD network contributing T1-weighted brain MRI samples. This pre-trained model served as the foundation for the subsequent data quality control method.
Once the data quality control is activated, metrics for monitoring model performance are used to identify images that are harmful to model learning. Therefore, this work utilizes common behavior-based metrics to assess the model’s performance following weight and optimizer updates39. As the name suggests, behavior-based metrics monitor the behavior of the model based on a pre-defined metric. Given the healthcare application scenario and the critical impact of misdiagnosis in computer-aided diagnosis tools, we specifically focus on the false-positive rate (FPR) and false-negative rate (FNR) to monitor the model’s behavior and facilitate the identification and removal of harmful imaging data. The baseline FPR and FNR for the pre-trained model are computed prior to the start of data quality control by aggregating these metrics from each center’s hold-out data via the server.
The data quality control method is integrated through three additional steps into the TM training process: data verification, data revisit, and data elimination (Fig. 1). After training on a batch of data, the training center sends the model back to the server, which then distributes a copy of the model to all participating centers for verification. During the verification step, each center computes the FPR and FNR using its local hold-out set and reports these metrics to the server. The server evaluates whether the batch has caused the FPR or FNR to exceed a pre-defined error threshold. If so, the image(s) responsible for the unacceptable error are identified, flagged as potentially harmful, removed from the training data pool, and added to a “revisit list.” The revisit step occurs after a pre-defined number of training cycles, where each flagged image on the revisit list is independently re-evaluated. If a flagged image no longer causes the FPR or FNR to exceed acceptable thresholds, it is returned to the local training data pool. However, if the image still negatively impacts these metrics, it is permanently removed from the local training pool for that center. Further details about each step are provided in the following section and illustrated in Fig. 7.
Blue represents the steps of a regular cycle, purple illustrates the steps of the revisit cycle, and pink shows the steps of the verification process. Created in BioRender: https://BioRender.com/w08f826.
Quality control steps
The verification step occurs after local training on each batch. Instead of a center training the model using the entire locally available dataset, it trains the model for one batch and sends it back to the server. The server then simultaneously distributes a copy of the updated model to every other center to verify model performance on their hold-out data (similar to a federated learning approach). The centers perform inference and send the FPR and FNR to the server that aggregates the metrics. If the aggregated FNR or FPR do not exceed an acceptable error threshold, the server updates the baseline metrics with the new FPR and FNR, keeps the updated model and optimizer, and sends it back to the center, which continues to train with local data if more batches are available. If the center has already finished training with all batches, the server sends the model to the next center in the sequence.
However, if the aggregated metrics increase above an acceptable error threshold, more actions are necessary. First, the server reverses the weights and optimizer updates in the model. Then, it sends the model back to the center, which retrains it using data from the same batch but employs a batch size of one to identify the exact image(s) that increased FPR or FNR. When the images(s) that exceed the error threshold is/are identified, the server reverses the weights and optimizer updates in the model, adds the image(s) to the revisit list, and sends the model back to the center to continue training if more batches are available. If the center has already finished training with all batches, the server sends the model to the next center in the sequence. This entire process is illustrated in blue on the left side of Fig. 7.
The revisit step occurs once the travelling sequence is completed, and the model has visited every center. At this time, the server determines which center has images in the revisit list that require review in that cycle. Then, the server sends the model to centers with flagged images following the order that the images were added to the revisit list. Each center trains the model with a batch size of one and the model undergoes verification after each update. If the image no longer leads to a downgrade of the model performance, it is reinstated into the center’s dataset pool as shown in purple in the revisit step of the flowchart. The revisiting process is essential to prevent images from being mistakenly eliminated, for example, due to the order in which centers are visited during each cycle. For instance, an image from a particular center might be incorrectly flagged as harmful if the centers visited earlier provided data with significantly different participant demographics, leading to a wide variance in data distribution. This variation could cause the model to incorrectly flag these images as harmful. However, revisiting these images later in the process, after the model has been further trained on data from all centers, makes it more likely that the model has a better understanding of the diverse data distribution. This allows the model to reassess and potentially recognize the value of these images during the review process. Conversely, if an image is still identified as being harmful, it is flagged for elimination.
In the elimination step, any image flagged for elimination is removed from the training pool of its respective center. The image will no longer be used for training or verification in future cycles. Moreover, if a center no longer has data in its training pool after the elimination step, it will be omitted from future travelling sequences. Finally, a new travelling sequence is generated to introduce variability, akin to the batch shuffling process commonly used in centralized ML training. This process also reduces the risk of catastrophic forgetting, which occurs when the model forgets patterns about the data from the centers placed earlier in the travelling sequence. The TM with quality control utilized the same batch size and optimizer configurations employed in the traditional TM setup.
Parkinson’s disease dataset
The set of data used in this work comprises 1817 T1-weighted 3D brain MRIs acquired in 83 centers worldwide5,6,50,51,52,53,54,55,56,57,58,59,60,61, each contributing diverse subject demographics, acquisition protocols, scanners, and varying quantities of local datasets. Table 2 summarizes the characteristics of each study involved in the data acquisition, while detailed center information is provided in Supplementary Table 3. All datasets were preprocessed as described in ref. 25, including skull-stripping, resampling to 1 mm isotropic resolution, bias field correction, affine registration to the PD25-T1-MPRAGE-1 mm atlas62, and background cropping. The current study received approval from the Conjoint Health Research Ethics Board at the University of Calgary (REB21-0454). Moreover, all participants provided written informed consent to the individual studies local ethics committee following the guidelines set forth in the declaration of Helsinki.
Experimental setup
The proposed method was evaluated using three different scenarios: (1) introducing three centers to the PD data network, each with the entirety of the local datasets composed of harmful data samples, (2) randomly incorporating a single harmful data sample into the datasets from nine of the 83 centers with large local datasets, and (3) using the original data from the 83 centers, providing only correct T1-weighted brain MRIs to identify any low-quality samples that survived manual quality control but harm the model performance.
In the real-world scenario (i.e., the third scenario), centers with more than 25 images allocated 80% of their data for training and 20% for hold-out testing. For centers with fewer than 25 images, the data was divided between training and hold-out testing with the goal of maintaining a balanced representation of sex and age in both sets, totalizing 1410 datasets for training and 407 for testing. For the first and second scenarios, harmful data types were included exclusively in the training sets. In the first scenario, three centers were deliberately introduced into the PD data network to provide harmful data: malicious center #1 provided three inverted T1-weighted images to simulate a different MRI modality; malicious center #2 contributed seven chest CT scans to simulate the wrong imaging modality and body part; and malicious center #3 provided five images of pure noise. These noise images were generated by adding Gaussian noise to T1-weighted data and removing brain tissue, simulating extremely poor image quality (see Fig. 2 for examples of each dataset). In the second scenario, nine centers were randomly selected to inject one harmful data sample into their pool of T1-weighted MRI images. Specifically, Calgary, UOA, and Montreal included inverted T1-weighted images; Neurocon, Japan, and OASIS provided one noise data sample each; while Hamburg, UKBB, and BIOCOG each contributed one chest CT dataset.
For the first and second scenarios, the combination of four error thresholds (2%, 3%, 4%, and 5%) and two revisit cycles (2 and 5) were investigated. The error threshold defines what constitutes a harmful image by specifying the allowable increase in false-positive and false-negative rates. Notably, the error threshold is applied to each image individually, meaning that the threshold value is scaled according to the number of images in a batch. For example, during the verification step with a 2% error threshold, if the batch contains five images, the acceptable error margin would be 10% (2% multiplied by 5). The revisit cycle determines for how many cycles a flagged harmful image remains in the revisit list, meaning it will be skipped in subsequent cycles until the revisit cycle limit is reached. As a result, eight experiments were conducted for each scenario involving harmful data types.
Additionally, six ablation studies (three for the first scenario and three for the second one) using the optimal hyperparameters were conducted to investigate if any one of the harmful data types is more challenging to identify than others. In these scenarios, the three centers denoted as “malicious-X” and the nine centers with large local databases contribute the same harmful data type as follows.
-
Experiment 1: The three malicious centers contributed noise images. Therefore, malicious #1 contributed three noise images, malicious #2 contributed seven noise images, and malicious #3 contributed five noise images. As a result, the network had 86 collaborators.
-
Experiment 2: The three malicious centers contributed CT imaging. Therefore, malicious #1 contributed three CT images, malicious #2 contributed seven CT images, and malicious #3 contributed five CT images. As a result, the network had 86 collaborators.
-
Experiment 3: The three malicious centers contributed inverted MR imaging. Therefore, malicious #1 contributed three inverted MR images, malicious #2 contributed seven inverted MR images, and malicious #3 contributed five inverted MR images. As a result, the network had 86 collaborators.
-
Experiment 4: included a single noise image to nine centers with large local datasets. Thus, the network had the original 83 collaborators.
-
Experiment 5: included a single chest CT imaging to nine centers with large local datasets. Thus, the network had the original 83 collaborators.
-
Experiment 6: included a single inverted MR imaging to nine centers with large local datasets. Thus, the network had the original 83 collaborators.
Finally, two additional experiments using the optimal hyperparameters, as identified by the first two scenarios and confirmed by the six ablation studies, were performed for the real-world scenario to identify potentially harmful data that was missed during manual quality control. Each model of these 16 experiments was trained for 16 cycles to match the 26 cycles of the best-performing “clean” baseline, as a model that had been pre-trained for ten cycles was used as the basis in all cases.
Baselines
Every baseline in this work followed the TM procedure without data quality control procedure. The “clean” baseline was comprised of the training data from the 83 centers providing T1-weighted MRI images and was iterated over 30 cycles to optimize the model’s performance. Furthermore, six models were trained with different seeds to establish the travelling sequences and determine its lower and upper bounds. Similarly, the “dirty” baselines underwent 30 cycles of iteration to optimize the model’s performance, but they included all harmful data samples. The pre-trained model underwent ten training cycles and included only the training data from the 83 centers providing T1-weighted MRI images, as described previously.
Evaluation metrics
Five evaluation metrics were used to quantitatively assess the performance of the models: accuracy, sensitivity, specificity, F1-score, and AUROC. Accuracy is the percentage of cases that were correctly diagnosed based on a threshold of 0.5. Sensitivity measures the proportion of patients with PD correctly identified by the model, while specificity measures the proportion of healthy participants correctly identified as healthy. The F1-score is the harmonic mean of sensitivity and precision, providing a balance between the two metrics and accounts for both false positives and false negatives. The AUROC measures the model’s ability to discriminate between patients with PD and healthy participants across various thresholds.
To illustrate the number of samples placed in the revisit list and eliminated during training, a heatmap was used. Here, the x-axis of the heatmap represents the number of training cycles, while the y-axis represents the collaborating centers, with centers containing harmful data highlighted in red. For each training cycle, the proportion of images moved to the revisit list or eliminated from the training pool was computed for each center. This proportion was calculated by dividing the number of images in the revisit list plus the number of images eliminated by the total number of images available for training at each center. In the heatmap, light green (0) indicates that the images available at that center are not harmful to the task (i.e., none of them are in the revisit list or eliminated). Conversely, dark blue (1) indicates that every image provided by that center does not contribute useful information for PD classification (i.e., they are either in the revisit list or eliminated). Other shades in the heatmap signify that some images were harmful while others were not.
To illustrate the impact of different acceptable error thresholds and revisit cycles, a bar plot was utilized. This bar plot specifically examined the harmful data intentionally added in the first and second scenarios. This allows us to observe the effectiveness of our methodology in identifying and removing harmful images for each combination of error thresholds and revisit cycles.
Data availability
Image data used were provided in part by the UK Biobank (application number 77508), by the PPMI-a, public-private partnership funded by Michael J. Fox Foundation by the OASIS-3 project (Principal Investigators: T. Benzinger, D. Marcus, J. Morris; NIH P50 AG00561, P30 NS09857781, P01 AG026276, P01 AG003991, R01 AG043434, UL1 TR000448, R01 EB009352), by the OpenfMRI database (accession number ds000245), and by the Alzheimer’s Disease Neuroimaging Initiative (ADNI), a partnership involving multiple centers across North America with the goal of tracking participants through periods of cognitive decline and dementia. Launched in 2003, ADNI continues to evaluate biomarker, neuroimaging, and neuropsychological status in participants. The data can be requested by contacting the corresponding host institution.
Code availability
The code used in this work is available at https://github.com/RaissaSouza/decentralized-quality-control.
References
Lo Vercio, L. et al. Supervised machine learning tools: a tutorial for clinicians. J. Neural Eng. 17, 062001 (2020).
Maceachern, S. J. & Forkert, N. D. Machine learning for precision medicine. Genome 64, 416–425 (2021).
Naylor, C. D. On the Prospects for a (Deep) Learning Health Care System. JAMA 320, 1099 (2018).
Mehrabi, N., Morstatter, F., Saxena, N., Lerman, K. & Galstyan, A. A Survey on Bias and Fairness in Machine Learning. ACM Comput Surv. 54, 1–35 (2022).
Thibeau-Sutre, E., Couvy-Duchesne, B., Dormont, D., Colliot, O. & Burgos, N. MRI Field Strength Predicts Alzheimer’s Disease: a Case Example of Bias in the ADNI Data Set. in 2022 IEEE 19th International Symposium on Biomedical Imaging (ISBI) vols 2022-March 1–4 (IEEE, 2022)
Parkinson’s Progression Markers Initiative. https://www.ppmi-info.org/. Accessed November 22, 2022.
Annas, G. J. HIPAA Regulations — A New Era of Medical-Record Privacy? N. Engl. J. Med. 348, 1486–1490 (2003).
Voigt, P. & von dem Bussche, A. The EU General Data Protection Regulation (GDPR). (Springer International Publishing, Cham, 2017). https://doi.org/10.1007/978-3-319-57959-7
Rieke, N. et al. The future of digital health with federated learning. npj Digital Med. 2020 3:1 3, 1–7 (2020).
Tuladhar, A., Rajashekar, D. & Forkert, N. D. Distributed learning in healthcare. In Trends of artificial intelligence and big data for E-health. Integrated Science, vol 9 (eds Sakly, H., et al.) 183–212 (Springer, Cham, 2022). https://doi.org/10.1007/978-3-031-11199-0_10.
McMahan, H. B., Moore, E., Ramage, D., Hampson, S. & Arcas, B. A. y. Communication-efficient learning of deep networks from decentralized data. In Proc. of the 20th International Conference on Artificial Intelligence and Statistics, AISTATS 2017 (2017). http://arxiv.org/abs/1602.05629.
Souza, R. et al. Multi-institutional travelling model for tumor segmentation in MRI Datasets. in Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics) 420–432 (Springer, 2022). https://doi.org/10.1007/978-3-031-09002-8_37.
Nielsen, C., Souza, R., Wilms, M. & Forkert, N. D. Foundation model-driven distributed learning for enhanced retinal age prediction. J. Am. Med. Inform. Assoc. 31, 2550–2559 (2024).
Souza, R. et al. An analysis of the effects of limited training data in distributed learning scenarios for brain age prediction. J. Am. Med. Inform. Assoc. 30, 112–119 (2022).
Darzi, E., Sijtsema, N. M. & van Ooijen, P. M. A. A comparative study of federated learning methods for COVID-19 detection. Sci. Rep. 14, 3944 (2024).
Dayan, I. et al. Federated learning for predicting clinical outcomes in patients with COVID-19. Nat. Med. 27, 1735–1743 (2021).
Feki, I., Ammar, S., Kessentini, Y. & Muhammad, K. Federated learning for COVID-19 screening from Chest X-ray images. Appl Soft Comput 106, 107330 (2021).
Yan, B. et al. Experiments of Federated Learning for COVID-19 Chest X-ray Images. in Communications in Computer and Information Science vol. 1423 41–53 (Springer Science and Business Media Deutschland GmbH, 2021).
Soltan, A. A. S. et al. A scalable federated learning solution for secondary care using low-cost microcomputing: privacy-preserving development and evaluation of a COVID-19 screening test in UK hospitals. Lancet Digit Health 6, e93–e104 (2024).
Nguyen, D. C., Ding, M., Pathirana, P. N., Seneviratne, A. & Zomaya, A. Y. Federated learning for COVID-19 detection with generative adversarial networks in edge cloud computing. IEEE Internet Things J. 9, 10257–10271 (2022).
Sheller, M. J. et al. Federated learning in medicine: facilitating multi-institutional collaborations without sharing patient data. Sci. Rep. 10, 12598 (2020).
Pati, S. et al. Federated learning enables big data for rare cancer boundary detection. Nat. Commun. 13, 7346 (2022).
Schmidt, K. et al. Fair evaluation of federated learning algorithms for automated breast density classification: The results of the 2022 ACR-NCI-NVIDIA federated learning challenge. Med. Image Anal. 95, 103206 (2024).
Souza, R. et al. A comparative analysis of the impact of data distribution on distributed learning with a traveling model for brain age prediction. in Medical Imaging 2022: Imaging Informatics for Healthcare, Research, and Applications (eds. Park, B. J. & Deserno, T. M.) vol. 12037 1 (SPIE, 2022).
Souza, R. et al. A multi-center distributed learning approach for Parkinson’s disease classification using the traveling model paradigm. Front. Artif. Intell. 7, https://doi.org/10.3389/frai.2024.1301997 (2024).
Gunesli, G. N., Bilal, M., Raza, S. E. A. & Rajpoot, N. M. FedDropoutAvg: Generalizable federated learning for histopathology image classification. http://arxiv.org/abs/2111.13230 (2021).
Lu, M. Y. et al. Federated learning for computational pathology on gigapixel whole slide images. Med. Image Anal. 76, 102298 (2022).
Liu, Z., Wu, F., Wang, Y., Yang, M. & Pan, X. FedCL: Federated contrastive learning for multi-center medical image classification. Pattern Recognit. 143, 109739 (2023).
Lei, K., Syed, A. B., Zhu, X., Pauly, J. M. & Vasanawala, S. S. Artifact- and content-specific quality assessment for MRI with image rulers. Med. Image Anal. 77, 102344 (2022).
Nikiforaki, K. et al. Image Quality Assessment Tool for Conventional and Dynamic Magnetic Resonance Imaging Acquisitions. J. Imaging 10, 115 (2024).
Gu, Y. & Bai, Y. LDIA: Label distribution inference attack against federated learning in edge computing. J. Inf. Security Appl. 74, 103475 (2023).
Lu, G. et al. DEFEAT: A decentralized federated learning against gradient attacks. High.-Confid. Comput. 3, 100128 (2023).
Jin, R. & Li, X. Backdoor attack and defense in federated generative adversarial network-based medical image synthesis. Med. Image Anal. 90, 102965 (2023).
Gupta, A., Luo, T., Ngo, M. V. & Das, S. K. Long-Short History of Gradients Is All You Need: Detecting Malicious and Unreliable Clients in Federated Learning. in Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics) vol. 13556 LNCS 445–465 (Springer Science and Business Media Deutschland GmbH, 2022).
Le, J. et al. Privacy-Preserving Federated Learning With Malicious Clients and Honest-but-Curious Servers. IEEE Trans. Inf. Forensics Security 18, 4329–4344 (2023).
Zhang, Z., Cao, X., Jia, J. & Gong, N. Z. FLDetector: Defending Federated Learning Against Model Poisoning Attacks via Detecting Malicious Clients. in Proceedings of the 28th ACM SIGKDD Conference on Knowledge Discovery and Data Mining vol. 22 2545–2555 (ACM, New York, NY, USA, 2022).
Cao, X., Jia, J. & Gong, N. Z. Provably secure federated learning against malicious clients. Proc. AAAI Conf. Artif. Intell. 35, 6885–6893 (2021).
Nielsen, C., Tuladhar, A. & Forkert, N. D. Investigating the Vulnerability of Federated Learning-Based Diabetic Retinopathy Grade Classification to Gradient Inversion Attacks. in Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics) vol. 13576 LNCS 183–192 (Springer Science and Business Media Deutschland GmbH, 2022).
Raynal, M. & Troncoso, C. On the Conflict of Robustness and Learning in Collaborative Machine Learning. http://arxiv.org/abs/2402.13700 (2024).
Blanchard, P., Mahdi El Mhamdi, E., Guerraoui, R. & Stainer, J. Machine Learning with Adversaries: Byzantine Tolerant Gradient Descent. in Advances in Neural Information Processing Systems vol. 30 (Curran Associates, Inc., 2017).
Lycklama, H., Burkhalter, L., Viand, A., Küchler, N. & Hithnawi, A. RoFL: Robustness of Secure Federated Learning. in 2023 IEEE Symposium on Security and Privacy (SP) vols 2023-May 453–476 (IEEE, 2023).
He, L., Karimireddy, S. P. & Jaggi, M. Byzantine-Robust Decentralized Learning via ClippedGossip. http://arxiv.org/abs/2202.01545 (2022).
Fang, M., Cao, X., Jia, J. & Gong, N. Z. Local Model Poisoning Attacks to Byzantine-Robust Federated Learning. Proceedings of the 29th USENIX Security Symposium 1623–1640 (2019).
VanBerlo, B., Hoey, J. & Wong, A. A survey of the impact of self-supervised pretraining for diagnostic tasks in medical X-ray, CT, MRI, and ultrasound. BMC Med. Imaging 24, 1–24 (2024).
Li, M. et al. IMFL: an incentive mechanism for federated learning with personalized protection. IEEE Int. Things J. 11, 23862–23877 (2024).
Jiang, M. et al. Fair Federated Medical Image Segmentation via Client Contribution Estimation. in 2023 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) vols 2023-June 16302–16311 (IEEE, 2023)
Xu, X. et al. Gradient-Driven Rewards to Guarantee Fairness in Collaborative Machine Learning. in dvances in Neural Information Processing Systems 16104–16117 (Curran Associates, Inc., 2021).
Ng, D., Lan, X., Yao, M. M.-S., Chan, W. P. & Feng, M. Federated learning: a collaborative effort to achieve better medical imaging models for individual sites that have small labelled datasets. Quant. Imaging Med. Surg. 11, 852–857 (2021).
Kirkpatrick, J. et al. Overcoming catastrophic forgetting in neural networks. Proc. Natl. Acad. Sci. 114, 3521–3526 (2017).
Wei, D. et al. Structural and functional brain scans from the cross-sectional Southwest University adult lifespan dataset. Sci. Data 5, 180134 (2018).
OpenNeuro. https://openneuro.org/datasets/ds000245/versions/00001. Accessed November 24, 2022.
Open Science, to accelerate discovery and deliver cures | The Neuro - McGill University. https://www.mcgill.ca/neuro/open-science/c-big-repository, https://www.mcgill.ca/neuro/open-science.
Badea, L., Onu, M., Wu, T., Roceanu, A. & Bajenaru, O. Exploring the reproducibility of functional connectivity alterations in Parkinson’s disease. PLoS One 12, e0188196 (2017).
LaMontagne, P. J. et al. OASIS-3: Longitudinal Neuroimaging, Clinical, and Cognitive Dataset for Normal Aging and Alzheimer Disease. medRxiv 2019.12.13.19014902 https://doi.org/10.1101/2019.12.13.19014902 (2019).
Lang, S. et al. Network basis of the dysexecutive and posterior cortical cognitive profiles in Parkinson’s disease. Mov. Disord. 34, 893–902 (2019).
Hanganu, A. et al. Mild cognitive impairment is linked with faster rate of cortical thinning in patients with Parkinson’s disease longitudinally. Brain 137, 1120–1129 (2014).
Duchesne, S. et al. The Canadian dementia imaging protocol: harmonizing national cohorts. J. Magn. Reson. Imaging 49, 456–465 (2019).
Acharya, H. J., Bouchard, T. P., Emery, D. J. & Camicioli, R. M. Axial signs and magnetic resonance imaging correlates in Parkinson’s disease. Can. J. Neurolog. Sci. / J. Canadien des. Sci. Neurologiques 34, 56–61 (2007).
Talai, A. S., Sedlacik, J., Boelmans, K. & Forkert, N. D. Utility of multi-modal MRI for differentiating of Parkinson’s disease and progressive supranuclear palsy using machine learning. Front Neurol. 12, 648548 (2021).
Sudlow, C. et al. UK Biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 12, e1001779 (2015).
Jack, C. R. et al. The Alzheimer’s disease neuroimaging initiative (ADNI): MRI methods. J. Magn. Reson. Imaging 27, 685–691 (2008).
Xiao, Y. et al. Multi-contrast unbiased MRI atlas of a Parkinson’s disease population. Int. J. Comput. Assist. Radio. Surg. 10, 329–341 (2015).
Acknowledgements
Data used in preparation of this article were obtained from the Alzheimer’s Disease Neuroimaging Initiative (ADNI) database (adni.loni.usc.edu). As such, the investigators within the ADNI contributed to the design and implementation of ADNI and/or provided data but did not participate in analysis or writing of this report. A complete listing of ADNI investigators can be found at: http://adni.loni.usc.edu/wp-content/uploads/how_to_apply/ADNI_Acknowledgement_List.pdf. This work was supported by the Parkinson Association of Alberta, the Hotchkiss Brain Institute, the Canadian Consortium on Neurodegeneration in Aging (CCNA), the Canadian Open Neuroscience Platform (CONP), the Natural Sciences and Engineering Research Council of Canada (NSERC), the Canada Research Chairs program, the River Fund at Calgary Foundation, the Canadian Institutes for Health Research, the Tourmaline Chair in Parkinson disease, and the Institut de valorisation des données (IVADO).
Author information
Authors and Affiliations
Contributions
R.S., E.A.M.S., A.J.W., M.W., and N.D.F. contributed to the study’s conception. R.C., O.M. and Z.I. contributed to data acquisition and data curation. R.S. implemented the model. R.S., E.A.M.S., A.J.W., C.K., K.A., E.Y.O., and G.D. analyzed the results and were involved in the visualizations. R.S. wrote the first draft of the article. All authors critically revised the previous versions of the article and approved the final article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Souza, R., Stanley, E.A.M., Winder, A.J. et al. Self-supervised identification and elimination of harmful datasets in distributed machine learning for medical image analysis. npj Digit. Med. 8, 104 (2025). https://doi.org/10.1038/s41746-025-01499-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-025-01499-0